Opinion Multi-species sequence comparison: the next frontier in genome annotation Inna Dubchak* and Kelly Frazer
|
|
|
- Annis Marjorie Baldwin
- 6 years ago
- Views:
Transcription
1 Opinion Multi-species sequence comparison: the next frontier in genome annotation Inna Dubchak* and Kelly Frazer Addresses: *Genomics Division, Lawrence Berkeley National Laboratory, Berkeley, CA 94720, USA. Perlegen Sciences, 2021 Stierlin Ct., Mountain View, CA 94043, USA. Correspondence: Inna Dubchak. Published: 27 November 2003 The electronic version of this article is the complete one and can be found online at BioMed Central Ltd Abstract Multi-species comparisons of DNA sequences are more powerful for discovering functional sequences than pairwise DNA sequence comparisons. Most current computational tools have been designed for pairwise comparisons, and efficient extension of these tools to multiple species will require knowledge of the ideal evolutionary distance to choose and the development of new algorithms for alignment, analysis of conservation, and visualization of results. Comparison of DNA sequences from different species is an extremely efficient way to identify functional DNA elements - both coding regions and transcriptional control regions that lie beyond the coding sequences of genes. Several recent reviews of comparative sequence analysis [1-5] describe this fast-growing field and the computational resources that are currently available for a wide range of biological investigations. Most of the large-scale comparative studies completed to date have been based on pairwise comparison of sequences; such studies have resulted in the identification of new genes [6-9], and have proved efficient at discovering functional elements in non-coding genomic intervals [10-12]. Several groups have aligned the entire human and mouse genomes [13-15] and have presented comprehensive statistical data on the patterns of DNA conservation between the two species. Recent comparative studies demonstrate that adding additional species to the analysis provides an even more powerful approach for detecting functionally important elements, because characteristic signatures - such as open reading frames and splice-site consensus sequences within genes, and motifs within regulatory elements - are easier to detect when they are conserved in multiple species [16]. For example, a recent large-scale study of over 12 megabases (Mb) of sequences from 12 species, derived from genomic regions orthologous to a 1.8 Mb region on human chromosome 7 that contains ten genes [17], demonstrated that some highly conserved elements revealed by multiple sequence alignments could not be reliably identified with any set of parameters in a pairwise human-mouse alignment. As the number of available complete genome sequences increases, there is a clear need to understand what we can learn from multiple-species sequence comparisons. Studies of this type will require the development of new comparative algorithms and computational tools, such as multi-genome alignment techniques, analysis of conservation and visualization of comparative results. Developing easy-to-use and efficient techniques is not trivial, however, given that the algorithms should be capable of handling a whole range of evolutionary distances between multiple species and of providing new insights into biology. Selecting multiple species for comparative analysis The comparison of DNA sequences between evolutionarily distantly related species, such as humans and pufferfish, which diverged approximately 450 million years ago, comment reviews reports deposited research refereed research interactions information
2 122.2 Genome Biology 2003, Volume 4, Issue 12, Article 122 Dubchak and Frazer primarily identifies the coding sequences as conserved [18] - because transcribed protein coding sequences are highly functionally constrained, and thus change very slowly during evolution. The comparison of DNA sequences between species that diverged from a common ancestor around million years ago - such as humans and mice, or two species of fruitflies (Drosophila melanogaster and Drosophila pseudoobscura), or two species of nematodes (Caenorhabditis elegans and Caenorhabditis briggsae) [19] - results in identifying as evolutionarily conserved both coding sequences and a significant number of noncoding sequences. Only a limited number of conserved noncoding sequences that have been identified through sequence comparisons between species at this evolutionary distance have been characterized functionally, however. Among those that have had their functions assigned are transcriptional regulatory elements of genes in close proximity [11] or genes as far away from each other as 200 kb [10]. Comparative analyses of genomic DNA from closely related species, such as humans and chimpanzees, on the other hand, identifies those sequences that have changed in recent evolutionary history [20,21]. Some of these sequence changes may have been partly responsible for the speciation of ancestral primates. Thus, comparison of a segment of DNA with the sequences of multiple species at different evolutionary distances allows one to identify coding sequences, conserved noncoding elements with regulatory functions, and those sequences that are unique for a given species. A recent report by Cooper et al. [22] proposed a method for quantitatively assessing the effectiveness of a comparative sequence analysis to identify new information in a genome: it uses the phylogenetic scope, representing the range of organisms that share a last common ancestor whose sequence can be inferred by adding each genome to the analysis. The comparative studies described below demonstrate that the evolutionary distance of the species in a sequence comparison analysis is critical for discovering potentially functional sequence elements. The stem cell leukemia genomic interval For many years the mouse and human genomes, which diverged from a common ancestor about million years ago, have been extensively used for comparative studies [9-12]; but it is still an open question as to which species should be added to this comparative analysis to derive the most information content. Among several recent studies providing guidance for selecting additional species is the investigation of the stem cell leukemia (SCL) genomic interval, originally based on a human-mouse sequence comparison [23], and later expanded to include three additional species: chicken, pufferfish and zebrafish [12]. This analysis demonstrates that mouse-human alignments show high levels of sequence similarity for all coding exons and for all eight known murine regulatory regions of the SCL locus. Human-mouse-chicken alignments identified the similarity of all coding exons and also discovered protein-binding motifs in five of the known regulatory regions. Thus, inclusion of the chicken DNA sequences allowed for superior functional annotation of a subset of the regulatory regions that had already been identified by the human-mouse comparison. Pairwise mouse-pufferfish and mouse-zebrafish sequence alignments identified only some of the coding exons, and found similarity for only two of the eight known regulatory regions in the pufferfish comparison; and no significant similarity was found for any known regulatory region in the zebrafish comparison [12]. This analysis suggests that comparative analysis of zebrafish and mammalian genomic sequences might be of limited value for the identification of functionally significant noncoding sequences in the SCL region; and these results are consistent with what is expected on the basis of the evolutionary distance of the species analyzed. Drosophila melanogaster compared with other species The analysis of conservation between Drosophila melanogaster and four other Drosophila species (D. erecta, D. pseudoobscura, D. willistoni, and D. littoralis) that have different divergence times (6-15, 46, 53 and million years, respectively) [24] has generated several important conclusions to guide further functional studies of these species [25]. One conclusion is that the addition of a third species could reveal functional constraints in otherwise nonsignificant pairwise exon comparisons. All D. melanogaster genes identified in divergent species show evidence of functional constraint; and including more distantly related species defines the exact position of short regulatory elements that are hard to find in the long regions of non-coding sequence conservation observed in closely related species. Non-coding conserved sequences have also been found to be spatially clustered, and these clusters can be used to predict enhancer sequences [25]. This work provided a solid basis for choosing species whose genome sequences would be most useful in aiding the functional annotation of coding and cis-regulatory sequences in D. melanogaster: D. pseudoobscura, which has recently been sequenced, was recognized as the best for discovery of functional genomic features, and adding D. willistoni to the comparison allows the dissection of regions of the Drosophila genome under different levels of functional constraint. Saccharomyces cerevisiae Using multiple alignments in the S. cerevisiae and related fungal genome annotation projects [26] provided a powerful demonstration of functional analysis, yielding results that would be difficult to obtain by other computational and experimental methods. A comprehensive comparison of the genome of the yeast S. cerevisiae with those of three related Saccharomyces species (S. paradoxus, S. mikatae and S. bayanus) [26] yielded a major revision to the yeast gene catalog, reducing the total count by about 500 genes. In addition, motif analysis automatically identified a
