Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank
|
|
|
- Lisa Harvey
- 5 years ago
- Views:
Transcription
1 Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank Peter Tiňo, Jakub Mažgút, Hong Yan, Mikael Bodén Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.1/32
2 Motivation Increasing number of data processing tasks involve manipulation of multi-dimensional objects - tensors. Applying pattern recognition or machine learning methods directly - high computational and memory requirements, as well as poor generalization. To address this curse of dimensionality - decomposition methods to compress the data while capturing the dominant trends. New methods for processing multi-dimensional tensors in their natural structure have been introduced - real-valued tensors - nonnegative tensors - symmetric tensors Not suitable for binary tensors. Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.2/32
3 An Example Source: [Li et al.:mpca, 2008] Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.3/32
4 Tensor N-th order tensor A R I 1 I 2... I N Addressed by N indices i n ranging from 1 to I n, n = 1,2,...,N. Rank-1 tensora R I 1 I 2... I N can be obtained as an outer product ofn non-zero vectors u (n) R I n, n = 1,2,...,N: A = u(1) u (2)... u (N). For a particular index setting i = (i 1,i 2,...,i N ) Υ = {1,2,...,I 1 } {1,2,...,I 2 }... {1,2,...,I N }, we have A i = A i1,i 2,...,i N = N n=1 u (n) i n, where u (n) i n is the i n -th component of the vector u (n). Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.4/32
5 Basic algebra A tensor can be multiplied by a matrix (2nd order tensor) using n-mode products. The n-mode product of a tensor A R I 1 I 2... I N by a matrix U R J I n is a tensor (A n U) with entries (A n U) i1,...,i n 1,j,i n+1,...,i N = I n in =1 A i 1,...,i n 1,i n,i n+1,...,i N U j,in. Orthonormal basis {u (n) 1, u(n) 2 Basis matrix U (n) = (u (n) 1, u(n) 2,..., u(n) I n } for the n-mode space R I n.,..., u(n) I n ). Any tensor A can be decomposed into the product A = Q 1 U (1) 2 U (2) 3... N U (N). The expansion coefficients stored in the Nth order tensor Q R I 1 I 2... I N. Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.5/32
6 Tucker model for decomposing real tensors The expansion can also be written as A = i Υ Q i (u (1) i 1 u (2) i 2... u (N) i N ). In other words, tensor A is expressed as a linear combination of ( N n=1 I n) (a lot!) rank-1 basis tensors (u (1) i 1 u (2) i 2... u (N) i N ). The rank-1 basis tensors obtained as outer products of the corresponding basis vectors. More restricted models available - e.g. PARAFAC Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.6/32
7 Reduced rank representations of tensors Assume a smaller number of basis tensors are sufficient to approximate all tensors in a given dataset: A i K Q i (u (1) i 1 u (2) i 2... u (N) i N ), where K Υ. Tensors can be found close to the K -dimensional hyperplane in the tensor space spanned by the basis tensors (u (1) i 1 u (2) i 2... u (N) i N ), i K. A can be represented through expansion coefficients Q i, i K. Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.7/32
8 The model Data: M tensors D = {A m } M m=1 Each element A m,i is assumed to be (independently) Bernoulli distributed with parameter (mean) p m,i : P(A m,i p m,i ) = pa m,i m,i (1 p m,i )1 A m,i. Parametrized through log-odds (natural parameter), θ m,i = log p m,i 1 p m,i. Link function is the logistic function p m,i = σ(θ m,i ) = 1 1+e θ m,i, 1 p m,i = σ( θ m,i ) Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.8/32
9 The model For each data tensor A m, m = 1,2,...,M, we have P(A m θ m ) = i ΥP(A m,i θ m,i ), where P(A m,i θ m,i ) = σ(θ m,i )A m,i σ( θ m,i )1 A m,i. We collect all the parameters θ m,i in a tensor Θ RM I 1 I 2... I N of order N +1. Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.9/32
10 The model So far the values in the parameter tensor Θ were unconstrained. Constrain the Nth order parameter tensors θ m R I 1 I 2... I N to lie in the subspace spanned by the reduced set of basis tensors (u (1) r 1 u (2) r 2... u (N) r N ), where r n {1,2,...,R n }, and R n I n, i = 1,2...,N. The indices r = (r 1,r 2,...,r N ) take values from the set ρ = {1,2,...,R 1 } {1,2,...,R 2 }... {1,2,...,R N }. There is no explicit pressure in the model to ensure independence of the basis vectors. However, in practice, the optimized model parameters always represented independent basis vectors, as dependent basis vectors would lead to dependent basis tensors, implying smaller than intended rank of the tensor decomposition Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.10/32
11 The model Allow for an N-th order bias tensor R I 1 I 2... I N Parameter tensors θ m are constrained onto an affine space. θ m = r ρ Q m,r (u (1) r 1 u (2) r 2... u (N) r N )+ θ m,i = r ρ Q m,r (u (1) r 1 u (2) r 2... u (N) r N ) i + i = r ρq m,r N n=1 u (n) r n,i n + i Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.11/32
12 The model log-likelihood: M L = A m,i logσ N m,r r ρq m=1 i Υ (1 A m,i )logσ m,r r ρq n=1 N n=1 r n,i n + i + u (n) u (n) r n,i n i Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.12/32
13 Parameter Estimation The log likelihood is not convex in the parameters, it is convex in any of these parameters, if the others are fixed. Analytical updates were derived from a lower bound on the likelihood, using a trick from [Schein et al., 2003]. The linear tensor structure gets through! Iterative estimation scheme: while(convergence criterion) 1. argmax Q L 2. argmax u L 3. argmax L Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.13/32
14 Parameter Estimation Messy derivations, but the main message is: Even though the original vector model [Schein 2003] is non-linear in parameters, the strong linear algebraic structure of the Tucker model for tensor decomposition can be superimposed on the parameter space of the tensor model, so that the efficient linear nature of parameter updates of [Schein 2003] can be preserved. Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.14/32
15 Parameter Estimation - Basis Vectors For n = 1,2,...,N, define Υ n = {1,2,...,I 1 }... {1,2,...,I n 1 } {1} {1,2,...,I n+1 }... {1,2,...,I N } Analogously for ρ n. Given i Υ n and an n-mode index j {1,2,...,I n }, the index N-tuple (i 1,...,i n 1,j,i n+1,...,i N ) formed by inserting j at the nth place of i is denoted by [i,j n]. Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.15/32
16 Parameter Estimation - Basis Vectors B (n) m,i,q = r ρ n Q m,[r,q n] N s=1,s n u (s) r s,i s S (n) q,j = M m=1 i Υ n (2A m,[i,j n] 1 T m,[i,j n] [i,j n] ) B(n) m,i,q K (n) q,t,j = M m=1 r ρ n Q m,[r,t n] i Υ n T m,[i,j n] B(n) m,i,q N s=1,s n u (s) r s,i s Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.16/32
17 Parameter Estimation - Basis Vectors For each n-mode coordinate j {1,2,...,I n }: Collect the j-th coordinate values of all n-mode basis vectors into a column vector u (n) :,j = (u (n) 1,j,u(n) 2,j,...,u(n) R n,j )T. Stack all the S (n) q,j values in a column vector S (n) :,j = (S (n) 1,j,S(n) 2,j,...,S(n) R n,j )T. Construct an R n R n matrix K (n) :,:,j whose q-th row is (K (n) q,1,j,k(n) q,2,j,...,k(n) q,r n,j ), q = 1,2,...,R n. The n-mode basis vectors are updated by solving I n linear systems of size R n R n : K (n) :,:,j u(n) :,j = S (n) :,j, j = 1,2,...,I n. Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.17/32
