Archaeal diversity and abundance within different layers of summer sea-ice and seawater from Prydz Bay, Antarctica
|
|
|
- Katrina Gardner
- 5 years ago
- Views:
Transcription
1 Article Advances in Polar Science doi: /j.advps March 2014 Vol. 25 No. 1: Archaeal diversity and abundance within different layers of summer sea-ice and seawater from Prydz Bay, Antarctica MA Jifei 1,2, DU Zongjun 2, LUO Wei 1, YU Yong 1, ZENG Yixin 1, CHEN Bo 1 & LI Huirong 1* 1 Polar Research Institute of China, Shanghai , China; 2 College of Marine Science, Shandong University, Weihai, Weihai , China Received 15 May 2013; accepted 30 October 2013 Abstract Fluorescence in-situ hybridization (FISH) and 16S rrna gene clone library analyses were used to determine the abundance and diversity of archaea in Prydz Bay, Antarctica. Correlation analysis was also performed to assess links between physicochemical parameters and archaeal abundance and diversity within the sea-ice. Samples of sea-ice and seawater were collected during the 26th Chinese National Antarctic Research Expedition. The results of FISH showed that archaea were relatively abundant within the top layer of the sea-ice, and correlation analysis suggested that the concentration of NH + 4 might be one of the main factors underlying this distribution pattern. However, using 16S rrna gene libraries, archaea were not detected in the top and middle layers of the sea-ice. All archaeal clones obtained from the bottom layer of the sea-ice were grouped into the Marine Group I Crenarchaeota while the archaeal clones from seawater were assigned to Marine Group I Crenarchaeota, Marine Group II Euryarchaeota, and Marine Group III Euryarchaeota. Overall, the findings of this study showed that the diversity of archaea in the sea-ice in Prydz Bay was low. Keywords Antarctica, summer sea-ice, archaea, diversity, abundance Citation: Ma J F, Du Z J, Luo W, et al. Archaeal diversity and abundance within different layers of summer sea-ice and seawater from Prydz Bay, Antarctica. Adv Polar Sci, 2014, 25: 54-60, doi: /j.advps Introduction Research has shown that archaea are widespread and abundant in many different environments [1-3]. The domain Archaea consists of five distinct phyla, Crenarchaeota, Euryarchaeota, Nanoarchaeota, Korarchaeota, and Thaumarchaeota [4-7]. Archaea were first identified in Antarctic ecosystems in 1988 [8], and to date archaea have been found in marine waters, frozen lakes, and other Antarctic marine environments [9-10]. Previous studies in Antarctic waters have found high archaeal abundance with archaea accounting for 34% of prokaryotes [11-12]. In Antarctic marine environments, Group I Crenarchaeota and Group II Euryarchaeota were predominant, and Group I was generally more abundant than * Corresponding author ( lihuirong@pric.gov.cn) Group II [13]. Archaea make up a sizable proportion of the biomass in Antarctic frazil ice [9]. However, archaea have not been identified in sea-ice until recently, although they are abundant in Arctic pelagic communities. Brown and Bowman [14] and Brinkmeyer and Knittel [15] found that archaea were absent from Arctic summer sea-ice and sea-ice near Antarctica. Junge et al. [16] investigated the abundance of archaea using fluorescence in-situ hybridization (FISH) and found that they accounted for 0 3.4% of the total cells in Arctic winter seaice. Collins et al. [17] showed that archaeal sequences were present in both Arctic winter sea-ice and seawater clone libraries, and the majority (91%) of sequences from the archaeal libraries belonged to Marine Group I [17]. Cowie et al. [10] found a low abundance of archaea in Antarctic seaice with archaea making up just 6.6% of the prokaryotic community. The majority (90.8%) of the sequences were journal.polar.gov.cn
2 Archaeal diversity and abundance within different layers of summer sea-ice and seawater from Prydz Bay, Antarctica 55 clustered within Marine Group I [10]. The main aim of this study was to quantify archaeal abundance using FISH and to describe archaeal diversity using 16S rrna gene molecular methods. In addition, correlation analysis was carried out to investigate the relationship between physicochemical parameters and archaeal diversity and abundance within the sea-ice. To our knowledge, this is the first study to investigate the abundance and diversity of archaea in different sea-ice layers. Therefore, this study provides important new information on archaea in Antarctic sea-ice ecosystems and forms a theoretical basis for future research. 2 Materials and methods 2.1 Sample collection and DNA extraction Sea-ice samples (Table 1) were collected from Prydz Bay, Antarctica, during the 26th Chinese National Antarctic Research Expedition (26th CHINARE; November 2008 to May 2009). Samples were collected using the method described by Li et al. [18] Sea-ice samples were kept frozen during transportation and were stored at 20 C until analysis. The fluorometric method was used to measure chlorophyll a (Chl a) concentrations. Total organic carbon (TOC) and total organic nitrogen (TON) were measured by researchers at Tongji University in Shanghai. The concentrations of nitrate nitrogen (NO 3 ), ammonium nitrogen, phosphate phosphorus, and silica were measured using a continuous flow nutrient analyzer (Skalar San++, Skalar UK (Ltd.), York, UK). The DNA extraction was carried out following the method described by Li et al. [18] 2.2 Fluorescence in-situ hybridization (FISH) The abundance of archaea was determined using FISH with probe ARCH915-Cy3 (5 -GTG CTC CCC CGC CAA TTC CT-3 ). Hybridization was carried out following the method described by Junge et al. [16] A Nikon 80i fluorescence microscope was used to visualize samples under excitation and emission filters of 360 and 420 nm, respectively, for DAPI-labeled cells (blue), and under excitation and emission filters of 550 and 570 nm, respectively, for Cy3-labeled cells (red). For each sample, cells in 20 randomly selected microscopic fields were counted and the concentration was then determined as follows: cells cm 3 = (the mean number of cells per field/the volume of the sample trapped on the filter) (the area of the filter/the area of the field). Table 1 Ice core collection stations and sea-ice layers Station Date (YYYY-MM-DD) Longitude Latitude Length of ice core /cm Layers /cm I E S 167 Top: 0 20 Middle: Bottom: I E S 168 Top: 0 20 Middle: Bottom: I E S 168 Top: 0 20 Middle: Bottom: I E S 183 Top: 0 20 Middle: Bottom: I E S 191 Top: 0 20 Middle: Bottom: Phylogenetic analysis Samples from station I02 were selected for the analysis of diversity. The 16S rrna gene was amplified with the primer pair Arch-21F, and Arch-958R [19]. Clone library construction, screening, and processing followed the methods described by DeLong [19]. Briefly, the PCR products were purified with a gel extraction kit (Watson, Shanghai, China) according to the manufacturer s instructions. Cloning was conducted with the pgem-t Vector (Promega, Madison, WI, USA) following the manufacturer s instructions. Libraries were screened for the 0.8 kb 16S rdna insert by PCR with M13 primers. The restriction endonuclease Alu I was used for amplified ribosomal DNA restriction analysis (ARDRA). The sequences that had different ARDRA patterns were sequenced on an ABI PRISM 3730 sequencer. Clustal X1.8, BioEdit Sequence Alignment Editor (version 5.0.9), and MEGA (version 4) software packages were used for the analysis of all nucleotide sequences. The 16S rrna gene sequences obtained in this study
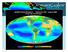
3 56 Ma J F, et al. Adv Polar Sci March(2014) Vol. 25 No. 1 were submitted to the GenBank database (accession nos. JQ JQ753094). 2.4 Estimation of operational taxonomic unit richness To estimate richness and to ensure rigorous comparison among the communities, sequences were placed into operational taxonomic units (OTUs) at a level of sequence similarity of 97%. All OTU richness and sample coverage calculations were performed with the program EstimateS (version 8.0). The OTU richness was calculated for each sample using the non-parametric estimator Chao 1 [20]. Extrapolation using best-fit regression analysis was performed to calculate the point at which 95% confidence intervals (CIs) did not overlap [21]. 2.5 Correlation analysis Correlation analysis was performed to investigate the Station Section T m / C Table 2 Physical and chemical characteristics of sea-ice Salinity/ TOC /(mg L -1 ) TON /(µg ml -1 ) relationship between physicochemical parameters and archaeal diversity and abundance within the sea-ice. The analysis was performed using SPSS Statistics 17 software package. 3 Results 3.1 Physicochemical parameters of the sea-ice Physical and chemical characteristics of the sea-ice samples are shown in Table 2. The highest concentrations of Chl a, TOC, TON, ammonium nitrogen, phosphate, nitrate nitrogen, and silicate were found within the bottom layer of the sea-ice. 3.2 Total cell numbers and archaeal abundance The fluorescent stain 4,6-diamidino-2-phenylindole 2HCl Chl a 3 PO 4 3 SiO 4 NH 4 + (NO 3 + NO 2) I01 I01T I01M I01B I02 I02T I02M ND I02B I03 I03T ND 9.87 I03M I03B ND I04 I04T I04M I04B I05 I05T I05M I05B (DAPI) for DNA was used to determine the abundance of total bacteria in sea-ice samples melted in the dark at 4 C (Figure 1a). The total number of bacterial cells in the seaice samples was cells cm 3. The mean numbers of bacterial cells in the top, middle, and bottom layers of the sea-ice were (8.84±0.66) 10 5, (2.78±1.02) 10 5, and (9.26±4.4) 10 5 cells cm 3, respectively. The results of FISH showed that the total number of archaeal cells in the sea-ice was cells cm 3 (Figure 1b), and the mean numbers of archaeal cells in the top, middle, and bottom layers were (2.5±2.2) 10 5, (1.05±0.97) 10 5, and (5.12±1.17) 10 4 cells cm 3, respectively. The total number of bacterial cells in seawater samples was cells cm 3 and the mean abundance of archaea in the seawater was (3.4±2.9) 10 4 cells cm 3. Figure 1 The abundance of total bacteria and archaea in sea-ice and seawater samples. T, M, and B represent the top, middle, and bottom layers of sea-ice, respectively, and W represents seawater.
