Representing Phenotypes in OWL
|
|
|
- Jocelin Hart
- 5 years ago
- Views:
Transcription
1 Representing Phenotypes in OWL Chris Mungall, Georgios Gkoutos, Nicole Washington and Suzanna Lewis Abstract. Accurate representation of phenotypes using ontologies is important in biology and biomedicine. This paper describes the OWL translation of our methodology for representing phenotypes using ontologies in the OBO Foundry. 1 Introduction: On Phenotypes In simple terms, a phenotype is a collection of characteristics that arise through the expression of the genes of an organism, in an environment. Some examples of phenotypes are the red eyes of a typical fruitfly, Drosophila melanogaster; the length of a body part such as the tail of a mouse; high blood pressure in the arteries of a human, sensitivity to a chemical or to light; reduced mass of an organ or part of an organism such as bone. In biology, we are often interested in how variations in genotype and environment can lead to variations in phenotype. There is a wealth of research to draw from, but much of the knowledge is currently encoded in natural language and free text in the literature and biological databases[?] - if this knowledge can be captured using ontologies and ontological formalisms we increase the range of computational methods that can be used to analyse this data[?]. This paper describes the OWL translation of a formalism for representing phenotypes making use of the ontologies that comprise Open Bio-Ontologies Foundry ( Many of these ontologies such as the Gene Ontology[?], the Foundational Model of Anatomy[?] and the Cell Ontology[?] represent canonical biology, which is to say the biology of normal, typical or healthy organisms. We use these ontologies as building blocks that can be combined with an ontology of qualities to construct descriptions of phenotypes, many of which deviate from canonical biology. This methodology can be used in a variety of different kinds of investigations. The reader of this paper should be aware that the end-user of the software systems and databases built around the formalism presented do not work directly at level of OWL constructs, or at their OBO-Formalism equivalents. They typically interact via intermediate representations and user interfaces such as Phenote 1 - more details can be found on the phenotype ontology website 2. We present the OWL version of the formalism in order to give it a grounding in logic, and as a means of leveraging OWL-based tools and reasoning engines, although space dictates we do not discuss these applications
2 1.1 Conventions Used We use the Manchester OWL syntax [?] for writing our OWL representations 3. Most of the OWL descriptions in this paper can also be translated to OBO- Format equivalents; we do not include these here for brevity, details can be found at and the OboInOwl page 4. Our examples are all drawn from OBO Foundry Ontologies[?] for the most part. OBO ontologies uniquely identify classes via a numeric identifier, but in the examples given we replace these with class labels in order to enhance readability. We do not always specify the ontology used in the examples given, the full list can be found on the OBO Foundry website 5. 2 OWL Representation The biological and biomedical literature contains descriptions of phenotypes such as the following: 1. red eye (a typical or wild-type specimen of the fruitfly Drosophila melanogaster has red eyes. 2. curvature of a wing 3. high blood pressure 4. high glucose concentration in the blood 5. short tail 6. invaginated cell membrane How should we go about representing phenotypes such as these in OWL? We will first review two existing W3C recommendations (instance and class partitions), and then describe our approach. 2.1 Enumeration of Individuals The W3C Working Group note Representing Specified Values in OWL: value partitions and value sets[?] provides guidelines for two different ways of representing what it calls specified values: (1) enumerations of individuals and (2) partitions of classes. These Specified Values are applicable to organismal phenotypes. The first method involves subdividing a quality such as curvature into owl Individuals, such as flat and curved, represented in Manchester Syntax as: Class: CurvatureValue EquivalentClass: {flat curved} Individual: flat TYPE CurvatureValue Individual: curved TYPE CurvatureValue 3 syntax/
3 We may also want to use differentfrom statements to state that flat and curved are distinct individuals: Individual: flat DifferentFrom: curved We can relate individuals to color hues using an OWL functional property 6 : FunctionalProperty: hascurvature Range: CurvatureValue So an instance of a curved fruitfly wing 7 could be represented as: Individual: fly-wing TYPE Wing Facts: hascurvature VALUE curved or the logically equivalent: Individual: fly-wing TYPE (Wing THAT hascurvature VALUE curved) The W3C document notes some problems with this approach, such as the inability to further partition the values. For example, we could not create fiat partitions dividing curved into highly curved, mildly curved, etc 8. This is a major disadvantage for representing phenotypes, as multiple levels of partitioning may be required to accurately represent the biology. On top of the listed problems, we would also add that this solution is ontologically unsatisfying, as we hold that these values correspond to types (also known as universals) and are instantiated multiple times in nature, and are best represented as owl Classes, not owl Individuals. 2.2 Value Partitions The second method described in the W3C technical note involves partitioning qualities into owl Classes. We can write the same example in Manchester Syntax as follows: Class: CurvatureValue EquivalentClass: Flat OR Curved This states that the class CurvatureValue is equivalent to the union of the 2 classes Flat and Curved - this also states the list is closed - there are no other classes that are also CurvatureValues. We could, however, create subclasses of Curved. In addition we can specify disjointness conditions: 6 for simplicity, we omit treatment of 3D objects that may have surfaces of different curvature, and assume any entity has a single curvature 7 Represented in the Fly Anatomy Ontologyhttp://obo.sourceforge.net/cgi-bin/detail.cgi?fly anatomy 8 in this example it may be possible to combine value partitions but this is inherently problematic
4 Disjoint: Flat Curved The functional property linking curvature individuals to individuals with curvature is the same as for the enumeration pattern: FunctionalProperty: hascurvature Range: CurvatureValue Manchester Syntax even has a macro feature for conveniently representing value partitions: ValuePartition: Curvature hascurvature [Flat Curved] This combines the unionof axioms with the disjointwith axioms into a handy syntactic idiom. So an instance of a curved fly wing could be represented as: Individual: fly-wing TYPE (Wing THAT hascurvature SOME Curved) We can also name the class expression: Class: CurvedWing EquivalentClass: Wing THAT hascurvature SOME Curved It is important to note that an important consequence of this pattern is that the existence of each individual of type CurvedWing also entails the existence of a distinct quality individual of type Curved. This is in contrast to the enumeration pattern, in which there is a single quality individual curved, which is shared by all curved things. This gives the class partition pattern various advantages over the enumeration of individuals pattern. It allows the quality classes to be further partitioned into more refined classes. We also hold that this pattern is ontologically preferable, and that distinct quality instances, such as my hair color or the shape of a particular organ, exist in reality. 2.3 Qualities and their Bearers Our approach, which has been dubbed the EQ model, is highly similar to the class partition pattern described above, but differs in a few key respects. Our treatment is based on a previously published formal analysis of qualities[?], and uses an ontology of qualities called PATO[?][?]. In our formalisation, a phenotype is one or more qualities which are borne by bearer entities in some organism 9. The relation between a quality and its bearer entity is one of inherence. An example of an instance of a phenotype is the particular instance of high curvature inhering in a particular instance of wing in a particular fruitfly. 9 the phenotype of an organism is influenced by the genetics of that organism, mediated by the environment; description of these crucial factors are omitted for reasons of space
5 C W i n g u r v e d i n s t a n c e _ o f i n s t a n c e _ o f i n h e r e s _ i n Fig. 1. The quality square: instances are depicted as rectangles, types as circles. Entity instances instantiate entity types (in this case, an eye). quality instances instantiate quality types (here the quality of curved, or high curvature). The quality instance inheres in the entity instance. The instances are unnamed. Note that sometimes qualities are referred to as properties, both colloquially and in the philosophical literature, but we avoid this usage, due to the contradictory use of the term property within the OWL community to denote what we would call a relation. Here we also distinguish between instances, the particular instances in reality, and the individuals that represent them. All qualities are linked to their bearers by means of the inherence relation, which can be represented in OWL as: FunctionalProperty: inheresin Domain: Quality Note that this approach has no need of relations such as hascolor (although these could optionally be added as sub-relations of inheresin). Rather than creating distinct class partitions for hascolor, hasshape and so on, we have a single subclass hierarchy, with all quality classes ultimately inheriting from the root Quality class. We can describe the fly wild-type phenotype high curvature wings using the class expression: A-1 Curved that inheresin some Wing We can also describe a curved wing: A-2 Wing that hasquality some Curved These two expressions are intuitively similar, but crucially distinct, in that the first represents a quality, and the second represents the three dimensional object that is the bearer of the quality. The existence of an instance of one entails the existence of some instance of the other at any given time. Thus either class is eligible as the description of the phenotype. We choose A-1 as the normal-form for phenotypes.
