Available online at
|
|
|
- Madeleine Barnett
- 5 years ago
- Views:
Transcription
1 Available online at Indian Journal of Advances in Chemical Science 4(1) (2016) Theoretical Approach on Structural Aspects of a Potent, Selective, Orally Bioavailable Hedgehog Antagonist, 2-chloro-N-[4-chloro-3-(pyridin-2-yl)phenyl]- 4-(methylsulfonyl)benzamide Amaku Friday James*, Otuokere Ifeanyi Edozie Department of Chemistry, Michael Okpara University of Agriculture, Umudike, Nigeria. Received 16 th September 2015; Revised 22 nd September 2015; Accepted 24 th October ABSTRACT The molecular mechanics potential energy function were evaluated in terms of energies associated with bonded interactions (bond length, bond angle, and dihedral angle) as well as non-bonded interactions (Vander Waals and electrostatic). Surfaces were created to visualize excited state properties such as highest occupied molecular orbitals, lowest unoccupied molecular orbitals, and electrostatic potential mapped density. The steric energy for 2-chloro-N-[4-chloro-3-(pyridin-2-yl)phenyl]-4-(methylsulfonyl)benzamide (vismodegib) was calculated to be au ( kcal/mol). The most energetically favorable conformation of vismodegib was found to have a heat of formation of kcal/mol. The self-consistent field energy was calculated by geometry convergence function using RHF/AM1 method with a net charge of 0 and valence electron of 124, in ArgusLab software. The most feasible position for vismodegib to act as a highly potent and totally selective orally bioavailable Hedgehog antagonist was found to be au ( kcal mol 1 ). Key words: Vismodegib, Molecular mechanics, ArgusLab software. Indian Journal of Advances in Chemical Science 1. INTRODUCTION 2-chloro-N-[4-chloro-3-(pyridin-2-yl)phenyl]-4- (methylsulfonyl)benzamide (vismodegib) is a potent, selective, orally bioavailable Hedgehog antagonist, which shows remarkable activity against basal cell carcinoma that has metastasized to other parts of the body, relapsed after surgery, or cannot be treated with surgery or radiation [1]. It therapeutic and preventive efficacy is of great value, this is as a result of factors contributing to its exceptionally rapid progress from first-in-human testing to regulatory approval include an acceptable toxicity profile, early recognition of achievement of a biologically effective concentration in the first dose cohort explored, and an early-phase development plan that focused on a disease strongly dependent on the pathway being targeted [2-4]. The substance acts as a cyclopamine-competitive antagonist of the smoothened (SMO) receptor, which is part of the Hedgehog signaling pathway. SMO inhibition causes the transcription factors GLI1 and GLI2 to remain inactive, which prevents the expression of tumor mediating genes within the Hedgehog pathway [5]. This pathway is pathogenetically relevant in more than 90% of basal-cell carcinomas and can be retarded *Corresponding Author: ifeanyiotuokere@gmail.com Tel.: by making some transcription factor to remain inactive [6]. A molecule is considered as a collection of atoms held together by classical forces. These forces are described by potential energy function of structural features such as bond lengths, bond angles, and torsion angles. The energy (E) of the molecule is calculated as a sum of terms as in Equation (1). E=E stretching+e bending+e torsion+e Vander Waals+E electrostatic+e hydrogen bond+cross term These terms are of importance for the accurate calculation of geometric properties of molecules. The set of energy functions and the corresponding parameters are called a force field and can be generated using ArgusLab. ArgusLab is the electronic structure program that is based on the quantum mechanics; it predicts the potential energies, molecular structures; geometry optimization of structure, vibration frequencies of coordinates of atoms, bond length, and bond angle. Local charges such as Mulliken charges and zero differential overlaps (ZDO) charges can also be generated from ArgusLab using the AM1 31
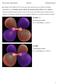
2 Table 1: Atomic coordinates of vismodegib. Figure 1: Prospective view of vismodegib by ACD/ Chemsketch. Figure 2: Self-consistent field energy of graph of vismodegib. a c e Figure 3: (a) Electron density cloud, (b) active conformation, (c) highest occupied, (d) lowest unoccupied molecular orbitals, (e) electrostatic potential mapped density vismodegib. parameterized method. In the ZDO approximation, the product of two deferent atomic orbitals is set to zero. The integral which survives the ZDO approximation was partly computed using the uniform charge sphere and the rest parameterized. The result produced is the integrated form of Hückel theory which takes into account electron repulsion [7]. Mulliken charges arise from the Mulliken population analysis and provide a means of estimating partial atomic charges b d Atom X Y Z C C C C C C C S O O C Cl N O C C C C C C Cl C N C C C C from calculations carried out by the methods of computational chemistry, particularly those based on the linear combination of atomic orbitals molecular orbital method, and are routinely used as variables in linear regression (quantitative structure-activity relationship) procedures [8,9]. We hereby present the computational study of the geometry optimization and excited state properties of vismodegib by ArgusLab software. 2. EXPERIMENTAL Vismodegib structure was sketched with ACD Lab Chem Sketch software and saved as MDL molfiles (*mol) [10]. The vismodegib structure was generated by ArgusLab, and minimization was performed with universal force field molecular mechanics method [11,12]. The minimum potential energy was calculated using geometry convergence function in ArgusLab software. Surfaces created to visualize ground state properties as well as excited 32
3 Table 2: Bond length of vismodegib. Atom Bond length C1 C C1 C C2 C C3 C C3 C C4 C C4 S C5 C C5 Cl C7 N C7 O S8 O S8 O S8 C N13 C C15 C C15 C C16 C C17 C C18 C C19 C C19 C C20 Cl C22 N C22 C N23 C C24 C C25 C C26 C state properties such as orbital, electron densities, electrostatic potentials (ESP), spin densities, and generated the grid data were used to make molecular orbital surfaces and electro static potential mapped on electron density surface [13-15]. The minimum potential energy was calculated for vismodegib through the geometry convergence map [16]. Mulliken atomic charges, ZDO atomic charges of vismodegib, and ground state dipole (debye) of vismodegib were determined using AM1 method. 3. RESULTS AND DISCUSSION Prospective view and calculated properties of vismodegib molecule are shown in Figure 1. Selfconsistent field (SCF) energy curve is shown in Figure 2. The active conformation, electron density cloud, highest occupied molecular orbital (HOMO), lowest unoccupied molecular orbital (LUMO), and Table 3: Bond angles of vismodegib. Atoms Bond angles Alternate angles C2 C1 C C1 C2 C C1 C3 C C1 C3 C C2 C4 C C2 C4 S C5 C3 C C3 C5 C C3 C5 Cl C3 C7 N C3 C7 O C6 C4 S C4 C6 C C4 S8 O C4 S8 O C4 S8 C C6 C5 Cl N13 C7 O C7 N13 C O9 S8 O O9 S8 C O10 S8 C N13 C15 C N13 C15 C C16 C15 C C15 C16 C C15 C17 C C16 C18 C C17 C19 C C17 C19 C C18 C20 C C18 C20 Cl C20 C19 C C19 C20 Cl C19 C22 N C19 C22 C N23 C22 C C22 N23 C C22 C24 C N23 C25 C C24 C26 C C25 C27 C ESP mapped density of vismodegib are shown in Figure 3. 33
