The secreted cell signal Folded Gastrulation regulates glial morphogenesis and axon guidance in Drosophila
|
|
|
- Juliana Cummings
- 5 years ago
- Views:
Transcription
1 Developmental Biology 308 (2007) The secreted cell signal Folded Gastrulation regulates glial morphogenesis and axon guidance in Drosophila Anuradha Ratnaparkhi 1, Kai Zinn Broad Center, Division of Biology, California Institute of Technology, Pasadena, CA 91125, USA Received for publication 14 February 2007; revised 11 May 2007; accepted 16 May 2007 Available online 24 May 2007 Abstract During gastrulation in Drosophila, ventral cells change shape, undergoing synchronous apical constriction, to create the ventral furrow (VF). This process is affected in mutant embryos lacking zygotic function of the folded gastrulation (fog) gene, which encodes a putative secreted protein. Fog is an essential autocrine signal that induces cytoskeletal changes in invaginating VF cells. Here we show that Fog is also required for nervous system development. Fog is expressed by longitudinal glia in the central nervous system (CNS), and reducing its expression in glia causes defects in process extension and axon ensheathment. Glial Fog overexpression produces a disorganized glial lattice. Fog has a distinct set of functions in CNS neurons. Our data show that reduction or overexpression of Fog in these neurons produces axon guidance phenotypes. Interestingly, these phenotypes closely resemble those seen in embryos with altered expression of the receptor tyrosine phosphatase PTP52F. We conducted epistasis experiments to define the genetic relationships between Fog and PTP52F, and the results suggest that PTP52F is a downstream component of the Fog signaling pathway in CNS neurons. We also found that Ptp52F mutants have early VF phenotypes like those seen in fog mutants Elsevier Inc. All rights reserved. Keywords: Drosophila nervous system development; Receptor tyrosine phosphatase; Glia; Motor axon guidance; Epistasis; Ventral furrow development; Autocrine signaling; Folded Gastrulation Introduction Corresponding author. Fax: addresses: aratnaparkhi@mednet.ucla.edu (A. Ratnaparkhi), zinnk@caltech.edu (K. Zinn). 1 Present address: Department of Neurology and Brain Research Institute, David Geffen School of Medicine at UCLA, 4335 Gonda Center, 695 Charles E. Young Drive South, Los Angeles, CA , USA. During gastrulation in Drosophila, two groups of cells invaginate and form the primordia of the mesoderm and endoderm. Cells located within a strip around the ventral midline change shape, undergoing synchronous apical constriction, to create the ventral furrow (VF). At the posterior end, apical constriction of a circular group of cells forms the posterior midgut (PMG) invagination, which encloses the pole cells (Sweeton et al., 1991; Costa et al., 1993). The mechanisms involved in the induction of cell shape changes during gastrulation are not well understood. However, several mutations have been identified that affect these processes. Embryos lacking zygotic function of the folded gastrulation (fog) gene undergo asynchronous changes in cell shape, resulting in an irregular VF, and do not form the PMG invagination (Costa et al., 1994). Identical phenotypes are observed in embryos lacking maternally contributed concertina (cta) gene function (Parks and Wieschaus, 1991). fog encodes a large protein with an N terminal hydrophobic region that resembles a signal sequence, suggesting that at least part of it is secreted. It is expressed by the cells that will invaginate to form the VF and PMG (Costa et al., 1994). Cta is a G protein alpha subunit, and it is uniformly distributed within the blastoderm embryo (Parks and Wieschaus, 1991). The similarities in the loss-of-function (LOF) and gain-of-function (GOF) phenotypes of fog and cta, and the observation that cta is epistatic to fog, have led to the proposal that Fog is a ligand for an as yet unidentified G protein-coupled receptor (GPCR) that couples to Cta (Morize et al., 1998). The mechanisms involved in Fog signaling have been examined primarily in the context of gastrulation. It has been /$ - see front matter 2007 Elsevier Inc. All rights reserved. doi: /j.ydbio
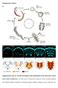
2 A. Ratnaparkhi, K. Zinn / Developmental Biology 308 (2007) shown that Fog is an autocrine signal that induces cytoskeletal changes in invaginating VF cells (Dawes-Hoang et al., 2005). Exposure to the Fog signal causes translocation of nonmuscle myosin from the basal to the apical cell surface, and this facilitates apical constriction of VF cells (Dawes-Hoang et al., 2005). Fog is also required for development of the salivary glands and Malphigian tubules (Lammel and Saumweber, 2000). Our laboratory has focused on examining the functions of receptor tyrosine phosphatases (RPTPs) in regulation of CNS and motor axon guidance during Drosophila embryogenesis. Like the other RPTPs, PTP52F is expressed primarily in the embryonic CNS. However, in early embryos, expression of the protein is also observed in cells surrounding the VF (Schindelholz et al., 2001). More recently, we discovered that Ptp52F mutants have fog-like VF phenotypes (Supplementary Fig. 2; A. Schmid, B. Schindelholz, and K.Z., unpublished). The similarity in VF phenotype between fog and Ptp52F led us to explore the possibility of an interaction between these two genes in the embryonic CNS. In this paper, we characterize the expression of Fog in the embryonic CNS. We show that Fog is expressed in glia and neurons and has distinct roles in the two cell types. Fog expressed by longitudinal glia is required for glial ensheathment of CNS axons. Neuronal Fog regulates axon guidance, and Fog signaling in neurons requires the PTP52F receptor. Materials and methods Fly stocks fog 4a6 and fog 4a6,hkb::fog flies were obtained from the laboratories of E. Wieschaus and M Leptin, respectively. EP52F, an EP insertion upstream of the Ptp52F gene, was obtained from Florenci Serras. Ptp52F 18.3, a null allele of Ptp52F (Schindelholz et al., 2001), was used for the epistasis experiment of Fig. 5. For the heat shock experiment of Fig. 1, an hs-gal4, UAS-GFP line was used. C155-GAL4, elav-gal4 and Repo-GAL4 were obtained from the Bloomington stock center. Htl-GAL4 was obtained from Alicia Hidalgo. Generation of transgenic lines The 1.6 kb full-length Fog cdna was cloned into the puast vector and the plasmid DNA was injected into 0 1 h embryos using standard methods. Genetic crosses were carried out to establish stable transgenic lines. To obtain UAS-fog dsrna transgenic lines, an inverted repeat sequence of the fog cdna corresponding to amino acids was first generated as follows: using two independent 5 primers and a common 3 primer, two kinds of PCR products were generated. The 5 primers were designed such that they carried either a XhoI oraxbai site at the 5 end. The common 3 primer was Fig. 1. Expression of fog mrna and protein in dissected stage 16 embryos. (A and B) fog mrna expression in the CNS, detected by in situ hybridization using alkaline phosphatase (AP) histochemistry. The CNS (anterior up) is vertically oriented at the center of the panels; four segments are shown. (A) fog mrna (blue) is expressed in a subset of CNS cells (arrows). Note that the pattern is not precisely bilaterally symmetric and is not entirely reproducible between segments. (B) fog in situ hybridization co-stained with anti-repo antibody (brown). Most fog positive cells also express Repo (arrows), but some are Repo-negative (asterisks). (C and D) The CNS in a wild-type (C and C ) and a hs-gal4::uas-dsfogrna (fog RNAi) (D and D ) embryo stained with anti-fog antibody (red) and mabbp102 (green), which is a marker for CNS axons. Fog is expressed at highest levels in the CNS axon ladder (C, arrow). Midline glia are indicated by an asterisk in penel C. Axonal staining is reduced in fog RNAi embryos (D, arrow). (E) fog mrna expression (blue, arrow) in peripheral chordotonal organs (also stained with mab22c10 (brown), which is a marker for peripheral sensory neurons (asterisks indicate chordotonal neuron cell bodies)). fog expression is observed in the region of the scolopale and cap cells (arrows). Anterior is to the left in panels E G. (F) Expression of Fog protein (red) in chordotonal organs, detected by immunofluorescence. Protein is observed in the dendrites of the sensory neurons (arrows), scolopale cells (bulb-like shapes at the end of the dendrites, yellow arrowheads), and cap cells (white arrowheads). (G) Chordotonal organs stained with anti-fog (red) in the absence of detergent. Expression in the dendritic shaft of the sensory neuron (arrows), the scolopales (yellow arrowheads), and the cap cells (white arrowheads) can be detected, indicating that Fog is present at the cell surface. The asterisks mark the cell bodies of the sensory neurons. Note that background staining is higher when this method is used. Scale bar for panels A and B is in panel A and corresponds to 10 μm, for panels C Fitisin panel C and corresponds to 10 μm, and the scale bar in panel G corresponds to 25 μm.