4 Archaeal diversity and abundance within different layers of summer sea-ice and seawater from Prydz Bay, Antarctica Correlation analysis Alu I enzyme pattern analysis was used in this study. Four phylotypes were found in the 16S rrna gene clone analysis of samples of the bottom layer of the sea-ice and nine phylotypes were detected in the seawater samples. Correlation analysis was performed to investigate the relationship between the number of phylotypes and the physicochemical parameters of the sea-ice and the seawater. The results presented in Table 3 show that archaeal diversity was significantly positively correlated with salinity (r=0.965, P=0.035) and the concentration of silicate (r=0.996, P=0.004). It was therefore suggested that salinity and the concentration of silicate could play an important role in determining the distribution of archaeal cells in sea-ice. At sites I02 and I03, there was a significant negative correlation between the abundance of archaea and the concentration of ammonium nitrogen (r= 1.000), but there was no obvious correlation between the abundance and other physicochemical parameters. 3.4 Analysis of archaeal 16S rrna gene clone libraries In total, 144 positive archaeal 16S rrna gene clones from the top, middle, and bottom layers of sea-ice from station I02 and from seawater were PCR screened with M13 primers. After sequencing and chimera checking, a total of 37 clones of 16S rrna gene sequences from the four archaeal clone libraries (12 from the bottom layer and 25 from seawater) were obtained for further analysis. The majority of sequences were similar to 16S rrna gene sequences in GenBank with similarities ranging from 90% 100%. Sequences having the same type after ARDRA were assigned to the same phylotype. Four and nine phylotypes were defined within the bottom layer of the sea-ice and the seawater libraries, respectively. The phylogenetic relationships among clones are displayed in Figure 2. All the clones from bottom layer libraries were affiliated with Marine Group I and most of the clones from the libraries were similar to environmental sequences recovered from Arctic or Antarctic sea-ice and seawater. Most clones from the seawater libraries were affiliated with Marine Group I and Marine Group III; only one clone was affiliated with Marine Group II. Six sequences detected in sea-ice and six sequences detected in seawater samples were associated with clones from Arctic winter sea-ice samples (99%) [17] and all were affiliated with Marine Group I. The archaeal sequence I02W106 affiliated with Marine Group II was associated with the clones from Arctic sea-ice samples (99%). Marine Group III, as a unique representative phylotype in the library, was associated with the clone SHZW623 from seawater. 3.5 Estimation of OTU richness of clone libraries As the clone libraries were constructed concurrently, the archaeal diversity of Antarctic sea-ice and seawater clone libraries underwent comparative analysis to extrapolate species richness. Plotting the cumulative number of OTUs estimated against the sampling effort generates species richness curves (Figure 3). The highest estimated number of species was present in the seawater samples, which contained seven OTUs (95% CI, 7 to 37), whereas the bottom sea-ice samples yielded three OTUs (95% CI, 2 to 6). The 95% CIs for the bottom sea-ice and seawater sample communities overlapped, therefore there was no significant difference in species richness among the samples (P>0.05). The Shannon s index of diversity for bottom sea-ice (I02B) and seawater (I02W) samples was 0.72 and 1.34, respectively. Table 3 Correlation analysis between physicochemical parameters and archaeal diversity and abundance within sea-ice 3 T m Salinity TOC TON Chl a PO 3 4 SiO4 4 NH + NO 3 + NO 2 Diversity I02 Correlation * ** Significance FISH I02 Correlation ** Significance I03 Correlation ** Significance I04 Correlation Significance I05 Correlation Significance Notes: ** significant correlation at the 0.01 level (double sided); * significant correlation at the 0.05 level (double sided).
5 58 Ma J F, et al. Adv Polar Sci March(2014) Vol. 25 No. 1 Figure 2 Phylogenetic tree of archaeal 16S rrna gene sequences obtained from the bottom sea-ice layer at station I02 and seawater under ice clone libraries. The trees were constructed using Jukes-Cantor distances and the neighbor-joining method. Bootstrap support values >55% (of 100 replicates) are shown. The numbers are the numbers of closely related sequences with similarity >99% in the same library and the numbers in parentheses are the accession numbers of sequences. The scale bar indicates the estimated number of base changes per nucleotide sequence position. I02B and I02W represent the bottom layer of sea-ice and the seawater samples, respectively. 4 Discussion Figure 3 OTU estimate curves derived from archaeal 16S rrna gene clone libraries data. I02B and I02W represent the bottom layer of sea-ice and the seawater samples, respectively. Our results showed that the mean abundance of archaea in sea-ice was cells cm 3, making up 6.6% of the total picoplankton abundance in the Antarctic sea-ice samples. Results of the present study were consistent with those of Cowie et al. [10], who also found that archaea were present in low abundance and accounted for 6.6% of the
6 Archaeal diversity and abundance within different layers of summer sea-ice and seawater from Prydz Bay, Antarctica 59 prokaryotic community in Antarctic sea-ice. The abundance of archaea was also low in Arctic sea-ice samples and Brinkmeyer and Knittel [15] showed that 93% 96% of DAPIstained cells hybridized with the general bacterial probe, and archaea were below the detection limit of FISH in Arctic and Antarctic pack-ice communities. Junge et al. [16] determined that archaeal cells made up 0 3.4% of total cells in Arctic winter sea-ice. In our study, the mean abundance of archaea was lowest in the bottom layer of the sea-ice, and archaeal abundance showed a significant negative correlation with the concentration of ammonium nitrogen, suggesting that the concentration of ammonium nitrogen might be identified as a major driver of this distribution pattern. Church et al. [13] reported that the abundance of archaea increased significantly with the depth of seawater, averaged cells cm 3, and accounted for 9% 39% of the total picoplankton abundance in Antarctic Peninsula water. These findings are consistent with our results; the mean abundance of archaea in our seawater sample was (3.4±2.9) 10 4 cells cm 3, making up 14.7% of the total picoplankton abundance. Kirchman et al. [22] found that Crenarchaeota increased from about 10% of prokaryotes in surface waters of the western Arctic waters to as much as 40% in samples from depths of m. Kirchman et al. [22] also found that the abundance of Crenarchaeota was correlated with ammonium nitrogen concentrations. The present study provides characterization of archaeal diversity in sea-ice and seawater collected from Antarctica, based on 16S rrna gene clone libraries. Overall, a total of 37 archaeal 16S rrna gene clones from the bottom layer of the sea-ice and seawater samples revealed low phylogenetic diversity. All sequences derived from the bottom layer of the sea-ice fell into one archaeal phylogenetic lineage, Marine Group I. Collins et al. [17] found that Marine Group I Crenarchaeota was the dominant archaeal phylotype in Arctic sea-ice and seawater libraries, and they also detected the rarer Marine Group II Euryarchaeota. Cowie et al. [10] obtained similar results with the majority (90.8%) of sequences clustered within Marine Group I. Archaeal abundance measured using quantitative PCR in Antarctic sea-ice samples was found to be low, accounting for just 6.6% of the prokaryotic community [10]. We did not detect Marine Group II in our Antarctic sea-ice samples. Marine Group I and Marine Group III were the predominant groups in our Antarctic seawater samples and our results suggested that Group II populations are not numerically abundant in Antarctic plankton communities. Our results were in agreement with previous studies. Galand et al. [23] reported an archaeal community dominated by Group III Euryarchaeota and demonstrated that Group III Euryarchaeota were more abundant in deep-water masses, representing the second most abundant tags after Group I Crenarchaeota in the deep Atlantic layer of the central Arctic Ocean. Group III Euryarchaeota were first identified in the Northeast Pacific [24] and were later detected in different marine environments. However, Group III is rarely represented in marine ecosystems, in which Marine Group I Crenarchaeota and Group II Euryarchaeota are the most abundant groups. Recently, metagenomic analysis revealed the presence of genes that resembled those found in ammonia-oxidizing bacteria in Group III DNA fragments, suggesting that at least some members of Group III Euryarchaeota may oxidize ammonia [23]. If this finding is confirmed it suggests that the group could play an important role in the nitrogen cycle of Antarctica. The objectives of this study were to quantify archaeal abundance and to investigate the distribution of archaea in Antarctic sea-ice and seawater. Our data showed that the mean abundance of archaea was higher in the top and middle layers of sea-ice. However, we did not detect archaea in the top and middle layer sea-ice samples on 16S