18 Experiments - Synthetic Data Goal: Evaluate the amount of preserved information in compressed tensor representations. Compare the performance with existing real-value tensor decomposition model (TensorLSI). Experiment sets of binary tensors were sampled from different Bernoulli natural parameter subspaces. 2. Each set contains 10,000 2nd-order binary tensors of size (30,250). 3. On each set, both models were used to find subspaces using 80% of the tensors. 4. The hold-out sets of tensors (20%) were projected onto the subspaces and then reconstructed back. 5. To evaluate the match between the real-valued predictions and the target binary values we employ AUC analysis. Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.18/32
19 Synthetic Data A sample of randomly generated binary tensors from the above Bernoulli natural parameter space: Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.19/32
20 ROC curve analysis {x 1,x 2...x J } - model prediction outputs for all nonzero elements of tensors from the test set {y 1,y 2...y K } - outputs for all zero elements AUC value for that prediction (reconstruction) of the test set tensors: AUC = J j=1 K k=1 C(x j,y k ), J K where J and K are the total number of nonzero and zero tensor elements in the test set, respectively, and C is a scoring function C(x j,y k ) = { 1 if xj > y k 0 otherwise. Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.20/32
21 Hold-out Binary Tensor Reconstructions AUC Num. of Principal Components GML PCA TensorLSI Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.21/32
22 Introns and Promoters in DNA Sequences Introns: nucleotide sequences within a gene that get removed by RNA splicing to generate the final RNA product of a gene. Sequences that are joined together in the final mature RNA after RNA splicing are exons. Promoters: a region of DNA that facilitates the transcription of a particular gene. Promoters are located near the genes they regulate. Promoters contain specific DNA sequences providing an initial binding site for RNA polymerase and for proteins - transcription factors - that recruit RNA polymerase. Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.22/32
23 Topographic Mapping of DNA Sequences Goal: Find a mapping that groups functionally similar sub-sequences. Underlying principle: DNA sub-sequences from different functional regions differ in local term composition. To capture the composition we propose a binary tensor representation of the DNA sub-sequences. As a dataset of DNA sequences we used 62,000 promoter and intronic subsequences employed in [Li et al., 2008]. Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.23/32
24 Representation of a Genomic Sequence DNA sub-sequence: cgctcccggggtggccccacgcccc ctctgagc gagcggcggcgcgggacggggacggctctggccgggaccagcaggcctcgggcatccgggacgccggggccgc gctccaggccaggggcgggggcgggaccggggcgggggccggcggcggggccgcgccctcggcctctccccggggcgaccgggcggctccacacgcgctgcgcccgcc gccggccccacgcgcggcccatgtcctccgcgc Term Corresponding term-posi on matrix representa on: aag agc acg at gat ggca ggcg ggct ggt gca gcg gcca gccg gcct gct gt ca cga cgcg cgt cca ccgc ccca cccg cct ctca tca tcg ttc tttc tttt Posi on Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.24/32
25 3D PCA Projections of 10-dim Tensor Subspaces 3D PCA Projection of Sequences Analyzed by GML-PCA 3D PCA Projection of Sequences Analyzed by TensorLSI 2 5 3rd Principal Component rd Principal Component nd Principal Component st Principal Component 5 Promoter sequences Intron sequences 2 2nd Principal Component st Principal Component 2 4 Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.25/32
26 Topographic Mapping of DNA sequences aag agc acg at gat ggca ggcg ggct ggt gca gcg gcca gccg gcct gct gt ca cga cgcg cgt cca ccgc ccca cccg cct ctca tca tcg ttc tttc tttt aag agc acg at gat ggca ggcg ggct ggt gca gcg gcca gccg gcct gct gt ca cga cgcg cgt cca ccgc ccca cccg cct ctca tca tcg ttc tttc tttt aag agc acg at gat ggca ggcg ggct ggt gca gcg gcca gccg gcct gct gt ca cga cgcg cgt cca ccgc ccca cccg cct ctca tca tcg ttc tttc tttt I-1 I-2 P-1 P-5 2nd Principal Component D PCA Projection of Expansion Coefficients from GML-PCA st Principal Component aag agc acg at gat ggca ggcg ggct ggt gca gcg gcca gccg gcct gct gt ca cga cgcg cgt cca ccgc ccca cccg cct ctca tca tcg ttc tttc tttt aag agc acg at gat ggca ggcg ggct ggt gca gcg gcca gccg gcct gct gt ca cga cgcg cgt cca ccgc ccca cccg cct ctca tca tcg ttc tttc tttt P-2 aag agc acg at gat ggca ggcg ggct ggt gca gcg gcca gccg gcct gct gt ca cga cgcg cgt cca ccgc ccca cccg cct ctca tca tcg ttc tttc tttt I-3 aag agc acg at gat ggca ggcg ggct ggt gca gcg gcca gccg gcct gct gt ca cga cgcg cgt cca ccgc ccca cccg cct ctca tca tcg ttc tttc tttt P-4 aag agc acg at gat ggca ggcg ggct ggt gca gcg gcca gccg gcct gct gt ca cga cgcg cgt cca ccgc ccca cccg cct ctca tca tcg ttc tttc tttt P-3 Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.26/32
27 Functional enrichment analysis of promoters DNA-binding sites of transcription factors are often characterized as relatively short (5-15 nucleotides) sequence patterns. They may occur multiple times in promoters of the genes the expression of which they modulate. To validate that our model picks up biologically meaningful patterns, we searched the compressed feature space of promoters for biologically relevant structure (including that left by transcription factors). Genes that are transcribed by the same factors are often functionally similar. Suitable representations of promoters should correlate with the roles assigned to their genes. If the projection to a compressed space highlights such features, it is testament to a method s utility for processing biological sequences. Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.27/32
28 Functional enrichment analysis of promoters Gene Ontology (GO) provides a controlled vocabulary for annotation of genes, broadly categorized into terms for cellular component, biological process and molecular function. Assign biologically meaningful labels to promoters: Sequences were mapped to gene identifiers. In cases of multiple gene identifiers for the same promoter sequence, we picked the identifier with the greatest number of annotations unique GO terms annotating promoters. Evaluate whether promoters deemed similar in the topographic mapping are also functionally similar. Need the notion of a distance between a pair of promoters. Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.28/32
29 Functional enrichment analysis of promoters Naive approach - use the Euclidean distance in the 10-dim coordinate space of natural parameters. Not correct: (1) the basis tensors are not orthogonal; (2) they span a space of Bernoulli natural parameters that have a nonlinear relationship with the data values. Model-based distance between two promoter sequences m and l - sum of average symmetrized Kullback-Leibler divergences between noise distributions for all corresponding tensor elements i Υ: ( KL[pm,i p l,i ]+KL[p l,i p m,i ] ) D(m,l) = i Υ 2. Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.29/32
30 Are the compressed promoter representations are biologically meaningful? In each test, select one promoter as a reference. Repeat for all promoters. Given a reference promoter m, we label the group of promoters S m = {l D(m,l) < D 0 } within a pre-specified distance D(m,l) < D 0 as positives and all others as negatives. D 0 = 25, usually rendering over one hundred positives. For each GO term Fisher s exact test resolves if it occurs more often amongst positives than would be expected by chance. Null hypothesis - the GO term is not attributed more often than by chance to the positives. A small p-value indicates that the term is enriched at the position of the reference promoter m. We repeated the tests after shuffling the points assigned to promoters. In no case did this permutation test identify a single GO term as significant. Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.30/32
