Amide I -II 2D IR spectroscopy provides enhanced protein secondary structural sensitivity
|
|
|
- Rodney May
- 6 years ago
- Views:
Transcription
1 Amide I -II 2D IR spectroscopy provides enhanced protein secondary structural sensitivity The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher DeFlores, Lauren P. et al. Amide III 2D IR Spectroscopy Provides Enhanced Protein Secondary Structural Sensitivity. Journal of the American Chemical Society (2009): American Chemical Society Version Author's final manuscript Accessed Tue Jan 02 16:01:37 EST 2018 Citable Link Terms of Use Detailed Terms Article is made available in accordance with the publisher's policy and may be subject to US copyright law. Please refer to the publisher's site for terms of use.
2 Amide I -II 2D IR spectroscopy provides enhanced protein secondary structural sensitivity Lauren P. DeFlores, Ziad Ganim, Rebecca A. Nicodemus, and Andrei Tokmakoff * Department of Chemistry, Massachusetts Institute of Technology, Cambridge, MA 02139, USA * Corresponding author: tokmakof@mit.edu Abstract We demonstrate how multi-mode 2D IR spectroscopy of the protein amide I and II vibrations can be used to distinguish protein secondary structure. Polarization-dependent amide I -II 2D IR experiments on poly-l-lysine in the -sheet, -helix, and random coil conformations show that a combination of amide I and II diagonal and cross peaks can effectively distinguish between secondary structural content where amide I infrared spectroscopy alone cannot. The enhanced sensitivity arises from frequency and amplitude correlations between amide II and amide I spectra that reflect the symmetry of secondary structures. 2D IR surfaces are used to parameterize an excitonic model for the amide I -II manifold suitable to predict protein amide I -II spectra. This model reveals that the dominant vibrational interaction contributing to this sensitivity is a combination of negative amide II -II through-bond coupling and amide I -II coupling within the peptide unit. The empirically determined amide II -II couplings do not significantly vary with secondary structure: -8.5 cm -1 for the sheet, -8.7 cm -1 for the helix, and -5 cm -1 for the coil. 1
3 Introduction Amide vibrational spectroscopy 1 is an important tool for characterizing protein structure and dynamics with picosecond time-resolution. 2 The amide vibrations are intrinsic to the protein backbone and have secondary structure sensitivity. Unlike traditional protein infrared spectroscopy, which has the practical limitation of congested lineshapes, emerging methods in two-dimensional infrared (2D IR) spectroscopy are providing substantially more detailed methods for analyzing the structural and dynamical information content of these vibrations. 1,3-11 2D IR spreads infrared transitions over two frequency axes and reveals cross peaks that quantify vibrational couplings and other correlations between vibrations. The combination of enhanced spectroscopic sensitivity and a new generation of structure-based amide spectroscopic models 3,12-15 is leading to a toolset that can be used for the quantitative study of time-dependent protein conformation. 16 While protein 2D IR spectroscopy has primarily targeted the structure sensitive amide I band, the use of multi-mode 2D IR spectroscopy that investigates crosscorrelations between different amide vibrations opens entirely new possibilities for enhancing structural information in protein vibrational spectra. In this paper, we report such an advance in which amide I -II 2D IR spectroscopy is used to give enhanced protein secondary structural sensitivity. The most studied protein vibration, amide I, is composed predominately of peptide carbonyl stretch and displays secondary structure sensitivity. Empirical frequency-structure correlations find that -sheets have a strong absorption band at cm -1 and a weaker band at cm -1. The -helix and random coil structure are located at cm -1 and cm -1, respectively. 17,18 The origin of these 2
4 band shifts is a strong coupling between individual amide I units which gives rise to delocalized or excitonic vibrational states that report on the symmetry and size of secondary structure 2,19. While anti-parallel sheets and aggregates give rise to distinct diagnostic amide I bands 8, other secondary structures are poorly resolved in an amide I line shape with multiple contributions. The -helical structure is predicted to have three IR active amide I modes, but their splittings are on the order of a few cm -1 and the features are not resolvable. The amide II vibration, due to its composition of NH bend and CN stretch, is predominately known for its sensitivity to the protonation state of the peptide unit. 20 Its 100 cm -1 frequency downshift between protonated (amide II) and deuterated (amide II ) forms can be used with kinetic H/D exchange experiments to probe water exposure of protein structure. 21 Despite early predictions of Miyazawa and Blout 22 that amide II would exhibit sensitivity similar to amide I, the amide II band is relatively unstructured and varies little with secondary structure. In the case of amide II, the congested nature of the linear IR spectrum, as well as its overlap with the HOD bend absorption, makes its use as a structural diagnostic impractical. This work uses 2D IR to resolve details of the 2D amide II lineshape and investigate the amide I -II cross-peaks in a model system that adopts different secondary structures, poly-l-lysine (PLL). 2D IR cross peaks are used to characterize secondary structure specific spectral features. It is found that the amide II 2D line shape gives rise to distinctive signatures for helices and random coil regions, which, when combined with the sheet sensitivity of the amide I region, allow for separation of all the common secondary structural motifs. Polarization dependent experiments are used to quantify 3
5 vibrational couplings and transition dipole orientations between amide II oscillators and between amide I and II, and also describe the symmetry of the modes involved. 2D IR spectra are interpreted with an excitonic model for the amide I -II manifold suitable to predict protein amide I -II FTIR and 2D IR spectra. The mechanism for vibrational coupling that leads to these structure-sensitive spectra is revealed to be a combination of through-bond coupling between amide II local modes in addition to a strong amide I -II coupling at a given peptide unit. This supports recent theoretical observations 13,23 and provides clear evidence against the widely made assumption that the amide vibrations are orthogonal and therefore energetically isolated. Methods and Materials Materials. All samples are purchased from Sigma-Aldrich (St. Louis, MO). Before use, samples were fully deuterated by hydrogen/deuterium exchange in D 2 O at 45 C for 1 hour, followed by lyophilization. Samples were held in a temperaturecontrolled cell with 1-mm thick calcium fluoride windows and a 50 m path length. Concentrations of the polypeptide forms are selected such that the infrared absorbance of the amide I band is <0.5; approximately 20 mg/ml. Poly-L-lysine samples are prepared under different conditions in order to generate the desired secondary structure motif. 24 The random coil species is prepared by dissolving the protein in neat D 2 O at 20 C and ph = The -helical conformation is prepared by dissolving PLL into neat D 2 O and raising the ph to 12 using 1M NaOD solution. The -sheet form of PLL is prepared by subsequently raising the temperature of the -helical solution to 46 C over 30 minutes. Spectra of the helical and sheet moieties were taken consecutively on identical samples. Protein samples were prepared in a 50 mm phosphate buffer. 4
6 2D IR Spectroscopy. 2D IR experiments are performed with bandwidth sufficient to cover the fundamental and overtone bands of amide I and amide II using a 2D FT spectrometer that is described in detail elsewhere. 26 The mid-infrared laser pulse was centered at 1550 cm -1 with a FWHM bandwidth of approximately 350 cm -1 and compressed to sub-80 fs in duration. 2D IR spectra are acquired in a two-beam geometry using a collinear pulse pair to pump and third pulse to serve as the probe and local oscillator. ZZYY spectra were collected by rotation of the probe polarization through a 90 turn periscope. The excitation dimension is obtained by step scanning the 1 time axis to 1.8 ps in steps of 2 fs giving a resolution of ~1 cm -1. The transmitted probe is dispersed in a monochromator using a 75-groove/mm grating and collected using a 64 pixel MCT array detector. The 1.9 cm -1 resolution in the 3 detection axis results in a 120 cm -1 single-scan detection bandwidth. 2D spectra are assembled from spectrograms centered at 1450 cm -1 and 1650 cm -1. Fourier transformation of the signal as a function of the timing between pump pulses ( 1 ) resolves the 2D spectrum along the 1 axis. Structure Based Modeling. Modeling of amide I -II spectra drew on idealized structures for an anti-parallel sheet, helix, and random coil. Molden 27 was used to generate an idealized anti-parallel sheet consisting of six strands with seven peptide units per strand by repeating =-139 and =135. The strands have co-planar C spaced 3.04 Å apart. A 20 residue helix was generated by repeating =-57 and =-47. Calculations for the random coil involved an average over 5000 snapshots of a GROMACS 28,29 molecular dynamics simulation of an 18-oscillator peptide with rigid bond lengths 30 and angles in vacuum, coupled to a 1000 C bath. 30 5
7 Analogous to previous work 12, amide I -II spectra are calculated by diagonalizing a local mode Hamiltonian developed in the basis of the amide I and II vibrations for the individual peptide units. The elements of this local amide Hamiltonian are assigned on the basis of the position of the peptide units within the structure. The amide II Hamiltonian parameters are experimentally constrained and appear in the Local Mode Modeling section. The amide I vibrations are characterized by anharmonically coupled oscillators with empirically chosen, degenerate site energies of 1675 cm -1 and anharmonicity of 16 cm Non-bonded amide I -I interactions were determined through transition dipole coupling. 31 The orientation of the amide I transition dipole was set 20 off the C=O bond, with the dipole center located Å along the carbonyl axis, 20 and a normalized unit dipole strength of 1.0. Spectral calculations involved a sum over transition pathways between eigenstates weighted by polarization factors including twoquantum states through a weak anharmonicity and harmonic scaling of the transition moments. 8 Calculation averaged over a static disordered system using a homogeneous linewidth (FWHM) of 14 cm -1. Results and Discussion Linear Spectroscopy of Amide I and II The FTIR spectra of the three forms of poly-l-lysine are shown in Fig. 1. Unlike the spectra of the single peptide unit in N-methylacetamide, 32 the amide I and amide II bands are structured. In random coil form, poly-l-lysine shows a small shoulder on the blue side of the amide I band maximum (1645 cm -1 ), and is broad and suggestive of overlapping resonances in the amide II band. In the -helical form, the amide I band is 6