rrna gene analysis, suggesting that the general archaeal primer may underestimate total archaeal copy numbers. The diversity of archaea in sea-ice was low and our findings were consistent with previous results. Acknowledgments This work was supported by the Natural Science Foundation of China (Grant nos , and ), the Chinese Polar Environment Comprehensive Investigation & Assessment Programs (Grant no. CHINARE ), and the State High-Tech Research and Development Project (863) of the Ministry of Science and Technology of China (Grant no. 2012AA021706). This work is also part of the Project Chinese National Basic Research Program. Data were issued by the Data-sharing Platform of Polar Science ( cn) maintained by Polar Research Institute of China (PRIC) and Chinese National Arctic & Antarctic Data Center (CN-NADC). We appreciate the assistance of the crew of the R/V XUE LONG icebreaker and the contribution of the scientists who collected samples for us on the 26th CHINARE cruise, especially Xiaoge Bai, Rongrong Wu, and Shanglin Xiong. References 1 Karner M B, DeLong E F, Karl D M. Archaeal dominance in the mesopelagic zone of the Pacific Ocean. Nature, 2001, 409(6819): Leininger S, Urich T, Schloter M, et al. Archaea predominate among ammonia-oxidizing prokaryotes in soils. Nature, 2006, 442(7104): Gillan D C, Danis B. The archaebacterial communities in Antarctic bathypelagic sediments. Deep Sea Res Part II, 2007, 54(16-17): Woese C R, Kandler O, Wheelis M L. Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proc Natl Acad Sci USA, 1990, 87(12): Huber H, Hohn M J, Rachel R, et al. A new phylum of Archaea represented by a nanosized hyperthermophilic symbiont. Nature, 2002, 417(6884): Elkins J G, Podarc M, Graham D E, et al. A korarchaeal genome reveals insights into the evolution of the Archaea. Proc Natl Acad Sci USA, 2008, 105(23): Brochier-Armanet C, Boussau B, Gribaldo S, et al. Mesophilic Crenarchaeota: proposal for a third archaeal phylum, the Thaumarchaeota. Nature Rev Microbiol, 2008, 6(3):
7 60 Ma J F, et al. Adv Polar Sci March(2014) Vol. 25 No. 1 8 R a n z m a n n P D, S t a c k e b r a n d t E, S a n d e r s o n K, e t a l. Halobacteriumlacusprofundi sp. nov., a halophilic bacterium isolated from Deep Lake, Antarctica. Syst Appl Microbiol, 1988, 11(1): DeLong E F, Wu K Y, Prézelin B B, et al. High abundance of Archaea in Antarctic marine picoplankton. Nature, 1994, 371(6499): Cowie R M, Maas E W, Ryan K G. Archaeal diversity revealed in Antarctic sea ice. Antar Sci Inst Sub, 2011, 23(6): Thomas D N, Dieckmann G S. Antarctic sea ice a habitat for extremophiles. Science, 2002, 295(5555): Bowman J P, Mccammon S A, Brown M V, et al. Diversity and association of psychrophilic bacteria in Antarctic sea ice. Appl Environ Microbiol, 1997, 63(8): Church M J, Delong E F, Ducklow H W, et al. Abundance and distribution of planktonic Archaea and Bacteria in the waters west of the Antarctic Peninsula. Limnol Oceanogr, 2003, 48(5): Brown M V, Bowman J P. A molecular phylogenetic survey of sea-ice microbial communities (SIMCO). Microbiol Ecol, 2001, 35(3): Brinkmeyer R, Knittel K. Diversity and structure of bacterial communities in Arctic versus Antarctic pack ice. Appl Environ Microbiol, 2003, 69(11): Junge K H, Eicken H, Deming J W. Bacterial activity at -2 to -20 C in Arctic wintertime sea ice. Appl Environ Microbiol, 2004, 70(1): Collins R E, Rocap G, Deming J W. Persistence of bacterial and archaeal communities in sea ice through an Arctic winter. Environ Microbiol, 2010, 12(7): Li H, Yu Y, Zeng Y, et al. Molecular genetic diversity of bacteria in the bottom section of Arctic sea ice from the Canada Basin. Acta Oceanol Sin, 2005, DeLong E F. Archaea in coastal marine environments. Proc Natl Acad Sci USA, 1992, 89(12): Chao A. Estimating the population size for capture-recapture data with unequal catchability. Biometrics, 1987, 43(4): Hughes J B, Hellmann J J, Ricketts T H, et al. Counting the uncountable: statistical approaches to estimating microbial diversity. Appl Environ Microbiol, 2001, 67(10): Kirchman D L, Elifantz H, Dittel A I, et al. Standing stocks and activity of Archaea and Bacteria in the western Arctic Ocean. Limnol Oceanogr, 2007, 52(2): Galand P E, Casamayor E, Kirchman D L, et al. Unique archaeal assemblages in the Arctic ocean unveiled by massively parallel tag sequencing. Inter Soc Microbial Ecol, 2009, 3(7): DeLong E F, Preston C M. Community genomics among stratified microbial assemblages in the ocean's interior. Science, 2006, 311(5760):
Exploring Microbes in the Sea. Alma Parada Postdoctoral Scholar Stanford University
 Exploring Microbes in the Sea Alma Parada Postdoctoral Scholar Stanford University Cruising the ocean to get us some microbes It s all about the Microbe! Microbes = microorganisms an organism that requires
Exploring Microbes in the Sea Alma Parada Postdoctoral Scholar Stanford University Cruising the ocean to get us some microbes It s all about the Microbe! Microbes = microorganisms an organism that requires
Microbiome: 16S rrna Sequencing 3/30/2018
 Microbiome: 16S rrna Sequencing 3/30/2018 Skills from Previous Lectures Central Dogma of Biology Lecture 3: Genetics and Genomics Lecture 4: Microarrays Lecture 12: ChIP-Seq Phylogenetics Lecture 13: Phylogenetics
Microbiome: 16S rrna Sequencing 3/30/2018 Skills from Previous Lectures Central Dogma of Biology Lecture 3: Genetics and Genomics Lecture 4: Microarrays Lecture 12: ChIP-Seq Phylogenetics Lecture 13: Phylogenetics
Abundance and distribution of planktonic Archaea and Bacteria in the waters west of the Antarctic Peninsula
 Limnol. Oceanogr., 48(5), 2003, 1893 1902 2003, by the American Society of Limnology and Oceanography, Inc. Abundance and distribution of planktonic Archaea and Bacteria in the waters west of the Antarctic
Limnol. Oceanogr., 48(5), 2003, 1893 1902 2003, by the American Society of Limnology and Oceanography, Inc. Abundance and distribution of planktonic Archaea and Bacteria in the waters west of the Antarctic
Chad Burrus April 6, 2010
 Chad Burrus April 6, 2010 1 Background What is UniFrac? Materials and Methods Results Discussion Questions 2 The vast majority of microbes cannot be cultured with current methods Only half (26) out of
Chad Burrus April 6, 2010 1 Background What is UniFrac? Materials and Methods Results Discussion Questions 2 The vast majority of microbes cannot be cultured with current methods Only half (26) out of
Standing stocks and activity of Archaea and Bacteria in the western Arctic Ocean
 Limnol. Oceanogr., 52(2), 2007, 495 507 E 2007, by the American Society of Limnology and Oceanography, Inc. Standing stocks and activity of Archaea and Bacteria in the western Arctic Ocean David L. Kirchman,
Limnol. Oceanogr., 52(2), 2007, 495 507 E 2007, by the American Society of Limnology and Oceanography, Inc. Standing stocks and activity of Archaea and Bacteria in the western Arctic Ocean David L. Kirchman,
concentration ( mol l -1 )
 concentration ( mol l -1 ) 8 10 0 20 40 60 80 100 120 140 160 180 methane sulfide ammonium oxygen sulfate (/10) b depth (m) 12 14 Supplementary Figure 1. Water column parameters from August 2011. Chemical
concentration ( mol l -1 ) 8 10 0 20 40 60 80 100 120 140 160 180 methane sulfide ammonium oxygen sulfate (/10) b depth (m) 12 14 Supplementary Figure 1. Water column parameters from August 2011. Chemical
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Detailed overview of the primer-free full-length SSU rrna library preparation.
 Supplementary Figure 1 Detailed overview of the primer-free full-length SSU rrna library preparation. Detailed overview of the primer-free full-length SSU rrna library preparation. Supplementary Figure
Supplementary Figure 1 Detailed overview of the primer-free full-length SSU rrna library preparation. Detailed overview of the primer-free full-length SSU rrna library preparation. Supplementary Figure
Abundance, Diversity, and Activity of Ammonia-Oxidizing Prokaryotes in the Coastal Arctic Ocean in Summer and Winter
 APPLIED AND ENVIRONMENTAL MICROBIOLOGY, Mar. 2011, p. 2026 2034 Vol. 77, No. 6 0099-2240/11/$12.00 doi:10.1128/aem.01907-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Abundance,
APPLIED AND ENVIRONMENTAL MICROBIOLOGY, Mar. 2011, p. 2026 2034 Vol. 77, No. 6 0099-2240/11/$12.00 doi:10.1128/aem.01907-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Abundance,
net production = 0 (how well is low light used) also quantum
 net production = 0 (how well is low light used) also quantum particulate organic matter = POM particulate organic carbon = POC particulate organic nitrogen = PON zooplankton POM POM fish CO 2 fixation
net production = 0 (how well is low light used) also quantum particulate organic matter = POM particulate organic carbon = POC particulate organic nitrogen = PON zooplankton POM POM fish CO 2 fixation
MiGA: The Microbial Genome Atlas
 December 12 th 2017 MiGA: The Microbial Genome Atlas Jim Cole Center for Microbial Ecology Dept. of Plant, Soil & Microbial Sciences Michigan State University East Lansing, Michigan U.S.A. Where I m From
December 12 th 2017 MiGA: The Microbial Genome Atlas Jim Cole Center for Microbial Ecology Dept. of Plant, Soil & Microbial Sciences Michigan State University East Lansing, Michigan U.S.A. Where I m From
Meat and two veg? Determining feeding selectivity of bivalve larvae in the Western English Channel with traditional and molecular techniques.