3 Genome Biology 2003, Volume 4, Issue 12, Article 122 Dubchak and Frazer number of genome-wide elements, including most known regulatory motifs and numerous new motifs suitable for biological study. Multiple primate analysis Another approach to multiple species sequence analysis, phylogenetic shadowing [21], is used for comparison of evolutionarily closely related species. It demonstrates the utility of sequence comparisons within the primate group for discovering common mammalian, as well as primatespecific, functional elements in the human genome, which could not be achieved by comparison of more evolutionarily distant species. Rubin and colleagues [21] showed that the high information content of comprehensive primate sequence comparisons could be captured with a small subset of phylogenetically close primates, such that sequence from as few as four or six primate species compared with humans might be sufficient for the identification of a large fraction of functional elements in the human genome, many of which are likely to be missed by humanmouse comparisons. While the number of multi-species comparative studies grows, it is becoming clear that reasonable selection of species for comparison of a particular genomic interval is still to a large extent an intuitive process, with some guidance from previous successful comparative studies. Multi-species sequence alignment and analysis of conservation As well as selecting a set of species that provide maximum functional content, the quality of the sequence alignment must also be sufficient to the task in hand. Single pairwise comparisons of sequences do not allow for the detection of conserved sequence strings with high precision, given that functional elements - such as transcriptional-regulator binding sites - are quite short compared to the surrounding nonfunctional sequence. Thus, functional signals can sometimes be indistinguishable from the noise that results from aligning divergent nonfunctional sequences. The hope is that multiple sequence alignment provides a way to increase the sensitivity of the search for regulatory signals. The area of sequence alignment is well developed, but many of its problems are far from being completely resolved, especially for multiple species [27]. Alignment methods can be roughly divided into local alignments, which produce optimal similarity scores between subregions of the two sequences, and global alignments, which generate optimal similarity scores over the entire length of the two sequences. Global alignments attempt to find a monotonically increasing map between the letters of each sequence, in the process rejecting alignments that overlap or cross over. A recently published review on comparative genomics gives more details of the various kinds of alignment [1]. Unfortunately, a comprehensive study of the strengths and weaknesses of different alignments algorithms applied to different biological problems has yet to appear. The local and global alignment methods that generate pairwise comparisons can also be used for multiple species, but multiple alignments are considerably more difficult to compute because of statistical complexity and the difficulties of scoring the results. Progressive multiple alignment is a heuristic technique that uses successive applications of a pairwise alignment algorithm. The best-known progressive alignment program, CLULSTALW [28], is very efficient in aligning proteins and short nucleotide sequences, but it is not suitable for long genomic regions [29]. Below we describe new alignment techniques that can handle long DNA sequences efficiently. Global alignments Two recently developed algorithms, MLAGAN [16,30] and MAVID [31,32], are designed for global alignment of both evolutionarily close and distant megabase-length genomic sequences. The MLAGAN [16,30] algorithm assumes that the phylogenetic tree is known, as is usually the case for large vertebrate genomes. The program is based on progressive alignment: a multiple alignment of K sequences is constructed in K-1 pairwise alignment steps, such that in each step two sequences, or intermediate multiple alignments, are aligned. MLAGAN uses LAGAN as the global pairwise-alignment subroutine, and introduces new methods for scoring and refining a multiple alignment. It also aligns the sequences in the order of the given phylogenetic tree. For example, MLAGAN aligns sequences from human, chimpanzee, mouse, rat, and chicken, in the following order: first, human-chimpanzee; second, mouse-rat; third, human-chimpanzee to mouse-rat; fourth, human-chimpanzee-mouse-rat to chicken. Each alignment step merges two sequences or alignments into a larger alignment, effectively building a profile of all the sequences. The results obtained with MLAGAN on the cystic fibrosis (CFTR) genomic region [16], suggest that multiple alignments are better than pairwise alignments at aligning conserved exons between distant species: it was precise enough to refine mis-annotated splicing sites. MAVID [31,32] is a progressive global alignment program that works by recursively aligning the alignments at ancestral nodes of the guide phylogenetic tree. At each internal node, ancestral sequences are inferred from the existing alignments using maximum likelihood, and these alignments are then aligned using the global aligner AVID [33]. The multiple alignment is used to build a phylogenetic tree for the sequences, which is subsequently used as a basis for identifying conserved regions in the alignment. Local alignments MultiPipMaker [34,35] uses multiple pairwise local alignments of secondary sequences against a reference sequence comment reviews reports deposited research refereed research interactions information
4 122.4 Genome Biology 2003, Volume 4, Issue 12, Article 122 Dubchak and Frazer to create a crude multiple alignment that is subsequently refined to generate a true multiple alignment. Analysis of multiple alignments generated by MultiPipMaker [35] allowed for discovery of regulatory elements in the mammalian WNT2 genomic region, and confirmed the phylogenetic inference that horses are evolutionarily more closely related to cats than to cows [17]. Alignments between the human sequence and the sequence of each of the other 12 species used in the analysis [17] showed, as expected, that the fraction of sequence that can be aligned generally decreases with increasing evolutionary distance from humans (except for mouse and rat). Another program, Multiz, developed for large-scale comparison of multiple sequences, takes BLASTZ/axtBest [35] as the pairwise input. This program has been used for the alignment of the mouse and the rat draft assemblies to the human genome [36]. Motif finding Phylogenetic footprinting [37] aims to discover specific protein-binding sites within regulatory regions of multiple sequences on the basis of phylogenetic relationships. It is a method that is mostly applied to promoter regions of orthologous genes. Sumiyama with coauthors [38] attained good results by using multiple sequence comparison combined with a small window size (where the window is the region analyzed in each sub-comparison). This high-resolution procedure can predict the binding sites of transcription factors and reveal polymorphisms in control elements between phylogenetic clades. Phylogenetic footprinting was applied to the Hoxc8 early enhancer region, where it successfully identified a known protein-binding cis-regulatory motif that had previously been analyzed in depth by functional methods [38]. The authors demonstrated that an eight-species analysis is clearly superior to the conventional two-species methodology for this type of study. Another group of specialized phylogenetic footprinting algorithms finds the most conserved motifs among the input sequences, as measured by a parsimony score of the underlying phylogenetic tree [39,40]. These algorithms have been used successfully to identify a variety of regulatory elements, some known and some novel, in sets of diverse vertebrate DNA sequences. Although phylogenetic footprinting methods show a lot of promising results, their use requires prior information about the location of orthologous regions in genomic intervals of interest. Multiple sequence alignments can help in defining these regions by finding longer conserved regions that can serve as guides to functionally important elements [10]. Analysis of conservation The most obvious but difficult question to ask in comparative studies is how to define a functionally significant level of sequence conservation between species. Although two-way comparison is effective for discovery of evolutionarily constrained elements, distinguishing them from conserved sequences that are present due to lack of sufficient divergence time is not straightforward and requires knowledge of the neutral substitution rate [13,41]. In the majority of comparative genomics studies the definition of a significant level of conservation between two species has been intuitive, or based on biological experience. For example, aligning sequences in divergent noncoding regions proved useful in analyzing the enhancer in the -globin locus-control region [42]. The conventional cutoff of 70-75% conservation over 100 base-pairs for the human-mouse comparison has produced discoveries of several important biologically functional elements [10,43]. One of the major obstacles to applying a single universal conservation criterion for potential regulatory regions is the substantial variation in the underlying mutation rates from region to region [13,41]. Conservation scores that incorporate the local neutral substitution rate are now available for the human and mouse genomes [13], and they can help to determine if a particular sequence is likely to be functional. A more detailed analysis of interspecies pairwise genomic sequence alignments, aiming to distinguish regulatory regions from neutrally evolving DNA, has appeared recently [44]. This study proposed scoring procedures that evaluate alignments for properties other than overall percentage identity, although highly conserved noncoding sequences have proven to be good indicators of regulatory elements; among these procedures are discrimination on the basis of frequencies of nucleotide pairs or gaps, in combination with scoring procedures that include the alignment context, using frequencies of short runs of alignment columns. This study [44] thus gave a good start for extensive testing of these measures. Adding genomic sequences from multiple vertebrates to the analysis makes the problem of estimating conservation even less trivial. Expanding pairwise analyses of conservation to multiple sequence alignments would require calculation of the neutral substitution rate between all pairs of sequences. That would give a weighted contribution of each sequence in the multiple alignment, but would also requires much more detailed evolutionary information than is available now. A three-way comparison makes it possible to enrich a pairwise alignment, and a simplified method for calculating a level of active non-coding conservation in such a comparison [45] is based on the supposition that actively conserved humanmouse noncoding sequences are likely to be present in additional mammals, whereas noncoding regions that are similar because of an insufficient accumulation of random mutations will not be present in other mammals. Visualization of results Visualization of results is a critical aspect of comparative sequence analysis, since manual examination of alignment
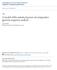
