Non-bonded interactions
|
|
|
- Gertrude Mitchell
- 5 years ago
- Views:
Transcription
1 speeding up the number-crunching continued Marcus Elstner and Tomáš Kubař December 3, 2013
2 why care? number of individual pair-wise interactions bonded interactions proportional to N: O(N) non-bonded interactions between every pair of atoms computational effort of O(N 2 ) take the largest part of computational time good target of optimization
3 Intro to electrostatic interaction Coulomb s law elstat. interaction energy of point charges q and Q on distance r: E el = 1 q Q 4πε 0 r electrostatic potential (ESP) induced at r by a number of point charges Φ( r) = 1 Q i 4πε 0 r r i if we know ESP at r, and q is placed there, elstat. energy follows as E el ( r) = Φ( r) q i
4 Intro to electrostatic interaction Coulomb s law continuous charge distribution charge density ρ = Q/ V = lim V Q/ V summing the potential induced by charge in volume elements V gives Φ( r) = 1 4πε 0 i ρ( r i ) V r r i for infinitesimal volume elements (with d 3 r = dv ): Φ( r) = 1 4πε 0 ρ( r 1 ) r r 1 d3 r 1
5 Intro to electrostatic interaction Poisson s equation needs to be solved to get ESP from charge distribution (differential equation for Φ as a function of r): 2 Φ( r) = ρ( r) ε example: ESP of Gaussian charge density around o with width σ: [ ] 1 r 2 ρ(r) = Q σ 3 2π 3 exp 2σ 2 solution of Poisson s equation: Φ(r) = 1 4πε Q r erf [ r 2σ ]
6 Intro to electrostatic interaction Poisson s equation solution for Gaussian charge density: Φ(r) = 1 4πε Q r erf [ r 2σ ]
7 Intro to electrostatic interaction Poisson s equation solution for Gaussian charge density: Φ(r) = 1 4πε Q r erf [ r 2σ ] if we move far from the center of charge density (r is large) erf converges to 1, ESP equals that of a point charge placed in o a point charge and a localized charge density look the same from large distance the distance need not be too large actually: erf[x] = already for x = 2.4
8 Periodic boundary conditions Biomolecule in solution typical setup of a biomolecular MD simulation a biomolecule / complex + several 1000s H 2 O molecules (as few as possible) in a cube n n n nm issue everything is close to the surface, while we are interested in a molecule in bulk solvent
9 Periodic boundary conditions Periodic boundary conditions molecular system placed in a regular-shaped box the box is virtually replicated in all spatial directions this way, the system is infinite no surface! the problems of a system in vacuo disappear
10 Periodic boundary conditions PBC example
11 Periodic boundary conditions PBC features positions / velocities of all atoms are identical in all replicas, so that we can keep only one copy in the memory an atom that leaves the box enters it on the other side the atoms near the wall of the simulation cell interact with the atoms in the neighboring replica carefull accounting of interactions between atoms is necessary
12 Periodic boundary conditions PBC box shape may be simple cubic or orthorhombic, parallelepiped (specially, rhombohedron), or hexagonal prism
13 Periodic boundary conditions PBC box shape... but also more complicated truncated octahedral or rhombic dodecahedral quite complex equations for interactions & eqns of motion advantage for simulation of spherical objects (globular proteins) no corners far from the molecule filled with unnecessary H 2 O
14 Periodic boundary conditions PBC box shape 2D objects phase interfaces, membrane systems usually treated in a slab geometry
15 Accelerating non-bonded interactions Cut-off simple idea non-bonded terms bottleneck of the calculation with PBC infinite number of interaction pairs, but the interaction decreases with increasing distance simplest and crudest approach to limit the number of calculations neglect interaction of atoms further apart than r c cut-off very good for rapidly decaying LJ interaction (1/r 6 ) (r c = 10 Å) not so good for slowly decaying electrostatics (1/r) discontinuity of potential energy and, even worse, the forces
16 Accelerating non-bonded interactions Cut-off better alternatives elstat. interaction energy of two unit positive charges shift / switch applied to either energy or force cut-off schemes based on functional groups, if applicable
17 Accelerating non-bonded interactions Neighbor lists cut-off we still have to calculate the distance for every two atoms (to compare it with the cut-off distance) we do not win much yet there are still O(N 2 ) distances observation: pick an atom A. the atoms that are within cut-off distance r c around A, remain within r c for several consecutive steps of dynamics, while no other atoms approach A that close idea: maybe it is only necessary to calculate the interactions between A and these close atoms neighbors
18 Accelerating non-bonded interactions Neighbor lists
19 Accelerating non-bonded interactions Neighbor lists what will we do? calculate the distances for every pair of atoms less frequently, i.e. every 10 or 20 steps of dynamics, and record the atoms within cut-off distance in a neighbor list then calculate the interaction for each atom only with for the atoms in the neighbor list formally O(N) note the build of the neighbor list itself is O(N 2 ), which can be reduced with further tricks ( cell lists )
20 Complete treatment of non-bonded interactions Accounting of all of the replicas cut-off often bad approximation, e.g. with highly charged molecular systems (DNA, some proteins) switching functions bring on artificial forces e.g. artificial accumulation of ions around cut-off the only solution abandon the cut-off sum up the long-range Coulomb interaction between all the replicas of the simulation cell
21 Complete treatment of non-bonded interactions Can we sum it up simply? sum of Coulomb interactions over all replicas: E Coul = 1 2 i,j replicas n q i q j r ij + n i and j run over all atoms in the unit cell (r ij their distance) infinite sum with special convergence problems alternating harmonic series n ( 1)n /n conditionally convergent: it converges i=1 a i <, but does not converge absolutely: i=1 a i = convergence is slow and dependent on the order of summation
22 Complete treatment of non-bonded interactions Sum it up not so simply Ewald any smart way to calculate ESP induced by all images of all atoms? Φ( r i ) = j replicas n q j r ij + n to get the Coulomb energy of charges q i in the unit cell E Coul = 1 q i Φ( r i ) 2 idea: pass to a sum of two series that will converge rapidly: 1 r = f (r) r i + 1 f (r) r may seem awkward, but will work well
23 Complete treatment of non-bonded interactions Sum it up not so simply Ewald summing over point charges difficult (convergence problem) Ewald method Gaussian densities of the same magnitude: ( ) α 3 q j q j exp [ α 2 r j 2] π obtain ESP with Poisson s equation for 1 Gaussian ESP has a form of error function erf[x] = 2 π x erfc[x] = 1 erf[x] 0 exp[ t 2 ] dt with Ewald: summing ESP induced by all charges, we obtain Φ( r i ) = j replicas n q j erf [α r ij + n ] r ij + n
24 Complete treatment of non-bonded interactions Sum it up not so simply Ewald the full ESP induced by all replicas of all charges: Φ( r i ) = j + j replicas n replicas n = Φ real ( r i ) + Φ rec ( r i ) q j erfc [α r ij + n ] r ij + n q j erf [α r ij + n ] r ij + n Φ real ( r i ) real-space contribution from a certain, quite small distance (depending on α): point charges and the charge densities cancel each other this contribution vanishes and we can use cut-off here
25 Complete treatment of non-bonded interactions Ewald two contributions real-space contribution to the Ewald sum original point charges (red) and Gaussian charge densities (blue) of the same magnitude but opposite sign a distant Gaussian looks much like a point charge, and the difference of ESP goes to zero cut-off is justified reciprocal-space contribution
26 Complete treatment of non-bonded interactions Ewald 2nd contribution the total charge density is periodic it may be meaningful to Fourier-transform the calculation to the reciprocal space (density) solve (hard) (potential) Fourier transform inverse Fourier transform (transformed density) solve (easy) (transformed potential) ˆf (ξ) = Re ˆf (ξ) = Im ˆf (ξ) = f (x) exp[ i x ξ] dx f (x) cos[x ξ] dx f (x) sin[x ξ] dx
27 Complete treatment of non-bonded interactions Ewald 2nd contribution the total charge density is periodic it may be meaningful to Fourier-transform the calculation to the reciprocal space (density) solve (hard) (potential) Fourier transform inverse Fourier transform (transformed density) solve (easy) (transformed potential) 2 Φ( r) = ρ( r) ε k 2 ˆΦ( k) = ˆρ( k) ε hard easy