8 red shifted and asymmetric. This lineshape is commonly interpreted as the overlapping contributions from the intense A band near 1630 cm -1 ( A ) and the weaker blue-shifted E bands ( E ). The amide II band shows a distinct two peak structure. As the temperature of the -helical form is increased, PLL undergoes a structural transition into a -sheet aggregate. In -sheet form, the amide I band splits into the signature high and low intensity -sheet modes. The 1610 cm -1 and 1690 cm -1 modes are termed and, respectively, to refer to whether the amide I oscillators of the mode vibrate in-phase perpendicular or parallel to the strands of the sheet. For the sheet, amide II does not exhibit such a drastic change, but rather loses structure. 2D IR Spectroscopy of Amide I and II The 2D IR spectra of PLL in the various secondary structure conditions are shown in Fig. 2 for parallel (ZZZZ) and perpendicular (ZZYY) polarized probing. Spectral regions of interest include the lineshapes of the amide I ( 1, 3 = cm -1 ) and II ( 1, 3 = cm -1 ) diagonal regions, and the amide I -II cross peaks. As expected from its low cross-section, no signal contributions from the HOD bend were observed in the amide II region. The secondary structural sensitivity revealed by describing the contours of the 2D lineshapes, which result from interference between positive (fundamental) and negative (overtone) transitions within the amide band. In addition, since the cross peak intensity scales quadratically with the dot product of the diagonal transition dipole moments, it encodes both the magnitude and the relative angle between the transition dipoles. Comparison of the cross peak amplitude as a function of polarization reveals the angle. 33 Qualitatively, the strong enhancement of a cross peak in 7
9 crossed polarization indicates a transition dipole projection angle close to orthogonal (>70º), while a decrease indicates angles closer to parallel. The 2D IR spectroscopy of the diagonal amide I line shape has been described previously. 1,34 The amide I region of Fig. 2a shows the characteristic 8-peak structure for extended -sheets, which arises from fundamental and overtone transitions of the and vibrations and cross-peaks between them. The amide I -I cross peaks are enhanced in the perpendicular spectra in Fig. 2b, and a comparison of the cross peak amplitudes in parallel and perpendicular spectra 35 gives an angle between their transition dipoles of I -I =67 7, consistent with our previous measurement. 1 Similar to amide I, the diagonal amide II region shows two peaks ( 1, 3 =1440 cm -1 and 1490 cm -1 ) and an enhancement of the cross peak between them in the perpendicular spectrum. Analysis of this cross peak amplitude variation is limited due to interference effects, but puts a constraint on their orientation II -II > 35. The amide I regions of the -helix and random coil 2D spectra are similar, and it is difficult to distinguish between the two moieties using only the amide I mode. The random coil lineshape is more diagonally elongated ( 1 = 3 ) than the helix. The amide I helix lineshape also shows slight off-diagonal broadening that is consistent with previous interpretations of two underlying A and E 1 symmetry modes However, examination of the diagonal amide II region of the parallel spectra in Fig. 2c and 2e reveals distinct differences; the random coil line shape is again diagonally elongated while the helix shows two peaks at 1470 cm -1 and 1440 cm -1. Surprisingly, both the helix and random coil show off-diagonal broadening in the perpendicular spectra. This arises from a plateau ( 3 =1480 cm -1 ) indicative of a cross-peak between two modes. The 8
10 presence of two peaks in the coil spectrum indicates that significant amide II vibrational couplings persist in this state. A clearer indication of the amide II structural sensitivity is seen in a projection of the perpendicular lineshape onto 3, shown in Fig. 3a. Two distinct peaks are observed for the sheet and coil conformations, and the helix resonance is peaked between them. The multimode structure present in the random coil spectrum is an indication that the amide II lineshape is reporting on local backbone configurational variation rather than the longer range structure of the entire chain. Amide I -II Cross Peaks Cross peaks between amide I and amide II allow the new spectral features of amide II to be assigned by correlation to understood amide I signatures. Both upward ( 1 < 3 ) and downward ( 1 > 3 ) cross peaks appear. The perpendicular -sheet spectrum shows cross peaks between both pairs of the amide I and II modes. The most intense cross-peak to amide I varies with polarization; inspection of the upward cross peaks along the slice 3 = 1611 cm -1 shows that in perpendicular polarization the more intense cross peak is to the 1447 cm -1 amide II band, while in the parallel polarization the cross peak to the 1470 cm -1 amide II band is more intense. This intensity flipping is visible, albeit more subtle, in the cross-peaks of the amide I band to amide II (along 3 = 1690 cm -1 ). The alternating intensity of the cross peaks between amide I and II indicates that the low frequency amide I (1611 cm -1 ) and the high frequency amide II (1470 cm -1 ) modes have aligned transition dipoles which are nearly orthogonal to those of the high frequency amide I (1690 cm -1 ) and low frequency amide II (1447 cm -1 ) modes. In the reference frame of the protein, if the transition dipole of the more intense 9
11 amide I mode is aligned perpendicular to the strands, then the transition dipole of the more intense amide II mode (1447 cm -1 ) is close to parallel with them. Therefore we assign the 1470 cm -1 band to the amide II mode and the 1447 cm -1 band to the amide II mode of the sheet. The amide I -II upward cross peak in -helical PLL is broad and shows an antidiagonal elongation. (Fig. 2d). This implies that the frequency shift of the amide II A and E modes are anti-correlated with the frequency shift in amide I. The dramatic intensity increase of the downward cross peak from parallel to perpendicular spectra (Fig. 2c to 2d) implies a near perpendicular projection angle between the dominant amide I and II transition dipoles of the helix. Since the A mode dominates the amide I spectrum, the amide II spectrum will be dominated by the helix mode with E 1 symmetry. Marsh and coworkers 39 tabulate the angles of amide I and II relative to the helix axis to be and 73-77, respectively. We can compare these results on amide II to our amide II measurements drawing on the previous characterization of dipole projection angles for NMA-h 7 and NMA-d These show a very small angle between amide I and I and a roughly 35 angle between amide II and II transition moments. From this, one would predict that the transition dipole of amide II lies between to the helix axis, giving a angle between amide I and II, which is consistent with the results observed here. Our results indicate that transition dipole of the dominant amide II band of -helical PLL lies nearly perpendicular to the helix axis. From these observations, we assign the 1470 cm -1 and 1440 cm -1 vibrations to the amide II E and A modes, respectively. In addition, the downward cross-peak of the helical PLL in both ZZZZ and 10
12 ZZYY 2D spectra show a weak negative transition. This is a distinguishing attribute relative to the random coil conformation. For the coil spectra, the presence of an antidiagonal tilt to the perpendicularly polarized cross peak region, the presence of two peaks in the amide II diagonal region, and a diagonally elongated amide I diagonal peak together indicate that there is are delocalized or excitonic amide I and II states in this species. Although the chain may be disordered, this should not be surprising, given the strength of amide I -I and amide II - II couplings between bonded peptide groups. While we expect a complex band structure to this feature, for preliminary assignment, we expect that the more intense red side of the amide II band (1460 cm -1 ) involves symmetric (+) motion between the CN stretches of bonded neighbors, whereas the weaker (1480 cm -1 ) has antisymmetric ( ) character. From the antidiagonal tilt of the amide I -II cross peak, we associate the red and blue sides of the amide I band (1650 and 1670 cm -1 ) with modes of + and character, respectively. A summary of the peak assignments to vibrations and the quantitative analysis of the projection angles between modes is presented in Table 1. These values of the projection angles and their constraints are determined using the ratio of peak amplitudes in parallel and perpendicular geometries assuming the rules for two coupled vibrations and that the ratio for the dominant diagonal peaks is 1/3. Interference effects reduce the ability to separate each of the possible angles, and typically lead to a lower bound on the projection angle. Of the PLL conformations, the sheet spectra provide the highest number of constraints. We note that the sheet projection angles cannot be easily reconciled with the constraint that they lay within a plane. This suggests that twisting of 11
13 the sheet and/or stacking of multiple sheets may be the origin of these angles. Additionally, we note that there is limited evidence that the sheet form of PLL is does indeed have antiparallel registry. It remains possible that the sheet for of PLL is an aggregate with the parallel in-register cross-beta structure seen in amyloid fibrils. The greatest complement that amide II adds to amide I 2D IR spectroscopy is the ability to separate -helix and random coil signatures. In addition to differences in the amide II 2D lineshape, the random coil amide I -II upward and downward cross peaks are both more anti-diagonally elongated and the downward cross peak shows a strong negative transition. All together, amide I -II 2D IR spectroscopy allows clear separation of sheets, helices, and random coil structural motifs. The enhanced secondary structure sensitivity of the amide I -II cross peak is more clearly seen as projections of the downward cross-peak lineshape onto the axis in Fig. 3b. These projections provide distinct spectral signatures for the three secondary structures. The sheet and coil both show a two-peak structure, but the more intense band is red shifted for the sheet and blue shifted for the coil. The helix is dominated by one resonance that lies between the others. As an indication of how amide I /II 2D IR spectra would be used in the analysis of proteins, we have acquired spectra for four proteins that vary in the secondary structure contest (Fig. 5). The presence of cross peaks in the amide I (~1650 cm - 1 )/II (~1450 cm -1 ) and amide I(~1650 cm -1 )/II(~1550 cm -1 ) regions of Concanavalin A, a -protein, and myoglobin, an -protein, indicate partial hydrogen-deuterium exchange. 21 Non-exchanging peptides within the Con A -sheets are seen by the vertically displaced doublet for the amide I/II cross peak. The random coil content has exchanged and is seen 12