5 Genome Biology 2003, Volume 4, Issue 12, Article 122 Dubchak and Frazer on the scale of long genomic regions presents a significant challenge and is not efficient. Alignment-browsing systems should identify regions that exhibit properties suggestive of a particular biological function, for example well-conserved segments within an alignment, or matching the consensus sequence for a specific transcription-factor-binding site [27]. There are several publicly available visualization tools for long pairwise DNA alignments. PipMaker [15,34] represents the level of conservation in ungapped regions of a BLASTZ local alignment as horizontal dashes. VISTA [45-47] displays comparative data in the form of a curve, where conservation is calculated in a sliding window of a gapped global alignment. SynPlot [23] also generates a curve plot calculated from a global alignment, but displays it slightly differently. All three tools can also be used to visualize multiple pairwise alignments [1,12,23,34], but one of the sequences needs to be selected as a reference, and the level of conservation is displayed on its scale. The same principle of selecting a reference sequence is utilized for whole genomes in the UCSC genome browser [36,48] and the VISTA browser [14,49]. Figure 1 shows a multiple pairwise VISTA display of a 5 kilobase fragment of the CFTR region aligned by MLAGAN [16]. This view is based on the coordinates of the human sequence and displays the level of conservation between human and all other sequences in the multiple alignment. The first exon of the CAV1 gene is clearly well conserved across all 11 species, including the pufferfish Fugu. The upstream region of the CAV1 gene (at 183 kb) has a distinct area of noncoding conservation across most of the pairwise comparisons, ranging from human/mouse to human/chicken. On the other hand, there are some peaks of non-coding conservation (at 181 kb) that are found in some mammalian species, but not others. Knowing the phylogenetic relationship among species is important for building and analyzing multiple alignments, so visualizing sequence alignment data while taking phylogenetic trees into account presents a significant advance. A recently developed new program from the VISTA family, Phylo-VISTA (short for Phylogenetic VISTA) [50], uses phylogenetic relationships as a guide to display and analyze the level of conservation across internal nodes of the phylogenetic tree. Using the entire multiple alignment, not a reference sequence, as a base in the x axis of the visualization allows for additional options, such as presentation of comparative data together with available annotations for all sequences, and computation of a measure of similarity for any node of the phylogenetic tree. In conclusion, pairwise sequence comparisons of the complete genomes of human and mouse have brought the revolutionary discovery that more than half of the functionally conserved sequences in the human genome are not proteinencoding [13]. Unfortunately, pairwise studies also make it CAV1 180, , , , , ,000 Non-coding Coding Non-coding Chimp Baboon Cat Dog Cow Pig Mouse Rat Chicken Fugu Zebrafish Figure 1 Multi-VISTA display of a 5 kilobase fragment of the MLAGAN alignment of the CFTR region [16]. This view is based on the coordinates of the human sequence and displays the level of conservation between human and all other sequences in the multiple alignment. A fragment of the CAV1 gene is shown as an arrow above the plots. The following cutoffs were used to show the conserved regions: above 80% over 100 bp for chimpanzee, baboon, cat, dog, cow, pig, mouse and rat; above 65% over 100 bp for chicken; above 50% over 100 bp for Fugu and zebrafish. clear that functional noncoding sequences are not easily distinguished from non-functional segments that happen to have accumulated very few mutations since the last common ancestor. Initial reports suggest that multi-species DNA comparisons have greater potential for filtering out evolutionarily neutral regions, and should therefore provide a more reliable basis for decoding and annotating genomic sequences at high resolution. This would improve our ability to discover non-protein-coding functional elements, which are currently poorly understood in comparison to their coding counterparts. Thus, we face the exciting prospect of discovering which species are the most informative in comparative studies, developing sophisticated algorithms for multi-sequence alignment and analysis of conservation, and building new effective visualization techniques for comparative data. Acknowledgements The authors are grateful to Nameeta Shah, Michael Brudno, Olivier Couronne, Shyam Prabhakar, Len Pennacchio and Alexander Poliakov for help and discussion. ID was partially supported by the Programs for Genomic Applications grant from the NHLBI/NIH. References 1. Frazer KA, Elnitski L, Church DM, Dubchak I, Hardison RC: Crossspecies sequence comparisons: a review of methods and available resources. Genome Res 2003, 13: Pennacchio LA, Rubin EM: Genomic strategies to identify mammalian regulatory sequences. Nat Rev Genet 2001, 2: Sidow A: Sequence first. Ask questions later. Cell 2002, 111: comment reviews reports deposited research refereed research interactions information
6 122.6 Genome Biology 2003, Volume 4, Issue 12, Article 122 Dubchak and Frazer 4. Ureta-Vidal A, Ettwiller L, Birney E: Comparative genomics: genome-wide analysis in metazoan eukaryotes. Nat Rev Genet 2003, 4: Wei L, Liu Y, Dubchak I, Shon J, Park J: Comparative genomics approaches to study organism similarities and differences. J Biomed Inform 2002, 35: Batzoglou S, Pachter L, Mesirov JP, Berger B, Lander ES: Human and mouse gene structure: comparative analysis and application to exon prediction. Genome Res 2000, 10: Korf I, Flicek P, Duan D, Brent MR: Integrating genomic homology into gene structure prediction. Bioinformatics 2001, 17 Suppl 1:S140-S Guigo R, Dermitzakis ET, Agarwal P, Ponting CP, Parra G, Reymond A, Abril JF, Keibler E, Lyle R, Ucla C, Antonarakis SE, Brent MR: Comparison of mouse and human genomes followed by experimental verification yields an estimated 1,019 additional genes. Proc Natl Acad Sci USA 2003, 100: Pennacchio LA, Olivier M, Hubacek JA, Cohen JC, Cox DR, Fruchart JC, Krauss RM, Rubin EM: An apolipoprotein influencing triglycerides in humans and mice revealed by comparative sequencing. Science 2001, 294: Loots GG, Locksley RM, Blankespoor CM, Wang ZE, Miller W, Rubin EM, Frazer KA: Identification of a coordinate regulator of interleukins 4, 13, and 5 by cross-species sequence comparisons. Science 2000, 288: Hardison RC, Oeltjen J, Miller W: Long human-mouse sequence alignments reveal novel regulatory elements: a reason to sequence the mouse genome. Genome Res 1997, 7: Gottgens B, Barton LM, Chapman MA, Sinclair AM, Knudsen B, Grafham D, Gilbert JG, Rogers J, Bentley DR, Green AR: Transcriptional regulation of the stem cell leukemia gene (SCL) - comparative analysis of five vertebrate SCL loci. Genome Res 2002, 12: Waterston RH, Lindblad-Toh K, Birney E, Rogers J, Abril, JF, Agarwal P, Agarwala R, Ainscough R, Alexandersson M, An P, et al.: Initial sequencing and comparative analysis of the mouse genome. Nature 2002, 420: Couronne O, Poliakov A, Bray N, Ishkhanov T, Ryaboy D, Rubin EM, Pachter L, Dubchak I: Strategies and tools for whole genome alignments. Genome Res, 2003, 13: Schwartz S, Kent WJ, Smit A, Zhang Z, Baertsch R, Hardison RC, Haussler D, Miller W: Human-mouse alignments with BLASTZ. Genome Res 2003, 13: Brudno M, Do CB, Cooper GM, Kim MF, Davydov E, Green ED, Sidow A, Batzoglou S, NISC Comparative Sequencing Program: LAGAN and Multi-LAGAN: efficient tools for large-scale multiple alignment of genomic DNA. Genome Res 2003, 13: Thomas JW, Touchman JW, Blakesley RW, Bouffard GG, Beckstrom-Sternberg SM, Margulies EH, Blanchette M, Siepel AC, Thomas PJ, McDowell JC et al.: Comparative analyses of multispecies sequences from targeted genomic regions. Nature 2003, 424: Aparicio S, Chapman J, Stupka E, Putnam N, Chia JM, Dehal P, Christoffels A, Rash S, Hoon S, Smit A, et al.: Whole-genome shotgun assembly and analysis of the genome of Fugu rubripes. Science 2002, 297: Kent WJ, Zahler AM: Conservation, regulation, synteny, and introns in a large-scale C. briggsae - C. elegans genomic alignment. Genome Res 2000, 10: Frazer KA, Chen X, Hinds DA, Pant PV, Patil N, Cox DR: Genomic DNA insertions and deletions occur frequently between humans and nonhuman primates. Genome Res 2003, 13: Boffelli D, McAuliffe J, Ovcharenko D, Lewis KD, Ovcharenko I, Pachter L, Rubin EM: Phylogenetic shadowing of primate sequences to find functional regions of the human genome. Science 2003, 299: Cooper GM, Brudno M, Green ED, Batzoglou S, Sidow A, NISC Comparative Sequencing Program: Quantitative estimates of sequence divergence for comparative analyses of mammalian genomes. Genome Res 2003, 13: Gottgens B, Gilbert JG, Barton LM, Grafham D, Rogers J, Bentley DR, Green AR: Long-range comparison of human and mouse SCL loci: localized regions of sensitivity to restriction endonucleases correspond precisely with peaks of conserved noncoding sequences. Genome Res 2001, 11: Powell JR: Progress and Prospects in Evolutionary Biology: The Drosophila Model. Oxford: Oxford University Press; Bergman CM, Pfeiffer BD, Rincon-Limas DE, Hoskins RA, Gnirke A, Mungall CJ, Wang AM, Kronmiller B, Pacleb J, Park S, et al.: Assessing the impact of comparative genomic sequence data on the functional annotation of the Drosophila genome. Genome Biol 2002, 3:research Kellis M, Patterson N, Endrizzi M, Birren B, Lander ES: Sequencing and comparison of yeast species to identify genes and regulatory elements. Nature 2003, 423: Miller W: Comparison of genomic DNA sequences: solved and unsolved problems. Bioinformatics 2001, 17: Thompson JD, Higgins DG, Gibson TJ: CLUSTALW: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position specific gap penalties and weight matrix choice. Nucleic Acids Res 1994, 22: Thompson JD, Plewniak F, Poch O: A comprehensive comparison of multiple sequence alignment programs. Nucleic Acids Res 1999, 27: MLAGAN [ 31. Bray N, Pachter L: The MAVID multiple alignment server. Nucleic Acids Res 2003, 31: MAVID [ 33. Bray N, Dubchak I, Pachter L: AVID: a global alignment program. Genome Res 2003, 13: PipMaker and MultiPipMaker [ 35. Schwartz S, Elnitski L, Li M, Weirauch M, Riemer C, Smit A, Green ED, Hardison RC, Miller W, NISC Comparative Sequencing Program: MultiPipMaker and supporting tools: alignments and analysis of multiple genomic DNA sequences. Nucleic Acids Res 2003, 31: Genome browser at UCSC [ 37. Tagle DA, Koop BF, Goodman M, Slightom JL, Hess DL, Jones RT: Embryonic epsilon and gamma globin genes of a prosimian primate (Galago crassicaudatus). Nucleotide and amino acid sequences, developmental regulation and phylogenetic footprints. J Mol Biol 1988, 203: Sumiyama K, Kim CB, Ruddle FH: An efficient cis-element discovery method using multiple sequence comparisons based on evolutionary relationships. Genomics 2001, 71: Tompa M: Identifying functional elements by comparative DNA sequence analysis. Genome Res 2001, 11: Blanchette M, Tompa M: Discovery of regulatory elements by a computational method for phylogenetic footprinting. Genome Res 2002, 12: Hardison RC: Conserved noncoding sequences are reliable guides to regulatory elements. Trends Genet 2000, 16: Hardison RC, Roskin KM, Yang S, Diekhans M, Kent WJ, Weber R, Elnitski L, Li J, O Connor M, Kolbe D, et al.: Covariation in frequencies of substitution, deletion, transposition, and recombination during eutherian evolution. Genome Res 2003, 13: Elnitski L, Miller W, Hardison R: Conserved E boxes function as part of the enhancer in hypersensitive site 2 of the -globin locus control region. Role of basic helix-loop-helix proteins. J Biol Chem 1997, 272: Elnitski L, Hardison RC, Li J, Yang S, Kolbe D, Eswara P, O Connor MJ, Schwartz S, Miller W, Chiaromonte F: Distinguishing regulatory DNA from neutral sites. Genome Res 2003, 13: Dubchak I, Brudno M, Pachter LS, Loots GG, Mayor C, Rubin EM, Frazer KA: Active conservation of noncoding sequences revealed by three-way species comparisons. Genome Res 2000, 10: Mayor C, Brudno M, Schwartz JR, Poliakov A, Rubin EM, Frazer KA, Pachter LS, Dubchak I: VISTA: visualizing global DNA sequence alignments of arbitrary length. Bioinformatics 2000, 16: VISTA [ 48. Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, Haussler D: The human genome browser at UCSC. Genome Res 2002, 12: VISTA Genome Browser [ 50. Shah N, Couronne O, Pennacchio LA, Brudno M, Batzoglou S, Bethel E, Rubin EM, Hamann B, Dubchak I: Phylo-VISTA: an interactive visualization tool for multiple DNA sequence alignments. Bioinformatics, in press.