31 Yes! At significance p < , 75 GO terms were enriched around one or more reference promoters. The observation that a subset of promoter sequences are functionally organized after decomposition into 10 basis tensors adds support to the methods ability to detect variation at an information-rich level. Found a number of terms that are specifically concerned with chromatin structure (that packages the DNA), e.g. GO: Nucleosome, GO: Chromatin assembly or disassembly and GO: Protein-DNA complex assembly. Found several enriched terms related to development, e.g. GO: Reproductive process and GO: Female pregnancy. Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.31/32
32 Promoter regions assigned to GO: GO: Biological process: Reproduction GO: biological process Reproduction Topographic Mapping and Dimensionality Reduction of Binary Tensor Data of Arbitrary Rank p.32/32
Dimensionality Reduction and Topographic Mapping of Binary Tensors
 Pattern Analysis and Applications manuscript No. (will be inserted by the editor) Dimensionality Reduction and Topographic Mapping of Binary Tensors Jakub Mažgut Peter Tiňo Mikael Bodén Hong Yan Received:
Pattern Analysis and Applications manuscript No. (will be inserted by the editor) Dimensionality Reduction and Topographic Mapping of Binary Tensors Jakub Mažgut Peter Tiňo Mikael Bodén Hong Yan Received:
O 3 O 4 O 5. q 3. q 4. Transition
 Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Practical Bioinformatics
 5/2/2017 Dictionaries d i c t i o n a r y = { A : T, T : A, G : C, C : G } d i c t i o n a r y [ G ] d i c t i o n a r y [ N ] = N d i c t i o n a r y. h a s k e y ( C ) Dictionaries g e n e t i c C o
5/2/2017 Dictionaries d i c t i o n a r y = { A : T, T : A, G : C, C : G } d i c t i o n a r y [ G ] d i c t i o n a r y [ N ] = N d i c t i o n a r y. h a s k e y ( C ) Dictionaries g e n e t i c C o
Supplementary Information for
 Supplementary Information for Evolutionary conservation of codon optimality reveals hidden signatures of co-translational folding Sebastian Pechmann & Judith Frydman Department of Biology and BioX, Stanford
Supplementary Information for Evolutionary conservation of codon optimality reveals hidden signatures of co-translational folding Sebastian Pechmann & Judith Frydman Department of Biology and BioX, Stanford
B553 Lecture 5: Matrix Algebra Review
 B553 Lecture 5: Matrix Algebra Review Kris Hauser January 19, 2012 We have seen in prior lectures how vectors represent points in R n and gradients of functions. Matrices represent linear transformations
B553 Lecture 5: Matrix Algebra Review Kris Hauser January 19, 2012 We have seen in prior lectures how vectors represent points in R n and gradients of functions. Matrices represent linear transformations
PCA & ICA. CE-717: Machine Learning Sharif University of Technology Spring Soleymani
 PCA & ICA CE-717: Machine Learning Sharif University of Technology Spring 2015 Soleymani Dimensionality Reduction: Feature Selection vs. Feature Extraction Feature selection Select a subset of a given
PCA & ICA CE-717: Machine Learning Sharif University of Technology Spring 2015 Soleymani Dimensionality Reduction: Feature Selection vs. Feature Extraction Feature selection Select a subset of a given
STA 414/2104: Lecture 8
 STA 414/2104: Lecture 8 6-7 March 2017: Continuous Latent Variable Models, Neural networks With thanks to Russ Salakhutdinov, Jimmy Ba and others Outline Continuous latent variable models Background PCA
STA 414/2104: Lecture 8 6-7 March 2017: Continuous Latent Variable Models, Neural networks With thanks to Russ Salakhutdinov, Jimmy Ba and others Outline Continuous latent variable models Background PCA
Crick s early Hypothesis Revisited
 Crick s early Hypothesis Revisited Or The Existence of a Universal Coding Frame Ryan Rossi, Jean-Louis Lassez and Axel Bernal UPenn Center for Bioinformatics BIOINFORMATICS The application of computer
Crick s early Hypothesis Revisited Or The Existence of a Universal Coding Frame Ryan Rossi, Jean-Louis Lassez and Axel Bernal UPenn Center for Bioinformatics BIOINFORMATICS The application of computer
SEQUENCE ALIGNMENT BACKGROUND: BIOINFORMATICS. Prokaryotes and Eukaryotes. DNA and RNA
 SEQUENCE ALIGNMENT BACKGROUND: BIOINFORMATICS 1 Prokaryotes and Eukaryotes 2 DNA and RNA 3 4 Double helix structure Codons Codons are triplets of bases from the RNA sequence. Each triplet defines an amino-acid.
SEQUENCE ALIGNMENT BACKGROUND: BIOINFORMATICS 1 Prokaryotes and Eukaryotes 2 DNA and RNA 3 4 Double helix structure Codons Codons are triplets of bases from the RNA sequence. Each triplet defines an amino-acid.
Advanced topics in bioinformatics
 Feinberg Graduate School of the Weizmann Institute of Science Advanced topics in bioinformatics Shmuel Pietrokovski & Eitan Rubin Spring 2003 Course WWW site: http://bioinformatics.weizmann.ac.il/courses/atib
Feinberg Graduate School of the Weizmann Institute of Science Advanced topics in bioinformatics Shmuel Pietrokovski & Eitan Rubin Spring 2003 Course WWW site: http://bioinformatics.weizmann.ac.il/courses/atib
Universität Potsdam Institut für Informatik Lehrstuhl Maschinelles Lernen PCA. Tobias Scheffer
 Universität Potsdam Institut für Informatik Lehrstuhl Maschinelles Lernen PCA Tobias Scheffer Overview Principal Component Analysis (PCA) Kernel-PCA Fisher Linear Discriminant Analysis t-sne 2 PCA: Motivation
Universität Potsdam Institut für Informatik Lehrstuhl Maschinelles Lernen PCA Tobias Scheffer Overview Principal Component Analysis (PCA) Kernel-PCA Fisher Linear Discriminant Analysis t-sne 2 PCA: Motivation
ECE 521. Lecture 11 (not on midterm material) 13 February K-means clustering, Dimensionality reduction
 ECE 521 Lecture 11 (not on midterm material) 13 February 2017 K-means clustering, Dimensionality reduction With thanks to Ruslan Salakhutdinov for an earlier version of the slides Overview K-means clustering
ECE 521 Lecture 11 (not on midterm material) 13 February 2017 K-means clustering, Dimensionality reduction With thanks to Ruslan Salakhutdinov for an earlier version of the slides Overview K-means clustering
Maximum variance formulation
 12.1. Principal Component Analysis 561 Figure 12.2 Principal component analysis seeks a space of lower dimensionality, known as the principal subspace and denoted by the magenta line, such that the orthogonal
12.1. Principal Component Analysis 561 Figure 12.2 Principal component analysis seeks a space of lower dimensionality, known as the principal subspace and denoted by the magenta line, such that the orthogonal
Statistical Machine Learning
 Statistical Machine Learning Christoph Lampert Spring Semester 2015/2016 // Lecture 12 1 / 36 Unsupervised Learning Dimensionality Reduction 2 / 36 Dimensionality Reduction Given: data X = {x 1,..., x
Statistical Machine Learning Christoph Lampert Spring Semester 2015/2016 // Lecture 12 1 / 36 Unsupervised Learning Dimensionality Reduction 2 / 36 Dimensionality Reduction Given: data X = {x 1,..., x
BME 5742 Biosystems Modeling and Control
 BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
Kernel Methods. Machine Learning A W VO
 Kernel Methods Machine Learning A 708.063 07W VO Outline 1. Dual representation 2. The kernel concept 3. Properties of kernels 4. Examples of kernel machines Kernel PCA Support vector regression (Relevance
Kernel Methods Machine Learning A 708.063 07W VO Outline 1. Dual representation 2. The kernel concept 3. Properties of kernels 4. Examples of kernel machines Kernel PCA Support vector regression (Relevance
Vectors To begin, let us describe an element of the state space as a point with numerical coordinates, that is x 1. x 2. x =
 Linear Algebra Review Vectors To begin, let us describe an element of the state space as a point with numerical coordinates, that is x 1 x x = 2. x n Vectors of up to three dimensions are easy to diagram.
