A binding site for Gli proteins is essential for HNF-3β floor plate enhancer activity in transgenics and can respond to Shh in vitro
|
|
|
- Sibyl Day
- 6 years ago
- Views:
Transcription
1 Development 124, (1997) Printed in Great Britain The Company of Biologists Limited 1997 DEV A binding site for Gli proteins is essential for HNF-3β floor plate enhancer activity in transgenics and can respond to Shh in vitro Hiroshi Sasaki 1, *, Chi-chung Hui 2, Masato Nakafuku 3 and Hisato Kondoh 1 1 Laboratory of Developmental Biology, Institute for Molecular and Cellular Biology, Osaka University, 1-3 Yamada-oka, Suita, Osaka 565, Japan 2 Program in Developmental Biology and Division of Endocrinology, Research Institute, The Hospital for Sick Children, 555 University Avenue, Toronto, Ontario M5G 1X8, Canada 3 Division of Signal Transduction and Metabolic Regulation in Animal Cells, Graduate School of Biological Sciences, Nara Advanced Institute of Science and Technology, Ikoma, Nara , Japan *Author for correspondence ( hsasaki@imcb.osaka-u.ac.jp) SUMMARY The floor plate plays important roles in ventral pattern formation and axonal guidance within the neural tube of vertebrate embryos. A critical event for floor plate development is the induction of a winged helix transcription factor, Hepatocyte Nuclear Factor-3β (HNF-3β). The enhancer for floor plate expression of HNF-3β is located 3 of the transcription unit and consists of multiple elements. HNF-3β induction depends on the notochord-derived signal, Sonic hedgehog (Shh). Genetic analysis in Drosophila has led to the identification of genes involved in the Hh signalling pathway, and cubitus interruptus (ci), encoding a protein with five zinc finger motifs, was placed downstream. In the present work, we test the involvement of Gli proteins, the mouse homologues of Ci, in activation of the floor plate enhancer of HNF-3β. Transgenic analysis shows that a Gli-binding site is required for the activity of the minimal floor plate enhancer of HNF-3β in vivo. Three Gli genes are differentially expressed in the developing neural tube. Gli expression is restricted to the ventral part, while Gli2 and Gli3 are expressed throughout the neural tube and dorsally, respectively. Strong Gli and Gli2, and weak Gli3 expressions transiently overlap with HNF-3β at the time of its induction. Consistent with ventrally localized expression, Gli expression can be up-regulated by Shh in a cell line. Finally, the Gli-binding site acts as a Shh responsive element, and human GLI, but not GLI3, can activate this binding site in tissue culture. Taken together, these findings suggest that Gli, and probably also Gli2, are good candidates for transcriptional activators of the HNF-3β floor plate enhancer, and the binding site for Gli proteins is a key element for response to Shh signalling. These results also support the idea that Gli/Ci are evolutionary conserved transcription factors in the Hedgehog signalling pathway. Key words: Gli, HNF-3β, floor plate, Sonic hedgehog, enhancer, pattern formation, Drosophila INTRODUCTION The floor plate is a group of specialized cells located in the ventral midline of the vertebrate neural tube. It plays important roles in neural tube development as a source of signalling molecules for dorsoventral patterning and axonal guidance (for reviews, see Jessell and Dodd, 1992; Dodd and Jessell, 1993). A floor plate-derived signalling molecule, Sonic hedgehog (Shh), which is a vertebrate homologue of Drosophila Hedgehog (Hh), induces differentiation of ventral type neurons (Ericson et al., 1995; Marti et al., 1995; Roelink et al., 1995). Another floor plate-derived signalling molecule, netrin-1, works as both a chemoattractant and a chemorepellent of axons, depending on neuronal cell types (Kennedy et al., 1994; Colamarino and Tessier-Lavigne, 1995). Differentiation of the floor plate is induced by a signal derived from the notochord or by Shh. High concentrations of Shh-N peptide or contact with the notochord are required for floor plate induction in neural explants (Marti et al., 1995; Roelink et al., 1995). A critical event of floor plate development is the expression of a winged-helix transcription factor, HNF-3β (Ang et al., 1993; Ruiz i Altaba et al., 1993; Sasaki and Hogan, 1993, 1994). Ectopic expression of HNF-3β in the neural tube induces ectopic expression of series of floor plate marker genes and prevents dorsal/lateral maker expression, indicating that HNF-3β acts as a regulator of floor plate development (Sasaki and Hogan, 1994; Ruiz i Altaba et al., 1995a). The HNF-3β expression in the floor plate is induced by a signal derived from the notochord or by Shh, and its induction does not require de novo protein synthesis (Roelink et al., 1995; Ruiz i Altaba et al., 1995b; Chiang et al., 1996). This makes HNF-3β a candidate for a direct target of Shh signalling. Previously, we have identified cis-regulatory regions of the mouse HNF-3β gene in transgenic mouse embryos. Enhancers for floor plate expression and node/notochord expression were
2 1314 H. Sasaki and others Fig. 1. Schematic presentation of the previous knowledge of mouse HNF-3β floor plate enhancer. The transcription unit and its direction are indicated by boxes and an arrow, respectively. Minimal floor plate enhancer activity requires combination of a 1.5 kb A fragment and a 400 bp HaeIII fragment. Analysis using internal deletion series within the 400 bp fragment identified multiple regions involved in regulation. The regions del-3 and del-4 are required for anterior floor plate gene expression, while regions del-5 and del-6 are essential for the enhancer activity (Sasaki and Hogan, 1996). 0 5kb 10kb 15kb 20kb K K A B E Xh B Xh Xh Sl E K B EB E B Hae III B Xh XbXh Sl A Dra I B Fok I 1kb floor plate enhancer Hae III 100b del-3 required for anterior expression del-4 del-5 del-6 essential for expression B = Bam HI E = Eco RI K = Kpn I Xh = Xho I Sl = Sal I minimal floor plate enhancer identified in separate regions 3 and 5 of the transcription unit, respectively. Deletion analysis of the minimal floor plate enhancer identified multiple elements required for its activity (Fig. 1) (Sasaki and Hogan, 1996). The present study was initiated by finding a Gli-protein-binding site within one of these essential elements. GLI is a gene that was originally identified as an amplified gene in a human glioblastoma (Kinzler et al., 1987). It encodes a nuclear protein, containing five zinc finger motifs, which binds to DNA in a sequence-specific manner (Kinzler et al., 1988; Kinzler and Vogelstein, 1990). In humans, there are three GLI genes (GLI, GLI2, GLI3) (Ruppert et al., 1988), which have closely related zinc fingers, and their mouse counterparts have also been identified (Gli, Gli2, Gli3) (Walterhouse et al., 1993; Hui et al., 1994). The Drosophila homologue of Gli genes is cubitus interruptus (ci), which is involved in the Hedgehog (Hh) signalling pathway (Orenic et al., 1990). According to genetic analysis, the Hh pathway also includes transmembrane Hh receptor Patched (Ptc), a seven-transmembrane protein Smoothened (Smo), a serine-threonine kinase Fused (Fu), camp-dependent protein kinase A (PKA) and a gene costal-2 (cos-2) (Alcedo et al., 1996; Chen and Struhl, 1996; van den Heuvel and Ingham, 1996; for reviews, see Forbes et al., 1993; Perrimon, 1995). Among them, Ci was proposed to be a transcription factor that works at the most downstream point of the pathway (Forbes et al., 1993; Domínguez et al., 1996). In fact, recent experiments showed that Ci acts as a transcriptional activator and that it activates one of the target genes, ptc, through GLI-binding sites (Alexandre et al., 1996). At least a part of the Hh signalling pathway seems to be conserved between Drosophila and vertebrates, while hh homologues appear to form a multigene family in vertebrates (Echelard et al., 1993; Ekker et al., 1995a,b; Currie and Ingham, 1996). For example, it has recently been shown that vertebrate Ptc (vptc) directly binds Shh and Dhh, suggesting its involvement as a receptor (Marigo et al., 1996a; Stone et al., 1996), and that vertebrate Smo (vsmo) makes a complex with vptc, suggesting involvement of vsmo in the Hh signalling pathway (Stone et al., 1996). In addition, injection of RNAs encoding Shh, Indian hedgehog (Ihh) or PKI, a dominant-negative regulatory subunit of PKA, into zebrafish embryos results in equivalent phenotypes, indicating that PKA acts as a negative regulator of Hh signalling in vertebrates (Hammerschmidt et al., 1996). By extension of such similarities, the three Gli proteins are good candidates for downstream transcription factors of the Hh pathway in vertebrates (Hui et al., 1994). In this study, the involvement of Gli proteins in activation of the HNF-3β floor plate enhancer was tested using transgenic embryos. A Gli-binding site within an essential region of the minimal enhancer was found to be essential for the enhancer activity. Three Gli genes are differentially expressed in the developing neural tube, and transiently overlap with HNF-3β. In addition, Shh and human GLI, but not GLI3, can activate this Gli-binding site in tissue culture. The results suggest that some of the Gli proteins are involved in induction of HNF-3β by Shh as transcription activators of the floor plate enhancer, and that downstream transcription factors of the Hh signalling pathway are conserved between the mouse and the fly. MATERIALS AND METHODS Production of GST fusion proteins Glutathione S-transferase (GST) fusion proteins were produced in E. coli using pgex expression vectors. The pgex-kg (Guan and Dixon, 1991) was used for synthesis of GST. The plasmid pgex- HNF-3β (a gift from Richard O Brien, Vanderbilt University, USA), which produces a fusion protein of GST with full length HNF-3β protein, was used for GST-HNF-3β. The plasmid pgex-gli, which was constructed by cloning HincII-XbaI fragment of human GLI cdna (pgli K12) (Kinzler et al., 1988) into SmaI and XbaI sites of pgex-kg, was used for GST-GLI. For protein production, bacterial culture was induced with 0.1 mm IPTG. After 3 hours of induction, bacteria were harvested by centrifugation and disrupted by sonication with a Branson sonifier in PBS containing 1% Tween-20. The sonicate was cleared by centrifugation at 12,000 g for 10 minutes and cleared lysate was used as GST fusion protein. Gel mobility shift assay The binding reaction was set up as follows: 1 µl of cleared E. coli lysate, 0.01 pmoles of 32 P-labeled probe, 1 µg poly(di-dc) with or without 0.5 pmoles of non-radioactive competitor were combined in 10 µl of 1 binding buffer (10 mm Tris HCl, ph 7.5, 50 mm NaCl,
