Feedback Loops Regulates Cdc42p Activity during Spontaneous Cell Polarization
|
|
|
- Noel Gallagher
- 5 years ago
- Views:
Transcription
1 Developmental Cell, Vol. 9, , October, 005, Copyright 005 by Elsevier Inc. DOI /j.devcel A System of Counteracting Short Article Feedback Loops Regulates Cdc4p Activity during Spontaneous Cell Polarization Ertugrul M. Ozbudak, 1 Attila Becskei, 1 and Alexander van Oudenaarden 1, * Department of Physics Massachusetts Institute of Technology Cambridge, Massachusetts 0139 Summary Cellular polarization is often a response to distinct extracellular or intracellular cues, such as nutrient gradients or cortical landmarks. However, in the absence of such cues, some cells can still select a polarization axis at random. Positive feedback loops promoting localized activation of the GTPase Cdc4p are central to this process in budding yeast. Here, we explore spontaneous polarization during bud site selection in mutant yeast cells that lack functional landmarks. We find that these cells do not select a single random polarization axis, but continuously change this axis during the G1 phase of the cell cycle. This is reflected in traveling waves of activated Cdc4p which randomly explore the cell periphery. Our integrated computational and in vivo analyses of these waves reveal a negative feedback loop that competes with the aforementioned positive feedback loops to regulate Cdc4p activity and confer dynamic responsiveness on the robust initiation of cell polarization. Introduction Establishment of cell polarity is an essential process both in uni- and multicellular organisms during cell division, chemotaxis, differentiation, morphogenesis, and cell migration (Nelson, 003). In general, the axis of cell polarity is selected either by cell extrinsic stimuli or intrinsic cues. The role of intrinsic cues is exemplified by the selection of the bud site during the cell division cycle of the yeast Saccharomyces cerevisiae. In this case, a new bud forms where landmark proteins were deposited during the previous cell division cycle. The Ras GTPase Rsr1p (Bud1p) landmark protein recruits the guanosine-nucleotide exchange factor (GEF) Cdc4p to the plasma membrane (Toenjes et al., 004), resulting in a local activation of Cdc4p (Kozminski et al., 003; Park et al., 1997). Subsequently, activated Cdc4p recruits its effectors forming a crescent-shaped polar cap on the cell periphery, which orchestrates rearrangements of the actin cytoskeleton and polarized secretion to initiate budding. Cells lacking functional landmarks initiate budding at a random location (Chant, 1999) by spontaneous symmetry breaking (Wedlich-Soldner et al., 003, 004; Irazoqui et al., 003). This reflects the efficiency of polarization mechanisms to amplify minute asymmetries in the absence of any spatial cues. In general, this may be accomplished *Correspondence: avano@mit.edu 1 Lab address: by positive feedback regulation of a polarization activator that is initially randomly distributed at the plasma membrane. Theoretical studies show that if activation requires a limiting diffusible substrate, an initially random activator distribution has the potential to self-organize into a highly polarized distribution characterized by a single, stable, and randomly oriented polarization axis (Gierer and Meinhardt, 197). Indeed, recent experimental studies suggest that the regulation of Cdc4p activity is part of a positive feedback loop (Wedlich- Soldner et al., 003; Irazoqui et al., 003). Potential molecular candidates for this positive feedback include actin-mediated transport of secretory vesicles containing Cdc4p to sites of enhanced activity (Wedlich- Soldner et al., 003) and a Cdc4p-induced local polymerization of scaffolding proteins at the plasma membrane that enhances the local Cdc4p recruitment rate (Irazoqui et al., 003). This symmetry breaking requires the scaffold protein Bem1p (Irazoqui et al., 003) and the cycling of Cdc4p between GTP and GDP-bound forms (Caviston et al., 00), which is catalyzed by the Cdc4p GEF and various Cdc4p-specific GTPase activating proteins (GAPs) including Rga1p and Bem3p. Here, we study the dynamics of this symmetry breaking in yeast cells with RSR1 deleted. In these cells lacking functional landmarks, we monitor the pool of activated Cdc4p at the plasma membrane. We find that cells in the early G1 phase randomly select a polarization axis. However, this axis is not static but gradually wanders during the rest of the G1 phase. This is experimentally reflected in traveling activity waves of Cdc4p that often traverse the circumference of a cell more than once before budding. This dynamic behavior cannot be explained by the positive feedback regulation of Cdc4p activity alone. We propose that negative feedback regulation of Cdc4p activity mediated through the actin cytoskeleton in parallel to a positive feedback provides the drive of the traveling activity waves. Consistently, the wave mobility is significantly reduced when the actin cytoskeleton is depolymerized or when the concentration of Cdc4p activity inhibitors (GAPs) is reduced. Results and Discussion Cdc4 Activity Is Organized into Spatial Waves In order to monitor the polarization dynamics of activated Cdc4p, a CRIB Gic -GFP fusion protein was monitored using fluorescence microscopy. The CRIB domain of Gicp, CRIB Gic, binds to the GTP-bound form of Cdc4p (Burbelo et al., 1995). This probe is expected to respond rapidly to changes in the Cdc4p-GTP location because photobleached areas of polar caps recovered their initial fluorescence intensity in less than one second (Figure S1). This response is considerably faster than the exchange rate of Cdc4p between the polar cap and the cytoplasm (Wedlich-Soldner et al., 004). Time-lapse experiments showed that, in wildtype cells, the location of the polar cap did not change
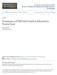
2 Developmental Cell 566 Figure 1. Dynamics of the Polar Cap Movement (A) Gicp (1 08) -GFP is used as a reporter for activated Cdc4p. Localized fluorescence signal in wild-type (strain ERT4.1; upper panel) or rsr1 (strain ERT5.1; lower panel) cells. Time is reported in minutes. (B) Bem1p-CFP localization is shown in an rsr1 (strain ERT73.) cell. Time is reported in minutes. (C) Time evolution of the polarization angle in a wild-type (open circles) or rsr1 (solid red circles) cell. The polarization angle is defined as the angle difference between the current and incipient polarization site. after the initial establishment of cellular polarity, although the cap displayed small displacements. Eventually, budding started adjacent to the previous division site (Figure 1A, upper panel; Movie S1). Similar behavior was observed in bem1d cells. This is expected because in these cells the budding landmark is fully functional and cells bud in a nonrandom, monopolar fashion. If dynamic polarization in the absence of landmark protein Rsr1p (Bud1p) is solely determined by autoamplification of fluctuations in Cdc4p regulators, we expected that a polar cap would be initiated at a random location followed by budding at this same location. Indeed, the polar cap formed at a single site, but contrary to the above expectations, it started to move around the cell periphery immediately after symmetry breaking. The direction of movement changed at random time intervals. Occasionally, polar caps traveled over the entire cell perimeter before settling down at a random location to initiate budding (Figure 1A, lower panel; Movies S and S3). Similar behavior was observed when the location of the polar cap components was monitored by Bem1p-CFP (Figure 1B), Gicp-GFP, and CRIB Cla4 -GFP (data not shown). The interval between the appearance of the polar cap and the inception of budding at room temperature is (60 ± 13) min (N = 13). We find that the wandering stops about 15 0 min before we can detect the appearance of the bud in the phase contrast image. The motion of traveling polar caps was quantified by their angular displacement with respect to the initial polarization site at the start of the experiment. Analysis of the polarization angle indicates that there is a significant difference in mobility of the polar caps between wild-type and rsr1 cells over the entire G1 stage of the cell cycle (Figure 1C). The mobility of the traveling activity waves was quantified by plotting the mean square angular displacement of polar caps q RMS as a function of time, averaged over multiple cells (Saxton and Jacobson, 1997). The linear dependence of q RMS with time for rsr1d cells reveals that traveling waves perform a random unconstrained walk at the cell periphery (Figure A). Deletion of other landmark proteins, such as Budp and Bud5p, also resulted in the appearance of traveling waves. The mobility of the polar caps as measured by the slope of the q RMS -time relation showed similar values for rsr1, bud, and bud5 cells (Figure 3A). This indicates that cycling between the GTP and GDP-bound state of Rsr1p is necessary to prevent the polar cap from traveling, because cells expressing constitutively active and inactive Rsr1p (bud and bud5 cells, respectively) display highly mobile waves. Dosage Dependence of Wave Mobility on Localizing Factors The above results suggest the existence of patterndestabilizing processes, which become dominant in the absence of localizing factors such as Rsr1p. Next, we reduced the Rsr1p expression level below the wild-type level to attain an intermediate level of polarizing strength and to moderate the effect of pattern destabilization. For this purpose, the weak P IXR1 and P GLN3 promoters (Holland, 00) were used to drive to expression of RSR1. The mobility of the polar cap was reduced in comparison to rsr1 cells as the concentration of Rsr1p increased (Figure A). When RSR1 is driven by the weak P GLN3 promoter, the increase of q RMS slowed down with time, indicative of confined random motion of the polar cap (Saxton and Jacobson, 1997). This is consistent with the model that large fluctuations in pattern destabilization cause the polar cap to escape from the area of activated Rsr1p localization at low expression level. However, on average, the polar cap spends more time in the area of landmark proteins. In order to examine how the wave mobility affects bud site selection, we measured the budding angle. The budding angle reflects the displacement of the settled, late G1 localization of the polar cap with respect to the previous budding site (see Experimental Procedures). The average budding angle correlates nonlinearly with the wave mo-