6 C H C M C Q S Our approach is most consistent with the second approach in the W3C note above, as we use owl Classes to denote our Quality universals. However, we differ from this approach in that we have no specific need for multiple relations such as hascolor, hassize, hasshape and so on. Our ontology of qualities is a single hierarchy, with all classes a subclass of the root class Quality. 3 Results 3.1 PATO: An Ontology of Qualities To support the description of phenotypes, we have developed an ontology of qualities called PATO 10. The ontology is constructed as far as possible according to OBO Foundry 11 principles, with each class in the ontology having a single is a (subclass) parent. The ontology is maintained in OBO format and is available from OBO, but is also available in OWL 12. s u b C l a s s O f h a p e u a l i t y u r v a t u r e s u b C l a s s O f s u b C l a s s O f C u r v e d F l a t s u b C l a s s O f s u b C l a s s O f i g h u r v a t u r e i l d u r v a t u r e Fig.2. Depiction of classes representing curvature in PATO 3.2 Granularity of Qualities The ontology is divided along granular levels [?] [?]. On the one hand we have physical qualities, such as mass, velocity and color; next we have cellular qualities such as ploidy (which inheres in a cell or cell nucleus by virtue of the number of chromosomes it has) and cellular potential (capability of differentiating into a range of other cell types; finally we have organismal qualities. Note that of course cells and organisms have physical qualities such as mass, by virtue of the physical qualities of their subparts. Physics is sufficient to give an account of 10 This originally stood for Phenotype and Trait Ontology. The full name is somewhat misleading as the classes in the ontology represent qualities, rather than full-blown phenotypes
7 physical qualities - however, other qualities are emergent and a description must be given at a higher level of granularity. Although the ontology is developed with the intent of use for phenotypes in biology and biomedicine, it is our hope that the physical quality sub-hierarchy of PATO will prove useful in other scientific realms. 3.3 Relational Qualities We follow [?] in distinguishing between monadic and relational qualities. The former are qualities that inhere in and depend on a single entity. The latter depend on some additional entity or entity type. For example, shape is a monadic quality because shapes exist in totality in a single shape-bearing. A quality such as sensitivity, by contrast, has a single bearer but is always with respect to some other entity or kind of entity (such as in, for example, sensitivity to sunlight). Neuhaus et al[?] represent relational qualities in a first-order logic framework using an inherence relation with a higher arity - the additional entity universals are referenced as additional arguments. As all relations in OWL are binary, we must introduce an additional relation to represent relational qualities - we call this the towards relation. Using this relation we can represent the phenotype skin photosensitivity to UV light using the class expression: A-1 Sensitive that towards some UltravioletLight and inheresin some Skin Note this is ontologically unsatisfactory, as any instance of photosensitivity is with respect to the universal UltravioletLight, rather than any one particular instance of a light wave or particle bundle. However, for the purposes of DL reasoning the representation is practical, and a reasoner such as Pellet will correctly reason that A-1 is subsumed by an expression such as A-2: A-2 Sensitive that towards some Radiation and inheresin some Organ We could instead use the UltravioletLight class as an individual, as in A-3: A-3 Sensitive that towards value UltravioletLight and inheresin some Skin However, this has the undesirable consequence of taking us to OWL Full, and will result in omissions in the reasoning process. 3.4 Temporal Durations Descriptions of phenotypes may be temporally qualified. The qualities in question may only inhere in the bearer entity over some interval - for example, organismal development involves changes in morphology of parts of organisms. Sometimes the quality may only be observed over a certain interval - for example, a particular assay of high blood pressure.
8 This raises interesting representation issues - all relations in OWL are binary, and there is no simple way to add a temporal index to binary instance-level relations, such as the relation between a quality and its bearer. There are a number of representation patterns for getting around this limitation [?] [?], such as reifying relations, but unfortunately they are all unsastifactory from an ontological and practical point of view. Consider an portion of tissue, such as the optic placode 13, that changes its shape from flat to curved or invaginated during a specific stage of development. How do we represent this using binary relations? We are unsatisfied with all the solutions afforded us within the constraints of binary relations - Our current compromise approach would be to represent this kind of anatomical entity as follows: A-4 OpticPlacode that hasquality some (Flat that during some Stage11) and hasquality some (Invaginated that during some Stage12) (note that here we have an object-centered representation rather than qualitycentered, using the hasquality relation, the inverse of inheresin) This approach is unsatisfactory because it involves stating artefactual classes such as flat during stage 11. This approach also causes problems when reasoning, as illustrated below: Assume also that Invaginated and Flat are disjoint (a reasonable assumption, given some fiat[?] quality boundary), and subclasses of Shape Class: Invaginated SubClassOf: Shape DisjointFrom: Flat Now we add the restriction that all organism parts have exactly one shape, as follows 14 : Class: OrganismPart SubClassOf: hasquality EXACTLY 1 Shape However, if flat and invaginated are disjoint classes, and we declare that each organism part can only assume a single shape, this will result A-4 being equivalent to the empty set, an undesirable result. An alternate representational paradigm involves representing time-slices of three dimensional objects (such as an organism part) at different times. These slices can then be related via binary relations to qualities that obtain over intervals. Each slice can then be linked back to the representation of the three dimensional object. This approach is sometimes called the perdurantist or 4- dimensionalist approach, and the time-slices are sometimes called histories or snapshots. 13 the part of an organism that will develop into the eye 14 the intent is to specify a Qualified Cardinality Constraint, as is allowed in OWL-1.1, but the authors are not sure if this is the correct for in the Manchester syntax
9 For example, we can represent an optic placode whose shape changes between flat and invaginated during a stage of development using the following class expression: A-5 OpticPlacode that hastimeslice some (Slice that hasquality some Flat and during Stage11) and hastimeslice some (Slice that hasquality some Invaginated and during Stage12) However, this solution is unsatisfactory from both an ontological and a practical perspective. On the one hand we must introduce additional artefacts, that have no place in upper ontologies such as BFO[?] and DOLCE. Our commitment to OBO Foundry principles leads us to reject the existence of unusual entities such as time-slices. In addition, the time slice paradigm must be anticipated in all ontologies used, such that for example the domain and range of relations can be 3D objects or time-slices of 3D objects. At best, the above construction is unwieldy and will require specific tool support if these constructs are to be generally useful. Another approach is to reify the relation in question, in this case inheresin, modeling it with an OWL class rather than an OWL property. We omit a discussion of this approach for reasons of space, but we find this solution even more unsatisfactory. An extension to OWL allowing would simplify this problem immensely. For example, if relations could take a third temporal argument as illustrated in the following expression: A-6 Organism that (hasquality some Flat at Stage11) and (hasquality some Invaginated at Stage12) The proposed AT operator could be used to index any relation at the class or instance level in OWL. We recognise that temporal extensions to DL expressivity would introduce many challenging problems, but we believe that this issue must be addressed sooner rather than later in order to accurately represent biology. One possibility is for 3-ary temporal expression such as the above to be compiled down to standard binary relation oriented description logics bu automatically introducing artefactual entities such as time-slices behind the scenes. 4 Discussion Representation of and reasoning over phenotypes is of huge importance to the life sciences and biomedicine. Our methodology for describing phenotypes through the composition of quality classes with classes of quality-bearers via the inherence relation works well for a variety of phenotypes, and promotes the reuse of ontologies in a modular way. The ontology of qualities, PATO, can be used across a variety of organisms for phenotypes across multiple levels of granularity. We find the ability to compose classes using description logic type expressions useful, and is a representational technique the general life sciences and