Contact 1 University of California, Davis, 2 Lawrence Berkeley National Laboratory, 3 Stanford University * Corresponding authors
 Phylo-VISTA: Interactive Visualization of Multiple DNA Sequence Alignments Nameeta Shah 1,*, Olivier Couronne 2,*, Len A. Pennacchio 2, Michael Brudno 3, Serafim Batzoglou 3, E. Wes Bethel 2, Edward M.
Phylo-VISTA: Interactive Visualization of Multiple DNA Sequence Alignments Nameeta Shah 1,*, Olivier Couronne 2,*, Len A. Pennacchio 2, Michael Brudno 3, Serafim Batzoglou 3, E. Wes Bethel 2, Edward M.
Multiple Alignment of Genomic Sequences
 Ross Metzger June 4, 2004 Biochemistry 218 Multiple Alignment of Genomic Sequences Genomic sequence is currently available from ENTREZ for more than 40 eukaryotic and 157 prokaryotic organisms. As part
Ross Metzger June 4, 2004 Biochemistry 218 Multiple Alignment of Genomic Sequences Genomic sequence is currently available from ENTREZ for more than 40 eukaryotic and 157 prokaryotic organisms. As part
Comparative Genomics. Primer. Ross C. Hardison
 Primer Comparative Genomics Ross C. Hardison A complete genome sequence of an organism can be considered to be the ultimate genetic map, in the sense that the heritable characteristics are encoded within
Primer Comparative Genomics Ross C. Hardison A complete genome sequence of an organism can be considered to be the ultimate genetic map, in the sense that the heritable characteristics are encoded within
Homolog. Orthologue. Comparative Genomics. Paralog. What is Comparative Genomics. What is Comparative Genomics
 Orthologue Orthologs are genes in different species that evolved from a common ancestral gene by speciation. Normally, orthologs retain the same function in the course of evolution. Identification of orthologs
Orthologue Orthologs are genes in different species that evolved from a common ancestral gene by speciation. Normally, orthologs retain the same function in the course of evolution. Identification of orthologs
Lawrence Berkeley National Laboratory Lawrence Berkeley National Laboratory
 Lawrence Berkeley National Laboratory Lawrence Berkeley National Laboratory Title Automated whole-genome multiple alignment of rat, mouse, and human Permalink https://escholarship.org/uc/item/1z58c37n
Lawrence Berkeley National Laboratory Lawrence Berkeley National Laboratory Title Automated whole-genome multiple alignment of rat, mouse, and human Permalink https://escholarship.org/uc/item/1z58c37n
Supplementary text and figures: Comparative assessment of methods for aligning multiple genome sequences
 Supplementary text and figures: Comparative assessment of methods for aligning multiple genome sequences Xiaoyu Chen Martin Tompa Department of Computer Science and Engineering Department of Genome Sciences
Supplementary text and figures: Comparative assessment of methods for aligning multiple genome sequences Xiaoyu Chen Martin Tompa Department of Computer Science and Engineering Department of Genome Sciences
Frazer et al. ago (Aparicio et al. 2002), conserved long-range sequence organization has not been reported for more distantly related species. Figure
 Review Cross-Species Sequence Comparisons: A Review of Methods and Available Resources Kelly A. Frazer, 1,6 Laura Elnitski, 2,3 Deanna M. Church, 4 Inna Dubchak, 5 and Ross C. Hardison 3 1 Perlegen Sciences,
Review Cross-Species Sequence Comparisons: A Review of Methods and Available Resources Kelly A. Frazer, 1,6 Laura Elnitski, 2,3 Deanna M. Church, 4 Inna Dubchak, 5 and Ross C. Hardison 3 1 Perlegen Sciences,
Supplemental Information for Pramila et al. Periodic Normal Mixture Model (PNM)
 Supplemental Information for Pramila et al. Periodic Normal Mixture Model (PNM) The data sets alpha30 and alpha38 were analyzed with PNM (Lu et al. 2004). The first two time points were deleted to alleviate
Supplemental Information for Pramila et al. Periodic Normal Mixture Model (PNM) The data sets alpha30 and alpha38 were analyzed with PNM (Lu et al. 2004). The first two time points were deleted to alleviate
Handling Rearrangements in DNA Sequence Alignment
 Handling Rearrangements in DNA Sequence Alignment Maneesh Bhand 12/5/10 1 Introduction Sequence alignment is one of the core problems of bioinformatics, with a broad range of applications such as genome
Handling Rearrangements in DNA Sequence Alignment Maneesh Bhand 12/5/10 1 Introduction Sequence alignment is one of the core problems of bioinformatics, with a broad range of applications such as genome
Multiple Genome Alignment by Clustering Pairwise Matches
 Multiple Genome Alignment by Clustering Pairwise Matches Jeong-Hyeon Choi 1,3, Kwangmin Choi 1, Hwan-Gue Cho 3, and Sun Kim 1,2 1 School of Informatics, Indiana University, IN 47408, USA, {jeochoi,kwchoi,sunkim}@bio.informatics.indiana.edu
Multiple Genome Alignment by Clustering Pairwise Matches Jeong-Hyeon Choi 1,3, Kwangmin Choi 1, Hwan-Gue Cho 3, and Sun Kim 1,2 1 School of Informatics, Indiana University, IN 47408, USA, {jeochoi,kwchoi,sunkim}@bio.informatics.indiana.edu
Ensembl focuses on metazoan (animal) genomes. The genomes currently available at the Ensembl site are:
 Comparative genomics and proteomics Species available Ensembl focuses on metazoan (animal) genomes. The genomes currently available at the Ensembl site are: Vertebrates: human, chimpanzee, mouse, rat,
Comparative genomics and proteomics Species available Ensembl focuses on metazoan (animal) genomes. The genomes currently available at the Ensembl site are: Vertebrates: human, chimpanzee, mouse, rat,
Reading the French version
 Journal Club Army Ants Trapped by Their Evolutionary History Frédéric Delsuc Reading the French version of Journey to the Ants by Bert Hölldobler and Edward O. Wilson (1996) during the course of my graduate
Journal Club Army Ants Trapped by Their Evolutionary History Frédéric Delsuc Reading the French version of Journey to the Ants by Bert Hölldobler and Edward O. Wilson (1996) during the course of my graduate
Whole Genome Alignments and Synteny Maps
 Whole Genome Alignments and Synteny Maps IINTRODUCTION It was not until closely related organism genomes have been sequenced that people start to think about aligning genomes and chromosomes instead of
Whole Genome Alignments and Synteny Maps IINTRODUCTION It was not until closely related organism genomes have been sequenced that people start to think about aligning genomes and chromosomes instead of
Comparative Genomics. Chapter for Human Genetics - Principles and Approaches - 4 th Edition
 Chapter for Human Genetics - Principles and Approaches - 4 th Edition Editors: Friedrich Vogel, Arno Motulsky, Stylianos Antonarakis, and Michael Speicher Comparative Genomics Ross C. Hardison Affiliations:
Chapter for Human Genetics - Principles and Approaches - 4 th Edition Editors: Friedrich Vogel, Arno Motulsky, Stylianos Antonarakis, and Michael Speicher Comparative Genomics Ross C. Hardison Affiliations:
Vertebrate genome sequencing: building a backbone for comparative genomics
 104 Forum Web Watch Vertebrate genome sequencing: building a backbone for comparative genomics James W. Thomas and Jeffrey W. Touchman The human genome sequence provides a reference point from which we
104 Forum Web Watch Vertebrate genome sequencing: building a backbone for comparative genomics James W. Thomas and Jeffrey W. Touchman The human genome sequence provides a reference point from which we
Eric A. Stone, 1,2 Gregory M. Cooper, 3 and Arend Sidow 2,3 INTRODUCTION
 Annu. Rev. Genomics Hum. Genet. 2005. 6:143 64 doi: 10.1146/annurev.genom.6.080604.162146 Copyright c 2005 by Annual Reviews. All rights reserved First published online as a Review in Advance on April
Annu. Rev. Genomics Hum. Genet. 2005. 6:143 64 doi: 10.1146/annurev.genom.6.080604.162146 Copyright c 2005 by Annual Reviews. All rights reserved First published online as a Review in Advance on April
Analyses of deep mammalian sequence alignments and constraint predictions for 1% of the human genome
 Article Analyses of deep mammalian sequence alignments and constraint predictions for 1% of the human genome Elliott H. Margulies, 2,7,8,21 Gregory M. Cooper, 2,3,9 George Asimenos, 2,10 Daryl J. Thomas,
Article Analyses of deep mammalian sequence alignments and constraint predictions for 1% of the human genome Elliott H. Margulies, 2,7,8,21 Gregory M. Cooper, 2,3,9 George Asimenos, 2,10 Daryl J. Thomas,
Distribution and intensity of constraint in mammalian genomic sequence
 Article Distribution and intensity of constraint in mammalian genomic sequence Gregory M. Cooper, 1 Eric A. Stone, 2,3 George Asimenos, 4 NISC Comparative Sequencing Program, 5 Eric D. Green, 5 Serafim
Article Distribution and intensity of constraint in mammalian genomic sequence Gregory M. Cooper, 1 Eric A. Stone, 2,3 George Asimenos, 4 NISC Comparative Sequencing Program, 5 Eric D. Green, 5 Serafim
Genomes and Their Evolution
 Chapter 21 Genomes and Their Evolution PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 21 Genomes and Their Evolution PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Evolution at the nucleotide level: the problem of multiple whole-genome alignment
 Human Molecular Genetics, 2006, Vol. 15, Review Issue 1 doi:10.1093/hmg/ddl056 R51 R56 Evolution at the nucleotide level: the problem of multiple whole-genome alignment Colin N. Dewey 1, * and Lior Pachter
Human Molecular Genetics, 2006, Vol. 15, Review Issue 1 doi:10.1093/hmg/ddl056 R51 R56 Evolution at the nucleotide level: the problem of multiple whole-genome alignment Colin N. Dewey 1, * and Lior Pachter
Analysis of Genome Evolution and Function, University of Toronto, Toronto, ON M5R 3G4 Canada
 Multiple Whole Genome Alignments Without a Reference Organism Inna Dubchak 1,2, Alexander Poliakov 1, Andrey Kislyuk 3, Michael Brudno 4* 1 Genome Sciences Division, Lawrence Berkeley National Laboratory,
Multiple Whole Genome Alignments Without a Reference Organism Inna Dubchak 1,2, Alexander Poliakov 1, Andrey Kislyuk 3, Michael Brudno 4* 1 Genome Sciences Division, Lawrence Berkeley National Laboratory,
Transcription Regulation and Gene Expression in Eukaryotes FS08 Pharmacenter/Biocenter Auditorium 1 Wednesdays 16h15-18h00.
