The Bio-chemical Information Processing Metaphor as a Programming Paradigm for Organic Computing (CHEMORG III)
|
|
|
- Duane Bailey
- 5 years ago
- Views:
Transcription
1 The Bio-chemical Information Processing Metaphor as a Programming Paradigm for Organic Computing (CHEMORG III) Gabi Escuela, Peter Kreyßig, and Peter Dittrich Friedrich-Schiller-University Jena, Department of Mathematics and Computer Science, Bio Systems Analysis Group 15. September 2011
2 Overview Summary New Results 1. Space: Reaction Flow Artificial Chemistries 2. Structured Molecules: Embodied Evolution 3. Theory: Decomposition Theorem and Stochastic Organizations Outlook Apoptosis
3 Aim Employ the (bio-)chemical principles of information processing as a programming approach for Organic Computing. How to program chemical-like systems? Current state: 45% of Phase III.
4 Main Results Organization-oriented chemical programing N. Matsumaru, F. Centler, P. Speroni d. F., P. Dittrich. Chemical Organization Theory as a Theoretical Base for Chemical Computing. Int. J. Unconv. Comput., 3(4), , 2007
5 Organizations for different inflows N. Matsumaru, F. Centler, P. Speroni d. F., P. Dittrich. Chemical Organization Theory as a Theoretical Base for Chemical Computing. Int. J. Unconv. Comput., 3(4), , 2007
6
7
8
9
10
11 Main Results Organization-oriented chemical programing N. Matsumaru, F. Centler, P. Speroni d. F., P. Dittrich. Chemical Organization Theory as a Theoretical Base for Chemical Computing. Int. J. Unconv. Comput., 3(4), , 2007 Analysis Simulation various examples N. Matsumaru, T. Hinze, P. Dittrich. Organization-Oriented Chemical Programming of Distributed Artefacts. Int. J. Nanotechnol. Mol. Comp., 1(4), 1-19, 2009 Compared with evolutionary design Theory S. Peter, P. Dittrich. On the Relation between Organizations and Limit Sets in Chemical Reaction Systems, Adv. Complex Syst., 14(1): 77-96, 2011 Tools F. Centler, C. Kaleta, P. Speroni di Fenizio, P. Dittrich. Computing Chemical Organizations in Biological Networks, Bioinformatics, 24(14), , 2008 (& 2010) Concept: Emergent Control P. Dittrich,P. Kreyssig. Emergent Control. In: C. Muller-Schloer, H. Schmeck, T. Ungerer (Eds.), Organic Computing A Paradigm Shift for Complex Systems, Autonomic Systems, Volume 1, Part 1, 67-78, Springer, Basel, 2011
12 micro level macro level Architecture for Emergent Control feedforward controller system to be controlled macro-to-micro translator macro goals micro rules or downward causation desired macro behavior P. Dittrich,P. Kreyssig. Emergent Control. In: C. Muller-Schloer, H. Schmeck, T. Ungerer (Eds.), Organic Computing A Paradigm Shift for Complex Systems, Autonomic Systems, Volume 1, Part 1, 67-78, Springer, Basel, 2011
13 Considering space: Reaction Flow Artificial Chemistry [P. Kreyssig and P. Dittrich. Reaction flow artificial chemistries. In: T. Lenaerts et al. (Eds.), Proc. of ECAL 2011, MIT Press, Boston, MA, p , 2011]
14 Reaction Flow Artificial Chemistries spatial distribution of molecules given by a flow. Get additional parameters to control and program the artificial chemistry. [P. Kreyssig and P. Dittrich. Reaction flow artificial chemistries. In: T. Lenaerts et al. (Eds.), Proc. of ECAL 2011, MIT Press, Boston, MA, p , 2011]
15 RFAC - Example [P. Kreyssig and P. Dittrich. Reaction flow artificial chemistries. In: T. Lenaerts et al. (Eds.), Proc. of ECAL 2011, MIT Press, Boston, MA, p , 2011]
16 RFAC Differential Model Let the speciesbem {m,...,m 1 M }. Concentrationof a speciesis[m ](x, i y, t), so[m ]:R R R i R. [m i] t 1 V ([m ] i V ),V R k,i ([m ],..., [m 1 M ]) The changeof directional derivative of [m ] moleculeconcentration due to movementis the inthe directionof #reactions k, i, with constantreactionrates k R i V The changecausedby the reactionsappearsin the reactionterms R [P. Kreyssig and P. Dittrich. Reaction flow artificial chemistries. In: T. Lenaerts et al. (Eds.), Proc. of ECAL 2011, MIT Press, Boston, MA, p , 2011]. V.
17 RFAC Analysis via Organizations Chemical organizations at different spatial scales. Functional units should be an organization. [P. Kreyssig and P. Dittrich. Reaction flow artificial chemistries. In: T. Lenaerts et al. (Eds.), Proc. of ECAL 2011, MIT Press, Boston, MA, p , 2011]
18 RFAC - Conclusion Spatial flow dynamics adds another level to influence the behavior Potential for a new programing mechanism for chemical information processing. Spatial chemical organizations helps in understanding spatial chemical computing. [P. Kreyssig and P. Dittrich. Reaction flow artificial chemistries. In: T. Lenaerts et al. (Eds.), Proc. of ECAL 2011, MIT Press, Boston, MA, p , 2011]
19 Structured Molecules embodied evolution [G. Gruenert, G. Escuela, P. Dittrich, T. Hinze. Morphological Algorithms: Membrane Receptor-ligand Interactions and Rule-based Molecule Graph Evolution for Exact Set Cover Problem. In Proc. of 12 th Conf. on Membrane Computing, LNCS, Springer, Berlin, 2011]
20 Structured Molecules Studied κ-expression four different fraglets inspired by biology of arbitrary size still want to describe and analyze it inspired by computer sciences interesting for implementing our algorithms [J. Feret, V. Danos, J. Krivine, R. Harmer, and W. Fontana. Internal coarse-graining of molecular systems. PNAS 2009] [C. Tschudin. Fraglets-a metabolistic execution model for communication protocols. AINS 2003]
21 Structured Molecules: Molecular EA Embodied evolution: Using molecules as substrate for Evolutionary Algorithms Rule-based implementation (even Selection and Reproduction) T(x~a,t!1).Trans(f!1~a,t!11~a,t!12~b,t!13~c,t,dock~p1).T(x~a,t!11).T(x~b,t!12).T(x~c,t!13) 30 T(x~b,t!1).Trans(f!1~b,t!11~d,t!12~e,t!13~f,t,dock~p1).T(x~d,t!11).T(x~e,t!12).T(x~f,t!13) 30 T(x~c,t!1).Trans(f!1~c,t!11~g,t!12~h,t,t,dock~p1).T(x~g,t!11).T(x~h,t!12) 30 T(x~d,t!1).Trans(f!1~d,t!11~i,t!12~j,t,t,dock~p1).T(x~i,t!11).T(x~j,t!12) 30 T(x~e,t!1).Trans(f!1~e,t!11~a,t,t,t,dock~p1).T(x~a,t!11) 30 T(x~f,t!1).Trans(f!1~f,t!11~d,t,t,t,dock~p1).T(x~d,t!11) 30 1 Sol(test~ok,t~a,eval!+) + T(x~a,t!5).Trans(f!5,dock~p1) -> Sol(test~ok,t~a!1,eval!+).T(x~a!1,t!5).Trans(f!5,dock~p1) kfastbind 1 Sol(test~ok,t~b,eval!+) + T(x~b,t!5).Trans(f!5,dock~p1) -> Sol(test~ok,t~b!1,eval!+).T(x~b!1,t!5).Trans(f!5,dock~p1) kfastbind 1 Sol(test~ok,t~c,eval!+) + T(x~c,t!5).Trans(f!5,dock~p1) -> Sol(test~ok,t~c!1,eval!+).T(x~c!1,t!5).Trans(f!5,dock~p1) kfastbind 1 Sol(test~ok,t~d,eval!+) + T(x~d,t!5).Trans(f!5,dock~p1) -> Sol(test~ok,t~d!1,eval!+).T(x~d!1,t!5).Trans(f!5,dock~p1) kfastbind 1 Sol(test~ok,t~e,eval!+) + T(x~e,t!5).Trans(f!5,dock~p1) -> Sol(test~ok,t~e!1,eval!