3 160 A. Ratnaparkhi, K. Zinn / Developmental Biology 308 (2007) designed with a BamHI site followed by an EcoRI site at the 5 end. The first PCR product was cloned into pbluescript using the XbaI and EcoRI sites; the second product was cloned in the opposite orientation using the BamHI present in the first PCR product and the XhoI site in pbluescript. The resulting inverted repeat sequence was then subcloned into puast using the XhoI and XbaI sites and transformed using Stbl-2 cells (Invitrogen). The puast construct was then injected into 0 1 h embryos using standard methods to obtain transgenic lines. Several such lines were analyzed, and all had equivalent phenotypes. The neuronal and glial RNAi experiments were carried out at room temperature (23 C) and 29 C, respectively. In situ hybridization and immunohistochemistry In situ hybridization was carried out using sense and anti-sense digoxigenin labeled RNA as probes. The hybridization and detection procedures used have been previously described in Kraut et al. (2001). No signal was detected with the sense probe. Immunohistochemical analysis was carried out on embryos collected from crosses between flies of the appropriate genotypes. Embryos of the desired genotype were identified by the absence of staining of rabbit anti-betagalactosidase (Invitrogen), which detects the presence of the lacz gene on the balancer chromosome. The following antibodies were used: anti-fog raised against the full-length protein (1:100 gift of N. Fuse), anti-fog (N terminal antibody) raised against the first 300 amino acids (1:50, gift of the Wieschaus lab), chicken anti-gfp (1:500, Chemicon), anti-twist (1:5000, gift of S. Roth). Monoclonal antibodies mab 1D4, mab 22C10 and anti-repo (Developmental Studies Hybridoma Bank, Iowa, United States) were used at 1:10, 1:50 and 1:50 respectively. Embryo fixation and immunohistochemistry using horseradish preoxidase and fluorescence were performed as described in Patel (1994). All embryos stained using horseradish peroxidase were photographed with a Magnafire digital camera on a Zeiss Axioplan microscope using Nomarski optics. For the side view of the CNS in Fig. 2, dissected CNS of stained embryos were mounted on their side and images were taken as a z-series. A projected
4 A. Ratnaparkhi, K. Zinn / Developmental Biology 308 (2007) image from the z-series was obtained using the Zeiss confocal software. Subsequent processing was done using Adobe Photoshop 7.0. Fog staining in the CNS was carried out on live-dissected and fixed embryos. The dissected embryos were fixed for 20 min with 4% paraformaldehyde on ice. Blocking was done with PBTX containing 5% normal goat serum. Antibody staining was carried out using 1:100 dilution of the antibody. To detect secreted Fog protein, live-dissected embryos were incubated with antibody for 2 h at room temperature. The antibody was diluted in PBS containing 0.5% normal goat serum. After incubation with the antibody, the embryos were washed at least 3 times with PBS and fixed with 4% paraformaldehyde. Subsequent detection of the antibody was carried out using standard procedures (Patel, 1994) embryos. The embryos were fixed for Alexa 488 or Alexa 568 secondary antibodies (Molecular Probes) were used for all fluorescent immunohistochemistry. Images were processed using Adobe Photoshop 7. Quantitation of glial cell surface areas was carried out using ImageJ software (NIH). Results fog is expressed in longitudinal glia and neurons To evaluate the expression pattern of the fog gene in the embryonic CNS, we performed in situ hybridization experiments using fog cdna probes. We found that fog mrna is uniformly expressed in germ band extended embryos (stages 10 and 11), but then becomes restricted to small groups of cells later in development. fog mrna expression within the CNS is most easily observed at stage 15 or later. Most of the cells expressing fog at high levels are located within narrow zones that flank the midline and extend from the anterior to the posterior of the embryo (Fig. 1A). These zones are two or three cells wide. Interestingly, however, not all cells within the zones express fog mrna at levels above the detection threshold, and the pattern is not identical between segments or between embryos. This suggests that expression levels in individual cells might be determined in a stochastic manner. To determine the identities of the fog-expressing cells, we double-stained embryos with fog probe (blue) and antibodies against glial and neuronal markers. We found that most of the prominent (high-level) fog-expressing cells in the CNS also stained for Repo, a transcription factor expressed in longitudinal glia (brown in Fig. 1B, arrows) (Xiong et al., 1994; Halter et al., 1995). There were also Repo-negative, fog-mrna-positive cells within the CNS (Fig. 1B, asterisk). Some of these are located near the midline and might be midline glia, which stain with antibodies against Fog protein (Fig. 1C). Our results on expression of Fog protein (Figs. 1C G, Supplementary Fig. 1) indicate that it is expressed by some CNS and peripheral neurons as well; however, the levels of fog mrna in neurons may be below the detection threshold for in situ hybridization. In the periphery, strong fog mrna expression was observed in the scolopale and cap cells, which ensheath the dendrites of the sensory neurons in the chordotonal organs (Fig. 1E). To study localization of Fog protein, we stained live-dissected stage 16 embryos using an antibody generated against full-length Fog (a gift from N. Fuse) and detected expression by immunofluorescence. We observed that Fog protein was localized to the scolopale and cap cells, consistent with the in situ hybridization results, and was also seen on the dendrites of chordotonal neurons (Fig. 1F). Background staining levels are high with this antibody. This is commonly observed with antibodies against secreted proteins, particularly when they must be used at high concentrations (1:100 in this case). Within the CNS, the strongest staining with anti-fog antibody was observed on the longitudinal axonal tracts, which are visualized with the axonal marker BP102 (Figs. 1C and C ). In order to confirm that this axonal staining represents genuine Fog protein, we utilized RNAi techniques, expressing fog double-stranded RNA (dsrna) in embryos using a heat shock promoter-gal4 driver (fog RNAi embryos; see below). Embryos that were heat shocked displayed decreased axonal staining with anti-fog (Figs. 1D and D ), while those that were not subjected to heat shock showed strong Fog expression in the CNS. We also expressed fog dsrna with pan-neuronal (Elav) and longitudinal glial (Repo) drivers and found that in neuronal fog RNAi embryos, CNS axonal staining was reduced, while glial fog RNAi did not strongly affect staining. In the periphery, neuronal fog RNAi eliminated staining of the dendritic shafts of Fig. 2. Glial morphology and organization are altered in fog glial RNAi, fog mutant, and Fog overexpression embryos. (A, B, E, F) Five segments of the CNS in late stage 16 embryos are shown, stained with anti-gfp to reveal the morphology of mcd8-gfp expressing glia; anterior is up. Panels C D and G I are lateral views of the CNS; anterior is to the left and dorsal is up. (A) Htl-GAL4::UAS-mCD8GFP. The longitudinal glia that express htl are organized in two rows on either side of the midline. The cells appear flat and spread out. (B) Htl-GAL4::UAS-mCD8GFP, UAS-dsfogRNA (glial fog RNAi). The glial lattice is disrupted, with gaps appearing in glial rows (arrow). Glia appear smaller and rounder, and fingerlike projections are observed around their edges; the gap between rows at the midline is wider, primarily because the glia are smaller. (C and D) Lateral view of the ventral nerve cord of late stage 16 embryos stained with mab BP102 (red), which marks the axon scaffold, and anti-gfp (green) to reveal the glial extensions of mcd8-gfp expressing glia. d, dorsal; v, ventral. (C) Htl-GAL4::UAS-mCD8GFP. Arrow indicates the longitudinal glial cell bodies overlying the axon scaffold (red). Glial processes are seen enveloping the axon bundles and glial staining is observed through the entire depth (d v axis) of the nerve cord. (D) Htl-GAL4::UAS-mCD8GFP, UAS-dsfogRNA (glial fog RNAi). Glial ensheathment of the nerve cord by longitudinal glia (arrow indicates cell bodies) is strongly affected. There is only a partial wrapping of glial processes around the red-labeled CNS axon scaffold. Glial staining in the region ventral to the axon layer is dramatically reduced (asterisk). (E) Repo-GAL4::UAS-mCD8GFP. The longitudinal glia have a somewhat different appearance than when mcd8-gfp is driven with Htl-GAL4, but the two rows of longitudinal glia are clearly observed on either side of the midline (arrows). Glial processes spanning the midline (asterisks) are prominent as are exit glia ensheathing the peripheral nerve roots (arrowhead). This driver provides better visualization of the exit glia than does Htl-GAL4. (F) Repo-GAL4::UAS-mCD8GFP, UAS-fog (glial Fog overexpression). The longitudinal glial lattice is disorganized, as are the exit glia (arrowhead). (G I) Lateral view of the ventral nerve cord of late stage 16 embryos stained with anti-htl antibody. (G) Wild type. Htl staining is observed in a dorsal zone (arrow) made up of longitudinal glia that ensheath the neuropil, and a ventral zone (asterisk) made up of glia and glial processes that extend below the nerve cord. (H) fog; hkb::fog. Htl staining in the dorsal zone (arrow) appears disorganized and its expression is almost absent in the ventral zone (asterisk). Overall Htl staining intensity is also reduced relative to wild type. (I) Htl-GAL4::UAS-mCD8GFP, UAS-dsfogRNA (glial fog RNAi). As in the fog mutant, Htl staining in the dorsal zone appears disorganized (arrow), Htl is very weak in the ventral zone (asterisk), and overall staining intensity is reduced. Scale bar in A and E is 10 μm. Scale bar in panel A applies to panels A D and G I. Scale bar in panel E applies to panels E and F.
5 162 A. Ratnaparkhi, K. Zinn / Developmental Biology 308 (2007) the chordotonal neurons (Supplementary Fig. 1). Expression in the scolopale and cap cells, however, appears to be resistant to both neuronal and heat-shock driven RNAi. It is also resistant to glial RNAi (not shown); this is expected since the scolopale and cap cells do not express Repo. In order to evaluate phenotypic effects observed with glial fog RNAi, we needed to demonstrate that glial RNAi could inhibit Fog protein expression. Since Fog staining in the CNS apparently derives from both glia and the neurons they ensheath, and Fog is a secreted protein, effects of glial RNAi on staining with anti-fog antibody are likely to be obscured by neuronal Fog expression. We thus examined Fog protein staining in the retinal basal glia of the larval eye disc and optic stalk, where we had observed that glial Fog expression can be spatially distinguished from expression in nearby neurons. We compared Fog staining between control larvae and glial fog RNAi larvae and observed that staining in retinal basal glia (visualized with Repo-GAL4-driven mcd8-gfp) was reduced or eliminated in RNAi larvae (Supplementary Fig. 1). The residual punctate staining in the glial region in the RNAi larvae probably represents the usual background observed with anti- Fog antibody. Since Repo-positive glia comprise most of the high-level fog mrna-expressing cells in the CNS, we wondered whether some of this axonal Fog might be transferred from glia to the axons. To examine this question, we stained gcm 7-4 mutant embryos (Jones et al., 1995), in which most CNS glia are absent, with anti-fog. We found that axonal Fog staining was unchanged (data not shown). Together with the RNAi data, these results indicate that axonal Fog is at least partially derived from Fog expressed in neurons. The N terminus of the predicted 730 amino acid (aa) Fog protein resembles a signal sequence, and it has no obvious transmembrane domain. This is consistent with the prediction that Fog is a secreted ligand (Costa et al., 1994). It has also been demonstrated that Fog is transported through the secretory pathway (Dawes-Hoang et al., 2005). However, it has never been directly shown that Fog is extracellularly localized. To determine if Fog protein can be detected at or outside of the cell surface, we carried out immunofluorescent staining of unfixed Fig. 3. Motor axon phenotypes in fog and Ptp52F mutants. Late stage 16 embryo fillets stained with mab1d4 (brown) to reveal motor axons and their projections. Two or three abdominal segments are shown in panels B E. (A) Schematic representation of SNa bifurcation. The SNa nerve (red) branches at the dorsal edge of muscle 12. The anterior (dorsal) branch innervates muscles 21 24, and the posterior (lateral) branch innervates muscles 5 and 8. The pink nerve on the left is the ISN. (B) Wild-type (Oregon R) embryo. Arrow indicates bifurcation point. (C) fog 4a6,hkb::fog mutant embryo. The posterior branch is missing in the right-hand segment (asterisk). An extra branch emerges from the anterior branch in the middle segment (arrow). (D) Elav-GAL4::UAS-dsfogRNA (Neuronal fog RNAi) embryo. The anterior branch is missing in the middle segment (asterisk). In the right-hand segment, the anterior branch is truncated (arrow). (E) Ptp52F 18.3 mutant embryo. The posterior branch is missing in the middle segment (asterisk). An extra branch is seen in the left-hand segment (arrow). Scale bar in panel B, 10 μm (applies to all panels).