3 Counteracting Feedback Loops Control Cdc4 Activity 567 Figure. The Root Mean Square of the Angular Displacement θ RMS of the Polar Cap Quantifies Wave Mobility θ RMS is plotted as a function of time with respect to the initial polarization angle at the start of the observation. θ RMS was obtained by averaging over at least ten different cells. The average slope, obtained by a linear least-square fit, of θ RMS-time relation defines the wave mobility. (A) Dosage-dependent effect of RSR1 expression on the motility of the polar cap shown for wild-type (strain ERT4.1; black squares), rsr1 (strain ERT5.1; red circles), rsr1 /P GLN3 RSR1 (strain ERT75.; green triangles), and rsr1 /P IXR1 RSR1 (strain ERT76.3; green triangles) cells. (B) Dosage-dependent effect of GAP expression on the motility of the polar cap for wild-type (strain ERT4.1; black squares), rsr1 (strain ERT5.1; red circles), rsr1 bem3 /P POL30 RGA1 (strain ERT8.3; blue triangles), rsr1 bem3 (strain ERT54.1; purple stars), and rsr1 rga1 (strain ERT55.1; green triangles) cells. bility (Figure 3B), revealing that confinement of waves results in a more deterministic bud site selection. Regulators of Cdc4p GTPase Activity Determine the Mobility of Waves Whereas landmark proteins localize the traveling waves, activities that enhance the mobility of the waves are likely to be related to restructuring of the polar cap. Cycling of Cdc4p between GTP and GDP-bound forms is crucial for recruitment of polar cap components (Irazoqui et al., 003, 004). Inhomogeneous recruitment of Cdc4p regulators within the polar cap may lead to a local decrease in the Cdc4p turnover and, consequently, the net dissociation rate of polar cap components would increase. As a result, the restructuring and mobility of the polar cap would be reduced. The spatial distribution of Cdc4p and its regulators bound to secretory or endocytotic vesicles is in part dependent on actin-based transport (Wedlich-Soldner et al., 004; Irazoqui et al., 005). Therefore, we hypothesized that impairment of actin polymerization or changing the turnover of the Cdc4p GTPase cycle would decrease the wave mobility. First, inhibition of actin polymerization by latrunculin A abolished wave mobility in rsr1 cells (Figure 3A). While initial pattern formation is independent of actin even in the absence of landmark proteins (Ayscough et al., 1997; Irazoqui et al., 003; Figure S), the mobility and the dynamic nature of polar caps is actin-dependent. The wave mobility in latrunculin A-treated rsr1 cells is very similar to that observed for wild-type cells. We do not observe the alternating appearance and disappearance of the polar cap as was observed when Cdc4p-GFP was used as a reporter in latrunculin A-treated cells (Wedlich-Soldner et al., 004). This difference might be attributed to use of a different reporter or different expression levels of Cdc4p or Rsr1p in these strains. Second, the turnover of the Cdc4p GTPase cycle was impaired by deleting BEM3, a gene encoding a GAP protein for Cdc4p. This deletion reduced the wave mobility substantially in rsr1 cells (Figures B and 3A). To verify whether this phenotype was specific to the deletion of BEM3 or whether it reflects the reduction of GAP activity in general, another GAP gene, RGA1, was deleted. The effect of this deletion was comparable to that observed for the BEM3 deletion. In addition, overexpression of Rga1p in rsr1 bem3 cells restored the mobility of the polar cap (Figures B and 3A). When BEM3 is overexpressed in rsr1 cells, the wave mobility does not change appreciably, indicating that the wave mobility in rsr1 cells is close to its maximum saturated value (Figure 3A). This might indicate that there is another limiting factor that determines the maximum recruitment rate of the GAPs to the polarization site. These data suggest that wave mobility depends on the dosage of GAP activity. Third, overexpression of a cell membrane-targeted form of the GEF Cdc4p also resulted in a decreased mobility (Figure 3A) of a more widely distributed polar cap (Supplemental Data). A Model of Feedback Regulation of Cdc4p Activity Explains Wave Formation The formation of Cdc4p activity waves can be modeled in terms of a feedback control of activators and
4 Developmental Cell 568 Figure 3. Regulation of Wave Mobility and Effect on the Budding Angle (A) Wave mobility (units of deg /min) of the polar cap in cells having different genetic backgrounds. The same strains were used as described in Figure. The last two bars were obtained from cells overexpressing myrga-cdc4 (strain ERT36.1) and myr-cdc4 (strain ERT37.1). (B) The wave mobility nonlinearly correlates with the average budding angle of cells expressing varying amounts of landmark proteins. In wild-type cells, new buds are excluded from the previous budding sites. This results in a minimum budding angle of around 40 for wild-type cells. An angle of about 10 corresponds to random budding. inhibitors of Cdc4p (Figure 4A; Supplemental Data). Interactions between Bem1p, Cdc4p, and Cdc4p are suggested to form an actin-independent, positive feedback loop that enhances the recruitment of activators to the polar cap (Bose et al., 001; Butty et al., 00; Irazoqui et al., 003; Wedlich-Soldner et al., 003, 004; Shimada et al., 004). We hypothesize that this positive feedback loop accounts for the initial symmetry breaking (Figure S). A second, actin-mediated, positive feedback could be mediated by the actin-based delivery of secretory vesicles containing, for example, Cdc4p or Cdc4p (Wedlich-Soldner et al., 003, 004). Indeed, we sometimes observe faint Gicp GFPlabeled dots moving toward the polar cap (Figure S4). Positive feedback loops allow an initially homogenous system to polarize in a random static direction (Gierer and Meinhardt, 197). Because there is a limited number of activated Cdc4p molecules distributed along the membrane, a uniform distribution will still exhibit small local concentration deviations about the average. Due to the positive feedback regulation, local concentration maximums will grow at the expense of the surrounding areas in the membrane. This mechanism can explain the initial symmetry breaking in, for example, latrunculin-treated cells (Figure 4B, middle panel), but cannot explain the traveling Cdc4p activity waves. We therefore propose that in addition to the positive regulation, a negative feedback loop must exist. Because we do not observe waves in latrunculin-treated cells, it is likely that the negative feedback loop is mediated by the actin cytoskeleton. A potential molecular implementation of the negative loop might be the delivery of vesicles containing GAPs along the actin cables. Alternatively, recent experiments show that actindependent endocytosis might disperse polarizing factors (Irazoqui et al., 005). Because actin patch components (associated with endocytosis) can be recruited by Cdc4p-GTP (Lechler et al., 000), this indicates that Cdc4p could stimulate dispersal of polarized factors in an actin-dependent manner, effectively establishing a negative feedback loop. The negative feedback loop provides the system with the potential to exhibit traveling waves (Meinhardt, 1999). To observe traveling waves, it is important that the negative feedback regulation operates at a slower rate than the positive regulation. We propose that the actin-independent positive feedback reacts faster to a change in Cdc4p activation state than the regulation mediated by the actin cytoskeleton. This is a reasonable assumption because nucleation and polymerization of new actin cables followed by vesicle transport might be a slower process than the formation of the Bem1p scaffolding complex that relies on relatively fast protein-protein interactions and rapid diffusion. We propose that the waves observed in rsr1 cells are a result of a competition between a fast, actinindependent, positive feedback regulation and a slow and therefore delayed, negative feedback regulation.
5 Counteracting Feedback Loops Control Cdc4 Activity 569 the absence of the cytoskeleton, only one positive feedback loop remains, explaining the absence of waves in latrunculin-treated cells (Figure 4B, middle panel). The proposed feedback system (Figure 4A) therefore qualitatively explains all the experimental data. A numerical model that explicitly models the dynamics in the different mutants is presented in Supplemental Data. The above model is likely to account for cellular polarization in other contexts, for example during cell shape changes upon exposure of yeast cells to pheromones. Interestingly, wandering polar caps were also observed in pheromone-treated mutant strains that lack landmark proteins and are defective in chemotropism (Nern and Arkowitz, 000). Conclusion Figure 4. Hypothetical Model of Wave Formation of Cdc4p Regulators (A) Three feedback loops regulate the recruitment of Cdc4p regulators: an actin-independent positive feedback loop (left), an actinmediated positive feedback loop (middle), and an actin-mediated negative feedback loop (right). (B) Initiation of waves (indicated by gray arrow) is expected in cells for which the actin-mediated negative feedback dominates the actin-mediated positive feedback (left). This net negative feedback can destabilize the polar cap and initiate cap motion. No wave mobility is expected when the positive feedback loops dominate. This can be achieved in the absence of the actin cytoskeleton (+lat A, middle) or in strains in which the GAP activity is reduced or the Cdc4p activity is enhanced (gap or GEF+, right). More details on the model are given in Supplemental Data. Because the inactivation always lags behind the spreading activation, this induces a wave motion (Meinhardt, 1999). This implies that in rsr1 cells, the actin-mediated negative feedback dominates the actin-mediated positive feedback. Wandering of the polar cap was not observed in earlier experiments in which the position of the polar cap was monitored by using a GFP fusion of wild-type Cdc4p or a constitutively activated version of Cdc4p (Wedlich-Soldner et al., 003, 004). The strains used in these experiments carried a functional RSR1 gene and expressed higher levels of activated Cdc4p compared to wild-type. This elevated Cdc4p activity leads to a stronger actin-mediated positive feedback, which might explain the absence of Cdc4p activity waves in these strains. In rsr1 cells, the effective feedback systems consist of one actin-independent positive feedback and one actin-mediated negative feedback (Figure 4B, left panel). The opposite holds for rsr1 cells in which the GAP activity has been reduced or the GEF activity has been increased. In these cells, the positive, actin-mediated loop dominates the negative feedback and therefore the feedback structure consists effectively of two parallel positive loops that allow symmetry breaking but do not support wave mobility (Figure 4B, right panel). In Cdc4 is essential for chemotaxis in higher eukaryotes such as Dictyostelium and neutrophils (Watanabe et al., 004; Weiner et al., 00). In these cells, interlocked positive and negative feedback loops have been identified among Cdc4, its regulators, and the actin network (Meili and Firtel, 003). Similar arguments that explain the Cdc4p activity waves in budding yeast might apply to the rotating pseudopod waves observed in Dictyostelium (Killich et al., 1993, 1994) or the mincde oscillatory waves in Escherichia coli (Meinhardt and de Boer, 001; Huang et al., 003). Mechanisms responsible for symmetry breaking are