4 Table 4: List of Mulliken atomic charges and ZDO atomic charges of vismodegib. Atoms ZDO atomic charges Mulliken atomic charges C C C C C C C S O O L N O C C C C C C Cl C N C ZDO=Zero differential overlap Table 5: Ground state dipole (debye) of vismodegib. X Y Z Length Table 6: Final energy evaluation. Force field Energy components (au) MM bond (Estr) MM angle (Ebend)+(Estrbend) MM dihedral (Etor) MM ImpTor (Eoop) MM vdw (EVdW) MM coulomb (Eqq) Total MM=Molecular mechanics au ( kcal/mol) The color map shows the ESP energy (in hartrees) for the various colors. The red end of the spectrum shows regions of highest stability for a positive test charge, magenta/blue show the regions of least stability for a positive test charge. The HOMO of the molecule and the LUMO, respectively, rendered in opaque forms. The positive and negative phases of the orbital are represented by two colors; the blue regions represent an increase in electron density, and the red regions show a decrease in electron density. This type of surface representations is useful to discuss drug receptor interaction. Atomic coordinates of molecule are given in Table 1 and bond length and bond angles are given in Tables 2 and 3, respectively, which are calculated after geometry optimization of the molecule from ArgusLab by using molecular mechanics methods. Tables 4-6 show Mulliken atomic charges, ZDO atomic charges of ground state dipole (debye), and calculated the steric energy of vismodegib molecule. The steric energy calculated for vismodegib molecule was au ( kcal/mol), and its SCF energy was found to be au ( kcal/mol) as calculated by RHF/AM1 method, by ArgusLab suite. 4. CONCLUSION The present work indicates that the best conformation of vismodegib is found to be au ( kcal/mol) which is the minimum potential energy by using ArgusLab software. At this point, vismodegib will be more active as a chemotherapy agent. 5. REFERENCES 1. J. M. Ng, T. Curran, (2011) The Hedgehog s tale: Developing strategies for targeting cancer, Nature Reviews Cancer, 11(7): J. Rodon Ahnert, J. Baselga, H. A. Tawbi, Y. Shou, C. Granvil, J. Dey, (2010) A phase I dose-escalation study of LDE225 a smoothened (SMO) antagonist, in patients with advanced solid tumors, Journal of Clinical Oncology, 28: L. L. Siu, K. Papadopoulos, S. R. Alberts, R. Kirchoff-Ross, B. Vakkalagadda, L. Lang, (2010) A first-in-human, phase I study of an oral Hedgehog pathway antagonist, BMS (XL139), in subjects with advanced or metastatic solid tumors, Journal of Clinical Oncology, 29: C. M. Rudin, A. Jimeno, W. H. Miller, B. J. Eigl, S. N. Gettinger, A. L. S. Chang, (2011) A phase I study of IPI-926, a novel Hedgehog pathway inhibitor, in patients with advanced or metastatic solid tumors, Journal of Clinical Oncology, 29:
5 5. H. Hahn, C. Wicking, P. G. Zaphiropoulous, M. R. Gailani, S. Shanley, A. Chidambaram, (1996) Mutations of the human homolog of drosophila patched in the nevoid basal cell carcinoma syndrome, Cell, 85: R. L, Johnson, A. L. Rothman, J. Xie, L. V. Goodrich, J. W. Bare, J. M. Bonifas, (1996) Human homolog of patched, a candidate gene for the basal cell nevus syndrome, Science, 272: R. S. Mulliken, (1955) Electronic population analysis on LCAO-MO molecular wave functions, The Journal of Chemical Physics, 23(10): A. R. Leach. (2001) Molecular Modelling: Principles and Applications, Englewood Cliffs, NJ: Prentice Hall, p S. William, P. E. Ohlinger, B. J. Klunzinger, J. H. Warren, (2009) Efficient calculation of heats of formation, The Journal of Physical Chemistry, 113(10): A. T. Mark. (2003) Seattle, WA: Planaria Software LLC. Available from: [Last accessed on 2014 Oct 09]. 11. I. I. I. Dunn, A. J. K. Hopfinger, (1998) Drug Discovery, London: Kluwer Academic Publisher, p G. Cruciani, S. Clementi, M. Pastor, (1998) GOLPE guided region selection, Perspectives in Drug Discovery and Design, 12-14(16): J. Simons, P. Jorgensen, H. Taylor, J. Ozment, (1983) Walking on potential energy surfaces, The Journal of Physical Chemistry, 87: I. G. Csizmadia, R. D. Enriz, (2001) Peptide and protein folding, Journal of Molecular Structure: Theochem, 543: I. E. Otuokere, C. O. Alisa, (2014) Molecular mechanics steric energy evaluation of reversible acetyl cholinesterase inhibitor, donepezil, (RS)-2- ([1-benzyl-4-piperidyl] methyl)-5,6-dimethoxy- 2,3-dihydroinden-1-one, Indian Journal of Advances in Chemical Science, 3: Y. C. Martin, (1998) Perspective in Drug Discovery and Design, USA: Springer Publisher, p3-23. *Bibliographical Sketch Amaku Friday James is a specialist in Physical Chemistry, with bias in adsorption and computational chemistry. He has published extensively in the area of adsorption and computational chemistry. He is a lecturer in the department of Chemistry, Michael Okpara University of Agriculture, Umudike, Nigeria. He is a member of the Chemical Society of Nigeria. 35
Pelagia Research Library
 Available online at www.pelagiaresearchlibrary.com Der Chemica Sinica, 2015, 6(11):1-6 ISSN: 0976-8505 CODEN (USA) CSHIA5 Computational study on the geometry optimization and excited-state properties triamterene
Available online at www.pelagiaresearchlibrary.com Der Chemica Sinica, 2015, 6(11):1-6 ISSN: 0976-8505 CODEN (USA) CSHIA5 Computational study on the geometry optimization and excited-state properties triamterene
Theoretical Approach on structural aspects of antiepileptic agent indoline-2,3- dione-3-oxime by arguslab 4 software
 Available online at www.scientiaresearchlibrary.com Scientia Research Library ISSN 2348-0408 USA CODEN: JACOGN Journal of Applied Chemistry, 2014, 2 (1):92-101 (http://www.scientiaresearchlibrary.com/arhcive.php)
Available online at www.scientiaresearchlibrary.com Scientia Research Library ISSN 2348-0408 USA CODEN: JACOGN Journal of Applied Chemistry, 2014, 2 (1):92-101 (http://www.scientiaresearchlibrary.com/arhcive.php)
CONFORMATIONAL ANALYSIS (GEOMETRY OPTIMIZATION) OF NUCLEOSIDIC ANTITUMOR ANTIBIOTIC SHOWDOMYCIN BY ARGUSLAB 4 SOFTWARE
 CONFORMATIONAL ANALYSIS (GEOMETRY OPTIMIZATION) OF NUCLEOSIDIC ANTITUMOR ANTIBIOTIC SHOWDOMYCIN BY ARGUSLAB 4 SOFTWARE AFSHAN NAZ, KHALIDA BANO, FARHAT BANO, NAJAF ABBAS GHAFOOR AND NAHEED AKHTAR Biophysics
CONFORMATIONAL ANALYSIS (GEOMETRY OPTIMIZATION) OF NUCLEOSIDIC ANTITUMOR ANTIBIOTIC SHOWDOMYCIN BY ARGUSLAB 4 SOFTWARE AFSHAN NAZ, KHALIDA BANO, FARHAT BANO, NAJAF ABBAS GHAFOOR AND NAHEED AKHTAR Biophysics
General Physical Chemistry II