28 Complete treatment of non-bonded interactions Ewald 2nd contribution Φ rec ( r i ) reciprocal-space contribution with reciprocal vector ( k = k x 2π L x, k y 2π L y, k z 2π L z ), k i Z best evaluated in the form Φ rec ( r i ) = 4π V [ ] 1 exp k 2 k 2 4α 2 q j exp[ i k r ij ] k o j terms decrease with increasing k quickly cut-off possible converges fast with large Gaussian width α therefore, the value of α is a compromise between the requirements of real- and reciprocal-space calcul. both contributions favorable convergence behavior we can evaluate electrostatic interactions with atoms in all periodic images
29 Complete treatment of non-bonded interactions Ewald the last contribution broadened charge density interacts with itself and this energy must be subtracted from the final result Coulomb self-energy of a broadened Gaussian: E self = j q j Φ( o) = j q j q j α π
30 Complete treatment of non-bonded interactions Ewald complete expression for energy 3 contributions: real-space E real = 1 q j Φ real ( r j ) 2 j reciprocal-space E rec = 1 q j Φ rec ( r j ) 2 self-energy E Ewald = E real + E rec E self j
31 Complete AND efficient treatment of electrostatics Thinking about Ewald Ewald summation correct Coulomb interaction energy at a quite high computational cost: scales with the number of atoms as O(N 2 ) with a better algorithm O(N 3 2 ) not efficient enough for large-scale simulations goal improved efficiency of the long-range sum (reciprocal-space contribution)
32 Complete AND efficient treatment of electrostatics Thinking about Ewald Ewald summation correct Coulomb interaction energy at a quite high computational cost: scales with the number of atoms as O(N 2 ) with a better algorithm O(N 3 2 ) not efficient enough for large-scale simulations goal improved efficiency of the long-range sum (reciprocal-space contribution) particle mesh Ewald method (1993) combines ideas from crystallography (Ewald method) and plasma physics (particle mesh method) key to success 3D fast Fourier transform technique
33 Complete AND efficient treatment of electrostatics Long range energy with PME PME works with a regular grid constructed in the simulation box step 1 convert the point charges to Gaussian charge densities and spread on the grid in the form of splines series of polynoms practically, we need to have charges discretized on the grid points if an atom is close to the edge of the box, a part of its charge must be put to the opposite side of the box (PBC) step 2 Fourier transform the charge density on the grid discrete 3D fast Fourier transform technique
34 Complete AND efficient treatment of electrostatics Long range energy with PME step 3 solve Poisson s eqn in the reciprocal space energy and Fourier transform of potential E rec = 1 2 K 1 1 K 2 1 K 3 1 k 1 =0 k 2 =0 k 3 =0 Q(k 1, k 2, k 3 ) (Θ rec Q)(k 1, k 2, k 3 ) gridded charge array Q( k) represents the transformed density ˆρ Θ rec is given by box size and character of splines easy to get 3D-FFT used to calculate the convolution Θ rec Q this gives the transformed ESP in recip. space, ˆΦ rec
35 Complete AND efficient treatment of electrostatics Long range energy with PME step 4 get the potential in real space (inverse Fourier transform), interpolate its derivative to calculate the forces expressed in terms of splines analytical calculation step 5 get E real and E self directly from the presented expressions step 6 attention the reciprocal energy/forces include contributions from atom pairs that are connected with bonds these have to be subtracted afterwards (excluded) list Eexcl rec = 1 q i q j erf [α r ij ] 4πε 0 r ij i,j
36 Complete AND efficient treatment of electrostatics Long range energy with PME PME parameters: spacing of grid ca. 1 Å, α 1 ca. 2.5 Å short-range cutoff 10 Å possible neighbor lists every nl steps, the following procedure: measure distances of every pair of atoms in system distance smaller than a pre-set value? put the pair on the list in the following nl steps, calculate short-ranged interactions only for the pairs on this neighbor list linear scaling of short-range interactions (O(N)) complexity of the long-range PME component: O(N log N) due to the efficiency of FFT modern implementations nearly as efficient as cut-off!
37 Explicit water models Water in biomolecular simulations most simulations something in aqueous solutions H 2 O usually (many) thousands molecules
38 Explicit water models Water in biomolecular simulations most simulations something in aqueous solutions H 2 O usually (many) thousand molecules example simulation of DNA decanucleotide: PBC box nm (smallest meaningful) 630 atoms in DNA, 8346 atoms in water and 18 Na + concentration of DNA: 18 mmol/l very high! of all pair interactions: 86 % are water water, most of the others involve water
39 Explicit water models Water models most interactions involve H 2 O necessary to pay attention to its description model of water must be simple enough (computational cost) and accurate enough, at the same time water models usually rigid bond lengths and angles do not vary constraints molecule with three sites (atoms in this case), or up to six sites three atoms and virtual sites corresponding to a center of electron density or lone electron pairs
40 Explicit water models Water models TIP3P (very similar is SPC) TIP4P TIP5P most frequently used 3 atoms with 3 rigid bonds, charge on every atom ( 0.834/+0.417) only the O possesses non-zero LJ parameters (optimization) negative charge placed on virtual site M rather than on the O electric field around the molecule described better 2 virtual sites L with negative charges near the O lone pairs better description of directionality of H-bonding etc. (radial distribution function, temperature of highest density)
Non-bonded interactions
 speeding up the number-crunching Marcus Elstner and Tomáš Kubař May 8, 2015 why care? key to understand biomolecular structure and function binding of a ligand efficiency of a reaction color of a chromophore
speeding up the number-crunching Marcus Elstner and Tomáš Kubař May 8, 2015 why care? key to understand biomolecular structure and function binding of a ligand efficiency of a reaction color of a chromophore
Biomolecular modeling II
 2015, December 16 System boundary and the solvent Biomolecule in solution typical MD simulations molecular system in aqueous solution task make the system as small as possible (reduce cost) straightforward
2015, December 16 System boundary and the solvent Biomolecule in solution typical MD simulations molecular system in aqueous solution task make the system as small as possible (reduce cost) straightforward
Computer simulation methods (2) Dr. Vania Calandrini
 Computer simulation methods (2) Dr. Vania Calandrini in the previous lecture: time average versus ensemble average MC versus MD simulations equipartition theorem (=> computing T) virial theorem (=> computing
Computer simulation methods (2) Dr. Vania Calandrini in the previous lecture: time average versus ensemble average MC versus MD simulations equipartition theorem (=> computing T) virial theorem (=> computing
Advanced Molecular Molecular Dynamics
 Advanced Molecular Molecular Dynamics Technical details May 12, 2014 Integration of harmonic oscillator r m period = 2 k k and the temperature T determine the sampling of x (here T is related with v 0
Advanced Molecular Molecular Dynamics Technical details May 12, 2014 Integration of harmonic oscillator r m period = 2 k k and the temperature T determine the sampling of x (here T is related with v 0
An FPGA Implementation of Reciprocal Sums for SPME
 An FPGA Implementation of Reciprocal Sums for SPME Sam Lee and Paul Chow Edward S. Rogers Sr. Department of Electrical and Computer Engineering University of Toronto Objectives Accelerate part of Molecular
An FPGA Implementation of Reciprocal Sums for SPME Sam Lee and Paul Chow Edward S. Rogers Sr. Department of Electrical and Computer Engineering University of Toronto Objectives Accelerate part of Molecular
Neighbor Tables Long-Range Potentials
 Neighbor Tables Long-Range Potentials Today we learn how we can handle long range potentials. Neighbor tables Long-range potential Ewald sums MSE485/PHY466/CSE485 1 Periodic distances Minimum Image Convention:
Neighbor Tables Long-Range Potentials Today we learn how we can handle long range potentials. Neighbor tables Long-range potential Ewald sums MSE485/PHY466/CSE485 1 Periodic distances Minimum Image Convention:
Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar.
 Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar. 25, 2002 Molecular Dynamics: Introduction At physiological conditions, the
Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar. 25, 2002 Molecular Dynamics: Introduction At physiological conditions, the
Simulations of bulk phases. Periodic boundaries. Cubic boxes
 Simulations of bulk phases ChE210D Today's lecture: considerations for setting up and running simulations of bulk, isotropic phases (e.g., liquids and gases) Periodic boundaries Cubic boxes In simulations
Simulations of bulk phases ChE210D Today's lecture: considerations for setting up and running simulations of bulk, isotropic phases (e.g., liquids and gases) Periodic boundaries Cubic boxes In simulations
Computational Molecular Biophysics. Computational Biophysics, GRS Jülich SS 2013
 Computational Molecular Biophysics Computational Biophysics, GRS Jülich SS 2013 Computational Considerations Number of terms for different energy contributions / E el ~N ~N ~N ~N 2 Computational cost for
Computational Molecular Biophysics Computational Biophysics, GRS Jülich SS 2013 Computational Considerations Number of terms for different energy contributions / E el ~N ~N ~N ~N 2 Computational cost for
Notes on Ewald summation techniques
 February 3, 011 Notes on Ewald summation techniques Adapted from similar presentation in PHY 71 he total electrostatic potential energy of interaction between point charges {q i } at the positions {r i
February 3, 011 Notes on Ewald summation techniques Adapted from similar presentation in PHY 71 he total electrostatic potential energy of interaction between point charges {q i } at the positions {r i
Tutorial on the smooth particle-mesh Ewald algorithm
 Tutorial on the smooth particle-mesh Ewald algorithm These notes explain the smooth particle-mesh Ewald algorithm (SPME) in detail. The accompanying library libpme6 is a fully-featured implementation of
Tutorial on the smooth particle-mesh Ewald algorithm These notes explain the smooth particle-mesh Ewald algorithm (SPME) in detail. The accompanying library libpme6 is a fully-featured implementation of
Lecture 12: Solvation Models: Molecular Mechanics Modeling of Hydration Effects
 Statistical Thermodynamics Lecture 12: Solvation Models: Molecular Mechanics Modeling of Hydration Effects Dr. Ronald M. Levy ronlevy@temple.edu Bare Molecular Mechanics Atomistic Force Fields: torsion
Statistical Thermodynamics Lecture 12: Solvation Models: Molecular Mechanics Modeling of Hydration Effects Dr. Ronald M. Levy ronlevy@temple.edu Bare Molecular Mechanics Atomistic Force Fields: torsion
MOLECULAR DYNAMICS SIMULATIONS OF BIOMOLECULES: Long-Range Electrostatic Effects
 Annu. Rev. Biophys. Biomol. Struct. 1999. 28:155 79 Copyright c 1999 by Annual Reviews. All rights reserved MOLECULAR DYNAMICS SIMULATIONS OF BIOMOLECULES: Long-Range Electrostatic Effects Celeste Sagui
Annu. Rev. Biophys. Biomol. Struct. 1999. 28:155 79 Copyright c 1999 by Annual Reviews. All rights reserved MOLECULAR DYNAMICS SIMULATIONS OF BIOMOLECULES: Long-Range Electrostatic Effects Celeste Sagui
Long-range interactions: P 3 M, MMMxD, ELC, MEMD and ICC
 Long-range interactions: P 3 M, MMMxD, ELC, MEMD and ICC Axel Arnold Institute for Computational Physics Universität Stuttgart October 10, 2012 Long-range interactions gravity Coulomb interaction dipolar
Long-range interactions: P 3 M, MMMxD, ELC, MEMD and ICC Axel Arnold Institute for Computational Physics Universität Stuttgart October 10, 2012 Long-range interactions gravity Coulomb interaction dipolar
Molecular Dynamics Simulations
 Molecular Dynamics Simulations Dr. Kasra Momeni www.knanosys.com Outline Long-range Interactions Ewald Sum Fast Multipole Method Spherically Truncated Coulombic Potential Speeding up Calculations SPaSM
Molecular Dynamics Simulations Dr. Kasra Momeni www.knanosys.com Outline Long-range Interactions Ewald Sum Fast Multipole Method Spherically Truncated Coulombic Potential Speeding up Calculations SPaSM
All-atom Molecular Mechanics. Trent E. Balius AMS 535 / CHE /27/2010
 All-atom Molecular Mechanics Trent E. Balius AMS 535 / CHE 535 09/27/2010 Outline Molecular models Molecular mechanics Force Fields Potential energy function functional form parameters and parameterization
All-atom Molecular Mechanics Trent E. Balius AMS 535 / CHE 535 09/27/2010 Outline Molecular models Molecular mechanics Force Fields Potential energy function functional form parameters and parameterization
Optimizing GROMACS for parallel performance
 Optimizing GROMACS for parallel performance Outline 1. Why optimize? Performance status quo 2. GROMACS as a black box. (PME) 3. How does GROMACS spend its time? (MPE) 4. What you can do What I want to
Optimizing GROMACS for parallel performance Outline 1. Why optimize? Performance status quo 2. GROMACS as a black box. (PME) 3. How does GROMACS spend its time? (MPE) 4. What you can do What I want to
Introduction to molecular dynamics
 1 Introduction to molecular dynamics Yves Lansac Université François Rabelais, Tours, France Visiting MSE, GIST for the summer Molecular Simulation 2 Molecular simulation is a computational experiment.
1 Introduction to molecular dynamics Yves Lansac Université François Rabelais, Tours, France Visiting MSE, GIST for the summer Molecular Simulation 2 Molecular simulation is a computational experiment.