inherently coupled to mechanisms that enable restructuring of patterns. A design of competing feedback regulation loops combines efficient polarization with the ability to dynamically respond to varying intracellular or environmental conditions. Experimental Procedures Construction of Plasmids and Strains The P GLN3,P IXR1,P MYO, and P POL30 promoter sequences correspond to the 600, 858, 677, and 600 base pair regions upstream of the start codon of the respective genes. P TET0 corresponds to the CYC1 TATA region with two upstream rtta binding sites. The T RSR1, T CDC4, and T CYC1 terminator sequences correspond to the 596, 373, and 04(187) base pair regions downstream of the stop codon of the respective genes. BamHI-RGA1 (1 60 stop codon)-noti fragment was cloned into prs401 backbone downstream of P POL30.A sequence encoding the MGCTVSTQTIGDESDP myristoylation signal or a mutated signal sequence (GA) was fused to the N terminus of CDC4 by PCR. The resulting BamHI-myr-CDC4-T CDC4 - NotI and BamHI-myr(GA)-CDC4-T CDC4 -NotI constructs were cloned into prs316 and prs401 backbones downstream of P POL30. KpnI-P MYO -rtta-t CYC1 -NotI was cloned into prs305 and prs401 backbones. KpnI-P GLN3 -RSR1-T RSR1 -NotI and KpnI-P IXR1 -RSR1- T RSR1 -NotI was cloned into prs401 backbone. XhoI-P TETO - (BamHI/BglII)-GIC (1 08) -(BglII/BamHI)-GFP-T CYC1 -NotI was cloned into prs306 backbone. All strains are derivatives of BY4741 (MATa his3 1 leu 0 met15 0 ura3 0). The strain list is given in Table S1. Plasmid MJ79 was a gift from M. Peter. Growth Conditions and Data Acquisition Yeast cells were grown overnight at 30 C in synthetic selective dextrose media. Expression of GIC(1 08)GFP was induced overnight by doxycycline addition (50 g/l) into the growth media. Cells were harvested at exponential growth. For time-lapse experiments, slides were covered with 1% agarose containing synthetic media supplemented with % glucose. Cells were spun down and resuspended in the same media and placed on top of a thin agar layer. Fluorescence signals of single cells were measured using a Nikon
6 Developmental Cell 570 TE000 microscope (Microvideo Instruments, Avon, MA) with automated stage and a cooled back-thinned CCD camera (Micromax; Roper Scientific, Duluth, GA). Imaging was performed at room temperature (T = C). Images were collected every 60 s to follow the movement of the polar cap. The site of maximum fluorescence intensity in the polar cap was marked as the center of the cap. The angle between the initial and subsequent centers of the moving polar cap was calculated using Metamorph (Universal Imaging; Research Precision Instruments, Natick, MA). On average, 30 time series were obtained to calculate the q RMS in each genetic background. The wave mobility was calculated from fitting the q RMS time plot between t = 0 and 10 min using a linear least square fit. Cells in the G1 stage had polar caps wandering around the cell periphery which stopped around 15 0 min before the onset of budding (S phase). In addition, a small fraction of cells in G1 stage (<0%) had polar caps which either did not move or the movement paused for a longer time period (around 0 5 min). Some of these cells did not bud at all, which may reflect that they entered a quiescent state. For calculating the mobility coefficients, cells were considered in which the polar caps did not stop or pause. For measuring the budding angle, cells were stained with ConA (Lew and Reed, 1993). The angle between a small bud and the birth scar in daughter and young mother cells was measured. Both rsr1 bem3 and rsr1 rga1 cells continued budding randomly like rsr1 cells, as assayed by budding angle (data not shown), although the stability of the position of the polar cap in these cells was similar to that in wild-type cells. For actin depolymerization, cells were treated with 100 M latrunculin A for 10 min. Cells suspended in media containing latrunculin A were placed on a thin agar layer and microscopy was performed as described above. For FRAP experiments, a small region in the middle of the polar cap was bleached and fluorescence recovery was followed by confocal microscopy. Supplemental Data Supplemental data including a computer simulation, discussion, figures, table, and movies are available at com/cgi/content/full/9/4/565/dc1/. Acknowledgments We thank J. Chabot for technical assistance, B. Tam for help with confocal microscopy, D. Pellman, J. Pedraza, and S. Oliferenko for helpful suggestions and critical reading of the manuscript, and M. Peter, D. Lew, E. Bi, and D. Pellman for kindly providing strains and plasmids. A.B. is a Long-Term Fellow of the Human Frontier Science Program. This work was supported by NSF-CAREER (PHY ) and NIH (GM068957) grants. Received: February 1, 005 Revised: August 3, 005 Accepted: August 9, 005 Published: October 3, 005 References Ayscough, K.R., Stryker, J., Pokala, N., Sanders, M., Crews, P., and Drubin, D.G. (1997). High rates of actin filament turnover in budding yeast and roles for actin in establishment and maintenance of cell polarity revealed using the actin inhibitor latrunculin-a. J. Cell Biol. 137, Bose, I., Irazoqui, J.E., Moskow, J.J., Bardes, E.S., Zyla, T.R., and Lew, D.J. (001). Assembly of scaffold-mediated complexes containing Cdc4p, the exchange factor Cdc4p, and the effector Cla4p required for cell cycle-regulated phosphorylation of Cdc4p. J. Biol. Chem. 76, Burbelo, P.D., Drechsel, D., and Hall, A. (1995). A conserved binding motif defines numerous candidate target proteins for both Cdc4 and Rac GTPases. J. Biol. Chem. 70, Butty, A.C., Perrinjaquet, N., Petit, A., Jaquenoud, M., Segall, J.E., Hofmann, K., Zwahlen, C., and Peter, M. (00). A positive feedback loop stabilizes the guanine-nucleotide exchange factor Cdc4 at sites of polarization. EMBO J. 1, Caviston, J.P., Tcheperegine, S.E., and Bi, E. (00). Singularity in budding: a role for the evolutionarily conserved small GTPase Cdc4p. Proc. Natl. Acad. Sci. USA 99, Chant, J. (1999). Cell polarity in yeast. Annu. Rev. Cell Dev. Biol. 15, Gierer, A., and Meinhardt, H. (197). A theory of biological pattern formation. Kybernetik 1, Holland, M.J. (00). Transcript abundance in yeast varies over six orders of magnitude. J. Biol. Chem. 77, Huang, K.C., Meir, Y., and Wingreen, N.S. (003). Dynamic structures in Escherichia coli: spontaneous formation of MinE rings and MinD polar zones. Proc. Natl. Acad. Sci. USA 100, Irazoqui, J.E., Gladfelter, A.S., and Lew, D.J. (003). Scaffold-mediated symmetry breaking by Cdc4p. Nat. Cell Biol. 5, Irazoqui, J.E., Gladfelter, A.S., and Lew, D.J. (004). Cdc4p, GTP hydrolysis, and the cell s sense of direction. Cell Cycle 3, Irazoqui, J.E., Howell, A.S., Theesfeld, C.L., and Lew, D.J. (005). Opposing roles for actin in Cdc4p polarization. Mol. Biol. Cell 16, Killich, T., Plath, P.J., Xiang, W., Bultmann, H., Rensing, L., and Vicker, M.G. (1993). The locomotion shape and pseudopodial dynamics of unstimulated Dictyostelium cells are not random. J. Cell Sci. 106, Killich, T., Plath, P.J., Hass, E.C., Xiang, W., Bultmann, H., Rensing, L., and Vicker, M.G. (1994). Cell-movement and shape are nonrandom and determined by intracellular, oscillatory rotating waves in Dictyostelium amebas. Biosystems 33, Kozminski, K.G., Beven, L., Angerman, E., Tong, A.H., Boone, C., and Park, H.O. (003). Interaction between a Ras and a Rho GTPase couples selection of a growth site to the development of cell polarity in yeast. Mol. Biol. Cell 14, Lechler, T., Shevchenko, A., and Li, R. (000). Direct involvement of yeast type I myosins in Cdc4-dependent actin polymerization. J. Cell Biol. 148, Lew, D.J., and Reed, S.I. (1993). Morphogenesis in the yeast cell cycle: regulation by Cdc8 and cyclins. J. Cell Biol. 10, Meili, R., and Firtel, R.A. (003). Two poles and a compass. Cell 114, Meinhardt, H. (1999). Orientation of chemotactic cells and growth cones: models and mechanisms. J. Cell Sci. 11, Meinhardt, H., and de Boer, P.A.J. (001). Pattern formation in E. coli: a model for the pole-to-pole oscillations of Min proteins and the localization of the division site. Proc. Natl. Acad. Sci. USA 98, Nelson, W.J. (003). Adaptation of core mechanisms to generate cell polarity. Nature 4, Nern, A., and Arkowitz, R.A. (000). G proteins mediate changes in cell shape by stabilizing the axis of polarity. Mol. Cell 5, Park, H.O., Bi, E., Pringle, J.R., and Herskowitz, I. (1997). Two active states of the Ras-related Bud1/Rsr1 protein bind to different effectors to determine yeast cell polarity. Proc. Natl. Acad. Sci. USA 94, Saxton, M.J., and Jacobson, K. (1997). Single-particle tracking: applications to membrane dynamics. Annu. Rev. Biophys. Biomol. Struct. 6, Shimada, Y., Wiget, P., Gulli, M.P., Bi, E., and Peter, M. (004). The nucleotide exchange factor Cdc4p may be regulated by auto-inhibition. EMBO J. 3, Toenjes, K.A., Simpson, D., and Johnson, D.I. (004). Separate membrane targeting and anchoring domains function in the localization of the S. cerevisiae Cdc4p guanine nucleotide exchange factor. Curr. Genet. 45, Watanabe, T., Wang, S., Noritake, J., Sato, K., Fukata, M., Takefuji, M., Nakagawa, M., Izumi, N., Akiyama, T., and Kaibuchi, K. (004). Interaction with IQGAP1 links APC to Rac1, Cdc4, and actin filaments during cell polarization and migration. Dev. Cell 7,
7 Counteracting Feedback Loops Control Cdc4 Activity 571 Wedlich-Soldner, R., Altschuler, S., Wu, L., and Li, R. (003). Spontaneous cell polarization through actomyosin-based delivery of the Cdc4 GTPase. Science 99, Wedlich-Soldner, R., Wai, S.C., Schmidt, T., and Li, R. (004). Robust cell polarity is a dynamic state established by coupling transport and GTPase signaling. J. Cell Biol. 166, Weiner, O.D., Neilsen, P.O., Prestwich, G.D., Kirschner, M.W., Cantley, L.C., and Bourne, H.R. (00). A PtdInsP(3)- and Rho GTPasemediated positive feedback loop regulates neutrophil polarity. Nat. Cell Biol. 4,
Central Roles of Small GTPases in the Development of Cell Polarity in Yeast and Beyond
 MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, Mar. 2007, p. 48 96 Vol. 71, No. 1 1092-2172/07/$08.00 0 doi:10.1128/mmbr.00028-06 Copyright 2007, American Society for Microbiology. All Rights Reserved. Central
MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, Mar. 2007, p. 48 96 Vol. 71, No. 1 1092-2172/07/$08.00 0 doi:10.1128/mmbr.00028-06 Copyright 2007, American Society for Microbiology. All Rights Reserved. Central
Symmetry breaking and the establishment of cell polarity in budding yeast Jayme M Johnson 1, Meng Jin 2 and Daniel J Lew 1
 Available online at www.sciencedirect.com Symmetry breaking and the establishment of cell polarity in budding yeast Jayme M Johnson 1, Meng Jin 2 and Daniel J Lew 1 Cell polarity is typically oriented
Available online at www.sciencedirect.com Symmetry breaking and the establishment of cell polarity in budding yeast Jayme M Johnson 1, Meng Jin 2 and Daniel J Lew 1 Cell polarity is typically oriented
NIH Public Access Author Manuscript Curr Opin Genet Dev. Author manuscript; available in PMC 2012 December 1.