 General Physical Chemistry II Lecture 13 Aleksey Kocherzhenko October 16, 2014" Last time " The Hückel method" Ø Used to study π systems of conjugated molecules" Ø π orbitals are treated separately from
General Physical Chemistry II Lecture 13 Aleksey Kocherzhenko October 16, 2014" Last time " The Hückel method" Ø Used to study π systems of conjugated molecules" Ø π orbitals are treated separately from
Session 1. Introduction to Computational Chemistry. Computational (chemistry education) and/or (Computational chemistry) education
 Session 1 Introduction to Computational Chemistry 1 Introduction to Computational Chemistry Computational (chemistry education) and/or (Computational chemistry) education First one: Use computational tools
Session 1 Introduction to Computational Chemistry 1 Introduction to Computational Chemistry Computational (chemistry education) and/or (Computational chemistry) education First one: Use computational tools
Semi-Empirical MO Methods
 Semi-Empirical MO Methods the high cost of ab initio MO calculations is largely due to the many integrals that need to be calculated (esp. two electron integrals) semi-empirical MO methods start with the
Semi-Empirical MO Methods the high cost of ab initio MO calculations is largely due to the many integrals that need to be calculated (esp. two electron integrals) semi-empirical MO methods start with the
AN EFFICIENT FeSO 4 MEDIATED SYNTHESIS OF METHYL-4-(ETHOXYMETHYL)-BENZOATE AND BASIC CONFORMATIONAL ANALYSIS OF THE SAME USING COMPUTATIONAL TOOLS
 29 Kragujevac J. Sci. 34 (2012) 29-37. UDC 541.27:541.67:543.42 AN EFFICIENT FeSO 4 MEDIATED SYNTHESIS OF METHYL-4-(ETHOXYMETHYL)-BENZOATE AND BASIC CONFORMATIONAL ANALYSIS OF THE SAME USING COMPUTATIONAL
29 Kragujevac J. Sci. 34 (2012) 29-37. UDC 541.27:541.67:543.42 AN EFFICIENT FeSO 4 MEDIATED SYNTHESIS OF METHYL-4-(ETHOXYMETHYL)-BENZOATE AND BASIC CONFORMATIONAL ANALYSIS OF THE SAME USING COMPUTATIONAL
CHEM 344 Molecular Modeling
 CHEM 344 Molecular Modeling The Use of Computational Chemistry to Support Experimental Organic Chemistry Part 1: Molecular Orbital Theory, Hybridization, & Formal Charge * all calculation data obtained
CHEM 344 Molecular Modeling The Use of Computational Chemistry to Support Experimental Organic Chemistry Part 1: Molecular Orbital Theory, Hybridization, & Formal Charge * all calculation data obtained
Structural biology and drug design: An overview
 Structural biology and drug design: An overview livier Taboureau Assitant professor Chemoinformatics group-cbs-dtu otab@cbs.dtu.dk Drug discovery Drug and drug design A drug is a key molecule involved
Structural biology and drug design: An overview livier Taboureau Assitant professor Chemoinformatics group-cbs-dtu otab@cbs.dtu.dk Drug discovery Drug and drug design A drug is a key molecule involved
Quantum Chemistry. NC State University. Lecture 5. The electronic structure of molecules Absorption spectroscopy Fluorescence spectroscopy
 Quantum Chemistry Lecture 5 The electronic structure of molecules Absorption spectroscopy Fluorescence spectroscopy NC State University 3.5 Selective absorption and emission by atmospheric gases (source:
Quantum Chemistry Lecture 5 The electronic structure of molecules Absorption spectroscopy Fluorescence spectroscopy NC State University 3.5 Selective absorption and emission by atmospheric gases (source:
An introduction to Molecular Dynamics. EMBO, June 2016
 An introduction to Molecular Dynamics EMBO, June 2016 What is MD? everything that living things do can be understood in terms of the jiggling and wiggling of atoms. The Feynman Lectures in Physics vol.
An introduction to Molecular Dynamics EMBO, June 2016 What is MD? everything that living things do can be understood in terms of the jiggling and wiggling of atoms. The Feynman Lectures in Physics vol.
hand and delocalization on the other, can be instructively exemplified and extended
 Text Related to Segment 8.0 00 Claude E. Wintner The ideas developed up to this point, concerning stereochemistry on the one hand and delocalization on the other, can be instructively exemplified and extended
Text Related to Segment 8.0 00 Claude E. Wintner The ideas developed up to this point, concerning stereochemistry on the one hand and delocalization on the other, can be instructively exemplified and extended
ENERGY MINIMIZATION AND CONFORMATION SEARCH ANALYSIS OF TYPE-2 ANTI-DIABETES DRUGS
 Int. J. Chem. Sci.: 6(2), 2008, 982-992 EERGY MIIMIZATI AD CFRMATI SEARC AALYSIS F TYPE-2 ATI-DIABETES DRUGS R. PRASAA LAKSMI a, C. ARASIMA KUMAR a, B. VASATA LAKSMI, K. AGA SUDA, K. MAJA, V. JAYA LAKSMI
Int. J. Chem. Sci.: 6(2), 2008, 982-992 EERGY MIIMIZATI AD CFRMATI SEARC AALYSIS F TYPE-2 ATI-DIABETES DRUGS R. PRASAA LAKSMI a, C. ARASIMA KUMAR a, B. VASATA LAKSMI, K. AGA SUDA, K. MAJA, V. JAYA LAKSMI
Lecture 1. Conformational Analysis in Acyclic Systems
 Lecture 1 Conformational Analysis in Acyclic Systems Learning Outcomes: by the end of this lecture and after answering the associated problems, you will be able to: 1. use Newman and saw-horse projections
Lecture 1 Conformational Analysis in Acyclic Systems Learning Outcomes: by the end of this lecture and after answering the associated problems, you will be able to: 1. use Newman and saw-horse projections
All-atom Molecular Mechanics. Trent E. Balius AMS 535 / CHE /27/2010
 All-atom Molecular Mechanics Trent E. Balius AMS 535 / CHE 535 09/27/2010 Outline Molecular models Molecular mechanics Force Fields Potential energy function functional form parameters and parameterization
All-atom Molecular Mechanics Trent E. Balius AMS 535 / CHE 535 09/27/2010 Outline Molecular models Molecular mechanics Force Fields Potential energy function functional form parameters and parameterization
Molecular Simulation I
 Molecular Simulation I Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences
Molecular Simulation I Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences
Chem120a : Exam 3 (Chem Bio) Solutions
 Chem10a : Exam 3 (Chem Bio) Solutions November 7, 006 Problem 1 This problem will basically involve us doing two Hückel calculations: one for the linear geometry, and one for the triangular geometry. We
Chem10a : Exam 3 (Chem Bio) Solutions November 7, 006 Problem 1 This problem will basically involve us doing two Hückel calculations: one for the linear geometry, and one for the triangular geometry. We
Tuning Color Through Substitution
 1 Tuning Color Through Substitution Introduction In this experiment, the effect of substituents on the absorbance spectra of molecules will be related to the structure of the molecular orbitals involved
1 Tuning Color Through Substitution Introduction In this experiment, the effect of substituents on the absorbance spectra of molecules will be related to the structure of the molecular orbitals involved
Lecture B6 Molecular Orbital Theory. Sometimes it's good to be alone.