10 biomedical informatics and database communities would find useful in general. In particular, the elegance of Manchester OWL Syntax renders complex ontological representations accessible to ordinary mortals 15. This is a significant result for taking OWL out of the ivory tower into the real world. To illustrate this, here is a class expression involving multiple levels of nesting utilising classes from multiple ontologies to represent the phenotype high permeability of mitochondrial cristae in the axons of pyramidal cells in the CA1 hippocampal field: A-1 HighPermeability that inheresin some (MitochondrialCristae that partof some (Axon that partof some PyramidalCell that partof some CA1Field)) The ability to attach these class expressions to named phenotypes, such as those that may be found in the Mammalian Phenotype Ontology[?] and the Plant Trait Ontology[?] will be important for comparing data across organisms, and we have commenced an effort to create these expressions 16. We have found that limiting expressions to binary relations has negative implications for ontologically sound and intuitive representations of biology, particularly with respect to time. For lack of space we did not address various other important topics such as the representation of experimental assays and quantitative measurements. We also avoided discussion of representation on the complex relationship between genotypes, environments and phenotypes. However, we note that the methodology employed in this paper for representing phenotypes is also applicable to representing environment types. 5 Acknowledgements The ideas in this paper arose through discussions with many individuals, including Fabian Neuhaus, Barry Smith, Alan Rector, Nigam Shah, William Bug, Mark Gibson, John Day-Richter, Michael Ashburner and the members of the FlyBase consortium, Monte Westerfield and the members of ZFIN Consortium, Pankaj Jaiswal and the participants on the obo-phenotype mailing list. References 1. M. Ashburner, C. A. Ball, J. A. Blake, D. Botstein, H. Butler, J. M. Cherry, A. P. Davis, K. Dolinski, S. S. Dwight, J. T. Eppig, M. A. Harris, D. P. Hill, L. Issel- Tarver, A. Kasarskis, S. Lewis, J. C. Matese, J. E. Richardson, M. Ringwald, G. M. Rubin, and G. Sherlock. Gene ontology: tool for the unification of biology. the gene ontology consortium. Nat Genet, 25(1):25 9, Journal Article. 15 of course, the end-user does not see Manchester Syntax or any other OWL-oriented syntax, they work with intermediate representations and user interfaces such as Phenote 16
11 2. M. Ashburner, C. J. Mungall, and S. E. Lewis. Ontologies for biologists: a community model for the annotation of genomic data. Cold Spring Harb Symp Quant Biol, 68:227 35, Journal Article Review Review, Tutorial. 3. J. Bard, S.Y. Rhee, and M. Ashburner. An ontology for cell types. Genome Biology, Manuscript submitted, R. Drysdale. Phenotypic data in flybase. Brief Bioinform, 2(1):68 80, Journal Article. 5. G. V. Gkoutos, E. C. Green, A. M. Mallon, J. M. Hancock, and D. Davidson. Building mouse phenotype ontologies. Pac Symp Biocomput, pages , Journal Article. 6. G. V. Gkoutos, E. C. Green, A. M. Mallon, J. M. Hancock, and D. Davidson. Using ontologies to describe mouse phenotypes. Genome Biol, 6(1):R8, Journal Article. 7. Pierre Grenon. Temporal qualification and change with first-order binary predicates. In International Conference on Formal Ontology in Information Systems (FOIS 2006), Baltimore, Maryland (USA), Pierre Grenon, Barry Smith, and Louis Goldberg. Biodynamic ontology: applying BFO in the biomedical domain. Stud Health Technol Inform, 102:20 38, Purvesh Khatri and Sorin Draghici. Ontological analysis of gene expression data: current tools, limitations, and open problems. Bioinformatics, 21(18): , Sep Anand Kumar, Barry Smith, and Daniel D. Novotny. Biomedical informatics and granularity: Conference papers. Comp. Funct. Genomics, 5(6-7): , M.Horridge, N.Drummond, J.Goodwin, A.Rector, R.Stevens, and H.Wan. The manchester owl syntax. OWL: Experience and Directions 2006, F. Neuhaus, P. Grenon, and B. Smith. A formal theory of substances, qualities, and universals. In A. Varzi and L. Vieu, editors, Formal Ontology in Information Systems (FOIS?04), page 49?59, Turin, Italy, IOS Press. 13. Alan Rector. Representing specified values in OWL: value partitions and value sets. W3C note, W3C, May Alan Rector and Natasha Noy. Defining n-ary relations on the semantic web. W3C note, W3C, April Alan Rector, Jeremy Rogers, and Thomas Bittner. Granularity, scale and collectivity: when size does and does not matter. J. of Biomedical Informatics, 39(3): , C. Rosse and J.L.V. Mejino. A reference ontology for bioinformatics: The foundational model of anatomy. Journal of Biomedical Informatics 36: Barry Smith and Achille C. Varzi. Fiat and bona fide boundaries: Towards on ontology of spatially extended objects. In Spatial Information Theory, pages , Cynthia L Smith, Carroll-Ann W Goldsmith, and Janan T Eppig. The Mammalian Phenotype Ontology as a tool for annotating, analyzing and comparing phenotypic information. Genome Biol, 6(1):R7, Yukiko Yamazaki and Pankaj Jaiswal. Biological ontologies in rice databases. An introduction to the activities in Gramene and Oryzabase. Plant Cell Physiol, 46(1):63 68, Jan 2005.
An OWL Ontology for Quantum Mechanics
 An OWL Ontology for Quantum Mechanics Marcin Skulimowski Faculty of Physics and Applied Informatics, University of Lodz Pomorska 149/153, 90-236 Lodz, Poland mskulim@uni.lodz.pl Abstract. An OWL ontology
An OWL Ontology for Quantum Mechanics Marcin Skulimowski Faculty of Physics and Applied Informatics, University of Lodz Pomorska 149/153, 90-236 Lodz, Poland mskulim@uni.lodz.pl Abstract. An OWL ontology
CARO The Common Anatomy Reference Ontology
 1 CARO The Common Anatomy Reference Ontology Haendel, M.A., Neuhaus, F., Osumi-Sutherland, D.S., Mabee, P.M., Mejino J.L.V., Mungall, C.J., and Smith, B. 1.1 Abstract The Common Anatomy Reference Ontology
1 CARO The Common Anatomy Reference Ontology Haendel, M.A., Neuhaus, F., Osumi-Sutherland, D.S., Mabee, P.M., Mejino J.L.V., Mungall, C.J., and Smith, B. 1.1 Abstract The Common Anatomy Reference Ontology
Gabriella Rustici EMBL-EBI
 Towards a cellular phenotype ontology Gabriella Rustici EMBL-EBI gabry@ebi.ac.uk Systems Microscopy NoE at a glance The Systems Microscopy NoE (2011 2015) is a FP7 funded project, involving 15 research
Towards a cellular phenotype ontology Gabriella Rustici EMBL-EBI gabry@ebi.ac.uk Systems Microscopy NoE at a glance The Systems Microscopy NoE (2011 2015) is a FP7 funded project, involving 15 research
The anatomy of phenotype ontologies: principles, properties and applications
 Briefings in Bioinformatics, 19(5), 2018, 1008 1021 doi: 10.1093/bib/bbx035 Advance Access Publication Date: 6 April 2017 Paper The anatomy of phenotype ontologies: principles, properties and applications
Briefings in Bioinformatics, 19(5), 2018, 1008 1021 doi: 10.1093/bib/bbx035 Advance Access Publication Date: 6 April 2017 Paper The anatomy of phenotype ontologies: principles, properties and applications
Ontologies for nanotechnology. Colin Batchelor
 Ontologies for nanotechnology Colin Batchelor batchelorc@rsc.org 2009-03-25 Ontologies for nanotechnology: Outline What is an ontology and how do they work? What do people use ontologies for? How are we
Ontologies for nanotechnology Colin Batchelor batchelorc@rsc.org 2009-03-25 Ontologies for nanotechnology: Outline What is an ontology and how do they work? What do people use ontologies for? How are we
Estimation of Identification Methods of Gene Clusters Using GO Term Annotations from a Hierarchical Cluster Tree
 Estimation of Identification Methods of Gene Clusters Using GO Term Annotations from a Hierarchical Cluster Tree YOICHI YAMADA, YUKI MIYATA, MASANORI HIGASHIHARA*, KENJI SATOU Graduate School of Natural
Estimation of Identification Methods of Gene Clusters Using GO Term Annotations from a Hierarchical Cluster Tree YOICHI YAMADA, YUKI MIYATA, MASANORI HIGASHIHARA*, KENJI SATOU Graduate School of Natural
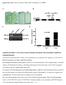 Supplemental Table 1. Primers used for qrt-pcr analysis Name abf2rtf abf2rtr abf3rtf abf3rtf abf4rtf abf4rtr ANAC096RTF ANAC096RTR abf2lp abf2rp abf3lp abf3lp abf4lp abf4rp ANAC096F(XbaI) ANAC096R(BamHI)
Supplemental Table 1. Primers used for qrt-pcr analysis Name abf2rtf abf2rtr abf3rtf abf3rtf abf4rtf abf4rtr ANAC096RTF ANAC096RTR abf2lp abf2rp abf3lp abf3lp abf4lp abf4rp ANAC096F(XbaI) ANAC096R(BamHI)
Introduction to the EMBL-EBI Ontology Lookup Service
 Introduction to the EMBL-EBI Ontology Lookup Service Simon Jupp jupp@ebi.ac.uk, @simonjupp Samples, Phenotypes and Ontologies Team European Bioinformatics Institute Cambridge, UK. Ontologies in the life
Introduction to the EMBL-EBI Ontology Lookup Service Simon Jupp jupp@ebi.ac.uk, @simonjupp Samples, Phenotypes and Ontologies Team European Bioinformatics Institute Cambridge, UK. Ontologies in the life
OWL Semantics COMP Sean Bechhofer Uli Sattler
 OWL Semantics COMP62342 Sean Bechhofer sean.bechhofer@manchester.ac.uk Uli Sattler uli.sattler@manchester.ac.uk 1 Toward Knowledge Formalization Acquisition Process Elicit tacit knowledge A set of terms/concepts
OWL Semantics COMP62342 Sean Bechhofer sean.bechhofer@manchester.ac.uk Uli Sattler uli.sattler@manchester.ac.uk 1 Toward Knowledge Formalization Acquisition Process Elicit tacit knowledge A set of terms/concepts
Document Navigation: Ontologies or Knowledge Organisation Systems
 Document Navigation: Ontologies or Knowledge Organisation Systems Simon Jupp - NETTAB 2007 BioHealth Informatics Group University of Manchester, UK Introduction Bioinformatics relies heavily on web for
Document Navigation: Ontologies or Knowledge Organisation Systems Simon Jupp - NETTAB 2007 BioHealth Informatics Group University of Manchester, UK Introduction Bioinformatics relies heavily on web for
GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón
 GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón What is GO? The Gene Ontology (GO) project is a collaborative effort to address the need for consistent descriptions of gene products in
GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón What is GO? The Gene Ontology (GO) project is a collaborative effort to address the need for consistent descriptions of gene products in
GOSAP: Gene Ontology Based Semantic Alignment of Biological Pathways
 GOSAP: Gene Ontology Based Semantic Alignment of Biological Pathways Jonas Gamalielsson and Björn Olsson Systems Biology Group, Skövde University, Box 407, Skövde, 54128, Sweden, [jonas.gamalielsson][bjorn.olsson]@his.se,
GOSAP: Gene Ontology Based Semantic Alignment of Biological Pathways Jonas Gamalielsson and Björn Olsson Systems Biology Group, Skövde University, Box 407, Skövde, 54128, Sweden, [jonas.gamalielsson][bjorn.olsson]@his.se,
Overview. An Introduction to applied ontology. Information Integration. The ideal situation. Intro and motivation: Why do we need ontologies?