 NIH Public Access Author Manuscript Published in final edited form as: Curr Opin Genet Dev. 2011 December ; 21(6): 740 746. doi:10.1016/j.gde.2011.09.007. Symmetry breaking and the establishment of cell
NIH Public Access Author Manuscript Published in final edited form as: Curr Opin Genet Dev. 2011 December ; 21(6): 740 746. doi:10.1016/j.gde.2011.09.007. Symmetry breaking and the establishment of cell
Cyclical Regulation of the Exocyst and Cell Polarity Determinants for Polarized Cell Growth
 Molecular Biology of the Cell Vol. 16, 1500 1512, March 2005 Cyclical Regulation of the Exocyst and Cell Polarity Determinants for Polarized Cell Growth Allison Zajac,* Xiaoli Sun,* Jian Zhang, and Wei
Molecular Biology of the Cell Vol. 16, 1500 1512, March 2005 Cyclical Regulation of the Exocyst and Cell Polarity Determinants for Polarized Cell Growth Allison Zajac,* Xiaoli Sun,* Jian Zhang, and Wei
Mechanisms for Precise Positional Information in Bacteria: The Min system in E. coli and B. subtilis
 Mechanisms for Precise Positional Information in Bacteria: The Min system in E. coli and B. subtilis Martin Howard Imperial College London Bacterial Organization Many processes where bacterial cell needs
Mechanisms for Precise Positional Information in Bacteria: The Min system in E. coli and B. subtilis Martin Howard Imperial College London Bacterial Organization Many processes where bacterial cell needs
Stochastic simulations
 Stochastic simulations Application to molecular networks Literature overview Noise in genetic networks Origins How to measure and distinguish between the two types of noise (intrinsic vs extrinsic)? What
Stochastic simulations Application to molecular networks Literature overview Noise in genetic networks Origins How to measure and distinguish between the two types of noise (intrinsic vs extrinsic)? What
1 Morphogenesis: Control of Cell Types and Shape
 1 Morphogenesis: Control of Cell Types and Shape K.J. Boyce 1,2,A.Andrianopoulos 1 CONTENTS I. Introduction Cell Types and Cell Shapes: a Diverse Array of Form... 3 II. Polarity... 3 A. Cytoskeleton...
1 Morphogenesis: Control of Cell Types and Shape K.J. Boyce 1,2,A.Andrianopoulos 1 CONTENTS I. Introduction Cell Types and Cell Shapes: a Diverse Array of Form... 3 II. Polarity... 3 A. Cytoskeleton...
SPATIOTEMPORAL DYNAMICS OF GRADIENT SENSING AND POLARIZATION IN YEAST
 SPATIOTEMPORAL DYNAMICS OF GRADIENT SENSING AND POLARIZATION IN YEAST Meng Jin A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of the requirements
SPATIOTEMPORAL DYNAMICS OF GRADIENT SENSING AND POLARIZATION IN YEAST Meng Jin A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of the requirements
Dominance of Old End Growth is Inherited in Fission Yeast
 University of Tennessee, Knoxville Trace: Tennessee Research and Creative Exchange University of Tennessee Honors Thesis Projects University of Tennessee Honors Program 5-2015 Dominance of Old End Growth
University of Tennessee, Knoxville Trace: Tennessee Research and Creative Exchange University of Tennessee Honors Thesis Projects University of Tennessee Honors Program 5-2015 Dominance of Old End Growth
Figure 1f, cell 1) and at the bud tip ( 30% large-budded Accepted: 9 April 2001
 Brief Communication 803 A localized GTPase exchange factor, Bud5, determines the orientation of division axes in yeast Adele L. Marston*, Tracy Chen*, Melody C. Yang*, Pierre Belhumeur and John Chant*
Brief Communication 803 A localized GTPase exchange factor, Bud5, determines the orientation of division axes in yeast Adele L. Marston*, Tracy Chen*, Melody C. Yang*, Pierre Belhumeur and John Chant*
1. The plasma membrane of eukaryotic cells is supported by a. actin filaments. b. microtubules. c. lamins. d. intermediate filaments.
 ANALYSIS AND MODELING OF CELL MECHANICS Homework #2 (due 1/30/13) This homework involves comprehension of key biomechanical concepts of the cytoskeleton, cell-matrix adhesions, and cellcell adhesions.
ANALYSIS AND MODELING OF CELL MECHANICS Homework #2 (due 1/30/13) This homework involves comprehension of key biomechanical concepts of the cytoskeleton, cell-matrix adhesions, and cellcell adhesions.
Cytoskeletal Dynamics and the Temporal Control of Yeast Morphogenesis. Hsin Chen. Department of Cell Biology Duke University.
 Cytoskeletal Dynamics and the Temporal Control of Yeast Morphogenesis by Hsin Chen Department of Cell Biology Duke University Date: Approved: Daniel J. Lew, Supervisor Harold Erickson, Committee Chair
Cytoskeletal Dynamics and the Temporal Control of Yeast Morphogenesis by Hsin Chen Department of Cell Biology Duke University Date: Approved: Daniel J. Lew, Supervisor Harold Erickson, Committee Chair
Modeling Yeast Cell Polarization Induced by Pheromone Gradients
 Journal of Statistical Physics ( C 27 ) DOI: 1.17/s1955-7-9285-1 Modeling Yeast Cell Polarization Induced by Pheromone Gradients Tau-Mu Yi, 1 Shanqin Chen, 2 Ching-Shan Chou 2 and Qing Nie 2 Received March
Journal of Statistical Physics ( C 27 ) DOI: 1.17/s1955-7-9285-1 Modeling Yeast Cell Polarization Induced by Pheromone Gradients Tau-Mu Yi, 1 Shanqin Chen, 2 Ching-Shan Chou 2 and Qing Nie 2 Received March
Neurite initiation. Neurite formation begins with a bud that sprouts from the cell body. One or several neurites can sprout at a time.