 Lecture B6 Molecular Orbital Theory Sometimes it's good to be alone. Covalent Bond Theories 1. VSEPR (valence shell electron pair repulsion model). A set of empirical rules for predicting a molecular geometry
Lecture B6 Molecular Orbital Theory Sometimes it's good to be alone. Covalent Bond Theories 1. VSEPR (valence shell electron pair repulsion model). A set of empirical rules for predicting a molecular geometry
Chem 1140; Molecular Modeling
 P. Wipf 1 Chem 1140 $E = -! (q a + q b )" ab S ab + ab!k
P. Wipf 1 Chem 1140 $E = -! (q a + q b )" ab S ab + ab!k
Physical Chemistry II Recommended Problems Chapter 12( 23)
 Physical Chemistry II Recommended Problems Chapter 1( 3) Chapter 1(3) Problem. Overlap of two 1s orbitals recprobchap1.odt 1 Physical Chemistry II Recommended Problems, Chapter 1(3), continued Chapter
Physical Chemistry II Recommended Problems Chapter 1( 3) Chapter 1(3) Problem. Overlap of two 1s orbitals recprobchap1.odt 1 Physical Chemistry II Recommended Problems, Chapter 1(3), continued Chapter
Molecular Orbital Theory. Molecular Orbital Theory: Electrons are located in the molecule, not held in discrete regions between two bonded atoms
 Molecular Orbital Theory Valence Bond Theory: Electrons are located in discrete pairs between specific atoms Molecular Orbital Theory: Electrons are located in the molecule, not held in discrete regions
Molecular Orbital Theory Valence Bond Theory: Electrons are located in discrete pairs between specific atoms Molecular Orbital Theory: Electrons are located in the molecule, not held in discrete regions
QSAR Studies of Some Pharmacological Schiff Base Compounds
 Asian Journal of Chemistry Vol. 20, o. 8 (2008), 594-5946 QSAR Studies of Some Pharmacological Schiff Base Compounds KISHOR ARORA Department of Chemistry, Government K.R.G. Autonomous Post Graduate College,
Asian Journal of Chemistry Vol. 20, o. 8 (2008), 594-5946 QSAR Studies of Some Pharmacological Schiff Base Compounds KISHOR ARORA Department of Chemistry, Government K.R.G. Autonomous Post Graduate College,
Learning to Use Scigress Wagner, Eugene P. (revised May 15, 2018)
 Learning to Use Scigress Wagner, Eugene P. (revised May 15, 2018) Abstract Students are introduced to basic features of Scigress by building molecules and performing calculations on them using semi-empirical
Learning to Use Scigress Wagner, Eugene P. (revised May 15, 2018) Abstract Students are introduced to basic features of Scigress by building molecules and performing calculations on them using semi-empirical
Semi-Empirical Methods CHEM 430
 Semi-Empirical Methods CHEM 430 Cost, Hartree%Fock, scales,as,n 4,(N=#, basis,funcfons), Due,to,two% electron, integrals, within,fock, matrix, Semi%empirical,cut, cost,by,reducing, number,of, integrals,
Semi-Empirical Methods CHEM 430 Cost, Hartree%Fock, scales,as,n 4,(N=#, basis,funcfons), Due,to,two% electron, integrals, within,fock, matrix, Semi%empirical,cut, cost,by,reducing, number,of, integrals,
QSAR of Microtubule Stabilizing Dictyostatins
 QSAR of Microtubule Stabilizing Dictyostatins Kia Montgomery BBSI 2007- University of Pittsburgh Department of Chemistry, Grambling State University Billy Day, Ph.D. Department of Pharmaceutical Sciences,
QSAR of Microtubule Stabilizing Dictyostatins Kia Montgomery BBSI 2007- University of Pittsburgh Department of Chemistry, Grambling State University Billy Day, Ph.D. Department of Pharmaceutical Sciences,
Ethene. Introduction. The ethene molecule is planar (i.e. all the six atoms lie in the same plane) and has a high degree of symmetry:
 FY1006 Innføring i kvantefysikk og TFY4215 Kjemisk fysikk og kvantemekanikk Spring 2012 Chemical Physics Exercise 1 To be delivered by Friday 27.04.12 Introduction Ethene. Ethylene, C 2 H 4, or ethene,
FY1006 Innføring i kvantefysikk og TFY4215 Kjemisk fysikk og kvantemekanikk Spring 2012 Chemical Physics Exercise 1 To be delivered by Friday 27.04.12 Introduction Ethene. Ethylene, C 2 H 4, or ethene,
( ) + ( ) + ( ) = 0.00
 Chem 3502/4502 Physical Chemistry II (Quantum Mechanics) 3 Credits Spring Semester 2006 Christopher J. Cramer Lecture 32, April 14, 2006 (Some material in this lecture has been adapted from Cramer, C.
Chem 3502/4502 Physical Chemistry II (Quantum Mechanics) 3 Credits Spring Semester 2006 Christopher J. Cramer Lecture 32, April 14, 2006 (Some material in this lecture has been adapted from Cramer, C.
Keywords: anti-coagulants, factor Xa, QSAR, Thrombosis. Introduction
 PostDoc Journal Vol. 2, No. 3, March 2014 Journal of Postdoctoral Research www.postdocjournal.com QSAR Study of Thiophene-Anthranilamides Based Factor Xa Direct Inhibitors Preetpal S. Sidhu Department
PostDoc Journal Vol. 2, No. 3, March 2014 Journal of Postdoctoral Research www.postdocjournal.com QSAR Study of Thiophene-Anthranilamides Based Factor Xa Direct Inhibitors Preetpal S. Sidhu Department
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015,
 Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
MOLECULAR STRUCTURE. Molecular Structure - B. Molecular Structure - B. Molecular Structure - B. Molecular Structure - B. Molecular Structure - B
 MOLECULAR STRUCTURE Molecular Orbital all orbitals of the appropriate symmetry contribute to a molecular orbital. Bundet Boekfa Chem Div, Faculty Lib Arts & Sci Kasetsart University Kamphaeng Saen Campus
MOLECULAR STRUCTURE Molecular Orbital all orbitals of the appropriate symmetry contribute to a molecular orbital. Bundet Boekfa Chem Div, Faculty Lib Arts & Sci Kasetsart University Kamphaeng Saen Campus
COMPUTER AIDED DRUG DESIGN (CADD) AND DEVELOPMENT METHODS
 COMPUTER AIDED DRUG DESIGN (CADD) AND DEVELOPMENT METHODS DRUG DEVELOPMENT Drug development is a challenging path Today, the causes of many diseases (rheumatoid arthritis, cancer, mental diseases, etc.)
COMPUTER AIDED DRUG DESIGN (CADD) AND DEVELOPMENT METHODS DRUG DEVELOPMENT Drug development is a challenging path Today, the causes of many diseases (rheumatoid arthritis, cancer, mental diseases, etc.)
Quantum Chemical Simulations and Descriptors. Dr. Antonio Chana, Dr. Mosè Casalegno
 Quantum Chemical Simulations and Descriptors Dr. Antonio Chana, Dr. Mosè Casalegno Classical Mechanics: basics It models real-world objects as point particles, objects with negligible size. The motion
Quantum Chemical Simulations and Descriptors Dr. Antonio Chana, Dr. Mosè Casalegno Classical Mechanics: basics It models real-world objects as point particles, objects with negligible size. The motion
CHEM3023: Spins, Atoms and Molecules
 CHEM3023: Spins, Atoms and Molecules Lecture 5 The Hartree-Fock method C.-K. Skylaris Learning outcomes Be able to use the variational principle in quantum calculations Be able to construct Fock operators
CHEM3023: Spins, Atoms and Molecules Lecture 5 The Hartree-Fock method C.-K. Skylaris Learning outcomes Be able to use the variational principle in quantum calculations Be able to construct Fock operators
3rd Advanced in silico Drug Design KFC/ADD Molecular mechanics intro Karel Berka, Ph.D. Martin Lepšík, Ph.D. Pavel Polishchuk, Ph.D.