Linear Algebra Review Vectors To begin, let us describe an element of the state space as a point with numerical coordinates, that is x 1 x x = 2. x n Vectors of up to three dimensions are easy to diagram.
High throughput near infrared screening discovers DNA-templated silver clusters with peak fluorescence beyond 950 nm
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2018 High throughput near infrared screening discovers DNA-templated silver clusters with peak fluorescence
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2018 High throughput near infrared screening discovers DNA-templated silver clusters with peak fluorescence
Linear & nonlinear classifiers
 Linear & nonlinear classifiers Machine Learning Hamid Beigy Sharif University of Technology Fall 1394 Hamid Beigy (Sharif University of Technology) Linear & nonlinear classifiers Fall 1394 1 / 34 Table
Linear & nonlinear classifiers Machine Learning Hamid Beigy Sharif University of Technology Fall 1394 Hamid Beigy (Sharif University of Technology) Linear & nonlinear classifiers Fall 1394 1 / 34 Table
Introduction to Machine Learning
 10-701 Introduction to Machine Learning PCA Slides based on 18-661 Fall 2018 PCA Raw data can be Complex, High-dimensional To understand a phenomenon we measure various related quantities If we knew what
10-701 Introduction to Machine Learning PCA Slides based on 18-661 Fall 2018 PCA Raw data can be Complex, High-dimensional To understand a phenomenon we measure various related quantities If we knew what
Computational Linear Algebra
 Computational Linear Algebra PD Dr. rer. nat. habil. Ralf-Peter Mundani Computation in Engineering / BGU Scientific Computing in Computer Science / INF Winter Term 2018/19 Part 6: Some Other Stuff PD Dr.
Computational Linear Algebra PD Dr. rer. nat. habil. Ralf-Peter Mundani Computation in Engineering / BGU Scientific Computing in Computer Science / INF Winter Term 2018/19 Part 6: Some Other Stuff PD Dr.
CPSC 340: Machine Learning and Data Mining. More PCA Fall 2017
 CPSC 340: Machine Learning and Data Mining More PCA Fall 2017 Admin Assignment 4: Due Friday of next week. No class Monday due to holiday. There will be tutorials next week on MAP/PCA (except Monday).
CPSC 340: Machine Learning and Data Mining More PCA Fall 2017 Admin Assignment 4: Due Friday of next week. No class Monday due to holiday. There will be tutorials next week on MAP/PCA (except Monday).
Lecture: Face Recognition and Feature Reduction
 Lecture: Face Recognition and Feature Reduction Juan Carlos Niebles and Ranjay Krishna Stanford Vision and Learning Lab Lecture 11-1 Recap - Curse of dimensionality Assume 5000 points uniformly distributed
Lecture: Face Recognition and Feature Reduction Juan Carlos Niebles and Ranjay Krishna Stanford Vision and Learning Lab Lecture 11-1 Recap - Curse of dimensionality Assume 5000 points uniformly distributed
Quick Introduction to Nonnegative Matrix Factorization
 Quick Introduction to Nonnegative Matrix Factorization Norm Matloff University of California at Davis 1 The Goal Given an u v matrix A with nonnegative elements, we wish to find nonnegative, rank-k matrices
Quick Introduction to Nonnegative Matrix Factorization Norm Matloff University of California at Davis 1 The Goal Given an u v matrix A with nonnegative elements, we wish to find nonnegative, rank-k matrices
Computational Genomics. Systems biology. Putting it together: Data integration using graphical models
 02-710 Computational Genomics Systems biology Putting it together: Data integration using graphical models High throughput data So far in this class we discussed several different types of high throughput
02-710 Computational Genomics Systems biology Putting it together: Data integration using graphical models High throughput data So far in this class we discussed several different types of high throughput
6.047 / Computational Biology: Genomes, Networks, Evolution Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Linear Algebra and Eigenproblems
 Appendix A A Linear Algebra and Eigenproblems A working knowledge of linear algebra is key to understanding many of the issues raised in this work. In particular, many of the discussions of the details
Appendix A A Linear Algebra and Eigenproblems A working knowledge of linear algebra is key to understanding many of the issues raised in this work. In particular, many of the discussions of the details
Unsupervised learning: beyond simple clustering and PCA
 Unsupervised learning: beyond simple clustering and PCA Liza Rebrova Self organizing maps (SOM) Goal: approximate data points in R p by a low-dimensional manifold Unlike PCA, the manifold does not have
Unsupervised learning: beyond simple clustering and PCA Liza Rebrova Self organizing maps (SOM) Goal: approximate data points in R p by a low-dimensional manifold Unlike PCA, the manifold does not have
CSEP 590A Summer Tonight MLE. FYI, re HW #2: Hemoglobin History. Lecture 4 MLE, EM, RE, Expression. Maximum Likelihood Estimators
 CSEP 59A Summer 26 Lecture 4 MLE, EM, RE, Expression FYI, re HW #2: Hemoglobin History 1 Alberts et al., 3rd ed.,pg389 2 Tonight MLE: Maximum Likelihood Estimators EM: the Expectation Maximization Algorithm
CSEP 59A Summer 26 Lecture 4 MLE, EM, RE, Expression FYI, re HW #2: Hemoglobin History 1 Alberts et al., 3rd ed.,pg389 2 Tonight MLE: Maximum Likelihood Estimators EM: the Expectation Maximization Algorithm
CSEP 590A Summer Lecture 4 MLE, EM, RE, Expression
 CSEP 590A Summer 2006 Lecture 4 MLE, EM, RE, Expression 1 FYI, re HW #2: Hemoglobin History Alberts et al., 3rd ed.,pg389 2 Tonight MLE: Maximum Likelihood Estimators EM: the Expectation Maximization Algorithm
CSEP 590A Summer 2006 Lecture 4 MLE, EM, RE, Expression 1 FYI, re HW #2: Hemoglobin History Alberts et al., 3rd ed.,pg389 2 Tonight MLE: Maximum Likelihood Estimators EM: the Expectation Maximization Algorithm
Vector Space Models. wine_spectral.r
 Vector Space Models 137 wine_spectral.r Latent Semantic Analysis Problem with words Even a small vocabulary as in wine example is challenging LSA Reduce number of columns of DTM by principal components
Vector Space Models 137 wine_spectral.r Latent Semantic Analysis Problem with words Even a small vocabulary as in wine example is challenging LSA Reduce number of columns of DTM by principal components