6 A. Ratnaparkhi, K. Zinn / Developmental Biology 308 (2007) Fig. 4. CNS phenotypes in fog and Ptp52F mutants. Each panel shows four segments of a late stage 16 embryo fillet stained with mab 1D4 (brown); anterior is up. Scale bar in panel A, 10 μm (applies to all panels). (A) Wild-type (Oregon R) embryo. Note the three straight 1D4-positive longitudinal axon bundles on each side of the midline. (B) fog 4a6,hkb::fog mutant embryo. There are breaks in the outer bundle (arrows), and fusion of bundles is sometimes observed (arrowhead). (C) Elav- GAL4::UAS-dsfogRNA (neuronal fog RNAi) embryo. A similar, but usually milder phenotype, is observed, in which the outer bundle is discontinuous (arrows). (D) Ptp52F 18.3 mutant embryo. The outer and middle 1D4-positive bundles are both disorganized. The outer bundle is also discontinuous (arrow) and is often fused to the middle bundle (arrowhead). live-dissected embryos in the absence of detergent. We observed expression of Fog on scolopale cells (Fig. 1F), indicating that Fog is indeed secreted or localized to the cell surface, at least in the periphery. Within the CNS, however, the high background staining observed under these conditions prevented us from clearly determining whether axonal Fog is also extracellular. Reduction or overexpression of Fog in glia affects morphology and axon ensheathment The results described above (Fig. 1) show that within the embryonic CNS fog mrna is expressed primarily by Repopositive glia. To evaluate the function of Fog in these glia, and to separate glial fog LOF phenotypes from those conferred by loss of Fog in neurons (see below), we used glial drivers to coexpress fog dsrna together with mcd8-gfp, a membranetethered form of GFP, so that we could directly visualize glial morphology. Several independent UAS-fog dsrna transgenic lines were examined, and all produced the same phenotypes. To confirm the results obtained using RNAi techniques, we also examined fog LOF mutants in which early embryonic lethality caused by lack of PMG invagination (the folded phenotype) is rescued via expression of Fog in the posterior midgut region (PMG) from the huckebein (hkb) promoter (fog, hkb::fog; a gift of Maria Leptin). These embryos have near normal morphology at later stages of development (M. Leptin, personal communication). In particular, the ventrally located CNS develops normally in these embryos. When Fog dsrna was expressed using heartless (Htl)- GAL4 or Repo-GAL4 drivers (glial fog RNAi), we observed distinct changes in the positioning and shape of the longitudinal glia. In wild-type embryos, longitudinal glia expressing both Htl and Repo are organized in two rows on either side of the CNS midline (Fig. 2A) (Halter et al., 1995; Shishido et al., 1997). In glial fog RNAi embryos, gaps were observed in these rows (Fig.
7 164 A. Ratnaparkhi, K. Zinn / Developmental Biology 308 (2007) B, arrow). The glial cells appeared smaller and rounder than normal, and their outlines were irregular, often bearing tiny finger-like projections. The distance between the inner glial rows was also greater than in wild-type. To quantitate these phenotypes, we measured the sizes of the GFP-positive glial profiles and observed that glial surface area was reduced by
8 A. Ratnaparkhi, K. Zinn / Developmental Biology 308 (2007) about 30% relative to controls (from 131 μm 2 (n=51) to 94 μm 2 (n=110)). To see if the change in glial shape also affected glial wrapping of axons, we examined the CNS of glial fog RNAi and fog, hkb:: fog embryos in side view. The glial fog RNAi embryos were costained with anti-gfp (in green) and the axon marker BP102 (in red). A series of confocal images (z stacks) were taken for both wild-type and RNAi embryos. Upon superimposing the images, we found that in glial fog RNAi embryos, the glial processes fail to extend ventrally and do not ensheath the axons (Figs. 2C D). Glial staining is localized to the dorsal edge of the CNS, while in control (Htl-GAL4 UAS-mCD8-GFP) embryos it extends ventrally beyond the axon layer. Glial processes were also visualized with anti-htl in both glial fog RNAi and fog, hkb::fog embryos. The normal Htl expression pattern is similar to that of Htl-GAL4-driven mcd8gfp (Fig. 2G). When fog expression is reduced, Htlpositive glial processes were not observed ventral to the CNS axon layer, and a disorganized pattern of axonal ensheathment was observed (Figs. 2H, I). Overall Htl staining intensity was also reduced. This is interesting since the fog phenotype we observed resembles the previously described Htl LOF phenotype (Shishido et al., 1997). To carry out overexpression experiments, we generated UAS-fog transgenes bearing the entire fog cdna cloned into the puast vector. We confirmed that Fog protein was overexpressed in these embryos. Several independent transgenic lines were examined, and all had the same phenotypes. While glial fog RNAi animals survive to adulthood, animals in which Fog is overexpressed using glial drivers die as 1st or 2nd instar larvae. Examination of glial morphology in Repo-GAL4:: UAS-mCD8GFP, UAS-fog embryos showed that they had severe glial defects. The longitudinal glia had a disorganized and clumped appearance, and extension of glial processes appeared abnormal (Figs. 2E F). Loss of neuronal Fog causes axon guidance defects Our examination of Fog protein expression and its inhibition by RNAi indicates that Fog is expressed by at least some neurons (Fig. 1, Supplementary Fig. 1). To evaluate whether neuronal Fog regulates axon guidance, we examined axons in two genetic contexts. First, we reduced Fog expression in neurons by driving fog dsrna with the Elav-GAL4 driver, which is expressed in all postmitotic neurons (neuronal fog RNAi). Several independent UAS-fog dsrna transgenic lines were examined, and all produced the same phenotypes. Second, we examined fog, hkb::fog embryos. In the abdominal segments of wild-type embryos, motor axons exit the CNS via the SN (segmental nerve) and ISN (intersegmental nerve) roots. These then split into 5 different nerves, which innervate 30 body wall muscle fibers (Keshishian et al., 1996). These can all be visualized by staining with the 1D4 mab (Van Vactor et al., 1993). SNa and SNc emerge from the SN root, while the ISN, ISNb and ISNd arise from the ISN root. The SNa nerve shows a characteristic pattern of bifurcation. The bifurcation occurs at the dorsal edge of muscle 12, at a point between muscles 22 and 23 (Figs. 3A B). The anterior (or dorsal) SNa branch innervates muscles 21 24, while the posterior (or lateral) branch innervates muscles 5 and 8. Both fog 4a6,hkb::fog and neuronal fog RNAi embryos had SNa motor axon guidance errors (scored in segments A2 A7). In most segments that displayed phenotypes, either the posterior or the anterior SNa branch was missing (Figs. 3C and D, asterisks). Occasionally, extra anterior branches were observed (Fig. 3C, arrow). In rare cases, the entire SNa appeared to stall at the bifurcation point. The penetrance of these defects was 13.2% and 12.5% in fog 4a6,hkb::fog and neuronal fog RNAi embryos, respectively. We did not observe any motor axon guidance phenotypes in glial fog RNAi embryos. Interestingly, the motor axon guidance defects in fog embryos were identical to those seen in mutants lacking expression of PTP52F (Fig. 3E) (Schindelholz et al., 2001). In Ptp52F embryos, the most common phenotype is also loss of the anterior or posterior SNa branch, and ectopic branches and stalling are observed at lower frequencies (Fig. 3E). The penetrance of these phenotypes is higher ( 40% in null mutants) than in fog embryos. However, we do not know if our fog phenotypes represent the null condition. RNAi usually does not completely eliminate protein expression. Furthermore, fog 4a6,hkb::fog may not be fog-null within the CNS since hkb is expressed in a subset of CNS neuroblasts in each segment, and secreted Fog expressed by these cells could diffuse within the CNS (Chu-LaGraff et al., 1995; Bossing et al., 1996). We also examined the three pairs of 1D4-positive longitudinal axon bundles in the CNS (Van Vactor et al., 1993)(Fig. 4A). In fog 4a6,hkb::fog embryos, the outer 1D4-positive bundle was discontinuous, and occasional breaks and fusions were observed in the inner two bundles (Fig. 4B, arrow). Similar, but milder, phenotypes were observed in neuronal fog RNAi embryos (Fig. 4C). Glial fog RNAi embryos did not have CNS axonal phenotypes. Interestingly, the fog 4a6,hkb::fog CNS axonal phenotype also resembles that seen in Ptp52F mutants (Schindelholz et al., 2001). However, in Ptp52F mutants there is more disorga- Fig. 5. Fog and PTP52F CNS overexpression phenotypes, and suppression of the Fog overexpression phenotype by loss of Ptp52F function. Late stage 16 embryos were stained with mab 1D4 (brown) and dissected. Four segments are shown; anterior is up. (A) Wild-type (Oregon R) embryo. Note that the three 1D4-positive longitudinal axon bundles do not cross the midline. (B) Elav-GAL4::UAS-fog. All three longitudinal bundles are disorganized and often fused to each other. Breaks in the bundles are often present, especially in the outer bundle (arrowhead). Frequent ectopic midline crossing by the inner bundle is observed (arrow). (C) Elav-GAL4:: EP52F. A phenotype similar to panel B is observed, with occasional bundle fusions and breaks (arrowhead), and ectopic midline crossing (arrow). (D) Elav-GAL4:: UAS-fog, Ptp52F 18.3 /Ptp52F Removal of Ptp52F function in Fog overexpression embryos suppresses ectopic midline crossing. (E) Repo-GAL4::UAS-fog. Ectopic midline crossing is also seen when Fog is overexpressed in glia (arrow). Breaks are frequently observed in the outer longitudinal bundles (arrowhead). (F) Bar graph showing percentages of segments with ectopic midline crossing observed in different genotypes. The numbers above the bars indicate the number of segments scored for each genotype. Scale bar in panel A, 10 μm (applies to all panels).