14 as intensity between this doublet of the amide I /II region. Myoglobin shows similar features to partially exchanged helical content. The intensity of the downward cross peak overtone shows a direct signature of exchanged random coil. Ubiquitin and RNaseA, mixed -sheet and helical proteins, show full exchange under these conditions. Cross peaks show a super-position of all secondary structure motifs. Local Mode Modeling of Amide I -II Spectra To gain further insight into the origin of secondary structure sensitivity in amide I -II spectra and a detailed molecular assignment of the spectral features, we have constructed a structure-based model suitable for calculating amide I -II spectra of proteins. This modeling expands the local amide Hamiltonian (LAH) approach that has been successful in predicting amide I 2D IR spectra and providing frequency-structure correlations. 3,12 The amide I -II manifold of vibrations is described as a linear combination of unit amide I and II oscillators on each peptide group. The LAH is constructed in this basis and the elements are assigned using a structure. The diagonal elements (site energy) correspond to the intrinsic vibrational frequency of that isolated oscillator, and off-diagonal terms (coupling between oscillators) depend on their separation and orientation. Diagonalizing the LAH allows us to calculate FTIR and 2D IR spectra for a given structure. The amide I block of the local amide Hamiltonian (LAH) is constructed as previously described 31 (details appear above) and amide II is added with the minimum number of extra variables. This reflects two assumptions: (1) For simplicity we assume the amide II oscillators couple with each other only as a result of through-bond interactions with their neighbors in the chain. This is expected given the small transition 13
15 dipole moment and unfavorable position for through space amide II couplings. (2) Similarly, we assume that amide I and II vibrations only couple within the same peptide unit. This assumption is reasonable since the strength of the on-site coupling in NMA is far greater than all other interactions involving amide I and II 13,32. Parameters for the LAH are obtained from prior amide I -II 2D IR experiments on NMA-d 7 in D 2 O. 32 We set the amide II site energy to II = 1450 cm -1, the amide II vibrational anharmonicity to II = 10 cm -1, the amide I -II on-site coupling to V I -II = 39 cm -1, and the transition dipole ratio AmII / AmI = To match the random coil lineshape, the amide I and amide II site energies are sampled from a normal distribution with standard deviation of 10 cm -1 about the assigned I and II values. Amide II -II coupling between nearest neighbor sites were set with empirically determined values to V II -II = 8.5 cm -1 for the sheet, 8.7 cm -1 for the helix, and sampled from a normal distribution ( V II -II = 5 cm -1 and = 3 cm -1 ) for the random coil. These idealized structures are our best approximations to the unknown atomic structures of poly-l-lysine, which may contain nonregular, fibrillar structures. 40 The sign and magnitude of V II -II were constrained by the splitting and intensity of the experimental cross peaks; positive coupling can be ruled out as it leads to the wrong intensity ratio between the high and low frequency features in each structural motif. Previous theoretical work gives evidence in favor of strong through-bond amide II-II coupling, but is of limited comparison because it has not considered deuterated amide groups. 13,41 Linear and 2D IR spectra for the AP sheet, helix and random coil were calculated for parallel and perpendicular polarizations, and appear in Fig. 4. FTIR spectra 14
16 capture the two mode structure of amide II with different intensity variations for sheets and helices. General trends in the position and amplitude of diagonal and cross peaks, and the interference patterns they participate in are largely reproduced. Experimental and calculated 2D IR spectra are also compared using projections along 1 in Fig. 3. For the random coil spectra, lineshapes and intensities of positive and negative contributions are well-reproduced by the calculation, indicating that the two peak structure is not an indicator of a random contour to the chain, but of through-bond coupling that has weak dependence on conformation. The intensity of the low frequency amide II helical peak is exaggerated in the calculation and appears as only a shoulder in the experiment. The splitting and intensities of the sheet projections match the experiment reasonably well. All together these results explain the intensity variation between positive and negative lobes of the peaks without invoking electrical anharmonicity, but rather the result of interference. Agreement between the experiment and calculations demonstrates two points. First, only nearest neighbor coupling is required to reproduce the splitting between amide II vibrations. Second, the sign and magnitude of this nearest neighbor coupling can be unambiguously obtained from the intensity ratio and splitting observed in the amide I -II 2D IR spectra. Control calculations of FTIR and 2D IR spectra with and without coupling were used to confirm the origin of spectral features (not shown). Amide I -II coupling is required for amide I -II cross peaks, but not to yield the observed structure in the amide II lineshape for any of the three structural motifs. Conversely, introducing amide I -II coupling without amide II -II coupling never causes appreciable splitting in amide II. Therefore we conclude that amide II oscillators couple directly to neighboring amide II 15
17 oscillators; a coupling mechanism where amide II oscillators are indirectly coupled through mutual coupling to the amide I manifold is insufficient to reproduce any peak structure. Using the amide II eigenstates of the LAH, we have visualized the symmetry of the vibrations contributing to the spectral features using doorway mode analysis. This method identifies bright states that carry a majority of the oscillator strength within a restricted frequency region, and color-codes the amplitude and phase of the oscillators involved. 42,43 These doorway modes are found to have the symmetries expected for the secondary structure. For amide II, two bright states are found that best describe the low and high frequency -sheet modes: the amide II sites oscillate in-phase along the strands (, ) or across the strands (, ), respectively. For the helix, the bright states also mirror those found for amide I with A and E 1 symmetry modes with phases aligned parallel and perpendicular to the helix axis, respectively. The relative angles between transition dipole moments of the doorway modes are calculated for the amide I and II modes of the sheet and helix and appear in Table 2. In regions where the density of states is too low (< 5 states), a single intense eigenstate was selected. These results correlate very well to the experimentally observed cross peak ratios of parallel and perpendicular 2D IR spectra from Figure 1. As expected the strongest transitions of the amide I and II transitions are nearly orthogonal, with a calculated angle of 87 between the transition dipoles. Similarly, the dominant -helix mode of amide I runs along the axis while the dominant amide II is orthogonal. This is observed in both the calculated angles of the amide I A mode relative to the E 1 modes of 16
18 amide II of 72 and 92 and in the experimental polarization spectra. These results are consistent with those presented by Marsh and co-workers. 39 Conclusion We have demonstrated the increased protein secondary structural sensitivity that can be obtained through multi-mode 2D IR spectroscopy. To this point, the amide I vibrational mode of proteins has been predominantly used to provide molecular insight to the protein structure and dynamics. While 2D amide I spectra improve sensitivity over traditional FTIR, they still cannot readily separate helix and coil absorptions. Amide I -II 2D IR experiments on the idealized system of poly-l-lysine in the -sheet, -helix, and random coil conformations shows that a combination of amide I and II diagonal and cross peaks can more effectively distinguish the secondary structural content. As shown by polarization-selective measurements, this added sensitivity arises from the nearorthogonal transition dipole moments of amide II relative to amide I, allowing modes of differing symmetry to be correlated. Modeling of the amide I -II FTIR and 2D IR spectra provides insight into the molecular origin of the observed amide II peaks. We find that negative amide II -II couplings between bonded residues and strong on-site amide I -II couplings are the two critical features to describe 2D IR spectra. The empirically obtained V II -II do not vary significantly with secondary structure: -8.5 cm -1 for the sheet, -8.7 cm -1 for the helix, and -5 cm -1 for the coil are found. Through-space amide II couplings are not required to obtain the basic features of the spectra. These observations suggest that somewhat more sophisticated treatments that allow the site energies and couplings to vary 44 can be used to obtain amide I /II spectra of proteins using MD simulation. 12,45,46 17
19 The combination of 2D IR spectra and structure-based modeling extends the capabilities of 2D IR spectroscopy to interpret protein conformational changes. Transient 2D IR experiments can be used to directly follow conformational dynamics and kinetics and interpreted with the aid of MD simulation, as demonstrated recently on ubiquitin unfolding using amide I. 47 Further, these methods can be used in conjunction with H/D exchange experiments to reveal those secondary structures that are solvent exposed. 21 Acknowledgements This work was supported by the National Science Foundation (CHE ), the U.S. Department of Energy (DE-FG02-99ER14988), and the Petroleum Research Fund. R. A. N. thanks the DOD for an NDSEG fellowship. Supporting Information Complete amide I -II 2D IR spectra of PLL as a function of polarization including expanded diagonal regions. Similar plots are presented for calculated spectra. Also, visualizations of the amide I and II modes of protein secondary structures and their transition dipole orientations. 18
20 Figure. 1 FTIR spectra of poly-l-lysine as a function of ph and temperature showing the -sheet (ph=12,t=46 C), -helix (ph=12,t<20 C) and random coil (ph=4,t=20 C) form of the homopolymer. 19
21 Figure. 2 ZZZZ and ZZYY Amide I -II 2D IR spectra of poly-l-lysine as a function of ph and temperature of preparation showing the -sheet (a,b; ph=12,t=46 C), -helix (c,d; ph=12,t<20 C) and random coil (e,f; ph=4,t=20 C) forms. Contours are plotted from +/- 80% of the amide I normalized peak at a 5% spacing. For the -sheet conformation, contours are plotted at 1.25% between +/- 20% to enhance subtle features of the cross peak region. 20
22 Figure. 3 Comparison of the experimental (a,b) and calculated (c,d) ZZYY 2D IR projections. Projections of the sheet, helix and random coil of the amide II diagonal ( cm -1 ) and the amide I -II downward cross peak ( cm -1 ). Projections show identical band structures with inclusion of negative nearest neighbor couplings. 21
23 Figure 4 Calculated 2D IR spectra 22
24 Figure 5: ZZZZ and ZZYY Amide I -II 2D IR spectra of concanavalin A, myoglobin, ubiquitin and RNaseA. Contours are plotted from +/- 80% of the amide I normalized peak at a 4% spacing. In addition for concanavalin A, extra contours are plotted at 1.25% between +/- 20%. The contours of the amide I/II and I /II regions are bolded to enhance subtle features of the cross peak regions. 23