Using Molecular Dynamics to Compute Properties CHEM 430
 Using Molecular Dynamics to Compute Properties CHEM 43 Heat Capacity and Energy Fluctuations Running an MD Simulation Equilibration Phase Before data-collection and results can be analyzed the system
Using Molecular Dynamics to Compute Properties CHEM 43 Heat Capacity and Energy Fluctuations Running an MD Simulation Equilibration Phase Before data-collection and results can be analyzed the system
A reciprocal space based method for treating long range interactions in ab initio and force-field-based calculations in clusters
 A reciprocal space based method for treating long range interactions in ab initio and force-field-based calculations in clusters Glenn J. Martyna and Mark E. Tuckerman Citation: The Journal of Chemical
A reciprocal space based method for treating long range interactions in ab initio and force-field-based calculations in clusters Glenn J. Martyna and Mark E. Tuckerman Citation: The Journal of Chemical
Gromacs Workshop Spring CSC
 Gromacs Workshop Spring 2007 @ CSC Erik Lindahl Center for Biomembrane Research Stockholm University, Sweden David van der Spoel Dept. Cell & Molecular Biology Uppsala University, Sweden Berk Hess Max-Planck-Institut
Gromacs Workshop Spring 2007 @ CSC Erik Lindahl Center for Biomembrane Research Stockholm University, Sweden David van der Spoel Dept. Cell & Molecular Biology Uppsala University, Sweden Berk Hess Max-Planck-Institut
N-body simulations. Phys 750 Lecture 10
 N-body simulations Phys 750 Lecture 10 N-body simulations Forward integration in time of strongly coupled classical particles or (rigid-body composite objects) dissipation Brownian motion Very general
N-body simulations Phys 750 Lecture 10 N-body simulations Forward integration in time of strongly coupled classical particles or (rigid-body composite objects) dissipation Brownian motion Very general
A Fast N-Body Solver for the Poisson(-Boltzmann) Equation
 A Fast N-Body Solver for the Poisson(-Boltzmann) Equation Robert D. Skeel Departments of Computer Science (and Mathematics) Purdue University http://bionum.cs.purdue.edu/2008december.pdf 1 Thesis Poisson(-Boltzmann)
A Fast N-Body Solver for the Poisson(-Boltzmann) Equation Robert D. Skeel Departments of Computer Science (and Mathematics) Purdue University http://bionum.cs.purdue.edu/2008december.pdf 1 Thesis Poisson(-Boltzmann)
Seminar in Particles Simulations. Particle Mesh Ewald. Theodora Konstantinidou
 Seminar in Particles Simulations Particle Mesh Ewald Theodora Konstantinidou This paper is part of the course work: seminar 2 Contents 1 Introduction 3 1.1 Role of Molecular Dynamics - MD.......................
Seminar in Particles Simulations Particle Mesh Ewald Theodora Konstantinidou This paper is part of the course work: seminar 2 Contents 1 Introduction 3 1.1 Role of Molecular Dynamics - MD.......................
Intermolecular Forces and Monte-Carlo Integration 열역학특수연구
 Intermolecular Forces and Monte-Carlo Integration 열역학특수연구 2003.3.28 Source of the lecture note. J.M.Prausnitz and others, Molecular Thermodynamics of Fluid Phase Equiliria Atkins, Physical Chemistry Lecture
Intermolecular Forces and Monte-Carlo Integration 열역학특수연구 2003.3.28 Source of the lecture note. J.M.Prausnitz and others, Molecular Thermodynamics of Fluid Phase Equiliria Atkins, Physical Chemistry Lecture
Free energy simulations
 Free energy simulations Marcus Elstner and Tomáš Kubař January 14, 2013 Motivation a physical quantity that is of most interest in chemistry? free energies Helmholtz F or Gibbs G holy grail of computational
Free energy simulations Marcus Elstner and Tomáš Kubař January 14, 2013 Motivation a physical quantity that is of most interest in chemistry? free energies Helmholtz F or Gibbs G holy grail of computational
Molecular mechanics. classical description of molecules. Marcus Elstner and Tomáš Kubař. April 29, 2016
 classical description of molecules April 29, 2016 Chemical bond Conceptual and chemical basis quantum effect solution of the SR numerically expensive (only small molecules can be treated) approximations
classical description of molecules April 29, 2016 Chemical bond Conceptual and chemical basis quantum effect solution of the SR numerically expensive (only small molecules can be treated) approximations
Developing Monovalent Ion Parameters for the Optimal Point Charge (OPC) Water Model. John Dood Hope College
 Developing Monovalent Ion Parameters for the Optimal Point Charge (OPC) Water Model John Dood Hope College What are MD simulations? Model and predict the structure and dynamics of large macromolecules.
Developing Monovalent Ion Parameters for the Optimal Point Charge (OPC) Water Model John Dood Hope College What are MD simulations? Model and predict the structure and dynamics of large macromolecules.
Molecular Modeling -- Lecture 15 Surfaces and electrostatics
 Molecular Modeling -- Lecture 15 Surfaces and electrostatics Molecular surfaces The Hydrophobic Effect Electrostatics Poisson-Boltzmann Equation Electrostatic maps Electrostatic surfaces in MOE 15.1 The
Molecular Modeling -- Lecture 15 Surfaces and electrostatics Molecular surfaces The Hydrophobic Effect Electrostatics Poisson-Boltzmann Equation Electrostatic maps Electrostatic surfaces in MOE 15.1 The
Multiphase Flow and Heat Transfer
 Multiphase Flow and Heat Transfer Liquid-Vapor Interface Sudheer Siddapuredddy sudheer@iitp.ac.in Department of Mechanical Engineering Indian Institution of Technology Patna Multiphase Flow and Heat Transfer
Multiphase Flow and Heat Transfer Liquid-Vapor Interface Sudheer Siddapuredddy sudheer@iitp.ac.in Department of Mechanical Engineering Indian Institution of Technology Patna Multiphase Flow and Heat Transfer
Coarse-Grained Models!
 Coarse-Grained Models! Large and complex molecules (e.g. long polymers) can not be simulated on the all-atom level! Requires coarse-graining of the model! Coarse-grained models are usually also particles
Coarse-Grained Models! Large and complex molecules (e.g. long polymers) can not be simulated on the all-atom level! Requires coarse-graining of the model! Coarse-grained models are usually also particles
Efficient Parallelization of Molecular Dynamics Simulations on Hybrid CPU/GPU Supercoputers
 Efficient Parallelization of Molecular Dynamics Simulations on Hybrid CPU/GPU Supercoputers Jaewoon Jung (RIKEN, RIKEN AICS) Yuji Sugita (RIKEN, RIKEN AICS, RIKEN QBiC, RIKEN ithes) Molecular Dynamics
Efficient Parallelization of Molecular Dynamics Simulations on Hybrid CPU/GPU Supercoputers Jaewoon Jung (RIKEN, RIKEN AICS) Yuji Sugita (RIKEN, RIKEN AICS, RIKEN QBiC, RIKEN ithes) Molecular Dynamics
Biomolecular modeling I
 2015, December 15 Biomolecular simulation Elementary body atom Each atom x, y, z coordinates A protein is a set of coordinates. (Gromacs, A. P. Heiner) Usually one molecule/complex of interest (e.g. protein,
2015, December 15 Biomolecular simulation Elementary body atom Each atom x, y, z coordinates A protein is a set of coordinates. (Gromacs, A. P. Heiner) Usually one molecule/complex of interest (e.g. protein,
Molecular Dynamics. A very brief introduction
 Molecular Dynamics A very brief introduction Sander Pronk Dept. of Theoretical Physics KTH Royal Institute of Technology & Science For Life Laboratory Stockholm, Sweden Why computer simulations? Two primary
Molecular Dynamics A very brief introduction Sander Pronk Dept. of Theoretical Physics KTH Royal Institute of Technology & Science For Life Laboratory Stockholm, Sweden Why computer simulations? Two primary
Lecture 11: Potential Energy Functions
 Lecture 11: Potential Energy Functions Dr. Ronald M. Levy ronlevy@temple.edu Originally contributed by Lauren Wickstrom (2011) Microscopic/Macroscopic Connection The connection between microscopic interactions
Lecture 11: Potential Energy Functions Dr. Ronald M. Levy ronlevy@temple.edu Originally contributed by Lauren Wickstrom (2011) Microscopic/Macroscopic Connection The connection between microscopic interactions
What is Classical Molecular Dynamics?