 3rd Advanced in silico Drug Design KFC/ADD Molecular mechanics intro Karel Berka, Ph.D. Martin Lepšík, Ph.D. Pavel Polishchuk, Ph.D. Thierry Langer, Ph.D. Jana Vrbková, Ph.D. UP Olomouc, 23.1.-26.1. 2018
3rd Advanced in silico Drug Design KFC/ADD Molecular mechanics intro Karel Berka, Ph.D. Martin Lepšík, Ph.D. Pavel Polishchuk, Ph.D. Thierry Langer, Ph.D. Jana Vrbková, Ph.D. UP Olomouc, 23.1.-26.1. 2018
Computational study on the geometry optimization and excited State properties of Riboflavin by ArgusLab 4.0.1
 Computational study on the geometry optimization and excited State properties of Riboflavin by ArgusLab 4.0.1 Ambreen Hafeez*, Afshan Naz, Sadaf Naeem, Khalida Bano and Naheed Akhtar Department of Biochemistry,
Computational study on the geometry optimization and excited State properties of Riboflavin by ArgusLab 4.0.1 Ambreen Hafeez*, Afshan Naz, Sadaf Naeem, Khalida Bano and Naheed Akhtar Department of Biochemistry,
Chem 3502/4502 Physical Chemistry II (Quantum Mechanics) 3 Credits Spring Semester Christopher J. Cramer. Lecture 30, April 10, 2006
 Chem 3502/4502 Physical Chemistry II (Quantum Mechanics) 3 Credits Spring Semester 20056 Christopher J. Cramer Lecture 30, April 10, 2006 Solved Homework The guess MO occupied coefficients were Occupied
Chem 3502/4502 Physical Chemistry II (Quantum Mechanics) 3 Credits Spring Semester 20056 Christopher J. Cramer Lecture 30, April 10, 2006 Solved Homework The guess MO occupied coefficients were Occupied
Excited States Calculations for Protonated PAHs
 52 Chapter 3 Excited States Calculations for Protonated PAHs 3.1 Introduction Protonated PAHs are closed shell ions. Their electronic structure should therefore be similar to that of neutral PAHs, but
52 Chapter 3 Excited States Calculations for Protonated PAHs 3.1 Introduction Protonated PAHs are closed shell ions. Their electronic structure should therefore be similar to that of neutral PAHs, but
Structural Bioinformatics (C3210) Molecular Mechanics
 Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
Lecture 5: Electrostatic Interactions & Screening
 Lecture 5: Electrostatic Interactions & Screening Lecturer: Prof. Brigita Urbanc (brigita@drexel.edu) PHYS 461 & 561, Fall 2009-2010 1 A charged particle (q=+1) in water, at the interface between water
Lecture 5: Electrostatic Interactions & Screening Lecturer: Prof. Brigita Urbanc (brigita@drexel.edu) PHYS 461 & 561, Fall 2009-2010 1 A charged particle (q=+1) in water, at the interface between water
LUMO + 1 LUMO. Tómas Arnar Guðmundsson Report 2 Reikniefnafræði G
 Q1: Display all the MOs for N2 in your report and classify each one of them as bonding, antibonding or non-bonding, and say whether the symmetry of the orbital is σ or π. Sketch a molecular orbital diagram
Q1: Display all the MOs for N2 in your report and classify each one of them as bonding, antibonding or non-bonding, and say whether the symmetry of the orbital is σ or π. Sketch a molecular orbital diagram
Chemistry 2000 Lecture 1: Introduction to the molecular orbital theory
 Chemistry 2000 Lecture 1: Introduction to the molecular orbital theory Marc R. Roussel January 5, 2018 Marc R. Roussel Introduction to molecular orbitals January 5, 2018 1 / 24 Review: quantum mechanics
Chemistry 2000 Lecture 1: Introduction to the molecular orbital theory Marc R. Roussel January 5, 2018 Marc R. Roussel Introduction to molecular orbitals January 5, 2018 1 / 24 Review: quantum mechanics
CHEM 2010 Symmetry, Electronic Structure and Bonding Winter Numbering of Chapters and Assigned Problems
 CHEM 2010 Symmetry, Electronic Structure and Bonding Winter 2011 Numbering of Chapters and Assigned Problems The following table shows the correspondence between the chapter numbers in the full book (Physical
CHEM 2010 Symmetry, Electronic Structure and Bonding Winter 2011 Numbering of Chapters and Assigned Problems The following table shows the correspondence between the chapter numbers in the full book (Physical
Lecture 4: Band theory
 Lecture 4: Band theory Very short introduction to modern computational solid state chemistry Band theory of solids Molecules vs. solids Band structures Analysis of chemical bonding in Reciprocal space
Lecture 4: Band theory Very short introduction to modern computational solid state chemistry Band theory of solids Molecules vs. solids Band structures Analysis of chemical bonding in Reciprocal space
Introduction to Hartree-Fock calculations in Spartan
 EE5 in 2008 Hannes Jónsson Introduction to Hartree-Fock calculations in Spartan In this exercise, you will get to use state of the art software for carrying out calculations of wavefunctions for molecues,
EE5 in 2008 Hannes Jónsson Introduction to Hartree-Fock calculations in Spartan In this exercise, you will get to use state of the art software for carrying out calculations of wavefunctions for molecues,
Chemical Bonding. Lewis Theory-VSEPR Valence Bond Theory Molecular Orbital Theory
 Chemical Bonding Lewis Theory-VSEPR Valence Bond Theory Molecular Orbital Theory Problems with Valence Bond Theory VB theory predicts properties better than Lewis theory bonding schemes, bond strengths,
Chemical Bonding Lewis Theory-VSEPR Valence Bond Theory Molecular Orbital Theory Problems with Valence Bond Theory VB theory predicts properties better than Lewis theory bonding schemes, bond strengths,
CHEM 344 Molecular Modeling
 CHEM 344 Molecular Modeling The Use of Computational Chemistry to Support Experimental Organic Chemistry Day 1 all calculation data obtained from Gaussian09 using B3LYP/6-31G(d) unless otherwise noted.
CHEM 344 Molecular Modeling The Use of Computational Chemistry to Support Experimental Organic Chemistry Day 1 all calculation data obtained from Gaussian09 using B3LYP/6-31G(d) unless otherwise noted.
Lecture 16, February 25, 2015 Metallic bonding
 Lecture 16, February 25, 2015 Metallic bonding Elements of Quantum Chemistry with Applications to Chemical Bonding and Properties of Molecules and Solids Course number: Ch125a; Room 115 BI Hours: 11-11:50am
Lecture 16, February 25, 2015 Metallic bonding Elements of Quantum Chemistry with Applications to Chemical Bonding and Properties of Molecules and Solids Course number: Ch125a; Room 115 BI Hours: 11-11:50am
Module17: Intermolecular Force between Surfaces and Particles. Lecture 23: Intermolecular Force between Surfaces and Particles
 Module17: Intermolecular Force between Surfaces and Particles Lecture 23: Intermolecular Force between Surfaces and Particles 1 We now try to understand the nature of spontaneous instability in a confined
Module17: Intermolecular Force between Surfaces and Particles Lecture 23: Intermolecular Force between Surfaces and Particles 1 We now try to understand the nature of spontaneous instability in a confined
Receptor Based Drug Design (1)
 Induced Fit Model For more than 100 years, the behaviour of enzymes had been explained by the "lock-and-key" mechanism developed by pioneering German chemist Emil Fischer. Fischer thought that the chemicals
Induced Fit Model For more than 100 years, the behaviour of enzymes had been explained by the "lock-and-key" mechanism developed by pioneering German chemist Emil Fischer. Fischer thought that the chemicals
Life Sciences 1a Lecture Slides Set 10 Fall Prof. David R. Liu. Lecture Readings. Required: Lecture Notes McMurray p , O NH
 Life ciences 1a Lecture lides et 10 Fall 2006-2007 Prof. David R. Liu Lectures 17-18: The molecular basis of drug-protein binding: IV protease inhibitors 1. Drug development and its impact on IV-infected
Life ciences 1a Lecture lides et 10 Fall 2006-2007 Prof. David R. Liu Lectures 17-18: The molecular basis of drug-protein binding: IV protease inhibitors 1. Drug development and its impact on IV-infected
PHARMA SCIENCE MONITOR
 PHARMA SCIENCE MONITOR AN INTERNATIONAL JOURNAL OF PHARMACEUTICAL SCIENCES EXTENDED HUCKEL PARTIAL ATOMIC CHARGES OF NITROIMIDAZOLE AND PREDICTION OF DNA DAMAGE B.R.Prajapati *, A.K.Seth 2, K.I.Molvi 3,
PHARMA SCIENCE MONITOR AN INTERNATIONAL JOURNAL OF PHARMACEUTICAL SCIENCES EXTENDED HUCKEL PARTIAL ATOMIC CHARGES OF NITROIMIDAZOLE AND PREDICTION OF DNA DAMAGE B.R.Prajapati *, A.K.Seth 2, K.I.Molvi 3,
Computational Chemistry. An Introduction to Molecular Dynamic Simulations
 Computational Chemistry An Introduction to Molecular Dynamic Simulations Computational chemistry simulates chemical structures and reactions numerically, based in full or in part on the fundamental laws
Computational Chemistry An Introduction to Molecular Dynamic Simulations Computational chemistry simulates chemical structures and reactions numerically, based in full or in part on the fundamental laws
Ping-Chiang Lyu. Institute of Bioinformatics and Structural Biology, Department of Life Science, National Tsing Hua University.