9 166 A. Ratnaparkhi, K. Zinn / Developmental Biology 308 (2007) nization of the outer 1D4 bundle, and it is often fused with the middle bundle (Fig. 4D, arrowhead). PTP52F is epistatic to Fog Overexpression of Fog in neurons using Elav-GAL4 produced strong CNS defects. The three pairs of 1D4-positive longitudinal axon bundles in the CNS displayed multiple breaks and fusions and failed to extend along straight pathways (Fig. 5B, arrowhead). Ectopic midline crossing by 1D4-positive longitudinal axons was observed in 27% of segments (Fig. 5B, arrow). In embryos carrying two copies of the GAL4 driver, the percentage of segments with ectopic midline crossing increased to 44% (Fig. 5F). SNa defects were also observed (Table 1). Glial Fog overexpression also caused axon guidance defects, producing ectopic midline crossing phenotypes like those seen for neuronal Fog overexpression embryos and with a similar penetrance (23%, Figs. 5E F). These data suggest that excess secreted Fog within the CNS can affect axons in a similar manner regardless of whether it is expressed by neurons or by glia. To see if PTP52F overexpression causes effects like those seen for Fog overexpression, we used Elav-GAL4 to drive expression from a UAS-bearing P element (EP element; Rorth et al., 1998) inserted immediately upstream of the Ptp52F gene (a gift of Florenci Serras). This produced CNS axon guidance defects that resembled those caused by excess Fog (Fig. 5C). The 1D4-positive axon bundles had breaks and defasiculated regions, and ectopic midline crossing by longitudinal axons was also observed in 18% (n = 161) of segments (Fig. 5F). These results show that fog and Ptp52F have similar LOF phenotypes in the neuromuscular system and CNS. They also have similar CNS overexpression phenotypes. Because of these phenotypic resemblances, we conducted an epistasis analysis to see if Fog and PTP52F might function within the same signaling pathway. We found that removal of Ptp52F function from Fog overexpression embryos resulted in a strong suppression of the CNS phenotype (Figs. 5D, F). Only 3% of segments had ectopic midline crossing, as compared to 44% for Fog overexpression embryos with a wild-type Ptp52F background (embryos with two copies of neuronal GAL4 driver). For embryos with one copy of the driver, the phenotype was suppressed from 27% to 5%. The longitudinal tracts also looked more normal in Fog overexpression, Ptp52F embryos, resembling those seen in Ptp52F single mutants. These results indicate that the generation of ectopic midline crossing phenotypes by excess neuronal Fog Table 1 SNa motor axon phenotypes in fog mutants Genotype Number of hemisegments Total % SNa defects elav-gal4 UAS-fog dsrna (RT) fog 4a6,hkb::fog (RT) elav-gal4 UAS-fog (25 ) elav c155 -GAL4; elav-gal4 UAS-fog (25 ) cta RC10 /cta RC10 (RT) Abdominal segments A2 A7 were scored for SNa guidance defect. signaling requires PTP52F and suggest that this RPTP functions downstream of Fog in CNS neurons. Discussion In this paper, we show that the secreted cell signal Fog, which is necessary for VF formation in early embryos, also functions during development of the nervous system. Fog is expressed by both glia and neurons within the CNS (Fig. 1, Supplementary Fig. 1), and phenotypic analysis indicates that it has distinct functions in the two cell types. The level of glial Fog expression is a critical element in regulating migration of glial cell bodies, extension of processes and ensheathment of axons (Fig. 2). Autocrine Fog reception by VF cells regulates cytoskeletal morphology and thereby cell shape (Dawes- Hoang et al., 2005), and Fog might function in a similar manner to control glial morphogenesis. Reduction of Fog in neurons produces subtle axon guidance phenotypes affecting both motor neurons and CNS interneurons (Figs. 3, 4). Overexpression of Fog in neurons produces strong CNS phenotypes in which longitudinal axons abnormally cross the midline (Fig. 5). The same phenotypes can be produced by overexpressing Fog in CNS longitudinal glia, which are in apposition to the axons. This result suggests that glial Fog causes cytoskeletal changes that alter axon guidance in neurons, implicating Fog as an exocrine as well as an autocrine signal during nervous system development. Studies of Fog signaling during gastrulation have indicated that the cytoskeletal changes produced by autocrine Fog involve maternal Cta and RhoGEF2 and nonmuscle myosin II (Dawes- Hoang et al., 2005). We tested whether these components participate in Fog signaling in the nervous system by examining the zygotic phenotypes of cta and RhoGEF2 mutants (germline clones do not develop to this stage). Cta may also be involved in Fog signaling during nervous system development because we found that cta zygotic mutant embryos display the same defects in the CNS and neuromuscular system as do fog embryos (Table 1). RhoGEF2 mutants, however, have no visible nervous system defects (data not shown). Myosin II (zipper) mutant embryos have a variety of generalized defects that preclude analysis of specific axon guidance phenotypes. Fog is an autocrine signal required for glial morphogenesis Most of the cells in the CNS of late embryos that express fog mrna at high levels are Repo-positive longitudinal glia (Fig. 1). These glia are required for normal morphogenesis of the CNS axon tracts (Hidalgo et al., 1995; Hidalgo and Booth, 2000), but we did not observe CNS axon phenotypes when Fog expression was reduced in glia. To evaluate Fog's functions in the glia, we examined glial morphology directly using a membrane-associated GFP marker. When Fog expression is reduced in glia, glial processes fail to extend normally and ensheath CNS axons (Fig. 2). There are gaps in the regular array of glia, glial surface areas are smaller than in wild-type, and the glia have a rounded appearance. These changes in cell shape could involve nonmuscle myosin.
10 A. Ratnaparkhi, K. Zinn / Developmental Biology 308 (2007) Overexpression of Fog in glia confers lethality during early larval phases. Glial morphogenesis is affected by overexpression, but the phenotypes are different from those seen when Fog is reduced. Glia appear to have normal shapes, but the glial lattice is quite disorganized. In wild-type embryos, lines of glia define the positions of the longitudinal tracts, commissural tracts and peripheral nerves; these regular arrays are not observed in Fog glial overexpression embryos (Fig. 2). Thus, both reduction and elevation of glial Fog produce a disorganized glial lattice, suggesting that a precise level of the Fog signal is necessary for normal glial development. Potential relationships between Fog and PTP52F The Fog receptor has not been identified, although it is speculated to be a GPCR because of the requirement of the G protein alpha subunit Cta for Fog signaling (Morize et al., 1998). However, existing genetic data do not show that Fog directly activates a GPCR; they are also consistent with models in which Fog regulates signaling through a GPCR-Cta pathway in an indirect manner by interacting with a non-gpcr receptor. PTP52F, like most RPTPs, is an orphan receptor. The motivation to conduct the experiments described in this paper arose from our observations that fog and Ptp52F embryos display similar VF phenotypes (Supplementary Fig. 2) and that PTP52F is expressed in ventral furrow cells during the gastrulation phase (Schindelholz et al., 2001). Based on these results, we wondered whether PTP52F could be the elusive Fog receptor. PTP52F is required for axon guidance in the embryonic CNS and neuromuscular system. Thus, to examine whether Fog and PTP52F might be part of the same signaling pathway, we examined fog axon guidance phenotypes and studied genetic interactions between the two molecules. Our data show that fog and Ptp52F have similar LOF and GOF phenotypes in the CNS (Figs. 4, 5). In the neuromuscular system, LOF mutations in both genes cause SNa bifurcation phenotypes (Fig. 3). Our definition of a fog GOF CNS phenotype allowed us to perform an epistasis experiment, and this showed that PTP52F is required for signaling downstream of Fog, at least in the context of this phenotype (Fig. 5). The genetic results we obtained indicate that PTP52F is involved in reception of the Fog signal by neurons, but do not prove that PTP52F is a Fog receptor. The results could also be explained if PTP52F positively regulates signaling through a Fog-GPCR-Cta pathway. For example, GPCRs are phosphorylated (on serine or threonine residues) and internalized; the activities of the relevant kinases and/or the proteins involved in internalization could be modulated by tyrosine phosphorylation. Tyrosine phosphorylation could also regulate effectors downstream of the G protein Cta. We have been unable to demonstrate a direct biochemical interaction between the PTP52F extracellular domain and several versions of the Fog protein. However, Fog might be processed in vivo to create a functional ligand, and this processing does not occur in heterologous systems. In our experiments, we found that Fog tagged at its N terminus is secreted from insect cells when expressed using the baculovirus system, but the protein is degraded to produce a ladder of bands ranging in size from 100 kda (the predicted size of glycosylated full-length Fog) to b20 kda. Fog tagged at its C terminus cannot be detected at all. Fog fused near its C terminus to human placental alkaline phosphatase (Fog-AP) can be expressed as a mixture of apparently full-length and degraded forms, but none of these proteins bound detectably to the tagged PTP52F extracellular domain (A.R., P.M. Snow, and K.Z., unpublished results). Taken together, our data suggest that the C terminal region of Fog is subject to degradation and that full-length Fog is unstable. There are several dibasic sequences in Fog which could represent proteolytic cleavage sites, and Wieschaus and colleagues (Costa et al., 1994) have previously proposed that Fog could be processed in vivo to generate an active fragment that binds to the receptor. In the CNS, such a fragment might derive from the middle region of Fog because we observed that antisera against full-length Fog stain late embryos (Fig. 1, Supplementary Fig. 1), while antisera against the first 300 amino acids of Fog (Dawes-Hoang et al., 2005) do not. Acknowledgments We are very grateful to the Wieschaus laboratory and Naoyuki Fuse for their generous gift of the Fog antibody and to Maria Leptin for sharing the fog 4a6,hkb::fog flies prior to publication. We also thank Florenci Serras for EP52F; the Caltech Biological Imaging Centre and the Gonda Imaging Centre, UCLA for use of their confocal microscopes. Aloisia Schmid and Benno Schindelholz (former Zinn group members) discovered the Ptp52F VF phenotype. We thank them and the other present and former members of the Zinn group for helpful discussions. A.R. particularly thanks Dr. George Jackson, UCLA, in whose laboratory some part of this work was done. This work was supported by an NIH RO1 grant, NS28182, to K. Zinn and by the Gosney fellowship from Caltech to A.R. Appendix A. Supplementary data Supplementary data associated with this article can be found, in the online version, at doi: /j.ydbio References Bossing, T., Technau, G.M., Doe, C.Q., huckebein is required for glial development and axon pathfinding in the neuroblast 1-1 and neuroblast 2-2 lineages in the Drosophila central nervous system. Mech. Dev. 55, Chu-LaGraff, Q., Schmid, A., Leidel, J., Bronner, G., Jackle, H., Doe, C.Q., huckebein specifies aspects of CNS precursor identity required for motoneuron axon pathfinding. Neuron 15, Costa, M., Sweeton, D., Wieschaus, E., Gastrulation in Drosophila: cellular mechanisms of morphogenetic movements. In: Bate, M., Martinez- Arias, A. (Eds.), The Development of Drosophila. Cold Spring Harbor Laboratories Press, New York, pp Costa, M., Wilson, E.T., Wieschaus, E., A putative cell signal encoded by the folded gastrulation gene coordinates cell shape changes during Drosophila gastrulation. Cell 76, Dawes-Hoang, R.E., Parmar, K.M., Christiansen, A.E., Phelps, C.B., Brand,