+).T(x~e!1,t!5).Trans(f!5,dock~p1) kfastbind # dissociate, if dock~bad 1 Sol(t!1).T(x!1,t!5).Trans(f!5,dock~bad) -> Sol(t) + T(x,t!5).Trans(f!5,dock~bad) ktdissociate # bind evaluation components: (if not bound to copyto) 2 Sol(test~ok,eval!2).Eval(t~a,sol!2) + T(x~a,t!5).Trans(t!5,dock~p1) -> Sol(test~ok,eval!2).Eval(t~a!1,sol!2).T(x~a!1,t!5).Trans(t!5,dock~p1) kfastbind 2 Sol(test~ok,eval!2).Eval(t~b,sol!2) + T(x~b,t!5).Trans(t!5,dock~p1) -> Sol(test~ok,eval!2).Eval(t~b!1,sol!2).T(x~b!1,t!5).Trans(t!5,dock~p1) kfastbind 2 Sol(test~ok,eval!2).Eval(t~c,sol!2) + T(x~c,t!5).Trans(t!5,dock~p1) -> Sol(test~ok,eval!2).Eval(t~c!1,sol!2).T(x~c!1,t!5).Trans(t!5,dock~p1) kfastbind 2 Sol(test~ok,eval!2).Eval(t~d,sol!2) + T(x~d,t!5).Trans(t!5,dock~p1) -> Sol(test~ok,eval!2).Eval(t~d!1,sol!2).T(x~d!1,t!5).Trans(t!5,dock~p1) kfastbind 2 Sol(test~ok,eval!2).Eval(t~e,sol!2) + T(x~e,t!5).Trans(t!5,dock~p1) -> Sol(test~ok,eval!2).Eval(t~e!1,sol!2).T(x~e!1,t!5).Trans(t!5,dock~p1) kfastbind 2 Sol(test~ok,eval!2).Eval(t~f,sol!2) + T(x~f,t!5).Trans(t!5,dock~p1) -> Sol(test~ok,eval!2).Eval(t~f!1,sol!2).T(x~f!1,t!5).Trans(t!5,dock~p1) kfastbind # dissociate if dock~bad 2 Eval(t!1).T(x!1,t!5).Trans(t!5,dock~bad) -> Eval(t) + T(x,t!5).Trans(t!5,dock~bad) ktdissociate [G. Gruenert, G. Escuela, P. Dittrich, T. Hinze. Morphological Algorithms: Membrane Receptor-ligand Interactions and Rule-based Molecule Graph Evolution for Exact Set Cover Problem. CMC 2011]
22 Structured Molecules: Molecular EA Embodied evolution: Using molecules as substrate for Evolutionary Algorithms Rule-based implementation (even Selection and Reproduction) T(x~a,t!1).Trans(f!1~a,t!11~a,t!12~b,t!13~c,t,dock~p1).T(x~a,t!11).T(x~b,t!12).T(x~c,t!13) 30 T(x~b,t!1).Trans(f!1~b,t!11~d,t!12~e,t!13~f,t,dock~p1).T(x~d,t!11).T(x~e,t!12).T(x~f,t!13) 30 T(x~c,t!1).Trans(f!1~c,t!11~g,t!12~h,t,t,dock~p1).T(x~g,t!11).T(x~h,t!12) 30 T(x~d,t!1).Trans(f!1~d,t!11~i,t!12~j,t,t,dock~p1).T(x~i,t!11).T(x~j,t!12) 30 T(x~e,t!1).Trans(f!1~e,t!11~a,t,t,t,dock~p1).T(x~a,t!11) 30 T(x~f,t!1).Trans(f!1~f,t!11~d,t,t,t,dock~p1).T(x~d,t!11) 30 1 Sol(test~ok,t~a,eval!+) + T(x~a,t!5).Trans(f!5,dock~p1) -> Sol(test~ok,t~a!1,eval!+).T(x~a!1,t!5).Trans(f!5,dock~p1) kfastbind 1 Sol(test~ok,t~b,eval!+) + T(x~b,t!5).Trans(f!5,dock~p1) -> Sol(test~ok,t~b!1,eval!+).T(x~b!1,t!5).Trans(f!5,dock~p1) kfastbind 1 Sol(test~ok,t~c,eval!+) + T(x~c,t!5).Trans(f!5,dock~p1) -> Sol(test~ok,t~c!1,eval!+).T(x~c!1,t!5).Trans(f!5,dock~p1) kfastbind 1 Sol(test~ok,t~d,eval!+) + T(x~d,t!5).Trans(f!5,dock~p1) -> Sol(test~ok,t~d!1,eval!+).T(x~d!1,t!5).Trans(f!5,dock~p1) kfastbind 1 Sol(test~ok,t~e,eval!+) + T(x~e,t!5).Trans(f!5,dock~p1) -> Sol(test~ok,t~e!1,eval!+).T(x~e!1,t!5).Trans(f!5,dock~p1) kfastbind # dissociate, if dock~bad 1 Sol(t!1).T(x!1,t!5).Trans(f!5,dock~bad) -> Sol(t) + T(x,t!5).Trans(f!5,dock~bad) ktdissociate # bind evaluation components: (if not bound to copyto) 2 Sol(test~ok,eval!2).Eval(t~a,sol!2) + T(x~a,t!5).Trans(t!5,dock~p1) -> Sol(test~ok,eval!2).Eval(t~a!1,sol!2).T(x~a!1,t!5).Trans(t!5,dock~p1) kfastbind 2 Sol(test~ok,eval!2).Eval(t~b,sol!2) + T(x~b,t!5).Trans(t!5,dock~p1) -> Sol(test~ok,eval!2).Eval(t~b!1,sol!2).T(x~b!1,t!5).Trans(t!5,dock~p1) kfastbind 2 Sol(test~ok,eval!2).Eval(t~c,sol!2) + T(x~c,t!5).Trans(t!5,dock~p1) -> Sol(test~ok,eval!2).Eval(t~c!1,sol!2).T(x~c!1,t!5).Trans(t!5,dock~p1) kfastbind 2 Sol(test~ok,eval!2).Eval(t~d,sol!2) + T(x~d,t!5).Trans(t!5,dock~p1) -> Sol(test~ok,eval!2).Eval(t~d!1,sol!2).T(x~d!1,t!5).Trans(t!5,dock~p1) kfastbind 2 Sol(test~ok,eval!2).Eval(t~e,sol!2) + T(x~e,t!5).Trans(t!5,dock~p1) -> Sol(test~ok,eval!2).Eval(t~e!1,sol!2).T(x~e!1,t!5).Trans(t!5,dock~p1) kfastbind 2 Sol(test~ok,eval!2).Eval(t~f,sol!2) + T(x~f,t!5).Trans(t!5,dock~p1) -> Sol(test~ok,eval!2).Eval(t~f!1,sol!2).T(x~f!1,t!5).Trans(t!5,dock~p1) kfastbind # dissociate if dock~bad 2 Eval(t!1).T(x!1,t!5).Trans(t!5,dock~bad) -> Eval(t) + T(x,t!5).Trans(t!5,dock~bad) ktdissociate [G. Gruenert, G. Escuela, P. Dittrich, T. Hinze. Morphological Algorithms: Membrane Receptor-ligand Interactions and Rule-based Molecule Graph Evolution for Exact Set Cover Problem. CMC 2011]
23 GEN PHE PHE GEN Structured Molecules - EAs T-A A trans a d T-a T-d T-A A B C a b c d e T-a T-d T-a T-d a b c d e A B C T-A Copier T-B B T-C C trans trans a b c d T-a T-c T-b T-d e T-e [G. Gruenert, G. Escuela, P. Dittrich, T. Hinze. Morphological Algorithms: Membrane Receptor-ligand Interactions and Rule-based Molecule Graph Evolution for Exact Set Cover Problem. CMC 2011]
24 Structured Molecules - EAs Medium mutation rate Extremely high mutation rate Low mutation rate Zero mutation rate Only local rules Asynchronous Dynamic optimisation Large populations Space [G. Gruenert, G. Escuela, P. Dittrich, T. Hinze. Morphological Algorithms: Membrane Receptor-ligand Interactions and Rule-based Molecule Graph Evolution for Exact Set Cover Problem. CMC 2011]
25 Theory - Decomposition theorem of chemical organizations - Stochastic systems [T. Veloz, B. Reynaert, D. Rojas-Camaggi and P. Dittrich. A Decomposition Theorem in Chemical Organizations. T. Lenaerts et al (Eds.), Proc. of ECAL 2011, MIT Press, Boston, MA, 2011] [C. Wozar, S. Peter, P. Kreyssig and P. Dittrich. Chemical Organization Theory in Discrete Systems. (in preparation) ]
26 Decomposition Theorem - Example A = N + E + F + C 1 + C C l 1. Non-reactive molecules N = { n } 2. (pure) Catalysts E = { e } 3. Overproduced molecules F = { o } 4. (pure) Cycle molecules C C = M (E+F+N) = { x, y, w, z } C = C 1 + C C l = {x, y} + {w, z} [T. Veloz, B. Reynaert, D. Rojas-Camaggi and P. Dittrich. A Decomposition Theorem in Chemical Organizations. Nürnberg, OC T. Lenaerts et al (Eds.), G. Proc. Escuela, of ECAL P. Kreyssig, 2011, MIT P. Dittrich Press, Boston, MA, 2011]
27 Decomposition Theorem - Conclusion better algorithms better understanding [T. Veloz, B. Reynaert, D. Rojas-Camaggi and P. Dittrich. A Decomposition Theorem in Chemical Organizations. T. Lenaerts et al (Eds.), Proc. of ECAL 2011, MIT Press, Boston, MA, 2011]
28 Theory II: Stochastic Organizations Link discrete stochastic dynamic models of (artificial) chemical systems to the theory of organizations. Markov chain [C. Wozar, S. Peter, P. Kreyssig and P. Dittrich. Chemical Organization Theory in Discrete Systems. (in preparation) ]
29 Stochastic Organizations - Example M R {A,B} {A B,B } Organizational part := set of strongly connected states such that no transition leaves the set from inside. [C. Wozar, S. Peter, P. Kreyssig and P. Dittrich. Chemical Organization Theory in Discrete Systems. (in preparation) ]
30 Outlook Artificial Apoptosis
31 Apoptosis vs Graceful Degradation
32 Apoptosis - Biology Apoptosis denotes the programmed cell death unlike the premature death of cells (necrosis). Can already be found in colonies of cells (not only in multi-cellular organisms) and in plants. Three functional units: Signaling pathways which can be activated from the inside or via receptors from the outside by other cells to induce the cell death. Apoptosis mechanism which breaks down the cell in small chunks (controlled demolition) without an inflammation (unlike necrosis). Notification of other cells of own death.