CS168: The Modern Algorithmic Toolbox Lecture #8: How PCA Works
 CS68: The Modern Algorithmic Toolbox Lecture #8: How PCA Works Tim Roughgarden & Gregory Valiant April 20, 206 Introduction Last lecture introduced the idea of principal components analysis (PCA). The
CS68: The Modern Algorithmic Toolbox Lecture #8: How PCA Works Tim Roughgarden & Gregory Valiant April 20, 206 Introduction Last lecture introduced the idea of principal components analysis (PCA). The
SUPPORTING INFORMATION FOR. SEquence-Enabled Reassembly of β-lactamase (SEER-LAC): a Sensitive Method for the Detection of Double-Stranded DNA
 SUPPORTING INFORMATION FOR SEquence-Enabled Reassembly of β-lactamase (SEER-LAC): a Sensitive Method for the Detection of Double-Stranded DNA Aik T. Ooi, Cliff I. Stains, Indraneel Ghosh *, David J. Segal
SUPPORTING INFORMATION FOR SEquence-Enabled Reassembly of β-lactamase (SEER-LAC): a Sensitive Method for the Detection of Double-Stranded DNA Aik T. Ooi, Cliff I. Stains, Indraneel Ghosh *, David J. Segal
Lecture: Face Recognition and Feature Reduction
 Lecture: Face Recognition and Feature Reduction Juan Carlos Niebles and Ranjay Krishna Stanford Vision and Learning Lab 1 Recap - Curse of dimensionality Assume 5000 points uniformly distributed in the
Lecture: Face Recognition and Feature Reduction Juan Carlos Niebles and Ranjay Krishna Stanford Vision and Learning Lab 1 Recap - Curse of dimensionality Assume 5000 points uniformly distributed in the
Linear Algebra for Machine Learning. Sargur N. Srihari
 Linear Algebra for Machine Learning Sargur N. srihari@cedar.buffalo.edu 1 Overview Linear Algebra is based on continuous math rather than discrete math Computer scientists have little experience with it
Linear Algebra for Machine Learning Sargur N. srihari@cedar.buffalo.edu 1 Overview Linear Algebra is based on continuous math rather than discrete math Computer scientists have little experience with it
Principal Component Analysis
 Machine Learning Michaelmas 2017 James Worrell Principal Component Analysis 1 Introduction 1.1 Goals of PCA Principal components analysis (PCA) is a dimensionality reduction technique that can be used
Machine Learning Michaelmas 2017 James Worrell Principal Component Analysis 1 Introduction 1.1 Goals of PCA Principal components analysis (PCA) is a dimensionality reduction technique that can be used
GCD3033:Cell Biology. Transcription
 Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
Principal Component Analysis -- PCA (also called Karhunen-Loeve transformation)
 Principal Component Analysis -- PCA (also called Karhunen-Loeve transformation) PCA transforms the original input space into a lower dimensional space, by constructing dimensions that are linear combinations
Principal Component Analysis -- PCA (also called Karhunen-Loeve transformation) PCA transforms the original input space into a lower dimensional space, by constructing dimensions that are linear combinations
Regulatory Sequence Analysis. Sequence models (Bernoulli and Markov models)
 Regulatory Sequence Analysis Sequence models (Bernoulli and Markov models) 1 Why do we need random models? Any pattern discovery relies on an underlying model to estimate the random expectation. This model
Regulatory Sequence Analysis Sequence models (Bernoulli and Markov models) 1 Why do we need random models? Any pattern discovery relies on an underlying model to estimate the random expectation. This model
Supplementary Information
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Directed self-assembly of genomic sequences into monomeric and polymeric branched DNA structures
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Directed self-assembly of genomic sequences into monomeric and polymeric branched DNA structures
COMP 551 Applied Machine Learning Lecture 13: Dimension reduction and feature selection
 COMP 551 Applied Machine Learning Lecture 13: Dimension reduction and feature selection Instructor: Herke van Hoof (herke.vanhoof@cs.mcgill.ca) Based on slides by:, Jackie Chi Kit Cheung Class web page:
COMP 551 Applied Machine Learning Lecture 13: Dimension reduction and feature selection Instructor: Herke van Hoof (herke.vanhoof@cs.mcgill.ca) Based on slides by:, Jackie Chi Kit Cheung Class web page:
Focus was on solving matrix inversion problems Now we look at other properties of matrices Useful when A represents a transformations.
 Previously Focus was on solving matrix inversion problems Now we look at other properties of matrices Useful when A represents a transformations y = Ax Or A simply represents data Notion of eigenvectors,
Previously Focus was on solving matrix inversion problems Now we look at other properties of matrices Useful when A represents a transformations y = Ax Or A simply represents data Notion of eigenvectors,
Linear Algebra & Geometry why is linear algebra useful in computer vision?
 Linear Algebra & Geometry why is linear algebra useful in computer vision? References: -Any book on linear algebra! -[HZ] chapters 2, 4 Some of the slides in this lecture are courtesy to Prof. Octavia
Linear Algebra & Geometry why is linear algebra useful in computer vision? References: -Any book on linear algebra! -[HZ] chapters 2, 4 Some of the slides in this lecture are courtesy to Prof. Octavia
20 Unsupervised Learning and Principal Components Analysis (PCA)
 116 Jonathan Richard Shewchuk 20 Unsupervised Learning and Principal Components Analysis (PCA) UNSUPERVISED LEARNING We have sample points, but no labels! No classes, no y-values, nothing to predict. Goal:
116 Jonathan Richard Shewchuk 20 Unsupervised Learning and Principal Components Analysis (PCA) UNSUPERVISED LEARNING We have sample points, but no labels! No classes, no y-values, nothing to predict. Goal:
ECE521 week 3: 23/26 January 2017
 ECE521 week 3: 23/26 January 2017 Outline Probabilistic interpretation of linear regression - Maximum likelihood estimation (MLE) - Maximum a posteriori (MAP) estimation Bias-variance trade-off Linear
ECE521 week 3: 23/26 January 2017 Outline Probabilistic interpretation of linear regression - Maximum likelihood estimation (MLE) - Maximum a posteriori (MAP) estimation Bias-variance trade-off Linear
Linear Algebra & Geometry why is linear algebra useful in computer vision?