 An Introduction to applied ontology Thomas Bittner bittner3@buffalo.edu Department of Philosophy Overview 1. Motivation for ontologies in information systems 2. What is an ontology? 3. Kinds of ontologies
An Introduction to applied ontology Thomas Bittner bittner3@buffalo.edu Department of Philosophy Overview 1. Motivation for ontologies in information systems 2. What is an ontology? 3. Kinds of ontologies
An evolutionary approach to Function
 JOURNAL OF BIOMEDICAL SEMANTICS PROCEEDINGS Open Access An evolutionary approach to Function Phillip Lord From Bio-Ontologies 2009: Knowledge in Biology Stockholm, Sweden. 28 June 2009 Correspondence:
JOURNAL OF BIOMEDICAL SEMANTICS PROCEEDINGS Open Access An evolutionary approach to Function Phillip Lord From Bio-Ontologies 2009: Knowledge in Biology Stockholm, Sweden. 28 June 2009 Correspondence:
Modeling Ontological Structures with Type Classes in Coq
 Modeling Ontological Structures with Type Classes in Coq Richard Dapoigny 1 Patrick Barlatier 1 1 LISTIC/Polytech Savoie University of Savoie, Annecy, (FRANCE) Cite as: Dapoigny, R., Barlatier, P. Modeling
Modeling Ontological Structures with Type Classes in Coq Richard Dapoigny 1 Patrick Barlatier 1 1 LISTIC/Polytech Savoie University of Savoie, Annecy, (FRANCE) Cite as: Dapoigny, R., Barlatier, P. Modeling
Using C-OWL for the Alignment and Merging of Medical Ontologies
 Using C-OWL for the Alignment and Merging of Medical Ontologies Heiner Stuckenschmidt 1, Frank van Harmelen 1 Paolo Bouquet 2,3, Fausto Giunchiglia 2,3, Luciano Serafini 3 1 Vrije Universiteit Amsterdam
Using C-OWL for the Alignment and Merging of Medical Ontologies Heiner Stuckenschmidt 1, Frank van Harmelen 1 Paolo Bouquet 2,3, Fausto Giunchiglia 2,3, Luciano Serafini 3 1 Vrije Universiteit Amsterdam
Evolution of a Foundational Model of Physiology: Symbolic Representation for Functional Bioinformatics
 MEDINFO 2004 M. Fieschi et al. (Eds) Amsterdam: IOS Press 2004 IMIA. All rights reserved Evolution of a Foundational Model of Physiology: Symbolic Representation for Functional Bioinformatics Daniel L.
MEDINFO 2004 M. Fieschi et al. (Eds) Amsterdam: IOS Press 2004 IMIA. All rights reserved Evolution of a Foundational Model of Physiology: Symbolic Representation for Functional Bioinformatics Daniel L.
An Upper Ontology of Event Classifications and Relations
 An Upper Ontology of Event Classifications and Relations Ken Kaneiwa and Michiaki Iwazume National Institute of Information and Communications Technology (NICT), Japan Ken Fukuda National Institute of
An Upper Ontology of Event Classifications and Relations Ken Kaneiwa and Michiaki Iwazume National Institute of Information and Communications Technology (NICT), Japan Ken Fukuda National Institute of
OWL Basics. Technologies for the Semantic Web. Building a Semantic Web. Ontology
 Technologies for the Semantic Web OWL Basics COMP60421 Sean Bechhofer University of Manchester sean.bechhofer@manchester.ac.uk Metadata Resources are marked-up with descriptions of their content. No good
Technologies for the Semantic Web OWL Basics COMP60421 Sean Bechhofer University of Manchester sean.bechhofer@manchester.ac.uk Metadata Resources are marked-up with descriptions of their content. No good
Extracting Modules from Ontologies: A Logic-based Approach
 Extracting Modules from Ontologies: A Logic-based Approach Bernardo Cuenca Grau, Ian Horrocks, Yevgeny Kazakov and Ulrike Sattler The University of Manchester School of Computer Science Manchester, M13
Extracting Modules from Ontologies: A Logic-based Approach Bernardo Cuenca Grau, Ian Horrocks, Yevgeny Kazakov and Ulrike Sattler The University of Manchester School of Computer Science Manchester, M13
The Role of Definitions in Biomedical Concept Representation
 The Role of Definitions in Biomedical Concept Representation Joshua Michael, José L. V. Mejino Jr., M.D., and Cornelius Rosse, M.D., D.Sc. Structural Informatics Group, Department of Biological Structure,
The Role of Definitions in Biomedical Concept Representation Joshua Michael, José L. V. Mejino Jr., M.D., and Cornelius Rosse, M.D., D.Sc. Structural Informatics Group, Department of Biological Structure,
Designing and Evaluating Generic Ontologies
 Designing and Evaluating Generic Ontologies Michael Grüninger Department of Industrial Engineering University of Toronto gruninger@ie.utoronto.ca August 28, 2007 1 Introduction One of the many uses of
Designing and Evaluating Generic Ontologies Michael Grüninger Department of Industrial Engineering University of Toronto gruninger@ie.utoronto.ca August 28, 2007 1 Introduction One of the many uses of
Modeling Ontologies Using OWL, Description Graphs, and Rules
 Modeling Ontologies Using OWL, Description Graphs, and Rules Boris Motik 1, Bernardo Cuenca Grau 1, Ian Horrocks 1, and Ulrike Sattler 2 1 University of Oxford, UK 2 University of Manchester, UK 1 Introduction
Modeling Ontologies Using OWL, Description Graphs, and Rules Boris Motik 1, Bernardo Cuenca Grau 1, Ian Horrocks 1, and Ulrike Sattler 2 1 University of Oxford, UK 2 University of Manchester, UK 1 Introduction
Integrating phenotype ontologies with PhenomeNET
 Integrating phenotype ontologies with PhenomeNET Miguel Angel Rodríguez García 1, Georgios V Gkoutos 2, Paul N Schofield 3, and Robert Hoehndorf 1 1 Computational Bioscience Research Center, King Abdullah
Integrating phenotype ontologies with PhenomeNET Miguel Angel Rodríguez García 1, Georgios V Gkoutos 2, Paul N Schofield 3, and Robert Hoehndorf 1 1 Computational Bioscience Research Center, King Abdullah
OWL Semantics. COMP60421 Sean Bechhofer University of Manchester
 OWL Semantics COMP60421 Sean Bechhofer University of Manchester sean.bechhofer@manchester.ac.uk 1 Technologies for the Semantic Web Metadata Resources are marked-up with descriptions of their content.
OWL Semantics COMP60421 Sean Bechhofer University of Manchester sean.bechhofer@manchester.ac.uk 1 Technologies for the Semantic Web Metadata Resources are marked-up with descriptions of their content.