 Pharmacophore-based Drug design Ping-Chiang Lyu Institute of Bioinformatics and Structural Biology, Department of Life Science, National Tsing Hua University 96/08/07 Outline Part I: Analysis The analytical
Pharmacophore-based Drug design Ping-Chiang Lyu Institute of Bioinformatics and Structural Biology, Department of Life Science, National Tsing Hua University 96/08/07 Outline Part I: Analysis The analytical
Conformation Searching Applications
 Chemistry 380.37 Fall 2015 Dr. Jean M. Standard September 23, 2015 Conformation Searching Applications Cycloheptadecane Redux So what would a conformation search on cycloheptadecane using Spartan find?
Chemistry 380.37 Fall 2015 Dr. Jean M. Standard September 23, 2015 Conformation Searching Applications Cycloheptadecane Redux So what would a conformation search on cycloheptadecane using Spartan find?
Tutorial I: IQ MOL and Basic DFT and MP2 Calculations 1 / 30
 Tutorial I: IQ MOL and Basic DFT and MP2 Calculations Q-Chem User Workshop, Denver March 21, 2015 1 / 30 2 / 30 Introduction to IQMOL DFT and MP2 Calculations 3 / 30 IQMOL and Q-CHEM IQMOL is an open-source
Tutorial I: IQ MOL and Basic DFT and MP2 Calculations Q-Chem User Workshop, Denver March 21, 2015 1 / 30 2 / 30 Introduction to IQMOL DFT and MP2 Calculations 3 / 30 IQMOL and Q-CHEM IQMOL is an open-source
Chemistry 200: Basic Chemistry and Applications Course Syllabus: Spring
 Chemistry 200: Basic Chemistry and Applications Course Syllabus: Spring 2017 2018 Course Instructors Faraj Hasanayn; Office of Faraj Hasanayn: Chem Bldg. Rm 522 Office Hours: TBA Email: fh19@aub.edu.lb.
Chemistry 200: Basic Chemistry and Applications Course Syllabus: Spring 2017 2018 Course Instructors Faraj Hasanayn; Office of Faraj Hasanayn: Chem Bldg. Rm 522 Office Hours: TBA Email: fh19@aub.edu.lb.
with the larger dimerization energy also exhibits the larger structural changes.
 A7. Looking at the image and table provided below, it is apparent that the monomer and dimer are structurally almost identical. Although angular and dihedral data were not included, these data are also
A7. Looking at the image and table provided below, it is apparent that the monomer and dimer are structurally almost identical. Although angular and dihedral data were not included, these data are also
Lecture 11: Protein Folding & Stability
 Structure - Function Protein Folding: What we know Lecture 11: Protein Folding & Stability 1). Amino acid sequence dictates structure. 2). The native structure represents the lowest energy state for a
Structure - Function Protein Folding: What we know Lecture 11: Protein Folding & Stability 1). Amino acid sequence dictates structure. 2). The native structure represents the lowest energy state for a
Protein Folding & Stability. Lecture 11: Margaret A. Daugherty. Fall Protein Folding: What we know. Protein Folding
 Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2003 Structure - Function Protein Folding: What we know 1). Amino acid sequence dictates structure. 2). The native structure represents
Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2003 Structure - Function Protein Folding: What we know 1). Amino acid sequence dictates structure. 2). The native structure represents
My additional comments, questions are colored in blue in the following slides.
 My additional comments, questions are colored in blue in the following slides. Do not forget to work the assigned HW from the text that is also posted on the boardlist. I will post my answers to these
My additional comments, questions are colored in blue in the following slides. Do not forget to work the assigned HW from the text that is also posted on the boardlist. I will post my answers to these
Advanced Medicinal Chemistry SLIDES B
 Advanced Medicinal Chemistry Filippo Minutolo CFU 3 (21 hours) SLIDES B Drug likeness - ADME two contradictory physico-chemical parameters to balance: 1) aqueous solubility 2) lipid membrane permeability
Advanced Medicinal Chemistry Filippo Minutolo CFU 3 (21 hours) SLIDES B Drug likeness - ADME two contradictory physico-chemical parameters to balance: 1) aqueous solubility 2) lipid membrane permeability
BIOC : Homework 1 Due 10/10
 Contact information: Name: Student # BIOC530 2012: Homework 1 Due 10/10 Department Email address The following problems are based on David Baker s lectures of forces and protein folding. When numerical
Contact information: Name: Student # BIOC530 2012: Homework 1 Due 10/10 Department Email address The following problems are based on David Baker s lectures of forces and protein folding. When numerical
ONETEP PB/SA: Application to G-Quadruplex DNA Stability. Danny Cole
 ONETEP PB/SA: Application to G-Quadruplex DNA Stability Danny Cole Introduction Historical overview of structure and free energy calculation of complex molecules using molecular mechanics and continuum
ONETEP PB/SA: Application to G-Quadruplex DNA Stability Danny Cole Introduction Historical overview of structure and free energy calculation of complex molecules using molecular mechanics and continuum
Lecture 10. Born-Oppenheimer approximation LCAO-MO application to H + The potential energy surface MOs for diatomic molecules. NC State University
 Chemistry 431 Lecture 10 Diatomic molecules Born-Oppenheimer approximation LCAO-MO application to H + 2 The potential energy surface MOs for diatomic molecules NC State University Born-Oppenheimer approximation
Chemistry 431 Lecture 10 Diatomic molecules Born-Oppenheimer approximation LCAO-MO application to H + 2 The potential energy surface MOs for diatomic molecules NC State University Born-Oppenheimer approximation
Protein Folding & Stability. Lecture 11: Margaret A. Daugherty. Fall How do we go from an unfolded polypeptide chain to a
 Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2004 How do we go from an unfolded polypeptide chain to a compact folded protein? (Folding of thioredoxin, F. Richards) Structure - Function
Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2004 How do we go from an unfolded polypeptide chain to a compact folded protein? (Folding of thioredoxin, F. Richards) Structure - Function
Computational Methods. Chem 561
 Computational Methods Chem 561 Lecture Outline 1. Ab initio methods a) HF SCF b) Post-HF methods 2. Density Functional Theory 3. Semiempirical methods 4. Molecular Mechanics Computational Chemistry " Computational
Computational Methods Chem 561 Lecture Outline 1. Ab initio methods a) HF SCF b) Post-HF methods 2. Density Functional Theory 3. Semiempirical methods 4. Molecular Mechanics Computational Chemistry " Computational
Chemical bonding in solids from ab-initio Calculations
 Chemical bonding in solids from ab-initio Calculations 1 Prof.P. Ravindran, Department of Physics, Central University of Tamil Nadu, India & Center for Materials Science and Nanotechnology, University
Chemical bonding in solids from ab-initio Calculations 1 Prof.P. Ravindran, Department of Physics, Central University of Tamil Nadu, India & Center for Materials Science and Nanotechnology, University
VSEPR Model. Valence-Shell Electron-Pair Repulsion Bonds (single or multiple) and lone pairs are thought of as charge clouds
 Molecular Shapes VSEPR Model Valence-Shell Electron-Pair Repulsion Bonds (single or multiple) and lone pairs are thought of as charge clouds They repel each other and stay as far away from each other as
Molecular Shapes VSEPR Model Valence-Shell Electron-Pair Repulsion Bonds (single or multiple) and lone pairs are thought of as charge clouds They repel each other and stay as far away from each other as
Atomic and molecular interaction forces in biology
 Atomic and molecular interaction forces in biology 1 Outline Types of interactions relevant to biology Van der Waals interactions H-bond interactions Some properties of water Hydrophobic effect 2 Types
Atomic and molecular interaction forces in biology 1 Outline Types of interactions relevant to biology Van der Waals interactions H-bond interactions Some properties of water Hydrophobic effect 2 Types
Biochemistry 675, Lecture 8 Electrostatic Interactions
 Biochemistry 675, Lecture 8 Electrostatic Interactions Previous Classes Hydrophobic Effect Hydrogen Bonds Today: Electrostatic Interactions Reading: Handout:P.R. Bergethon & E.R. Simons, Biophysical Chemistry,
Biochemistry 675, Lecture 8 Electrostatic Interactions Previous Classes Hydrophobic Effect Hydrogen Bonds Today: Electrostatic Interactions Reading: Handout:P.R. Bergethon & E.R. Simons, Biophysical Chemistry,
Chemistry 3211 Coordination Chemistry Part 3 Ligand Field and Molecular Orbital Theory
 Chemistry 3211 Coordination Chemistry Part 3 Ligand Field and Molecular Orbital Theory Electronic Structure of Six and Four-Coordinate Complexes Using Crystal Field Theory, we can generate energy level
Chemistry 3211 Coordination Chemistry Part 3 Ligand Field and Molecular Orbital Theory Electronic Structure of Six and Four-Coordinate Complexes Using Crystal Field Theory, we can generate energy level
The protein folding problem consists of two parts:
 Energetics and kinetics of protein folding The protein folding problem consists of two parts: 1)Creating a stable, well-defined structure that is significantly more stable than all other possible structures.