 Transcription Regulation and Gene Expression in Eukaryotes FS08 Pharmacenter/Biocenter Auditorium 1 Wednesdays 16h15-18h00. Promoters and Enhancers Systematic discovery of transcriptional regulatory motifs
Transcription Regulation and Gene Expression in Eukaryotes FS08 Pharmacenter/Biocenter Auditorium 1 Wednesdays 16h15-18h00. Promoters and Enhancers Systematic discovery of transcriptional regulatory motifs
A model of the statistical power of comparative genome sequence analysis
 Washington University School of Medicine Digital Commons@Becker Open Access Publications 2005 A model of the statistical power of comparative genome sequence analysis Sean R. Eddy Washington University
Washington University School of Medicine Digital Commons@Becker Open Access Publications 2005 A model of the statistical power of comparative genome sequence analysis Sean R. Eddy Washington University
Computation and Analysis of Genomic Multi-Sequence Alignments
 Annu. Rev. Genomics Hum. Genet. 2007. 8:193 213 First published online as a Review in Advance on May 9, 2007. The Annual Review of Genomics and Human Genetics is online at genom.annualreviews.org This
Annu. Rev. Genomics Hum. Genet. 2007. 8:193 213 First published online as a Review in Advance on May 9, 2007. The Annual Review of Genomics and Human Genetics is online at genom.annualreviews.org This
Alignment Strategies for Large Scale Genome Alignments
 Alignment Strategies for Large Scale Genome Alignments CSHL Computational Genomics 9 November 2003 Algorithms for Biological Sequence Comparison algorithm value scoring gap time calculated matrix penalty
Alignment Strategies for Large Scale Genome Alignments CSHL Computational Genomics 9 November 2003 Algorithms for Biological Sequence Comparison algorithm value scoring gap time calculated matrix penalty
Phylogenomic Resources at the UCSC Genome Browser
 9 Phylogenomic Resources at the UCSC Genome Browser Kate Rosenbloom, James Taylor, Stephen Schaeffer, Jim Kent, David Haussler, and Webb Miller Summary The UC Santa Cruz Genome Browser provides a number
9 Phylogenomic Resources at the UCSC Genome Browser Kate Rosenbloom, James Taylor, Stephen Schaeffer, Jim Kent, David Haussler, and Webb Miller Summary The UC Santa Cruz Genome Browser provides a number
Algorithms in Bioinformatics FOUR Pairwise Sequence Alignment. Pairwise Sequence Alignment. Convention: DNA Sequences 5. Sequence Alignment
 Algorithms in Bioinformatics FOUR Sami Khuri Department of Computer Science San José State University Pairwise Sequence Alignment Homology Similarity Global string alignment Local string alignment Dot
Algorithms in Bioinformatics FOUR Sami Khuri Department of Computer Science San José State University Pairwise Sequence Alignment Homology Similarity Global string alignment Local string alignment Dot
17 Non-collinear alignment Motivation A B C A B C A B C A B C D A C. This exposition is based on:
 17 Non-collinear alignment This exposition is based on: 1. Darling, A.E., Mau, B., Perna, N.T. (2010) progressivemauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS One 5(6):e11147.
17 Non-collinear alignment This exposition is based on: 1. Darling, A.E., Mau, B., Perna, N.T. (2010) progressivemauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS One 5(6):e11147.
Sequence analysis and comparison
 The aim with sequence identification: Sequence analysis and comparison Marjolein Thunnissen Lund September 2012 Is there any known protein sequence that is homologous to mine? Are there any other species
The aim with sequence identification: Sequence analysis and comparison Marjolein Thunnissen Lund September 2012 Is there any known protein sequence that is homologous to mine? Are there any other species
Small RNA in rice genome
 Vol. 45 No. 5 SCIENCE IN CHINA (Series C) October 2002 Small RNA in rice genome WANG Kai ( 1, ZHU Xiaopeng ( 2, ZHONG Lan ( 1,3 & CHEN Runsheng ( 1,2 1. Beijing Genomics Institute/Center of Genomics and
Vol. 45 No. 5 SCIENCE IN CHINA (Series C) October 2002 Small RNA in rice genome WANG Kai ( 1, ZHU Xiaopeng ( 2, ZHONG Lan ( 1,3 & CHEN Runsheng ( 1,2 1. Beijing Genomics Institute/Center of Genomics and
NIH Public Access Author Manuscript Pac Symp Biocomput. Author manuscript; available in PMC 2009 October 6.
 NIH Public Access Author Manuscript Published in final edited form as: Pac Symp Biocomput. 2009 ; : 162 173. SIMULTANEOUS HISTORY RECONSTRUCTION FOR COMPLEX GENE CLUSTERS IN MULTIPLE SPECIES * Yu Zhang,
NIH Public Access Author Manuscript Published in final edited form as: Pac Symp Biocomput. 2009 ; : 162 173. SIMULTANEOUS HISTORY RECONSTRUCTION FOR COMPLEX GENE CLUSTERS IN MULTIPLE SPECIES * Yu Zhang,
Early History up to Schedule. Proteins DNA & RNA Schwann and Schleiden Cell Theory Charles Darwin publishes Origin of Species
 Schedule Bioinformatics and Computational Biology: History and Biological Background (JH) 0.0 he Parsimony criterion GKN.0 Stochastic Models of Sequence Evolution GKN 7.0 he Likelihood criterion GKN 0.0
Schedule Bioinformatics and Computational Biology: History and Biological Background (JH) 0.0 he Parsimony criterion GKN.0 Stochastic Models of Sequence Evolution GKN 7.0 he Likelihood criterion GKN 0.0
Bio 1B Lecture Outline (please print and bring along) Fall, 2007
 Bio 1B Lecture Outline (please print and bring along) Fall, 2007 B.D. Mishler, Dept. of Integrative Biology 2-6810, bmishler@berkeley.edu Evolution lecture #5 -- Molecular genetics and molecular evolution
Bio 1B Lecture Outline (please print and bring along) Fall, 2007 B.D. Mishler, Dept. of Integrative Biology 2-6810, bmishler@berkeley.edu Evolution lecture #5 -- Molecular genetics and molecular evolution
Information Theoretic Distance Measures in Phylogenomics
 Information Theoretic Distance Measures in Phylogenomics Pavol Hanus, Janis Dingel, Juergen Zech, Joachim Hagenauer and Jakob C. Mueller Institute for Communications Engineering Technical University, 829
Information Theoretic Distance Measures in Phylogenomics Pavol Hanus, Janis Dingel, Juergen Zech, Joachim Hagenauer and Jakob C. Mueller Institute for Communications Engineering Technical University, 829
10-810: Advanced Algorithms and Models for Computational Biology. microrna and Whole Genome Comparison
 10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
Computational Identification of Evolutionarily Conserved Exons
 Computational Identification of Evolutionarily Conserved Exons Adam Siepel Center for Biomolecular Science and Engr. University of California Santa Cruz, CA 95064, USA acs@soe.ucsc.edu David Haussler Howard
Computational Identification of Evolutionarily Conserved Exons Adam Siepel Center for Biomolecular Science and Engr. University of California Santa Cruz, CA 95064, USA acs@soe.ucsc.edu David Haussler Howard
Supplementary Material
 Supplementary Material 1 Sequence Data and Multiple Alignments The five vertebrate, four insect, two worm, and seven yeast genomes used in the analysis are summarized in Table S1, and the four genome-wide
Supplementary Material 1 Sequence Data and Multiple Alignments The five vertebrate, four insect, two worm, and seven yeast genomes used in the analysis are summarized in Table S1, and the four genome-wide
Effective Cluster-Based Seed Design for Cross-Species Sequence Comparisons
 Effective Cluster-Based Seed Design for Cross-Species Sequence Comparisons Leming Zhou and Liliana Florea 1 Methods Supplementary Materials 1.1 Cluster-based seed design 1. Determine Homologous Genes.