33 Apoptosis vs Graceful Degradation First purposes of apoptosis is protection from further damage if cell is dysfunctional or could spread diseases. Different from graceful degradation which is prevalent in most technical systems.
34 Apoptosis for Morphogenesis Second purposes of apoptosis is the shaping of an organism (morphogenesis). Artificial development uses it for homeostasis rather than for giving form to the organism. Olsen et.al. showed the importance of the feedback (notification) before cell death in cancer growth simulations. [M.M. Olsen, N. Siegelmann-Danieli, H.T. Siegelmann. Dynamic Computational Model Suggests that Cellular Citizenship is Fundamental for Selective Tumor Apoptosis. PLoS One 2010]
35 Apoptosis in AIS Artificial Immune Systems (AIS) for Wireless Sensor Networks use apoptosis. Switch-off nodes which are malicious and disturbing other nodes.
36 Artificial Apoptosis
37 Artificial Apoptosis
38 Artificial Apoptosis
39 Acknowledgements Naoki Matsumaru Pietro Speroni di Fenizio Stephan Peter Tomas Veloz Christoph Kaleta Funding: DFG, SPP Organic Computing
Chemical Organization Theory
 Chemical Organization Theory How are living systems organized? How do they evolve? Peter Dittrich Bio Systems Analysis Group FSU Jena Friedrich-Schiller-Universität Jena Jena Centre for Bioinformatics
Chemical Organization Theory How are living systems organized? How do they evolve? Peter Dittrich Bio Systems Analysis Group FSU Jena Friedrich-Schiller-Universität Jena Jena Centre for Bioinformatics
Reaction Flow Artificial Chemistries
 Reaction Flow Artificial Chemistries Peter Kreyssig 1 and Peter Dittrich 1 1 Bio Systems Analysis Group, Department of Mathematics and Computer Science, Friedrich Schiller University Jena, 07743 Jena,
Reaction Flow Artificial Chemistries Peter Kreyssig 1 and Peter Dittrich 1 1 Bio Systems Analysis Group, Department of Mathematics and Computer Science, Friedrich Schiller University Jena, 07743 Jena,
Blank for whiteboard 1
 Blank for whiteboard 1 Biological Computation Artificial Chemistries Life through Artificial Chemistries? part one Dave Dembeck 2 Questions How did we get from simple molecules to self-replicating systems?
Blank for whiteboard 1 Biological Computation Artificial Chemistries Life through Artificial Chemistries? part one Dave Dembeck 2 Questions How did we get from simple molecules to self-replicating systems?
Formal Quantitative Analysis of Reaction Networks Using Chemical Organisation Theory Mu, Chunyan; Dittrich, Peter ; Parker, David; Rowe, Jonathan
 Formal Quantitative Analysis of Reaction Networks Using Chemical Organisation Theory Mu, Chunyan; Dittrich, Peter ; Parker, David; Rowe, Jonathan DOI: 0.007/78---577-0_5 License: None: All rights reserved
Formal Quantitative Analysis of Reaction Networks Using Chemical Organisation Theory Mu, Chunyan; Dittrich, Peter ; Parker, David; Rowe, Jonathan DOI: 0.007/78---577-0_5 License: None: All rights reserved
A Signal Processing Approach to the Analysis of Chemical Networking Protocols
 Laurea Specialistica 19 July 2010 A Signal Processing Approach to the Analysis of Chemical Networking Protocols Author Supervisors Prof. Marco Luise Prof. Filippo Giannetti (University of Pisa) (University
Laurea Specialistica 19 July 2010 A Signal Processing Approach to the Analysis of Chemical Networking Protocols Author Supervisors Prof. Marco Luise Prof. Filippo Giannetti (University of Pisa) (University
Reaction Networks as a Language for Systemic Modeling: On the Study of Structural Changes
 systems Article Reaction Networks as a Language for Systemic Modeling: On the Study of Structural Changes Tomas Veloz 1,2, * and Pablo Razeto-Barry 2,3 1 Center Leo Apostel, Brussels Free University, 1050
systems Article Reaction Networks as a Language for Systemic Modeling: On the Study of Structural Changes Tomas Veloz 1,2, * and Pablo Razeto-Barry 2,3 1 Center Leo Apostel, Brussels Free University, 1050
biologically-inspired computing lecture 5 Informatics luis rocha 2015 biologically Inspired computing INDIANA UNIVERSITY
 lecture 5 -inspired Sections I485/H400 course outlook Assignments: 35% Students will complete 4/5 assignments based on algorithms presented in class Lab meets in I1 (West) 109 on Lab Wednesdays Lab 0 :
lecture 5 -inspired Sections I485/H400 course outlook Assignments: 35% Students will complete 4/5 assignments based on algorithms presented in class Lab meets in I1 (West) 109 on Lab Wednesdays Lab 0 :
An Application of Genetic Algorithms to Membrane Computing
 An Application of Genetic Algorithms to Membrane Computing Gabi Escuela 1, Miguel A. Gutiérrez-Naranjo 2 1 Bio Systems Analysis Group Friedrich Schiller University Jena gabi.escuela@uni-jena.de 2 Research
An Application of Genetic Algorithms to Membrane Computing Gabi Escuela 1, Miguel A. Gutiérrez-Naranjo 2 1 Bio Systems Analysis Group Friedrich Schiller University Jena gabi.escuela@uni-jena.de 2 Research
Reception The target cell s detection of a signal coming from outside the cell May Occur by: Direct connect Through signal molecules
 Why Do Cells Communicate? Regulation Cells need to control cellular processes In multicellular organism, cells signaling pathways coordinate the activities within individual cells that support the function
Why Do Cells Communicate? Regulation Cells need to control cellular processes In multicellular organism, cells signaling pathways coordinate the activities within individual cells that support the function
AP Biology UNIT 1: CELL BIOLOGY. Advanced Placement
 Advanced Placement AP Biology builds students' understanding of biology on both the micro and macro scales. After studying cell biology, students move on to understand how evolution drives the diversity
Advanced Placement AP Biology builds students' understanding of biology on both the micro and macro scales. After studying cell biology, students move on to understand how evolution drives the diversity