 Meat and two veg? Determining feeding selectivity of bivalve larvae in the Western English Channel with traditional and molecular techniques. Pennie Lindeque, Elaine Fileman, Claudia Halsband-Lenk, Helen
Meat and two veg? Determining feeding selectivity of bivalve larvae in the Western English Channel with traditional and molecular techniques. Pennie Lindeque, Elaine Fileman, Claudia Halsband-Lenk, Helen
This supplementary information includes the data and corresponding citations
 S1 (table) This supplementary information includes the data and corresponding citations presented in Box 1 Figure for the proportion of inactive cells in different systems. Active cells were assessed using
S1 (table) This supplementary information includes the data and corresponding citations presented in Box 1 Figure for the proportion of inactive cells in different systems. Active cells were assessed using
Dr Mike Dyall-Smith. Archaea: Main points. Archaea: Discovery. Archaea: Discovery. Discovery of the Archaea. Lecture: Archaeal diversity
 Lecture: Archaeal diversity Dr Mike Dyall-Smith Haloarchaea Research Lab., Lab 3.07 mlds@unimelb.edu.au Reference: Microbiology (Prescott et al., 6th). Chapter 20. Archaea: Main points Discovery of a third
Lecture: Archaeal diversity Dr Mike Dyall-Smith Haloarchaea Research Lab., Lab 3.07 mlds@unimelb.edu.au Reference: Microbiology (Prescott et al., 6th). Chapter 20. Archaea: Main points Discovery of a third
Outline Classes of diversity measures. Species Divergence and the Measurement of Microbial Diversity. How do we describe and compare diversity?
 Species Divergence and the Measurement of Microbial Diversity Cathy Lozupone University of Colorado, Boulder. Washington University, St Louis. Outline Classes of diversity measures α vs β diversity Quantitative
Species Divergence and the Measurement of Microbial Diversity Cathy Lozupone University of Colorado, Boulder. Washington University, St Louis. Outline Classes of diversity measures α vs β diversity Quantitative
Hydrologic CTD Dataset Obtained through Chinese Antarctic Expeditions
 Journal of Global Change Data & Discovery. 2017, 1(2): 157-164 DOI:10.3974/geodp.2017.02.04 www.geodoi.ac.cn 2017 GCdataPR Global Change Research Data Publishing & Repository Hydrologic CTD Dataset Obtained
Journal of Global Change Data & Discovery. 2017, 1(2): 157-164 DOI:10.3974/geodp.2017.02.04 www.geodoi.ac.cn 2017 GCdataPR Global Change Research Data Publishing & Repository Hydrologic CTD Dataset Obtained
Microbes and mountains: metagenetics on Mount Fuji, Japan. Jonathan Adams, Biology Department, SNU, Korea
 Microbes and mountains: metagenetics on Mount Fuji, Japan Jonathan Adams, Biology Department, SNU, Korea jonadams@snu.ac.kr Until about a decade ago, culturing could only yield 8,000 described species
Microbes and mountains: metagenetics on Mount Fuji, Japan Jonathan Adams, Biology Department, SNU, Korea jonadams@snu.ac.kr Until about a decade ago, culturing could only yield 8,000 described species
2001 State of the Ocean: Chemical and Biological Oceanographic Conditions in the Newfoundland Region
 Stock Status Report G2-2 (2) 1 State of the Ocean: Chemical and Biological Oceanographic Conditions in the Background The Altantic Zone Monitoring Program (AZMP) was implemented in 1998 with the aim of
Stock Status Report G2-2 (2) 1 State of the Ocean: Chemical and Biological Oceanographic Conditions in the Background The Altantic Zone Monitoring Program (AZMP) was implemented in 1998 with the aim of
Microbes usually have few distinguishing properties that relate them, so a hierarchical taxonomy mainly has not been possible.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy. Slowly evolving molecules (e.g., rrna) used for large-scale structure; "fast- clock" molecules for fine-structure.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Heterogeneous archaeal communities in the particle-rich environment of an arctic shelf ecosystem
 Available online at www.sciencedirect.com Journal of Marine Systems 74 (2008) 774 782 www.elsevier.com/locate/jmarsys Heterogeneous archaeal communities in the particle-rich environment of an arctic shelf
Available online at www.sciencedirect.com Journal of Marine Systems 74 (2008) 774 782 www.elsevier.com/locate/jmarsys Heterogeneous archaeal communities in the particle-rich environment of an arctic shelf
Proteorhodopsin phototrophy in the ocean
 Nature 411, 786-789 (14 June 2001) doi:10.1038/35081051; Received 3 January 2001; Accepted 26 March 2001 Proteorhodopsin phototrophy in the ocean Oded Béjà, Elena N. Spudich, John L. Spudich, Marion Leclerc
Nature 411, 786-789 (14 June 2001) doi:10.1038/35081051; Received 3 January 2001; Accepted 26 March 2001 Proteorhodopsin phototrophy in the ocean Oded Béjà, Elena N. Spudich, John L. Spudich, Marion Leclerc
Estimating prokaryotic diversity: When are 16S rdna libraries large enough?
 LIMNOLOGY and OCEANOGRAPHY: METHODS Limnol. Oceanogr.: Methods 2, 2004, 114 125 2004, by the American Society of Limnology and Oceanography, Inc. Estimating prokaryotic diversity: When are 16S rdna libraries
LIMNOLOGY and OCEANOGRAPHY: METHODS Limnol. Oceanogr.: Methods 2, 2004, 114 125 2004, by the American Society of Limnology and Oceanography, Inc. Estimating prokaryotic diversity: When are 16S rdna libraries
Archaeal Diversity and the Prevalence of Crenarchaeota in Salt Marsh Sediments
 APPLIED AND ENVIRONMENTAL MICROBIOLOGY, June 2009, p. 4211 4215 Vol. 75, No. 12 0099-2240/09/$08.00 0 doi:10.1128/aem.00201-09 Copyright 2009, American Society for Microbiology. All Rights Reserved. Archaeal
APPLIED AND ENVIRONMENTAL MICROBIOLOGY, June 2009, p. 4211 4215 Vol. 75, No. 12 0099-2240/09/$08.00 0 doi:10.1128/aem.00201-09 Copyright 2009, American Society for Microbiology. All Rights Reserved. Archaeal
Microbial Taxonomy. Microbes usually have few distinguishing properties that relate them, so a hierarchical taxonomy mainly has not been possible.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Wuhan, Hubei , China 3 School of Life Sciences, University of Nevada at Las Vegas, 4505 Maryland Parkway, Las Vegas, NV , USA
 Supplementary Material Deep-sea piezosphere and piezophiles: geomicrobiology and biogeochemistry Jiasong Fang 1, Li Zhang 2, and Dennis A. Bazylinski 3 1 Department of Natural Sciences, Hawaii Pacific
Supplementary Material Deep-sea piezosphere and piezophiles: geomicrobiology and biogeochemistry Jiasong Fang 1, Li Zhang 2, and Dennis A. Bazylinski 3 1 Department of Natural Sciences, Hawaii Pacific
Proposal for a SCOR Working Group to Investigate the Role of Viruses in Marine Ecosystems
 Proposal for a SCOR Working Group to Investigate the Role of Viruses in Marine Ecosystems Abstract Viruses are a crucial component affecting the trophodynamics and composition of marine food webs. While
Proposal for a SCOR Working Group to Investigate the Role of Viruses in Marine Ecosystems Abstract Viruses are a crucial component affecting the trophodynamics and composition of marine food webs. While
Title ghost-tree: creating hybrid-gene phylogenetic trees for diversity analyses
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 Title ghost-tree: creating hybrid-gene phylogenetic trees for diversity analyses
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 Title ghost-tree: creating hybrid-gene phylogenetic trees for diversity analyses
Results of oceanographic analyses conducted under JARPA and possible evidence of environmental changes.