25 Amide II' Amide I' Amide I' Amide II' Amide I' Table 1. Experimentally determined projection angle between amide I and II transition dipole moments in poly-l-lysine for sheet, helix, and coil samples. Vibrational frequency assignments to mode symmetry are as discussed in text. and constraints on this value are obtained from the ratio of cross peak amplitudes for perpendicular and parallel polarization, using the rules for two coupled vibrations.[ref] Where destructive interference effects limit an accurate determination on angles, the lower bound is given. sheet Amide II' Amide I' 1447 cm cm cm cm > cm -1 > cm -1 > p 1447 cm helix Amide II' A E cm cm -1 E 1 > cm -1 * 62 5? A cm -1 * 90 10? *The dominance of the amide I A mode and amide II E 1 modes limits the ability to determine these angles. Random Coil Amide II' (+) (-) 1440 cm cm -1 (+) 1650 cm -1 > (-) 1645 cm >56 (+) 1470 cm -1 >
26 Amide I' Table 2. Calculated orientation of the amide I and II transition dipole moments for idealized secondary structures. defines the angles of the transition dipole relative to the dominant axis of the secondary structure: Along the -strands for -sheet and along the -helix axis. For each mode, the central frequency of the doorway mode or eigenstate is given. Bolded angles represent modes with the strongest cross peaks observed in the perpendicular 2D IR spectra. Amide II' p perp A E 1 E cm cm cm cm cm ll 1680 cm perp 1640 cm A 1645 cm E cm E cm
27 References (1) Demirdöven, N.; Cheatum, C. M.; Chung, H. S.; Khalil, M.; Knoester, J.; Tokmakoff, A. J. Am. Chem. Soc. 2004, 126, (2) Barth, A.; Zscherp, C. Q. Rev. Biophys 2002, 35, (3) Hamm, P.; Lim, M.; Hochstrasser, R. M. J. Phys. Chem. B 1998, 102, (4) Woutersen, S.; Hamm, P. J. Phys.: Condens. Mat. 2002, 14, (5) Zanni, M. T.; Hochstrasser, R. M. Curr. Opin. Struct. Biol. 2001, 11, (6) Scheurer, C.; Piryatinski, A.; Mukamel, S. J. Am. Chem. Soc. 2001, 123, (7) DeCamp, M. F.; DeFlores, L. P.; McCracken, J. M.; Tokmakoff, A.; Kwac, K.; Cho, M. J. Phys. Chem. B 2005, 109, (8) Cheatum, C. M.; Tokmakoff, A.; Knoester, J. J. Chem. Phys. 2004, 120, (9) Chung, H. S.; Khalil, M.; Tokmakoff, A. J. Phys. Chem. B 2004, 108, (10) Chung, H. S.; Khalil, M.; Smith, A. W.; Ganim, Z.; Tokmakoff, A. Proc. Natl. Acad. Sci. USA 2005, 102, (11) Chung, H. S.; Tokmakoff, A. J. Phys. Chem. B 2005, in press. (12) Ganim, Z.; Tokmakoff, A. Biophys J. 2006, 91, (13) Hayashi, T.; Mukamel, S. J. Phys. Chem. B. 2007, 111, (14) Jansen, T. l. C.; Knoester, J. J. Phys. Chem. B 2006, 110, (15) Choi, J.-H.; Lee, H.; Lee, K.-K.; Hahn, S.; Cho, M. J. Chem. Phys. 2007, 126, (16) Chung, H. S.; Khalil, M.; Smith, A. W.; Tokmakoff, A. Rev. Sci. Inst. 2007, 78. (17) Jackson, M.; Mantsch, H. H. Crit. Rev. Biochem. Mol. 1995, 30, (18) Byler, D. M.; Susi, H. Biopolymers 1986, 25, (19) Ganim, Z.; Chung, H. S.; Smith, A. W.; DeFlores, L. P.; Jones, K. C.; Tokmakoff, A. Acc. Chem. Res. 2008, 31, (20) Krimm, S.; Bandekar, J. Adv. Protein Chem. 1986, 38, (21) DeFlores, L. P.; Tokmakoff, A. J. Am. Chem. Soc. 2006, 128, (22) Miyazawa, T.; Blout, E. R. J. Am. Chem. Soc. 1961, 83, (23) Dijkstra, A.; Jansen, T. l. C.; Bloem, R.; Knoester, J. J. Chem. Phys. 2007, 127, (24) Rosenheck, K.; Doty, P. Proc. Nat. Acad. Sci. 1961, 47, (25) Davidson, B.; Fasman, G. D. Biochemistry 1967, 6, (26) DeFlores, L.; Nicodemus, R. A.; Tokmakoff, A. Opt. Lett. 2007, 32,
28 (27) G.Schaftenaar; Noordik, J. H. J. Comput.-Aided Mol. Design 2000, 14, (28) Lindahl, E.; Hess, B.; Spoel, D. v. d. J. Mol. Mod. 2001, 7, (29) Berendsen, H. J. C.; Spoel, D. v. d.; Drunen, R. v. Comp. Phys. Comm. 1995, 91, (30) Hess, B.; Bekker, H.; Berendsen, H. J. C.; Fraaije, J. G. E. M. J. Comp. Chem. 1997, 18, (31) Torii, H.; Tasumi, M. J. Chem. Phys. 1992, 96, (32) DeFlores, L. P.; Ganim, Z.; Ackley, S. F.; Chung, H. S.; Tokmakoff, A. J. Phys. Chem. B 2006, 110, (33) Golonzka, O.; Tokmakoff, A. J. Chem. Phys. 2001, 115, (34) Ganim, Z.; Chung, H. S.; Smith, A.; DeFlores, L.; Jones, K. C.; Tokmakoff, A. Acc. Chem. Res (35) Khalil, M.; Demirdoven, N.; Tokmakoff, A. J. Phys. Chem. A 2003, 107, (36) Woutersen, S.; Hamm, P. J. Chem. Phys. 2001, 115, (37) Fang, C.; Wang, J.; Charnley, A. K.; Barber-Armstrong, W.; Smith, A. B.; Decatur, S. M.; Hochstrasser, R. M. Chem. Phys. Lett. 2003, 382, (38) Ham, S.; Hahn, S.; Lee, C.; Kim, T.-K.; Kwak, K.; Cho, M. J. Phys. Chem. B 2004, 108, (39) Marsh, D.; Muller, M.; Schmitt, F. Biophys J. 2000, 78, (40) Fändrich, M.; Dobson, C. M. EMBO J. 2002, 21, (41) Moran, A.; Mukamel, S. Proc. Natl. Acad. Sci. USA 2004, 101, (42) Chung, H. S.; Tokmakoff, A. Journal of Physical Chemistry B 2006, 110, (43) Torii, H.; Tasumi, M. J. Chem. Phys. 1992, 97, (44) Bloem, R.; Dijkstra, A. D.; Jansen, T. l. C.; Knoester, J. J. Chem. Phys. 2008, 129, (45) Ham, S.; Cho, M. J. Chem. Phys. 2003, 118, (46) Jansen, T. l. C.; Knoester, J. J. Chem. Phys. 2006, in press. (47) Chung, H. S.; Ganim, Z.; Jones, K. C.; Tokmakoff, A. Proc. Nat. Acad. Sci. 2007, 104,
5.74 Introductory Quantum Mechanics II
 MIT OpenCourseWare http://ocw.mit.edu 5.74 Introductory Quantum Mechanics II Spring 009 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms. Andrei Tokmakoff,
MIT OpenCourseWare http://ocw.mit.edu 5.74 Introductory Quantum Mechanics II Spring 009 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms. Andrei Tokmakoff,
Correlation spectroscopy
 1 TWO-DIMENSIONAL SPECTROSCOPY Correlation spectroscopy What is two-dimensional spectroscopy? This is a method that will describe the underlying correlations between two spectral features. Our examination
1 TWO-DIMENSIONAL SPECTROSCOPY Correlation spectroscopy What is two-dimensional spectroscopy? This is a method that will describe the underlying correlations between two spectral features. Our examination
University of Groningen
 University of Groningen Two-dimensional infrared spectroscopy of antiparallel beta-sheet secondary structure Demirdoven, N; Cheatum, CM; Chung, HS; Khalil, M; Knoester, Jasper; Tokmakoff, A; Demirdöven,
University of Groningen Two-dimensional infrared spectroscopy of antiparallel beta-sheet secondary structure Demirdoven, N; Cheatum, CM; Chung, HS; Khalil, M; Knoester, Jasper; Tokmakoff, A; Demirdöven,
Secondary structural characteristics of proteins may be studied
 The origin of vibrational mode couplings in various secondary structural motifs of polypeptides Andrew Moran* and Shaul Mukamel *Department of Chemistry, Northwestern University, 2145 Sheridan Road, Evanston,
The origin of vibrational mode couplings in various secondary structural motifs of polypeptides Andrew Moran* and Shaul Mukamel *Department of Chemistry, Northwestern University, 2145 Sheridan Road, Evanston,
Anharmonic Vibrational Modes of Nucleic Acid Bases Revealed by 2D IR Spectroscopy
 Anharmonic Vibrational Modes of Nucleic Acid Bases Revealed by 2D IR Spectroscopy The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters.
Anharmonic Vibrational Modes of Nucleic Acid Bases Revealed by 2D IR Spectroscopy The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters.
Additional information on J. Chem. Phys. Maksim Grechko and Martin T. Zanni
 Quantification of transition dipole strengths using 1D and D spectroscopy for the identification of molecular structures via exciton delocalization: Application to α-helices Maksim Grechko and Martin T.
Quantification of transition dipole strengths using 1D and D spectroscopy for the identification of molecular structures via exciton delocalization: Application to α-helices Maksim Grechko and Martin T.
11.6. TWO-DIMENSIONAL CORRELATION SPECTROSCOPY
 Andrei Tokmakoff, MIT Department of Chemistry, 6/5/009 p. -54.6. TWO-DIMENSIONAL CORRELATION SPECTROSCOPY Our examination of pump-probe experiments indicates that the third-order response reports on the
Andrei Tokmakoff, MIT Department of Chemistry, 6/5/009 p. -54.6. TWO-DIMENSIONAL CORRELATION SPECTROSCOPY Our examination of pump-probe experiments indicates that the third-order response reports on the
Journal of Molecular Liquids
 Journal of Molecular Liquids 141 (2008) 149 154 Contents lists available at ScienceDirect Journal of Molecular Liquids journal homepage: www.elsevier.com/locate/molliq Two-dimensional vibrational lineshapes
Journal of Molecular Liquids 141 (2008) 149 154 Contents lists available at ScienceDirect Journal of Molecular Liquids journal homepage: www.elsevier.com/locate/molliq Two-dimensional vibrational lineshapes
SUPPLEMENTARY INFORMATION An Empirical IR Frequency Map for Ester C=O Stretching Vibrations
 SUPPLEMENTARY INFORMATION An Empirical IR Frequency Map for Ester C=O Stretching Vibrations Sean C. Edington, Jennifer C. Flanagan, Carlos R. Baiz* Department of Chemistry, University of Texas at Austin
SUPPLEMENTARY INFORMATION An Empirical IR Frequency Map for Ester C=O Stretching Vibrations Sean C. Edington, Jennifer C. Flanagan, Carlos R. Baiz* Department of Chemistry, University of Texas at Austin
THE JOURNAL OF CHEMICAL PHYSICS 125,
 THE JOURNAL OF CHEMICAL PHYSICS 125, 194510 2006 Correlated hydrogen bonding fluctuations and vibrational cross peaks in N-methyl acetamide: Simulation based on a complete electrostatic density functional
THE JOURNAL OF CHEMICAL PHYSICS 125, 194510 2006 Correlated hydrogen bonding fluctuations and vibrational cross peaks in N-methyl acetamide: Simulation based on a complete electrostatic density functional
Identification of Lactam-Lactim Tautomers of Aromatic. Heterocycles in Aqueous Solution Using 2D IR Spectroscopy
 Identification of Lactam-Lactim Tautomers of Aromatic Heterocycles in Aqueous Solution Using 2D IR Spectroscopy Chunte Sam Peng and Andrei Tokmakoff* Department of Chemistry, Massachusetts Institute of
Identification of Lactam-Lactim Tautomers of Aromatic Heterocycles in Aqueous Solution Using 2D IR Spectroscopy Chunte Sam Peng and Andrei Tokmakoff* Department of Chemistry, Massachusetts Institute of
University of Groningen
 University of Groningen Signatures of beta-sheet secondary structures in linear and two-dimensional infrared spectroscopy Cheatum, Christopher M.; Tokmakoff, Andrei; Knoester, Jasper Published in: Journal
University of Groningen Signatures of beta-sheet secondary structures in linear and two-dimensional infrared spectroscopy Cheatum, Christopher M.; Tokmakoff, Andrei; Knoester, Jasper Published in: Journal
Page 1. Introduction to Protein 2D IR Spectroscopy INTRODUCTION
 Introduction to Protein 2D IR Spectroscopy Carlos R. Baiz, Mike Reppert, and Andrei Tokmakoff Massachusetts Institute of Technology, Cambridge Massachusetts 02139 June 1, 2012 INTRODUCTION Proteins are
Introduction to Protein 2D IR Spectroscopy Carlos R. Baiz, Mike Reppert, and Andrei Tokmakoff Massachusetts Institute of Technology, Cambridge Massachusetts 02139 June 1, 2012 INTRODUCTION Proteins are
Chiral Sum Frequency Generation for In Situ Probing Proton Exchange in Antiparallel β-sheets at Interfaces
 Supporting Information for Chiral Sum Freuency Generation for In Situ Probing Proton Exchange in Antiparallel β-sheets at Interfaces Li Fu, Deuan Xiao, Zhuguang Wang, Victor S. Batista *, and Elsa C. Y.
Supporting Information for Chiral Sum Freuency Generation for In Situ Probing Proton Exchange in Antiparallel β-sheets at Interfaces Li Fu, Deuan Xiao, Zhuguang Wang, Victor S. Batista *, and Elsa C. Y.
University of Groningen
 University of Groningen Simulation of vibrational energy transfer in two-dimensional infrared spectroscopy of amide I and amide II modes in solution Bloem, Robbert; Dijkstra, Arend G.; Jansen, Thomas L.
University of Groningen Simulation of vibrational energy transfer in two-dimensional infrared spectroscopy of amide I and amide II modes in solution Bloem, Robbert; Dijkstra, Arend G.; Jansen, Thomas L.