A. Incorrect! The Cell Cycle contains 4 distinct phases: (1) G 1, (2) S Phase, (3) G 2 and (4) M Phase.
 Molecular Cell Biology - Problem Drill 21: Cell Cycle and Cell Death Question No. 1 of 10 1. Which of the following statements about the cell cycle is correct? Question #1 (A) The Cell Cycle contains 3
Molecular Cell Biology - Problem Drill 21: Cell Cycle and Cell Death Question No. 1 of 10 1. Which of the following statements about the cell cycle is correct? Question #1 (A) The Cell Cycle contains 3
Types of biological networks. I. Intra-cellurar networks
 Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Cybergenetics: Control theory for living cells
 Department of Biosystems Science and Engineering, ETH-Zürich Cybergenetics: Control theory for living cells Corentin Briat Joint work with Ankit Gupta and Mustafa Khammash Introduction Overview Cybergenetics:
Department of Biosystems Science and Engineering, ETH-Zürich Cybergenetics: Control theory for living cells Corentin Briat Joint work with Ankit Gupta and Mustafa Khammash Introduction Overview Cybergenetics:
Life Sciences 1a: Section 3B. The cell division cycle Objectives Understand the challenges to producing genetically identical daughter cells
 Life Sciences 1a: Section 3B. The cell division cycle Objectives Understand the challenges to producing genetically identical daughter cells Understand how a simple biochemical oscillator can drive the
Life Sciences 1a: Section 3B. The cell division cycle Objectives Understand the challenges to producing genetically identical daughter cells Understand how a simple biochemical oscillator can drive the
CLOSURE IN ARTIFICIAL CELL SIGNALLING NETWORKS Investigating the Emergence of Cognition in Collectively Autocatalytic Reaction Networks
 CLOSURE IN ARTIFICIAL CELL SIGNALLING NETWORKS Investigating the Emergence of Cognition in Collectively Autocatalytic Reaction Networks James Decraene Artificial Life Laboratory, Research Institute for
CLOSURE IN ARTIFICIAL CELL SIGNALLING NETWORKS Investigating the Emergence of Cognition in Collectively Autocatalytic Reaction Networks James Decraene Artificial Life Laboratory, Research Institute for
Programmed Cell Death
 Programmed Cell Death Dewajani Purnomosari Department of Histology and Cell Biology Faculty of Medicine Universitas Gadjah Mada d.purnomosari@ugm.ac.id What is apoptosis? a normal component of the development
Programmed Cell Death Dewajani Purnomosari Department of Histology and Cell Biology Faculty of Medicine Universitas Gadjah Mada d.purnomosari@ugm.ac.id What is apoptosis? a normal component of the development
Domain 6: Communication
 Domain 6: Communication 6.1: Cell communication processes share common features that reflect a shared evolutionary history. (EK3.D.1) 1. Introduction to Communication Communication requires the generation,
Domain 6: Communication 6.1: Cell communication processes share common features that reflect a shared evolutionary history. (EK3.D.1) 1. Introduction to Communication Communication requires the generation,
Apoptosis in Mammalian Cells
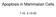 Apoptosis in Mammalian Cells 7.16 2-10-05 Apoptosis is an important factor in many human diseases Cancer malignant cells evade death by suppressing apoptosis (too little apoptosis) Stroke damaged neurons
Apoptosis in Mammalian Cells 7.16 2-10-05 Apoptosis is an important factor in many human diseases Cancer malignant cells evade death by suppressing apoptosis (too little apoptosis) Stroke damaged neurons
Chapter 6- An Introduction to Metabolism*
 Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Genetic Algorithm: introduction
 1 Genetic Algorithm: introduction 2 The Metaphor EVOLUTION Individual Fitness Environment PROBLEM SOLVING Candidate Solution Quality Problem 3 The Ingredients t reproduction t + 1 selection mutation recombination
1 Genetic Algorithm: introduction 2 The Metaphor EVOLUTION Individual Fitness Environment PROBLEM SOLVING Candidate Solution Quality Problem 3 The Ingredients t reproduction t + 1 selection mutation recombination
Introduction to Bioinformatics
 Systems biology Introduction to Bioinformatics Systems biology: modeling biological p Study of whole biological systems p Wholeness : Organization of dynamic interactions Different behaviour of the individual
Systems biology Introduction to Bioinformatics Systems biology: modeling biological p Study of whole biological systems p Wholeness : Organization of dynamic interactions Different behaviour of the individual
From cell biology to Petri nets. Rainer Breitling, Groningen, NL David Gilbert, London, UK Monika Heiner, Cottbus, DE
 From cell biology to Petri nets Rainer Breitling, Groningen, NL David Gilbert, London, UK Monika Heiner, Cottbus, DE Biology = Concentrations Breitling / 2 The simplest chemical reaction A B irreversible,
From cell biology to Petri nets Rainer Breitling, Groningen, NL David Gilbert, London, UK Monika Heiner, Cottbus, DE Biology = Concentrations Breitling / 2 The simplest chemical reaction A B irreversible,
biologically-inspired computing lecture 6 Informatics luis rocha 2015 INDIANA UNIVERSITY biologically Inspired computing
 lecture 6 -inspired Sections I485/H400 course outlook Assignments: 35% Students will complete 4/5 assignments based on algorithms presented in class Lab meets in I1 (West) 109 on Lab Wednesdays Lab 0 :
lecture 6 -inspired Sections I485/H400 course outlook Assignments: 35% Students will complete 4/5 assignments based on algorithms presented in class Lab meets in I1 (West) 109 on Lab Wednesdays Lab 0 :
Termination in Rule Based Approach to Signaling Systems
 Termination in Rule Based Approach to Signaling Systems Davis Buenger and Jim Faeder Bioengineering & Bioinformatics Summer Institute Dept of Bioengineering & Bioinformatics Summer Institute, Dept. of
Termination in Rule Based Approach to Signaling Systems Davis Buenger and Jim Faeder Bioengineering & Bioinformatics Summer Institute Dept of Bioengineering & Bioinformatics Summer Institute, Dept. of
Host-Pathogen Interaction. PN Sharma Department of Plant Pathology CSK HPKV, Palampur
 Host-Pathogen Interaction PN Sharma Department of Plant Pathology CSK HPKV, Palampur-176062 PATHOGEN DEFENCE IN PLANTS A BIOLOGICAL AND MOLECULAR VIEW Two types of plant resistance response to potential
Host-Pathogen Interaction PN Sharma Department of Plant Pathology CSK HPKV, Palampur-176062 PATHOGEN DEFENCE IN PLANTS A BIOLOGICAL AND MOLECULAR VIEW Two types of plant resistance response to potential
Bio 101 General Biology 1
 Revised: Fall 2016 Bio 101 General Biology 1 COURSE OUTLINE Prerequisites: Prerequisite: Successful completion of MTE 1, 2, 3, 4, and 5, and a placement recommendation for ENG 111, co-enrollment in ENF
Revised: Fall 2016 Bio 101 General Biology 1 COURSE OUTLINE Prerequisites: Prerequisite: Successful completion of MTE 1, 2, 3, 4, and 5, and a placement recommendation for ENG 111, co-enrollment in ENF
Evolutionary dynamics on graphs
 Evolutionary dynamics on graphs Laura Hindersin May 4th 2015 Max-Planck-Institut für Evolutionsbiologie, Plön Evolutionary dynamics Main ingredients: Fitness: The ability to survive and reproduce. Selection
Evolutionary dynamics on graphs Laura Hindersin May 4th 2015 Max-Planck-Institut für Evolutionsbiologie, Plön Evolutionary dynamics Main ingredients: Fitness: The ability to survive and reproduce. Selection
Creative Genomic Webs -Kapil Rajaraman PHY 498BIO, HW 4
 Creative Genomic Webs -Kapil Rajaraman (rajaramn@uiuc.edu) PHY 498BIO, HW 4 Evolutionary progress is generally considered a result of successful accumulation of mistakes in replication of the genetic code.
Creative Genomic Webs -Kapil Rajaraman (rajaramn@uiuc.edu) PHY 498BIO, HW 4 Evolutionary progress is generally considered a result of successful accumulation of mistakes in replication of the genetic code.