Effective Cluster-Based Seed Design for Cross-Species Sequence Comparisons Leming Zhou and Liliana Florea 1 Methods Supplementary Materials 1.1 Cluster-based seed design 1. Determine Homologous Genes.
Chromosomal rearrangements in mammalian genomes : characterising the breakpoints. Claire Lemaitre
 PhD defense Chromosomal rearrangements in mammalian genomes : characterising the breakpoints Claire Lemaitre Laboratoire de Biométrie et Biologie Évolutive Université Claude Bernard Lyon 1 6 novembre 2008
PhD defense Chromosomal rearrangements in mammalian genomes : characterising the breakpoints Claire Lemaitre Laboratoire de Biométrie et Biologie Évolutive Université Claude Bernard Lyon 1 6 novembre 2008
Benchmarking tools for the alignment of functional
 Benchmarking tools for the alignment of functional noncoding DNA. Daniel A. Pollard (dpollard@socrates.berkeley.edu) 1, Casey M. Bergman (cbergman@gen.cam.ac.uk) 2,3,,*, Jens Stoye (stoye@techfak.uni-bielefeld.de)
Benchmarking tools for the alignment of functional noncoding DNA. Daniel A. Pollard (dpollard@socrates.berkeley.edu) 1, Casey M. Bergman (cbergman@gen.cam.ac.uk) 2,3,,*, Jens Stoye (stoye@techfak.uni-bielefeld.de)
Cladistics and Bioinformatics Questions 2013
 AP Biology Name Cladistics and Bioinformatics Questions 2013 1. The following table shows the percentage similarity in sequences of nucleotides from a homologous gene derived from five different species
AP Biology Name Cladistics and Bioinformatics Questions 2013 1. The following table shows the percentage similarity in sequences of nucleotides from a homologous gene derived from five different species
Multiple Whole Genome Alignment
 Multiple Whole Genome Alignment BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 206 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by
Multiple Whole Genome Alignment BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 206 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by
HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM
 I529: Machine Learning in Bioinformatics (Spring 2017) HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington
I529: Machine Learning in Bioinformatics (Spring 2017) HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington
I519 Introduction to Bioinformatics, Genome Comparison. Yuzhen Ye School of Informatics & Computing, IUB
 I519 Introduction to Bioinformatics, 2015 Genome Comparison Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Whole genome comparison/alignment Build better phylogenies Identify polymorphism
I519 Introduction to Bioinformatics, 2015 Genome Comparison Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Whole genome comparison/alignment Build better phylogenies Identify polymorphism
Sequence Alignment: A General Overview. COMP Fall 2010 Luay Nakhleh, Rice University
 Sequence Alignment: A General Overview COMP 571 - Fall 2010 Luay Nakhleh, Rice University Life through Evolution All living organisms are related to each other through evolution This means: any pair of
Sequence Alignment: A General Overview COMP 571 - Fall 2010 Luay Nakhleh, Rice University Life through Evolution All living organisms are related to each other through evolution This means: any pair of
Evolution s cauldron: Duplication, deletion, and rearrangement in the mouse and human genomes
 Evolution s cauldron: Duplication, deletion, and rearrangement in the mouse and human genomes W. James Kent*, Robert Baertsch*, Angie Hinrichs*, Webb Miller, and David Haussler *Center for Biomolecular
Evolution s cauldron: Duplication, deletion, and rearrangement in the mouse and human genomes W. James Kent*, Robert Baertsch*, Angie Hinrichs*, Webb Miller, and David Haussler *Center for Biomolecular
Homology Modeling. Roberto Lins EPFL - summer semester 2005
 Homology Modeling Roberto Lins EPFL - summer semester 2005 Disclaimer: course material is mainly taken from: P.E. Bourne & H Weissig, Structural Bioinformatics; C.A. Orengo, D.T. Jones & J.M. Thornton,
Homology Modeling Roberto Lins EPFL - summer semester 2005 Disclaimer: course material is mainly taken from: P.E. Bourne & H Weissig, Structural Bioinformatics; C.A. Orengo, D.T. Jones & J.M. Thornton,
Motivating the need for optimal sequence alignments...
 1 Motivating the need for optimal sequence alignments... 2 3 Note that this actually combines two objectives of optimal sequence alignments: (i) use the score of the alignment o infer homology; (ii) use
1 Motivating the need for optimal sequence alignments... 2 3 Note that this actually combines two objectives of optimal sequence alignments: (i) use the score of the alignment o infer homology; (ii) use
3/1/17. Content. TWINSCAN model. Example. TWINSCAN algorithm. HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM
 I529: Machine Learning in Bioinformatics (Spring 2017) Content HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM Yuzhen Ye School of Informatics and Computing Indiana University,
I529: Machine Learning in Bioinformatics (Spring 2017) Content HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM Yuzhen Ye School of Informatics and Computing Indiana University,
One of most striking discoveries to arise from comparative
 A large family of ancient repeat elements in the human genome is under strong selection Michael Kamal*, Xiaohui Xie*, and Eric S. Lander* *Broad Institute of Massachusetts Institute of Technology and Harvard
A large family of ancient repeat elements in the human genome is under strong selection Michael Kamal*, Xiaohui Xie*, and Eric S. Lander* *Broad Institute of Massachusetts Institute of Technology and Harvard
Kernels for gene regulatory regions
 Kernels for gene regulatory regions Jean-Philippe Vert Geostatistics Center Ecole des Mines de Paris - ParisTech Jean-Philippe.Vert@ensmp.fr Robert Thurman Division of Medical Genetics University of Washington
Kernels for gene regulatory regions Jean-Philippe Vert Geostatistics Center Ecole des Mines de Paris - ParisTech Jean-Philippe.Vert@ensmp.fr Robert Thurman Division of Medical Genetics University of Washington
BMI/CS 776 Lecture #20 Alignment of whole genomes. Colin Dewey (with slides adapted from those by Mark Craven)
 BMI/CS 776 Lecture #20 Alignment of whole genomes Colin Dewey (with slides adapted from those by Mark Craven) 2007.03.29 1 Multiple whole genome alignment Input set of whole genome sequences genomes diverged
BMI/CS 776 Lecture #20 Alignment of whole genomes Colin Dewey (with slides adapted from those by Mark Craven) 2007.03.29 1 Multiple whole genome alignment Input set of whole genome sequences genomes diverged
InDel 3-5. InDel 8-9. InDel 3-5. InDel 8-9. InDel InDel 8-9
 Lecture 5 Alignment I. Introduction. For sequence data, the process of generating an alignment establishes positional homologies; that is, alignment provides the identification of homologous phylogenetic
Lecture 5 Alignment I. Introduction. For sequence data, the process of generating an alignment establishes positional homologies; that is, alignment provides the identification of homologous phylogenetic
The Phylo- HMM approach to problems in comparative genomics, with examples.