 SC/D06/J30 Results of oceanographic analyses conducted under JARPA and possible evidence of environmental changes. Tomowo Watanabe*, Takashi Yabuki**, Toshio Suga**, Kimio Hanawa**, Koji Matsuoka*** and
SC/D06/J30 Results of oceanographic analyses conducted under JARPA and possible evidence of environmental changes. Tomowo Watanabe*, Takashi Yabuki**, Toshio Suga**, Kimio Hanawa**, Koji Matsuoka*** and
Microbiota: Its Evolution and Essence. Hsin-Jung Joyce Wu "Microbiota and man: the story about us
 Microbiota: Its Evolution and Essence Overview q Define microbiota q Learn the tool q Ecological and evolutionary forces in shaping gut microbiota q Gut microbiota versus free-living microbe communities
Microbiota: Its Evolution and Essence Overview q Define microbiota q Learn the tool q Ecological and evolutionary forces in shaping gut microbiota q Gut microbiota versus free-living microbe communities
Comparing Prokaryotic and Eukaryotic Cells
 A prokaryotic cell Basic unit of living organisms is the cell; the smallest unit capable of life. Features found in all cells: Ribosomes Cell Membrane Genetic Material Cytoplasm ATP Energy External Stimuli
A prokaryotic cell Basic unit of living organisms is the cell; the smallest unit capable of life. Features found in all cells: Ribosomes Cell Membrane Genetic Material Cytoplasm ATP Energy External Stimuli
ABUNDANCE, DIVERSITY, AND ACTIVITY OF AMMONIA-OXIDIZING PROKARYOTES IN THE COASTAL ARCTIC OCEAN IN SUMMER AND WINTER. Glenn D.
 ABUNDANCE, DIVERSITY, AND ACTIVITY OF AMMONIA-OXIDIZING PROKARYOTES IN THE COASTAL ARCTIC OCEAN IN SUMMER AND WINTER by Glenn D. Christman A thesis submitted to the Faculty of the University of Delaware
ABUNDANCE, DIVERSITY, AND ACTIVITY OF AMMONIA-OXIDIZING PROKARYOTES IN THE COASTAL ARCTIC OCEAN IN SUMMER AND WINTER by Glenn D. Christman A thesis submitted to the Faculty of the University of Delaware
Taxonomy. Content. How to determine & classify a species. Phylogeny and evolution
 Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
Distributions of dissolved inorganic carbon and total alkalinity in the Western Arctic Ocean
 Article Advances in Polar Science doi:10.3724/sp.j.1085.2011.00246 December 2011 Vol.22 No.4 246 252 Distributions of dissolved inorganic carbon and total alkalinity in the Western Arctic Ocean SUN Heng
Article Advances in Polar Science doi:10.3724/sp.j.1085.2011.00246 December 2011 Vol.22 No.4 246 252 Distributions of dissolved inorganic carbon and total alkalinity in the Western Arctic Ocean SUN Heng
This is a repository copy of Microbiology: Mind the gaps in cellular evolution.
 This is a repository copy of Microbiology: Mind the gaps in cellular evolution. White Rose Research Online URL for this paper: http://eprints.whiterose.ac.uk/114978/ Version: Accepted Version Article:
This is a repository copy of Microbiology: Mind the gaps in cellular evolution. White Rose Research Online URL for this paper: http://eprints.whiterose.ac.uk/114978/ Version: Accepted Version Article:
Phylogenetic diversity and ecology of environmental Archaea Charles E Robertson, J Kirk Harris, John R Spear and Norman R Pace
 Phylogenetic diversity and ecology of environmental Archaea Charles E Robertson, J Kirk Harris, John R Spear and Norman R Pace On the basis of culture studies, Archaea were thought to be synonymous with
Phylogenetic diversity and ecology of environmental Archaea Charles E Robertson, J Kirk Harris, John R Spear and Norman R Pace On the basis of culture studies, Archaea were thought to be synonymous with
Archaea Ancient Oddities
 Archaea Ancient Oddities Death in Yellowstone If the waters of Yellowstone are so deadly, can anything survive in them? Animal Bones in hot spring Dead Trees in Mammoth Hot Springs Yes, but how? A History
Archaea Ancient Oddities Death in Yellowstone If the waters of Yellowstone are so deadly, can anything survive in them? Animal Bones in hot spring Dead Trees in Mammoth Hot Springs Yes, but how? A History
A Novel Ribosomal-based Method for Studying the Microbial Ecology of Environmental Engineering Systems
 A Novel Ribosomal-based Method for Studying the Microbial Ecology of Environmental Engineering Systems Tao Yuan, Asst/Prof. Stephen Tiong-Lee Tay and Dr Volodymyr Ivanov School of Civil and Environmental
A Novel Ribosomal-based Method for Studying the Microbial Ecology of Environmental Engineering Systems Tao Yuan, Asst/Prof. Stephen Tiong-Lee Tay and Dr Volodymyr Ivanov School of Civil and Environmental
SEAWIFS VALIDATION AT THE CARIBBEAN TIME SERIES STATION (CATS)
 SEAWIFS VALIDATION AT THE CARIBBEAN TIME SERIES STATION (CATS) Jesús Lee-Borges* and Roy Armstrong Department of Marine Science, University of Puerto Rico at Mayagüez, Mayagüez, Puerto Rico 00708 Fernando
SEAWIFS VALIDATION AT THE CARIBBEAN TIME SERIES STATION (CATS) Jesús Lee-Borges* and Roy Armstrong Department of Marine Science, University of Puerto Rico at Mayagüez, Mayagüez, Puerto Rico 00708 Fernando
Supplementary Information
 Supplementary Information Altitudinal patterns of diversity and functional traits of metabolically active microorganisms in stream biofilms Linda Wilhelm 1, Katharina Besemer 2, Lena Fragner 3, Hannes
Supplementary Information Altitudinal patterns of diversity and functional traits of metabolically active microorganisms in stream biofilms Linda Wilhelm 1, Katharina Besemer 2, Lena Fragner 3, Hannes
Prokaryotes Vs. Eukaryotes
 The Microbial World Prokaryotes Vs. Eukaryotes Mircrobes of the Ocean Primary Producers Are the organisms that produce bio-mass from inorganic compounds (autotrophs). -Photosynthetic autotrophs Phytoplankton
The Microbial World Prokaryotes Vs. Eukaryotes Mircrobes of the Ocean Primary Producers Are the organisms that produce bio-mass from inorganic compounds (autotrophs). -Photosynthetic autotrophs Phytoplankton
Contribution of Archaea to Total Prokaryotic Production in the Deep Atlantic Ocean
 APPLIED AND ENVIRONMENTAL MICROBIOLOGY, May 2005, p. 2303 2309 Vol. 71, No. 5 0099-2240/05/$08.00 0 doi:10.1128/aem.71.5.2303 2309.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved.
APPLIED AND ENVIRONMENTAL MICROBIOLOGY, May 2005, p. 2303 2309 Vol. 71, No. 5 0099-2240/05/$08.00 0 doi:10.1128/aem.71.5.2303 2309.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved.
Biogeography of major bacterial groups in the Delaware Estuary
 Limnol. Oceanogr., 50(5), 2005, 697 706 2005, by the American Society of Limnology and Oceanography, Inc. Biogeography of major bacterial groups in the Delaware Estuary David L. Kirchman, Ana I. Dittel,
Limnol. Oceanogr., 50(5), 2005, 697 706 2005, by the American Society of Limnology and Oceanography, Inc. Biogeography of major bacterial groups in the Delaware Estuary David L. Kirchman, Ana I. Dittel,
Hiromi Nishida. 1. Introduction. 2. Materials and Methods
 Evolutionary Biology Volume 212, Article ID 342482, 5 pages doi:1.1155/212/342482 Research Article Comparative Analyses of Base Compositions, DNA Sizes, and Dinucleotide Frequency Profiles in Archaeal
Evolutionary Biology Volume 212, Article ID 342482, 5 pages doi:1.1155/212/342482 Research Article Comparative Analyses of Base Compositions, DNA Sizes, and Dinucleotide Frequency Profiles in Archaeal
OCN621: Biological Oceanography- Sediment Microbiology. Guangyi Wang POST 103B
 OCN621: Biological Oceanography- Sediment Microbiology Guangyi Wang POST 103B guangyi@hawaii.edu Three Domains of Life 1) Unrooted phylogenetic tree constructed based on small-subunit rrna genes; 2) Members
OCN621: Biological Oceanography- Sediment Microbiology Guangyi Wang POST 103B guangyi@hawaii.edu Three Domains of Life 1) Unrooted phylogenetic tree constructed based on small-subunit rrna genes; 2) Members
Interpreting the Molecular Tree of Life: What Happened in Early Evolution? Norm Pace MCD Biology University of Colorado-Boulder
 Interpreting the Molecular Tree of Life: What Happened in Early Evolution? Norm Pace MCD Biology University of Colorado-Boulder nrpace@colorado.edu Outline What is the Tree of Life? -- Historical Conceptually
Interpreting the Molecular Tree of Life: What Happened in Early Evolution? Norm Pace MCD Biology University of Colorado-Boulder nrpace@colorado.edu Outline What is the Tree of Life? -- Historical Conceptually
Supplementary Figure S1
 Supplementary Figure S1 KRO STO R-p R-p3 RUD R-nt L-n L-h L-l ph 7 WEON [mg/kg dw soil] 3 WEOC [mg/kg dw soil] 3 Soil type sandy sandy loam silty clay GSF B77V B77T E16 Figure S1:
Supplementary Figure S1 KRO STO R-p R-p3 RUD R-nt L-n L-h L-l ph 7 WEON [mg/kg dw soil] 3 WEOC [mg/kg dw soil] 3 Soil type sandy sandy loam silty clay GSF B77V B77T E16 Figure S1:
Arctic Ocean Biology. from the surface to the deep sea
 Arctic Ocean Biology from the surface to the deep sea Christina Bienhold Helmholtz Max Planck Research Group for Deep Sea Ecology and Technology cbienhol@mpi-bremen.de ACCESS Summerschool, Bremen, Germany
Arctic Ocean Biology from the surface to the deep sea Christina Bienhold Helmholtz Max Planck Research Group for Deep Sea Ecology and Technology cbienhol@mpi-bremen.de ACCESS Summerschool, Bremen, Germany
A Microscopic Approach to Investigate Bacteria under In-Situ Conditions in Arctic Lake Ice:
 A Microscopic Approach to Investigate Bacteria under In-Situ Conditions in Arctic Lake Ice: Initial Comparisons to Sea Ice Karen Junge 1, School of Oceanography, Astrobiology Program, University of Washington,
A Microscopic Approach to Investigate Bacteria under In-Situ Conditions in Arctic Lake Ice: Initial Comparisons to Sea Ice Karen Junge 1, School of Oceanography, Astrobiology Program, University of Washington,
POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics
 POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics - in deriving a phylogeny our goal is simply to reconstruct the historical relationships between a group of taxa. - before we review the
POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics - in deriving a phylogeny our goal is simply to reconstruct the historical relationships between a group of taxa. - before we review the
Assigning Taxonomy to Marker Genes. Susan Huse Brown University August 7, 2014
 Assigning Taxonomy to Marker Genes Susan Huse Brown University August 7, 2014 In a nutshell Taxonomy is assigned by comparing your DNA sequences against a database of DNA sequences from known taxa Marker
Assigning Taxonomy to Marker Genes Susan Huse Brown University August 7, 2014 In a nutshell Taxonomy is assigned by comparing your DNA sequences against a database of DNA sequences from known taxa Marker
SUPPLEMENTARY INFORMATION
 Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
How volcanic heat affects biodiversity in Antarctica
 How volcanic heat affects biodiversity in Antarctica Report on field trip to Deception Island, February 2016, supported by Antarctic Science Ceridwen Fraser, Fenner School of Environment and Society, Australian
How volcanic heat affects biodiversity in Antarctica Report on field trip to Deception Island, February 2016, supported by Antarctic Science Ceridwen Fraser, Fenner School of Environment and Society, Australian
Ammonia-oxidizing Archaea in the Arctic Ocean and Antarctic coastal watersemi_
 Environmental Microbiology (2009) 11(9), 2434 2445 doi:10.1111/j.1462-2920.2009.01974.x Ammonia-oxidizing Archaea in the Arctic Ocean and Antarctic coastal watersemi_1974 2434..2445 Karen M. Kalanetra,
Environmental Microbiology (2009) 11(9), 2434 2445 doi:10.1111/j.1462-2920.2009.01974.x Ammonia-oxidizing Archaea in the Arctic Ocean and Antarctic coastal watersemi_1974 2434..2445 Karen M. Kalanetra,
Single-cell genomics applied to the picobiliphytes using next-generation sequencing
 Department of Ecology, Evolution and Natural Resources and Institute of Marine and Coastal Sciences Rutgers University, NJ 08901 Single-cell genomics applied to the picobiliphytes using next-generation
Department of Ecology, Evolution and Natural Resources and Institute of Marine and Coastal Sciences Rutgers University, NJ 08901 Single-cell genomics applied to the picobiliphytes using next-generation
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha
 Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Variations in pelagic bacterial communities in the North Atlantic Ocean coincide with water bodies
 The following supplement accompanies the article Variations in pelagic bacterial communities in the North Atlantic Ocean coincide with water bodies Richard L. Hahnke 1, Christina Probian 1, Bernhard M.
The following supplement accompanies the article Variations in pelagic bacterial communities in the North Atlantic Ocean coincide with water bodies Richard L. Hahnke 1, Christina Probian 1, Bernhard M.
The role of sea ice algae in Arctic marine ecosystems and oceanic emissions of dimethylsulfide. Hakase Hayashida Candidacy Exam August 19, 2015
 The role of sea ice algae in Arctic marine ecosystems and oceanic emissions of dimethylsulfide Hakase Hayashida Candidacy Exam August 19, 2015 Introduction: Role of sea ice algae in Arctic marine ecosystems
The role of sea ice algae in Arctic marine ecosystems and oceanic emissions of dimethylsulfide Hakase Hayashida Candidacy Exam August 19, 2015 Introduction: Role of sea ice algae in Arctic marine ecosystems
Why is there so much microbial diversity in NJ and Beyond? Rutgers University, Lee Kerkhof
 Institute of Marine and Coastal Sciences Why is there so much microbial diversity in NJ and Beyond? Rutgers University, Lee Kerkhof Overview of the seminar Background on how we do oceanography and a small
Institute of Marine and Coastal Sciences Why is there so much microbial diversity in NJ and Beyond? Rutgers University, Lee Kerkhof Overview of the seminar Background on how we do oceanography and a small
Size scaling deviation in phytoplankton photosynthesis and the energy flow through a
 ICES CM2004/Q:04 Size scaling deviation in phytoplankton photosynthesis and the energy flow through a coastal ecosystem. Pedro Cermeño, Emilio Marañón, Jaime Rodríguez, Emilio Fernández, Francisco Jiménez
ICES CM2004/Q:04 Size scaling deviation in phytoplankton photosynthesis and the energy flow through a coastal ecosystem. Pedro Cermeño, Emilio Marañón, Jaime Rodríguez, Emilio Fernández, Francisco Jiménez
Microbial Taxonomy and Phylogeny: Extending from rrnas to Genomes
 Microbial Taxonomy and Phylogeny: Extending from rrnas to Genomes Dr. Kostas Konstantinidis Department of Civil and Environmental Engineering & Department of Biology (Adjunct), Center for Bioinformatics
Microbial Taxonomy and Phylogeny: Extending from rrnas to Genomes Dr. Kostas Konstantinidis Department of Civil and Environmental Engineering & Department of Biology (Adjunct), Center for Bioinformatics
Package SPECIES. R topics documented: April 23, Type Package. Title Statistical package for species richness estimation. Version 1.
 Package SPECIES April 23, 2011 Type Package Title Statistical package for species richness estimation Version 1.0 Date 2010-01-24 Author Ji-Ping Wang, Maintainer Ji-Ping Wang
Package SPECIES April 23, 2011 Type Package Title Statistical package for species richness estimation Version 1.0 Date 2010-01-24 Author Ji-Ping Wang, Maintainer Ji-Ping Wang
The problem Lineage model Examples. The lineage model
 The lineage model A Bayesian approach to inferring community structure and evolutionary history from whole-genome metagenomic data Jack O Brien Bowdoin College with Daniel Falush and Xavier Didelot Cambridge,
The lineage model A Bayesian approach to inferring community structure and evolutionary history from whole-genome metagenomic data Jack O Brien Bowdoin College with Daniel Falush and Xavier Didelot Cambridge,
Seasonal cycle of phytoplankton community composition off Newport, Oregon, in 2009
 Seasonal cycle of phytoplankton community composition off Newport, Oregon, in 29 Xiuning Du 1, William Peterson 2 1 College of Environmental science and Engineering, Ocean University of China, Qingdao,
Seasonal cycle of phytoplankton community composition off Newport, Oregon, in 29 Xiuning Du 1, William Peterson 2 1 College of Environmental science and Engineering, Ocean University of China, Qingdao,
Silicate to Nitrate Ratio of the Upper Sub-Arctic Pacific and the Bering Sea Basin in Summer: Its Implication for Phytoplankton Dynamics
 Journal of Oceanography, Vol. 57, pp. 253 to 260, 2001 Silicate to Nitrate Ratio of the Upper Sub-Arctic Pacific and the Bering Sea Basin in Summer: Its Implication for Phytoplankton Dynamics ISAO KOIKE*,
Journal of Oceanography, Vol. 57, pp. 253 to 260, 2001 Silicate to Nitrate Ratio of the Upper Sub-Arctic Pacific and the Bering Sea Basin in Summer: Its Implication for Phytoplankton Dynamics ISAO KOIKE*,
Satellite-derived environmental drivers for top predator hotspots
 Satellite-derived environmental drivers for top predator hotspots Peter Miller @PeterM654 South West Marine Ecosystems 2017 21 Apr. 2017, Plymouth University Satellite environmental drivers for hotspots
Satellite-derived environmental drivers for top predator hotspots Peter Miller @PeterM654 South West Marine Ecosystems 2017 21 Apr. 2017, Plymouth University Satellite environmental drivers for hotspots
Bacteria, Friends or Foes?