ADVANCED PLACEMENT BIOLOGY
 ADVANCED PLACEMENT BIOLOGY Description Advanced Placement Biology is designed to be the equivalent of a two-semester college introductory course for Biology majors. The course meets seven periods per week
ADVANCED PLACEMENT BIOLOGY Description Advanced Placement Biology is designed to be the equivalent of a two-semester college introductory course for Biology majors. The course meets seven periods per week
A A A A B B1
 LEARNING OBJECTIVES FOR EACH BIG IDEA WITH ASSOCIATED SCIENCE PRACTICES AND ESSENTIAL KNOWLEDGE Learning Objectives will be the target for AP Biology exam questions Learning Objectives Sci Prac Es Knowl
LEARNING OBJECTIVES FOR EACH BIG IDEA WITH ASSOCIATED SCIENCE PRACTICES AND ESSENTIAL KNOWLEDGE Learning Objectives will be the target for AP Biology exam questions Learning Objectives Sci Prac Es Knowl
Autocatalytic sets and chemical organizations: modeling self-sustaining reaction networks at the origin of life
 PAPER OPEN ACCESS Autocatalytic sets and chemical organizations: modeling self-sustaining reaction networks at the origin of life To cite this article: Wim Hordijk et al 2018 New J. Phys. 20 015011 View
PAPER OPEN ACCESS Autocatalytic sets and chemical organizations: modeling self-sustaining reaction networks at the origin of life To cite this article: Wim Hordijk et al 2018 New J. Phys. 20 015011 View
A Database of human biological pathways
 A Database of human biological pathways Steve Jupe - sjupe@ebi.ac.uk 1 Rationale Journal information Nature 407(6805):770-6.The Biochemistry of Apoptosis. Caspase-8 is the key initiator caspase in the
A Database of human biological pathways Steve Jupe - sjupe@ebi.ac.uk 1 Rationale Journal information Nature 407(6805):770-6.The Biochemistry of Apoptosis. Caspase-8 is the key initiator caspase in the
AP Biology Gene Regulation and Development Review
 AP Biology Gene Regulation and Development Review 1. What does the regulatory gene code for? 2. Is the repressor by default active/inactive? 3. What changes the repressor activity? 4. What does repressor
AP Biology Gene Regulation and Development Review 1. What does the regulatory gene code for? 2. Is the repressor by default active/inactive? 3. What changes the repressor activity? 4. What does repressor
Engineering Self-Organization and Emergence: issues and directions
 5/0/ Engineering Self-Organization and Emergence: issues and directions Franco Zambonelli franco.zambonelli@unimore.it Agents and Pervasive Computing Group Università di Modena e Reggio Emilia SOAS 005
5/0/ Engineering Self-Organization and Emergence: issues and directions Franco Zambonelli franco.zambonelli@unimore.it Agents and Pervasive Computing Group Università di Modena e Reggio Emilia SOAS 005
Bioinformatics 2. Yeast two hybrid. Proteomics. Proteomics
 GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
Capacity of Discrete Molecular Diffusion Channels
 211 IEEE International Symposium on Information Theory Proceedings Capacity of Discrete Molecular Diffusion Channels Arash nolghozati, Mohsen Sardari, Ahmad Beirami, Faramarz Fekri School of Electrical
211 IEEE International Symposium on Information Theory Proceedings Capacity of Discrete Molecular Diffusion Channels Arash nolghozati, Mohsen Sardari, Ahmad Beirami, Faramarz Fekri School of Electrical
Big Idea #2. Energy. Types of Potential Energy. Kinetic Energy. Chemical Potential Energy. Metabolism
 Big Idea #2 Biological Systems utilize free energy and molecular building blocks to grow, to reproduce and to maintain dynamic homeostasis Life runs on chemical reactions rearranging atoms transforming
Big Idea #2 Biological Systems utilize free energy and molecular building blocks to grow, to reproduce and to maintain dynamic homeostasis Life runs on chemical reactions rearranging atoms transforming
Proteomics. Yeast two hybrid. Proteomics - PAGE techniques. Data obtained. What is it?
 Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
Biological Pathways Representation by Petri Nets and extension
 Biological Pathways Representation by and extensions December 6, 2006 Biological Pathways Representation by and extension 1 The cell Pathways 2 Definitions 3 4 Biological Pathways Representation by and
Biological Pathways Representation by and extensions December 6, 2006 Biological Pathways Representation by and extension 1 The cell Pathways 2 Definitions 3 4 Biological Pathways Representation by and
Biology New Jersey 1. NATURE OF LIFE 2. THE CHEMISTRY OF LIFE. Tutorial Outline
 Tutorial Outline New Jersey Tutorials are designed specifically for the New Jersey Core Curriculum Content Standards to prepare students for the PARCC assessments, the New Jersey Biology Competency Test
Tutorial Outline New Jersey Tutorials are designed specifically for the New Jersey Core Curriculum Content Standards to prepare students for the PARCC assessments, the New Jersey Biology Competency Test
BIOLOGY STANDARDS BASED RUBRIC
 BIOLOGY STANDARDS BASED RUBRIC STUDENTS WILL UNDERSTAND THAT THE FUNDAMENTAL PROCESSES OF ALL LIVING THINGS DEPEND ON A VARIETY OF SPECIALIZED CELL STRUCTURES AND CHEMICAL PROCESSES. First Semester Benchmarks:
BIOLOGY STANDARDS BASED RUBRIC STUDENTS WILL UNDERSTAND THAT THE FUNDAMENTAL PROCESSES OF ALL LIVING THINGS DEPEND ON A VARIETY OF SPECIALIZED CELL STRUCTURES AND CHEMICAL PROCESSES. First Semester Benchmarks:
BIRKBECK COLLEGE (University of London)
 BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
VCE BIOLOGY Relationship between the key knowledge and key skills of the Study Design and the Study Design
 VCE BIOLOGY 2006 2014 Relationship between the key knowledge and key skills of the 2000 2005 Study Design and the 2006 2014 Study Design The following table provides a comparison of the key knowledge (and
VCE BIOLOGY 2006 2014 Relationship between the key knowledge and key skills of the 2000 2005 Study Design and the 2006 2014 Study Design The following table provides a comparison of the key knowledge (and
 1 of 13 8/11/2014 10:32 AM Units: Teacher: APBiology, CORE Course: APBiology Year: 2012-13 Chemistry of Life Chapters 1-4 Big Idea 1, 2 & 4 Change in the genetic population over time is feedback mechanisms
1 of 13 8/11/2014 10:32 AM Units: Teacher: APBiology, CORE Course: APBiology Year: 2012-13 Chemistry of Life Chapters 1-4 Big Idea 1, 2 & 4 Change in the genetic population over time is feedback mechanisms
Map of AP-Aligned Bio-Rad Kits with Learning Objectives
 Map of AP-Aligned Bio-Rad Kits with Learning Objectives Cover more than one AP Biology Big Idea with these AP-aligned Bio-Rad kits. Big Idea 1 Big Idea 2 Big Idea 3 Big Idea 4 ThINQ! pglo Transformation
Map of AP-Aligned Bio-Rad Kits with Learning Objectives Cover more than one AP Biology Big Idea with these AP-aligned Bio-Rad kits. Big Idea 1 Big Idea 2 Big Idea 3 Big Idea 4 ThINQ! pglo Transformation
MEDLINE Clinical Laboratory Sciences Journals
 Source Type Publication Name ISSN Peer-Reviewed Academic Journal Acta Biochimica et Biophysica Sinica 1672-9145 Y Academic Journal Acta Physiologica 1748-1708 Y Academic Journal Aging Cell 1474-9718 Y
Source Type Publication Name ISSN Peer-Reviewed Academic Journal Acta Biochimica et Biophysica Sinica 1672-9145 Y Academic Journal Acta Physiologica 1748-1708 Y Academic Journal Aging Cell 1474-9718 Y
Understanding Science Through the Lens of Computation. Richard M. Karp Nov. 3, 2007
 Understanding Science Through the Lens of Computation Richard M. Karp Nov. 3, 2007 The Computational Lens Exposes the computational nature of natural processes and provides a language for their description.
Understanding Science Through the Lens of Computation Richard M. Karp Nov. 3, 2007 The Computational Lens Exposes the computational nature of natural processes and provides a language for their description.