3 Gli proteins in the HNF-3β enhancer mm MgCl 2, 0.5 mm DTT, 4% glycerol) on ice. After incubation for 30 minutes on ice, protein-dna complexes were separated on a 6% acrylamide gel (acrylamide:bis-acrylamide = 60:1) containing 0.25 TBE (22 mm Tris-borate ph 8.3, 0.5 mm EDTA). Immunoblot analysis 0.5 µl of a cleared E. coli lysate was electrophoresed in a 12% SDS polyacrylamide gel. After transfer of separated proteins onto a nitrocellulose membrane, GST fusion proteins were detected by a combination of anti-gst antibody (Z-5) (Santa Cruz Biotechnology), protein A-HRP (Amersham) and ECL (Amersham), following the manufacturers protocols. Production of transgenic mouse embryos Mutations were introduced in the 400 bp HaeIII fragment of the enhancer by a PCR-based in vitro mutagenesis method using Pfu DNA polymerase (Stratagene). The sequence was verified using a ABI 373 DNA sequencer. Transgenes were constructed by introducing enhancer fragments upstream of the hsp68 promoter-lacz-poly(a) cassette (ASShsplacZpA vector) (Sasaki and Hogan, 1996). Transgenic embryo production, genotyping and lacz staining of embryos were performed as described (Sasaki and Hogan, 1996). Transfection assay To construct reporter plasmids, eight directly repeated copies of 3 Gli- BS or m3 Gli-BS were cloned into the BamHI site of the plasmid pδ51lucii (Kamachi and Kondoh, 1993). pjt4/shh, which expresses chick Shh cdna under the control of the CMV promoter, was a gift from Dr Sumihare Noji (Tokushima University). pjt4 was created by removing Shh cdna from pjt4/shh. psrα-gli and psrα-gli3, which express human GLI and GLI3 cdnas under the control of the SRα promoter (Takebe et al., 1988), respectively, were gifts from Dr Shun-suke Ishii (RIKEN). psrα was a gift from Dr Ryohei Sekido (Osaka University). A rat neural stem cell line MNS70 was maintained as described (Nakafuku and Nakamura, 1995; Nakagawa et al., 1996). For transfection, MNS70 cells were plated into 6 cm Primaria dish (Falcon) at a density of cells/plate 1 day before. Medium was changed 1 hour before DNA addition. The CaPO 4/DNA mixture, containing effector (3 µg), reporter (3 µg) and reference (SV-β-gal, 1 µg) plasmids, was added to the MNS70 cells and incubated for 10 hours. In the experiments shown in Fig. 5C, the CaPO 4/DNA mixture contains effector 1 (2 µg), effector 2 (2 µg), reporter (2 µg) and reference (1 µg). Cells were washed twice with PBS and re-fed with fresh medium. Cells were harvested 2 days after DNA addition. Preparation of lysates, luciferase and β-galactosidase assays were as described (Fiering et al., 1991; Kamachi and Kondoh, 1993). Luciferase activities were normalized by β-galactosidase activities. All transfection experiments were independently repeated at least twice and the results were reproducible. RT-PCR For the Gli induction study, 7 µg of effector plasmid were transfected into MNS70 cells by the CaPO 4 method, as described above. RNA was prepared 2 days after DNA addition using Ultraspec RNA (Biotex). One fifth of the RNA from each plate was subjected to reverse transcription using Ready To Go You-Prime First-Strand Beads (Pharmacia) and oligo dt primer (Pharmacia). 1 µl of the reverse transcript was used for each PCR reaction. Primers for mouse Gli (5 -CGCGAATTCATGTGTGAGCAAGAAGGTTGC-3, 5 - GCCTCTAGAAGTCGAGGACACTGGCTATAGG-3 ) (Walterhouse et al., 1993) were used for amplification, and authenticity of the products was verified by their size and Southern hybridization with a mouse Gli cdna fragment. The conditions of the PCR were 95 C for 45 seconds, 58 C for 30 seconds, 72 C for 1.5 minutes for 30 cycles, followed by 72 C for 10 minutes. The radioactivity of each band was quantitated by BAS2000 (Fuji Film). β-actin primers were as described (Nakagawa et al., 1996). In situ hybridization A partial mouse Gli cdna was cloned by RT-PCR from a mixture of total RNAs of 8.5- and 12.5-day-old mouse embryos using primers, as described above. A partial mouse Gli3 cdna was obtained by screening a mouse 8.5-day post coitum (d.p.c.) cdna library (a gift from Kathy Mahon, Baylor College of Medicine, USA) with human GLI3 cdna (Ruppert et al., 1990) as a probe. Mouse Gli2, HNF-3β cdnas were as described (Sasaki and Hogan, 1993; Hui et al., 1994). A Shh cdna was a gift from Dr Hiroshi Hamada (Osaka University). In situ hybridizations using digoxigenin-labeled RNA probes on sections were performed as described (Sasaki and Hogan, 1994). Two or three embryos were analyzed for gene expression at each stage, and similar results were obtained reproducibly. RESULTS Identification of a Gli-binding site within an essential region of the minimal HNF-3β floor plate enhancer Previously, we identified a floor plate-specific enhancer of HNF-3β downstream of the transcription unit. The genetic elements responsible for the enhancer activity were refined to a combination of a 1.5 kb A fragment and a 0.4 kb HaeIII fragment (Fig. 1) (Sasaki and Hogan, 1996). By making an internal deletion series within the 0.4 kb fragment, two regions (del-5 and del-6) were found to be essential (Sasaki and Hogan, 1996). Therefore, it is possible that one of these regions contains an element responsive to the Shh signal from the notochord. Within one of the essential regions, del-5, a sequence motif (3 Gli-BS: 5 -GAACACCCA-3 ) that resembles a known consensus sequence of human GLI-binding sites (hgli-bs: 5 - GACCACCCA-3 ; different base underlined) was found (Fig. 2A) (Kinzler and Vogelstein, 1990). To test if the 3 Gli-BS motif is actually bound by Gli, a gel mobility shift DNAbinding assay was performed (Fig. 2B). Bacterial lysate containing a fusion protein of glutathione-s-transferase and human GLI (GST-GLI) was used as a GLI protein. Bacterial lysates containing either GST or GST-HNF-3β were used as negative controls. Each lysate contains approximately equal amounts of GST fusion proteins, as shown by immunoblotting (Fig. 2C). Only GST-GLI bound to the 3 Gli-BS probe as well as the hgli-bs probe in a specific manner (Fig. 2B lanes 3, 4, 7, 8, 10). Sequence alteration within the 3 Gli-BS motif (m3 Gli- BS: 5 -GAAGTGGGA-3 : modified bases are underlined) abolished GST-GLI binding in a competition assay, confirming that the fusion protein bound to this motif within the probe (Fig. 2B, lane 9). The apparent binding affinity of GST-GLI to the 3 Gli-BS is slightly lower than that of the hgli-bs, probably because 3 Gli-BS deviates from the consensus sequence (hgli-bs) (Fig. 2B and data not shown). These results suggest that Gli protein can bind to 3 Gli-BS in a specific manner. Considering that GLI and GLI3 have similar DNA-binding activities (Vortkamp et al., 1995) and that all three Gli proteins have very similar DNA-binding domains (Hui et al., 1994), it is likely that other Gli proteins can also bind to the 3 Gli-BS. Because of this, the phrase Gli-binding site will be used with the meaning of generic Gli-binding site throughout this paper. Since Gli genes are mouse homologues of Drosophila ci, and since Ci is a downstream transcription factor of the Hh signalling pathway, it is possible that this Glibinding site mediates the responsiveness of HNF-3β to a Shh signal derived from the notochord.
4 1316 H. Sasaki and others A B C hgli-bs TCGACTCCCGAAGACCACCCACAATGATGGTTC (A1) GAGGGCTTCTGGTGGGTGTTACTACCAAGAGCT del-5 TTATGACGGAGGCTAACAAGCAGGGAACACCCAAGTAGAAGCTGGCTGTC 3'Gli-BS m3'gli-bs TCGACAAGCAGGGAACACCCAAGTAGAAGCTC GTTCGTCCCTTGTGGGTTCATCTTCGAGAGCT TCGACAAGCAGGGAAgtgggAAGTAGAAGCTC GTTCGTCCCTTcacccTTCATCTTCGAGAGCT Fig. 2. Identification of a Gli-binding site within an essential region of the HNF-3β minimal floor plate enhancer. (A) Sequences of the oligonucleotides used in this study. A human GLI-binding site (hgli-bs) was adapted from A1 of the reference (Kinzler and Vogelstein, 1990). 3 Gli-BS was adapted from one of the essential regions, del-5. Gli-binding sites are underlined. Mutated bases in m3 Gli-BS are shown in lower case. (B) GLI protein binds to 3 Gli- BS in vitro (arrow). Combinations of probe, competitor and protein are indicated above each lane. (C) Western blot analysis of GST fusion protein production. The presence of multiple bands and their smaller sizes compared to the expected sizes (dots) indicate that the proteins are partially degraded. These degradations did not affect DNA-binding activities (B and data not shown). A 0 5kb 10kb 15kb 20kb K K B E Xh B Xh Xh Sl E K B EB E B B = Bam HI E = Eco RI K = Kpn I Xh = Xho I Sl = Sal I B Xh XbXh Sl A B 1kb Tg lacz floor plate wild type A Hae III Dra I Fok I Hae III 100b m3'gli-bs A Fig. 3. Gli-binding site is required for minimal floor plate enhancer activity. (A) Summary of a transgenic mouse experiment. Tg, number of transgenic embryos identified by PCR of yolk sac DNAs; lacz, number of embryos with lacz staining; floor plate, number of embryos with lacz staining in the floor plate. (B,C) Transgenic embryos with the wild-type minimal enhancer express β- galactosidase in the floor plate with some lateral ectopic expression (B,C and data not shown). (D) On the other hand, transgenic embryos with mutant enhancer having base alterations of the 3 Gli- BS into m3 Gli-BS show either no β-galactosidase expression (D and data not shown) or unrelated expression, probably caused by positional effects (data not shown).