 What is Classical Molecular Dynamics? Simulation of explicit particles (atoms, ions,... ) Particles interact via relatively simple analytical potential functions Newton s equations of motion are integrated
What is Classical Molecular Dynamics? Simulation of explicit particles (atoms, ions,... ) Particles interact via relatively simple analytical potential functions Newton s equations of motion are integrated
CS 542G: The Poisson Problem, Finite Differences
 CS 542G: The Poisson Problem, Finite Differences Robert Bridson November 10, 2008 1 The Poisson Problem At the end last time, we noticed that the gravitational potential has a zero Laplacian except at
CS 542G: The Poisson Problem, Finite Differences Robert Bridson November 10, 2008 1 The Poisson Problem At the end last time, we noticed that the gravitational potential has a zero Laplacian except at
Welcome. to Electrostatics
 Welcome to Electrostatics Outline 1. Coulomb s Law 2. The Electric Field - Examples 3. Gauss Law - Examples 4. Conductors in Electric Field Coulomb s Law Coulomb s law quantifies the magnitude of the electrostatic
Welcome to Electrostatics Outline 1. Coulomb s Law 2. The Electric Field - Examples 3. Gauss Law - Examples 4. Conductors in Electric Field Coulomb s Law Coulomb s law quantifies the magnitude of the electrostatic
Partial differential equations
 Partial differential equations Many problems in science involve the evolution of quantities not only in time but also in space (this is the most common situation)! We will call partial differential equation
Partial differential equations Many problems in science involve the evolution of quantities not only in time but also in space (this is the most common situation)! We will call partial differential equation
Review: Power series define functions. Functions define power series. Taylor series of a function. Taylor polynomials of a function.
 Taylor Series (Sect. 10.8) Review: Power series define functions. Functions define power series. Taylor series of a function. Taylor polynomials of a function. Review: Power series define functions Remarks:
Taylor Series (Sect. 10.8) Review: Power series define functions. Functions define power series. Taylor series of a function. Taylor polynomials of a function. Review: Power series define functions Remarks:
Biomolecular modeling. Theoretical Chemistry, TU Braunschweig (Dated: December 10, 2010)
 Biomolecular modeling Marcus Elstner and Tomáš Kubař Theoretical Chemistry, TU Braunschweig (Dated: December 10, 2010) IX. FREE ENERGY SIMULATIONS When searching for a physical quantity that is of most
Biomolecular modeling Marcus Elstner and Tomáš Kubař Theoretical Chemistry, TU Braunschweig (Dated: December 10, 2010) IX. FREE ENERGY SIMULATIONS When searching for a physical quantity that is of most
1. Poisson-Boltzmann 1.1. Poisson equation. We consider the Laplacian. which is given in spherical coordinates by (2)
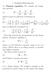 1. Poisson-Boltzmann 1.1. Poisson equation. We consider the Laplacian operator (1) 2 = 2 x + 2 2 y + 2 2 z 2 which is given in spherical coordinates by (2) 2 = 1 ( r 2 ) + 1 r 2 r r r 2 sin θ θ and in
1. Poisson-Boltzmann 1.1. Poisson equation. We consider the Laplacian operator (1) 2 = 2 x + 2 2 y + 2 2 z 2 which is given in spherical coordinates by (2) 2 = 1 ( r 2 ) + 1 r 2 r r r 2 sin θ θ and in
CS 273 Prof. Serafim Batzoglou Prof. Jean-Claude Latombe Spring Lecture 12 : Energy maintenance (1) Lecturer: Prof. J.C.
 CS 273 Prof. Serafim Batzoglou Prof. Jean-Claude Latombe Spring 2006 Lecture 12 : Energy maintenance (1) Lecturer: Prof. J.C. Latombe Scribe: Neda Nategh How do you update the energy function during the
CS 273 Prof. Serafim Batzoglou Prof. Jean-Claude Latombe Spring 2006 Lecture 12 : Energy maintenance (1) Lecturer: Prof. J.C. Latombe Scribe: Neda Nategh How do you update the energy function during the
Notes for Lecture 10
 February 2, 26 Notes for Lecture Introduction to grid-based methods for solving Poisson and Laplace Equations Finite difference methods The basis for most grid-based finite difference methods is the Taylor
February 2, 26 Notes for Lecture Introduction to grid-based methods for solving Poisson and Laplace Equations Finite difference methods The basis for most grid-based finite difference methods is the Taylor
Infinite series, improper integrals, and Taylor series
 Chapter 2 Infinite series, improper integrals, and Taylor series 2. Introduction to series In studying calculus, we have explored a variety of functions. Among the most basic are polynomials, i.e. functions
Chapter 2 Infinite series, improper integrals, and Taylor series 2. Introduction to series In studying calculus, we have explored a variety of functions. Among the most basic are polynomials, i.e. functions
Notes: Most of the material presented in this chapter is taken from Jackson, Chap. 2, 3, and 4, and Di Bartolo, Chap. 2. 2π nx i a. ( ) = G n.
 Chapter. Electrostatic II Notes: Most of the material presented in this chapter is taken from Jackson, Chap.,, and 4, and Di Bartolo, Chap... Mathematical Considerations.. The Fourier series and the Fourier
Chapter. Electrostatic II Notes: Most of the material presented in this chapter is taken from Jackson, Chap.,, and 4, and Di Bartolo, Chap... Mathematical Considerations.. The Fourier series and the Fourier
SYDE 112, LECTURE 7: Integration by Parts
 SYDE 112, LECTURE 7: Integration by Parts 1 Integration By Parts Consider trying to take the integral of xe x dx. We could try to find a substitution but would quickly grow frustrated there is no substitution
SYDE 112, LECTURE 7: Integration by Parts 1 Integration By Parts Consider trying to take the integral of xe x dx. We could try to find a substitution but would quickly grow frustrated there is no substitution
Supporting Information
 Supporting Information A: Calculation of radial distribution functions To get an effective propagator in one dimension, we first transform 1) into spherical coordinates: x a = ρ sin θ cos φ, y = ρ sin
Supporting Information A: Calculation of radial distribution functions To get an effective propagator in one dimension, we first transform 1) into spherical coordinates: x a = ρ sin θ cos φ, y = ρ sin
This is an electronic reprint of the original article. This reprint may differ from the original in pagination and typographic detail.
 This is an electronic reprint of the original article. This reprint may differ from the original in pagination and typographic detail. Author(s): Hub, Jochen S.; Groot, Bert L. De; Grubmüller, Helmut;
This is an electronic reprint of the original article. This reprint may differ from the original in pagination and typographic detail. Author(s): Hub, Jochen S.; Groot, Bert L. De; Grubmüller, Helmut;
UNIT I SOLID STATE PHYSICS
 UNIT I SOLID STATE PHYSICS CHAPTER 1 CRYSTAL STRUCTURE 1.1 INTRODUCTION When two atoms are brought together, two kinds of forces: attraction and repulsion come into play. The force of attraction increases
UNIT I SOLID STATE PHYSICS CHAPTER 1 CRYSTAL STRUCTURE 1.1 INTRODUCTION When two atoms are brought together, two kinds of forces: attraction and repulsion come into play. The force of attraction increases
Physics 200 Lecture 4. Integration. Lecture 4. Physics 200 Laboratory
 Physics 2 Lecture 4 Integration Lecture 4 Physics 2 Laboratory Monday, February 21st, 211 Integration is the flip-side of differentiation in fact, it is often possible to write a differential equation
Physics 2 Lecture 4 Integration Lecture 4 Physics 2 Laboratory Monday, February 21st, 211 Integration is the flip-side of differentiation in fact, it is often possible to write a differential equation
Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water?
 Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water? Ruhong Zhou 1 and Bruce J. Berne 2 1 IBM Thomas J. Watson Research Center; and 2 Department of Chemistry,
Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water? Ruhong Zhou 1 and Bruce J. Berne 2 1 IBM Thomas J. Watson Research Center; and 2 Department of Chemistry,
EECS 117 Lecture 7: Electrostatics Review
 EECS 117 Lecture 7: Electrostatics Review Prof. Niknejad University of California, Berkeley University of California, Berkeley EECS 117 Lecture 7 p. 1/19 Existence of Charge Charge, like mass, is an intrinsic
EECS 117 Lecture 7: Electrostatics Review Prof. Niknejad University of California, Berkeley University of California, Berkeley EECS 117 Lecture 7 p. 1/19 Existence of Charge Charge, like mass, is an intrinsic
Analysis of the simulation
 Analysis of the simulation Marcus Elstner and Tomáš Kubař January 7, 2014 Thermodynamic properties time averages of thermodynamic quantites correspond to ensemble averages (ergodic theorem) some quantities
Analysis of the simulation Marcus Elstner and Tomáš Kubař January 7, 2014 Thermodynamic properties time averages of thermodynamic quantites correspond to ensemble averages (ergodic theorem) some quantities
X-ray Crystallography
 2009/11/25 [ 1 ] X-ray Crystallography Andrew Torda, wintersemester 2009 / 2010 X-ray numerically most important more than 4/5 structures Goal a set of x, y, z coordinates different properties to NMR History
2009/11/25 [ 1 ] X-ray Crystallography Andrew Torda, wintersemester 2009 / 2010 X-ray numerically most important more than 4/5 structures Goal a set of x, y, z coordinates different properties to NMR History
This semester. Books
 Models mostly proteins from detailed to more abstract models Some simulation methods This semester Books None necessary for my group and Prof Rarey Molecular Modelling: Principles and Applications Leach,
Models mostly proteins from detailed to more abstract models Some simulation methods This semester Books None necessary for my group and Prof Rarey Molecular Modelling: Principles and Applications Leach,
1. Hydrogen atom in a box
 1. Hydrogen atom in a box Recall H atom problem, V(r) = -1/r e r exact answer solved by expanding in Gaussian basis set, had to solve secular matrix involving matrix elements of basis functions place atom
1. Hydrogen atom in a box Recall H atom problem, V(r) = -1/r e r exact answer solved by expanding in Gaussian basis set, had to solve secular matrix involving matrix elements of basis functions place atom
Time scales. Notes. Mixed Implicit/Explicit. Newmark Methods [ ] [ ] "!t = O 1 %
![Time scales. Notes. Mixed Implicit/Explicit. Newmark Methods [ ] [ ] !t = O 1 % Time scales. Notes. Mixed Implicit/Explicit. Newmark Methods [ ] [ ] !t = O 1 %](/thumbs/95/124165137.jpg) Notes Time scales! For using Pixie (the renderer) make sure you type use pixie first! Assignment 1 questions?! [work out]! For position dependence, characteristic time interval is "!t = O 1 % $ ' # K &!
Notes Time scales! For using Pixie (the renderer) make sure you type use pixie first! Assignment 1 questions?! [work out]! For position dependence, characteristic time interval is "!t = O 1 % $ ' # K &!
Chapter 24. Gauss s Law
 Chapter 24 Gauss s Law Let s return to the field lines and consider the flux through a surface. The number of lines per unit area is proportional to the magnitude of the electric field. This means that
Chapter 24 Gauss s Law Let s return to the field lines and consider the flux through a surface. The number of lines per unit area is proportional to the magnitude of the electric field. This means that
Plasmonics: elementary excitation of a plasma (gas of free charges) nano-scale optics done with plasmons at metal interfaces
 Plasmonics Plasmon: Plasmonics: elementary excitation of a plasma (gas of free charges) nano-scale optics done with plasmons at metal interfaces Femius Koenderink Center for Nanophotonics AMOLF, Amsterdam
Plasmonics Plasmon: Plasmonics: elementary excitation of a plasma (gas of free charges) nano-scale optics done with plasmons at metal interfaces Femius Koenderink Center for Nanophotonics AMOLF, Amsterdam
Fast Ewald Summation based on NFFT with Mixed Periodicity
 Fast Ewald Summation based on NFFT with Mixed Periodicity Franziska Nestler, Michael Pippig and Daniel Potts In this paper we develop new fast Fourier-based methods for the Coulomb problem. We combine
Fast Ewald Summation based on NFFT with Mixed Periodicity Franziska Nestler, Michael Pippig and Daniel Potts In this paper we develop new fast Fourier-based methods for the Coulomb problem. We combine
An introduction to Molecular Dynamics. EMBO, June 2016
 An introduction to Molecular Dynamics EMBO, June 2016 What is MD? everything that living things do can be understood in terms of the jiggling and wiggling of atoms. The Feynman Lectures in Physics vol.
An introduction to Molecular Dynamics EMBO, June 2016 What is MD? everything that living things do can be understood in terms of the jiggling and wiggling of atoms. The Feynman Lectures in Physics vol.
Multiple time step Monte Carlo simulations: Application to charged systems with Ewald summation
 JOURNAL OF CHEMICAL PHYSICS VOLUME 11, NUMBER 1 1 JULY 004 Multiple time step Monte Carlo simulations: Application to charged systems with Ewald summation Katarzyna Bernacki a) Department of Chemistry
JOURNAL OF CHEMICAL PHYSICS VOLUME 11, NUMBER 1 1 JULY 004 Multiple time step Monte Carlo simulations: Application to charged systems with Ewald summation Katarzyna Bernacki a) Department of Chemistry
The Fourier spectral method (Amath Bretherton)
 The Fourier spectral method (Amath 585 - Bretherton) 1 Introduction The Fourier spectral method (or more precisely, pseudospectral method) is a very accurate way to solve BVPs with smooth solutions on
The Fourier spectral method (Amath 585 - Bretherton) 1 Introduction The Fourier spectral method (or more precisely, pseudospectral method) is a very accurate way to solve BVPs with smooth solutions on
Applications of Molecular Dynamics
 June 4, 0 Molecular Modeling and Simulation Applications of Molecular Dynamics Agricultural Bioinformatics Research Unit, Graduate School of Agricultural and Life Sciences, The University of Tokyo Tohru
June 4, 0 Molecular Modeling and Simulation Applications of Molecular Dynamics Agricultural Bioinformatics Research Unit, Graduate School of Agricultural and Life Sciences, The University of Tokyo Tohru
3 Chapter. Gauss s Law
 3 Chapter Gauss s Law 3.1 Electric Flux... 3-2 3.2 Gauss s Law (see also Gauss s Law Simulation in Section 3.10)... 3-4 Example 3.1: Infinitely Long Rod of Uniform Charge Density... 3-9 Example 3.2: Infinite
3 Chapter Gauss s Law 3.1 Electric Flux... 3-2 3.2 Gauss s Law (see also Gauss s Law Simulation in Section 3.10)... 3-4 Example 3.1: Infinitely Long Rod of Uniform Charge Density... 3-9 Example 3.2: Infinite
Salmon: Lectures on partial differential equations
 6. The wave equation Of the 3 basic equations derived in the previous section, we have already discussed the heat equation, (1) θ t = κθ xx + Q( x,t). In this section we discuss the wave equation, () θ
6. The wave equation Of the 3 basic equations derived in the previous section, we have already discussed the heat equation, (1) θ t = κθ xx + Q( x,t). In this section we discuss the wave equation, () θ
 A Nobel Prize for Molecular Dynamics and QM/MM What is Classical Molecular Dynamics? Simulation of explicit particles (atoms, ions,... ) Particles interact via relatively simple analytical potential
A Nobel Prize for Molecular Dynamics and QM/MM What is Classical Molecular Dynamics? Simulation of explicit particles (atoms, ions,... ) Particles interact via relatively simple analytical potential
INTRODUCTION. Introduction. Discrete charges: Electric dipole. Continuous charge distributions. Flux of a vector field
 Chapter 2 THE ELECTRC FELD ntroduction Discrete charges: Electric dipole Continuous charge distributions Flux of a vector field Flux of an electric field for a spherical Gaussian surface concentric to
Chapter 2 THE ELECTRC FELD ntroduction Discrete charges: Electric dipole Continuous charge distributions Flux of a vector field Flux of an electric field for a spherical Gaussian surface concentric to
Conductors and Insulators
 Conductors and Insulators Lecture 11: Electromagnetic Theory Professor D. K. Ghosh, Physics Department, I.I.T., Bombay Self Energy of a Charge Distribution : In Lecture 1 we briefly discussed what we called
Conductors and Insulators Lecture 11: Electromagnetic Theory Professor D. K. Ghosh, Physics Department, I.I.T., Bombay Self Energy of a Charge Distribution : In Lecture 1 we briefly discussed what we called
Biomolecular modeling I
 2016, December 6 Biomolecular structure Structural elements of life Biomolecules proteins, nucleic acids, lipids, carbohydrates... Biomolecular structure Biomolecules biomolecular complexes aggregates...