11 168 A. Ratnaparkhi, K. Zinn / Developmental Biology 308 (2007) A.H., Weichaus, E.F., Folded gastrulation, cell shape change and the control of myosin localization. Development 132, Halter, D.A., Urban, J., Rickert, C., Ner, S.S., Ito, K., Travers, A.A., Technau, G.M., The homeobox gene repo is required for the differentiation and maintenance of glia function in the embryonic nervous system of Drosophila melanogaster. Development 121, Hidalgo, A., Booth, G.E., Glia dictate pioneer axon trajectories in the Drosophila embryonic CNS. Development 127, Hidalgo, A., Urban, J., Brand, A.H., Targeted ablation of glia disrupts axon tract formation in the Drosophila CNS. Development 121, Jones, B.W., Fetter, R.D., Tear, G., Goodman, C.S., Glial cells missing: a genetic switch that controls glial versus neuronal fate. Cell 82, Keshishian, H., Broadie, K., Chiba, A., Bate, M., The Drosophila neuromuscular junction: a model system for studying synaptic development and function. Annu. Rev. Neurosci. 19, Kraut, R., Menon, K., Zinn, K., A gain-of-function screen for genes controlling motor axon guidance and synaptogenesis in Drosophila. Curr. Biol. 11, Lammel, U., Saumweber, H., X-linked loci of Drosophila melanogaster causing defects in the morphology of the embryonic salivary glands. Dev. Genes Evol. 210, Morize, P., Christiansen, A.E., Costa, M., Parks, S., Wieschaus, E., Hyperactivation of the folded gastrulation pathway induces specific cell shape changes. Development 125, Parks, S., Wieschaus, E., The Drosophila gastrulation gene concertina encodes a G alpha-like protein. Cell 64, Patel, N.H., Imaging neuronal subsets and other cell types in whole-mount Drosophila embryos and larvae using antibody probes. In: Fryberg, E., Goldstein, L.S.B (Eds.), Drosophila melanogaster: Practical Uses in Cell and Molecular Biology, vol. 44. Academic Press, San Diego, CA, pp Rorth, P., Szabo, K., Bailey, A., Laverty, T., Rehm, J., Rubin, G.M., Weigmann, K., Milan, M., Benes, V., Ansorge, W., Cohen, S.M., Systematic gainof-function genetics in Drosophila. Development 125, Schindelholz, B., Knirr, M., Warrior, R., Zinn, K., Regulation of CNS and motor axon guidance in Drosophila by the receptor tyrosine phosphatase DPTP52F. Development 128, Shishido, E., Ono, N., Kojima, T., Saigo, K., Requirements of DFR1/ Heartless, a mesoderm-specific Drosophila FGF-receptor, for the formation of heart, visceral and somatic muscles, and ensheathing of longitudinal axon tracts in CNS. Development 124, Sweeton, D., Parks, S., Costa, M., Wieschaus, E., Gastrulation in Drosophila: the formation of the ventral furrow and posterior midgut invaginations. Development 112, Van Vactor, D., Sink, H., Fambrough, D., Tsoo, R., Goodman, C.S., Genes that control neuromuscular specificity in Drosophila. Cell 73, Xiong, W.C., Okano, H., Patel, N.H., Blendy, J.A., Montell, C., repo encodes a glial-specific homeo domain protein required in the Drosophila nervous system. Genes Dev. 8,
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
MCN. Complex Genetic Interactions among Four Receptor Tyrosine Phosphatases Regulate Axon Guidance in Drosophila
 MCN Molecular and Cellular Neuroscience 17, 274 291 (2001) doi:10.1006/mcne.2000.0939, available online at http://www.idealibrary.com on Complex Genetic Interactions among Four Receptor Tyrosine Phosphatases
MCN Molecular and Cellular Neuroscience 17, 274 291 (2001) doi:10.1006/mcne.2000.0939, available online at http://www.idealibrary.com on Complex Genetic Interactions among Four Receptor Tyrosine Phosphatases
Systematic Screening of Drosophila Deficiency Mutations for Embryonic Phenotypes and Orphan Receptor Ligands
 Systematic Screening of Drosophila Deficiency Mutations for Embryonic Phenotypes and Orphan Receptor Ligands Ashley P. Wright 1, A. Nicole Fox 1, Karl G. Johnson 2, Kai Zinn 1 * 1 Division of Biology,
Systematic Screening of Drosophila Deficiency Mutations for Embryonic Phenotypes and Orphan Receptor Ligands Ashley P. Wright 1, A. Nicole Fox 1, Karl G. Johnson 2, Kai Zinn 1 * 1 Division of Biology,
Baz, Par-6 and apkc are not required for axon or dendrite specification in Drosophila
 Baz, Par-6 and apkc are not required for axon or dendrite specification in Drosophila Melissa M. Rolls and Chris Q. Doe, Inst. Neurosci and Inst. Mol. Biol., HHMI, Univ. Oregon, Eugene, Oregon 97403 Correspondence
Baz, Par-6 and apkc are not required for axon or dendrite specification in Drosophila Melissa M. Rolls and Chris Q. Doe, Inst. Neurosci and Inst. Mol. Biol., HHMI, Univ. Oregon, Eugene, Oregon 97403 Correspondence
Fig. S1. Expression pattern of moody-gal4 in third instar. Maximum projection illustrating a dissected moody-gal4>ngfp L3 larva stained for Repo
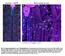 Fig. S1. Expression pattern of moody-gal4 in third instar. Maximum projection illustrating a dissected moody-gal4>ngfp L3 larva stained for Repo (magenta), Fas2 (blue) and GFP (green) in overview (A) and
Fig. S1. Expression pattern of moody-gal4 in third instar. Maximum projection illustrating a dissected moody-gal4>ngfp L3 larva stained for Repo (magenta), Fas2 (blue) and GFP (green) in overview (A) and
Attractive and repulsive functions of Slit are mediated by different receptors in the Drosophila trachea
 Development 129, 4941-4951 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV7966 4941 Attractive and repulsive functions of Slit are mediated by different receptors in the Drosophila
Development 129, 4941-4951 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV7966 4941 Attractive and repulsive functions of Slit are mediated by different receptors in the Drosophila
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Overexpression of YFP::GPR-1 in the germline.
 Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
Chapter 4 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays.
 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Peripheral Glia Direct Axon Guidance across the CNS/PNS Transition Zone
 Developmental Biology 238, 47 63 (2001) doi:10.1006/dbio.2001.0411, available online at http://www.idealibrary.com on Peripheral Glia Direct Axon Guidance across the CNS/PNS Transition Zone Katharine J.
Developmental Biology 238, 47 63 (2001) doi:10.1006/dbio.2001.0411, available online at http://www.idealibrary.com on Peripheral Glia Direct Axon Guidance across the CNS/PNS Transition Zone Katharine J.
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10923 Supplementary Figure 1 Ten-a and Ten-m antibody and cell type specificities. a c, Representative single confocal sections of a Drosophila NMJ stained with antibodies to Ten-a (red),
doi:10.1038/nature10923 Supplementary Figure 1 Ten-a and Ten-m antibody and cell type specificities. a c, Representative single confocal sections of a Drosophila NMJ stained with antibodies to Ten-a (red),
Midterm 1. Average score: 74.4 Median score: 77
 Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3267 Supplementary Figure 1 A group of genes required for formation or orientation of annular F-actin bundles and aecm ridges: RNAi phenotypes and their validation by standard mutations.
DOI: 10.1038/ncb3267 Supplementary Figure 1 A group of genes required for formation or orientation of annular F-actin bundles and aecm ridges: RNAi phenotypes and their validation by standard mutations.
Segment boundary formation in Drosophila embryos
 Segment boundary formation in Drosophila embryos Development 130, August 2003 Camilla W. Larsen, Elizabeth Hirst, Cyrille Alexandre and Jean Paul Vincent 1. Introduction: - Segment boundary formation:
Segment boundary formation in Drosophila embryos Development 130, August 2003 Camilla W. Larsen, Elizabeth Hirst, Cyrille Alexandre and Jean Paul Vincent 1. Introduction: - Segment boundary formation:
Nature Neuroscience: doi: /nn.2662
 Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Chapter 11. Development: Differentiation and Determination
 KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
Unicellular: Cells change function in response to a temporal plan, such as the cell cycle.
 Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Conclusions. The experimental studies presented in this thesis provide the first molecular insights
 C h a p t e r 5 Conclusions 5.1 Summary The experimental studies presented in this thesis provide the first molecular insights into the cellular processes of assembly, and aggregation of neural crest and
C h a p t e r 5 Conclusions 5.1 Summary The experimental studies presented in this thesis provide the first molecular insights into the cellular processes of assembly, and aggregation of neural crest and
The Transmembrane Semaphorin Sema I Is Required in Drosophila for Embryonic Motor and CNS Axon Guidance
 Neuron, Vol. 20, 207 220, February, 1998, Copyright 1998 by Cell Press The Transmembrane Semaphorin Sema I Is Required in Drosophila for Embryonic Motor and CNS Axon Guidance Hung-Hsiang Yu, Houmam H.
Neuron, Vol. 20, 207 220, February, 1998, Copyright 1998 by Cell Press The Transmembrane Semaphorin Sema I Is Required in Drosophila for Embryonic Motor and CNS Axon Guidance Hung-Hsiang Yu, Houmam H.