5 Gli proteins in the HNF-3β enhancer 1317 The Gli-binding site is essential for the floor plate enhancer activity The significance of the 3 Gli-BS in HNF-3β regulation in vivo was investigated with transgenic mouse embryos. The sequence of the 3 Gli-BS in the minimal enhancer was changed into the m3 Gli-BS so as to abolish binding of Gli proteins (Fig. 3A), and transgenic embryos were produced using the mutated enhancer. With the wild-type enhancer, three out of four transgenic embryos showed gene expression in the floor plate (Fig. 3A-C). On the contrary, with the mutant enhancer, none out of 11 transgenic embryos showed β-galactosidase expression in the floor plate (Fig. 3A,D). Transgenic embryos showed either no β-galactosidase expression (Fig. 3D and data not shown) or expression in unrelated sites, probably caused by integration site position effects (data not shown). The gene expression patterns observed with the mutant enhancer were different among the transgenic embryos. The results indicate that the 3 Gli-BS is essential for the minimal floor plate enhancer activity. Three Gli genes are differentially expressed in the neural tube/plate and transiently overlap with HNF- 3β In order to study the possible involvement of Gli genes in HNF- 3β induction, the expression patterns of Gli, Gli2 and Gli3 in the neural tube/plate were re-examined in comparison with HNF-3β and Shh by in situ hybridization of serial sections of the mouse embryos at 8.5 and 9.5 d.p.c. (Fig. 4). Three Gli genes show specific and dynamic changes of expression patterns along the anteroposterior axis of the neural tube/plate of 8.5 d.p.c. embryos (9-11 somites). Gli transcripts were abundant in the ventral region of the neural tube/plate and gradually lost dorsally (Fig. 4C,H). In the posterior part, where HNF-3β but not yet Shh is transcribed in the ventral midline (Fig. 4A,B), its expression overlaps with HNF-3β (Fig. 4C). However, in the anterior part, where both HNF-3β and Shh are co-expressed in the floor plate, Gli expression is excluded from the floor plate (Fig. 4H). Gli2 transcripts were rather evenly distributed throughout neural tube/plate (Fig. 4D,I). As observed with Gli, its expression also overlaps with HNF-3β in the posterior floor plate (Fig. 4D), but is excluded from the anterior floor plate (Fig. 4I). On the other hand, Gli3 seems to be expressed at a very low level throughout the neural tube in the posterior part, since signals above background levels were observed reproducibly (Fig. 4E). In the anterior part, strong Gli3 expression is widely observed but in an opposite gradient to Gli, i.e. it is strong in the dorsal region and weak in the Fig. 4. Comparison of Gli, Gli-2 and Gli-3 expressions with HNF- 3β and Shh in developing neural tubes. In situ hybridization of HNF-3β (A,F,K), Shh (B,G,L), Gli (C,H,M), Gli-2 (D,I,N) and Gli-3 (E,J,O) on serial cross sections of 8.5 (A-J) and 9.5 d.p.c. (K-O) mouse embryos. Sections through posterior part (A-E) and hindbrain level (F-J) of an 8.5 d.p.c. embryo. Left side somites of D and E are damaged. Sections through posterior forelimb bud level of a 9.5 d.p.c. embryo (K-O). f, floor plate; nc, notochord; n, neural tube/fold; g, gut. Scale bar, 0.2 mm.
6 1318 H. Sasaki and others ventral (Fig. 4J). Again, no signal is seen in the floor plate. Differential expression of the three Gli genes similar to Fig. 4H- J, which are essentially the same as our previous observation (Hui et al., 1994), were also observed in more anterior regions, up to the level of the diencephalon (data not shown). However, in the mid- and forebrains, where HNF-3β expression domains are clearly broader than Shh, the ventral boundary of Gli expression apparently coincides with the Shh boundary (data not shown). It is likely that the differential expression of the three Gli genes along the anteroposterior axis of the neural tube/plate correlates with the differentiation of the neuroectoderm rather than with its position along the body axis per se. Supporting this idea, sections through posterior forelimb bud level (around 11-13th somites) of a 9.5 d.p.c. embryo (Fig. 4K-O), which is equivalent in position to the section of the posterior level of the 8.5 d.p.c. embryo shown in Fig. 4A-E, now show gene expression patterns similar to those in the anterior of the 8.5 d.p.c. embryo shown in Fig. 4F-J. In addition, the expression patterns of three Gli genes in the brain level of the 7.5 d.p.c. embryo (Hui et al., 1994) are similar to those in the posterior of the 8.5 d.p.c. embryo (Fig. 4C-E). Taken together, strong expression of Gli and Gli2 and very weak Gli3 expression, transiently overlap with HNF-3β at the time of its induction. These expression patterns of the three Gli genes are consistent with the model of Gli proteins being activators of the floor plate enhancer of HNF-3β, although the contribution of Gli3 may be minimal. The transient nature of the expression of Gli genes suggests involvement of Gli proteins in induction of HNF-3β rather than in the maintenance of its expression. Similar differential expression of the three Gli genes was also observed in paraxial mesoderm, including presomitic mesoderm, somites and head mesenchyme (Fig. 4 and data not shown). The expression patterns of Gli, Gli2 and Gli3 in the somites are ventromedial, throughout the somite, and dorsal, respectively. Such differential expression of the three Gli genes in paraxial mesoderm also suggests the involvement of Gli genes in dorsoventral patterning of the mesoderm. Gli expression can be up-regulated by Shh The restricted expression of Gli in tissues surrounding Shhexpressing tissues such as the notochord and the floor plate (Fig. 4) suggests that the expression of Gli itself is controlled by Shh signalling. Therefore, this possibility was tested using a multipotential rat neuronal cell, MNS70 (Nakagawa et al., 1996). This cell line is derived from the embryonic forebrain and expresses various ventral neuroepithelium-specific genes upon induction by Shh (Nakagawa et al., 1996). Either a Shh expression plasmid or a control plasmid was transfected into MNS70 cells and the endogenous Gli RNA level was monitored by RT-PCR. Gli RNA was up-regulated by transfection of the Shh expression plasmid (Fig. 5). Although Gli2 and Gli3 were also expressed in this cell line, no obvious change was observed for these transcript levels by transfection of a Shh expression plasmid (data not shown). This result, together with the Gli expression patterns in developing mouse embryos (Fig. 4), suggest that it is likely that the expression of Gli is controlled by Shh signalling in vivo. It is possible that a similar Gli induction by Shh also occurs in the ventral midline of the neural plate, and that this is a part Fig. 5. Expression of Gli is controlled by Shh signalling. MNS70 cells were transfected with pjt4/shh (lanes 3 and 4) or pjt4 (lanes 5 and6), and then analyzed for Gli expression by RT-PCR. Lanes 1 and 2 are mock transfected. Expression of Gli was increased by transfection of Shh expression plasmid. β-actin was used as a control. Duplicates are shown for each plasmid. No obvious induction of Isl-1 by Shh was observed under these conditions (data not shown). of the scenario responsible for HNF-3β induction in the floor plate. The Gli-binding site is a Shh responsive element Since floor plate expression of HNF-3β is induced by Shh, and since a Gli homologue, ci, is involved in Hh signalling pathway in Drosophila, involvement of the 3 Gli-BS motif of the floor plate enhancer within the Shh signalling pathway was tested by transient transfection assay in MNS70 cells. Although it does not express HNF-3β in response to Shh treatment, the basic mechanism of Hh-induced gene expression appears to be common in various tissues, at least in part (see Introduction). Therefore, MNS70 was used to characterize the isolated 3 Gli- BS element in the Shh pathway. A luciferase reporter plasmid having eight copies of either the 3 Gli-BS or the m3 Gli-BS motif was co-transfected with either a Shh expression plasmid or a control expression plasmid into MNS70 cells, expecting autocrine and/or paracrine effects of Shh (Fig. 6A). The expression of the reporter with the 3 Gli-BS motif was increased by co-transfection of the Shh expression vector (Fig. 6B). On the other contrary, the expression of the reporter with the m3 Gli-BS motif was not changed, confirming that the response to Shh was mediated by the Gli-binding site (Fig. 6B). Therefore, the Gli-binding site acts as a Shh responsive element. This is the first demonstration of a Shh responsive element in vitro. GLI, but not GLI3, can activate the Gli-binding site Since Shh activates the 3 Gli-BS, we tested whether any Gli proteins can activate the same reporter. Human GLI and GLI3 were used instead of mouse Gli and Gli3, respectively. Gli2 was not tested because of the present unavailability of fulllength cdna. The expression of the reporter with the 3 Gli-BS motif was increased by co-transfection of the GLI expression vector (Fig. 7A,B). The expression of the reporter with the m3 Gli-BS motif was not changed, confirming that the GLI activates reporter gene expression through the Gli-binding site (Fig. 7B). On the other hand, co-transfection of the GLI3 expression vector did not activate, but rather repressed, reporter expression in the 3 Gli-BS-dependent manner (Fig. 7B and