Modular Reuse of Ontologies: Theory and Practice
 Journal of Artificial Intelligence Research 31 (2008) 273-318 Submitted 07/07; published 02/08 Modular Reuse of Ontologies: Theory and Practice Bernardo Cuenca Grau Ian Horrocks Yevgeny Kazakov Oxford
Journal of Artificial Intelligence Research 31 (2008) 273-318 Submitted 07/07; published 02/08 Modular Reuse of Ontologies: Theory and Practice Bernardo Cuenca Grau Ian Horrocks Yevgeny Kazakov Oxford
Week 4. COMP62342 Sean Bechhofer, Uli Sattler
 Week 4 COMP62342 Sean Bechhofer, Uli Sattler sean.bechhofer@manchester.ac.uk, uli.sattler@manchester.ac.uk Today Some clarifications from last week s coursework More on reasoning: extension of the tableau
Week 4 COMP62342 Sean Bechhofer, Uli Sattler sean.bechhofer@manchester.ac.uk, uli.sattler@manchester.ac.uk Today Some clarifications from last week s coursework More on reasoning: extension of the tableau
BSC 1010C Biology I. Themes in the Study of Life Chapter 1
 BSC 1010C Biology I Themes in the Study of Life Chapter 1 Objectives Distinguish among the three domains of life. Distinguish between the Levels of Biological Organization. Note the differences in the
BSC 1010C Biology I Themes in the Study of Life Chapter 1 Objectives Distinguish among the three domains of life. Distinguish between the Levels of Biological Organization. Note the differences in the
Knowledge Sharing. A conceptualization is a map from the problem domain into the representation. A conceptualization specifies:
 Knowledge Sharing A conceptualization is a map from the problem domain into the representation. A conceptualization specifies: What sorts of individuals are being modeled The vocabulary for specifying
Knowledge Sharing A conceptualization is a map from the problem domain into the representation. A conceptualization specifies: What sorts of individuals are being modeled The vocabulary for specifying
Classification of Gene Expression Profiles: Comparison of K-means and Expectation Maximization Algorithms
 Eighth International Conference on Hybrid Intelligent Systems Classification of Gene Expression Profiles: Comparison of K-means and Expectation Maximization Algorithms Cristina Rubio-Escudero Dpto. Lenguajes
Eighth International Conference on Hybrid Intelligent Systems Classification of Gene Expression Profiles: Comparison of K-means and Expectation Maximization Algorithms Cristina Rubio-Escudero Dpto. Lenguajes
Student Handout Fruit Fly Ethomics & Genomics
 Student Handout Fruit Fly Ethomics & Genomics Summary of Laboratory Exercise In this laboratory unit, students will connect behavioral phenotypes to their underlying genes and molecules in the model genetic
Student Handout Fruit Fly Ethomics & Genomics Summary of Laboratory Exercise In this laboratory unit, students will connect behavioral phenotypes to their underlying genes and molecules in the model genetic
Web Ontology Language (OWL)
 Web Ontology Language (OWL) Need meaning beyond an object-oriented type system RDF (with RDFS) captures the basics, approximating an object-oriented type system OWL provides some of the rest OWL standardizes
Web Ontology Language (OWL) Need meaning beyond an object-oriented type system RDF (with RDFS) captures the basics, approximating an object-oriented type system OWL provides some of the rest OWL standardizes
A theory of granular parthood based on qualitative cardinality and size measures
 A theory of granular parthood based on qualitative cardinality and size measures Thomas BITTNER 1,2,3,4 and Maureen DONNELLY 1,3 1 Department of Philosophy, 2 Department of Geography 3 New York State Center
A theory of granular parthood based on qualitative cardinality and size measures Thomas BITTNER 1,2,3,4 and Maureen DONNELLY 1,3 1 Department of Philosophy, 2 Department of Geography 3 New York State Center
Axioms for parthood and containment relations in bio-ontologies
 Axioms for parthood and containment relations in bio-ontologies Thomas Bittner Institute for Formal Ontology and Medical Information Science University of Leipzig thomas.bittner@ifomis.uni-leipzig.de Abstract
Axioms for parthood and containment relations in bio-ontologies Thomas Bittner Institute for Formal Ontology and Medical Information Science University of Leipzig thomas.bittner@ifomis.uni-leipzig.de Abstract
Computational Biology Course Descriptions 12-14
 Computational Biology Course Descriptions 12-14 Course Number and Title INTRODUCTORY COURSES BIO 311C: Introductory Biology I BIO 311D: Introductory Biology II BIO 325: Genetics CH 301: Principles of Chemistry
Computational Biology Course Descriptions 12-14 Course Number and Title INTRODUCTORY COURSES BIO 311C: Introductory Biology I BIO 311D: Introductory Biology II BIO 325: Genetics CH 301: Principles of Chemistry
Knowledge Evolution, Inc Seaton Street NW Washington DC 20009, United States
 Supporting Information Toward Automated Inventory Modeling in Life Cycle Assessment: The Utility of Semantic Data Modeling to Predict Real-World Chemical Production Vinit K. Mittal 1, Sidney C. Bailin
Supporting Information Toward Automated Inventory Modeling in Life Cycle Assessment: The Utility of Semantic Data Modeling to Predict Real-World Chemical Production Vinit K. Mittal 1, Sidney C. Bailin
86 Part 4 SUMMARY INTRODUCTION
 86 Part 4 Chapter # AN INTEGRATION OF THE DESCRIPTIONS OF GENE NETWORKS AND THEIR MODELS PRESENTED IN SIGMOID (CELLERATOR) AND GENENET Podkolodny N.L. *1, 2, Podkolodnaya N.N. 1, Miginsky D.S. 1, Poplavsky
86 Part 4 Chapter # AN INTEGRATION OF THE DESCRIPTIONS OF GENE NETWORKS AND THEIR MODELS PRESENTED IN SIGMOID (CELLERATOR) AND GENENET Podkolodny N.L. *1, 2, Podkolodnaya N.N. 1, Miginsky D.S. 1, Poplavsky
Object Oriented Knowledge Bases in Logic Programming. Vinay K. Chaudhri Stijn Heymans Michael Weseel Son Cao Tran
 Object Oriented Knowledge Bases in Logic Programming Vinay K. Chaudhri Stijn Heymans Michael Weseel Son Cao Tran 1 Acknowledgment This work is funded by Vulcan Inc. and SRI International See http://www.vulcan.com
Object Oriented Knowledge Bases in Logic Programming Vinay K. Chaudhri Stijn Heymans Michael Weseel Son Cao Tran 1 Acknowledgment This work is funded by Vulcan Inc. and SRI International See http://www.vulcan.com
Knowledge Representation for the Semantic Web Lecture 2: Description Logics I
 Knowledge Representation for the Semantic Web Lecture 2: Description Logics I Daria Stepanova slides based on Reasoning Web 2011 tutorial Foundations of Description Logics and OWL by S. Rudolph Max Planck
Knowledge Representation for the Semantic Web Lecture 2: Description Logics I Daria Stepanova slides based on Reasoning Web 2011 tutorial Foundations of Description Logics and OWL by S. Rudolph Max Planck
The Systems Biology Graphical Notation
 http://www.sbgn.org/ The Systems Biology Graphical Notation Nicolas Le Novère, EMBL-EBI (on the behalf of SBGN editors, authors and contributors) Different representations of a pathway No temporal sequence
http://www.sbgn.org/ The Systems Biology Graphical Notation Nicolas Le Novère, EMBL-EBI (on the behalf of SBGN editors, authors and contributors) Different representations of a pathway No temporal sequence
Can Vector Space Bases Model Context?
 Can Vector Space Bases Model Context? Massimo Melucci University of Padua Department of Information Engineering Via Gradenigo, 6/a 35031 Padova Italy melo@dei.unipd.it Abstract Current Information Retrieval
Can Vector Space Bases Model Context? Massimo Melucci University of Padua Department of Information Engineering Via Gradenigo, 6/a 35031 Padova Italy melo@dei.unipd.it Abstract Current Information Retrieval
F1 Parent Cell R R. Name Period. Concept 15.1 Mendelian inheritance has its physical basis in the behavior of chromosomes
 Name Period Concept 15.1 Mendelian inheritance has its physical basis in the behavior of chromosomes 1. What is the chromosome theory of inheritance? 2. Explain the law of segregation. Use two different
Name Period Concept 15.1 Mendelian inheritance has its physical basis in the behavior of chromosomes 1. What is the chromosome theory of inheritance? 2. Explain the law of segregation. Use two different
First Order Logic (1A) Young W. Lim 11/18/13
 Copyright (c) 2013. Permission is granted to copy, distribute and/or modify this document under the terms of the GNU Free Documentation License, Version 1.2 or any later version published by the Free Software
Copyright (c) 2013. Permission is granted to copy, distribute and/or modify this document under the terms of the GNU Free Documentation License, Version 1.2 or any later version published by the Free Software
AP Biology Essential Knowledge Cards BIG IDEA 1
 AP Biology Essential Knowledge Cards BIG IDEA 1 Essential knowledge 1.A.1: Natural selection is a major mechanism of evolution. Essential knowledge 1.A.4: Biological evolution is supported by scientific
AP Biology Essential Knowledge Cards BIG IDEA 1 Essential knowledge 1.A.1: Natural selection is a major mechanism of evolution. Essential knowledge 1.A.4: Biological evolution is supported by scientific
SPA for quantitative analysis: Lecture 6 Modelling Biological Processes