Collective oscillations and the linear and two-dimensional infrared spectra of inhomogeneous beta-sheets Dijkstra, AG; Knoester, Jasper
 University of Groningen Collective oscillations and the linear and two-dimensional infrared spectra of inhomogeneous beta-sheets Dijkstra, AG; Knoester, Jasper Published in: Journal of Physical Chemistry
University of Groningen Collective oscillations and the linear and two-dimensional infrared spectra of inhomogeneous beta-sheets Dijkstra, AG; Knoester, Jasper Published in: Journal of Physical Chemistry
TRANSIENT 2D-IR SPECTROSCOPY: TOWARDS MEASURING ULTRAFAST STRUCTURAL DYNAMICS
 TRANSIENT 2DIR SPECTROSCOPY: TOWARDS MEASURING ULTRAFAST STRUCTURAL DYNAMICS JENS BREDENBECK 1, JAN HELBING 1, JOSEF WACHTVEITL 2 AND PETER HAMM 1 The technique of transient twodimensional infrared (T2DIR)
TRANSIENT 2DIR SPECTROSCOPY: TOWARDS MEASURING ULTRAFAST STRUCTURAL DYNAMICS JENS BREDENBECK 1, JAN HELBING 1, JOSEF WACHTVEITL 2 AND PETER HAMM 1 The technique of transient twodimensional infrared (T2DIR)
The two-dimensional IR nonlinear spectroscopy of a cyclic penta-peptide in relation to its three-dimensional structure
 Proc. Natl. Acad. Sci. USA Vol. 96, pp. 2036 2041, March 1999 Biophysics The two-dimensional IR nonlinear spectroscopy of a cyclic penta-peptide in relation to its three-dimensional structure PETER HAMM,
Proc. Natl. Acad. Sci. USA Vol. 96, pp. 2036 2041, March 1999 Biophysics The two-dimensional IR nonlinear spectroscopy of a cyclic penta-peptide in relation to its three-dimensional structure PETER HAMM,
Supporting Information
 Supporting Information C-terminal truncated -synuclein fibrils contain strongly twisted β-sheets Aditya Iyer, Steven J. Roeters ǂ, Vladimir Kogan Ί, Sander Woutersen *ǂ, Mireille M.A.E Claessens * and
Supporting Information C-terminal truncated -synuclein fibrils contain strongly twisted β-sheets Aditya Iyer, Steven J. Roeters ǂ, Vladimir Kogan Ί, Sander Woutersen *ǂ, Mireille M.A.E Claessens * and
Physical principles of IR and Raman. Infrared Spectroscopy
 Physical principles of IR and Raman IR results from the absorption of energy by vibrating chemical bonds. Raman scattering results from the same types of transitions, but the selection rules are different
Physical principles of IR and Raman IR results from the absorption of energy by vibrating chemical bonds. Raman scattering results from the same types of transitions, but the selection rules are different
Probing protein structure and folding pathways is one of the
 Dissecting coherent vibrational spectra of small proteins into secondary structural elements by sensitivity analysis Wei Zhuang, Darius Abramavicius, and Shaul Mukamel* Department of Chemistry, University
Dissecting coherent vibrational spectra of small proteins into secondary structural elements by sensitivity analysis Wei Zhuang, Darius Abramavicius, and Shaul Mukamel* Department of Chemistry, University
Accidental vibrational degeneracy in vibrational excited states observed with ultrafast two-dimensional IR vibrational echo spectroscopy
 THE JOURNAL OF CHEMICAL PHYSICS 13, 164301 005 Accidental vibrational degeneracy in vibrational excited states observed with ultrafast two-dimensional IR vibrational echo spectroscopy Junrong Zheng, Kyungwon
THE JOURNAL OF CHEMICAL PHYSICS 13, 164301 005 Accidental vibrational degeneracy in vibrational excited states observed with ultrafast two-dimensional IR vibrational echo spectroscopy Junrong Zheng, Kyungwon
Supporting Information
 1 Supporting Information 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 Materials and Methods Experiment In this study, an alloy IR probe which allowed us to get access to spectral
1 Supporting Information 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 Materials and Methods Experiment In this study, an alloy IR probe which allowed us to get access to spectral
Computing Infrared Spectra of Proteins Using the Exciton Model
 FULL PAPER WWW.C-CHEM.ORG Computing Infrared Spectra of Proteins Using the Exciton Model Fouad S. Husseini, [a] David Robinson, [a] Neil T. Hunt, [b] Anthony W. Parker, [c] and Jonathan D. Hirst* [a] The
FULL PAPER WWW.C-CHEM.ORG Computing Infrared Spectra of Proteins Using the Exciton Model Fouad S. Husseini, [a] David Robinson, [a] Neil T. Hunt, [b] Anthony W. Parker, [c] and Jonathan D. Hirst* [a] The
Infrared analogs of heteronuclear nuclear magnetic resonance coherence transfer experiments in peptides
 JOURNAL OF CHEMICAL PHYSICS VOLUME 116, NUMBER 15 15 APRIL 2002 Infrared analogs of heteronuclear nuclear magnetic resonance coherence transfer experiments in peptides C. Scheurer a) and S. Mukamel b)
JOURNAL OF CHEMICAL PHYSICS VOLUME 116, NUMBER 15 15 APRIL 2002 Infrared analogs of heteronuclear nuclear magnetic resonance coherence transfer experiments in peptides C. Scheurer a) and S. Mukamel b)
Supplementary Figures
 Supplementary Figures iso ( =2900 cm -1 ) 1.0 0.8 0.6 0.4 0.2 0.0-0.2-0.4 pump cm -1 3450 cm -1 cm -1 cm -1-0.5 0.0 0.5 1.0 1.5 2.0 2.5 delay [ps] Supplementary Figure 1: Raw infrared pump-probe traces.
Supplementary Figures iso ( =2900 cm -1 ) 1.0 0.8 0.6 0.4 0.2 0.0-0.2-0.4 pump cm -1 3450 cm -1 cm -1 cm -1-0.5 0.0 0.5 1.0 1.5 2.0 2.5 delay [ps] Supplementary Figure 1: Raw infrared pump-probe traces.
1 Introduction FOURIER TRANSFORM INFRARED STUDIES IN SOLID EGG WHITE LYSOZYME. 1C/94/403 INTERNAL REPORT (Limited Distribution)
 1C/94/403 INTERNAL REPORT (Limited Distribution) International Atomic Energy Agency and Nations Educational Scientific and Cultural Organization INTERNATIONAL CENTRE FOR THEORETICAL PHYSICS 1 Introduction
1C/94/403 INTERNAL REPORT (Limited Distribution) International Atomic Energy Agency and Nations Educational Scientific and Cultural Organization INTERNATIONAL CENTRE FOR THEORETICAL PHYSICS 1 Introduction
Two-dimensional infrared spectroscopy displays signatures of structural odering in peptide aggregates
 Haverford College Haverford Scholarship Faculty Publications Chemistry 2006 Two-dimensional infrared spectroscopy displays signatures of structural odering in peptide aggregates Casey H. Londergan Haverford
Haverford College Haverford Scholarship Faculty Publications Chemistry 2006 Two-dimensional infrared spectroscopy displays signatures of structural odering in peptide aggregates Casey H. Londergan Haverford
Useful background reading
 Overview of lecture * General comment on peptide bond * Discussion of backbone dihedral angles * Discussion of Ramachandran plots * Description of helix types. * Description of structures * NMR patterns
Overview of lecture * General comment on peptide bond * Discussion of backbone dihedral angles * Discussion of Ramachandran plots * Description of helix types. * Description of structures * NMR patterns
I690/B680 Structural Bioinformatics Spring Protein Structure Determination by NMR Spectroscopy
 I690/B680 Structural Bioinformatics Spring 2006 Protein Structure Determination by NMR Spectroscopy Suggested Reading (1) Van Holde, Johnson, Ho. Principles of Physical Biochemistry, 2 nd Ed., Prentice
I690/B680 Structural Bioinformatics Spring 2006 Protein Structure Determination by NMR Spectroscopy Suggested Reading (1) Van Holde, Johnson, Ho. Principles of Physical Biochemistry, 2 nd Ed., Prentice
Since the first structures of proteins became available, it was
 Picosecond dynamics of a membrane protein revealed by 2D IR Prabuddha Mukherjee, Itamar Kass, Isaiah Arkin, and Martin T. Zanni Department of Chemistry, University of Wisconsin, Madison, WI 53706-1396;
Picosecond dynamics of a membrane protein revealed by 2D IR Prabuddha Mukherjee, Itamar Kass, Isaiah Arkin, and Martin T. Zanni Department of Chemistry, University of Wisconsin, Madison, WI 53706-1396;
ion mobility spectrometry IR spectroscopy
 Debasmita Gho 29.10.2016 Introducti on Owing to its accuracy, sensitivity, and speed, mass spectrometry (MS) coupled to fragmentation techniques is the method of choice for determining the primary structure
Debasmita Gho 29.10.2016 Introducti on Owing to its accuracy, sensitivity, and speed, mass spectrometry (MS) coupled to fragmentation techniques is the method of choice for determining the primary structure
Supporting Materials
 Supporting Materials Figure S1 Experimental Setup Page Figure S (a) (b) (c) Feynman Diagrams Page 3-6 Figure S3 D IR Spectra Page 7 Figure S4 Kinetic Model Page 8 Figure S5 Van t Hoff Plots Page 9 1 k
Supporting Materials Figure S1 Experimental Setup Page Figure S (a) (b) (c) Feynman Diagrams Page 3-6 Figure S3 D IR Spectra Page 7 Figure S4 Kinetic Model Page 8 Figure S5 Van t Hoff Plots Page 9 1 k
Use of Periodic Boundary Conditions to Calculate Accurate β-sheet Frequencies Using Density Functional Theory
 Use of Periodic Boundary Conditions to Calculate Accurate β-sheet Frequencies Using Density Functional Theory Stefan Franzen Department of Chemistry North Carolina State University Raleigh, NC 27695 Abstract
Use of Periodic Boundary Conditions to Calculate Accurate β-sheet Frequencies Using Density Functional Theory Stefan Franzen Department of Chemistry North Carolina State University Raleigh, NC 27695 Abstract
FEMTOSECOND MID-INFRARED SPECTROSCOPY OF HYDROGEN-BONDED LIQUIDS
 Laser Chem., 1999, Vol. 19, pp. 83-90 Reprints available directly from the publisher Photocopying permitted by license only (C) 1999 OPA (Overseas Publishers Association) N.V. Published by license under
Laser Chem., 1999, Vol. 19, pp. 83-90 Reprints available directly from the publisher Photocopying permitted by license only (C) 1999 OPA (Overseas Publishers Association) N.V. Published by license under
SUPPLEMENTARY INFORMATION
 Hydrogen bonding at the water surface revealed by isotopic dilution spectroscopy Igor V. Stiopkin, 1,2 Champika Weeraman, 1,3 Piotr A. Pieniazek, 4 Fadel Y. Shalhout, 1,5 James L. Skinner, 4 and Alexander
Hydrogen bonding at the water surface revealed by isotopic dilution spectroscopy Igor V. Stiopkin, 1,2 Champika Weeraman, 1,3 Piotr A. Pieniazek, 4 Fadel Y. Shalhout, 1,5 James L. Skinner, 4 and Alexander
THE TANGO ALGORITHM: SECONDARY STRUCTURE PROPENSITIES, STATISTICAL MECHANICS APPROXIMATION
 THE TANGO ALGORITHM: SECONDARY STRUCTURE PROPENSITIES, STATISTICAL MECHANICS APPROXIMATION AND CALIBRATION Calculation of turn and beta intrinsic propensities. A statistical analysis of a protein structure
THE TANGO ALGORITHM: SECONDARY STRUCTURE PROPENSITIES, STATISTICAL MECHANICS APPROXIMATION AND CALIBRATION Calculation of turn and beta intrinsic propensities. A statistical analysis of a protein structure
Matthias Lütgens, Frank Friedriszik, and Stefan Lochbrunner* 1 Concentration dependent CARS and Raman spectra of acetic acid in carbon tetrachloride
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2014 SUPPORTING INFORMATION Direct observation of the cyclic dimer in liquid acetic
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2014 SUPPORTING INFORMATION Direct observation of the cyclic dimer in liquid acetic
2. Infrared spectroscopy
 2. Infrared spectroscopy 2-1Theoretical principles An important tool of the organic chemist is Infrared Spectroscopy, or IR. IR spectra are acquired on a special instrument, called an IR spectrometer.