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila July 9, 2008 Drosophila Development Overview Fertilization Cleavage Gastrulation Drosophila body plan Oocyte formation Genetic control
Developmental Biology Biology 4361 Axis Specification in Drosophila July 9, 2008 Drosophila Development Overview Fertilization Cleavage Gastrulation Drosophila body plan Oocyte formation Genetic control
Crk-associated substrate (Cas) signaling protein functions with integrins to specify axon guidance during development
 RESEARCH ARTICLE 2337 Development 134, 2337-2347 (2007) doi:10.1242/dev.004242 Crk-associated substrate (Cas) signaling protein functions with integrins to specify axon guidance during development Zhiyu
RESEARCH ARTICLE 2337 Development 134, 2337-2347 (2007) doi:10.1242/dev.004242 Crk-associated substrate (Cas) signaling protein functions with integrins to specify axon guidance during development Zhiyu
Drosophila Life Cycle
 Drosophila Life Cycle 1 Early Drosophila Cleavage Nuclei migrate to periphery after 10 nuclear divisions. Cellularization occurs when plasma membrane folds in to divide nuclei into cells. Drosophila Superficial
Drosophila Life Cycle 1 Early Drosophila Cleavage Nuclei migrate to periphery after 10 nuclear divisions. Cellularization occurs when plasma membrane folds in to divide nuclei into cells. Drosophila Superficial
10/2/2015. Chapter 4. Determination and Differentiation. Neuroanatomical Diversity
 Chapter 4 Determination and Differentiation Neuroanatomical Diversity 1 Neurochemical diversity: another important aspect of neuronal fate Neurotransmitters and their receptors Excitatory Glutamate Acetylcholine
Chapter 4 Determination and Differentiation Neuroanatomical Diversity 1 Neurochemical diversity: another important aspect of neuronal fate Neurotransmitters and their receptors Excitatory Glutamate Acetylcholine
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature12791 Supplementary Figure 1 (1/3) WWW.NATURE.COM/NATURE 1 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 1 (2/3) 2 WWW.NATURE.COM/NATURE SUPPLEMENTARY
SUPPLEMENTARY INFORMATION doi:10.1038/nature12791 Supplementary Figure 1 (1/3) WWW.NATURE.COM/NATURE 1 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 1 (2/3) 2 WWW.NATURE.COM/NATURE SUPPLEMENTARY
Chapter 18 Lecture. Concepts of Genetics. Tenth Edition. Developmental Genetics
 Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Positional Cues in the Drosophila Nerve Cord: Semaphorins Pattern the Dorso-Ventral Axis
 Positional Cues in the Drosophila Nerve Cord: Semaphorins Pattern the Dorso-Ventral Axis Marta Zlatic 1,2,3 *, Feng Li 1, Maura Strigini 4, Wesley Grueber 2, Michael Bate 1 * 1 Department of Zoology, University
Positional Cues in the Drosophila Nerve Cord: Semaphorins Pattern the Dorso-Ventral Axis Marta Zlatic 1,2,3 *, Feng Li 1, Maura Strigini 4, Wesley Grueber 2, Michael Bate 1 * 1 Department of Zoology, University
MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning
 MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning Learning Goals for Lecture 29 4.1 Describe the contributions of early developmental events in the embryo to the formation
MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning Learning Goals for Lecture 29 4.1 Describe the contributions of early developmental events in the embryo to the formation
Axis determination in flies. Sem 9.3.B.5 Animal Science
 Axis determination in flies Sem 9.3.B.5 Animal Science All embryos are in lateral view (anterior to the left). Endoderm, midgut; mesoderm; central nervous system; foregut, hindgut and pole cells in yellow.
Axis determination in flies Sem 9.3.B.5 Animal Science All embryos are in lateral view (anterior to the left). Endoderm, midgut; mesoderm; central nervous system; foregut, hindgut and pole cells in yellow.
Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and
 Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and Lifeact-Ruby (red) were imaged in vivo to visualize secretion
Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and Lifeact-Ruby (red) were imaged in vivo to visualize secretion
Drosophila Plexin B is a Sema-2a receptor required for axon guidance
 RESEARCH ARTICLE 2125 Development 133, 2125-2135 (2006) doi:10.1242/dev.02380 Drosophila Plexin B is a Sema-2a receptor required for axon guidance Joseph C. Ayoob 1, Jonathan R. Terman 1,2 and Alex L.
RESEARCH ARTICLE 2125 Development 133, 2125-2135 (2006) doi:10.1242/dev.02380 Drosophila Plexin B is a Sema-2a receptor required for axon guidance Joseph C. Ayoob 1, Jonathan R. Terman 1,2 and Alex L.
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8
 Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
The Control of Semaphorin-1a-Mediated Reverse Signaling by Opposing Pebble and RhoGAPp190 Functions in Drosophila
 Article The Control of Semaphorin-1a-Mediated Reverse Signaling by Opposing Pebble and RhoGAPp190 Functions in Drosophila Sangyun Jeong, 1 Katarina Juhaszova, 1 and Alex L. Kolodkin 1, 1 Solomon H. Snyder
Article The Control of Semaphorin-1a-Mediated Reverse Signaling by Opposing Pebble and RhoGAPp190 Functions in Drosophila Sangyun Jeong, 1 Katarina Juhaszova, 1 and Alex L. Kolodkin 1, 1 Solomon H. Snyder
Developmental Dynamics of Peripheral Glia in Drosophila melanogaster
 GLIA 30:122 133 (2000) Developmental Dynamics of Peripheral Glia in Drosophila melanogaster KATHARINE J. SEPP, JOOST SCHULTE, AND VANESSA J. AULD* Department of Zoology, University of British Columbia,
GLIA 30:122 133 (2000) Developmental Dynamics of Peripheral Glia in Drosophila melanogaster KATHARINE J. SEPP, JOOST SCHULTE, AND VANESSA J. AULD* Department of Zoology, University of British Columbia,
Why Flies? stages of embryogenesis. The Fly in History
 The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
Supporting Information
 Supporting Information Cao et al. 10.1073/pnas.1306220110 Gram - bacteria Hemolymph Cytoplasm PGRP-LC TAK1 signalosome Imd dfadd Dredd Dnr1 Ikk signalosome P Relish Nucleus AMP and effector genes Fig.
Supporting Information Cao et al. 10.1073/pnas.1306220110 Gram - bacteria Hemolymph Cytoplasm PGRP-LC TAK1 signalosome Imd dfadd Dredd Dnr1 Ikk signalosome P Relish Nucleus AMP and effector genes Fig.
The Midline Protein Regulates Axon Guidance by Blocking the Reiteration of Neuroblast Rows within the Drosophila Ventral Nerve Cord
 The Midline Protein Regulates Axon Guidance by Blocking the Reiteration of Neuroblast Rows within the Drosophila Ventral Nerve Cord Mary Ann Manavalan, Ivana Gaziova, Krishna Moorthi Bhat* Department of
The Midline Protein Regulates Axon Guidance by Blocking the Reiteration of Neuroblast Rows within the Drosophila Ventral Nerve Cord Mary Ann Manavalan, Ivana Gaziova, Krishna Moorthi Bhat* Department of
Regulation of CNS and motor axon guidance in Drosophila by the receptor tyrosine phosphatase DPTP52F
 Development 128, 4371-4382 (2001) Printed in Great Britain The Company of Biologists Limited 2001 DEV8821 4371 Regulation of CNS and motor axon guidance in Drosophila by the receptor tyrosine phosphatase
Development 128, 4371-4382 (2001) Printed in Great Britain The Company of Biologists Limited 2001 DEV8821 4371 Regulation of CNS and motor axon guidance in Drosophila by the receptor tyrosine phosphatase
Interactions between a Receptor Tyrosine Phosphatase and a Cell Surface Ligand Regulate Axon Guidance and Glial-Neuronal Communication
 Article Interactions between a Receptor Tyrosine Phosphatase and a Cell Surface Ligand Regulate Axon Guidance and Glial-Neuronal Communication Hyung-Kook (Peter) Lee, 1 Amy Cording, 1,2 Jost Vielmetter,
Article Interactions between a Receptor Tyrosine Phosphatase and a Cell Surface Ligand Regulate Axon Guidance and Glial-Neuronal Communication Hyung-Kook (Peter) Lee, 1 Amy Cording, 1,2 Jost Vielmetter,
Slit Is the Midline Repellent for the Robo Receptor in Drosophila
 Cell, Vol. 96, 785 794, March 19, 1999, Copyright 1999 by Cell Press Slit Is the Midline Repellent for the Robo Receptor in Drosophila Thomas Kidd, Kimberly S. Bland, and Corey S. Goodman* Howard Hughes
Cell, Vol. 96, 785 794, March 19, 1999, Copyright 1999 by Cell Press Slit Is the Midline Repellent for the Robo Receptor in Drosophila Thomas Kidd, Kimberly S. Bland, and Corey S. Goodman* Howard Hughes
Developmental genetics: finding the genes that regulate development
 Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Development of Drosophila
 Development of Drosophila Hand-out CBT Chapter 2 Wolpert, 5 th edition March 2018 Introduction 6. Introduction Drosophila melanogaster, the fruit fly, is found in all warm countries. In cooler regions,
Development of Drosophila Hand-out CBT Chapter 2 Wolpert, 5 th edition March 2018 Introduction 6. Introduction Drosophila melanogaster, the fruit fly, is found in all warm countries. In cooler regions,
Axon guidance I. Paul Garrity March 15, /9.013
 Axon guidance I Paul Garrity March 15, 2004 7.68/9.013 Neuronal Wiring: Functional Framework of the Nervous System Stretch reflex circuit Early theories of axonogenesis Schwann: many neurons link to form