Energetics and kinetics of protein folding The protein folding problem consists of two parts: 1)Creating a stable, well-defined structure that is significantly more stable than all other possible structures.
BCMP 201 Protein biochemistry
 BCMP 201 Protein biochemistry BCMP 201 Protein biochemistry with emphasis on the interrelated roles of protein structure, catalytic activity, and macromolecular interactions in biological processes. The
BCMP 201 Protein biochemistry BCMP 201 Protein biochemistry with emphasis on the interrelated roles of protein structure, catalytic activity, and macromolecular interactions in biological processes. The
Fundamental Interactions: 6 Forces
 Fundamental Interactions: 6 Forces In nuclear and high-energy physics 6 fundamental forces are recognized, which describe the structure of matter. - the strong interaction - the weak interaction act inside
Fundamental Interactions: 6 Forces In nuclear and high-energy physics 6 fundamental forces are recognized, which describe the structure of matter. - the strong interaction - the weak interaction act inside
Contents. 1. Building a molecular model 2. Additional Procedures and Options: 3. Selecting a calculation method
 Getting started with HyperChem. Program description. In this project you will create simple organic molecules and learn how to manipulate them. However, before starting you should create your home directory
Getting started with HyperChem. Program description. In this project you will create simple organic molecules and learn how to manipulate them. However, before starting you should create your home directory
2. A(nother) BRIEF TOUR OF COMPUTATIONAL CHEMISTRY METHODS
 1. PURPOSE OF THE EXPERIMET This project will further your exposure to PC-Spartan for modeling molecular properties. In particular, you will investigate the electronic structure of polyenes and porphyrins
1. PURPOSE OF THE EXPERIMET This project will further your exposure to PC-Spartan for modeling molecular properties. In particular, you will investigate the electronic structure of polyenes and porphyrins
Plan. Day 2: Exercise on MHC molecules.
 Plan Day 1: What is Chemoinformatics and Drug Design? Methods and Algorithms used in Chemoinformatics including SVM. Cross validation and sequence encoding Example and exercise with herg potassium channel:
Plan Day 1: What is Chemoinformatics and Drug Design? Methods and Algorithms used in Chemoinformatics including SVM. Cross validation and sequence encoding Example and exercise with herg potassium channel:
Randa AbiRafi Jaber Office: Chem Bldg. Rm 212 Office Hrs: Tue, Wed 11:00 Thu: 2:00 3:00 pm
 Faraj Hasanayn Office Chem Bldg. Rm 522 Office Hrs: Mon: 10:30 11:30 am Tuesday: 1:30 2:30 pm, and by appointment Email: fh19@aub.edu.lb, Phone Ext: 3994 Randa AbiRafi Jaber Office: Chem Bldg. Rm 212 Office
Faraj Hasanayn Office Chem Bldg. Rm 522 Office Hrs: Mon: 10:30 11:30 am Tuesday: 1:30 2:30 pm, and by appointment Email: fh19@aub.edu.lb, Phone Ext: 3994 Randa AbiRafi Jaber Office: Chem Bldg. Rm 212 Office
Dihedral Angles. Homayoun Valafar. Department of Computer Science and Engineering, USC 02/03/10 CSCE 769
 Dihedral Angles Homayoun Valafar Department of Computer Science and Engineering, USC The precise definition of a dihedral or torsion angle can be found in spatial geometry Angle between to planes Dihedral
Dihedral Angles Homayoun Valafar Department of Computer Science and Engineering, USC The precise definition of a dihedral or torsion angle can be found in spatial geometry Angle between to planes Dihedral
Physical Chemistry - Problem Drill 01: Chemistry and Physics Review
 Physical Chemistry - Problem Drill 01: Chemistry and Physics Review No. 1 of 10 1. Chemical bonds are considered to be the interaction of their electronic structures of bonding atoms involved, with the
Physical Chemistry - Problem Drill 01: Chemistry and Physics Review No. 1 of 10 1. Chemical bonds are considered to be the interaction of their electronic structures of bonding atoms involved, with the
MOLECULAR DRUG TARGETS
 MOLECULAR DRUG TARGETS LEARNING OUTCOMES At the end of this session student shall be able to: List different types of druggable targets Describe forces involved in drug-receptor interactions Describe theories
MOLECULAR DRUG TARGETS LEARNING OUTCOMES At the end of this session student shall be able to: List different types of druggable targets Describe forces involved in drug-receptor interactions Describe theories
Journal of Chemical and Pharmaceutical Research
 Available on line www.jocpr.com Journal of Chemical and Pharmaceutical Research ISSN No: 0975-7384 CODEN(USA): JCPRC5 J. Chem. Pharm. Res., 2011, 3(1):572-583 Ground state properties: Gross orbital charges,
Available on line www.jocpr.com Journal of Chemical and Pharmaceutical Research ISSN No: 0975-7384 CODEN(USA): JCPRC5 J. Chem. Pharm. Res., 2011, 3(1):572-583 Ground state properties: Gross orbital charges,
Molecular Modeling Study of Some Anthelmintic 2-phenyl Benzimidazole-1- Acetamides as β-tubulin Inhibitor
 Sawant et al : Molecular Modeling Study of Some Anthelmintic 2-phenyl Benzimidazole-1-Acetamides as -tubulin Inhibitor 1269 International Journal of Drug Design and Discovery Volume 5 Issue 1 January March
Sawant et al : Molecular Modeling Study of Some Anthelmintic 2-phenyl Benzimidazole-1-Acetamides as -tubulin Inhibitor 1269 International Journal of Drug Design and Discovery Volume 5 Issue 1 January March
In silico pharmacology for drug discovery
 In silico pharmacology for drug discovery In silico drug design In silico methods can contribute to drug targets identification through application of bionformatics tools. Currently, the application of
In silico pharmacology for drug discovery In silico drug design In silico methods can contribute to drug targets identification through application of bionformatics tools. Currently, the application of
BIBC 100. Structural Biochemistry
 BIBC 100 Structural Biochemistry http://classes.biology.ucsd.edu/bibc100.wi14 Papers- Dialogue with Scientists Questions: Why? How? What? So What? Dialogue Structure to explain function Knowledge Food
BIBC 100 Structural Biochemistry http://classes.biology.ucsd.edu/bibc100.wi14 Papers- Dialogue with Scientists Questions: Why? How? What? So What? Dialogue Structure to explain function Knowledge Food