 1/ 223 SPA for quantitative analysis: Lecture 6 Modelling Biological Processes Jane Hillston LFCS, School of Informatics The University of Edinburgh Scotland 7th March 2013 Outline 2/ 223 1 Introduction
1/ 223 SPA for quantitative analysis: Lecture 6 Modelling Biological Processes Jane Hillston LFCS, School of Informatics The University of Edinburgh Scotland 7th March 2013 Outline 2/ 223 1 Introduction
Computational Structural Bioinformatics
 Computational Structural Bioinformatics ECS129 Instructor: Patrice Koehl http://koehllab.genomecenter.ucdavis.edu/teaching/ecs129 koehl@cs.ucdavis.edu Learning curve Math / CS Biology/ Chemistry Pre-requisite
Computational Structural Bioinformatics ECS129 Instructor: Patrice Koehl http://koehllab.genomecenter.ucdavis.edu/teaching/ecs129 koehl@cs.ucdavis.edu Learning curve Math / CS Biology/ Chemistry Pre-requisite
HAWAII CONTENT AND PERFORMANCE STANDARDS BIOLOGICAL SCIENCE
 HAWAII CONTENT AND PERFORMANCE STANDARDS BIOLOGICAL SCIENCE Correlated to BIOLOGY: CYCLES OF LIFE 2006 5910 Rice Creek Parkway, Suite 1000 Shoreview, Minnesota 55126 Telephone (800) 328-2560 www.agsglobe.com
HAWAII CONTENT AND PERFORMANCE STANDARDS BIOLOGICAL SCIENCE Correlated to BIOLOGY: CYCLES OF LIFE 2006 5910 Rice Creek Parkway, Suite 1000 Shoreview, Minnesota 55126 Telephone (800) 328-2560 www.agsglobe.com
Standards Map Basic Comprehensive Program Science Grade Seven Focus on Life Sciences SE/TE: , SE/TE: ,
 Publisher: Pearson Program Title: Biology: Exploring Life 2006 Components: 0-13-250-925-3 (SE), 0-13-250-883-4 (TE) Grade Level(s): 7-12 s Map Basic Comprehensive Program Science Grade Seven Focus on Life
Publisher: Pearson Program Title: Biology: Exploring Life 2006 Components: 0-13-250-925-3 (SE), 0-13-250-883-4 (TE) Grade Level(s): 7-12 s Map Basic Comprehensive Program Science Grade Seven Focus on Life
BMD645. Integration of Omics
 BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
Description logics. Description Logics. Applications. Outline. Syntax - AL. Tbox and Abox
 Description Logics Description logics A family of KR formalisms, based on FOPL decidable, supported by automatic reasoning systems Used for modelling of application domains Classification of concepts and
Description Logics Description logics A family of KR formalisms, based on FOPL decidable, supported by automatic reasoning systems Used for modelling of application domains Classification of concepts and
Probabilistic Ontologies: Logical Approach
 Probabilistic Ontologies: Logical Approach Pavel Klinov Applied Artificial Intelligence Lab ECE Department University of Cincinnati Agenda Why do we study ontologies? Uncertainty Probabilistic ontologies
Probabilistic Ontologies: Logical Approach Pavel Klinov Applied Artificial Intelligence Lab ECE Department University of Cincinnati Agenda Why do we study ontologies? Uncertainty Probabilistic ontologies
Affordances in Representing the Behaviour of Event-Based Systems
 Affordances in Representing the Behaviour of Event-Based Systems Fahim T. IMAM a,1, Thomas R. DEAN b a School of Computing, Queen s University, Canada b Department of Electrical and Computer Engineering,
Affordances in Representing the Behaviour of Event-Based Systems Fahim T. IMAM a,1, Thomas R. DEAN b a School of Computing, Queen s University, Canada b Department of Electrical and Computer Engineering,
Description Logics. Foundations of Propositional Logic. franconi. Enrico Franconi
 (1/27) Description Logics Foundations of Propositional Logic Enrico Franconi franconi@cs.man.ac.uk http://www.cs.man.ac.uk/ franconi Department of Computer Science, University of Manchester (2/27) Knowledge
(1/27) Description Logics Foundations of Propositional Logic Enrico Franconi franconi@cs.man.ac.uk http://www.cs.man.ac.uk/ franconi Department of Computer Science, University of Manchester (2/27) Knowledge
Enduring understanding 1.A: Change in the genetic makeup of a population over time is evolution.
 The AP Biology course is designed to enable you to develop advanced inquiry and reasoning skills, such as designing a plan for collecting data, analyzing data, applying mathematical routines, and connecting
The AP Biology course is designed to enable you to develop advanced inquiry and reasoning skills, such as designing a plan for collecting data, analyzing data, applying mathematical routines, and connecting
7.014 Problem Set 6. Question 1. MIT Department of Biology Introductory Biology, Spring 2004
 MIT Department of Biology 7.014 Introductory Biology, Spring 2004 Name: 7.014 Problem Set 6 Please print out this problem set and record your answers on the printed copy. Problem sets will not be accepted
MIT Department of Biology 7.014 Introductory Biology, Spring 2004 Name: 7.014 Problem Set 6 Please print out this problem set and record your answers on the printed copy. Problem sets will not be accepted
An Approach to the Anatomical Correlation of Species through the Foundational Model of Anatomy
 An Approach to the Anatomical Correlation of Species through the Foundational Model of Anatomy Ravensara S. Travillian MA ; Cornelius Rosse MD, DSc ; Linda G. Shapiro PhD Structural Informatics Group,
An Approach to the Anatomical Correlation of Species through the Foundational Model of Anatomy Ravensara S. Travillian MA ; Cornelius Rosse MD, DSc ; Linda G. Shapiro PhD Structural Informatics Group,
A Logical Formulation of the Granular Data Model
 2008 IEEE International Conference on Data Mining Workshops A Logical Formulation of the Granular Data Model Tuan-Fang Fan Department of Computer Science and Information Engineering National Penghu University
2008 IEEE International Conference on Data Mining Workshops A Logical Formulation of the Granular Data Model Tuan-Fang Fan Department of Computer Science and Information Engineering National Penghu University
A Logical Framework for Modularity of Ontologies
 A Logical Framework for Modularity of Ontologies Bernardo Cuenca Grau, Ian Horrocks, Yevgeny Kazakov and Ulrike Sattler The University of Manchester School of Computer Science Manchester, M13 9PL, UK {bcg,
A Logical Framework for Modularity of Ontologies Bernardo Cuenca Grau, Ian Horrocks, Yevgeny Kazakov and Ulrike Sattler The University of Manchester School of Computer Science Manchester, M13 9PL, UK {bcg,
Francisco M. Couto Mário J. Silva Pedro Coutinho
 Francisco M. Couto Mário J. Silva Pedro Coutinho DI FCUL TR 03 29 Departamento de Informática Faculdade de Ciências da Universidade de Lisboa Campo Grande, 1749 016 Lisboa Portugal Technical reports are
Francisco M. Couto Mário J. Silva Pedro Coutinho DI FCUL TR 03 29 Departamento de Informática Faculdade de Ciências da Universidade de Lisboa Campo Grande, 1749 016 Lisboa Portugal Technical reports are
An Introduction to the Use of Ontologies in Linking Evolutionary Phenotypes to Genetics. Paula Mabee University of South Dakota
 An Introduction to the Use of Ontologies in Linking Evolutionary Phenotypes to Genetics Paula Mabee University of South Dakota Shared ontologies & syntax connects models to humans Model organism Human
An Introduction to the Use of Ontologies in Linking Evolutionary Phenotypes to Genetics Paula Mabee University of South Dakota Shared ontologies & syntax connects models to humans Model organism Human
Practice Test on Cell Biology (the REAL test is on Friday the 17th) KEY LO: Describe and explain the Central Dogma. SLE: Meet NGSS.
 Practice Test on Cell Biology (the REAL test is on Friday the 17th) KEY LO: Describe and explain the Central Dogma. SLE: Meet NGSS. On the real test, you will be given this chart of the genetic code (from
Practice Test on Cell Biology (the REAL test is on Friday the 17th) KEY LO: Describe and explain the Central Dogma. SLE: Meet NGSS. On the real test, you will be given this chart of the genetic code (from
A conceptualization is a map from the problem domain into the representation. A conceptualization specifies:
 Knowledge Sharing A conceptualization is a map from the problem domain into the representation. A conceptualization specifies: What sorts of individuals are being modeled The vocabulary for specifying
Knowledge Sharing A conceptualization is a map from the problem domain into the representation. A conceptualization specifies: What sorts of individuals are being modeled The vocabulary for specifying
When and Why to use a Classifier?
 1 When and Why to use a Classifier? Alan Rector with acknowledgement to Jeremy Rogers, Pieter Zanstra,, & the GALEN Consortium Nick Drummond, Matthew Horridge, Hai Wang in CO-ODE/ ODE/HyOntUSE Information
1 When and Why to use a Classifier? Alan Rector with acknowledgement to Jeremy Rogers, Pieter Zanstra,, & the GALEN Consortium Nick Drummond, Matthew Horridge, Hai Wang in CO-ODE/ ODE/HyOntUSE Information
USING BLAST TO IDENTIFY PROTEINS THAT ARE EVOLUTIONARILY RELATED ACROSS SPECIES
 USING BLAST TO IDENTIFY PROTEINS THAT ARE EVOLUTIONARILY RELATED ACROSS SPECIES HOW CAN BIOINFORMATICS BE USED AS A TOOL TO DETERMINE EVOLUTIONARY RELATIONSHPS AND TO BETTER UNDERSTAND PROTEIN HERITAGE?
USING BLAST TO IDENTIFY PROTEINS THAT ARE EVOLUTIONARILY RELATED ACROSS SPECIES HOW CAN BIOINFORMATICS BE USED AS A TOOL TO DETERMINE EVOLUTIONARY RELATIONSHPS AND TO BETTER UNDERSTAND PROTEIN HERITAGE?