7 Gli proteins in the HNF-3β enhancer 1319 A effector pjt4/shh CMV intron chick Shh cdna poly A promoter signal B 800 pjt4 reporter 8 x 3'Gli-BS Luc -crystallin basal promoter luciferase luciferase activity x m3'gli-bs Luc Fig. 6. Gli-binding site is a Shh responsive element. (A) Diagram of effector and reporter plasmids. Each dark and light circle of the reporters indicates 3 Gli-BS and m3 Gli-BS sequences, respectively, listed in Fig. 2A. (B) Stimulation of the transcription of 8 3 Gli-BS by Shh. Gene expression of 8 3 Gli-BS Luc was up-regulated by co-transfection with pjt4/shh, while no response was observed with 8 m3 Gli-BS Luc. Each combination is shown in duplicate. effector reporter 0 pjt4/shh pjt4 8 x 3'Gli-BS 8 x m3'gli-bs data not shown). The repression by GLI3 was also tested by co-transfection of both GLI and GLI3 expression vectors. Consistent with the idea of GLI3 being a repressor, GLI3 counteracted the effect of GLI (Fig. 7C). Considering the strong similarities of mouse Gli and Gli3 to human GLI and GLI3, respectively, these results suggest that the Gli, but not Gli3, is a transcriptional activator and can directly activate the 3 Gli- BS. DISCUSSION Transcriptional regulation of HNF-3β in the floor plate The initiation of HNF-3β expression in the floor plate is induced by a signal, Shh, from the notochord. This study showed that the HNF-3β floor plate enhancer contains a binding site for Gli-like proteins (3 Gli-BS) that is essential for floor plate gene expression in transgenic mouse embryos, and that the 3 Gli-BS is activated by Shh and GLI, but not by GLI3, in a cultured cell line. Finally, strong Gli and Gli2, and very weak Gli3 expression, transiently overlap with that of HNF-3β in the ventral midline of the neural tube/plate at the time of floor plate induction. These results suggest that the Shh signalling from the notochord turns on HNF-3β via all or some of the three Gli transcription factors. Among the three Gli proteins, Gli is a better candidate than Gli3 for activation of the 3 Gli-BS, for the following reasons. Judging from the facts that all three Gli genes are co-expressed with HNF-3β at the time of floor plate induction (Fig. 4), and that GLI and GLI3 have similar DNA-binding activities in vitro (Vortkamp et al., 1995), it is equally possible for any of the three Gli proteins to be involved in the activation of the 3 Gli- BS. However, the results of the co-transfection assay of GLI and GLI3 (Fig. 7) support the model of 3 Gli-BS activation by Gli but not by Gli3. Although the results of our co-transfection assay do not completely rule out the possibility of transcriptional activation by GLI3 in certain situations, at least co-transfection of the GLI3 and Shh expression vectors did not convert GLI3 into an activator (H.S., unpublished observation). This model is also consistent with the fact that in the region of overlap with HNF-3β the levels of Gli and Gli3 are very high and very low, respectively. The human GLI cdna used in the transfection assay was isolated from the glioblastoma cell line, and hence may not be wild type (Kinzler et al., 1987). However, the hypothesis of GLI being a transcriptional activator while GLI3 being a repressor has also been independently proposed by Marigo et al. (1996b), based on analysis with chick limb buds. Although the role of Gli2 as a transcriptional regulator is not yet known, the phenotype of homozygous Gli2 null mutant mouse embryos, which have lost HNF-3β expression in the floor plate except in the head region (C.-c. H., unpublished observation), suggests that Gli2 functions as an activator. Finally, Gli proteins are involved only in the process of HNF- 3β induction and not in its maintenance, since none of these three Gli genes are expressed in the differentiated floor plate (Fig. 4) (Hui et al., 1994). Initial Shh signal transduction to Gli proteins probably involves post-translational control, since HNF-3β induction by the notochord does not require de novo protein synthesis in the chick neural explant assay (Ruiz i Altaba et al., 1995b). Therefore, the induction or maintenance of a transcriptional activator, Gli, by Shh signalling (Fig. 5) may be a rather late event that may function to back up or stabilize Shh target gene activation by increasing the Gli protein level. Alternatively, the induction or maintenance of Gli in cells adjacent to the floor plate may contribute to the later inductive events, such as the induction of oligodendrocyte precursors and motor neuron differentiation (Ericson et al., 1996; Orentas and Miller, 1996; Pringle et al., 1996). Although the Gli-binding site (3 Gli-BS) is essential for enhancer activity, and can respond to Shh signalling in transfection assays in a cell line, Gli proteins cannot be the only factors involved in floor plate gene expression in vivo. In fact, multiple copies of the 3 Gli-BS alone did not efficiently activate gene expression in transgenic mouse embryos. Only one out of 11 transgenic embryos showed β-galactosidase
8 1320 H. Sasaki and others A effectors psr -GLI psr -GLI3 psr reporters 8 x 3'Gli-BS Luc 8 x m3'gli-bs Luc SR promoter intron human GLI cdna human GLI3 cdna -crystallin basal promoter luciferase poly A signal Fig. 7. GLI, but not GLI-3, is a transcriptional activator. (A) Diagram of effector and reporter plasmids. (B) Transcriptional activation of 8 Gli-BS by GLI. Gene expression of 8 3 Gli-BS Luc was up-regulated by cotransfection with psrα-gli, while no activation was observed either with 8 m3 Gli- BS Luc reporter or by co-transfection with psrα-gli-3 effector. Each combination is shown in duplicate. (C) Transcriptional repression of 8 Gli-BS by GLI-3. Transcriptional activation of 8 3 Gli-BS Luc by co-transfection with psrα-gli was partially repressed by addition of psrα-gli- 3. The amount of each effector was kept constant by addition of same amount of empty effector plasmid (psrα). Each combination is shown in duplicate. B 4000 C luciferase activity 2000 luciferase activity reporter effector 0 8 x 3'Gli-BS 8 x m3'gli-bs psrα-gli psrα-gli3 psrα reporter effector 8 x 3'Gli-BS 8 x m3'gli-bs psrα-gli psrα-gli3 psrα expression related to Shh (H.S., unpublished observation). Therefore, Gli proteins probably co-operate with other transcription factors that bind to other essential elements of the floor plate enhancer, e.g. the del-6 and A fragment regions, to achieve Shh-inducible and floor plate-restricted gene expression (Fig. 1) (Sasaki and Hogan, 1996). Possible roles of Gli proteins for dorsoventral patterning of the neural tube and paraxial mesoderm mediated by Shh signalling Shh plays important roles in the dorsoventral patterning of the neural tube and somites (Johnson et al., 1994; Ericson et al., 1995; Hynes et al., 1995; Roelink et al., 1995; Yang and Niswander, 1995; Chiang et al., 1996). The present study suggests involvement of Gli proteins in the Shh signalling pathway as transcriptional regulators (see below). The present study also describes the differential expression patterns of the three Gli genes in the neural tube/plate and paraxial mesoderm (Fig. 4). Briefly, Gli is strongly expressed in the ventral and medial parts of the neural tube/plate and paraxial mesoderm, which are supposed to receive the Shh signal from notochord and/or floor plate. Gli2 is rather evenly distributed, and the Gli3 distribution pattern is complementary to Gli, i.e. strongly expressed in the dorsal and lateral parts of the neural tube and paraxial mesoderm, which are supposed to barely receive the Shh signal. These expression patterns, taken together with the results of GLI but not GLI3 being an activator (Fig. 7), and up-regulation of Gli by Shh in cultured cell lines (Fig. 5), suggest that ventrally expressed Gli is involved in the activation of Shh target genes, while dorsally expressed Gli3 has the opposite effect. This model is also supported by the facts that GLI and GLI3 are up- and
9 Gli proteins in the HNF-3β enhancer 1321 down-regulated by Shh in chick limb bud, respectively, and that GLI-VP16 can activate another target of Shh, Ptc (Marigo et al., 1996b). Although no biochemical information is available at present about the role of Gli2, which is evenly expressed throughout the neural tube, analysis of mutant embryos has shown that Gli-2 and Gli-3 have specific and partially redundant functions in skeletal development (Mo et al., 1997). Possible evolutionary conservation of Hedgehog signalling pathway between vertebrates and Drosophila As described in the Introduction, multiple components of the Hh signalling pathway are conserved between Drosophila and vertebrates. Briefly, Drosophila Ptc and vptc are Hh receptors, Drosophila Smo and vsmo are receptor-coupled seven-transmembrane proteins, and PKAs are negative regulators of Hh signalling. The present study suggests the additional conservation of a downstream transcription factor, Ci. In vertebrates, there are three ci-related genes, Gli, Gli2 and Gli3, which encode proteins with closely related five-zinc finger motifs (Hui et al., 1994). The present analysis showed that a binding site for Gli proteins is essential for the enhancer activity of a target gene, that Shh can activate a binding site for Gli proteins, and that GLI is a transcriptional activator. These findings are consistent with a role for Gli proteins in the Shh signalling pathway as transcriptional regulators. However, there seem to be some differences, too. In Drosophila, there is only one ci, and its transcript level is not changed in cells receiving the Hh signal (Domínguez et al., 1996). In mouse, however, there are three Gli genes, and expression of Gli can be up-regulated by Shh signalling. Therefore, the general scheme of Gli/Ci as downstream transcription factors of the Hh signalling pathways seems to be conserved between vertebrates and Drosophila, although the details may be different. I thank Professor Hisato Kondoh for his generous support, encouragement and discussions; members of the laboratory, especially Drs Yujiro Higashi, Yusuke Kamachi and Ryohei Sekido for discussions, technical advice and reagents; Dr Sumihare Noji for pjt4/shh; Dr Yusuke Kamachi for pδ51luc II and SV-β-galactosidase; Dr Bert Vogelstein, The Johns Hopkins University and the Howard Hughes Medical Institution for human GLI and GLI3 cdnas; Dr Shun-suke Ishii for psrα-gli and psrα-gli3; Dr Ryohei Sekido for psrα. Richard O Brien for pgex-hnf-3β; Dr Kathy Mahon for 8.5 d.p.c. mouse embryo cdna library. I also thank Drs Hisato Kondoh and Brigid L.M. Hogan for critically reading the manuscript. Part of this work was supported by a grant-in-aid from the Ministry of Education, Science and Culture of Japan to H.S. REFERENCES Alcedo, J., Ayzenzon, M., Von Ohlen, T., Noll, M. and Hooper, J. E. (1996). The drosophila smoothened gene encodes a seven-pass membrane protein, a putative receptor for the hedgehog signal. Cell 86, Alexandre, C., Jacinto, A. and Ingham, P. W. (1996). Transcriptional activation of hedgehog target genes in Drosophila is mediated directly by the Cubitus interruptus protein, a member of the GLI family of zinc finger DNAbinding proteins. Genes Dev. 10, Ang, S.