 Linear Algebra & Geometry why is linear algebra useful in computer vision? References: -Any book on linear algebra! -[HZ] chapters 2, 4 Some of the slides in this lecture are courtesy to Prof. Octavia
Linear Algebra & Geometry why is linear algebra useful in computer vision? References: -Any book on linear algebra! -[HZ] chapters 2, 4 Some of the slides in this lecture are courtesy to Prof. Octavia
Biology 644: Bioinformatics
 A stochastic (probabilistic) model that assumes the Markov property Markov property is satisfied when the conditional probability distribution of future states of the process (conditional on both past
A stochastic (probabilistic) model that assumes the Markov property Markov property is satisfied when the conditional probability distribution of future states of the process (conditional on both past
STA 414/2104: Lecture 8
 STA 414/2104: Lecture 8 6-7 March 2017: Continuous Latent Variable Models, Neural networks Delivered by Mark Ebden With thanks to Russ Salakhutdinov, Jimmy Ba and others Outline Continuous latent variable
STA 414/2104: Lecture 8 6-7 March 2017: Continuous Latent Variable Models, Neural networks Delivered by Mark Ebden With thanks to Russ Salakhutdinov, Jimmy Ba and others Outline Continuous latent variable
Singular Value Decomposition and Principal Component Analysis (PCA) I
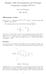 Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Degenerate Perturbation Theory. 1 General framework and strategy
 Physics G6037 Professor Christ 12/22/2015 Degenerate Perturbation Theory The treatment of degenerate perturbation theory presented in class is written out here in detail. The appendix presents the underlying
Physics G6037 Professor Christ 12/22/2015 Degenerate Perturbation Theory The treatment of degenerate perturbation theory presented in class is written out here in detail. The appendix presents the underlying
Fantope Regularization in Metric Learning
 Fantope Regularization in Metric Learning CVPR 2014 Marc T. Law (LIP6, UPMC), Nicolas Thome (LIP6 - UPMC Sorbonne Universités), Matthieu Cord (LIP6 - UPMC Sorbonne Universités), Paris, France Introduction
Fantope Regularization in Metric Learning CVPR 2014 Marc T. Law (LIP6, UPMC), Nicolas Thome (LIP6 - UPMC Sorbonne Universités), Matthieu Cord (LIP6 - UPMC Sorbonne Universités), Paris, France Introduction
Machine learning for pervasive systems Classification in high-dimensional spaces
 Machine learning for pervasive systems Classification in high-dimensional spaces Department of Communications and Networking Aalto University, School of Electrical Engineering stephan.sigg@aalto.fi Version
Machine learning for pervasive systems Classification in high-dimensional spaces Department of Communications and Networking Aalto University, School of Electrical Engineering stephan.sigg@aalto.fi Version
PCA, Kernel PCA, ICA
 PCA, Kernel PCA, ICA Learning Representations. Dimensionality Reduction. Maria-Florina Balcan 04/08/2015 Big & High-Dimensional Data High-Dimensions = Lot of Features Document classification Features per
PCA, Kernel PCA, ICA Learning Representations. Dimensionality Reduction. Maria-Florina Balcan 04/08/2015 Big & High-Dimensional Data High-Dimensions = Lot of Features Document classification Features per
Multiple Choice Review- Eukaryotic Gene Expression
 Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Machine Learning. CUNY Graduate Center, Spring Lectures 11-12: Unsupervised Learning 1. Professor Liang Huang.
 Machine Learning CUNY Graduate Center, Spring 2013 Lectures 11-12: Unsupervised Learning 1 (Clustering: k-means, EM, mixture models) Professor Liang Huang huang@cs.qc.cuny.edu http://acl.cs.qc.edu/~lhuang/teaching/machine-learning
Machine Learning CUNY Graduate Center, Spring 2013 Lectures 11-12: Unsupervised Learning 1 (Clustering: k-means, EM, mixture models) Professor Liang Huang huang@cs.qc.cuny.edu http://acl.cs.qc.edu/~lhuang/teaching/machine-learning
Lecture 9: Numerical Linear Algebra Primer (February 11st)
 10-725/36-725: Convex Optimization Spring 2015 Lecture 9: Numerical Linear Algebra Primer (February 11st) Lecturer: Ryan Tibshirani Scribes: Avinash Siravuru, Guofan Wu, Maosheng Liu Note: LaTeX template
10-725/36-725: Convex Optimization Spring 2015 Lecture 9: Numerical Linear Algebra Primer (February 11st) Lecturer: Ryan Tibshirani Scribes: Avinash Siravuru, Guofan Wu, Maosheng Liu Note: LaTeX template
Predicting Protein Functions and Domain Interactions from Protein Interactions
 Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Methods for sparse analysis of high-dimensional data, II
 Methods for sparse analysis of high-dimensional data, II Rachel Ward May 23, 2011 High dimensional data with low-dimensional structure 300 by 300 pixel images = 90, 000 dimensions 2 / 47 High dimensional
Methods for sparse analysis of high-dimensional data, II Rachel Ward May 23, 2011 High dimensional data with low-dimensional structure 300 by 300 pixel images = 90, 000 dimensions 2 / 47 High dimensional
Phylogenetic Tree Reconstruction
 I519 Introduction to Bioinformatics, 2011 Phylogenetic Tree Reconstruction Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Evolution theory Speciation Evolution of new organisms is driven
I519 Introduction to Bioinformatics, 2011 Phylogenetic Tree Reconstruction Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Evolution theory Speciation Evolution of new organisms is driven
MACHINE LEARNING ADVANCED MACHINE LEARNING
 MACHINE LEARNING ADVANCED MACHINE LEARNING Recap of Important Notions on Estimation of Probability Density Functions 2 2 MACHINE LEARNING Overview Definition pdf Definition joint, condition, marginal,
MACHINE LEARNING ADVANCED MACHINE LEARNING Recap of Important Notions on Estimation of Probability Density Functions 2 2 MACHINE LEARNING Overview Definition pdf Definition joint, condition, marginal,
March 27 Math 3260 sec. 56 Spring 2018
 March 27 Math 3260 sec. 56 Spring 2018 Section 4.6: Rank Definition: The row space, denoted Row A, of an m n matrix A is the subspace of R n spanned by the rows of A. We now have three vector spaces associated
March 27 Math 3260 sec. 56 Spring 2018 Section 4.6: Rank Definition: The row space, denoted Row A, of an m n matrix A is the subspace of R n spanned by the rows of A. We now have three vector spaces associated
Systems of Linear Equations
 LECTURE 6 Systems of Linear Equations You may recall that in Math 303, matrices were first introduced as a means of encapsulating the essential data underlying a system of linear equations; that is to
LECTURE 6 Systems of Linear Equations You may recall that in Math 303, matrices were first introduced as a means of encapsulating the essential data underlying a system of linear equations; that is to
Systems of Linear Equations
 Systems of Linear Equations Math 108A: August 21, 2008 John Douglas Moore Our goal in these notes is to explain a few facts regarding linear systems of equations not included in the first few chapters
Systems of Linear Equations Math 108A: August 21, 2008 John Douglas Moore Our goal in these notes is to explain a few facts regarding linear systems of equations not included in the first few chapters
Chapter 2. Error Correcting Codes. 2.1 Basic Notions
 Chapter 2 Error Correcting Codes The identification number schemes we discussed in the previous chapter give us the ability to determine if an error has been made in recording or transmitting information.
Chapter 2 Error Correcting Codes The identification number schemes we discussed in the previous chapter give us the ability to determine if an error has been made in recording or transmitting information.
Example Linear Algebra Competency Test
 Example Linear Algebra Competency Test The 4 questions below are a combination of True or False, multiple choice, fill in the blank, and computations involving matrices and vectors. In the latter case,
Example Linear Algebra Competency Test The 4 questions below are a combination of True or False, multiple choice, fill in the blank, and computations involving matrices and vectors. In the latter case,
Linear & nonlinear classifiers
 Linear & nonlinear classifiers Machine Learning Hamid Beigy Sharif University of Technology Fall 1396 Hamid Beigy (Sharif University of Technology) Linear & nonlinear classifiers Fall 1396 1 / 44 Table
Linear & nonlinear classifiers Machine Learning Hamid Beigy Sharif University of Technology Fall 1396 Hamid Beigy (Sharif University of Technology) Linear & nonlinear classifiers Fall 1396 1 / 44 Table
A Modular NMF Matching Algorithm for Radiation Spectra
 A Modular NMF Matching Algorithm for Radiation Spectra Melissa L. Koudelka Sensor Exploitation Applications Sandia National Laboratories mlkoude@sandia.gov Daniel J. Dorsey Systems Technologies Sandia
A Modular NMF Matching Algorithm for Radiation Spectra Melissa L. Koudelka Sensor Exploitation Applications Sandia National Laboratories mlkoude@sandia.gov Daniel J. Dorsey Systems Technologies Sandia
Machine Learning for Signal Processing Sparse and Overcomplete Representations
 Machine Learning for Signal Processing Sparse and Overcomplete Representations Abelino Jimenez (slides from Bhiksha Raj and Sourish Chaudhuri) Oct 1, 217 1 So far Weights Data Basis Data Independent ICA
Machine Learning for Signal Processing Sparse and Overcomplete Representations Abelino Jimenez (slides from Bhiksha Raj and Sourish Chaudhuri) Oct 1, 217 1 So far Weights Data Basis Data Independent ICA
Statistical Machine Learning from Data
 Samy Bengio Statistical Machine Learning from Data 1 Statistical Machine Learning from Data Support Vector Machines Samy Bengio IDIAP Research Institute, Martigny, Switzerland, and Ecole Polytechnique
Samy Bengio Statistical Machine Learning from Data 1 Statistical Machine Learning from Data Support Vector Machines Samy Bengio IDIAP Research Institute, Martigny, Switzerland, and Ecole Polytechnique
Practical considerations of working with sequencing data
 Practical considerations of working with sequencing data File Types Fastq ->aligner -> reference(genome) coordinates Coordinate files SAM/BAM most complete, contains all of the info in fastq and more!