Gene Ontology. Shifra Ben-Dor. Weizmann Institute of Science
 Gene Ontology Shifra Ben-Dor Weizmann Institute of Science Outline of Session What is GO (Gene Ontology)? What tools do we use to work with it? Combination of GO with other analyses What is Ontology? 1700s
Gene Ontology Shifra Ben-Dor Weizmann Institute of Science Outline of Session What is GO (Gene Ontology)? What tools do we use to work with it? Combination of GO with other analyses What is Ontology? 1700s
The Bayesian Ontology Language BEL
 Journal of Automated Reasoning manuscript No. (will be inserted by the editor) The Bayesian Ontology Language BEL İsmail İlkan Ceylan Rafael Peñaloza Received: date / Accepted: date Abstract We introduce
Journal of Automated Reasoning manuscript No. (will be inserted by the editor) The Bayesian Ontology Language BEL İsmail İlkan Ceylan Rafael Peñaloza Received: date / Accepted: date Abstract We introduce
Propositional Logic. Testing, Quality Assurance, and Maintenance Winter Prof. Arie Gurfinkel
 Propositional Logic Testing, Quality Assurance, and Maintenance Winter 2018 Prof. Arie Gurfinkel References Chpater 1 of Logic for Computer Scientists http://www.springerlink.com/content/978-0-8176-4762-9/
Propositional Logic Testing, Quality Assurance, and Maintenance Winter 2018 Prof. Arie Gurfinkel References Chpater 1 of Logic for Computer Scientists http://www.springerlink.com/content/978-0-8176-4762-9/
Completing Description Logic Knowledge Bases using Formal Concept Analysis
 Completing Description Logic Knowledge Bases using Formal Concept Analysis Franz Baader 1, Bernhard Ganter 1, Ulrike Sattler 2 and Barış Sertkaya 1 1 TU Dresden, Germany 2 The University of Manchester,
Completing Description Logic Knowledge Bases using Formal Concept Analysis Franz Baader 1, Bernhard Ganter 1, Ulrike Sattler 2 and Barış Sertkaya 1 1 TU Dresden, Germany 2 The University of Manchester,
AN EVIDENCE ONTOLOGY FOR USE IN PATHWAY/GENOME DATABASES
 AN EVIDENCE ONTOLOGY FOR USE IN PATHWAY/GENOME DATABASES Appears in Pacific Symposium on Biocomputing 2004 Pages 190-201, World Scientific Publishers, Singapore P.D. KARP, S. PALEY, C.J. KRIEGER SRI International,
AN EVIDENCE ONTOLOGY FOR USE IN PATHWAY/GENOME DATABASES Appears in Pacific Symposium on Biocomputing 2004 Pages 190-201, World Scientific Publishers, Singapore P.D. KARP, S. PALEY, C.J. KRIEGER SRI International,
Gene Ontology and overrepresentation analysis
 Gene Ontology and overrepresentation analysis Kjell Petersen J Express Microarray analysis course Oslo December 2009 Presentation adapted from Endre Anderssen and Vidar Beisvåg NMC Trondheim Overview How
Gene Ontology and overrepresentation analysis Kjell Petersen J Express Microarray analysis course Oslo December 2009 Presentation adapted from Endre Anderssen and Vidar Beisvåg NMC Trondheim Overview How
185.A09 Advanced Mathematical Logic
 185.A09 Advanced Mathematical Logic www.volny.cz/behounek/logic/teaching/mathlog13 Libor Běhounek, behounek@cs.cas.cz Lecture #1, October 15, 2013 Organizational matters Study materials will be posted
185.A09 Advanced Mathematical Logic www.volny.cz/behounek/logic/teaching/mathlog13 Libor Běhounek, behounek@cs.cas.cz Lecture #1, October 15, 2013 Organizational matters Study materials will be posted
CEX and MEX: Logical Diff and Semantic Module Extraction in a Fragment of OWL
 CEX and MEX: Logical Diff and Semantic Module Extraction in a Fragment of OWL Boris Konev 1, Carsten Lutz 2, Dirk Walther 1, and Frank Wolter 1 1 University of Liverpool, UK, {konev, dwalther, wolter}@liverpool.ac.uk
CEX and MEX: Logical Diff and Semantic Module Extraction in a Fragment of OWL Boris Konev 1, Carsten Lutz 2, Dirk Walther 1, and Frank Wolter 1 1 University of Liverpool, UK, {konev, dwalther, wolter}@liverpool.ac.uk
TEST SUMMARY AND FRAMEWORK TEST SUMMARY
 Washington Educator Skills Tests Endorsements (WEST E) TEST SUMMARY AND FRAMEWORK TEST SUMMARY BIOLOGY Copyright 2014 by the Washington Professional Educator Standards Board 1 Washington Educator Skills
Washington Educator Skills Tests Endorsements (WEST E) TEST SUMMARY AND FRAMEWORK TEST SUMMARY BIOLOGY Copyright 2014 by the Washington Professional Educator Standards Board 1 Washington Educator Skills
Essential knowledge 1.A.2: Natural selection
 Appendix C AP Biology Concepts at a Glance Big Idea 1: The process of evolution drives the diversity and unity of life. Enduring understanding 1.A: Change in the genetic makeup of a population over time
Appendix C AP Biology Concepts at a Glance Big Idea 1: The process of evolution drives the diversity and unity of life. Enduring understanding 1.A: Change in the genetic makeup of a population over time
Big Idea 1: The process of evolution drives the diversity and unity of life.
 Big Idea 1: The process of evolution drives the diversity and unity of life. understanding 1.A: Change in the genetic makeup of a population over time is evolution. 1.A.1: Natural selection is a major
Big Idea 1: The process of evolution drives the diversity and unity of life. understanding 1.A: Change in the genetic makeup of a population over time is evolution. 1.A.1: Natural selection is a major
Conceptual Modeling of Formal and Material Relations Applied to Ontologies
 Conceptual Modeling of Formal and Material Relations Applied to Ontologies Ricardo Ramos Linck, Guilherme Schievelbein and Mara Abel Institute of Informatics Universidade Federal do Rio Grande do Sul (UFRGS)
Conceptual Modeling of Formal and Material Relations Applied to Ontologies Ricardo Ramos Linck, Guilherme Schievelbein and Mara Abel Institute of Informatics Universidade Federal do Rio Grande do Sul (UFRGS)
Heredity and Genetics WKSH
 Chapter 6, Section 3 Heredity and Genetics WKSH KEY CONCEPT Mendel s research showed that traits are inherited as discrete units. Vocabulary trait purebred law of segregation genetics cross MAIN IDEA:
Chapter 6, Section 3 Heredity and Genetics WKSH KEY CONCEPT Mendel s research showed that traits are inherited as discrete units. Vocabulary trait purebred law of segregation genetics cross MAIN IDEA:
7. Propositional Logic. Wolfram Burgard and Bernhard Nebel
 Foundations of AI 7. Propositional Logic Rational Thinking, Logic, Resolution Wolfram Burgard and Bernhard Nebel Contents Agents that think rationally The wumpus world Propositional logic: syntax and semantics
Foundations of AI 7. Propositional Logic Rational Thinking, Logic, Resolution Wolfram Burgard and Bernhard Nebel Contents Agents that think rationally The wumpus world Propositional logic: syntax and semantics
AP Curriculum Framework with Learning Objectives
 Big Ideas Big Idea 1: The process of evolution drives the diversity and unity of life. AP Curriculum Framework with Learning Objectives Understanding 1.A: Change in the genetic makeup of a population over
Big Ideas Big Idea 1: The process of evolution drives the diversity and unity of life. AP Curriculum Framework with Learning Objectives Understanding 1.A: Change in the genetic makeup of a population over
Knowledge Representation and Reasoning
 Knowledge Representation and Reasoning Geraint A. Wiggins Professor of Computational Creativity Department of Computer Science Vrije Universiteit Brussel Objectives Knowledge Representation in Logic The
Knowledge Representation and Reasoning Geraint A. Wiggins Professor of Computational Creativity Department of Computer Science Vrije Universiteit Brussel Objectives Knowledge Representation in Logic The
Structures and Functions of Living Organisms (LS1)
 EALR 4: Big Idea: Core Content: Life Science Structures and Functions of Living Organisms (LS1) Processes Within Cells In prior grades students learned that all living systems are composed of cells which
EALR 4: Big Idea: Core Content: Life Science Structures and Functions of Living Organisms (LS1) Processes Within Cells In prior grades students learned that all living systems are composed of cells which
Modal logics: an introduction
 Modal logics: an introduction Valentin Goranko DTU Informatics October 2010 Outline Non-classical logics in AI. Variety of modal logics. Brief historical remarks. Basic generic modal logic: syntax and
Modal logics: an introduction Valentin Goranko DTU Informatics October 2010 Outline Non-classical logics in AI. Variety of modal logics. Brief historical remarks. Basic generic modal logic: syntax and
Knowledge Representation