-L., Wierda, A., Wong, D., Stevens, K. A., Cascio, S., Rossant, J. and Zaret, K. S. (1993). The formation and maintenance of the definitive endoderm lineage in the mouse: involvement of HNF3/fork head proteins. Development 119, Chen, Y. and Struhl, G. (1996). Dual roles for patched in sequestering and transducing hedgehog. Cell 87, Chiang, C., Litingtung, Y., Lee, E., Young, K. E., Corden, J. L., Westphal, H. and Beachy, P. A. (1996). Cyclopia and defective axial patterning in mice lacking Sonic hedgehog gene function. Nature 383, Colamarino, S. A. and Tessier-Lavigne, M. (1995). The axonal chemoattractant Netrin-1 is also a chemorepellent for trochlear motor axons. Cell 81, Currie, P. D. and Ingham, P. W. (1996). Induction of a specific muscle cell type by a hedgehog-like protein in zebrafish. Nature 382, Dodd, J. and Jessell, T. M. (1993). Axon guidance in the mammalian spinal cord. In Cell-cell Signaling in Vertebrate Development (ed. E. J. Robertson, F. R. Maxfieled and H. J. Vogel), pp Academic Press, Inc., San Diego. Domínguez, M., Brunner, M., Hafen, E. and Basler, K. (1996). Sending and recieveing the hedgehog signal: control by the Drosophila Gli protein Cubitus interruptus. Science 272, Echelard, Y., D.J., E., St-Jacques, B., Shen, L., Mohler, J., McMahon, J. A. and McMahon, A. P. (1993). Sonic hedgenog, a member of a family of putative signaling molecules, is implicated in the regulation of CNS polarity. Cell 75, Ekker, S. C., McGrew, L. L., Lai, C. J., Lee, J. J., von Kessler, D. P., Moon, R. T. and Beachy, P. A. (1995a). Distinct expression and shared activities of the hedgehog gene family of Xenopus laevis. Development 121, Ekker, S. C., Ungar, A. R., Greenstein, P., von Kessler, D. P., Porter, J. A., Moon, R. T. and Beachy, P. A. (1995b). Patterning activities of vertebrate hedgehog proteins in the developing eye and brain. Curr. Biol. 5., Ericson, J., Morton, S., Kawakami, A., Roelink, H. and Jessell, T. M. (1996). Two critical periods of Sonic hedgehog signalling required for the specification of motor neuron identity. Cell 87, Ericson, J., Muhr, J., Placzek, M., Lints, T., Jessell, T. M. and Edlund, T. (1995). Sonic hedgehog induces the differentiation of ventral forbrain neurons: a common signal for ventral patterning within the neural tube. Cell 81, Fiering, S. N., Roederer, M., Nolan, G. P., Micklem, D. R., Parks, D. R. and L.A., H. (1991). Improved FACS-Gal:flow cytometric analysis and sorting of viable eukaryotic cells expressing reporter gene constructs. Cytometry 12, Forbes, A. J., Nakano, Y., Taylor, A. M. and Ingham, P. W. (1993). Genetic analysis of hedgehog signalling in the Drosophila embryo. Development supplement, Guan, K. and Dixon, J. E. (1991). Eukaryotic proteins expressed in Escherichia coli: an improved thrombin cleavage and purification procedure of fusion proteins with glutathione S-transferase. Anal. Biochem. 192, Hammerschmidt, M., Bitgood, M. J. and McMahon, A. P. (1996). Protein kinase A is a common negative regulator of Hedgehog signalling in the vertebrate embryo. Genes Dev. 10, Hui, C.-C., Slusarski, D., Platt, K. A., Holmgreen, R. and Joyner, A. L. (1994). Expression of three mouse homologs of the Drosophila segment polarity gene cubitus interruptus, Gli, Gli-2 and Gli-3, in ectoderm- and mesoderm-derived tissues suggests multiple roles during postimplantation development. Dev. Biol. 162, Hynes, M., Porter, J. A., Chiang, C., Chang, D., Tessier-Lavigne, M., Beachy, P. A. and Rosenthal, A. (1995). Induction of midbrain dopaminergic neurons by Sonic hedgehog. Neuron 15, Jessell, T. M. and Dodd, J. (1992). Floor plate- derived signals and the control of neural cell pattern in vertebrates. Harvey Lect. 86, Johnson, R. L., Laufer, E., Riddle, R. D. and Tabin, C. (1994). Ectopic expression of sonic hedgehog alters dorsal-ventral patterning of somites. Cell 79, Kamachi, Y. and Kondoh, H. (1993). Overlapping positive and negative regulatory elements determine lens-specific activity of the δ1-crystallin enhancer. Mol. Cell. Biol. 13, Kennedy, T. E., Serafini, T., Torre, J. R. d. l. and Tessier-Lavigne, M. (1994). Netrins are diffusible chemotropic factors for commissural axons in the embryonic spinal cord. Cell 74, Kinzler, K. W., Bigner, S. H., Bigner, D. D., Trent, J. M., Law, M. L., O Brien, S. J., Wong, A. J. and Vogelstein, B. (1987). Identification of an amplified, highly expressed gene in a human glioma. Science 236, Kinzler, K. W., Ruppert, J. M., Bigner, S. H. and Vogelstein, B. (1988). The GLI gene is a member of Krüppel family of zinc finger proteins. Nature 332, Kinzler, K. W. and Vogelstein, B. (1990). The GLI gene encoles a nuclear protein which binds specific sequences in the human genome. Mol. Cell. Biol. 10,
10 1322 H. Sasaki and others Marigo, V., Davey, R. A., Zuo, Y., Cunninghan, J. M. and Tabin, C. J. (1996a). Biochemical evidence that Patched is the Hedgehog receptor. Nature 384, Marigo, V., Johnson, R. L., Vortkamp, A. and Tabin, C. J. (1996b). Sonic hedgehog differentially regulates expression of GLI and GLI3 during limb development. Dev. Biol. 180, Marti, E., Bumcrot, D. A., Takada, R. and McMahon, A. P. (1995). Requirement of 19K form of Sonic hedgehog for inductionn of distinct ventral cell types in CNS explants. Nature 375, Mo, R., Freer, A. M., Zinyk, D., Crackower, M. A., Heng, H. H.-Q., Chik, K. W., Shi, X. M., Tsui, L.-C., Cheng, S. H., Joyner, A. L. and Hui, C.-c. (1997). Specific and redundant functions of Gli-2 and Gli-3 zinc finger genes in skeletal patterning and development. Development (in press). Nakafuku, M. and Nakamura, S. (1995). Establishment and characterization of a multipotential neural cell line that can conditionally generate neurons, astrocytes, and oligodendrocytes in vitro. J. Neurosci. Res. 41, Nakagawa, Y., Kanako, T., Ogura, T., Suzuki, T., Torii, M., Kaibuchi, K., Arai, K.-I., Nakamura, S. and Nakafuku, M. (1996). Roles of cellautonomous mechanisms for differential expression of region-specific transcription factors in neuroepithelial cells. Development 122, Orenic, T. V., Slusarski, D. C., Kroll, K. L. and Holmgren, R. A. (1990). Cloning and characterization of the segment polarity gene cubitus interruptus Dominant of Drosophila. Genes Dev. 4, Orentas, D. M. and Miller, R. H. (1996). The origin of spinal cord oligodendrycytes is dependent on local influences from notochord. Dev. Biol. 177, Perrimon, N. (1995). Hedgehog and beyond. Cell 80, Pringle, N. P., Yu, W.-P., Guthrie, S., Roelink, H., Lumsden, A., Peterson, A. C. and Richardson, W. D. (1996). Determination of neuroepithelial cell fate: induction of the oligodendrocyte lineage by ventral midline cells and Sonic hedgehog. Dev. Biol. 177, Roelink, H., Porter, J., Chiang, C., Tanabe, Y., Chang, D. T., Beachy, P. A. and Jessell, T. M. (1995). Floor plate and motor neuron induction by different concentrations of the amino-terminal cleavage product of Sonic hedgehog autoproteolysis. Cell 81, Ruiz i Altaba, A., Jessell, T. M. and Roelink, H. (1995a). Restrictions to floor plate induction by hedgehog and winged-helix genes in the neural tube of frog embryos. Mol. Cell. Neurosci. 6, Ruiz i Altaba, A., Placzek, M., Baldassare, M., Dodd, J. and Jessell, T. (1995b). Early stages of notochord and floor plate development in the chick embryo defined by normal and induced expression of HNF-3β. Dev. Biol. 170, Ruiz i Altaba, A., Prezioso, V. R., Darvell, J. E. and Jessell, T. M. (1993). Sequential expression of HNF-3β and HNF-3α by embryonic organizing centers: the dorsal lip/node, notochord and floor plate. Mech. Dev. 44, Ruppert, J. M., Kinzler, K. W., Wong, A. J., Bigner, S. H., Kao, F., Law, M. L., Seuanez, H. N., O Brien, J. O. and Vogelstein, B. (1988). The GLI- Kruppel family of human genes. Mol. Cell. Biol. 8, Ruppert, J. M., Vogelstein, B., Arheden, K. and Kinzler, K. W. (1990). GLI3 encodes a 190-kilodalton protein with multiple regions of GLI similarity. Mol. Cell. Biol 10, Sasaki, H. and Hogan, B. L. M. (1993). Differential expression of multiple fork head related genes during gastrulation and axial pattern formation in the mouse embryo. Development 118, Sasaki, H. and Hogan, B. L. M. (1994). HNF-3β as a regulator of floor plate development. Cell 76, Sasaki, H. and Hogan, B. L. M. (1996). Enhancer analysis of the mouse HNF- 3β gene: regulatory elements for node/notochord and floor plate are independent and consist of multiple sub-elements. Genes to Cells 1, Stone, D. M., Hynes, M., Armanini, M., Swanson, T. A., Gu, Q., Johnson, R. L., Scott, M. P., Pennica, D., Goddard, A., Phillips, H., Noll, M., Hooper, J. E., de Sauvage, F. and Rosenthal, A. (1996). The tumour-suppressor gene patched encodes a candidate receptor for Sonic hedgehog. Nature 384, Takebe, Y., Seki, M., Fujisawa, J., Hoy, P., Yokota, K., Arai, K., Yoshida, M. and Arai, N. (1988). SRα promoter: an efficient and versatile mammalian cdna expression system composed of teh simian virus 40 early promoter and the R-U5 segment of human T-cell leukemia virus type 1 long terminal repeat. Mol. Cell. Biol. 8, van den Heuvel, M. and Ingham, P. W. (1996). smoothened encodes a receptor-like serpentine protein required for hedgehog signalling. Nature 382, Vortkamp, A., Gessler, M. and Grzeschik, K.-H. (1995). Identification of optimized target sequences for the GLI3 zinc finger protein. DNA Cell Biol. 14, Walterhouse, D., Ahmed, M., Slusarski, D., Kalamaras, J., Boucher, D., Holmgren, R. and Iannaccone, P. (1993). gli, a zinc finger transcription factor and oncogene, is expressed during normal mouse development. Dev. Dyn. 196, Yang, Y. and Niswander, L. (1995). Interaction between the signaling molecules WNT7a and SHH during vertebrate limb development: dorsal signals regulate anteroposterior patterning. Cell 80, (Accepted 31 January 1997)
Sonic hedgehog (Shh) signalling in the rabbit embryo
 Sonic hedgehog (Shh) signalling in the rabbit embryo In the first part of this thesis work the physical properties of cilia-driven leftward flow were characterised in the rabbit embryo. Since its discovery