 Neurite initiation. Neuronal maturation initiation f-actin polarization and maturation tubulin stage 1: "spherical" neuron stage 2: neurons extend several neurites stage 3: one neurite accelerates its
Neurite initiation. Neuronal maturation initiation f-actin polarization and maturation tubulin stage 1: "spherical" neuron stage 2: neurons extend several neurites stage 3: one neurite accelerates its
Neurite formation & neuronal polarization
 Neurite formation & neuronal polarization Paul Letourneau letou001@umn.edu Chapter 16; The Cytoskeleton; Molecular Biology of the Cell, Alberts et al. 1 An immature neuron in cell culture first sprouts
Neurite formation & neuronal polarization Paul Letourneau letou001@umn.edu Chapter 16; The Cytoskeleton; Molecular Biology of the Cell, Alberts et al. 1 An immature neuron in cell culture first sprouts
Cellular individuality in directional sensing. Azadeh Samadani (Brandeis University) Jerome Mettetal (MIT) Alexander van Oudenaarden (MIT)
 Cellular individuality in directional sensing Azadeh Samadani (Brandeis University) Jerome Mettetal (MIT) Alexander van Oudenaarden (MIT) How do cells make a decision? A cell makes many decisions based
Cellular individuality in directional sensing Azadeh Samadani (Brandeis University) Jerome Mettetal (MIT) Alexander van Oudenaarden (MIT) How do cells make a decision? A cell makes many decisions based
Neurite formation & neuronal polarization. The cytoskeletal components of neurons have characteristic distributions and associations
 Mechanisms of neuronal migration & Neurite formation & neuronal polarization Paul Letourneau letou001@umn.edu Chapter 16; The Cytoskeleton; Molecular Biology of the Cell, Alberts et al. 1 The cytoskeletal
Mechanisms of neuronal migration & Neurite formation & neuronal polarization Paul Letourneau letou001@umn.edu Chapter 16; The Cytoskeleton; Molecular Biology of the Cell, Alberts et al. 1 The cytoskeletal
Signal Transduction. Dr. Chaidir, Apt
 Signal Transduction Dr. Chaidir, Apt Background Complex unicellular organisms existed on Earth for approximately 2.5 billion years before the first multicellular organisms appeared.this long period for
Signal Transduction Dr. Chaidir, Apt Background Complex unicellular organisms existed on Earth for approximately 2.5 billion years before the first multicellular organisms appeared.this long period for
Cytokinesis in fission yeast: Modeling the assembly of the contractile ring
 Cytokinesis in fission yeast: Modeling the assembly of the contractile ring Nikola Ojkic, Dimitrios Vavylonis Department of Physics, Lehigh University Damien Laporte, Jian-Qiu Wu Department of Molecular
Cytokinesis in fission yeast: Modeling the assembly of the contractile ring Nikola Ojkic, Dimitrios Vavylonis Department of Physics, Lehigh University Damien Laporte, Jian-Qiu Wu Department of Molecular
Multistability in the lactose utilization network of Escherichia coli
 Multistability in the lactose utilization network of Escherichia coli Lauren Nakonechny, Katherine Smith, Michael Volk, Robert Wallace Mentor: J. Ruby Abrams Agenda Motivation Intro to multistability Purpose
Multistability in the lactose utilization network of Escherichia coli Lauren Nakonechny, Katherine Smith, Michael Volk, Robert Wallace Mentor: J. Ruby Abrams Agenda Motivation Intro to multistability Purpose
Abstract. Author Summary. Yougan Cheng 1, Hans Othmer 1,* 1 School of Mathematics, University of Minnesota, Minneapolis, MN, USA. *
 A Model for Direction Sensing in Dictyostelium Discoideum: Ras Activity and Symmetry Breaking Driven by a G βγ - Mediated, G α2 -Ric8 Dependent Signal Transduction Network - Submission to PLOS Computational
A Model for Direction Sensing in Dictyostelium Discoideum: Ras Activity and Symmetry Breaking Driven by a G βγ - Mediated, G α2 -Ric8 Dependent Signal Transduction Network - Submission to PLOS Computational
What are some of the major questions in cell biology? (That require quantitative methods and reasoning)
 Introduction What are some of the major questions in cell biology? (That require quantitative methods and reasoning) Big questions How does a cell know when to divide? How does it coordinate the process
Introduction What are some of the major questions in cell biology? (That require quantitative methods and reasoning) Big questions How does a cell know when to divide? How does it coordinate the process
ENDO-EXOCYTIC TRAFFICKING IN REGULATION OF CDC42 POLARITY. Leah Joy Watson
 ENDO-EXOCYTIC TRAFFICKING IN REGULATION OF CDC42 POLARITY Leah Joy Watson A dissertation submitted to the faculty at the University of North Carolina at Chapel Hill in partial fulfillment of the requirements
ENDO-EXOCYTIC TRAFFICKING IN REGULATION OF CDC42 POLARITY Leah Joy Watson A dissertation submitted to the faculty at the University of North Carolina at Chapel Hill in partial fulfillment of the requirements
13-3. Synthesis-Secretory pathway: Sort lumenal proteins, Secrete proteins, Sort membrane proteins
 13-3. Synthesis-Secretory pathway: Sort lumenal proteins, Secrete proteins, Sort membrane proteins Molecular sorting: specific budding, vesicular transport, fusion 1. Why is this important? A. Form and
13-3. Synthesis-Secretory pathway: Sort lumenal proteins, Secrete proteins, Sort membrane proteins Molecular sorting: specific budding, vesicular transport, fusion 1. Why is this important? A. Form and
Saccharomyces cerevisiae Cdc42p Localizes to Cellular Membranes and Clusters at Sites of Polarized Growth
 EUKARYOTIC CELL, June 2002, p. 458 468 Vol. 1, No. 3 1535-9778/02/$04.00 0 DOI: 10.1128/EC.1.3.458 468.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Saccharomyces cerevisiae
EUKARYOTIC CELL, June 2002, p. 458 468 Vol. 1, No. 3 1535-9778/02/$04.00 0 DOI: 10.1128/EC.1.3.458 468.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Saccharomyces cerevisiae
Mechanical Simulations of cell motility
 Mechanical Simulations of cell motility What are the overarching questions? How is the shape and motility of the cell regulated? How do cells polarize, change shape, and initiate motility? How do they
Mechanical Simulations of cell motility What are the overarching questions? How is the shape and motility of the cell regulated? How do cells polarize, change shape, and initiate motility? How do they
Sequential and Distinct Roles of the Cadherin Domain-containing Protein Axl2p in
 Sequential and Distinct Roles of the Cadherin Domain-containing Protein Axl2p in Cell Polarization in Yeast Cell Cycle Xiang-Dong Gao *, Lauren M. Sperber *, Steven A. Kane *, Zongtian Tong *, Amy Hin
Sequential and Distinct Roles of the Cadherin Domain-containing Protein Axl2p in Cell Polarization in Yeast Cell Cycle Xiang-Dong Gao *, Lauren M. Sperber *, Steven A. Kane *, Zongtian Tong *, Amy Hin
arxiv: v1 [q-bio.cb] 31 Dec 2015
![arxiv: v1 [q-bio.cb] 31 Dec 2015 arxiv: v1 [q-bio.cb] 31 Dec 2015](/thumbs/77/76028420.jpg) A Model for Direction Sensing in Dictyostelium Discoideum: Ras Activity and Symmetry Breaking Driven by a G βγ - Mediated, G α2 -Ric8 Dependent Signal Transduction Network arxiv:52.09233v [q-bio.cb] 3
A Model for Direction Sensing in Dictyostelium Discoideum: Ras Activity and Symmetry Breaking Driven by a G βγ - Mediated, G α2 -Ric8 Dependent Signal Transduction Network arxiv:52.09233v [q-bio.cb] 3
Cell Biology Review. The key components of cells that concern us are as follows: 1. Nucleus
 Cell Biology Review Development involves the collective behavior and activities of cells, working together in a coordinated manner to construct an organism. As such, the regulation of development is intimately
Cell Biology Review Development involves the collective behavior and activities of cells, working together in a coordinated manner to construct an organism. As such, the regulation of development is intimately
Activation of a receptor. Assembly of the complex
 Activation of a receptor ligand inactive, monomeric active, dimeric When activated by growth factor binding, the growth factor receptor tyrosine kinase phosphorylates the neighboring receptor. Assembly
Activation of a receptor ligand inactive, monomeric active, dimeric When activated by growth factor binding, the growth factor receptor tyrosine kinase phosphorylates the neighboring receptor. Assembly
Phosphorylation of Bem2p and Bem3p may contribute to local activation of Cdc42p at bud emergence
 The EMBO Journal (27) 26, 451 4513 & 27 European Molecular Biology Organization All Rights Reserved 261-4189/7 www.embojournal.org Phosphorylation of Bem2p and Bem3p may contribute to local activation
The EMBO Journal (27) 26, 451 4513 & 27 European Molecular Biology Organization All Rights Reserved 261-4189/7 www.embojournal.org Phosphorylation of Bem2p and Bem3p may contribute to local activation
16 The Cell Cycle. Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization
 The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
BE/APh161 Physical Biology of the Cell. Rob Phillips Applied Physics and Bioengineering California Institute of Technology
 BE/APh161 Physical Biology of the Cell Rob Phillips Applied Physics and Bioengineering California Institute of Technology Cells Decide: Where to Go The Hunters of the Immune Response (Berman et al.) There
BE/APh161 Physical Biology of the Cell Rob Phillips Applied Physics and Bioengineering California Institute of Technology Cells Decide: Where to Go The Hunters of the Immune Response (Berman et al.) There
Spatial Stochastic Dynamics Enable Robust Cell Polarization
 Spatial Stochastic Dynamics Enable Robust Cell Polarization Michael J. Lawson 1., Brian Drawert 2., Mustafa Khammash 3,4", Linda Petzold 2,3", Tau-Mu Yi 5"* 1 Department of BioMolecular Science and Engineering,
Spatial Stochastic Dynamics Enable Robust Cell Polarization Michael J. Lawson 1., Brian Drawert 2., Mustafa Khammash 3,4", Linda Petzold 2,3", Tau-Mu Yi 5"* 1 Department of BioMolecular Science and Engineering,
Regulation and signaling. Overview. Control of gene expression. Cells need to regulate the amounts of different proteins they express, depending on
 Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
The Rsr1/Bud1 GTPase interacts with itself and the Cdc42 GTPase during bud-site selection and polarity establishment in budding yeast
 The Rsr1/Bud1 GTPase interacts with itself and the Cdc42 GTPase during bud-site selection and polarity establishment in budding yeast Pil Jung Kang *, Laure Béven * =, Seethalakshmi Hariharan and Hay-Oak
The Rsr1/Bud1 GTPase interacts with itself and the Cdc42 GTPase during bud-site selection and polarity establishment in budding yeast Pil Jung Kang *, Laure Béven * =, Seethalakshmi Hariharan and Hay-Oak
Muscle regulation and Actin Topics: Tropomyosin and Troponin, Actin Assembly, Actin-dependent Movement
 1 Muscle regulation and Actin Topics: Tropomyosin and Troponin, Actin Assembly, Actin-dependent Movement In the last lecture, we saw that a repeating alternation between chemical (ATP hydrolysis) and vectorial
1 Muscle regulation and Actin Topics: Tropomyosin and Troponin, Actin Assembly, Actin-dependent Movement In the last lecture, we saw that a repeating alternation between chemical (ATP hydrolysis) and vectorial
The PXL1 Gene of Saccharomyces cerevisiae Encodes a Paxillin-like Protein Functioning in Polarized Cell Growth
 Molecular Biology of the Cell Vol. 15, 1904 1917, April 2004 The PXL1 Gene of Saccharomyces cerevisiae Encodes a Paxillin-like Protein Functioning in Polarized Cell Growth Nancy A. Mackin, Tarek J. Sousou,
Molecular Biology of the Cell Vol. 15, 1904 1917, April 2004 The PXL1 Gene of Saccharomyces cerevisiae Encodes a Paxillin-like Protein Functioning in Polarized Cell Growth Nancy A. Mackin, Tarek J. Sousou,
CELB40060 Membrane Trafficking in Animal Cells. Prof. Jeremy C. Simpson. Lecture 2 COPII and export from the ER
 CELB40060 Membrane Trafficking in Animal Cells Prof. Jeremy C. Simpson Lecture 2 COPII and export from the ER Today s lecture... The COPII coat - localisation and subunits Formation of the COPII coat at
CELB40060 Membrane Trafficking in Animal Cells Prof. Jeremy C. Simpson Lecture 2 COPII and export from the ER Today s lecture... The COPII coat - localisation and subunits Formation of the COPII coat at
The molecular genetics of chemotaxis: sensing and responding to chemoattractant gradients
 The molecular genetics of chemotaxis: sensing and responding to chemoattractant gradients Richard A. Firtel* and Chang Y. Chung Summary Chemotaxis plays a central role in various biological processes,
The molecular genetics of chemotaxis: sensing and responding to chemoattractant gradients Richard A. Firtel* and Chang Y. Chung Summary Chemotaxis plays a central role in various biological processes,
Edinburgh Research Explorer
 Edinburgh Research Explorer Many roads to symmetry breaking: Molecular mechanisms and theoretical models of yeast cell polarity Citation for published version: Goryachev, A & Leda, M 2017, 'Many roads
Edinburgh Research Explorer Many roads to symmetry breaking: Molecular mechanisms and theoretical models of yeast cell polarity Citation for published version: Goryachev, A & Leda, M 2017, 'Many roads
Pheromone Gradient Tracking Mechanisms During Yeast Mating. Allison Wolff McClure. Department of Pharmacology and Cancer Biology Duke University
 Pheromone Gradient Tracking Mechanisms During Yeast Mating by Allison Wolff McClure Department of Pharmacology and Cancer Biology Duke University Date: Approved: Daniel Lew, Supervisor Marc Caron Timothy
Pheromone Gradient Tracking Mechanisms During Yeast Mating by Allison Wolff McClure Department of Pharmacology and Cancer Biology Duke University Date: Approved: Daniel Lew, Supervisor Marc Caron Timothy
7.32/7.81J/8.591J. Rm Rm (under construction) Alexander van Oudenaarden Jialing Li. Bernardo Pando. Rm.