The Origin Of Life: A Network Oriented View
 The Origin Of Life: A Network Oriented View : A Chris Gordon-Smith Project www.simsoup.info Presentation To Life And Mind Group Sussex University 18 June 2010 Copyright Chris Gordon-Smith 2010 Presentation
The Origin Of Life: A Network Oriented View : A Chris Gordon-Smith Project www.simsoup.info Presentation To Life And Mind Group Sussex University 18 June 2010 Copyright Chris Gordon-Smith 2010 Presentation
Answer Key. Cell Growth and Division
 Cell Growth and Division Answer Key SECTION 1. THE CELL CYCLE Cell Cycle: (1) Gap1 (G 1): cells grow, carry out normal functions, and copy their organelles. (2) Synthesis (S): cells replicate DNA. (3)
Cell Growth and Division Answer Key SECTION 1. THE CELL CYCLE Cell Cycle: (1) Gap1 (G 1): cells grow, carry out normal functions, and copy their organelles. (2) Synthesis (S): cells replicate DNA. (3)
Towards a framework for multi-level modelling in Computational Biology
 Towards a framework for multi-level modelling in Computational Biology Sara Montagna sara.montagna@unibo.it Alma Mater Studiorum Università di Bologna PhD in Electronics, Computer Science and Telecommunications
Towards a framework for multi-level modelling in Computational Biology Sara Montagna sara.montagna@unibo.it Alma Mater Studiorum Università di Bologna PhD in Electronics, Computer Science and Telecommunications
Enduring understanding 1.A: Change in the genetic makeup of a population over time is evolution.
 The AP Biology course is designed to enable you to develop advanced inquiry and reasoning skills, such as designing a plan for collecting data, analyzing data, applying mathematical routines, and connecting
The AP Biology course is designed to enable you to develop advanced inquiry and reasoning skills, such as designing a plan for collecting data, analyzing data, applying mathematical routines, and connecting
Biological Networks. Gavin Conant 163B ASRC
 Biological Networks Gavin Conant 163B ASRC conantg@missouri.edu 882-2931 Types of Network Regulatory Protein-interaction Metabolic Signaling Co-expressing General principle Relationship between genes Gene/protein/enzyme
Biological Networks Gavin Conant 163B ASRC conantg@missouri.edu 882-2931 Types of Network Regulatory Protein-interaction Metabolic Signaling Co-expressing General principle Relationship between genes Gene/protein/enzyme
On the Evolution of Decoys in Plant Immune Systems
 On the Evolution of Decoys in Plant Immune Systems Iaroslav Ispolatov and Michael Doebeli Departments of Mathematics and Zoology University of British Columbia Vancouver, BC, Canada slava@math.ubc.ca doebeli@zoology.ubc.ca
On the Evolution of Decoys in Plant Immune Systems Iaroslav Ispolatov and Michael Doebeli Departments of Mathematics and Zoology University of British Columbia Vancouver, BC, Canada slava@math.ubc.ca doebeli@zoology.ubc.ca
I. Molecules and Cells: Cells are the structural and functional units of life; cellular processes are based on physical and chemical changes.
 I. Molecules and Cells: Cells are the structural and functional units of life; cellular processes are based on physical and chemical changes. A. Chemistry of Life B. Cells 1. Water How do the unique chemical
I. Molecules and Cells: Cells are the structural and functional units of life; cellular processes are based on physical and chemical changes. A. Chemistry of Life B. Cells 1. Water How do the unique chemical
The application of an artificial immune system for solving the identification problem in Ecology
 The application of an artificial immune system for solving the identification problem in Ecology IRINA ASTACHOVA 1 *, STANISLAV USHAKOV 1, ANDREI SELEMENEV 1, and JULIYA HITSKOVA 1 Department of Applied
The application of an artificial immune system for solving the identification problem in Ecology IRINA ASTACHOVA 1 *, STANISLAV USHAKOV 1, ANDREI SELEMENEV 1, and JULIYA HITSKOVA 1 Department of Applied
Objective 1: The student will demonstrate an understanding of the nature of science.
 August 2003 Objective 1: The student will demonstrate an understanding of the nature of science. Biology (1) and Integrated Physics and Chemistry (1) Scientific Processes. The student, for at least 40%
August 2003 Objective 1: The student will demonstrate an understanding of the nature of science. Biology (1) and Integrated Physics and Chemistry (1) Scientific Processes. The student, for at least 40%
Sample Questions for the Chemistry of Life Topic Test
 Sample Questions for the Chemistry of Life Topic Test 1. Enzymes play a crucial role in biology by serving as biological catalysts, increasing the rates of biochemical reactions by decreasing their activation
Sample Questions for the Chemistry of Life Topic Test 1. Enzymes play a crucial role in biology by serving as biological catalysts, increasing the rates of biochemical reactions by decreasing their activation
Introduction. ECE/CS/BioEn 6760 Modeling and Analysis of Biological Networks. Adventures in Synthetic Biology. Synthetic Biology.
 Introduction ECE/CS/BioEn 6760 Modeling and Analysis of Biological Networks Chris J. Myers Lecture 17: Genetic Circuit Design Electrical engineering principles can be applied to understand the behavior
Introduction ECE/CS/BioEn 6760 Modeling and Analysis of Biological Networks Chris J. Myers Lecture 17: Genetic Circuit Design Electrical engineering principles can be applied to understand the behavior
Systems Biology Across Scales: A Personal View XIV. Intra-cellular systems IV: Signal-transduction and networks. Sitabhra Sinha IMSc Chennai
 Systems Biology Across Scales: A Personal View XIV. Intra-cellular systems IV: Signal-transduction and networks Sitabhra Sinha IMSc Chennai Intra-cellular biochemical networks Metabolic networks Nodes:
Systems Biology Across Scales: A Personal View XIV. Intra-cellular systems IV: Signal-transduction and networks Sitabhra Sinha IMSc Chennai Intra-cellular biochemical networks Metabolic networks Nodes:
Big Idea 1: The process of evolution drives the diversity and unity of life.
 Big Idea 1: The process of evolution drives the diversity and unity of life. understanding 1.A: Change in the genetic makeup of a population over time is evolution. 1.A.1: Natural selection is a major
Big Idea 1: The process of evolution drives the diversity and unity of life. understanding 1.A: Change in the genetic makeup of a population over time is evolution. 1.A.1: Natural selection is a major
Regulation and signaling. Overview. Control of gene expression. Cells need to regulate the amounts of different proteins they express, depending on
 Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
AP Curriculum Framework with Learning Objectives
 Big Ideas Big Idea 1: The process of evolution drives the diversity and unity of life. AP Curriculum Framework with Learning Objectives Understanding 1.A: Change in the genetic makeup of a population over
Big Ideas Big Idea 1: The process of evolution drives the diversity and unity of life. AP Curriculum Framework with Learning Objectives Understanding 1.A: Change in the genetic makeup of a population over
Analysis of correlated mutations in Ras G-domain
 www.bioinformation.net Volume 13(6) Hypothesis Analysis of correlated mutations in Ras G-domain Ekta Pathak * Bioinformatics Department, MMV, Banaras Hindu University. Ekta Pathak - E-mail: ektavpathak@gmail.com;
www.bioinformation.net Volume 13(6) Hypothesis Analysis of correlated mutations in Ras G-domain Ekta Pathak * Bioinformatics Department, MMV, Banaras Hindu University. Ekta Pathak - E-mail: ektavpathak@gmail.com;
CHAPTER 2 THE CHEMICAL BASIS OF LIFE
 CHAPTER 2 THE CHEMICAL BASIS OF LIFE CHAPTER OVERVIEW This chapter introduces very basic concepts of chemistry, emphasizing the structure of atoms and how they combine (form bonds). The types of bonds,
CHAPTER 2 THE CHEMICAL BASIS OF LIFE CHAPTER OVERVIEW This chapter introduces very basic concepts of chemistry, emphasizing the structure of atoms and how they combine (form bonds). The types of bonds,
Evolutionary Games and Computer Simulations
 Evolutionary Games and Computer Simulations Bernardo A. Huberman and Natalie S. Glance Dynamics of Computation Group Xerox Palo Alto Research Center Palo Alto, CA 94304 Abstract The prisoner s dilemma
Evolutionary Games and Computer Simulations Bernardo A. Huberman and Natalie S. Glance Dynamics of Computation Group Xerox Palo Alto Research Center Palo Alto, CA 94304 Abstract The prisoner s dilemma
Evaluate evidence provided by data from many scientific disciplines to support biological evolution. [LO 1.9, SP 5.3]
![Evaluate evidence provided by data from many scientific disciplines to support biological evolution. [LO 1.9, SP 5.3] Evaluate evidence provided by data from many scientific disciplines to support biological evolution. [LO 1.9, SP 5.3]](/thumbs/75/72965665.jpg) Learning Objectives Evaluate evidence provided by data from many scientific disciplines to support biological evolution. [LO 1.9, SP 5.3] Refine evidence based on data from many scientific disciplines
Learning Objectives Evaluate evidence provided by data from many scientific disciplines to support biological evolution. [LO 1.9, SP 5.3] Refine evidence based on data from many scientific disciplines
Valley Central School District 944 State Route 17K Montgomery, NY Telephone Number: (845) ext Fax Number: (845)
 Valley Central School District 944 State Route 17K Montgomery, NY 12549 Telephone Number: (845)457-2400 ext. 18121 Fax Number: (845)457-4254 Advance Placement Biology Presented to the Board of Education
Valley Central School District 944 State Route 17K Montgomery, NY 12549 Telephone Number: (845)457-2400 ext. 18121 Fax Number: (845)457-4254 Advance Placement Biology Presented to the Board of Education
I. Molecules & Cells. A. Unit One: The Nature of Science. B. Unit Two: The Chemistry of Life. C. Unit Three: The Biology of the Cell.