Sonic hedgehog (Shh) signalling in the rabbit embryo In the first part of this thesis work the physical properties of cilia-driven leftward flow were characterised in the rabbit embryo. Since its discovery
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8
 Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
RNA Synthesis and Processing
 RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles
 Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
Regulation of Patched by Sonic hedgehog in the developing
 Proc. Natl. Acad. Sci. USA Vol. 93, pp. 9346-9351, September 1996 Colloquium Paper This paper was presented at a colloquium entitled "Biology of Developmental Transcription Control, " organized by Eric
Proc. Natl. Acad. Sci. USA Vol. 93, pp. 9346-9351, September 1996 Colloquium Paper This paper was presented at a colloquium entitled "Biology of Developmental Transcription Control, " organized by Eric
Induction of motor neurons by Sonic hedgehog is independent of floor plate differentiation
 Induction of motor neurons by Sonic hedgehog is independent of floor plate differentiation Yasuto Tanabe, Henk Roelink and Thomas M. Jessell Howard Hughes Medical Institute, Deptartment of Biochemistry
Induction of motor neurons by Sonic hedgehog is independent of floor plate differentiation Yasuto Tanabe, Henk Roelink and Thomas M. Jessell Howard Hughes Medical Institute, Deptartment of Biochemistry
Biology 218, practise Exam 2, 2011
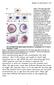 Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Axon guidance I. Paul Garrity March 15, /9.013
 Axon guidance I Paul Garrity March 15, 2004 7.68/9.013 Neuronal Wiring: Functional Framework of the Nervous System Stretch reflex circuit Early theories of axonogenesis Schwann: many neurons link to form
Axon guidance I Paul Garrity March 15, 2004 7.68/9.013 Neuronal Wiring: Functional Framework of the Nervous System Stretch reflex circuit Early theories of axonogenesis Schwann: many neurons link to form
Question Set # 4 Answer Key 7.22 Nov. 2002
 Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio
 Urun 1 Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio Fatma Rabia URUN ilkent University, nkara - TURKEY High mobility group domain containing transcription
Urun 1 Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio Fatma Rabia URUN ilkent University, nkara - TURKEY High mobility group domain containing transcription
3/8/ Complex adaptations. 2. often a novel trait
 Chapter 10 Adaptation: from genes to traits p. 302 10.1 Cascades of Genes (p. 304) 1. Complex adaptations A. Coexpressed traits selected for a common function, 2. often a novel trait A. not inherited from
Chapter 10 Adaptation: from genes to traits p. 302 10.1 Cascades of Genes (p. 304) 1. Complex adaptations A. Coexpressed traits selected for a common function, 2. often a novel trait A. not inherited from
Developmental Biology 3230 Midterm Exam 1 March 2006
 Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its
 Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Role of Organizer Chages in Late Frog Embryos
 Ectoderm Germ Layer Frog Fate Map Frog Fate Map Role of Organizer Chages in Late Frog Embryos Organizer forms three distinct regions Notochord formation in chick Beta-catenin localization How does beta-catenin
Ectoderm Germ Layer Frog Fate Map Frog Fate Map Role of Organizer Chages in Late Frog Embryos Organizer forms three distinct regions Notochord formation in chick Beta-catenin localization How does beta-catenin
PRACTICE EXAM. 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos.
 PRACTICE EXAM 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos. No Low [] Fly Embryo Embryo Non-neural Genes Neuroectoderm Genes
PRACTICE EXAM 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos. No Low [] Fly Embryo Embryo Non-neural Genes Neuroectoderm Genes
The role of Transcriptional Repression in Shh Signalling and Patterning of the Ventral Neural Tube
 From the Department of Cell and Molecular Biology The Medical Nobel Institute Karolinska Institutet, Stockholm, Sweden The role of Transcriptional Repression in Shh Signalling and Patterning of the Ventral
From the Department of Cell and Molecular Biology The Medical Nobel Institute Karolinska Institutet, Stockholm, Sweden The role of Transcriptional Repression in Shh Signalling and Patterning of the Ventral
purpose of this Chapter is to highlight some problems that will likely provide new
 119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Lecture Outlines
 Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Neural development its all connected
 Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
Midterm 1. Average score: 74.4 Median score: 77
 Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
Chapter 4 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays.
 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
!!!!!!!! DB3230 Midterm 2 12/13/2013 Name:
 1. (10 pts) Draw or describe the fate map of a late blastula stage sea urchin embryo. Draw or describe the corresponding fate map of the pluteus stage larva. Describe the sequence of gastrulation events
1. (10 pts) Draw or describe the fate map of a late blastula stage sea urchin embryo. Draw or describe the corresponding fate map of the pluteus stage larva. Describe the sequence of gastrulation events
Segment boundary formation in Drosophila embryos
 Segment boundary formation in Drosophila embryos Development 130, August 2003 Camilla W. Larsen, Elizabeth Hirst, Cyrille Alexandre and Jean Paul Vincent 1. Introduction: - Segment boundary formation:
Segment boundary formation in Drosophila embryos Development 130, August 2003 Camilla W. Larsen, Elizabeth Hirst, Cyrille Alexandre and Jean Paul Vincent 1. Introduction: - Segment boundary formation:
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Tuesday, December 27, 16
 Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Conclusions. The experimental studies presented in this thesis provide the first molecular insights
 C h a p t e r 5 Conclusions 5.1 Summary The experimental studies presented in this thesis provide the first molecular insights into the cellular processes of assembly, and aggregation of neural crest and
C h a p t e r 5 Conclusions 5.1 Summary The experimental studies presented in this thesis provide the first molecular insights into the cellular processes of assembly, and aggregation of neural crest and
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Bio 127 Section I Introduction to Developmental Biology. Cell Cell Communication in Development. Developmental Activities Coordinated in this Way
 Bio 127 Section I Introduction to Developmental Biology Cell Cell Communication in Development Gilbert 9e Chapter 3 It has to be EXTREMELY well coordinated for the single celled fertilized ovum to develop
Bio 127 Section I Introduction to Developmental Biology Cell Cell Communication in Development Gilbert 9e Chapter 3 It has to be EXTREMELY well coordinated for the single celled fertilized ovum to develop
AP Biology Gene Regulation and Development Review
 AP Biology Gene Regulation and Development Review 1. What does the regulatory gene code for? 2. Is the repressor by default active/inactive? 3. What changes the repressor activity? 4. What does repressor
AP Biology Gene Regulation and Development Review 1. What does the regulatory gene code for? 2. Is the repressor by default active/inactive? 3. What changes the repressor activity? 4. What does repressor
Unit 4 Evaluation Question 1:
 Name: Unit 4 Evaluation Question 1: /7 points A naturally occurring dominant mutant in mice is the Doublefoot (Dbf) mutant. Below is an image of the bones from a wildtype (wt) and Doublefoot mutant mouse.
Name: Unit 4 Evaluation Question 1: /7 points A naturally occurring dominant mutant in mice is the Doublefoot (Dbf) mutant. Below is an image of the bones from a wildtype (wt) and Doublefoot mutant mouse.
Introduction. Gene expression is the combined process of :
 1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
CHAPTER : Prokaryotic Genetics
 CHAPTER 13.3 13.5: Prokaryotic Genetics 1. Most bacteria are not pathogenic. Identify several important roles they play in the ecosystem and human culture. 2. How do variations arise in bacteria considering
CHAPTER 13.3 13.5: Prokaryotic Genetics 1. Most bacteria are not pathogenic. Identify several important roles they play in the ecosystem and human culture. 2. How do variations arise in bacteria considering
Chapter 18 Lecture. Concepts of Genetics. Tenth Edition. Developmental Genetics
 Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
THE Hedgehog (Hh) signaling pathway regulates
 Copyright Ó 2006 by the Genetics Society of America DOI: 10.1534/genetics.106.061036 The Contributions of Protein Kinase A and Smoothened Phosphorylation to Hedgehog Signal Transduction in Drosophila melanogaster
Copyright Ó 2006 by the Genetics Society of America DOI: 10.1534/genetics.106.061036 The Contributions of Protein Kinase A and Smoothened Phosphorylation to Hedgehog Signal Transduction in Drosophila melanogaster
Three types of RNA polymerase in eukaryotic nuclei
 Three types of RNA polymerase in eukaryotic nuclei Type Location RNA synthesized Effect of α-amanitin I Nucleolus Pre-rRNA for 18,.8 and 8S rrnas Insensitive II Nucleoplasm Pre-mRNA, some snrnas Sensitive
Three types of RNA polymerase in eukaryotic nuclei Type Location RNA synthesized Effect of α-amanitin I Nucleolus Pre-rRNA for 18,.8 and 8S rrnas Insensitive II Nucleoplasm Pre-mRNA, some snrnas Sensitive
UNIVERSITY OF YORK BIOLOGY. Developmental Biology
 Examination Candidate Number: UNIVERSITY OF YORK BSc Stage 2 Degree Examinations 2017-18 Department: BIOLOGY Title of Exam: Developmental Biology Desk Number: Time allowed: 1 hour and 30 minutes Total
Examination Candidate Number: UNIVERSITY OF YORK BSc Stage 2 Degree Examinations 2017-18 Department: BIOLOGY Title of Exam: Developmental Biology Desk Number: Time allowed: 1 hour and 30 minutes Total
CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON
 PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
1. What are the three general areas of the developing vertebrate limb? 2. What embryonic regions contribute to the developing limb bud?
 Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
10/15/09. Tetrapod Limb Development & Pattern Formation. Developing limb region is an example of a morphogenetic field
 Tetrapod Limb Development & Pattern Formation Figure 16.5(1) Limb Bud Formation derived from lateral plate (somatic) & paraxial (myotome) Fig. 16.2 Prospective Forelimb Field of Salamander Ambystoma maculatum
Tetrapod Limb Development & Pattern Formation Figure 16.5(1) Limb Bud Formation derived from lateral plate (somatic) & paraxial (myotome) Fig. 16.2 Prospective Forelimb Field of Salamander Ambystoma maculatum
Nature Neuroscience: doi: /nn.2662
 Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Lecture 18 June 2 nd, Gene Expression Regulation Mutations
 Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
Goosecoid and HNF-3beta genetically interact to regulate neural tube patterning during mouse embryogenesis
 University of Massachusetts Medical School escholarship@umms Rivera Lab Publications Pediatrics 7-1-1997 Goosecoid and HNF-3beta genetically interact to regulate neural tube patterning during mouse embryogenesis
University of Massachusetts Medical School escholarship@umms Rivera Lab Publications Pediatrics 7-1-1997 Goosecoid and HNF-3beta genetically interact to regulate neural tube patterning during mouse embryogenesis
BIS &003 Answers to Assigned Problems May 23, Week /18.6 How would you distinguish between an enhancer and a promoter?
 Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
Hedgehog Signal Transduction: From Flies to Vertebrates
 Experimental Cell Research 253, 25 33 (1999) Article ID excr.1999.4676, available online at http://www.idealibrary.com on Hedgehog Signal Transduction: From Flies to Vertebrates Maximilien Murone,* Arnon
Experimental Cell Research 253, 25 33 (1999) Article ID excr.1999.4676, available online at http://www.idealibrary.com on Hedgehog Signal Transduction: From Flies to Vertebrates Maximilien Murone,* Arnon
Chapter 11. Development: Differentiation and Determination
 KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
Life Sciences For NET & SLET Exams Of UGC-CSIR. Section B and C. Volume-08. Contents A. BASIC CONCEPT OF DEVELOPMENT 1
 Section B and C Volume-08 Contents 5. DEVELOPMENTAL BIOLOGY A. BASIC CONCEPT OF DEVELOPMENT 1 B. GAMETOGENESIS, FERTILIZATION AND EARLY DEVELOPMENT 23 C. MORPHOGENESIS AND ORGANOGENESIS IN ANIMALS 91 0
Section B and C Volume-08 Contents 5. DEVELOPMENTAL BIOLOGY A. BASIC CONCEPT OF DEVELOPMENT 1 B. GAMETOGENESIS, FERTILIZATION AND EARLY DEVELOPMENT 23 C. MORPHOGENESIS AND ORGANOGENESIS IN ANIMALS 91 0
Types of biological networks. I. Intra-cellurar networks
 Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Exam 4 ID#: July 7, 2008
 Biology 4361 Name: KEY Exam 4 ID#: July 7, 2008 Multiple choice (one point each; indicate the best answer) 1. RNA polymerase II is not able to transcribe RNA unless a. it is first bound to TFIIB. b. its
Biology 4361 Name: KEY Exam 4 ID#: July 7, 2008 Multiple choice (one point each; indicate the best answer) 1. RNA polymerase II is not able to transcribe RNA unless a. it is first bound to TFIIB. b. its
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature1237 a b retinol retinal RA OH RDH (retinol dehydrogenase) O H Raldh2 O R/R.6.4.2 (retinaldehyde dehydrogenase 2) RA retinal retinol..1.1 1 Concentration (nm)
SUPPLEMENTARY INFORMATION doi:1.138/nature1237 a b retinol retinal RA OH RDH (retinol dehydrogenase) O H Raldh2 O R/R.6.4.2 (retinaldehyde dehydrogenase 2) RA retinal retinol..1.1 1 Concentration (nm)
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1. HSP21 expression in 35S:HSP21 and hsp21 knockdown plants. (a) Since no T- DNA insertion line for HSP21 is available in the publicly available T-DNA collections,
Supplementary Figure 1 Supplementary Figure 1. HSP21 expression in 35S:HSP21 and hsp21 knockdown plants. (a) Since no T- DNA insertion line for HSP21 is available in the publicly available T-DNA collections,
16 CONTROL OF GENE EXPRESSION
 16 CONTROL OF GENE EXPRESSION Chapter Outline 16.1 REGULATION OF GENE EXPRESSION IN PROKARYOTES The operon is the unit of transcription in prokaryotes The lac operon for lactose metabolism is transcribed
16 CONTROL OF GENE EXPRESSION Chapter Outline 16.1 REGULATION OF GENE EXPRESSION IN PROKARYOTES The operon is the unit of transcription in prokaryotes The lac operon for lactose metabolism is transcribed
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
Developmental genetics: finding the genes that regulate development
 Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning
 MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning Learning Goals for Lecture 29 4.1 Describe the contributions of early developmental events in the embryo to the formation
MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning Learning Goals for Lecture 29 4.1 Describe the contributions of early developmental events in the embryo to the formation
10/2/2015. Chapter 4. Determination and Differentiation. Neuroanatomical Diversity
 Chapter 4 Determination and Differentiation Neuroanatomical Diversity 1 Neurochemical diversity: another important aspect of neuronal fate Neurotransmitters and their receptors Excitatory Glutamate Acetylcholine
Chapter 4 Determination and Differentiation Neuroanatomical Diversity 1 Neurochemical diversity: another important aspect of neuronal fate Neurotransmitters and their receptors Excitatory Glutamate Acetylcholine
Floor Plate and Motor Neuron Induction by Different Concentrations of the Amino-Terminal Cleavage Product of Sonic Hedgehog Autoproteolysis
 Cell, Vol. 81,445-455, May 5, 1995, Copyright 1995 by Cell Press Floor Plate and Motor euron Induction by Different Concentrations of the Amino-Terminal Cleavage Product of Sonic Hedgehog Autoproteolysis
Cell, Vol. 81,445-455, May 5, 1995, Copyright 1995 by Cell Press Floor Plate and Motor euron Induction by Different Concentrations of the Amino-Terminal Cleavage Product of Sonic Hedgehog Autoproteolysis
A Direct Contact between the Dorsal rel Homology Domain and Twist May Mediate Transcriptional Synergy
 MOLECULAR AND CELLULAR BIOLOGY, June 1997, p. 3345 3355 Vol. 17, No. 6 0270-7306/97/$04.00 0 Copyright 1997, American Society for Microbiology A Direct Contact between the Dorsal rel Homology Domain and
MOLECULAR AND CELLULAR BIOLOGY, June 1997, p. 3345 3355 Vol. 17, No. 6 0270-7306/97/$04.00 0 Copyright 1997, American Society for Microbiology A Direct Contact between the Dorsal rel Homology Domain and
Cell Cell Communication in Development
 Biology 4361 Developmental Biology Cell Cell Communication in Development June 25, 2008 Cell Cell Communication Concepts Cells in developing organisms develop in the context of their environment, including
Biology 4361 Developmental Biology Cell Cell Communication in Development June 25, 2008 Cell Cell Communication Concepts Cells in developing organisms develop in the context of their environment, including
Drosophila. Cubitus interruptus-independent transduction of the Hedgehog signal in
 Development 127, 5509-5522 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV7818 5509 Cubitus interruptus-independent transduction of the Hedgehog signal in Drosophila Armel Gallet
Development 127, 5509-5522 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV7818 5509 Cubitus interruptus-independent transduction of the Hedgehog signal in Drosophila Armel Gallet
Honors Biology Reading Guide Chapter 11
 Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Dorsoventral patterning of the vertebrate neural tube is conserved in a protochordate
 Development 124, 2335-2344 (1997) Printed in Great Britain The Company of Biologists Limited 1997 DEV9537 2335 Dorsoventral patterning of the vertebrate neural tube is conserved in a protochordate Joseph
Development 124, 2335-2344 (1997) Printed in Great Britain The Company of Biologists Limited 1997 DEV9537 2335 Dorsoventral patterning of the vertebrate neural tube is conserved in a protochordate Joseph
Axis determination in flies. Sem 9.3.B.5 Animal Science
 Axis determination in flies Sem 9.3.B.5 Animal Science All embryos are in lateral view (anterior to the left). Endoderm, midgut; mesoderm; central nervous system; foregut, hindgut and pole cells in yellow.
Axis determination in flies Sem 9.3.B.5 Animal Science All embryos are in lateral view (anterior to the left). Endoderm, midgut; mesoderm; central nervous system; foregut, hindgut and pole cells in yellow.
Unicellular: Cells change function in response to a temporal plan, such as the cell cycle.
 Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Lecture 7. Development of the Fruit Fly Drosophila
 BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION
 MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
Bypass and interaction suppressors; pathway analysis
 Bypass and interaction suppressors; pathway analysis The isolation of extragenic suppressors is a powerful tool for identifying genes that encode proteins that function in the same process as a gene of
Bypass and interaction suppressors; pathway analysis The isolation of extragenic suppressors is a powerful tool for identifying genes that encode proteins that function in the same process as a gene of
The Eukaryotic Genome and Its Expression. The Eukaryotic Genome and Its Expression. A. The Eukaryotic Genome. Lecture Series 11
 The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
Drosophila Life Cycle
 Drosophila Life Cycle 1 Early Drosophila Cleavage Nuclei migrate to periphery after 10 nuclear divisions. Cellularization occurs when plasma membrane folds in to divide nuclei into cells. Drosophila Superficial
Drosophila Life Cycle 1 Early Drosophila Cleavage Nuclei migrate to periphery after 10 nuclear divisions. Cellularization occurs when plasma membrane folds in to divide nuclei into cells. Drosophila Superficial
Exam 1 ID#: October 4, 2007
 Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Chapter 20. Initiation of transcription. Eukaryotic transcription initiation
 Chapter 20. Initiation of transcription Eukaryotic transcription initiation 2003. 5.22 Prokaryotic vs eukaryotic Bacteria = one RNA polymerase Eukaryotes have three RNA polymerases (I, II, and III) in
Chapter 20. Initiation of transcription Eukaryotic transcription initiation 2003. 5.22 Prokaryotic vs eukaryotic Bacteria = one RNA polymerase Eukaryotes have three RNA polymerases (I, II, and III) in
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018
 Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
Developmental processes Differential gene expression Introduction to determination The model organisms used to study developmental processes
 Date Title Topic(s) Learning Outcomes: Sept 28 Oct 3 1. What is developmental biology and why should we care? 2. What is so special about stem cells and gametes? Developmental processes Differential gene
Date Title Topic(s) Learning Outcomes: Sept 28 Oct 3 1. What is developmental biology and why should we care? 2. What is so special about stem cells and gametes? Developmental processes Differential gene
Novel retinoic acid generating activities in the neural tube and heart identified by conditional rescue of Raldh2 null mutant mice
 Development 129, 2271-2282 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV9819 2271 Novel retinoic acid generating activities in the neural tube and heart identified by conditional
Development 129, 2271-2282 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV9819 2271 Novel retinoic acid generating activities in the neural tube and heart identified by conditional
Dorsalization of the neural tube by the non-neural ectoderm
 Development 121, 2099-2106 (1995) Printed in Great Britain The Company of Biologists Limited 1995 2099 Dorsalization of the neural tube by the non-neural ectoderm Mary E. Dickinson 1, Mark A. J. Selleck
Development 121, 2099-2106 (1995) Printed in Great Britain The Company of Biologists Limited 1995 2099 Dorsalization of the neural tube by the non-neural ectoderm Mary E. Dickinson 1, Mark A. J. Selleck
3.B.1 Gene Regulation. Gene regulation results in differential gene expression, leading to cell specialization.