 Bacteria, Friends or Foes? This unit integrates molecular biology techniques with the role of bacteria in our environment, specifically in the marine environment. The unit starts with introductory activities
Bacteria, Friends or Foes? This unit integrates molecular biology techniques with the role of bacteria in our environment, specifically in the marine environment. The unit starts with introductory activities
Supplementary File 4: Methods Summary and Supplementary Methods
 Supplementary File 4: Methods Summary and Supplementary Methods Methods Summary Constructing and implementing an SDM model requires local measurements of community composition and rasters of environmental
Supplementary File 4: Methods Summary and Supplementary Methods Methods Summary Constructing and implementing an SDM model requires local measurements of community composition and rasters of environmental
Taxonomy and Clustering of SSU rrna Tags. Susan Huse Josephine Bay Paul Center August 5, 2013
 Taxonomy and Clustering of SSU rrna Tags Susan Huse Josephine Bay Paul Center August 5, 2013 Primary Methods of Taxonomic Assignment Bayesian Kmer Matching RDP http://rdp.cme.msu.edu Wang, et al (2007)
Taxonomy and Clustering of SSU rrna Tags Susan Huse Josephine Bay Paul Center August 5, 2013 Primary Methods of Taxonomic Assignment Bayesian Kmer Matching RDP http://rdp.cme.msu.edu Wang, et al (2007)
SPECIES OF ARCHAEA ARE MORE CLOSELY RELATED TO EUKARYOTES THAN ARE SPECIES OF PROKARYOTES.
 THE TERMS RUN AND TUMBLE ARE GENERALLY ASSOCIATED WITH A) cell wall fluidity. B) cell membrane structures. C) taxic movements of the cell. D) clustering properties of certain rod-shaped bacteria. A MAJOR
THE TERMS RUN AND TUMBLE ARE GENERALLY ASSOCIATED WITH A) cell wall fluidity. B) cell membrane structures. C) taxic movements of the cell. D) clustering properties of certain rod-shaped bacteria. A MAJOR
Life on Earth Topic Test
 Life on Earth Topic Test Multiple Choice Questions Select the best alternative and indicate your response on the answer sheet. (1 mark each) 1. The list below contains common substances that exist in Earth
Life on Earth Topic Test Multiple Choice Questions Select the best alternative and indicate your response on the answer sheet. (1 mark each) 1. The list below contains common substances that exist in Earth
something about srna in archaea
 something about srna in archaea or: Processed Small RNAs in Archaea and BHB Elements Sarah Berkemer Bioinformatics Vienzig Archaea? Sarah Berkemer (Bioinformatics Vienzig) BHB elements in Archaea 2 / 23
something about srna in archaea or: Processed Small RNAs in Archaea and BHB Elements Sarah Berkemer Bioinformatics Vienzig Archaea? Sarah Berkemer (Bioinformatics Vienzig) BHB elements in Archaea 2 / 23
Catastrophic reduction of seaice in the Arctic Ocean - its impact on the marine ecosystems in the polar region-
 1/12 Catastrophic reduction of seaice in the Arctic Ocean - its impact on the marine ecosystems in the polar region- KAKENHI No.22221003 Naomi Harada (JAMSTEC) J. Onodera, E. Watanabe, K. Matsuno, K. Kimoto,
1/12 Catastrophic reduction of seaice in the Arctic Ocean - its impact on the marine ecosystems in the polar region- KAKENHI No.22221003 Naomi Harada (JAMSTEC) J. Onodera, E. Watanabe, K. Matsuno, K. Kimoto,
Marine Ecology I: Phytoplankton and Primary production
 Marine Ecology I: Phytoplankton and Primary production Osvaldo Ulloa University of Concepcion, Chile oulloa@profc.udec.cl From SOLAS Science Plan Phytoplankton, biogeochemistry and climate I Uptake (through
Marine Ecology I: Phytoplankton and Primary production Osvaldo Ulloa University of Concepcion, Chile oulloa@profc.udec.cl From SOLAS Science Plan Phytoplankton, biogeochemistry and climate I Uptake (through
Fluorometry Project Chlorophyll Temperature Time Series
 Fluorometry Project Ocean Institute + Scripps Institution of Oceanography Chlorophyll Temperature Time Series The California Current Long Term Ecological Research (CCE LTER) Phytoplankton Phytoplankton
Fluorometry Project Ocean Institute + Scripps Institution of Oceanography Chlorophyll Temperature Time Series The California Current Long Term Ecological Research (CCE LTER) Phytoplankton Phytoplankton
Latitudinal trends of Crenarchaeota and Bacteria in the meso- and bathypelagic water masses of the Eastern North Atlantic
 Environmental Microbiology (2008) 10(1), 110 124 doi:10.1111/j.1462-2920.2007.01437.x Latitudinal trends of Crenarchaeota and Bacteria in the meso- and bathypelagic water masses of the Eastern North Atlantic
Environmental Microbiology (2008) 10(1), 110 124 doi:10.1111/j.1462-2920.2007.01437.x Latitudinal trends of Crenarchaeota and Bacteria in the meso- and bathypelagic water masses of the Eastern North Atlantic
Alexander H. Frank, Juan A. Garcia, Gerhard J. Herndl, Thomas Reinthaler
 Supplementary Information Connectivity between surface and deep waters determines prokaryotic diversity in the North Atlantic Deep Water Alexander H. Frank, Juan A. Garcia, Gerhard J. Herndl, Thomas Reinthaler
Supplementary Information Connectivity between surface and deep waters determines prokaryotic diversity in the North Atlantic Deep Water Alexander H. Frank, Juan A. Garcia, Gerhard J. Herndl, Thomas Reinthaler
Supporting Information. Simple Bacterial Detection and High-Throughput Drug Screening. Based on Graphene-Enzyme Complex
 Supporting Information Simple Bacterial Detection and High-Throughput Drug Screening Based on Graphene-Enzyme Complex Juan-Li, Ling-Jie Wu, Shan-Shan Guo, Hua-E Fu, Guo-Nan Chen* and Huang-Hao Yang* The
Supporting Information Simple Bacterial Detection and High-Throughput Drug Screening Based on Graphene-Enzyme Complex Juan-Li, Ling-Jie Wu, Shan-Shan Guo, Hua-E Fu, Guo-Nan Chen* and Huang-Hao Yang* The
A. Incorrect! In the binomial naming convention the Kingdom is not part of the name.
 Microbiology Problem Drill 08: Classification of Microorganisms No. 1 of 10 1. In the binomial system of naming which term is always written in lowercase? (A) Kingdom (B) Domain (C) Genus (D) Specific
Microbiology Problem Drill 08: Classification of Microorganisms No. 1 of 10 1. In the binomial system of naming which term is always written in lowercase? (A) Kingdom (B) Domain (C) Genus (D) Specific
Observation system for early warning of HAB events
 Observation system for early warning of HAB events Vera L. Trainer, NOAA Fisheries Northwest Fisheries Science Center Marine Biotoxins Program Seattle, Washington, USA Juan de Fuca eddy Regional HAB OOS
Observation system for early warning of HAB events Vera L. Trainer, NOAA Fisheries Northwest Fisheries Science Center Marine Biotoxins Program Seattle, Washington, USA Juan de Fuca eddy Regional HAB OOS
Life in an unusual intracellular niche a bacterial symbiont infecting the nucleus of amoebae
 Life in an unusual intracellular niche a bacterial symbiont infecting the nucleus of amoebae Frederik Schulz, Ilias Lagkouvardos, Florian Wascher, Karin Aistleitner, Rok Kostanjšek, Matthias Horn Supplementary
Life in an unusual intracellular niche a bacterial symbiont infecting the nucleus of amoebae Frederik Schulz, Ilias Lagkouvardos, Florian Wascher, Karin Aistleitner, Rok Kostanjšek, Matthias Horn Supplementary
ATOC OUR CHANGING ENVIRONMENT Class 19 (Chp 6) Objectives of Today s Class: The Cryosphere [1] Components, time scales; [2] Seasonal snow
![ATOC OUR CHANGING ENVIRONMENT Class 19 (Chp 6) Objectives of Today s Class: The Cryosphere [1] Components, time scales; [2] Seasonal snow ATOC OUR CHANGING ENVIRONMENT Class 19 (Chp 6) Objectives of Today s Class: The Cryosphere [1] Components, time scales; [2] Seasonal snow](/thumbs/95/126309683.jpg) ATOC 1060-002 OUR CHANGING ENVIRONMENT Class 19 (Chp 6) Objectives of Today s Class: The Cryosphere [1] Components, time scales; [2] Seasonal snow cover, permafrost, river and lake ice, ; [3]Glaciers and
ATOC 1060-002 OUR CHANGING ENVIRONMENT Class 19 (Chp 6) Objectives of Today s Class: The Cryosphere [1] Components, time scales; [2] Seasonal snow cover, permafrost, river and lake ice, ; [3]Glaciers and
CHAPTER 6 & 7 VOCABULARY
 CHAPTER 6 & 7 VOCABULARY 1. Biome 2. Climate 3. Latitude 4. Altitude 5. Emergent layer 6. Epiphyte 7. Understory 8. Permafrost 9. Wetland 10.Plankton 11.Nekton 12.Benthos 13.Littoral zone 14.Benthic zone
CHAPTER 6 & 7 VOCABULARY 1. Biome 2. Climate 3. Latitude 4. Altitude 5. Emergent layer 6. Epiphyte 7. Understory 8. Permafrost 9. Wetland 10.Plankton 11.Nekton 12.Benthos 13.Littoral zone 14.Benthic zone
Effects of Gap Open and Gap Extension Penalties
 Brigham Young University BYU ScholarsArchive All Faculty Publications 200-10-01 Effects of Gap Open and Gap Extension Penalties Hyrum Carroll hyrumcarroll@gmail.com Mark J. Clement clement@cs.byu.edu See
Brigham Young University BYU ScholarsArchive All Faculty Publications 200-10-01 Effects of Gap Open and Gap Extension Penalties Hyrum Carroll hyrumcarroll@gmail.com Mark J. Clement clement@cs.byu.edu See
Censusing the Sea in the 21 st Century
 Censusing the Sea in the 21 st Century Nancy Knowlton & Matthieu Leray Photo: Ove Hoegh-Guldberg Smithsonian s National Museum of Natural History Estimates of Marine/Reef Species Numbers (Millions) Marine
Censusing the Sea in the 21 st Century Nancy Knowlton & Matthieu Leray Photo: Ove Hoegh-Guldberg Smithsonian s National Museum of Natural History Estimates of Marine/Reef Species Numbers (Millions) Marine
Microbial diversity pa1erns in rela3on to hydrocarbon seepage
 Microbial diversity pa1erns in rela3on to hydrocarbon seepage I. Microbial mat pushcores II. Sediment gravity cores The MC118 subsurface microbial community microbial players within and outside of subsurface
Microbial diversity pa1erns in rela3on to hydrocarbon seepage I. Microbial mat pushcores II. Sediment gravity cores The MC118 subsurface microbial community microbial players within and outside of subsurface
Physiography Ocean Provinces p. 1 Dimensions p. 1 Physiographic Provinces p. 2 Continental Margin Province p. 2 Deep-Ocean Basin Province p.