 I. Molecules & Cells A. Unit One: The Nature of Science a. How is the scientific method used to solve problems? b. What is the importance of controls? c. How does Darwin s theory of evolution illustrate
I. Molecules & Cells A. Unit One: The Nature of Science a. How is the scientific method used to solve problems? b. What is the importance of controls? c. How does Darwin s theory of evolution illustrate
AP* Biology Prep Course
 AP* Biology Prep Course SYLLABUS Welcome to the FlinnPREP AP* Biology Online Prep Course! Your enrollment in this course is your first step toward a 5 on the AP* Biology exam. FlinnPREP covers fundamental
AP* Biology Prep Course SYLLABUS Welcome to the FlinnPREP AP* Biology Online Prep Course! Your enrollment in this course is your first step toward a 5 on the AP* Biology exam. FlinnPREP covers fundamental
Biology Teach Yourself Series Topic 2: Cells
 Biology Teach Yourself Series Topic 2: Cells A: Level 14, 474 Flinders Street Melbourne VIC 3000 T: 1300 134 518 W: tssm.com.au E: info@tssm.com.au TSSM 2013 Page 1 of 14 Contents Cells... 3 Prokaryotic
Biology Teach Yourself Series Topic 2: Cells A: Level 14, 474 Flinders Street Melbourne VIC 3000 T: 1300 134 518 W: tssm.com.au E: info@tssm.com.au TSSM 2013 Page 1 of 14 Contents Cells... 3 Prokaryotic
A game-theoretical approach to the analysis of microbial metabolic pathways
 A game-theoretical approach to the analysis of microbial metabolic pathways Stefan Schuster Dept. of Bioinformatics Friedrich Schiller University Jena, Germany Introduction Widely used hypothesis: During
A game-theoretical approach to the analysis of microbial metabolic pathways Stefan Schuster Dept. of Bioinformatics Friedrich Schiller University Jena, Germany Introduction Widely used hypothesis: During
Signal Transduction: Ethylene PSI AP Biology
 Signal Transduction: Ethylene PSI AP Biology Name: Objective Students will analyze the role ethylene plays as a plant hormone in the signal transduction pathway of fruit ripening. Students will use their
Signal Transduction: Ethylene PSI AP Biology Name: Objective Students will analyze the role ethylene plays as a plant hormone in the signal transduction pathway of fruit ripening. Students will use their
Chemistry Chapter 26
 Chemistry 2100 Chapter 26 The Central Dogma! The central dogma of molecular biology: Information contained in DNA molecules is expressed in the structure of proteins. Gene expression is the turning on
Chemistry 2100 Chapter 26 The Central Dogma! The central dogma of molecular biology: Information contained in DNA molecules is expressed in the structure of proteins. Gene expression is the turning on
Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes. - Supplementary Information -
 Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes - Supplementary Information - Martin Bartl a, Martin Kötzing a,b, Stefan Schuster c, Pu Li a, Christoph Kaleta b a
Dynamic optimisation identifies optimal programs for pathway regulation in prokaryotes - Supplementary Information - Martin Bartl a, Martin Kötzing a,b, Stefan Schuster c, Pu Li a, Christoph Kaleta b a
Biology Unit Overview and Pacing Guide
 This document provides teachers with an overview of each unit in the Biology curriculum. The Curriculum Engine provides additional information including knowledge and performance learning targets, key
This document provides teachers with an overview of each unit in the Biology curriculum. The Curriculum Engine provides additional information including knowledge and performance learning targets, key
The application of an artificial immune system for solving the identification problem
 IT Web of Conferences 9, 3 (17) DOI: 1.151/ itmconf/1793 ACSE 16 The application of an artificial immune system for solving the identification problem Irina Astachova 1, Stanislav Ushakov 1, Andrei Selemenev
IT Web of Conferences 9, 3 (17) DOI: 1.151/ itmconf/1793 ACSE 16 The application of an artificial immune system for solving the identification problem Irina Astachova 1, Stanislav Ushakov 1, Andrei Selemenev
Range of Competencies
 BIOLOGY Content Domain Range of Competencies l. Nature of Science 0001 0003 20% ll. Biochemistry and Cell Biology 0004 0005 13% lll. Genetics and Evolution 0006 0009 27% lv. Biological Unity and Diversity
BIOLOGY Content Domain Range of Competencies l. Nature of Science 0001 0003 20% ll. Biochemistry and Cell Biology 0004 0005 13% lll. Genetics and Evolution 0006 0009 27% lv. Biological Unity and Diversity
Course Syllabus. Department: Science and Technology. Date: February 3, I. Course Prefix and Number: BIO 122. Course Name: General Biology II
 Department: Science and Technology Date: February 3, 2012 I. Course Prefix and Number: BIO 122 Course Name: General Biology II Course Syllabus Credit Hours and Contact Hours: 4 Credit hours and 5 Contact
Department: Science and Technology Date: February 3, 2012 I. Course Prefix and Number: BIO 122 Course Name: General Biology II Course Syllabus Credit Hours and Contact Hours: 4 Credit hours and 5 Contact
West Windsor-Plainsboro Regional School District AP Biology Grades 11-12
 West Windsor-Plainsboro Regional School District AP Biology Grades 11-12 Unit 1: Chemistry of Life Content Area: Science Course & Grade Level: AP Biology, 11 12 Summary and Rationale The structural levels
West Windsor-Plainsboro Regional School District AP Biology Grades 11-12 Unit 1: Chemistry of Life Content Area: Science Course & Grade Level: AP Biology, 11 12 Summary and Rationale The structural levels
TEACHER CERTIFICATION STUDY GUIDE. Table of Contents I. BASIC PRINCIPLES OF SCIENCE (HISTORY AND NATURAL SCIENCE)
 Table of Contents I. BASIC PRINCIPLES OF SCIENCE (HISTORY AND NATURAL SCIENCE) A. Nature of scientific knowledge, inquiry, and historical perspectives 1. Scientific methods...1 2. Processes involved in
Table of Contents I. BASIC PRINCIPLES OF SCIENCE (HISTORY AND NATURAL SCIENCE) A. Nature of scientific knowledge, inquiry, and historical perspectives 1. Scientific methods...1 2. Processes involved in
Section 1: Discovering Cells
 Section 1: Discovering Cells Cells Definition: The basic unit of structure and function in living things All cells have a membrane protecting them All cells contain DNA and RNA Cells die through apoptosis
Section 1: Discovering Cells Cells Definition: The basic unit of structure and function in living things All cells have a membrane protecting them All cells contain DNA and RNA Cells die through apoptosis
Page 1. Name: UNIT: PHOTOSYNTHESIS AND RESPIRATION TOPIC: PHOTOSYNTHESIS
 Name: 4667-1 - Page 1 UNIT: PHOTOSYNTHESIS AND RESPIRATION TOPIC: PHOTOSYNTHESIS 1) The diagram below illustrates the movement of materials involved in a process that is vital for the energy needs of organisms.
Name: 4667-1 - Page 1 UNIT: PHOTOSYNTHESIS AND RESPIRATION TOPIC: PHOTOSYNTHESIS 1) The diagram below illustrates the movement of materials involved in a process that is vital for the energy needs of organisms.