 Introducing... 7.32/7.81J/8.591J Systems Biology modeling biological networks Lectures: Recitations: ti TR 1:00-2:30 PM W 4:00-5:00 PM Rm. 6-120 Rm. 26-204 (under construction) Alexander van Oudenaarden
Introducing... 7.32/7.81J/8.591J Systems Biology modeling biological networks Lectures: Recitations: ti TR 1:00-2:30 PM W 4:00-5:00 PM Rm. 6-120 Rm. 26-204 (under construction) Alexander van Oudenaarden
Supporting online material
 Supporting online material Materials and Methods Target proteins All predicted ORFs in the E. coli genome (1) were downloaded from the Colibri data base (2) (http://genolist.pasteur.fr/colibri/). 737 proteins
Supporting online material Materials and Methods Target proteins All predicted ORFs in the E. coli genome (1) were downloaded from the Colibri data base (2) (http://genolist.pasteur.fr/colibri/). 737 proteins
Lecture 4: Transcription networks basic concepts
 Lecture 4: Transcription networks basic concepts - Activators and repressors - Input functions; Logic input functions; Multidimensional input functions - Dynamics and response time 2.1 Introduction The
Lecture 4: Transcription networks basic concepts - Activators and repressors - Input functions; Logic input functions; Multidimensional input functions - Dynamics and response time 2.1 Introduction The
MAP Kinase Pathways in the Yeast Saccharomyces cerevisiae
 MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, Dec. 1998, p. 1264 1300 Vol. 62, No. 4 1092-2172/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. MAP Kinase Pathways in the
MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, Dec. 1998, p. 1264 1300 Vol. 62, No. 4 1092-2172/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. MAP Kinase Pathways in the
Three Myosins Contribute Uniquely to the Assembly and Constriction of the Fission Yeast Cytokinetic Contractile Ring
 Current Biology Supplemental Information Three Myosins Contribute Uniquely to the Assembly and Constriction of the Fission Yeast Cytokinetic Contractile Ring Caroline Laplante, Julien Berro, Erdem Karatekin,
Current Biology Supplemental Information Three Myosins Contribute Uniquely to the Assembly and Constriction of the Fission Yeast Cytokinetic Contractile Ring Caroline Laplante, Julien Berro, Erdem Karatekin,
Supporting Information
 Supporting Information Fleissner et al. 10.1073/pnas.0907039106 Fig. S1. (A) MAK-2-GFP localized to CATs tips is not bound by membrane. his-3::pccg1 mak-2-gfp; mak-2 strain labeled with membrane dye FM4
Supporting Information Fleissner et al. 10.1073/pnas.0907039106 Fig. S1. (A) MAK-2-GFP localized to CATs tips is not bound by membrane. his-3::pccg1 mak-2-gfp; mak-2 strain labeled with membrane dye FM4
BME Engineering Molecular Cell Biology. Basics of the Diffusion Theory. The Cytoskeleton (I)
 BME 42-620 Engineering Molecular Cell Biology Lecture 07: Basics of the Diffusion Theory The Cytoskeleton (I) BME42-620 Lecture 07, September 20, 2011 1 Outline Diffusion: microscopic theory Diffusion:
BME 42-620 Engineering Molecular Cell Biology Lecture 07: Basics of the Diffusion Theory The Cytoskeleton (I) BME42-620 Lecture 07, September 20, 2011 1 Outline Diffusion: microscopic theory Diffusion:
BME Engineering Molecular Cell Biology. Review: Basics of the Diffusion Theory. The Cytoskeleton (I)
 BME 42-620 Engineering Molecular Cell Biology Lecture 08: Review: Basics of the Diffusion Theory The Cytoskeleton (I) BME42-620 Lecture 08, September 22, 2011 1 Outline Background: FRAP & SPT Review: microscopic
BME 42-620 Engineering Molecular Cell Biology Lecture 08: Review: Basics of the Diffusion Theory The Cytoskeleton (I) BME42-620 Lecture 08, September 22, 2011 1 Outline Background: FRAP & SPT Review: microscopic
A NOVEL ROLE FOR THE RGS PROTEIN SST2 IN YEAST SIGNALING REVEALED BY MATHEMATICAL MODELING AND LIVE CELL IMAGING. Sai Phanindra Venkatapurapu
 A NOVEL ROLE FOR THE RGS PROTEIN SST2 IN YEAST SIGNALING REVEALED BY MATHEMATICAL MODELING AND LIVE CELL IMAGING Sai Phanindra Venkatapurapu A dissertation submitted to the faculty of the University of
A NOVEL ROLE FOR THE RGS PROTEIN SST2 IN YEAST SIGNALING REVEALED BY MATHEMATICAL MODELING AND LIVE CELL IMAGING Sai Phanindra Venkatapurapu A dissertation submitted to the faculty of the University of
Localized Ras signaling at the leading edge regulates PI3K, cell polarity, and directional cell movement
 JCB: ARTICLE Localized Ras signaling at the leading edge regulates PI3K, cell polarity, and directional cell movement Atsuo T. Sasaki, Cheryl Chun, Kosuke Takeda, and Richard A. Firtel Section of Cell
JCB: ARTICLE Localized Ras signaling at the leading edge regulates PI3K, cell polarity, and directional cell movement Atsuo T. Sasaki, Cheryl Chun, Kosuke Takeda, and Richard A. Firtel Section of Cell
56:198:582 Biological Networks Lecture 8
 56:198:582 Biological Networks Lecture 8 Course organization Two complementary approaches to modeling and understanding biological networks Constraint-based modeling (Palsson) System-wide Metabolism Steady-state
56:198:582 Biological Networks Lecture 8 Course organization Two complementary approaches to modeling and understanding biological networks Constraint-based modeling (Palsson) System-wide Metabolism Steady-state
Analysis and Simulation of Biological Systems
 Analysis and Simulation of Biological Systems Dr. Carlo Cosentino School of Computer and Biomedical Engineering Department of Experimental and Clinical Medicine Università degli Studi Magna Graecia Catanzaro,
Analysis and Simulation of Biological Systems Dr. Carlo Cosentino School of Computer and Biomedical Engineering Department of Experimental and Clinical Medicine Università degli Studi Magna Graecia Catanzaro,
return in class, or Rm B
 Last lectures: Genetic Switches and Oscillators PS #2 due today bf before 3PM return in class, or Rm. 68 371B Naturally occurring: lambda lysis-lysogeny decision lactose operon in E. coli Engineered: genetic
Last lectures: Genetic Switches and Oscillators PS #2 due today bf before 3PM return in class, or Rm. 68 371B Naturally occurring: lambda lysis-lysogeny decision lactose operon in E. coli Engineered: genetic
7.06 Problem Set #4, Spring 2005
 7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
Phosphorylation of the Cdc42 Exchange Factor Cdc24 by the PAK-like Kinase Cla4 May Regulate Polarized Growth in Yeast
 Molecular Cell, Vol. 6, 1155 1167, November, 2000, Copyright 2000 by Cell Press Phosphorylation of the Cdc42 Exchange Factor Cdc24 by the PAK-like Kinase Cla4 May Regulate Polarized Growth in Yeast Marie-Pierre
Molecular Cell, Vol. 6, 1155 1167, November, 2000, Copyright 2000 by Cell Press Phosphorylation of the Cdc42 Exchange Factor Cdc24 by the PAK-like Kinase Cla4 May Regulate Polarized Growth in Yeast Marie-Pierre
Signaling regulated endocytosis and exocytosis lead to mating pheromone concentration dependent morphologies in yeast
 FES Letters 586 (22) 428 424 journal homepage: www.fesletters.org Signaling regulated endocytosis and exocytosis lead to mating pheromone concentration dependent morphologies in yeast Ching-Shan Chou a,,,
FES Letters 586 (22) 428 424 journal homepage: www.fesletters.org Signaling regulated endocytosis and exocytosis lead to mating pheromone concentration dependent morphologies in yeast Ching-Shan Chou a,,,
Mechanisms of Cell Proliferation
 Mechanisms of Cell Proliferation Cell Cycle G 2 S G 1 Multi-cellular organisms depend on cell division/proliferation; Each organism has a developmental plan that determines its behavior and properties;
Mechanisms of Cell Proliferation Cell Cycle G 2 S G 1 Multi-cellular organisms depend on cell division/proliferation; Each organism has a developmental plan that determines its behavior and properties;
Stochastic simulations!