A Computer Study of Molecular Electronic Structure
 A Computer Study of Molecular Electronic Structure The following exercises are designed to give you a brief introduction to some of the types of information that are now readily accessible from electronic
A Computer Study of Molecular Electronic Structure The following exercises are designed to give you a brief introduction to some of the types of information that are now readily accessible from electronic
SCH 102. Dr Albert Ndakala. Mobile: Office: Basement of Chemistry (B-6)
 SCH 102 (INTRODUCTION TO ORGANIC CHEMISTRY, CHEMISTRY OF ALKANES AND CYCLOALKANES) Dr Albert Ndakala Email: andakala@uonbi.ac.ke Mobile: 0720443386 Office: Basement of Chemistry (B-6) B.Pharm Lecture Timetable
SCH 102 (INTRODUCTION TO ORGANIC CHEMISTRY, CHEMISTRY OF ALKANES AND CYCLOALKANES) Dr Albert Ndakala Email: andakala@uonbi.ac.ke Mobile: 0720443386 Office: Basement of Chemistry (B-6) B.Pharm Lecture Timetable
Next Generation Computational Chemistry Tools to Predict Toxicity of CWAs
 Next Generation Computational Chemistry Tools to Predict Toxicity of CWAs William (Bill) Welsh welshwj@umdnj.edu Prospective Funding by DTRA/JSTO-CBD CBIS Conference 1 A State-wide, Regional and National
Next Generation Computational Chemistry Tools to Predict Toxicity of CWAs William (Bill) Welsh welshwj@umdnj.edu Prospective Funding by DTRA/JSTO-CBD CBIS Conference 1 A State-wide, Regional and National
4 th Advanced in silico Drug Design KFC/ADD Molecular Modelling Intro. Karel Berka, Ph.D.
 4 th Advanced in silico Drug Design KFC/ADD Molecular Modelling Intro Karel Berka, Ph.D. UP Olomouc, 21.1.-25.1. 2019 Motto A theory is something nobody believes, except the person who made it An experiment
4 th Advanced in silico Drug Design KFC/ADD Molecular Modelling Intro Karel Berka, Ph.D. UP Olomouc, 21.1.-25.1. 2019 Motto A theory is something nobody believes, except the person who made it An experiment
Chem 3502/4502 Physical Chemistry II (Quantum Mechanics) 3 Credits Spring Semester 2006 Christopher J. Cramer. Lecture 31, April 12, 2006
 Chem 3502/4502 Physical Chemistry II (Quantum Mechanics) 3 Credits Spring Semester 2006 Christopher J. Cramer Lecture 31, April 12, 2006 (Some material in this lecture has been adapted from Cramer, C.
Chem 3502/4502 Physical Chemistry II (Quantum Mechanics) 3 Credits Spring Semester 2006 Christopher J. Cramer Lecture 31, April 12, 2006 (Some material in this lecture has been adapted from Cramer, C.
International Journal of Chemistry and Pharmaceutical Sciences
 Jain eha et al IJCPS, 2014, Vol.2(10): 1203-1210 Research Article ISS: 2321-3132 International Journal of Chemistry and Pharmaceutical Sciences www.pharmaresearchlibrary.com/ijcps Efficient Computational
Jain eha et al IJCPS, 2014, Vol.2(10): 1203-1210 Research Article ISS: 2321-3132 International Journal of Chemistry and Pharmaceutical Sciences www.pharmaresearchlibrary.com/ijcps Efficient Computational
Chapter 6- An Introduction to Metabolism*
 Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Supplementary Tables and Figures
 Supplementary Tables Supplementary Tables and Figures Supplementary Table 1: Tumor types and samples analyzed. Supplementary Table 2: Genes analyzed here. Supplementary Table 3: Statistically significant
Supplementary Tables Supplementary Tables and Figures Supplementary Table 1: Tumor types and samples analyzed. Supplementary Table 2: Genes analyzed here. Supplementary Table 3: Statistically significant
Molecular Orbitals for Ozone
 Molecular Orbitals for Ozone Purpose: In this exercise you will do semi-empirical molecular orbital calculations on ozone with the goal of understanding the molecular orbital print out provided by Spartan
Molecular Orbitals for Ozone Purpose: In this exercise you will do semi-empirical molecular orbital calculations on ozone with the goal of understanding the molecular orbital print out provided by Spartan
Medicinal Chemistry and Chemical Biology
 Medicinal Chemistry and Chemical Biology Activities Drug Discovery Imaging Chemical Biology Computational Chemistry Natural Product Synthesis Current Staff Mike Waring Professor of Medicinal Chemistry
Medicinal Chemistry and Chemical Biology Activities Drug Discovery Imaging Chemical Biology Computational Chemistry Natural Product Synthesis Current Staff Mike Waring Professor of Medicinal Chemistry
QSAR Study of Quinazoline Derivatives as Inhibitor of Epidermal Growth Factor Receptor-Tyrosine Kinase (EGFR-TK)
 3rd International Conference on Computation for cience and Technology (ICCT-3) QAR tudy of Quinazoline Derivatives as Inhibitor of Epidermal Growth Factor Receptor-Tyrosine Kinase (EGFR-TK) La de Aman1*,
3rd International Conference on Computation for cience and Technology (ICCT-3) QAR tudy of Quinazoline Derivatives as Inhibitor of Epidermal Growth Factor Receptor-Tyrosine Kinase (EGFR-TK) La de Aman1*,
Computational Biology 1
 Computational Biology 1 Protein Function & nzyme inetics Guna Rajagopal, Bioinformatics Institute, guna@bii.a-star.edu.sg References : Molecular Biology of the Cell, 4 th d. Alberts et. al. Pg. 129 190
Computational Biology 1 Protein Function & nzyme inetics Guna Rajagopal, Bioinformatics Institute, guna@bii.a-star.edu.sg References : Molecular Biology of the Cell, 4 th d. Alberts et. al. Pg. 129 190
74 these states cannot be reliably obtained from experiments. In addition, the barriers between the local minima can also not be obtained reliably fro
 73 Chapter 5 Development of Adiabatic Force Field for Polyvinyl Chloride (PVC) and Chlorinated PVC (CPVC) 5.1 Introduction Chlorinated polyvinyl chloride has become an important specialty polymer due to
73 Chapter 5 Development of Adiabatic Force Field for Polyvinyl Chloride (PVC) and Chlorinated PVC (CPVC) 5.1 Introduction Chlorinated polyvinyl chloride has become an important specialty polymer due to
General Chemistry I (2012) Lecture by B. H. Hong
 3.8 The Limitations of Lewis's Theory 3.9 Molecular Orbitals The valence-bond (VB) and molecular orbital (MO) theories are both procedures for constructing approximate wavefunctions of electrons. The MO
3.8 The Limitations of Lewis's Theory 3.9 Molecular Orbitals The valence-bond (VB) and molecular orbital (MO) theories are both procedures for constructing approximate wavefunctions of electrons. The MO