 INF5390 Kunstig intelligens Knowledge Representation Roar Fjellheim Outline General ontology Categories and objects Events and processes Reasoning systems Internet shopping world Summary Extracts from
INF5390 Kunstig intelligens Knowledge Representation Roar Fjellheim Outline General ontology Categories and objects Events and processes Reasoning systems Internet shopping world Summary Extracts from
Semantics and Inference for Probabilistic Ontologies
 Semantics and Inference for Probabilistic Ontologies Fabrizio Riguzzi, Elena Bellodi, Evelina Lamma, and Riccardo Zese ENDIF University of Ferrara, Italy, email: {fabrizio.riguzzi, elena.bellodi, evelina.lamma}@unife.it,
Semantics and Inference for Probabilistic Ontologies Fabrizio Riguzzi, Elena Bellodi, Evelina Lamma, and Riccardo Zese ENDIF University of Ferrara, Italy, email: {fabrizio.riguzzi, elena.bellodi, evelina.lamma}@unife.it,
Description Logics. an introduction into its basic ideas
 Description Logics an introduction into its basic ideas A. Heußner WS 2003/2004 Preview: Basic Idea: from Network Based Structures to DL AL : Syntax / Semantics Enhancements of AL Terminologies (TBox)
Description Logics an introduction into its basic ideas A. Heußner WS 2003/2004 Preview: Basic Idea: from Network Based Structures to DL AL : Syntax / Semantics Enhancements of AL Terminologies (TBox)
From Constructibility and Absoluteness to Computability and Domain Independence
 From Constructibility and Absoluteness to Computability and Domain Independence Arnon Avron School of Computer Science Tel Aviv University, Tel Aviv 69978, Israel aa@math.tau.ac.il Abstract. Gödel s main
From Constructibility and Absoluteness to Computability and Domain Independence Arnon Avron School of Computer Science Tel Aviv University, Tel Aviv 69978, Israel aa@math.tau.ac.il Abstract. Gödel s main
RDF and Logic: Reasoning and Extension
 RDF and Logic: Reasoning and Extension Jos de Bruijn Faculty of Computer Science, Free University of Bozen-Bolzano, Italy debruijn@inf.unibz.it Stijn Heymans Digital Enterprise Research Institute (DERI),
RDF and Logic: Reasoning and Extension Jos de Bruijn Faculty of Computer Science, Free University of Bozen-Bolzano, Italy debruijn@inf.unibz.it Stijn Heymans Digital Enterprise Research Institute (DERI),
Georgia Performance Standards Framework for Science GRADE 7
 The following instructional plan is part of a GaDOE collection of Unit Frameworks, Performance Tasks, examples of Student Work, and Teacher Commentary. Many more GaDOE approved instructional plans are
The following instructional plan is part of a GaDOE collection of Unit Frameworks, Performance Tasks, examples of Student Work, and Teacher Commentary. Many more GaDOE approved instructional plans are
Chemical Knowledge for the Semantic Web
 Chemical Knowledge for the Semantic Web Mykola Konyk 1, Alexander De Leon 1, and Michel Dumontier 1,2,3 1 School of Computer Science 2 Department of Biology 3 Institute of Biochemistry, Carleton University,
Chemical Knowledge for the Semantic Web Mykola Konyk 1, Alexander De Leon 1, and Michel Dumontier 1,2,3 1 School of Computer Science 2 Department of Biology 3 Institute of Biochemistry, Carleton University,
Name Class Date. biosphere biology metabolism biodiversity organism DNA. MAIN IDEA: Earth is home to an incredible diversity of life.
 Section 1: The Study of Life KEY CONCEPT Biologists study life in all its forms. VOCABULARY biosphere biology metabolism biodiversity organism DNA species cell MAIN IDEA: Earth is home to an incredible
Section 1: The Study of Life KEY CONCEPT Biologists study life in all its forms. VOCABULARY biosphere biology metabolism biodiversity organism DNA species cell MAIN IDEA: Earth is home to an incredible
Completeness Proof Strategies for Euler Diagram Logics
 Completeness Proof Strategies for Euler Diagram Logics Jim Burton, Gem Stapleton, and John Howse Visual Modelling Group, University of Brighton, UK {j.burton,g.e.stapleton,john.howse}@brighton.ac.uk Abstract.
Completeness Proof Strategies for Euler Diagram Logics Jim Burton, Gem Stapleton, and John Howse Visual Modelling Group, University of Brighton, UK {j.burton,g.e.stapleton,john.howse}@brighton.ac.uk Abstract.
Cell Biology 1.1- Introduction to Cells
 Essential idea: Cell Biology 1.1- Introduction to Cells The evolution of multicellular organisms allowed cell specialization and cell replacement. Nature of science: Looking for trends and discrepancies
Essential idea: Cell Biology 1.1- Introduction to Cells The evolution of multicellular organisms allowed cell specialization and cell replacement. Nature of science: Looking for trends and discrepancies
Introduction to Logic and Axiomatic Set Theory
 Introduction to Logic and Axiomatic Set Theory 1 Introduction In mathematics, we seek absolute rigor in our arguments, and a solid foundation for all of the structures we consider. Here, we will see some
Introduction to Logic and Axiomatic Set Theory 1 Introduction In mathematics, we seek absolute rigor in our arguments, and a solid foundation for all of the structures we consider. Here, we will see some
Logical properties of foundational mereogeometrical relations in bio-ontologies
 Applied Ontology 0 (2009) 1 0 1 IOS Press Logical properties of foundational mereogeometrical relations in bio-ontologies Thomas Bittner Departments of Philosophy and Geography, State University of New
Applied Ontology 0 (2009) 1 0 1 IOS Press Logical properties of foundational mereogeometrical relations in bio-ontologies Thomas Bittner Departments of Philosophy and Geography, State University of New
 1 of 13 8/11/2014 10:32 AM Units: Teacher: APBiology, CORE Course: APBiology Year: 2012-13 Chemistry of Life Chapters 1-4 Big Idea 1, 2 & 4 Change in the genetic population over time is feedback mechanisms
1 of 13 8/11/2014 10:32 AM Units: Teacher: APBiology, CORE Course: APBiology Year: 2012-13 Chemistry of Life Chapters 1-4 Big Idea 1, 2 & 4 Change in the genetic population over time is feedback mechanisms
cis32-ai lecture # 18 mon-3-apr-2006
 cis32-ai lecture # 18 mon-3-apr-2006 today s topics: propositional logic cis32-spring2006-sklar-lec18 1 Introduction Weak (search-based) problem-solving does not scale to real problems. To succeed, problem
cis32-ai lecture # 18 mon-3-apr-2006 today s topics: propositional logic cis32-spring2006-sklar-lec18 1 Introduction Weak (search-based) problem-solving does not scale to real problems. To succeed, problem
Biology 211 (1) Exam 4! Chapter 12!
 Biology 211 (1) Exam 4 Chapter 12 1. Why does replication occurs in an uncondensed state? 1. 2. A is a single strand of DNA. When DNA is added to associated protein molecules, it is referred to as. 3.
Biology 211 (1) Exam 4 Chapter 12 1. Why does replication occurs in an uncondensed state? 1. 2. A is a single strand of DNA. When DNA is added to associated protein molecules, it is referred to as. 3.
A classification of spatio-temporal entities based on their location in space-time
 A classification of spatio-temporal entities based on their location in space-time Thomas Bittner 1,2,3,4 and Maureen Donnelly 1,3 1 Department of Philosophy, 2 Department of Geography 3 New York State
A classification of spatio-temporal entities based on their location in space-time Thomas Bittner 1,2,3,4 and Maureen Donnelly 1,3 1 Department of Philosophy, 2 Department of Geography 3 New York State
Encoding formulas with partially constrained weights in a possibilistic-like many-sorted propositional logic
 Encoding formulas with partially constrained weights in a possibilistic-like many-sorted propositional logic Salem Benferhat CRIL-CNRS, Université d Artois rue Jean Souvraz 62307 Lens Cedex France benferhat@criluniv-artoisfr
Encoding formulas with partially constrained weights in a possibilistic-like many-sorted propositional logic Salem Benferhat CRIL-CNRS, Université d Artois rue Jean Souvraz 62307 Lens Cedex France benferhat@criluniv-artoisfr
Logic in AI Chapter 7. Mausam (Based on slides of Dan Weld, Stuart Russell, Subbarao Kambhampati, Dieter Fox, Henry Kautz )
 Logic in AI Chapter 7 Mausam (Based on slides of Dan Weld, Stuart Russell, Subbarao Kambhampati, Dieter Fox, Henry Kautz ) 2 Knowledge Representation represent knowledge about the world in a manner that
Logic in AI Chapter 7 Mausam (Based on slides of Dan Weld, Stuart Russell, Subbarao Kambhampati, Dieter Fox, Henry Kautz ) 2 Knowledge Representation represent knowledge about the world in a manner that