 3.B.1 Gene Regulation Gene regulation results in differential gene expression, leading to cell specialization. We will focus on gene regulation in prokaryotes first. Gene regulation accounts for some of
3.B.1 Gene Regulation Gene regulation results in differential gene expression, leading to cell specialization. We will focus on gene regulation in prokaryotes first. Gene regulation accounts for some of
Behavior of DNA-lacking mitochondria in Entamoeba histolytica revealed by organelle transplant
 9 10 11 Behavior of DNA-lacking mitochondria in Entamoeba histolytica revealed by organelle transplant Makoto Kazama 1 *, Sanae Ogiwara, Takashi Makiuchi 1, Kazuhiro Yoshida 1, Kumiko Nakada-Tsukui, Tomoyoshi
9 10 11 Behavior of DNA-lacking mitochondria in Entamoeba histolytica revealed by organelle transplant Makoto Kazama 1 *, Sanae Ogiwara, Takashi Makiuchi 1, Kazuhiro Yoshida 1, Kumiko Nakada-Tsukui, Tomoyoshi
Prokaryotic Regulation
 Prokaryotic Regulation Control of transcription initiation can be: Positive control increases transcription when activators bind DNA Negative control reduces transcription when repressors bind to DNA regulatory
Prokaryotic Regulation Control of transcription initiation can be: Positive control increases transcription when activators bind DNA Negative control reduces transcription when repressors bind to DNA regulatory
Cell-Cell Communication in Development
 Biology 4361 - Developmental Biology Cell-Cell Communication in Development October 2, 2007 Cell-Cell Communication - Topics Induction and competence Paracrine factors inducer molecules Signal transduction
Biology 4361 - Developmental Biology Cell-Cell Communication in Development October 2, 2007 Cell-Cell Communication - Topics Induction and competence Paracrine factors inducer molecules Signal transduction
7.013 Spring 2005 Problem Set 4
 MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel NAME TA 7.013 Spring 2005 Problem Set 4 FRIDAY April 8th,
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel NAME TA 7.013 Spring 2005 Problem Set 4 FRIDAY April 8th,
Midline signalling is required for Pax gene regulation and patterning of the eyes
 Development 121, 3267-3278 (1995) Printed in Great Britain The Company of Biologists Limited 1995 3267 Midline signalling is required for Pax gene regulation and patterning of the eyes Rachel Macdonald
Development 121, 3267-3278 (1995) Printed in Great Britain The Company of Biologists Limited 1995 3267 Midline signalling is required for Pax gene regulation and patterning of the eyes Rachel Macdonald
Why Flies? stages of embryogenesis. The Fly in History
 The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
Wan-Ju Liu 1,2, John S Reece-Hoyes 3, Albertha JM Walhout 3 and David M Eisenmann 1*
 Liu et al. BMC Developmental Biology 2014, 14:17 RESEARCH ARTICLE Open Access Multiple transcription factors directly regulate Hox gene lin-39 expression in ventral hypodermal cells of the C. elegans embryo
Liu et al. BMC Developmental Biology 2014, 14:17 RESEARCH ARTICLE Open Access Multiple transcription factors directly regulate Hox gene lin-39 expression in ventral hypodermal cells of the C. elegans embryo
Regulation and signaling. Overview. Control of gene expression. Cells need to regulate the amounts of different proteins they express, depending on
 Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
MBios 401/501: Lecture 14.2 Cell Differentiation I. Slide #1. Cell Differentiation
 MBios 401/501: Lecture 14.2 Cell Differentiation I Slide #1 Cell Differentiation Cell Differentiation I -Basic principles of differentiation (p1305-1320) -C-elegans (p1321-1327) Cell Differentiation II
MBios 401/501: Lecture 14.2 Cell Differentiation I Slide #1 Cell Differentiation Cell Differentiation I -Basic principles of differentiation (p1305-1320) -C-elegans (p1321-1327) Cell Differentiation II
From DNA to Diversity
 From DNA to Diversity Molecular Genetics and the Evolution of Animal Design Sean B. Carroll Jennifer K. Grenier Scott D. Weatherbee Howard Hughes Medical Institute and University of Wisconsin Madison,
From DNA to Diversity Molecular Genetics and the Evolution of Animal Design Sean B. Carroll Jennifer K. Grenier Scott D. Weatherbee Howard Hughes Medical Institute and University of Wisconsin Madison,
Initiation of translation in eukaryotic cells:connecting the head and tail
 Initiation of translation in eukaryotic cells:connecting the head and tail GCCRCCAUGG 1: Multiple initiation factors with distinct biochemical roles (linking, tethering, recruiting, and scanning) 2: 5
Initiation of translation in eukaryotic cells:connecting the head and tail GCCRCCAUGG 1: Multiple initiation factors with distinct biochemical roles (linking, tethering, recruiting, and scanning) 2: 5
Differential patterning of ventral midline cells by axial mesoderm is regulated by BMP7 and chordin
 Development 126, 397-408 (1999) Printed in Great Britain The Company of Biologists Limited 1998 DEV4103 397 Differential patterning of ventral midline cells by axial mesoderm is regulated by BMP7 and chordin
Development 126, 397-408 (1999) Printed in Great Britain The Company of Biologists Limited 1998 DEV4103 397 Differential patterning of ventral midline cells by axial mesoderm is regulated by BMP7 and chordin
REVIEW SESSION. Wednesday, September 15 5:30 PM SHANTZ 242 E
 REVIEW SESSION Wednesday, September 15 5:30 PM SHANTZ 242 E Gene Regulation Gene Regulation Gene expression can be turned on, turned off, turned up or turned down! For example, as test time approaches,
REVIEW SESSION Wednesday, September 15 5:30 PM SHANTZ 242 E Gene Regulation Gene Regulation Gene expression can be turned on, turned off, turned up or turned down! For example, as test time approaches,
Chapter 10, 11, 14: Gene Expression, Regulation, and Development Exam
 Chapter 10, 11, 14: Gene Expression, Regulation, and Development Exam Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Why did the original one-gene, one-enzyme
Chapter 10, 11, 14: Gene Expression, Regulation, and Development Exam Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Why did the original one-gene, one-enzyme
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila July 9, 2008 Drosophila Development Overview Fertilization Cleavage Gastrulation Drosophila body plan Oocyte formation Genetic control
Developmental Biology Biology 4361 Axis Specification in Drosophila July 9, 2008 Drosophila Development Overview Fertilization Cleavage Gastrulation Drosophila body plan Oocyte formation Genetic control
Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc
 OPTIMIZATION OF IMMUNOBLOT PROTOCOL 121 Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc Jacqueline Bjornton and John Wheeler Faculty Sponsor: Anne
OPTIMIZATION OF IMMUNOBLOT PROTOCOL 121 Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc Jacqueline Bjornton and John Wheeler Faculty Sponsor: Anne
Principles of Genetics
 Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Mouse dispatched mutants fail to distribute hedgehog proteins and are defective in hedgehog signaling
 Development 129, 5753-5765 2002 The Company of Biologists Ltd doi:10.1242/dev.00178 5753 Mouse dispatched mutants fail to distribute hedgehog proteins and are defective in hedgehog signaling Takatoshi
Development 129, 5753-5765 2002 The Company of Biologists Ltd doi:10.1242/dev.00178 5753 Mouse dispatched mutants fail to distribute hedgehog proteins and are defective in hedgehog signaling Takatoshi
Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus:
 m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
2. Yeast two-hybrid system
 2. Yeast two-hybrid system I. Process workflow a. Mating of haploid two-hybrid strains on YPD plates b. Replica-plating of diploids on selective plates c. Two-hydrid experiment plating on selective plates
2. Yeast two-hybrid system I. Process workflow a. Mating of haploid two-hybrid strains on YPD plates b. Replica-plating of diploids on selective plates c. Two-hydrid experiment plating on selective plates
The Gene The gene; Genes Genes Allele;
 Gene, genetic code and regulation of the gene expression, Regulating the Metabolism, The Lac- Operon system,catabolic repression, The Trp Operon system: regulating the biosynthesis of the tryptophan. Mitesh
Gene, genetic code and regulation of the gene expression, Regulating the Metabolism, The Lac- Operon system,catabolic repression, The Trp Operon system: regulating the biosynthesis of the tryptophan. Mitesh
Axon Guidance. Multiple decision points along a growing axon s trajectory Different types of axon guidance cues:
 Axon Guidance Multiple decision points along a growing axon s trajectory Different types of axon guidance cues: Contact mediated - requires direct contact by growth cone Long range - growth cone responds
Axon Guidance Multiple decision points along a growing axon s trajectory Different types of axon guidance cues: Contact mediated - requires direct contact by growth cone Long range - growth cone responds
Name: SBI 4U. Gene Expression Quiz. Overall Expectation:
 Gene Expression Quiz Overall Expectation: - Demonstrate an understanding of concepts related to molecular genetics, and how genetic modification is applied in industry and agriculture Specific Expectation(s):
Gene Expression Quiz Overall Expectation: - Demonstrate an understanding of concepts related to molecular genetics, and how genetic modification is applied in industry and agriculture Specific Expectation(s):