2. Infrared spectroscopy 2-1Theoretical principles An important tool of the organic chemist is Infrared Spectroscopy, or IR. IR spectra are acquired on a special instrument, called an IR spectrometer.
( ) x10 8 m. The energy in a mole of 400 nm photons is calculated by: ' & sec( ) ( & % ) 6.022x10 23 photons' E = h! = hc & 6.
 Introduction to Spectroscopy Spectroscopic techniques are widely used to detect molecules, to measure the concentration of a species in solution, and to determine molecular structure. For proteins, most
Introduction to Spectroscopy Spectroscopic techniques are widely used to detect molecules, to measure the concentration of a species in solution, and to determine molecular structure. For proteins, most
Multi-Dimensional IR Spectroscopy of Acetic Acid Dimers and Liquid Water
 Multi-Dimensional IR Spectroscopy of Acetic Acid Dimers and Liquid Water N. Huse 1, J. Dreyer 1, E.T.J.Nibbering 1, T. Elsaesser 1 B.D. Bruner 2, M.L. Cowan 2, J.R. Dwyer 2, B. Chugh 2, R.J.D. Miller 2
Multi-Dimensional IR Spectroscopy of Acetic Acid Dimers and Liquid Water N. Huse 1, J. Dreyer 1, E.T.J.Nibbering 1, T. Elsaesser 1 B.D. Bruner 2, M.L. Cowan 2, J.R. Dwyer 2, B. Chugh 2, R.J.D. Miller 2
Molecular dynamics simulations of anti-aggregation effect of ibuprofen. Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov
 Biophysical Journal, Volume 98 Supporting Material Molecular dynamics simulations of anti-aggregation effect of ibuprofen Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov Supplemental
Biophysical Journal, Volume 98 Supporting Material Molecular dynamics simulations of anti-aggregation effect of ibuprofen Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov Supplemental
THE VIBRATIONAL SPECTRA OF A POLYATOMIC MOLECULE (Revised 3/27/2006)
 THE VIBRATIONAL SPECTRA OF A POLYATOMIC MOLECULE (Revised 3/27/2006) 1) INTRODUCTION The vibrational motion of a molecule is quantized and the resulting energy level spacings give rise to transitions in
THE VIBRATIONAL SPECTRA OF A POLYATOMIC MOLECULE (Revised 3/27/2006) 1) INTRODUCTION The vibrational motion of a molecule is quantized and the resulting energy level spacings give rise to transitions in
Supporting Information for
 Supporting Information for Molecular Rectification in Conjugated Block Copolymer Photovoltaics Christopher Grieco 1, Melissa P. Aplan 2, Adam Rimshaw 1, Youngmin Lee 2, Thinh P. Le 2, Wenlin Zhang 2, Qing
Supporting Information for Molecular Rectification in Conjugated Block Copolymer Photovoltaics Christopher Grieco 1, Melissa P. Aplan 2, Adam Rimshaw 1, Youngmin Lee 2, Thinh P. Le 2, Wenlin Zhang 2, Qing
(002)(110) (004)(220) (222) (112) (211) (202) (200) * * 2θ (degree)
 Supplementary Figures. (002)(110) Tetragonal I4/mcm Intensity (a.u) (004)(220) 10 (112) (211) (202) 20 Supplementary Figure 1. X-ray diffraction (XRD) pattern of the sample. The XRD characterization indicates
Supplementary Figures. (002)(110) Tetragonal I4/mcm Intensity (a.u) (004)(220) 10 (112) (211) (202) 20 Supplementary Figure 1. X-ray diffraction (XRD) pattern of the sample. The XRD characterization indicates
Using ultrafast infrared multidimensional correlation spectroscopy to aid in vibrational spectral peak assignments
 Chemical Physics Letters 381 (2003) 139 146 www.elsevier.com/locate/cplett Using ultrafast infrared multidimensional correlation spectroscopy to aid in vibrational spectral peak assignments John B. Asbury,
Chemical Physics Letters 381 (2003) 139 146 www.elsevier.com/locate/cplett Using ultrafast infrared multidimensional correlation spectroscopy to aid in vibrational spectral peak assignments John B. Asbury,
Amide vibrations are delocalized across the hydrophobic interface of a transmembrane helix dimer
 Amide vibrations are delocalized across the hydrophobic interface of a transmembrane helix dimer Chong Fang, Alessandro Senes, Lidia Cristian, William F. DeGrado, and Robin M. Hochstrasser Department of
Amide vibrations are delocalized across the hydrophobic interface of a transmembrane helix dimer Chong Fang, Alessandro Senes, Lidia Cristian, William F. DeGrado, and Robin M. Hochstrasser Department of
Timescales of Protein Dynamics
 Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Summary of 1D Experiment time domain data Fourier Transform (FT) frequency domain data or Transverse Relaxation Ensemble of Nuclear
Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Summary of 1D Experiment time domain data Fourier Transform (FT) frequency domain data or Transverse Relaxation Ensemble of Nuclear
Timescales of Protein Dynamics
 Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Dynamics from NMR Show spies Amide Nitrogen Spies Report On Conformational Dynamics Amide Hydrogen Transverse Relaxation Ensemble
Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Dynamics from NMR Show spies Amide Nitrogen Spies Report On Conformational Dynamics Amide Hydrogen Transverse Relaxation Ensemble
PRINCIPLES OF NONLINEAR OPTICAL SPECTROSCOPY
 PRINCIPLES OF NONLINEAR OPTICAL SPECTROSCOPY Shaul Mukamel University of Rochester Rochester, New York New York Oxford OXFORD UNIVERSITY PRESS 1995 Contents 1. Introduction 3 Linear versus Nonlinear Spectroscopy
PRINCIPLES OF NONLINEAR OPTICAL SPECTROSCOPY Shaul Mukamel University of Rochester Rochester, New York New York Oxford OXFORD UNIVERSITY PRESS 1995 Contents 1. Introduction 3 Linear versus Nonlinear Spectroscopy
Many proteins spontaneously refold into native form in vitro with high fidelity and high speed.
 Macromolecular Processes 20. Protein Folding Composed of 50 500 amino acids linked in 1D sequence by the polypeptide backbone The amino acid physical and chemical properties of the 20 amino acids dictate
Macromolecular Processes 20. Protein Folding Composed of 50 500 amino acids linked in 1D sequence by the polypeptide backbone The amino acid physical and chemical properties of the 20 amino acids dictate
Chemistry 2. Assumed knowledge
 Chemistry 2 Lecture 8 IR Spectroscopy of Polyatomic Molecles Assumed knowledge There are 3N 6 vibrations in a non linear molecule and 3N 5 vibrations in a linear molecule. Only modes that lead to a change
Chemistry 2 Lecture 8 IR Spectroscopy of Polyatomic Molecles Assumed knowledge There are 3N 6 vibrations in a non linear molecule and 3N 5 vibrations in a linear molecule. Only modes that lead to a change
Significant effort has been dedicated to elucidating the
 pubs.acs.org/jpcl Terms of Use CC-BY Protein Denaturation with Guanidinium: A 2D-IR Study Adriana Huerta-Viga and Sander Woutersen* Van t Hoff Institute for Molecular Sciences (HIMS), University of Amsterdam,
pubs.acs.org/jpcl Terms of Use CC-BY Protein Denaturation with Guanidinium: A 2D-IR Study Adriana Huerta-Viga and Sander Woutersen* Van t Hoff Institute for Molecular Sciences (HIMS), University of Amsterdam,
Fourier Transform IR Spectroscopy
 Fourier Transform IR Spectroscopy Absorption peaks in an infrared absorption spectrum arise from molecular vibrations Absorbed energy causes molecular motions which create a net change in the dipole moment.
Fourier Transform IR Spectroscopy Absorption peaks in an infrared absorption spectrum arise from molecular vibrations Absorbed energy causes molecular motions which create a net change in the dipole moment.
A Quantitative Anharmonic Analysis of the Amide A Band in -Helical Poly(L-Alanine)
 Sang-Ho Lee Noemi G. Mirkin Samuel Krimm Biophysics Research Division and Department of Physics, The University of Michigan, Ann Arbor, Michigan 48109 A Quantitative Anharmonic Analysis of the Amide A
Sang-Ho Lee Noemi G. Mirkin Samuel Krimm Biophysics Research Division and Department of Physics, The University of Michigan, Ann Arbor, Michigan 48109 A Quantitative Anharmonic Analysis of the Amide A
The energy landscape theory provides an essential framework. Two-Dimensional Infrared (2DIR) Spectroscopy of the Peptide Beta3s Folding
 pubs.acs.org/jpcl Two-Dimensional Infrared (2DIR) Spectroscopy of the Peptide Beta3s Folding Zaizhi Lai, Nicholas K Preketes, Jun Jiang, Shaul Mukamel, and Jin Wang*,, Department of Chemistry and Department
pubs.acs.org/jpcl Two-Dimensional Infrared (2DIR) Spectroscopy of the Peptide Beta3s Folding Zaizhi Lai, Nicholas K Preketes, Jun Jiang, Shaul Mukamel, and Jin Wang*,, Department of Chemistry and Department
Insights on Interfacial Structure, Dynamics and. Proton Transfer from Ultrafast Vibrational Sum. Frequency Generation Spectroscopy of the
 Insights on Interfacial Structure, Dynamics and Proton Transfer from Ultrafast Vibrational Sum Frequency Generation Spectroscopy of the Alumina(0001)/Water Interface Aashish Tuladhar, Stefan M. Piontek,
Insights on Interfacial Structure, Dynamics and Proton Transfer from Ultrafast Vibrational Sum Frequency Generation Spectroscopy of the Alumina(0001)/Water Interface Aashish Tuladhar, Stefan M. Piontek,
Supplementary Figure 1 Level structure of a doubly charged QDM (a) PL bias map acquired under 90 nw non-resonant excitation at 860 nm.