Practical considerations of working with sequencing data File Types Fastq ->aligner -> reference(genome) coordinates Coordinate files SAM/BAM most complete, contains all of the info in fastq and more!
Degenerate Perturbation Theory
 Physics G6037 Professor Christ 12/05/2014 Degenerate Perturbation Theory The treatment of degenerate perturbation theory presented in class is written out here in detail. 1 General framework and strategy
Physics G6037 Professor Christ 12/05/2014 Degenerate Perturbation Theory The treatment of degenerate perturbation theory presented in class is written out here in detail. 1 General framework and strategy
Lecture 7: Con3nuous Latent Variable Models
 CSC2515 Fall 2015 Introduc3on to Machine Learning Lecture 7: Con3nuous Latent Variable Models All lecture slides will be available as.pdf on the course website: http://www.cs.toronto.edu/~urtasun/courses/csc2515/
CSC2515 Fall 2015 Introduc3on to Machine Learning Lecture 7: Con3nuous Latent Variable Models All lecture slides will be available as.pdf on the course website: http://www.cs.toronto.edu/~urtasun/courses/csc2515/
Statistical Pattern Recognition
 Statistical Pattern Recognition Feature Extraction Hamid R. Rabiee Jafar Muhammadi, Alireza Ghasemi, Payam Siyari Spring 2014 http://ce.sharif.edu/courses/92-93/2/ce725-2/ Agenda Dimensionality Reduction
Statistical Pattern Recognition Feature Extraction Hamid R. Rabiee Jafar Muhammadi, Alireza Ghasemi, Payam Siyari Spring 2014 http://ce.sharif.edu/courses/92-93/2/ce725-2/ Agenda Dimensionality Reduction
PROTEIN SYNTHESIS INTRO
 MR. POMERANTZ Page 1 of 6 Protein synthesis Intro. Use the text book to help properly answer the following questions 1. RNA differs from DNA in that RNA a. is single-stranded. c. contains the nitrogen
MR. POMERANTZ Page 1 of 6 Protein synthesis Intro. Use the text book to help properly answer the following questions 1. RNA differs from DNA in that RNA a. is single-stranded. c. contains the nitrogen
BMD645. Integration of Omics
 BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
Machine Learning - MT & 14. PCA and MDS
 Machine Learning - MT 2016 13 & 14. PCA and MDS Varun Kanade University of Oxford November 21 & 23, 2016 Announcements Sheet 4 due this Friday by noon Practical 3 this week (continue next week if necessary)
Machine Learning - MT 2016 13 & 14. PCA and MDS Varun Kanade University of Oxford November 21 & 23, 2016 Announcements Sheet 4 due this Friday by noon Practical 3 this week (continue next week if necessary)
Algebraic Statistics progress report
 Algebraic Statistics progress report Joe Neeman December 11, 2008 1 A model for biochemical reaction networks We consider a model introduced by Craciun, Pantea and Rempala [2] for identifying biochemical
Algebraic Statistics progress report Joe Neeman December 11, 2008 1 A model for biochemical reaction networks We consider a model introduced by Craciun, Pantea and Rempala [2] for identifying biochemical
Unsupervised Machine Learning and Data Mining. DS 5230 / DS Fall Lecture 7. Jan-Willem van de Meent
 Unsupervised Machine Learning and Data Mining DS 5230 / DS 4420 - Fall 2018 Lecture 7 Jan-Willem van de Meent DIMENSIONALITY REDUCTION Borrowing from: Percy Liang (Stanford) Dimensionality Reduction Goal:
Unsupervised Machine Learning and Data Mining DS 5230 / DS 4420 - Fall 2018 Lecture 7 Jan-Willem van de Meent DIMENSIONALITY REDUCTION Borrowing from: Percy Liang (Stanford) Dimensionality Reduction Goal:
EE613 Machine Learning for Engineers. Kernel methods Support Vector Machines. jean-marc odobez 2015
 EE613 Machine Learning for Engineers Kernel methods Support Vector Machines jean-marc odobez 2015 overview Kernel methods introductions and main elements defining kernels Kernelization of k-nn, K-Means,
EE613 Machine Learning for Engineers Kernel methods Support Vector Machines jean-marc odobez 2015 overview Kernel methods introductions and main elements defining kernels Kernelization of k-nn, K-Means,
Cheng Soon Ong & Christian Walder. Canberra February June 2018
 Cheng Soon Ong & Christian Walder Research Group and College of Engineering and Computer Science Canberra February June 2018 Outlines Overview Introduction Linear Algebra Probability Linear Regression
Cheng Soon Ong & Christian Walder Research Group and College of Engineering and Computer Science Canberra February June 2018 Outlines Overview Introduction Linear Algebra Probability Linear Regression
EECS 275 Matrix Computation
 EECS 275 Matrix Computation Ming-Hsuan Yang Electrical Engineering and Computer Science University of California at Merced Merced, CA 95344 http://faculty.ucmerced.edu/mhyang Lecture 22 1 / 21 Overview
EECS 275 Matrix Computation Ming-Hsuan Yang Electrical Engineering and Computer Science University of California at Merced Merced, CA 95344 http://faculty.ucmerced.edu/mhyang Lecture 22 1 / 21 Overview
MATH Linear Algebra
 MATH 304 - Linear Algebra In the previous note we learned an important algorithm to produce orthogonal sequences of vectors called the Gramm-Schmidt orthogonalization process. Gramm-Schmidt orthogonalization
MATH 304 - Linear Algebra In the previous note we learned an important algorithm to produce orthogonal sequences of vectors called the Gramm-Schmidt orthogonalization process. Gramm-Schmidt orthogonalization
Logistic Regression. Will Monroe CS 109. Lecture Notes #22 August 14, 2017
 1 Will Monroe CS 109 Logistic Regression Lecture Notes #22 August 14, 2017 Based on a chapter by Chris Piech Logistic regression is a classification algorithm1 that works by trying to learn a function
1 Will Monroe CS 109 Logistic Regression Lecture Notes #22 August 14, 2017 Based on a chapter by Chris Piech Logistic regression is a classification algorithm1 that works by trying to learn a function
Preliminary/Qualifying Exam in Numerical Analysis (Math 502a) Spring 2012
 Instructions Preliminary/Qualifying Exam in Numerical Analysis (Math 502a) Spring 2012 The exam consists of four problems, each having multiple parts. You should attempt to solve all four problems. 1.