2016, December 6 Biomolecular structure Structural elements of life Biomolecules proteins, nucleic acids, lipids, carbohydrates... Biomolecular structure Biomolecules biomolecular complexes aggregates...
Physics 2B. Lecture 24B. Gauss 10 Deutsche Mark
 Physics 2B Lecture 24B Gauss 10 Deutsche Mark Electric Flux Flux is the amount of something that flows through a given area. Electric flux, Φ E, measures the amount of electric field lines that passes
Physics 2B Lecture 24B Gauss 10 Deutsche Mark Electric Flux Flux is the amount of something that flows through a given area. Electric flux, Φ E, measures the amount of electric field lines that passes
Parallelization of Molecular Dynamics (with focus on Gromacs) SeSE 2014 p.1/29
 Parallelization of Molecular Dynamics (with focus on Gromacs) SeSE 2014 p.1/29 Outline A few words on MD applications and the GROMACS package The main work in an MD simulation Parallelization Stream computing
Parallelization of Molecular Dynamics (with focus on Gromacs) SeSE 2014 p.1/29 Outline A few words on MD applications and the GROMACS package The main work in an MD simulation Parallelization Stream computing
Bioengineering 215. An Introduction to Molecular Dynamics for Biomolecules
 Bioengineering 215 An Introduction to Molecular Dynamics for Biomolecules David Parker May 18, 2007 ntroduction A principal tool to study biological molecules is molecular dynamics simulations (MD). MD
Bioengineering 215 An Introduction to Molecular Dynamics for Biomolecules David Parker May 18, 2007 ntroduction A principal tool to study biological molecules is molecular dynamics simulations (MD). MD
1. Overview of the relations among charge, field and potential Gauss law Integrate charge to get potential More about energy Laplace and Poisson
 1. Overview of the relations among charge, field and potential Gauss law Integrate charge to get potential More about energy Laplace and Poisson equations 2. Intro to conductors Field inside is zero BEFORE
1. Overview of the relations among charge, field and potential Gauss law Integrate charge to get potential More about energy Laplace and Poisson equations 2. Intro to conductors Field inside is zero BEFORE
The Hydrogen Atom. Dr. Sabry El-Taher 1. e 4. U U r
 The Hydrogen Atom Atom is a 3D object, and the electron motion is three-dimensional. We ll start with the simplest case - The hydrogen atom. An electron and a proton (nucleus) are bound by the central-symmetric
The Hydrogen Atom Atom is a 3D object, and the electron motion is three-dimensional. We ll start with the simplest case - The hydrogen atom. An electron and a proton (nucleus) are bound by the central-symmetric
Lecture 17 - The Secrets we have Swept Under the Rug
 1.0 0.5 0.0-0.5-0.5 0.0 0.5 1.0 Lecture 17 - The Secrets we have Swept Under the Rug A Puzzle... What makes 3D Special? Example (1D charge distribution) A stick with uniform charge density λ lies between
1.0 0.5 0.0-0.5-0.5 0.0 0.5 1.0 Lecture 17 - The Secrets we have Swept Under the Rug A Puzzle... What makes 3D Special? Example (1D charge distribution) A stick with uniform charge density λ lies between
Homework 4: Hard-Copy Homework Due Wednesday 2/17
 Homework 4: Hard-Copy Homework Due Wednesday 2/17 Special instructions for this homework: Please show all work necessary to solve the problems, including diagrams, algebra, calculus, or whatever else may
Homework 4: Hard-Copy Homework Due Wednesday 2/17 Special instructions for this homework: Please show all work necessary to solve the problems, including diagrams, algebra, calculus, or whatever else may
2 Structure. 2.1 Coulomb interactions
 2 Structure 2.1 Coulomb interactions While the information needed for reproduction of living systems is chiefly maintained in the sequence of macromolecules, any practical use of this information must
2 Structure 2.1 Coulomb interactions While the information needed for reproduction of living systems is chiefly maintained in the sequence of macromolecules, any practical use of this information must
RWTH Aachen University
 IPCC @ RWTH Aachen University Optimization of multibody and long-range solvers in LAMMPS Rodrigo Canales William McDoniel Markus Höhnerbach Ahmed E. Ismail Paolo Bientinesi IPCC Showcase November 2016
IPCC @ RWTH Aachen University Optimization of multibody and long-range solvers in LAMMPS Rodrigo Canales William McDoniel Markus Höhnerbach Ahmed E. Ismail Paolo Bientinesi IPCC Showcase November 2016
ONE AND MANY ELECTRON ATOMS Chapter 15
 See Week 8 lecture notes. This is exactly the same as the Hamiltonian for nonrigid rotation. In Week 8 lecture notes it was shown that this is the operator for Lˆ 2, the square of the angular momentum.
See Week 8 lecture notes. This is exactly the same as the Hamiltonian for nonrigid rotation. In Week 8 lecture notes it was shown that this is the operator for Lˆ 2, the square of the angular momentum.
PHY102 Electricity Topic 3 (Lectures 4 & 5) Gauss s Law
 PHY1 Electricity Topic 3 (Lectures 4 & 5) Gauss s Law In this topic, we will cover: 1) Electric Flux ) Gauss s Law, relating flux to enclosed charge 3) Electric Fields and Conductors revisited Reading
PHY1 Electricity Topic 3 (Lectures 4 & 5) Gauss s Law In this topic, we will cover: 1) Electric Flux ) Gauss s Law, relating flux to enclosed charge 3) Electric Fields and Conductors revisited Reading
n 1 f n 1 c 1 n+1 = c 1 n $ c 1 n 1. After taking logs, this becomes
 Root finding: 1 a The points {x n+1, }, {x n, f n }, {x n 1, f n 1 } should be co-linear Say they lie on the line x + y = This gives the relations x n+1 + = x n +f n = x n 1 +f n 1 = Eliminating α and
Root finding: 1 a The points {x n+1, }, {x n, f n }, {x n 1, f n 1 } should be co-linear Say they lie on the line x + y = This gives the relations x n+1 + = x n +f n = x n 1 +f n 1 = Eliminating α and
University of Illinois at Chicago Department of Physics. Electricity & Magnetism Qualifying Examination
 University of Illinois at Chicago Department of Physics Electricity & Magnetism Qualifying Examination January 7, 28 9. am 12: pm Full credit can be achieved from completely correct answers to 4 questions.