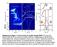 Supplementary Figure 1 Level structure of a doubly charged QDM (a) PL bias map acquired under 90 nw non-resonant excitation at 860 nm. Charging steps are labeled by the vertical dashed lines. Intensity
Supplementary Figure 1 Level structure of a doubly charged QDM (a) PL bias map acquired under 90 nw non-resonant excitation at 860 nm. Charging steps are labeled by the vertical dashed lines. Intensity
CHINESE JOURNAL OF CHEMICAL PHYSICS VOLUME 20, NUMBER 4 AUGUST 27, 2007
 CHINESE JOURNAL OF CHEMICAL PHYSICS VOLUME 20, NUMBER 4 AUGUST 27, 2007 ARTICLE Intermolecular Hydrogen Bonds Formed Between Amino Acid Molecules in Aqueous Solution Investigated by Temperature-jump Nanosecond
CHINESE JOURNAL OF CHEMICAL PHYSICS VOLUME 20, NUMBER 4 AUGUST 27, 2007 ARTICLE Intermolecular Hydrogen Bonds Formed Between Amino Acid Molecules in Aqueous Solution Investigated by Temperature-jump Nanosecond
/2Mα 2 α + V n (R)] χ (R) = E υ χ υ (R)
![/2Mα 2 α + V n (R)] χ (R) = E υ χ υ (R) /2Mα 2 α + V n (R)] χ (R) = E υ χ υ (R)](/thumbs/75/72875547.jpg) Spectroscopy: Engel Chapter 18 XIV 67 Vibrational Spectroscopy (Typically IR and Raman) Born-Oppenheimer approx. separate electron-nuclear Assume elect-nuclear motion separate, full wave fct. ψ (r,r) =
Spectroscopy: Engel Chapter 18 XIV 67 Vibrational Spectroscopy (Typically IR and Raman) Born-Oppenheimer approx. separate electron-nuclear Assume elect-nuclear motion separate, full wave fct. ψ (r,r) =
Vibrational Spectroscopies. C-874 University of Delaware
 Vibrational Spectroscopies C-874 University of Delaware Vibrational Spectroscopies..everything that living things do can be understood in terms of the jigglings and wigglings of atoms.. R. P. Feymann Vibrational
Vibrational Spectroscopies C-874 University of Delaware Vibrational Spectroscopies..everything that living things do can be understood in terms of the jigglings and wigglings of atoms.. R. P. Feymann Vibrational
Headspace Raman Spectroscopy
 ELECTRONICALLY REPRINTED FROM SEPTEMBER 2014 Molecular Spectroscopy Workbench Raman Spectroscopy We examine vapor-phase Raman spectroscopy through the acquisition of spectra from gas molecules confined
ELECTRONICALLY REPRINTED FROM SEPTEMBER 2014 Molecular Spectroscopy Workbench Raman Spectroscopy We examine vapor-phase Raman spectroscopy through the acquisition of spectra from gas molecules confined
Nanoscale IR spectroscopy of organic contaminants
 The nanoscale spectroscopy company The world leader in nanoscale IR spectroscopy Nanoscale IR spectroscopy of organic contaminants Application note nanoir uniquely and unambiguously identifies organic
The nanoscale spectroscopy company The world leader in nanoscale IR spectroscopy Nanoscale IR spectroscopy of organic contaminants Application note nanoir uniquely and unambiguously identifies organic
Dave S. Walker and Geraldine L. Richmond*
 J. Phys. Chem. C 2007, 111, 8321-8330 8321 Understanding the Effects of Hydrogen Bonding at the Vapor-Water Interface: Vibrational Sum Frequency Spectroscopy of H 2 O/HOD/D 2 O Mixtures Studied Using Molecular
J. Phys. Chem. C 2007, 111, 8321-8330 8321 Understanding the Effects of Hydrogen Bonding at the Vapor-Water Interface: Vibrational Sum Frequency Spectroscopy of H 2 O/HOD/D 2 O Mixtures Studied Using Molecular
k n (ω 01 ) k 2 (ω 01 ) k 3 (ω 01 ) k 1 (ω 01 )
 e (ω 01 ) n (ω 01 ) 3 (ω 01 ) 01 (ω 01 ) 1 (ω 01 ) (ω 01 ) 3 (ω 01 ) 1 (ω 01 ) 1 0 t 3 T w τ Rephasing R 1 R e = 1 + + 3 Nonrephasing n = 1 + 3 e (ω 1-1+1 ) n (ω 1-1+1 ) 3 (ω 1-1+1 ) (ω 01 ) t 1+1 1 3
e (ω 01 ) n (ω 01 ) 3 (ω 01 ) 01 (ω 01 ) 1 (ω 01 ) (ω 01 ) 3 (ω 01 ) 1 (ω 01 ) 1 0 t 3 T w τ Rephasing R 1 R e = 1 + + 3 Nonrephasing n = 1 + 3 e (ω 1-1+1 ) n (ω 1-1+1 ) 3 (ω 1-1+1 ) (ω 01 ) t 1+1 1 3
Basics of protein structure
 Today: 1. Projects a. Requirements: i. Critical review of one paper ii. At least one computational result b. Noon, Dec. 3 rd written report and oral presentation are due; submit via email to bphys101@fas.harvard.edu
Today: 1. Projects a. Requirements: i. Critical review of one paper ii. At least one computational result b. Noon, Dec. 3 rd written report and oral presentation are due; submit via email to bphys101@fas.harvard.edu
Coherent Three-Pulse Spectroscopy of Coupled Vibrations in a Rigid Dipeptide: Density Functional Theory Simulations
 J. Phys. Chem. B 2003, 107, 5967-5985 5967 Coherent Three-Pulse Spectroscopy of Coupled Vibrations in a Rigid Dipeptide: Density Functional Theory Simulations Jens Dreyer,*, Andrew M. Moran, and Shaul
J. Phys. Chem. B 2003, 107, 5967-5985 5967 Coherent Three-Pulse Spectroscopy of Coupled Vibrations in a Rigid Dipeptide: Density Functional Theory Simulations Jens Dreyer,*, Andrew M. Moran, and Shaul
Infrared Spectroscopy: Identification of Unknown Substances
 Infrared Spectroscopy: Identification of Unknown Substances Suppose a white powder is one of the four following molecules. How can they be differentiated? H N N H H H H Na H H H H H A technique that is
Infrared Spectroscopy: Identification of Unknown Substances Suppose a white powder is one of the four following molecules. How can they be differentiated? H N N H H H H Na H H H H H A technique that is
Secondary Structure. Bioch/BIMS 503 Lecture 2. Structure and Function of Proteins. Further Reading. Φ, Ψ angles alone determine protein structure
 Bioch/BIMS 503 Lecture 2 Structure and Function of Proteins August 28, 2008 Robert Nakamoto rkn3c@virginia.edu 2-0279 Secondary Structure Φ Ψ angles determine protein structure Φ Ψ angles are restricted
Bioch/BIMS 503 Lecture 2 Structure and Function of Proteins August 28, 2008 Robert Nakamoto rkn3c@virginia.edu 2-0279 Secondary Structure Φ Ψ angles determine protein structure Φ Ψ angles are restricted
Lecture 8. Assumed knowledge
 Chemistry 2 Lecture 8 IR Spectroscopy of Polyatomic Molecles Assumed knowledge There are 3N 6 vibrations in a non linear molecule and 3N 5 vibrations in a linear molecule. Only modes that lead to a change
Chemistry 2 Lecture 8 IR Spectroscopy of Polyatomic Molecles Assumed knowledge There are 3N 6 vibrations in a non linear molecule and 3N 5 vibrations in a linear molecule. Only modes that lead to a change
BMB/Bi/Ch 173 Winter 2018
 BMB/Bi/Ch 173 Winter 2018 Homework Set 8.1 (100 Points) Assigned 2-27-18, due 3-6-18 by 10:30 a.m. TA: Rachael Kuintzle. Office hours: SFL 220, Friday 3/2 4:00-5:00pm and SFL 229, Monday 3/5 4:00-5:30pm.
BMB/Bi/Ch 173 Winter 2018 Homework Set 8.1 (100 Points) Assigned 2-27-18, due 3-6-18 by 10:30 a.m. TA: Rachael Kuintzle. Office hours: SFL 220, Friday 3/2 4:00-5:00pm and SFL 229, Monday 3/5 4:00-5:30pm.
Chemistry 524--Final Exam--Keiderling Dec. 12, pm SES
 Chemistry 524--Final Exam--Keiderling Dec. 12, 2002 --4-8 pm -- 238 SES Please answer all questions in the answer book provided. Calculators, rulers, pens and pencils are permitted plus one 8.5 x 11 sheet
Chemistry 524--Final Exam--Keiderling Dec. 12, 2002 --4-8 pm -- 238 SES Please answer all questions in the answer book provided. Calculators, rulers, pens and pencils are permitted plus one 8.5 x 11 sheet
Peptide folding in non-aqueous environments investigated with molecular dynamics simulations Soto Becerra, Patricia
 University of Groningen Peptide folding in non-aqueous environments investigated with molecular dynamics simulations Soto Becerra, Patricia IMPORTANT NOTE: You are advised to consult the publisher's version
University of Groningen Peptide folding in non-aqueous environments investigated with molecular dynamics simulations Soto Becerra, Patricia IMPORTANT NOTE: You are advised to consult the publisher's version
Spectroscopy in Inorganic Chemistry. Vibration and Rotation Spectroscopy
 Spectroscopy in Inorganic Chemistry Vibrational energy levels in a diatomic molecule f = k r r V = ½kX 2 Force constant r Displacement from equilibrium point 2 X= r=r-r eq V = ½kX 2 Fundamental Vibrational
Spectroscopy in Inorganic Chemistry Vibrational energy levels in a diatomic molecule f = k r r V = ½kX 2 Force constant r Displacement from equilibrium point 2 X= r=r-r eq V = ½kX 2 Fundamental Vibrational
Advanced Pharmaceutical Analysis
 Lecture 2 Advanced Pharmaceutical Analysis IR spectroscopy Dr. Baraa Ramzi Infrared Spectroscopy It is a powerful tool for identifying pure organic and inorganic compounds. Every molecular compound has
Lecture 2 Advanced Pharmaceutical Analysis IR spectroscopy Dr. Baraa Ramzi Infrared Spectroscopy It is a powerful tool for identifying pure organic and inorganic compounds. Every molecular compound has
THE VIBRATIONAL SPECTRUM OF A POLYATOMIC MOLECULE (Revised 4/7/2004)
 INTRODUCTION THE VIBRATIONAL SPECTRUM OF A POLYATOMIC MOLECULE (Revised 4/7/2004) The vibrational motion of a molecule is quantized and the resulting energy level spacings give rise to transitions in the
INTRODUCTION THE VIBRATIONAL SPECTRUM OF A POLYATOMIC MOLECULE (Revised 4/7/2004) The vibrational motion of a molecule is quantized and the resulting energy level spacings give rise to transitions in the
IR Spectrography - Absorption. Raman Spectrography - Scattering. n 0 n M - Raman n 0 - Rayleigh
 RAMAN SPECTROSCOPY Scattering Mid-IR and NIR require absorption of radiation from a ground level to an excited state, requires matching of radiation from source with difference in energy states. Raman
RAMAN SPECTROSCOPY Scattering Mid-IR and NIR require absorption of radiation from a ground level to an excited state, requires matching of radiation from source with difference in energy states. Raman
Formed by a TTR Mutant. Supporting Information
 Possible Existence of α-sheets in the Amyloid Fibrils Formed by a TTR 105-115 Mutant Supporting Information Mary Rose Hilaire, 1, Bei Ding, 1,2, Debopreeti Mukherjee, 1, Jianxin Chen, 1,2 and Feng Gai
Possible Existence of α-sheets in the Amyloid Fibrils Formed by a TTR 105-115 Mutant Supporting Information Mary Rose Hilaire, 1, Bei Ding, 1,2, Debopreeti Mukherjee, 1, Jianxin Chen, 1,2 and Feng Gai
Supplementary Information for. Vibrational Spectroscopy at Electrolyte Electrode Interfaces with Graphene Gratings
 Supplementary Information for Vibrational Spectroscopy at Electrolyte Electrode Interfaces with Graphene Gratings Supplementary Figure 1. Simulated from pristine graphene gratings at different Fermi energy
Supplementary Information for Vibrational Spectroscopy at Electrolyte Electrode Interfaces with Graphene Gratings Supplementary Figure 1. Simulated from pristine graphene gratings at different Fermi energy
Model Answer (Paper code: AR-7112) M. Sc. (Physics) IV Semester Paper I: Laser Physics and Spectroscopy
 Model Answer (Paper code: AR-7112) M. Sc. (Physics) IV Semester Paper I: Laser Physics and Spectroscopy Section I Q1. Answer (i) (b) (ii) (d) (iii) (c) (iv) (c) (v) (a) (vi) (b) (vii) (b) (viii) (a) (ix)
Model Answer (Paper code: AR-7112) M. Sc. (Physics) IV Semester Paper I: Laser Physics and Spectroscopy Section I Q1. Answer (i) (b) (ii) (d) (iii) (c) (iv) (c) (v) (a) (vi) (b) (vii) (b) (viii) (a) (ix)
Spectroscopy: Tinoco Chapter 10 (but vibration, Ch.9)
 Spectroscopy: Tinoco Chapter 10 (but vibration, Ch.9) XIV 67 Vibrational Spectroscopy (Typical for IR and Raman) Born-Oppenheimer separate electron-nuclear motion ψ (rr) = χ υ (R) φ el (r,r) -- product
Spectroscopy: Tinoco Chapter 10 (but vibration, Ch.9) XIV 67 Vibrational Spectroscopy (Typical for IR and Raman) Born-Oppenheimer separate electron-nuclear motion ψ (rr) = χ υ (R) φ el (r,r) -- product
SUPPLEMENTARY INFORMATION
 doi:1.138/nature9829 Supplementary Information S1: Movie of the photo-induced phase transition: Figures 2b-e show four selected XUV ARPES snapshots illustrating the most pronounced changes in the course
doi:1.138/nature9829 Supplementary Information S1: Movie of the photo-induced phase transition: Figures 2b-e show four selected XUV ARPES snapshots illustrating the most pronounced changes in the course
Probing and Driving Molecular Dynamics with Femtosecond Pulses
 Miroslav Kloz Probing and Driving Molecular Dynamics with Femtosecond Pulses (wavelengths above 200 nm, energies below mj) Why femtosecond lasers in biology? Scales of size and time are closely rerated!