Instructions Preliminary/Qualifying Exam in Numerical Analysis (Math 502a) Spring 2012 The exam consists of four problems, each having multiple parts. You should attempt to solve all four problems. 1.
Bayes methods for categorical data. April 25, 2017
 Bayes methods for categorical data April 25, 2017 Motivation for joint probability models Increasing interest in high-dimensional data in broad applications Focus may be on prediction, variable selection,
Bayes methods for categorical data April 25, 2017 Motivation for joint probability models Increasing interest in high-dimensional data in broad applications Focus may be on prediction, variable selection,
Chapter 3 Transformations
 Chapter 3 Transformations An Introduction to Optimization Spring, 2014 Wei-Ta Chu 1 Linear Transformations A function is called a linear transformation if 1. for every and 2. for every If we fix the bases
Chapter 3 Transformations An Introduction to Optimization Spring, 2014 Wei-Ta Chu 1 Linear Transformations A function is called a linear transformation if 1. for every and 2. for every If we fix the bases
CS281 Section 4: Factor Analysis and PCA
 CS81 Section 4: Factor Analysis and PCA Scott Linderman At this point we have seen a variety of machine learning models, with a particular emphasis on models for supervised learning. In particular, we
CS81 Section 4: Factor Analysis and PCA Scott Linderman At this point we have seen a variety of machine learning models, with a particular emphasis on models for supervised learning. In particular, we
SUPPLEMENTARY DATA - 1 -
 - 1 - SUPPLEMENTARY DATA Construction of B. subtilis rnpb complementation plasmids For complementation, the B. subtilis rnpb wild-type gene (rnpbwt) under control of its native rnpb promoter and terminator
- 1 - SUPPLEMENTARY DATA Construction of B. subtilis rnpb complementation plasmids For complementation, the B. subtilis rnpb wild-type gene (rnpbwt) under control of its native rnpb promoter and terminator
c Springer, Reprinted with permission.
 Zhijian Yuan and Erkki Oja. A FastICA Algorithm for Non-negative Independent Component Analysis. In Puntonet, Carlos G.; Prieto, Alberto (Eds.), Proceedings of the Fifth International Symposium on Independent
Zhijian Yuan and Erkki Oja. A FastICA Algorithm for Non-negative Independent Component Analysis. In Puntonet, Carlos G.; Prieto, Alberto (Eds.), Proceedings of the Fifth International Symposium on Independent
PCA FACE RECOGNITION
 PCA FACE RECOGNITION The slides are from several sources through James Hays (Brown); Srinivasa Narasimhan (CMU); Silvio Savarese (U. of Michigan); Shree Nayar (Columbia) including their own slides. Goal
PCA FACE RECOGNITION The slides are from several sources through James Hays (Brown); Srinivasa Narasimhan (CMU); Silvio Savarese (U. of Michigan); Shree Nayar (Columbia) including their own slides. Goal
Latent Variable models for GWAs
 Latent Variable models for GWAs Oliver Stegle Machine Learning and Computational Biology Research Group Max-Planck-Institutes Tübingen, Germany September 2011 O. Stegle Latent variable models for GWAs
Latent Variable models for GWAs Oliver Stegle Machine Learning and Computational Biology Research Group Max-Planck-Institutes Tübingen, Germany September 2011 O. Stegle Latent variable models for GWAs
Machine Learning. Support Vector Machines. Manfred Huber
 Machine Learning Support Vector Machines Manfred Huber 2015 1 Support Vector Machines Both logistic regression and linear discriminant analysis learn a linear discriminant function to separate the data
Machine Learning Support Vector Machines Manfred Huber 2015 1 Support Vector Machines Both logistic regression and linear discriminant analysis learn a linear discriminant function to separate the data
Dimensionality Reduction:
 Dimensionality Reduction: From Data Representation to General Framework Dong XU School of Computer Engineering Nanyang Technological University, Singapore What is Dimensionality Reduction? PCA LDA Examples:
Dimensionality Reduction: From Data Representation to General Framework Dong XU School of Computer Engineering Nanyang Technological University, Singapore What is Dimensionality Reduction? PCA LDA Examples:
A vector from the origin to H, V could be expressed using:
 Linear Discriminant Function: the linear discriminant function: g(x) = w t x + ω 0 x is the point, w is the weight vector, and ω 0 is the bias (t is the transpose). Two Category Case: In the two category
Linear Discriminant Function: the linear discriminant function: g(x) = w t x + ω 0 x is the point, w is the weight vector, and ω 0 is the bias (t is the transpose). Two Category Case: In the two category
L11: Pattern recognition principles
 L11: Pattern recognition principles Bayesian decision theory Statistical classifiers Dimensionality reduction Clustering This lecture is partly based on [Huang, Acero and Hon, 2001, ch. 4] Introduction
L11: Pattern recognition principles Bayesian decision theory Statistical classifiers Dimensionality reduction Clustering This lecture is partly based on [Huang, Acero and Hon, 2001, ch. 4] Introduction
Assignment 1 Math 5341 Linear Algebra Review. Give complete answers to each of the following questions. Show all of your work.
 Assignment 1 Math 5341 Linear Algebra Review Give complete answers to each of the following questions Show all of your work Note: You might struggle with some of these questions, either because it has
Assignment 1 Math 5341 Linear Algebra Review Give complete answers to each of the following questions Show all of your work Note: You might struggle with some of these questions, either because it has
Linear Algebra 1 Exam 2 Solutions 7/14/3
 Linear Algebra 1 Exam Solutions 7/14/3 Question 1 The line L has the symmetric equation: x 1 = y + 3 The line M has the parametric equation: = z 4. [x, y, z] = [ 4, 10, 5] + s[10, 7, ]. The line N is perpendicular
Linear Algebra 1 Exam Solutions 7/14/3 Question 1 The line L has the symmetric equation: x 1 = y + 3 The line M has the parametric equation: = z 4. [x, y, z] = [ 4, 10, 5] + s[10, 7, ]. The line N is perpendicular
r=1 r=1 argmin Q Jt (20) After computing the descent direction d Jt 2 dt H t d + P (x + d) d i = 0, i / J
 7 Appendix 7. Proof of Theorem Proof. There are two main difficulties in proving the convergence of our algorithm, and none of them is addressed in previous works. First, the Hessian matrix H is a block-structured
7 Appendix 7. Proof of Theorem Proof. There are two main difficulties in proving the convergence of our algorithm, and none of them is addressed in previous works. First, the Hessian matrix H is a block-structured
Uncorrelated Multilinear Principal Component Analysis through Successive Variance Maximization
 Uncorrelated Multilinear Principal Component Analysis through Successive Variance Maximization Haiping Lu 1 K. N. Plataniotis 1 A. N. Venetsanopoulos 1,2 1 Department of Electrical & Computer Engineering,
Uncorrelated Multilinear Principal Component Analysis through Successive Variance Maximization Haiping Lu 1 K. N. Plataniotis 1 A. N. Venetsanopoulos 1,2 1 Department of Electrical & Computer Engineering,
PCA and admixture models
 PCA and admixture models CM226: Machine Learning for Bioinformatics. Fall 2016 Sriram Sankararaman Acknowledgments: Fei Sha, Ameet Talwalkar, Alkes Price PCA and admixture models 1 / 57 Announcements HW1
PCA and admixture models CM226: Machine Learning for Bioinformatics. Fall 2016 Sriram Sankararaman Acknowledgments: Fei Sha, Ameet Talwalkar, Alkes Price PCA and admixture models 1 / 57 Announcements HW1