Axon guidance I Paul Garrity March 15, 2004 7.68/9.013 Neuronal Wiring: Functional Framework of the Nervous System Stretch reflex circuit Early theories of axonogenesis Schwann: many neurons link to form
Question Set # 4 Answer Key 7.22 Nov. 2002
 Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.1244624/dc1 Supplementary Materials for Cytoneme-Mediated Contact-Dependent Transport of the Drosophila Decapentaplegic Signaling Protein Sougata Roy, Hai Huang,
www.sciencemag.org/cgi/content/full/science.1244624/dc1 Supplementary Materials for Cytoneme-Mediated Contact-Dependent Transport of the Drosophila Decapentaplegic Signaling Protein Sougata Roy, Hai Huang,
Report. Functional Diversity of Robo Receptor Immunoglobulin Domains Promotes Distinct Axon Guidance Decisions
 Current Biology 20, 567 572, March 23, 2010 ª2010 Elsevier Ltd All rights reserved DOI 10.1016/j.cub.2010.02.021 Functional Diversity of Robo Receptor Immunoglobulin Domains Promotes Distinct Axon Guidance
Current Biology 20, 567 572, March 23, 2010 ª2010 Elsevier Ltd All rights reserved DOI 10.1016/j.cub.2010.02.021 Functional Diversity of Robo Receptor Immunoglobulin Domains Promotes Distinct Axon Guidance
IN the Drosophila ventral nerve cord, there are 20
 Copyright Ó 2007 by the Genetics Society of America DOI: 10.1534/genetics.107.075085 Regulation of Axon Guidance by Slit and Netrin Signaling in the Drosophila Ventral Nerve Cord Krishna Moorthi Bhat,
Copyright Ó 2007 by the Genetics Society of America DOI: 10.1534/genetics.107.075085 Regulation of Axon Guidance by Slit and Netrin Signaling in the Drosophila Ventral Nerve Cord Krishna Moorthi Bhat,
Neural development its all connected
 Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
Cell biology: Death drags down the neighbourhood
 Cell biology: Death drags down the neighbourhood The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher Vasquez,
Cell biology: Death drags down the neighbourhood The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation As Published Publisher Vasquez,
split ends encodes large nuclear proteins that regulate neuronal cell fate and
 Development 127, 1517-1529 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV8693 1517 split ends encodes large nuclear proteins that regulate neuronal cell fate and axon extension
Development 127, 1517-1529 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV8693 1517 split ends encodes large nuclear proteins that regulate neuronal cell fate and axon extension
SUPPLEMENTARY INFORMATION
 Supplementary Figure 1 Sns and Duf co-localise in embryonic nephrocytes a-c, Wild-type stage 11 (a),14 (b) and 16 (c) embryos stained with anti-duf (green) and anti-sns (red). Higher magnification images
Supplementary Figure 1 Sns and Duf co-localise in embryonic nephrocytes a-c, Wild-type stage 11 (a),14 (b) and 16 (c) embryos stained with anti-duf (green) and anti-sns (red). Higher magnification images
Nature Neuroscience: doi: /nn.2717
 Supplementary Fig. 1. Dendrite length is not secondary to body length. Dendrite growth proceeds independently of the rate of body growth and decreases in rate in adults. n 20 on dendrite measurement, n
Supplementary Fig. 1. Dendrite length is not secondary to body length. Dendrite growth proceeds independently of the rate of body growth and decreases in rate in adults. n 20 on dendrite measurement, n
Connectin Mediates Adhesion in Drosophila
 Neuron, Vol. 18, 873 880, June, 1997, Copyright 1997 by Cell Press Connectin Mediates Adhesion in Drosophila Srikala Raghavan and Robert A. H. White in little observable perturbation of the normal axonal
Neuron, Vol. 18, 873 880, June, 1997, Copyright 1997 by Cell Press Connectin Mediates Adhesion in Drosophila Srikala Raghavan and Robert A. H. White in little observable perturbation of the normal axonal
A Screen of Cell-Surface Molecules Identifies Leucine-Rich Repeat Proteins as Key Mediators of Synaptic Target Selection
 Article A Screen of Cell-Surface Molecules Identifies Leucine-Rich Repeat Proteins as Key Mediators of Synaptic Target Selection Mitsuhiko Kurusu, 1,2, * Amy Cording, 1 Misako Taniguchi, 2 Kaushiki Menon,
Article A Screen of Cell-Surface Molecules Identifies Leucine-Rich Repeat Proteins as Key Mediators of Synaptic Target Selection Mitsuhiko Kurusu, 1,2, * Amy Cording, 1 Misako Taniguchi, 2 Kaushiki Menon,
Lecture 7. Development of the Fruit Fly Drosophila
 BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
Nirupama Deshpande, Rainer Dittrich, Gerhard M. Technau and Joachim Urban* SUMMARY
 Development 128, 3253-3261 (2001) Printed in Great Britain The Company of Biologists Limited 2001 DEV8801 3253 Successive specification of Drosophila neuroblasts NB 6-4 and NB 7-3 depends on interaction
Development 128, 3253-3261 (2001) Printed in Great Britain The Company of Biologists Limited 2001 DEV8801 3253 Successive specification of Drosophila neuroblasts NB 6-4 and NB 7-3 depends on interaction
Supplementary Figures
 Supplementary Figures Supplementary Fig. S1: Normal development and organization of the embryonic ventral nerve cord in Platynereis. (A) Life cycle of Platynereis dumerilii. (B-F) Axonal scaffolds and
Supplementary Figures Supplementary Fig. S1: Normal development and organization of the embryonic ventral nerve cord in Platynereis. (A) Life cycle of Platynereis dumerilii. (B-F) Axonal scaffolds and
Varicose and cheerio collaborate with pebble to mediate semaphorin-1a reverse signaling in Drosophila
 Varicose and cheerio collaborate with pebble to mediate semaphorin-1a reverse signaling in Drosophila Sangyun Jeong a,1, Da-som Yang a, Young Gi Hong a, Sarah P. Mitchell b,c, Matthew P. rown b,c, and
Varicose and cheerio collaborate with pebble to mediate semaphorin-1a reverse signaling in Drosophila Sangyun Jeong a,1, Da-som Yang a, Young Gi Hong a, Sarah P. Mitchell b,c, Matthew P. rown b,c, and
Drosophila homeodomain protein Nkx6 coordinates motoneuron
 Research article 5233 Drosophila homeodomain protein Nkx6 coordinates motoneuron subtype identity and axonogenesis Heather T. Broihier 1, *,,, Alexander Kuzin 2,, Yi Zhu 1, Ward Odenwald 2 and James B.
Research article 5233 Drosophila homeodomain protein Nkx6 coordinates motoneuron subtype identity and axonogenesis Heather T. Broihier 1, *,,, Alexander Kuzin 2,, Yi Zhu 1, Ward Odenwald 2 and James B.
SUPPLEMENTARY INFORMATION
 Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
Expression of a set of glial cell-specific markers in the Drosophila embryonic central nervous system
 BMB Reports BMB Rep. 2014; 47(6): 354-359 www.bmbreports.org Expression of a set of glial cell-specific markers in the Drosophila embryonic central nervous system Hui Jeong Ahn 1, Sang-Hak Jeon 2 & Sang
BMB Reports BMB Rep. 2014; 47(6): 354-359 www.bmbreports.org Expression of a set of glial cell-specific markers in the Drosophila embryonic central nervous system Hui Jeong Ahn 1, Sang-Hak Jeon 2 & Sang
Cell division genes promote asymmetric interaction between Numb and Notch in the Drosophila CNS
 Development 126, 2759-2770 (1999) Printed in Great Britain The Company of Biologists Limited 1999 DEV5300 2759 Cell division genes promote asymmetric interaction between Numb and Notch in the Drosophila
Development 126, 2759-2770 (1999) Printed in Great Britain The Company of Biologists Limited 1999 DEV5300 2759 Cell division genes promote asymmetric interaction between Numb and Notch in the Drosophila
Drosophila HB9 Is Expressed in a Subset of Motoneurons and Interneurons, Where It Regulates Gene Expression and Axon Pathfinding
 The Journal of Neuroscience, November 1, 2002, 22(21):9143 9149 Brief Communication Drosophila HB9 Is Expressed in a Subset of Motoneurons and Interneurons, Where It Regulates Gene Expression and Axon
The Journal of Neuroscience, November 1, 2002, 22(21):9143 9149 Brief Communication Drosophila HB9 Is Expressed in a Subset of Motoneurons and Interneurons, Where It Regulates Gene Expression and Axon
Caenorhabditis elegans
 Caenorhabditis elegans Why C. elegans? Sea urchins have told us much about embryogenesis. They are suited well for study in the lab; however, they do not tell us much about the genetics involved in embryogenesis.
Caenorhabditis elegans Why C. elegans? Sea urchins have told us much about embryogenesis. They are suited well for study in the lab; however, they do not tell us much about the genetics involved in embryogenesis.
Reading: Chapter 5, pp ; Reference chapter D, pp Problem set F
 Mosaic Analysis Reading: Chapter 5, pp140-141; Reference chapter D, pp820-823 Problem set F Twin spots in Drosophila Although segregation and recombination in mitosis do not occur at the same frequency
Mosaic Analysis Reading: Chapter 5, pp140-141; Reference chapter D, pp820-823 Problem set F Twin spots in Drosophila Although segregation and recombination in mitosis do not occur at the same frequency
9/4/2015 INDUCTION CHAPTER 1. Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology. Fig 1.