Miroslav Kloz Probing and Driving Molecular Dynamics with Femtosecond Pulses (wavelengths above 200 nm, energies below mj) Why femtosecond lasers in biology? Scales of size and time are closely rerated!
Chapter 12 Mass Spectrometry and Infrared Spectroscopy
 Organic Chemistry, 6 th Edition L. G. Wade, Jr. Chapter 12 Mass Spectrometry and Infrared Spectroscopy Jo Blackburn Richland College, Dallas, TX Dallas County Community College District 2006, Prentice
Organic Chemistry, 6 th Edition L. G. Wade, Jr. Chapter 12 Mass Spectrometry and Infrared Spectroscopy Jo Blackburn Richland College, Dallas, TX Dallas County Community College District 2006, Prentice
Specialized Raman Techniques. Strictly speaking the radiation-induced dipole moment should be expressed as
 Nonlinear effects Specialized Raman Techniques Strictly speaking the radiation-induced dipole moment should be expressed as M = E + ½E 2 + (1/6)E 3 +... Where and are the first and second hyperpolarizabilities.
Nonlinear effects Specialized Raman Techniques Strictly speaking the radiation-induced dipole moment should be expressed as M = E + ½E 2 + (1/6)E 3 +... Where and are the first and second hyperpolarizabilities.
Vibrational Analysis of Peptides, Polypeptides and Proteins XII-Fermi Resonance Analysis of the Unperturbed ND Stretching Fundamental in Polypeptides
 DEDICATED TO T. SHIMANOUCHI Vibrational Analysis of Peptides, Polypeptides and Proteins XII-Fermi Resonance Analysis of the Unperturbed ND Stretching Fundamental in Polypeptides S. Krimm and Anil M. Dwivedi
DEDICATED TO T. SHIMANOUCHI Vibrational Analysis of Peptides, Polypeptides and Proteins XII-Fermi Resonance Analysis of the Unperturbed ND Stretching Fundamental in Polypeptides S. Krimm and Anil M. Dwivedi
Supporting information for the manuscript. Excited state structural evolution during charge-transfer reactions in Betaine-30
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2015 Supporting information for the manuscript Excited state structural evolution during
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2015 Supporting information for the manuscript Excited state structural evolution during
16.1 Introduction to NMR Spectroscopy. Spectroscopy. Spectroscopy. Spectroscopy. Spectroscopy. Spectroscopy 4/11/2013
 What is spectroscopy? NUCLEAR MAGNETIC RESONANCE (NMR) spectroscopy may be the most powerful method of gaining structural information about organic compounds. NMR involves an interaction between electromagnetic
What is spectroscopy? NUCLEAR MAGNETIC RESONANCE (NMR) spectroscopy may be the most powerful method of gaining structural information about organic compounds. NMR involves an interaction between electromagnetic
Control and Characterization of Intramolecular Dynamics with Chirped Femtosecond Three-Pulse Four-Wave Mixing
 106 J. Phys. Chem. A 1999, 103, 106-1036 Control and Characterization of Intramolecular Dynamics with Chirped Femtosecond Three-Pulse Four-Wave Mixing Igor Pastirk, Vadim V. Lozovoy, Bruna I. Grimberg,
106 J. Phys. Chem. A 1999, 103, 106-1036 Control and Characterization of Intramolecular Dynamics with Chirped Femtosecond Three-Pulse Four-Wave Mixing Igor Pastirk, Vadim V. Lozovoy, Bruna I. Grimberg,
Mike Towrie Central Laser Facility Rutherford Appleton Laboratory. Diamond DIAMOND. Tony Parker, Pavel Matousek
 Ultrafast deactivation of the electronic excited states of DNA bases and polynucleotides following 267 nm laser excitation explored using picosecond time-resolved infrared spectroscopy 1 Mike Towrie (m.towrie@rl.ac.uk)
Ultrafast deactivation of the electronic excited states of DNA bases and polynucleotides following 267 nm laser excitation explored using picosecond time-resolved infrared spectroscopy 1 Mike Towrie (m.towrie@rl.ac.uk)
Supplementary Information
 Supplementary Information Supplementary Figure 1. Schematic diagram of the RF magneton sputtering system used to deposit a thin SiO2 film on CaF2 surface. 1 Supplementary Figure 2. A Li + DEC complex structure.
Supplementary Information Supplementary Figure 1. Schematic diagram of the RF magneton sputtering system used to deposit a thin SiO2 film on CaF2 surface. 1 Supplementary Figure 2. A Li + DEC complex structure.
MASSACHUSETTS INSTITUTE OF TECHNOLOGY Quantum Mechanics II Spring, 2004 Professor Robert W. Field. Problem Sets #8 and #9
 MIT Department of Chemistry 5.74 Spring 2004: Introductory Quantum Mechanics II Course Instructors: Professor Robert Field and Professor Andrei Tokmakoff MASSACHUSETTS INSTITUTE OF TECHNOLOGY 5.74 Quantum
MIT Department of Chemistry 5.74 Spring 2004: Introductory Quantum Mechanics II Course Instructors: Professor Robert Field and Professor Andrei Tokmakoff MASSACHUSETTS INSTITUTE OF TECHNOLOGY 5.74 Quantum
Nature Structural & Molecular Biology: doi: /nsmb.3194
 Supplementary Figure 1 Mass spectrometry and solution NMR data for -syn samples used in this study. (a) Matrix-assisted laser-desorption and ionization time-of-flight (MALDI-TOF) mass spectrum of uniformly-
Supplementary Figure 1 Mass spectrometry and solution NMR data for -syn samples used in this study. (a) Matrix-assisted laser-desorption and ionization time-of-flight (MALDI-TOF) mass spectrum of uniformly-
Spectroscopic techniques: why, when, where,and how Dr. Roberto GIANGIACOMO
 Spectroscopic techniques: why, when, where,and how Dr. Roberto GIANGIACOMO BASIC INFORMATION Spectroscopy uses light to analyze substances or products by describing the energy transfer between light and
Spectroscopic techniques: why, when, where,and how Dr. Roberto GIANGIACOMO BASIC INFORMATION Spectroscopy uses light to analyze substances or products by describing the energy transfer between light and
4. Circular Dichroism - Spectroscopy
 4. Circular Dichroism - Spectroscopy The optical rotatory dispersion (ORD) and the circular dichroism (CD) are special variations of absorption spectroscopy in the UV and VIS region of the spectrum. The
4. Circular Dichroism - Spectroscopy The optical rotatory dispersion (ORD) and the circular dichroism (CD) are special variations of absorption spectroscopy in the UV and VIS region of the spectrum. The
Two-dimensional vibrational optical probes for peptide fast folding investigation
 Two-dimensional vibrational optical probes for peptide fast folding investigation Wei Zhuang, Darius Abramavicius, and Shaul Mukamel PNAS published online Dec 4, 2006; doi:10.1073/pnas.0606912103 This
Two-dimensional vibrational optical probes for peptide fast folding investigation Wei Zhuang, Darius Abramavicius, and Shaul Mukamel PNAS published online Dec 4, 2006; doi:10.1073/pnas.0606912103 This
Introduction to Molecular Vibrations and Infrared Spectroscopy
 hemistry 362 Spring 2017 Dr. Jean M. Standard February 15, 2017 Introduction to Molecular Vibrations and Infrared Spectroscopy Vibrational Modes For a molecule with N atoms, the number of vibrational modes
hemistry 362 Spring 2017 Dr. Jean M. Standard February 15, 2017 Introduction to Molecular Vibrations and Infrared Spectroscopy Vibrational Modes For a molecule with N atoms, the number of vibrational modes
CONFOCHECK. Innovation with Integrity. Infrared Protein Analysis FT-IR
 CONFOCHECK Infrared Protein Analysis Innovation with Integrity FT-IR CONFOCHECK: FT-IR System for Protein Analytics FT-IR Protein Analysis Infrared spectroscopy measures molecular vibrations due to the
CONFOCHECK Infrared Protein Analysis Innovation with Integrity FT-IR CONFOCHECK: FT-IR System for Protein Analytics FT-IR Protein Analysis Infrared spectroscopy measures molecular vibrations due to the
As a partial differential equation, the Helmholtz equation does not lend itself easily to analytical
 Aaron Rury Research Prospectus 21.6.2009 Introduction: The Helmhlotz equation, ( 2 +k 2 )u(r)=0 1, serves as the basis for much of optical physics. As a partial differential equation, the Helmholtz equation
Aaron Rury Research Prospectus 21.6.2009 Introduction: The Helmhlotz equation, ( 2 +k 2 )u(r)=0 1, serves as the basis for much of optical physics. As a partial differential equation, the Helmholtz equation
Ultrafast heterodyne detected infrared multidimensional vibrational stimulated echo studies of hydrogen bond dynamics
 Chemical Physics Letters 374 (2003) 362 371 www.elsevier.com/locate/cplett Ultrafast heterodyne detected infrared multidimensional vibrational stimulated echo studies of hydrogen bond dynamics John B.
Chemical Physics Letters 374 (2003) 362 371 www.elsevier.com/locate/cplett Ultrafast heterodyne detected infrared multidimensional vibrational stimulated echo studies of hydrogen bond dynamics John B.