Biological networks CS449 BIOINFORMATICS
 CS449 BIOINFORMATICS Biological networks Programming today is a race between software engineers striving to build bigger and better idiot-proof programs, and the Universe trying to produce bigger and better
CS449 BIOINFORMATICS Biological networks Programming today is a race between software engineers striving to build bigger and better idiot-proof programs, and the Universe trying to produce bigger and better
SINGLE-ELECTRON CIRCUITS PERFORMING DENDRITIC PATTERN FORMATION WITH NATURE-INSPIRED CELLULAR AUTOMATA
 International Journal of Bifurcation and Chaos, Vol. 7, No. (7) 365 3655 c World Scientific Publishing Company SINGLE-ELECTRON CIRCUITS PERFORMING DENDRITIC PATTERN FORMATION WITH NATURE-INSPIRED CELLULAR
International Journal of Bifurcation and Chaos, Vol. 7, No. (7) 365 3655 c World Scientific Publishing Company SINGLE-ELECTRON CIRCUITS PERFORMING DENDRITIC PATTERN FORMATION WITH NATURE-INSPIRED CELLULAR
Computer Simulation and Data Analysis in Molecular Biology and Biophysics
 Victor Bloomfield Computer Simulation and Data Analysis in Molecular Biology and Biophysics An Introduction Using R Springer Contents Part I The Basics of R 1 Calculating with R 3 1.1 Installing R 3 1.1.1
Victor Bloomfield Computer Simulation and Data Analysis in Molecular Biology and Biophysics An Introduction Using R Springer Contents Part I The Basics of R 1 Calculating with R 3 1.1 Installing R 3 1.1.1
Tutorials are designed specifically for the Virginia Standards of Learning to prepare students for the Standards of Learning tests.
 Tutorial Outline Tutorials are designed specifically for the Virginia Standards of Learning to prepare students for the Standards of Learning tests. Biology Tutorials offer targeted instruction, practice,
Tutorial Outline Tutorials are designed specifically for the Virginia Standards of Learning to prepare students for the Standards of Learning tests. Biology Tutorials offer targeted instruction, practice,
AP Biology Essential Knowledge Cards BIG IDEA 1
 AP Biology Essential Knowledge Cards BIG IDEA 1 Essential knowledge 1.A.1: Natural selection is a major mechanism of evolution. Essential knowledge 1.A.4: Biological evolution is supported by scientific
AP Biology Essential Knowledge Cards BIG IDEA 1 Essential knowledge 1.A.1: Natural selection is a major mechanism of evolution. Essential knowledge 1.A.4: Biological evolution is supported by scientific
Text of objective. Investigate and describe the structure and functions of cells including: Cell organelles
 This document is designed to help North Carolina educators teach the s (Standard Course of Study). NCDPI staff are continually updating and improving these tools to better serve teachers. Biology 2009-to-2004
This document is designed to help North Carolina educators teach the s (Standard Course of Study). NCDPI staff are continually updating and improving these tools to better serve teachers. Biology 2009-to-2004
TEST SUMMARY AND FRAMEWORK TEST SUMMARY
 Washington Educator Skills Tests Endorsements (WEST E) TEST SUMMARY AND FRAMEWORK TEST SUMMARY BIOLOGY Copyright 2014 by the Washington Professional Educator Standards Board 1 Washington Educator Skills
Washington Educator Skills Tests Endorsements (WEST E) TEST SUMMARY AND FRAMEWORK TEST SUMMARY BIOLOGY Copyright 2014 by the Washington Professional Educator Standards Board 1 Washington Educator Skills
Science Course Descriptions
 BIOLOGY I (L) 3024 (BIO I) Biology I is a course based on the following core topics: cellular chemistry, structure and reproduction; matter cycles and energy transfer; interdependence of organisms; molecular
BIOLOGY I (L) 3024 (BIO I) Biology I is a course based on the following core topics: cellular chemistry, structure and reproduction; matter cycles and energy transfer; interdependence of organisms; molecular
Course Descriptions Biology
 Course Descriptions Biology BIOL 1010 (F/S) Human Anatomy and Physiology I. An introductory study of the structure and function of the human organ systems including the nervous, sensory, muscular, skeletal,
Course Descriptions Biology BIOL 1010 (F/S) Human Anatomy and Physiology I. An introductory study of the structure and function of the human organ systems including the nervous, sensory, muscular, skeletal,
Information Dynamics Foundations and Applications
 Gustavo Deco Bernd Schürmann Information Dynamics Foundations and Applications With 89 Illustrations Springer PREFACE vii CHAPTER 1 Introduction 1 CHAPTER 2 Dynamical Systems: An Overview 7 2.1 Deterministic
Gustavo Deco Bernd Schürmann Information Dynamics Foundations and Applications With 89 Illustrations Springer PREFACE vii CHAPTER 1 Introduction 1 CHAPTER 2 Dynamical Systems: An Overview 7 2.1 Deterministic
East Penn School District Curriculum and Instruction
 East Penn School District Curriculum and Instruction Curriculum for: Biology 1, Applied/CP/Honors Course(s): Biology 1 Grades: 9 and 10 Department: Science Length of Period (average minutes): 42 Periods
East Penn School District Curriculum and Instruction Curriculum for: Biology 1, Applied/CP/Honors Course(s): Biology 1 Grades: 9 and 10 Department: Science Length of Period (average minutes): 42 Periods
Advances in Cyclic Structural Causal Models
 Advances in Cyclic Structural Causal Models Joris Mooij j.m.mooij@uva.nl June 1st, 2018 Joris Mooij (UvA) Rotterdam 2018 2018-06-01 1 / 41 Part I Introduction to Causality Joris Mooij (UvA) Rotterdam 2018
Advances in Cyclic Structural Causal Models Joris Mooij j.m.mooij@uva.nl June 1st, 2018 Joris Mooij (UvA) Rotterdam 2018 2018-06-01 1 / 41 Part I Introduction to Causality Joris Mooij (UvA) Rotterdam 2018
Grade Level: AP Biology may be taken in grades 11 or 12.
 ADVANCEMENT PLACEMENT BIOLOGY COURSE SYLLABUS MRS. ANGELA FARRONATO Grade Level: AP Biology may be taken in grades 11 or 12. Course Overview: This course is designed to cover all of the material included
ADVANCEMENT PLACEMENT BIOLOGY COURSE SYLLABUS MRS. ANGELA FARRONATO Grade Level: AP Biology may be taken in grades 11 or 12. Course Overview: This course is designed to cover all of the material included
Deterministic Nonlinear Modeling of Ant Algorithm with Logistic Multi-Agent System
 Deterministic Nonlinear Modeling of Ant Algorithm with Logistic Multi-Agent System Rodolphe Charrier, Christine Bourjot, François Charpillet To cite this version: Rodolphe Charrier, Christine Bourjot,
Deterministic Nonlinear Modeling of Ant Algorithm with Logistic Multi-Agent System Rodolphe Charrier, Christine Bourjot, François Charpillet To cite this version: Rodolphe Charrier, Christine Bourjot,
PREREQUISITE CHECKLIST
 PREREQUISITE CHECKLIST UNIVERSITY OF CALIFORNIA, BERKELEY SCHOOL OF OPTOMETRY ADMISSIONS AND STUDENT AFFAIRS OFFICE Name: Date: Email: Status (complete, in progress, or planned) Prerequisite Course Requirements
PREREQUISITE CHECKLIST UNIVERSITY OF CALIFORNIA, BERKELEY SCHOOL OF OPTOMETRY ADMISSIONS AND STUDENT AFFAIRS OFFICE Name: Date: Email: Status (complete, in progress, or planned) Prerequisite Course Requirements
Miller & Levine Biology 2014
 A Correlation of Miller & Levine Biology To the Essential Standards for Biology High School Introduction This document demonstrates how meets the North Carolina Essential Standards for Biology, grades
A Correlation of Miller & Levine Biology To the Essential Standards for Biology High School Introduction This document demonstrates how meets the North Carolina Essential Standards for Biology, grades
How data assimilation helps to illuminate complex biology
 How data assimilation helps to illuminate complex biology The dynamic elastic-net Maik Kschischo Department of Mathematics and Technology University of Applied Sciences Koblenz RheinAhrCampus Remagen Joseph-Rovan-Allee
How data assimilation helps to illuminate complex biology The dynamic elastic-net Maik Kschischo Department of Mathematics and Technology University of Applied Sciences Koblenz RheinAhrCampus Remagen Joseph-Rovan-Allee
Basic Biology. Content Skills Learning Targets Assessment Resources & Technology
 Teacher: Lynn Dahring Basic Biology August 2014 Basic Biology CEQ (tri 1) 1. What are the parts of the biological scientific process? 2. What are the essential molecules and elements in living organisms?
Teacher: Lynn Dahring Basic Biology August 2014 Basic Biology CEQ (tri 1) 1. What are the parts of the biological scientific process? 2. What are the essential molecules and elements in living organisms?