University of Illinois at Chicago Department of Physics Electricity & Magnetism Qualifying Examination January 7, 28 9. am 12: pm Full credit can be achieved from completely correct answers to 4 questions.
Castep & Real Space Integrals
 Castep & Real Space Integrals MJR March 3, 213 1 Real and Reciprocal Representations The wavefunction in Castep is expressed in a plane wave basis. That the system is periodic means that the permitted
Castep & Real Space Integrals MJR March 3, 213 1 Real and Reciprocal Representations The wavefunction in Castep is expressed in a plane wave basis. That the system is periodic means that the permitted
Molecular dynamics simulation of Aquaporin-1. 4 nm
 Molecular dynamics simulation of Aquaporin-1 4 nm Molecular Dynamics Simulations Schrödinger equation i~@ t (r, R) =H (r, R) Born-Oppenheimer approximation H e e(r; R) =E e (R) e(r; R) Nucleic motion described
Molecular dynamics simulation of Aquaporin-1 4 nm Molecular Dynamics Simulations Schrödinger equation i~@ t (r, R) =H (r, R) Born-Oppenheimer approximation H e e(r; R) =E e (R) e(r; R) Nucleic motion described
Electronic Structure Theory for Periodic Systems: The Concepts. Christian Ratsch
 Electronic Structure Theory for Periodic Systems: The Concepts Christian Ratsch Institute for Pure and Applied Mathematics and Department of Mathematics, UCLA Motivation There are 10 20 atoms in 1 mm 3
Electronic Structure Theory for Periodic Systems: The Concepts Christian Ratsch Institute for Pure and Applied Mathematics and Department of Mathematics, UCLA Motivation There are 10 20 atoms in 1 mm 3
CE 530 Molecular Simulation
 1 CE 530 Molecular Simulation Lecture 14 Molecular Models David A. Kofke Department of Chemical Engineering SUNY Buffalo kofke@eng.buffalo.edu 2 Review Monte Carlo ensemble averaging, no dynamics easy
1 CE 530 Molecular Simulation Lecture 14 Molecular Models David A. Kofke Department of Chemical Engineering SUNY Buffalo kofke@eng.buffalo.edu 2 Review Monte Carlo ensemble averaging, no dynamics easy
The generalized Born model: its foundation, applications, and limitations
 The generalized Born model: its foundation, applications, and limitations Alexey Onufriev September 8, 2010 Abstract The generalized Born (GB) approximation is introduced in the context of the implicit
The generalized Born model: its foundation, applications, and limitations Alexey Onufriev September 8, 2010 Abstract The generalized Born (GB) approximation is introduced in the context of the implicit
Multiscale Modelling, taking into account Collisions
 Multiscale Modelling, taking into account Collisions Andreas Adelmann (Paul Scherrer Institut) October 28, 2017 Multiscale Modelling, taking into account Collisions October 28, 2017 Page 1 / 22 1 Motivation
Multiscale Modelling, taking into account Collisions Andreas Adelmann (Paul Scherrer Institut) October 28, 2017 Multiscale Modelling, taking into account Collisions October 28, 2017 Page 1 / 22 1 Motivation
Multiple time step Monte Carlo
 JOURNAL OF CHEMICAL PHYSICS VOLUME 117, NUMBER 18 8 NOVEMBER 2002 Multiple time step Monte Carlo Balázs Hetényi a) Department of Chemistry, Princeton University, Princeton, NJ 08544 and Department of Chemistry
JOURNAL OF CHEMICAL PHYSICS VOLUME 117, NUMBER 18 8 NOVEMBER 2002 Multiple time step Monte Carlo Balázs Hetényi a) Department of Chemistry, Princeton University, Princeton, NJ 08544 and Department of Chemistry
Classical Electrostatics for Biomolecular Simulations
 pubs.acs.org/cr Classical Electrostatics for Biomolecular Simulations G. Andreś Cisneros, Mikko Karttunen, Pengyu Ren, and Celeste Sagui*, Department of Chemistry, Wayne State University, Detroit, Michigan
pubs.acs.org/cr Classical Electrostatics for Biomolecular Simulations G. Andreś Cisneros, Mikko Karttunen, Pengyu Ren, and Celeste Sagui*, Department of Chemistry, Wayne State University, Detroit, Michigan
Integrals in Electrostatic Problems
 PHYS 119 Integrals in Electrostatic Problems Josh McKenney University of North Carolina at Chapel Hill (Dated: January 6, 2016) 1 FIG. 1. Three positive charges positioned at equal distances around an
PHYS 119 Integrals in Electrostatic Problems Josh McKenney University of North Carolina at Chapel Hill (Dated: January 6, 2016) 1 FIG. 1. Three positive charges positioned at equal distances around an
13 - ELECTROSTATICS Page 1 ( Answers at the end of all questions )
 3 - ELECTROSTATICS Page ) Two point charges 8 and - are located at x = 0 and x = L respectively. The location of a point on the x axis at which the net electric field due to these two point charges is
3 - ELECTROSTATICS Page ) Two point charges 8 and - are located at x = 0 and x = L respectively. The location of a point on the x axis at which the net electric field due to these two point charges is
Smoothed Particle Hydrodynamics (SPH) 4. May 2012
 Smoothed Particle Hydrodynamics (SPH) 4. May 2012 Calculating density SPH density estimator Weighted summation over nearby particles: ρ(r) = N neigh b=1 m bw (r r b, h) W weight function with dimension
Smoothed Particle Hydrodynamics (SPH) 4. May 2012 Calculating density SPH density estimator Weighted summation over nearby particles: ρ(r) = N neigh b=1 m bw (r r b, h) W weight function with dimension
Where, ε 0 = Permittivity of free space and = Nm 2 C 2 Therefore, force
 Exercises Question.: What is the force between two small charged spheres having charges of 2 0 7 C and 3 0 7 C placed 30 cm apart in air? Answer.: Repulsive force of magnitude 6 0 3 N Charge on the first
Exercises Question.: What is the force between two small charged spheres having charges of 2 0 7 C and 3 0 7 C placed 30 cm apart in air? Answer.: Repulsive force of magnitude 6 0 3 N Charge on the first
Plasmas as fluids. S.M.Lea. January 2007
 Plasmas as fluids S.M.Lea January 2007 So far we have considered a plasma as a set of non intereacting particles, each following its own path in the electric and magnetic fields. Now we want to consider
Plasmas as fluids S.M.Lea January 2007 So far we have considered a plasma as a set of non intereacting particles, each following its own path in the electric and magnetic fields. Now we want to consider
Geometry of Crystal Lattice
 0 Geometry of Crystal Lattice 0.1 Translational Symmetry The crystalline state of substances is different from other states (gaseous, liquid, amorphous) in that the atoms are in an ordered and symmetrical
0 Geometry of Crystal Lattice 0.1 Translational Symmetry The crystalline state of substances is different from other states (gaseous, liquid, amorphous) in that the atoms are in an ordered and symmetrical
Efficient multiple time step method for use with Ewald and particle mesh Ewald for large biomolecular systems
 JOURNAL OF CHEMICAL PHYSICS VOLUME 115, NUMBER 5 1 AUGUST 2001 Efficient multiple time step method for use with Ewald and particle mesh Ewald for large biomolecular systems Ruhong Zhou IBM Thomas J. Watson
JOURNAL OF CHEMICAL PHYSICS VOLUME 115, NUMBER 5 1 AUGUST 2001 Efficient multiple time step method for use with Ewald and particle mesh Ewald for large biomolecular systems Ruhong Zhou IBM Thomas J. Watson