 INDUCTION CHAPTER 1 Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology Fig 1.1 1 EVOLUTION OF METAZOAN BRAINS GASTRULATION MAKING THE 3 RD GERM LAYER
INDUCTION CHAPTER 1 Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology Fig 1.1 1 EVOLUTION OF METAZOAN BRAINS GASTRULATION MAKING THE 3 RD GERM LAYER
The Origin, Location, and Projections of the Embryonic Abdominal Motorneurons of Drosophila
 The Journal of Neuroscience, December 15, 1997, 17(24):9642 9655 The Origin, Location, and Projections of the Embryonic Abdominal Motorneurons of Drosophila Matthias Landgraf, 1 Torsten Bossing, 2 Gerd
The Journal of Neuroscience, December 15, 1997, 17(24):9642 9655 The Origin, Location, and Projections of the Embryonic Abdominal Motorneurons of Drosophila Matthias Landgraf, 1 Torsten Bossing, 2 Gerd
Role of Organizer Chages in Late Frog Embryos
 Ectoderm Germ Layer Frog Fate Map Frog Fate Map Role of Organizer Chages in Late Frog Embryos Organizer forms three distinct regions Notochord formation in chick Beta-catenin localization How does beta-catenin
Ectoderm Germ Layer Frog Fate Map Frog Fate Map Role of Organizer Chages in Late Frog Embryos Organizer forms three distinct regions Notochord formation in chick Beta-catenin localization How does beta-catenin
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its
 Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Early Development in Invertebrates
 Developmental Biology Biology 4361 Early Development in Invertebrates October 25, 2006 Early Development Overview Cleavage rapid cell divisions divisions of fertilized egg into many cells Gastrulation
Developmental Biology Biology 4361 Early Development in Invertebrates October 25, 2006 Early Development Overview Cleavage rapid cell divisions divisions of fertilized egg into many cells Gastrulation
Developmental Biology 3230 Midterm Exam 1 March 2006
 Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A)
 Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the
Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the
7.06 Problem Set #4, Spring 2005
 7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018
 Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
Reading. Lecture VI. Making Connections 9/17/12. Bio 3411 Lecture VI. Making Connections. Bio 3411 Monday September 17, 2012
 Lecture VI. Making Connections Bio 3411 Monday September 17, 2012!! 1! Reading NEUROSCIENCE: 5 th ed, pp!507?536! 4 th ed, pp 577-609 Bentley, D., & Caudy, M. (1983). Nature, 304(5921), 62-65. Dickson,
Lecture VI. Making Connections Bio 3411 Monday September 17, 2012!! 1! Reading NEUROSCIENCE: 5 th ed, pp!507?536! 4 th ed, pp 577-609 Bentley, D., & Caudy, M. (1983). Nature, 304(5921), 62-65. Dickson,
DETERMINING THE DIFFERENTIAL ROLES OF THE DOCK FAMILY OF GEFS IN DROSOPHILA DEVELOPMENT
 DETERMINING THE DIFFERENTIAL ROLES OF THE DOCK FAMILY OF GEFS IN DROSOPHILA DEVELOPMENT A DISSERTATION IN Cell Biology & Biophysics and Molecular Biology & Biochemistry Presented to the Faculty of the
DETERMINING THE DIFFERENTIAL ROLES OF THE DOCK FAMILY OF GEFS IN DROSOPHILA DEVELOPMENT A DISSERTATION IN Cell Biology & Biophysics and Molecular Biology & Biochemistry Presented to the Faculty of the
Follow this and additional works at: Part of the Medical Sciences Commons
 Bucknell University Bucknell Digital Commons Master s Theses Student Theses 2010 The overexpression of homeotic complex gene Ultrabithorax in the post-embryonic neuronal lineages of the ventral nervous
Bucknell University Bucknell Digital Commons Master s Theses Student Theses 2010 The overexpression of homeotic complex gene Ultrabithorax in the post-embryonic neuronal lineages of the ventral nervous
18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis
 18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis An organism arises from a fertilized egg cell as the result of three interrelated processes: cell division, cell
18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis An organism arises from a fertilized egg cell as the result of three interrelated processes: cell division, cell
Drosophila melanogaster- Morphogen Gradient
 NPTEL Biotechnology - Systems Biology Drosophila melanogaster- Morphogen Gradient Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by
NPTEL Biotechnology - Systems Biology Drosophila melanogaster- Morphogen Gradient Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles
 Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
178 Part 3.2 SUMMARY INTRODUCTION
 178 Part 3.2 Chapter # DYNAMIC FILTRATION OF VARIABILITY WITHIN EXPRESSION PATTERNS OF ZYGOTIC SEGMENTATION GENES IN DROSOPHILA Surkova S.Yu. *, Samsonova M.G. St. Petersburg State Polytechnical University,
178 Part 3.2 Chapter # DYNAMIC FILTRATION OF VARIABILITY WITHIN EXPRESSION PATTERNS OF ZYGOTIC SEGMENTATION GENES IN DROSOPHILA Surkova S.Yu. *, Samsonova M.G. St. Petersburg State Polytechnical University,
Developmental Biology Lecture Outlines
 Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Reciprocal Interactions between Neurons and Glia Are Required for Drosophila Peripheral Nervous System Development
 The Journal of Neuroscience, September 10, 2003 23(23):8221 8230 8221 Development/Plasticity/Repair Reciprocal Interactions between Neurons and Glia Are Required for Drosophila Peripheral Nervous System
The Journal of Neuroscience, September 10, 2003 23(23):8221 8230 8221 Development/Plasticity/Repair Reciprocal Interactions between Neurons and Glia Are Required for Drosophila Peripheral Nervous System
Anterograde Activin Signaling Regulates Postsynaptic Membrane Potential and GluRIIA/B Abundance at the Drosophila Neuromuscular Junction
 Anterograde Activin Signaling Regulates Postsynaptic Membrane Potential and GluRIIA/B Abundance at the Drosophila Neuromuscular Junction Myung-Jun Kim, Michael B. O Connor* Department of Genetics, Cell
Anterograde Activin Signaling Regulates Postsynaptic Membrane Potential and GluRIIA/B Abundance at the Drosophila Neuromuscular Junction Myung-Jun Kim, Michael B. O Connor* Department of Genetics, Cell
ATranscriptionFactorNetworkCoordinates Attraction, Repulsion, and Adhesion Combinatorially to Control Motor Axon Pathway Selection
 Article ATranscriptionFactorNetworkCoordinates Attraction, Repulsion, and Adhesion Combinatorially to Control Motor Axon Pathway Selection Aref Arzan Zarin, 1,2 Jamshid Asadzadeh, 1,2 Karsten Hokamp, 1
Article ATranscriptionFactorNetworkCoordinates Attraction, Repulsion, and Adhesion Combinatorially to Control Motor Axon Pathway Selection Aref Arzan Zarin, 1,2 Jamshid Asadzadeh, 1,2 Karsten Hokamp, 1
BIS &003 Answers to Assigned Problems May 23, Week /18.6 How would you distinguish between an enhancer and a promoter?
 Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
BILD7: Problem Set. 2. What did Chargaff discover and why was this important?
 BILD7: Problem Set 1. What is the general structure of DNA? 2. What did Chargaff discover and why was this important? 3. What was the major contribution of Rosalind Franklin? 4. How did solving the structure
BILD7: Problem Set 1. What is the general structure of DNA? 2. What did Chargaff discover and why was this important? 3. What was the major contribution of Rosalind Franklin? 4. How did solving the structure
CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON
 PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
Supplementary Information
 Supplementary Information Supplementary Figure 1. JAK/STAT in early wing development (a-f) Wing primordia of second instar larvae of the indicated genotypes labeled to visualize expression of upd mrna
Supplementary Information Supplementary Figure 1. JAK/STAT in early wing development (a-f) Wing primordia of second instar larvae of the indicated genotypes labeled to visualize expression of upd mrna
Neuronal cell death in grasshopper embryos: variable patterns in different species, clutches, and clones
 J. Embryol. exp. Morph. 78, 169-182 (1983) Printed in Great Britain The Company of Biologists Limited 1983 Neuronal cell death in grasshopper embryos: variable patterns in different species, clutches,
J. Embryol. exp. Morph. 78, 169-182 (1983) Printed in Great Britain The Company of Biologists Limited 1983 Neuronal cell death in grasshopper embryos: variable patterns in different species, clutches,
MCB 141 Midterm I Feb. 19, 2009
 Write your name and student ID# on EVERY PAGE of your exam MCB 141 Midterm I Feb. 19, 2009 Circle the name of your TA Jessica Lyons Alberto Stolfi Question #1 Question #2 Question #3 Question #4 TOTAL
Write your name and student ID# on EVERY PAGE of your exam MCB 141 Midterm I Feb. 19, 2009 Circle the name of your TA Jessica Lyons Alberto Stolfi Question #1 Question #2 Question #3 Question #4 TOTAL
Tyrosine kinase inhibition produces specific alterations in axon guidance in the grasshopper embryo
 Development 125, 4121-4131 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV8542 4121 Tyrosine kinase inhibition produces specific alterations in axon guidance in the grasshopper
Development 125, 4121-4131 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV8542 4121 Tyrosine kinase inhibition produces specific alterations in axon guidance in the grasshopper
Distinct functions of the leucine-rich repeat transmembrane proteins Capricious and Tartan in the Drosophila tracheal morphogenesis
 Developmental Biology 296 (2006) 253 264 www.elsevier.com/locate/ydbio Distinct functions of the leucine-rich repeat transmembrane proteins Capricious and Tartan in the Drosophila tracheal morphogenesis
Developmental Biology 296 (2006) 253 264 www.elsevier.com/locate/ydbio Distinct functions of the leucine-rich repeat transmembrane proteins Capricious and Tartan in the Drosophila tracheal morphogenesis
Single-Cell Analysis of Drosophila Larval Neuromuscular Synapses
 Developmental Biology 229, 55 70 (2001) doi:10.1006/dbio.2000.9983, available online at http://www.idealibrary.com on Single-Cell Analysis of Drosophila Larval Neuromuscular Synapses Bao Hoang 1 and Akira
Developmental Biology 229, 55 70 (2001) doi:10.1006/dbio.2000.9983, available online at http://www.idealibrary.com on Single-Cell Analysis of Drosophila Larval Neuromuscular Synapses Bao Hoang 1 and Akira
Bypass and interaction suppressors; pathway analysis
 Bypass and interaction suppressors; pathway analysis The isolation of extragenic suppressors is a powerful tool for identifying genes that encode proteins that function in the same process as a gene of
Bypass and interaction suppressors; pathway analysis The isolation of extragenic suppressors is a powerful tool for identifying genes that encode proteins that function in the same process as a gene of
SUPPLEMENTARY INFORMATION
 Supplementary Notes Downregulation of atlastin does not affect secretory traffic We investigated whether Atlastin might play a role in secretory traffic. Traffic impairment results in disruption of Golgi
Supplementary Notes Downregulation of atlastin does not affect secretory traffic We investigated whether Atlastin might play a role in secretory traffic. Traffic impairment results in disruption of Golgi
Supplemental Data. Wang et al. (2014). Plant Cell /tpc
 Supplemental Figure1: Mock and NPA-treated tomato plants. (A) NPA treated tomato (cv. Moneymaker) developed a pin-like inflorescence (arrowhead). (B) Comparison of first and second leaves from mock and
Supplemental Figure1: Mock and NPA-treated tomato plants. (A) NPA treated tomato (cv. Moneymaker) developed a pin-like inflorescence (arrowhead). (B) Comparison of first and second leaves from mock and
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION
 MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
Signal Transduction. Dr. Chaidir, Apt
 Signal Transduction Dr. Chaidir, Apt Background Complex unicellular organisms existed on Earth for approximately 2.5 billion years before the first multicellular organisms appeared.this long period for
Signal Transduction Dr. Chaidir, Apt Background Complex unicellular organisms existed on Earth for approximately 2.5 billion years before the first multicellular organisms appeared.this long period for