 Stochastic simulations! Application to biomolecular networks! Literature overview Noise in genetic networks! Origins! How to measure the noise and distinguish between the two sources of noise (intrinsic
Stochastic simulations! Application to biomolecular networks! Literature overview Noise in genetic networks! Origins! How to measure the noise and distinguish between the two sources of noise (intrinsic
56:198:582 Biological Networks Lecture 9
 56:198:582 Biological Networks Lecture 9 The Feed-Forward Loop Network Motif Subgraphs in random networks We have discussed the simplest network motif, self-regulation, a pattern with one node We now consider
56:198:582 Biological Networks Lecture 9 The Feed-Forward Loop Network Motif Subgraphs in random networks We have discussed the simplest network motif, self-regulation, a pattern with one node We now consider
Bacterial Chemotaxis
 Bacterial Chemotaxis Bacteria can be attracted/repelled by chemicals Mechanism? Chemoreceptors in bacteria. attractant Adler, 1969 Science READ! This is sensing, not metabolism Based on genetic approach!!!
Bacterial Chemotaxis Bacteria can be attracted/repelled by chemicals Mechanism? Chemoreceptors in bacteria. attractant Adler, 1969 Science READ! This is sensing, not metabolism Based on genetic approach!!!
Plant Molecular and Cellular Biology Lecture 8: Mechanisms of Cell Cycle Control and DNA Synthesis Gary Peter
 Plant Molecular and Cellular Biology Lecture 8: Mechanisms of Cell Cycle Control and DNA Synthesis Gary Peter 9/10/2008 1 Learning Objectives Explain why a cell cycle was selected for during evolution
Plant Molecular and Cellular Biology Lecture 8: Mechanisms of Cell Cycle Control and DNA Synthesis Gary Peter 9/10/2008 1 Learning Objectives Explain why a cell cycle was selected for during evolution
Multistability in the lactose utilization network of E. coli. Lauren Nakonechny, Katherine Smith, Michael Volk, Robert Wallace Mentor: J.
 Multistability in the lactose utilization network of E. coli Lauren Nakonechny, Katherine Smith, Michael Volk, Robert Wallace Mentor: J. Ruby Abrams Motivation Understanding biological switches in the
Multistability in the lactose utilization network of E. coli Lauren Nakonechny, Katherine Smith, Michael Volk, Robert Wallace Mentor: J. Ruby Abrams Motivation Understanding biological switches in the
STOCHASTICITY IN GENE EXPRESSION: FROM THEORIES TO PHENOTYPES
 STOCHASTICITY IN GENE EXPRESSION: FROM THEORIES TO PHENOTYPES Mads Kærn*, Timothy C. Elston, William J. Blake and James J. Collins Abstract Genetically identical cells exposed to the same environmental
STOCHASTICITY IN GENE EXPRESSION: FROM THEORIES TO PHENOTYPES Mads Kærn*, Timothy C. Elston, William J. Blake and James J. Collins Abstract Genetically identical cells exposed to the same environmental
In Escherichia coli, two systems are known to regulate the
 Dynamic structures in Escherichia coli: Spontaneous formation of MinE rings and MinD polar zones Kerwyn Casey Huang*, Yigal Meir, and Ned S. Wingreen *Department of Physics, Massachusetts Institute of
Dynamic structures in Escherichia coli: Spontaneous formation of MinE rings and MinD polar zones Kerwyn Casey Huang*, Yigal Meir, and Ned S. Wingreen *Department of Physics, Massachusetts Institute of
Mechanisms of Cell Proliferation
 Mechanisms of Cell Proliferation Cell Cycle G 2 S G 1 Multi-cellular organisms depend on cell division/proliferation; Each organism has a developmental plan that determines its behavior and properties;
Mechanisms of Cell Proliferation Cell Cycle G 2 S G 1 Multi-cellular organisms depend on cell division/proliferation; Each organism has a developmental plan that determines its behavior and properties;
Lecture 8: Temporal programs and the global structure of transcription networks. Chap 5 of Alon. 5.1 Introduction
 Lecture 8: Temporal programs and the global structure of transcription networks Chap 5 of Alon 5. Introduction We will see in this chapter that sensory transcription networks are largely made of just four
Lecture 8: Temporal programs and the global structure of transcription networks Chap 5 of Alon 5. Introduction We will see in this chapter that sensory transcription networks are largely made of just four
HHS Public Access Author manuscript IET Syst Biol. Author manuscript; available in PMC 2015 May 28.
 Alternative cell polarity behaviours arise from changes in G- protein spatial dynamics Ching-Shan Chou 1, Travis I. Moore 2, Qing Nie 3, and Tau-Mu Yi 4 Tau-Mu Yi: taumu.yi@lifesci.ucsb.edu 1 Department
Alternative cell polarity behaviours arise from changes in G- protein spatial dynamics Ching-Shan Chou 1, Travis I. Moore 2, Qing Nie 3, and Tau-Mu Yi 4 Tau-Mu Yi: taumu.yi@lifesci.ucsb.edu 1 Department
A Mathematical Model for Neutrophil Gradient Sensing and Polarization
 A Mathematical Model for Neutrophil Gradient Sensing and Polarization Matthew Onsum 1, Christopher V. Rao 2* 1 AstraZeneca R&D Boston, Waltham, Massachusetts, United States of America, 2 Department of
A Mathematical Model for Neutrophil Gradient Sensing and Polarization Matthew Onsum 1, Christopher V. Rao 2* 1 AstraZeneca R&D Boston, Waltham, Massachusetts, United States of America, 2 Department of
56:198:582 Biological Networks Lecture 10
 56:198:582 Biological Networks Lecture 10 Temporal Programs and the Global Structure The single-input module (SIM) network motif The network motifs we have studied so far all had a defined number of nodes.
56:198:582 Biological Networks Lecture 10 Temporal Programs and the Global Structure The single-input module (SIM) network motif The network motifs we have studied so far all had a defined number of nodes.
Actin-based Motility during Endocytosis in Budding Yeast V
 Molecular Biology of the Cell Vol. 17, 1354 1363, March 2006 Actin-based Motility during Endocytosis in Budding Yeast V Kyoungtae Kim,* Brian J. Galletta,* Kevin O. Schmidt, Fanny S. Chang, Kendall J.
Molecular Biology of the Cell Vol. 17, 1354 1363, March 2006 Actin-based Motility during Endocytosis in Budding Yeast V Kyoungtae Kim,* Brian J. Galletta,* Kevin O. Schmidt, Fanny S. Chang, Kendall J.
A complementation test would be done by crossing the haploid strains and scoring the phenotype in the diploids.
 Problem set H answers 1. To study DNA repair mechanisms, geneticists isolated yeast mutants that were sensitive to various types of radiation; for example, mutants that were more sensitive to UV light.
Problem set H answers 1. To study DNA repair mechanisms, geneticists isolated yeast mutants that were sensitive to various types of radiation; for example, mutants that were more sensitive to UV light.
Possible mechanisms for initiating macroscopic left-right asymmetry in developing organisms
 Possible mechanisms for initiating macroscopic left-right asymmetry in developing organisms Chris Henley, Ricky Chachra, Jimmy Shen Cornell U. [Support: U.S. Dept. of Energy] APS March Meeting, Mar. 2,
Possible mechanisms for initiating macroscopic left-right asymmetry in developing organisms Chris Henley, Ricky Chachra, Jimmy Shen Cornell U. [Support: U.S. Dept. of Energy] APS March Meeting, Mar. 2,
Cellular Biophysics SS Prof. Manfred Radmacher
 SS 20007 Manfred Radmacher Ch. 12 Systems Biology Let's recall chemotaxis in Dictiostelium synthesis of camp excretion of camp external camp gradient detection cell polarity cell migration 2 Single cells
SS 20007 Manfred Radmacher Ch. 12 Systems Biology Let's recall chemotaxis in Dictiostelium synthesis of camp excretion of camp external camp gradient detection cell polarity cell migration 2 Single cells
Active mechanics of cells. Tetsuya Hiraiwa The University of Tokyo
 Active mechanics of cells Tetsuya Hiraiwa The University of Tokyo 100μm Active mechanics of cells Tetsuya Hiraiwa The University of Tokyo (HeLa cells) Cellular scale (~ several 10μm) Subcellular scale
Active mechanics of cells Tetsuya Hiraiwa The University of Tokyo 100μm Active mechanics of cells Tetsuya Hiraiwa The University of Tokyo (HeLa cells) Cellular scale (~ several 10μm) Subcellular scale
Measuring TF-DNA interactions
 Measuring TF-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Regulation of Gene Expression by Transcription Factors TF trans-acting factors TF TF TF TF
Measuring TF-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Regulation of Gene Expression by Transcription Factors TF trans-acting factors TF TF TF TF
Under the Radar Screen: How Bugs Trick Our Immune Defenses
 Under the Radar Screen: How Bugs Trick Our Immune Defenses Session 2: Phagocytosis Marie-Eve Paquet and Gijsbert Grotenbreg Whitehead Institute for Biomedical Research Salmonella Gram negative bacteria
Under the Radar Screen: How Bugs Trick Our Immune Defenses Session 2: Phagocytosis Marie-Eve Paquet and Gijsbert Grotenbreg Whitehead Institute for Biomedical Research Salmonella Gram negative bacteria
Advanced Higher Biology. Unit 1- Cells and Proteins 2c) Membrane Proteins
 Advanced Higher Biology Unit 1- Cells and Proteins 2c) Membrane Proteins Membrane Structure Phospholipid bilayer Transmembrane protein Integral protein Movement of Molecules Across Membranes Phospholipid
Advanced Higher Biology Unit 1- Cells and Proteins 2c) Membrane Proteins Membrane Structure Phospholipid bilayer Transmembrane protein Integral protein Movement of Molecules Across Membranes Phospholipid
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06072 SUPPLEMENTARY INFORMATION Molecular noise and size control: origins of variability in the budding yeast cell cycle. Stefano Di Talia 1,2, Jan M. Skotheim 2, James M. Bean 1,*,
doi: 10.1038/nature06072 SUPPLEMENTARY INFORMATION Molecular noise and size control: origins of variability in the budding yeast cell cycle. Stefano Di Talia 1,2, Jan M. Skotheim 2, James M. Bean 1,*,
Chapter 6- An Introduction to Metabolism*
 Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
The Alpha-Factor Receptor C-terminus Is Important for Mating Projection Formation and Orientation in Saccharomyces cerevisiae
 The Alpha-Factor Receptor C-terminus Is Important for Mating Projection Formation and Orientation in Saccharomyces cerevisiae Laura G. Vallier, 1 Jeffrey E. Segall, 2 and Michael Snyder 1 * Cell Motility
The Alpha-Factor Receptor C-terminus Is Important for Mating Projection Formation and Orientation in Saccharomyces cerevisiae Laura G. Vallier, 1 Jeffrey E. Segall, 2 and Michael Snyder 1 * Cell Motility
Lecture 10: Cyclins, cyclin kinases and cell division
 Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
RGS Proteins and Septins Cooperate to Promote Chemotropism by Regulating Polar Cap Mobility
 Article RGS Proteins and Septins Cooperate to Promote Chemotropism by Regulating Polar Cap Mobility Graphical Abstract Authors Joshua B. Kelley, Gauri Dixit,..., Timothy C. Elston, Henrik G. Dohlman Correspondence
Article RGS Proteins and Septins Cooperate to Promote Chemotropism by Regulating Polar Cap Mobility Graphical Abstract Authors Joshua B. Kelley, Gauri Dixit,..., Timothy C. Elston, Henrik G. Dohlman Correspondence
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3267 Supplementary Figure 1 A group of genes required for formation or orientation of annular F-actin bundles and aecm ridges: RNAi phenotypes and their validation by standard mutations.