 The Phylo- HMM approach to problems in comparative genomics, with examples. Keith Bettinger Introduction The theory of evolution explains the diversity of organisms on Earth by positing that earlier species
The Phylo- HMM approach to problems in comparative genomics, with examples. Keith Bettinger Introduction The theory of evolution explains the diversity of organisms on Earth by positing that earlier species
Lecture 14: Multiple Sequence Alignment (Gene Finding, Conserved Elements) Scribe: John Ekins
 Lecture 14: Multiple Sequence Alignment (Gene Finding, Conserved Elements) 2 19 2015 Scribe: John Ekins Multiple Sequence Alignment Given N sequences x 1, x 2,, x N : Insert gaps in each of the sequences
Lecture 14: Multiple Sequence Alignment (Gene Finding, Conserved Elements) 2 19 2015 Scribe: John Ekins Multiple Sequence Alignment Given N sequences x 1, x 2,, x N : Insert gaps in each of the sequences
Module: Sequence Alignment Theory and Applications Session: Introduction to Searching and Sequence Alignment
 Module: Sequence Alignment Theory and Applications Session: Introduction to Searching and Sequence Alignment Introduction to Bioinformatics online course : IBT Jonathan Kayondo Learning Objectives Understand
Module: Sequence Alignment Theory and Applications Session: Introduction to Searching and Sequence Alignment Introduction to Bioinformatics online course : IBT Jonathan Kayondo Learning Objectives Understand
Evolution and functional classification of vertebrate gene deserts
 Chicken Special/Letter Evolution and functional classification of vertebrate gene deserts Ivan Ovcharenko, 1,7 Gabriela G. Loots, 2 Marcelo A. Nobrega, 3 Ross C. Hardison, 4 Webb Miller, 5,6 and Lisa Stubbs
Chicken Special/Letter Evolution and functional classification of vertebrate gene deserts Ivan Ovcharenko, 1,7 Gabriela G. Loots, 2 Marcelo A. Nobrega, 3 Ross C. Hardison, 4 Webb Miller, 5,6 and Lisa Stubbs
A Browser for Pig Genome Data
 A Browser for Pig Genome Data Thomas Mailund January 2, 2004 This report briefly describe the blast and alignment data available at http://www.daimi.au.dk/ mailund/pig-genome/ hits.html. The report describes
A Browser for Pig Genome Data Thomas Mailund January 2, 2004 This report briefly describe the blast and alignment data available at http://www.daimi.au.dk/ mailund/pig-genome/ hits.html. The report describes
Phylogenetics in the Age of Genomics: Prospects and Challenges
 Phylogenetics in the Age of Genomics: Prospects and Challenges Antonis Rokas Department of Biological Sciences, Vanderbilt University http://as.vanderbilt.edu/rokaslab http://pubmed2wordle.appspot.com/
Phylogenetics in the Age of Genomics: Prospects and Challenges Antonis Rokas Department of Biological Sciences, Vanderbilt University http://as.vanderbilt.edu/rokaslab http://pubmed2wordle.appspot.com/
Conservation of Human Microsatellites across 450 Million Years of Evolution
 Conservation of Human Microsatellites across 450 Million Years of Evolution Emmanuel Buschiazzo*,1,2 and Neil J. Gemmell 1,3 1 School of Biological Sciences, University of Canterbury, Christchurch, New
Conservation of Human Microsatellites across 450 Million Years of Evolution Emmanuel Buschiazzo*,1,2 and Neil J. Gemmell 1,3 1 School of Biological Sciences, University of Canterbury, Christchurch, New
Effects of Gap Open and Gap Extension Penalties
 Brigham Young University BYU ScholarsArchive All Faculty Publications 200-10-01 Effects of Gap Open and Gap Extension Penalties Hyrum Carroll hyrumcarroll@gmail.com Mark J. Clement clement@cs.byu.edu See
Brigham Young University BYU ScholarsArchive All Faculty Publications 200-10-01 Effects of Gap Open and Gap Extension Penalties Hyrum Carroll hyrumcarroll@gmail.com Mark J. Clement clement@cs.byu.edu See
Outline. Genome Evolution. Genome. Genome Architecture. Constraints on Genome Evolution. New Evolutionary Synthesis 11/8/16
 Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Comparative Gene Finding. BMI/CS 776 Spring 2015 Colin Dewey
 Comparative Gene Finding BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2015 Colin Dewey cdewey@biostat.wisc.edu Goals for Lecture the key concepts to understand are the following: using related genomes
Comparative Gene Finding BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2015 Colin Dewey cdewey@biostat.wisc.edu Goals for Lecture the key concepts to understand are the following: using related genomes
8.2% of the Human Genome Is Constrained: Variation in Rates of Turnover across Functional Element Classes in the Human Lineage
 8.2% of the Human Genome Is Constrained: Variation in Rates of Turnover across Functional Element Classes in the Human Lineage Chris M. Rands 1, Stephen Meader 1, Chris P. Ponting 1 *, Gerton Lunter 2
8.2% of the Human Genome Is Constrained: Variation in Rates of Turnover across Functional Element Classes in the Human Lineage Chris M. Rands 1, Stephen Meader 1, Chris P. Ponting 1 *, Gerton Lunter 2
Drosophila melanogaster and D. simulans, two fruit fly species that are nearly
 Comparative Genomics: Human versus chimpanzee 1. Introduction The chimpanzee is the closest living relative to humans. The two species are nearly identical in DNA sequence (>98% identity), yet vastly different
Comparative Genomics: Human versus chimpanzee 1. Introduction The chimpanzee is the closest living relative to humans. The two species are nearly identical in DNA sequence (>98% identity), yet vastly different
Comparative genomics. Lucy Skrabanek ICB, WMC 6 May 2008
 Comparative genomics Lucy Skrabanek ICB, WMC 6 May 2008 What does it encompass? Genome conservation transfer knowledge gained from model organisms to non-model organisms Genome evolution understand how
Comparative genomics Lucy Skrabanek ICB, WMC 6 May 2008 What does it encompass? Genome conservation transfer knowledge gained from model organisms to non-model organisms Genome evolution understand how
Graph Alignment and Biological Networks
 Graph Alignment and Biological Networks Johannes Berg http://www.uni-koeln.de/ berg Institute for Theoretical Physics University of Cologne Germany p.1/12 Networks in molecular biology New large-scale
Graph Alignment and Biological Networks Johannes Berg http://www.uni-koeln.de/ berg Institute for Theoretical Physics University of Cologne Germany p.1/12 Networks in molecular biology New large-scale
Bioinformatics Exercises
 Bioinformatics Exercises AP Biology Teachers Workshop Susan Cates, Ph.D. Evolution of Species Phylogenetic Trees show the relatedness of organisms Common Ancestor (Root of the tree) 1 Rooted vs. Unrooted
Bioinformatics Exercises AP Biology Teachers Workshop Susan Cates, Ph.D. Evolution of Species Phylogenetic Trees show the relatedness of organisms Common Ancestor (Root of the tree) 1 Rooted vs. Unrooted
Sequence Alignment Techniques and Their Uses
 Sequence Alignment Techniques and Their Uses Sarah Fiorentino Since rapid sequencing technology and whole genomes sequencing, the amount of sequence information has grown exponentially. With all of this
Sequence Alignment Techniques and Their Uses Sarah Fiorentino Since rapid sequencing technology and whole genomes sequencing, the amount of sequence information has grown exponentially. With all of this
Outline. Genome Evolution. Genome. Genome Architecture. Constraints on Genome Evolution. New Evolutionary Synthesis 11/1/18
 Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Alignment. Peak Detection
 ChIP seq ChIP Seq Hongkai Ji et al. Nature Biotechnology 26: 1293-1300. 2008 ChIP Seq Analysis Alignment Peak Detection Annotation Visualization Sequence Analysis Motif Analysis Alignment ELAND Bowtie
ChIP seq ChIP Seq Hongkai Ji et al. Nature Biotechnology 26: 1293-1300. 2008 ChIP Seq Analysis Alignment Peak Detection Annotation Visualization Sequence Analysis Motif Analysis Alignment ELAND Bowtie
Supplemental Data. Perea-Resa et al. Plant Cell. (2012) /tpc
 Supplemental Data. Perea-Resa et al. Plant Cell. (22)..5/tpc.2.3697 Sm Sm2 Supplemental Figure. Sequence alignment of Arabidopsis LSM proteins. Alignment of the eleven Arabidopsis LSM proteins. Sm and
Supplemental Data. Perea-Resa et al. Plant Cell. (22)..5/tpc.2.3697 Sm Sm2 Supplemental Figure. Sequence alignment of Arabidopsis LSM proteins. Alignment of the eleven Arabidopsis LSM proteins. Sm and
CONCEPT OF SEQUENCE COMPARISON. Natapol Pornputtapong 18 January 2018
 CONCEPT OF SEQUENCE COMPARISON Natapol Pornputtapong 18 January 2018 SEQUENCE ANALYSIS - A ROSETTA STONE OF LIFE Sequence analysis is the process of subjecting a DNA, RNA or peptide sequence to any of
CONCEPT OF SEQUENCE COMPARISON Natapol Pornputtapong 18 January 2018 SEQUENCE ANALYSIS - A ROSETTA STONE OF LIFE Sequence analysis is the process of subjecting a DNA, RNA or peptide sequence to any of
METHODS FOR DETERMINING PHYLOGENY. In Chapter 11, we discovered that classifying organisms into groups was, and still is, a difficult task.
 Chapter 12 (Strikberger) Molecular Phylogenies and Evolution METHODS FOR DETERMINING PHYLOGENY In Chapter 11, we discovered that classifying organisms into groups was, and still is, a difficult task. Modern
Chapter 12 (Strikberger) Molecular Phylogenies and Evolution METHODS FOR DETERMINING PHYLOGENY In Chapter 11, we discovered that classifying organisms into groups was, and still is, a difficult task. Modern
Supporting Information
 Supporting Information Weghorn and Lässig 10.1073/pnas.1210887110 SI Text Null Distributions of Nucleosome Affinity and of Regulatory Site Content. Our inference of selection is based on a comparison of
Supporting Information Weghorn and Lässig 10.1073/pnas.1210887110 SI Text Null Distributions of Nucleosome Affinity and of Regulatory Site Content. Our inference of selection is based on a comparison of
Understanding relationship between homologous sequences
 Molecular Evolution Molecular Evolution How and when were genes and proteins created? How old is a gene? How can we calculate the age of a gene? How did the gene evolve to the present form? What selective
Molecular Evolution Molecular Evolution How and when were genes and proteins created? How old is a gene? How can we calculate the age of a gene? How did the gene evolve to the present form? What selective
Conserved noncoding elements (CNEs) represent 3.5% of
 A family of conserved noncoding elements derived from an ancient transposable element Xiaohui Xie*, Michael Kamal*, and Eric S. Lander* *Broad Institute of Massachusetts Institute of Technology and Harvard
A family of conserved noncoding elements derived from an ancient transposable element Xiaohui Xie*, Michael Kamal*, and Eric S. Lander* *Broad Institute of Massachusetts Institute of Technology and Harvard
O 3 O 4 O 5. q 3. q 4. Transition
 Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Practical considerations of working with sequencing data
 Practical considerations of working with sequencing data File Types Fastq ->aligner -> reference(genome) coordinates Coordinate files SAM/BAM most complete, contains all of the info in fastq and more!