 Physiography Ocean Provinces p. 1 Dimensions p. 1 Physiographic Provinces p. 2 Continental Margin Province p. 2 Deep-Ocean Basin Province p. 2 Mid-Ocean Ridge Province p. 3 Benthic and Pelagic Provinces
Physiography Ocean Provinces p. 1 Dimensions p. 1 Physiographic Provinces p. 2 Continental Margin Province p. 2 Deep-Ocean Basin Province p. 2 Mid-Ocean Ridge Province p. 3 Benthic and Pelagic Provinces
Supporting Information
 Supporting Information Farrant et al. 1.173/pnas.1524865113 Prochlorococcus % % 8% % % 5% 4% 3% 2% 1% % A Assignment level: Phylum Genus SubCluster Clade Synechococcus % % 8% % % 5% 4% 3% 2% 1% % 1-26-125
Supporting Information Farrant et al. 1.173/pnas.1524865113 Prochlorococcus % % 8% % % 5% 4% 3% 2% 1% % A Assignment level: Phylum Genus SubCluster Clade Synechococcus % % 8% % % 5% 4% 3% 2% 1% % 1-26-125
Diversity of Halophilic Archaea from Ezzemoul Sabkha in Algeria
 Óbuda University e-bulletin Vol. 5, No. 1, 2015 Diversity of Halophilic Archaea from Ezzemoul Sabkha in Algeria Karima Kharroub 1, Amine Mohamed Gomri 1, Mercedes Monteoliva-Sanchez 2 1 Department of Biotechnology/
Óbuda University e-bulletin Vol. 5, No. 1, 2015 Diversity of Halophilic Archaea from Ezzemoul Sabkha in Algeria Karima Kharroub 1, Amine Mohamed Gomri 1, Mercedes Monteoliva-Sanchez 2 1 Department of Biotechnology/
Other resources. Greengenes (bacterial) Silva (bacteria, archaeal and eukarya)
 General QIIME resources http://qiime.org/ Blog (news, updates): http://qiime.wordpress.com/ Support/forum: https://groups.google.com/forum/#!forum/qiimeforum Citing QIIME: Caporaso, J.G. et al., QIIME
General QIIME resources http://qiime.org/ Blog (news, updates): http://qiime.wordpress.com/ Support/forum: https://groups.google.com/forum/#!forum/qiimeforum Citing QIIME: Caporaso, J.G. et al., QIIME
Supplementary Information
 Supplementary Information Supplementary Figure 1. Schematic pipeline for single-cell genome assembly, cleaning and annotation. a. The assembly process was optimized to account for multiple cells putatively
Supplementary Information Supplementary Figure 1. Schematic pipeline for single-cell genome assembly, cleaning and annotation. a. The assembly process was optimized to account for multiple cells putatively
Impact of changing Siberian land-shelf-basin on the Arctic Ocean biogeochemical dynamics
 Impact of changing Siberian land-shelf-basin on the Arctic Ocean biogeochemical dynamics PI: Shigeto Nishino (JAMSTEC) Co-PI: Igor Semiletov (IARC/UAF) Other Investigators: Natalia Shakhova (IARC/UAF),
Impact of changing Siberian land-shelf-basin on the Arctic Ocean biogeochemical dynamics PI: Shigeto Nishino (JAMSTEC) Co-PI: Igor Semiletov (IARC/UAF) Other Investigators: Natalia Shakhova (IARC/UAF),
Correction to Evaluation of the simulation of the annual cycle of Arctic and Antarctic sea ice coverages by 11 major global climate models
 JOURNAL OF GEOPHYSICAL RESEARCH, VOL. 111,, doi:10.1029/2006jc003949, 2006 Correction to Evaluation of the simulation of the annual cycle of Arctic and Antarctic sea ice coverages by 11 major global climate
JOURNAL OF GEOPHYSICAL RESEARCH, VOL. 111,, doi:10.1029/2006jc003949, 2006 Correction to Evaluation of the simulation of the annual cycle of Arctic and Antarctic sea ice coverages by 11 major global climate
Time-series observations in the Northern Indian Ocean V.V.S.S. Sarma National Institute of Oceanography Visakhapatnam, India
 The Second GEOSS Asia-Pacific Symposium, Tokyo, 14-16 th April 28 Time-series observations in the Northern Indian Ocean V.V.S.S. Sarma National Institute of Oceanography Visakhapatnam, India Seasonal variations
The Second GEOSS Asia-Pacific Symposium, Tokyo, 14-16 th April 28 Time-series observations in the Northern Indian Ocean V.V.S.S. Sarma National Institute of Oceanography Visakhapatnam, India Seasonal variations
Physical Oceanography
 Physical Oceanography SECTION 15.1 The Oceans In your textbook, read about modern oceanography. For each item in Column A, write the letter of the matching item in Column B. Column A 1. German research
Physical Oceanography SECTION 15.1 The Oceans In your textbook, read about modern oceanography. For each item in Column A, write the letter of the matching item in Column B. Column A 1. German research
Progress report on development of a spatially explicit operating model for tropical tuna populations.
 IOTC-2018-WPTT20-27 Progress report on development of a spatially explicit operating model for tropical tuna populations. Prepared for Indian Ocean Tuna Commission September 2018 Prepared by: Simon Hoyle
IOTC-2018-WPTT20-27 Progress report on development of a spatially explicit operating model for tropical tuna populations. Prepared for Indian Ocean Tuna Commission September 2018 Prepared by: Simon Hoyle
Research Programme Polar, Marine and Coastal Systems. Current and future Arctic research priorities of Germany Nicole Biebow, AWI
 Research Programme Polar, Marine and Coastal Systems Current and future Arctic research priorities of Germany Nicole Biebow, AWI Arctic Science and Technology (S&T) Collaboration and Engagement Workshop,
Research Programme Polar, Marine and Coastal Systems Current and future Arctic research priorities of Germany Nicole Biebow, AWI Arctic Science and Technology (S&T) Collaboration and Engagement Workshop,
State of the Ocean 2003: Physical Oceanographic Conditions in the Gulf of St. Lawrence
 Ecosystem Status Report 24/2 Oceanographic sampling gear State of the Ocean 23: Physical Oceanographic Conditions in the Gulf of St. Lawrence Background The physical oceanographic environment influences
Ecosystem Status Report 24/2 Oceanographic sampling gear State of the Ocean 23: Physical Oceanographic Conditions in the Gulf of St. Lawrence Background The physical oceanographic environment influences
Vertical distribution and temporal variation of marine planktonic archaea in the Gerlache Strait, Antarctica, during early spring
 Vertical distribution and temporal variation of marine planktonic archaea in the Gerlache Strait, Antarctica, during early spring Ramon Massana, Lance T. Taylor, Alison E. Murray, Ke Y. Wu, Wade H. Jefsyey,
Vertical distribution and temporal variation of marine planktonic archaea in the Gerlache Strait, Antarctica, during early spring Ramon Massana, Lance T. Taylor, Alison E. Murray, Ke Y. Wu, Wade H. Jefsyey,
53 contributors for 35 individual reports in 2009 show 5% of figures today
 A Group Approach to Understanding Ecosystem Dynamics in the Northeast Pacific Ocean William Crawford and James Irvine, Fisheries and Oceans Canada (DFO) * * * 53 contributors for 35 individual reports
A Group Approach to Understanding Ecosystem Dynamics in the Northeast Pacific Ocean William Crawford and James Irvine, Fisheries and Oceans Canada (DFO) * * * 53 contributors for 35 individual reports
(review) Organization of life
 Marine life: the plankton Production & Energy Transfer Part of Chapter 12, Chapter 13 (review) Organization of life Prokaryotes (usually no nucleus simple life forms) Domain Archaea: : most are extremophiles
Marine life: the plankton Production & Energy Transfer Part of Chapter 12, Chapter 13 (review) Organization of life Prokaryotes (usually no nucleus simple life forms) Domain Archaea: : most are extremophiles