DOI: 10.1038/ncb3267 Supplementary Figure 1 A group of genes required for formation or orientation of annular F-actin bundles and aecm ridges: RNAi phenotypes and their validation by standard mutations.
Dr. Fred Cross, Rockefeller (KITP Bio Networks 3/26/2003) 1
 Outline Cell growth as the driver for cell cycle (in microbes): coordination of growth and division A basic principle organizing cell cycle control: why cyclin-dependent kinase activity must oscillate
Outline Cell growth as the driver for cell cycle (in microbes): coordination of growth and division A basic principle organizing cell cycle control: why cyclin-dependent kinase activity must oscillate
Adventures in Multicellularity
 Adventures in Multicellularity The social amoeba (a.k.a. slime molds) Dictyostelium discoideum Dictyostelium discoideum the most studied of the social amoebae / cellular slime molds predatory soil amoeba
Adventures in Multicellularity The social amoeba (a.k.a. slime molds) Dictyostelium discoideum Dictyostelium discoideum the most studied of the social amoebae / cellular slime molds predatory soil amoeba
Heather Currinn, Benjamin Guscott, Zita Balklava, Alice Rothnie and Thomas Wassmer*
 Online Resources APP controls the formation of PI(3,5)P 2 vesicles through its binding of the PIKfyve complex. Cellular and Molecular Life Sciences Heather Currinn, Benjamin Guscott, Zita Balklava, Alice
Online Resources APP controls the formation of PI(3,5)P 2 vesicles through its binding of the PIKfyve complex. Cellular and Molecular Life Sciences Heather Currinn, Benjamin Guscott, Zita Balklava, Alice
Cell polarity models
 Cell polarity models Universal features of polarizing cells 1. Ability to sense both steep and shallow external gradients (as small as 1% 2%) in vast range of concentrations. Polarization leads to an amplification
Cell polarity models Universal features of polarizing cells 1. Ability to sense both steep and shallow external gradients (as small as 1% 2%) in vast range of concentrations. Polarization leads to an amplification
monomer polymer polymeric network cell
 3.1 Motivation 3.2 Polymerization The structural stability of the cell is provided by the cytoskeleton. Assembling and disassembling dynamically, the cytoskeleton enables cell movement through a highly
3.1 Motivation 3.2 Polymerization The structural stability of the cell is provided by the cytoskeleton. Assembling and disassembling dynamically, the cytoskeleton enables cell movement through a highly
Mechanistic basis of equal plasmid spacing in E. coli: a general mechanism for sizing and spacing
 Mechanistic basis of equal plasmid spacing in E. coli: a general mechanism for sizing and spacing CANES, Kings College London November 11 th, 2015 Martin Howard Computational & Systems Biology John Innes
Mechanistic basis of equal plasmid spacing in E. coli: a general mechanism for sizing and spacing CANES, Kings College London November 11 th, 2015 Martin Howard Computational & Systems Biology John Innes
Molecular Cell Biology 5068 In Class Exam 1 September 30, Please print your name:
 Molecular Cell Biology 5068 In Class Exam 1 September 30, 2014 Exam Number: Please print your name: Instructions: Please write only on these pages, in the spaces allotted and not on the back. Write your
Molecular Cell Biology 5068 In Class Exam 1 September 30, 2014 Exam Number: Please print your name: Instructions: Please write only on these pages, in the spaces allotted and not on the back. Write your
Correct Targeting of Plant ARF GTPases Relies on Distinct Protein Domains
 Traffic 2008; 9: 103 120 Blackwell Munksgaard # 2007 The Authors Journal compilation # 2007 Blackwell Publishing Ltd doi: 10.1111/j.1600-0854.2007.00671.x Correct Targeting of Plant ARF GTPases Relies
Traffic 2008; 9: 103 120 Blackwell Munksgaard # 2007 The Authors Journal compilation # 2007 Blackwell Publishing Ltd doi: 10.1111/j.1600-0854.2007.00671.x Correct Targeting of Plant ARF GTPases Relies
BCH 4054 Spring 2001 Chapter 33 Lecture Notes
 BCH 4054 Spring 2001 Chapter 33 Lecture Notes Slide 1 The chapter covers degradation of proteins as well. We will not have time to get into that subject. Chapter 33 Protein Synthesis Slide 2 Prokaryotic
BCH 4054 Spring 2001 Chapter 33 Lecture Notes Slide 1 The chapter covers degradation of proteins as well. We will not have time to get into that subject. Chapter 33 Protein Synthesis Slide 2 Prokaryotic
Student Learning Outcomes: Nucleus distinguishes Eukaryotes from Prokaryotes
 9 The Nucleus Student Learning Outcomes: Nucleus distinguishes Eukaryotes from Prokaryotes Explain general structures of Nuclear Envelope, Nuclear Lamina, Nuclear Pore Complex Explain movement of proteins
9 The Nucleus Student Learning Outcomes: Nucleus distinguishes Eukaryotes from Prokaryotes Explain general structures of Nuclear Envelope, Nuclear Lamina, Nuclear Pore Complex Explain movement of proteins
DISCOVERIES OF MACHINERY REGULATING VESICLE TRAFFIC, A MAJOR TRANSPORT SYSTEM IN OUR CELLS. Scientific Background on the Nobel Prize in Medicine 2013
 DISCOVERIES OF MACHINERY REGULATING VESICLE TRAFFIC, A MAJOR TRANSPORT SYSTEM IN OUR CELLS Scientific Background on the Nobel Prize in Medicine 2013 Daniela Scalet 6/12/2013 The Nobel Prize in Medicine
DISCOVERIES OF MACHINERY REGULATING VESICLE TRAFFIC, A MAJOR TRANSPORT SYSTEM IN OUR CELLS Scientific Background on the Nobel Prize in Medicine 2013 Daniela Scalet 6/12/2013 The Nobel Prize in Medicine
Chapter 3 Part 1! 10 th ed.: pp ! 11 th ed.: pp !! Cellular Transport Mechanisms! The Cell Cycle!
 Chapter 3 Part 1! 10 th ed.: pp. 87 105! 11 th ed.: pp. 90 107!! Cellular Transport Mechanisms! The Cell Cycle! Transport Processes: Passive and Active (1 of 2)! 1. Passive transport! Does not use ATP!
Chapter 3 Part 1! 10 th ed.: pp. 87 105! 11 th ed.: pp. 90 107!! Cellular Transport Mechanisms! The Cell Cycle! Transport Processes: Passive and Active (1 of 2)! 1. Passive transport! Does not use ATP!
Chapter 3 Part 1! 10 th ed.: pp ! 11 th ed.: pp !! Cellular Transport Mechanisms! The Cell Cycle!
 Chapter 3 Part 1! 10 th ed.: pp. 87 105! 11 th ed.: pp. 90 107!! Cellular Transport Mechanisms! The Cell Cycle! Transport Processes: Passive and Active (1 of 2)! 1. Passive transport! Does not use ATP!
Chapter 3 Part 1! 10 th ed.: pp. 87 105! 11 th ed.: pp. 90 107!! Cellular Transport Mechanisms! The Cell Cycle! Transport Processes: Passive and Active (1 of 2)! 1. Passive transport! Does not use ATP!
Chapter 16. Cellular Movement: Motility and Contractility. Lectures by Kathleen Fitzpatrick Simon Fraser University Pearson Education, Inc.
 Chapter 16 Cellular Movement: Motility and Contractility Lectures by Kathleen Fitzpatrick Simon Fraser University Two eukaryotic motility systems 1. Interactions between motor proteins and microtubules
Chapter 16 Cellular Movement: Motility and Contractility Lectures by Kathleen Fitzpatrick Simon Fraser University Two eukaryotic motility systems 1. Interactions between motor proteins and microtubules
Protein Pattern Formation
 Protein Pattern Formation Erwin Frey 1, Jacob Halatek 1, Simon Kretschmer 2,3, and Petra Schwille 2 1 Arnold-Sommerfeld-Center for Theoretical Physics and Center for NanoScience, Department of Physics,
Protein Pattern Formation Erwin Frey 1, Jacob Halatek 1, Simon Kretschmer 2,3, and Petra Schwille 2 1 Arnold-Sommerfeld-Center for Theoretical Physics and Center for NanoScience, Department of Physics,