Practical considerations of working with sequencing data File Types Fastq ->aligner -> reference(genome) coordinates Coordinate files SAM/BAM most complete, contains all of the info in fastq and more!
C3020 Molecular Evolution. Exercises #3: Phylogenetics
 C3020 Molecular Evolution Exercises #3: Phylogenetics Consider the following sequences for five taxa 1-5 and the known outgroup O, which has the ancestral states (note that sequence 3 has changed from
C3020 Molecular Evolution Exercises #3: Phylogenetics Consider the following sequences for five taxa 1-5 and the known outgroup O, which has the ancestral states (note that sequence 3 has changed from
Multiple Sequence Alignment, Gunnar Klau, December 9, 2005, 17:
 Multiple Sequence Alignment, Gunnar Klau, December 9, 2005, 17:50 5001 5 Multiple Sequence Alignment The first part of this exposition is based on the following sources, which are recommended reading:
Multiple Sequence Alignment, Gunnar Klau, December 9, 2005, 17:50 5001 5 Multiple Sequence Alignment The first part of this exposition is based on the following sources, which are recommended reading:
Abstract. comment reviews reports deposited research refereed research interactions information
 http://genomebiology.com/2002/3/12/research/0086.1 Research Assessing the impact of comparative genomic sequence data on the functional annotation of the Drosophila genome Casey M Bergman*, Barret D Pfeiffer*,
http://genomebiology.com/2002/3/12/research/0086.1 Research Assessing the impact of comparative genomic sequence data on the functional annotation of the Drosophila genome Casey M Bergman*, Barret D Pfeiffer*,
How to usefully compare homologous plant genes and chromosomes as DNA sequences
 The Plant Journal (2008) 53, 661 673 doi: 10.1111/j.1365-313X.2007.03326.x TECHNIQUES FOR MOLECULAR ANALYSIS How to usefully compare homologous plant genes and chromosomes as DNA sequences Eric Lyons *
The Plant Journal (2008) 53, 661 673 doi: 10.1111/j.1365-313X.2007.03326.x TECHNIQUES FOR MOLECULAR ANALYSIS How to usefully compare homologous plant genes and chromosomes as DNA sequences Eric Lyons *
Bustamante et al., Supplementary Nature Manuscript # 1 out of 9 Information #
 Bustamante et al., Supplementary Nature Manuscript # 1 out of 9 Details of PRF Methodology In the Poisson Random Field PRF) model, it is assumed that non-synonymous mutations at a given gene are either
Bustamante et al., Supplementary Nature Manuscript # 1 out of 9 Details of PRF Methodology In the Poisson Random Field PRF) model, it is assumed that non-synonymous mutations at a given gene are either
Lecture 4: Evolutionary Models and Substitution Matrices (PAM and BLOSUM)
 Bioinformatics II Probability and Statistics Universität Zürich and ETH Zürich Spring Semester 2009 Lecture 4: Evolutionary Models and Substitution Matrices (PAM and BLOSUM) Dr Fraser Daly adapted from
Bioinformatics II Probability and Statistics Universität Zürich and ETH Zürich Spring Semester 2009 Lecture 4: Evolutionary Models and Substitution Matrices (PAM and BLOSUM) Dr Fraser Daly adapted from
Comparison of Cost Functions in Sequence Alignment. Ryan Healey
 Comparison of Cost Functions in Sequence Alignment Ryan Healey Use of Cost Functions Used to score and align sequences Mathematically model how sequences mutate and evolve. Evolution and mutation can be
Comparison of Cost Functions in Sequence Alignment Ryan Healey Use of Cost Functions Used to score and align sequences Mathematically model how sequences mutate and evolve. Evolution and mutation can be
Some Problems from Enzyme Families
 Some Problems from Enzyme Families Greg Butler Department of Computer Science Concordia University, Montreal www.cs.concordia.ca/~faculty/gregb gregb@cs.concordia.ca Abstract I will discuss some problems
Some Problems from Enzyme Families Greg Butler Department of Computer Science Concordia University, Montreal www.cs.concordia.ca/~faculty/gregb gregb@cs.concordia.ca Abstract I will discuss some problems
Dr. Amira A. AL-Hosary
 Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Sequence Database Search Techniques I: Blast and PatternHunter tools
 Sequence Database Search Techniques I: Blast and PatternHunter tools Zhang Louxin National University of Singapore Outline. Database search 2. BLAST (and filtration technique) 3. PatternHunter (empowered
Sequence Database Search Techniques I: Blast and PatternHunter tools Zhang Louxin National University of Singapore Outline. Database search 2. BLAST (and filtration technique) 3. PatternHunter (empowered
A bioinformatics approach to the structural and functional analysis of the glycogen phosphorylase protein family
 A bioinformatics approach to the structural and functional analysis of the glycogen phosphorylase protein family Jieming Shen 1,2 and Hugh B. Nicholas, Jr. 3 1 Bioengineering and Bioinformatics Summer
A bioinformatics approach to the structural and functional analysis of the glycogen phosphorylase protein family Jieming Shen 1,2 and Hugh B. Nicholas, Jr. 3 1 Bioengineering and Bioinformatics Summer
Phylogenetic inference
 Phylogenetic inference Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, March 7 th 016 After this lecture, you can discuss (dis-) advantages of different information types
Phylogenetic inference Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, March 7 th 016 After this lecture, you can discuss (dis-) advantages of different information types
Biological Networks: Comparison, Conservation, and Evolution via Relative Description Length By: Tamir Tuller & Benny Chor
 Biological Networks:,, and via Relative Description Length By: Tamir Tuller & Benny Chor Presented by: Noga Grebla Content of the presentation Presenting the goals of the research Reviewing basic terms
Biological Networks:,, and via Relative Description Length By: Tamir Tuller & Benny Chor Presented by: Noga Grebla Content of the presentation Presenting the goals of the research Reviewing basic terms
Overview Multiple Sequence Alignment
 Overview Multiple Sequence Alignment Inge Jonassen Bioinformatics group Dept. of Informatics, UoB Inge.Jonassen@ii.uib.no Definition/examples Use of alignments The alignment problem scoring alignments
Overview Multiple Sequence Alignment Inge Jonassen Bioinformatics group Dept. of Informatics, UoB Inge.Jonassen@ii.uib.no Definition/examples Use of alignments The alignment problem scoring alignments
Introduction to Bioinformatics Online Course: IBT
 Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec1 Building a Multiple Sequence Alignment Learning Outcomes 1- Understanding Why multiple
Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec1 Building a Multiple Sequence Alignment Learning Outcomes 1- Understanding Why multiple
Research Proposal. Title: Multiple Sequence Alignment used to investigate the co-evolving positions in OxyR Protein family.
 Research Proposal Title: Multiple Sequence Alignment used to investigate the co-evolving positions in OxyR Protein family. Name: Minjal Pancholi Howard University Washington, DC. June 19, 2009 Research
Research Proposal Title: Multiple Sequence Alignment used to investigate the co-evolving positions in OxyR Protein family. Name: Minjal Pancholi Howard University Washington, DC. June 19, 2009 Research
Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p
 Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p.110-114 Arrangement of information in DNA----- requirements for RNA Common arrangement of protein-coding genes in prokaryotes=
Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p.110-114 Arrangement of information in DNA----- requirements for RNA Common arrangement of protein-coding genes in prokaryotes=
I519 Introduction to Bioinformatics, Genome Comparison. Yuzhen Ye School of Informatics & Computing, IUB
 I519 Introduction to Bioinformatics, 2011 Genome Comparison Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Whole genome comparison/alignment Build better phylogenies Identify polymorphism
I519 Introduction to Bioinformatics, 2011 Genome Comparison Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Whole genome comparison/alignment Build better phylogenies Identify polymorphism
Quantifying sequence similarity
 Quantifying sequence similarity Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 16 th 2016 After this lecture, you can define homology, similarity, and identity
Quantifying sequence similarity Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 16 th 2016 After this lecture, you can define homology, similarity, and identity
3. SEQUENCE ANALYSIS BIOINFORMATICS COURSE MTAT
 3. SEQUENCE ANALYSIS BIOINFORMATICS COURSE MTAT.03.239 25.09.2012 SEQUENCE ANALYSIS IS IMPORTANT FOR... Prediction of function Gene finding the process of identifying the regions of genomic DNA that encode
3. SEQUENCE ANALYSIS BIOINFORMATICS COURSE MTAT.03.239 25.09.2012 SEQUENCE ANALYSIS IS IMPORTANT FOR... Prediction of function Gene finding the process of identifying the regions of genomic DNA that encode
In-Depth Assessment of Local Sequence Alignment
 2012 International Conference on Environment Science and Engieering IPCBEE vol.3 2(2012) (2012)IACSIT Press, Singapoore In-Depth Assessment of Local Sequence Alignment Atoosa Ghahremani and Mahmood A.
2012 International Conference on Environment Science and Engieering IPCBEE vol.3 2(2012) (2012)IACSIT Press, Singapoore In-Depth Assessment of Local Sequence Alignment Atoosa Ghahremani and Mahmood A.
