Supplementary Materials for
|
|
|
- Everett McCarthy
- 6 years ago
- Views:
Transcription
1 advances.sciencemag.org/cgi/content/full/1/9/e /dc1 Supplementary Materials for Contractility parameters of human -cardiac myosin with the hypertrophic cardiomyopathy mutation R403Q show loss of motor function Suman Nag, Ruth F. Sommese, Zoltan Ujfalusi, Ariana Combs, Stephen Langer, Shirley Sutton, Leslie A. Leinwand, Michael A. Geeves, Kathleen M. Ruppel, James A. Spudich The PDF file includes: Published 9 October 2015, Sci. Adv. 1, e (2015) DOI: /sciadv Fig. S1. Schematic of the different constructs of the human -cardiac ss1 used in our experiments and relative purity of the wild type and R403Q protein preparations. Fig. S2. Representative experimental traces of wild-type and R403Q ss1 unloaded motility. Fig. S3. Single-molecule stroke size measurements of wild type and R403Q human -cardiac ss1. Table S1. Rate constants and equilibrium constants for the various kinetic steps of wild type and R403Q human -cardiac ss1. Reaction schemes Scheme 1. Kinetic schemes for interaction of myosin only with nucleotide. Scheme 2. Kinetic schemes for interaction of myosin with actin and nucleotide. Scheme 3. A combined equilibrium scheme for myosin, actin, and ADP binding to each other. Reference (70)
2 WT-eGFP R403Q-eGFP WT-affinity clamp R403Q-affinity clamp Supplementary materials Supplementary Figures A B GSG egfp ss1 100 kd Human b-cardiac ss1 50 kd ss1 GSG RGSIDTWV 25 kd Human ELC Figure S1. Expression of recombinant human β-cardiac ss1. (A) A schematic of two different C- terminally-tagged constructs of human β-cardiac short Subfragment 1 (ss1, residues) used in our experiments. For all ATPase and single molecule trap experiments a C-terminal egfp tag on the ss1 was used (top). For all other experiments, except stopped-flow experiments, a C-terminal eight residue peptide (RGSIDTWV) was used (bottom), which specifically binds to the SNAP-PDZ18 system (70). We call this an 'affinity clamp' construct. For both of these constructs, a flexible GSG linker separates the N-terminal portion of the ss1 from the C-terminal tag. Both constructs have an N-terminal FLAGtagged (DYKDDDDK) human ventricular essential light chain (ELC) containing a TEV-precision protease site for affinity purification of the ss1. For most of the stopped flow experiments, we used ss1 with no C-terminal heavy chain tag, but with a 6X-His tag on the ELC; we duplicated the ADP-release kinetic experiments with the C-terminal egfp tagged version and found no differences compared to the untagged version (Table S1). (B) An overloaded SDS/PAGE of purified recombinant human β-cardiac ss1. Lane 1 contains the Invitrogen BenchMark Ladder. Three different molecular weights are marked for convenience. Lanes 2, 3, 4 and 5 contain wild-type (WT ss1-egfp), R403Q ss1-egfp, wild type (WT ss1-affinity clamp) and R403Q ss1-affinity clamp, respectively. The C-terminal egfp constructs migrate around 110 kda and the affinity clamp construct migrates around 90 kda. Both ss1 fragments co-purify with the human ventricular ELC, which migrates around 24 kda.
3 Number of events Number of events A B Filtered Unfiltered Filtered Unfiltered Actin filament length (mm) % Tolerance Actin filament length (mm) % Tolerance Figure S2. A representative unloaded motility output of human β-cardiac ss1. Actin filaments were tracked for at least 30 consecutive frames, actin velocities and actin filament lengths were averaged over 5 consecutive frames, a 20%-tolerance filter was applied to eliminate intermittently moving filaments with velocities fluctuating higher than 20 percent of the mean velocity, and stuck filaments were excluded from the scatter plot. Scatter plot of actin filament length and velocities for (A) wild-type and (B) R403Q β-cardiac ss1. Top left panel of (A) and (B): Maximum velocity of each filament is in triangles (black for wild-type and red for R403Q) and lower velocities are in gray circles. The solid line is the fit to a single exponential decay function to the maximum velocities to obtain the PLATEAU velocity. The dashed line marks the mean of the top 5% velocities (TOP5%). Top center panel of (A) and (B): A 20%-tolerance-filtered filament velocity distribution excluding stuck filaments. The solid line denotes the mean of the velocity distribution (MVEL20). Top right panel of (A) and (B): The unfiltered filament velocity distribution including stuck filaments. The solid line marks the mean of the distribution (MVEL). Filaments with a mean velocity less than 1 pixel length per second (80 nm/s) were classified as stuck. %STUCK represents the time-weighted fraction of stuck filaments. Bottom left panel of (A) and (B): Actin filament length distribution. The solid line marks the mean of the distribution (<FIL-LENGTH>). Bottom right panel of (A) and (B): MVEL (solid line) and TOP5% velocity (dashed line) from the data subjected to different levels of tolerance filtering. For more details see Aksel et al. (37).
4 Fraction of events Fraction of events Step size Step (nm) size (nm) Figure S3. Single-molecule stroke size measurements. Combined histogram of stroke size distribution from several single-molecules of wild-type (grey bars) and R403Q (red bars) human β-cardiac ss1 at 0.5 µm ATP. A Gaussian function using the least square method was used to fit the histogram (solid black line for wild-type and red for R403Q) to give a mean stroke size value.
5 Supplementary Table R403Q ss1 wild-type ss1 ATP-binding to 20 o C, Scheme 1 ATP binding rate constant K 1k +2 (µm -1 s -1 ) 5.4 ± ± 0.4 Rate constant of conformational change upon ATP binding k +2 (s -1 ) ± ± 1.8 Equilibrium dissociation constant 1/K 1 (µm) 20.8 ± ± 1.4 ADP-binding to 20 o C, Scheme 1,3 Equilibrium dissociation constant K D =K 6K 7 (µm) 0.49 ± ± 0.07 Rate constant for ADP release k +6 (s -1 ) 0.6 ± ± 0.03 ATP-binding to actin 20 o C, Scheme 2 ATP induced actin ss1dissociation rate constant K' 1k' +2 (µm -1 s -1 ) 4.9 ± ± 0.2 Equilibrium dissociation constant 1/K' 1 (µm) n.d n.d Rate constant of strong-to-weak binding transition of actin ss1 k' +2 (s -1 ) n.d n.d Nucleotide pocket isomerization equilibrium constant K α1 n.d n.d Nucleotide pocket isomerization forward rate constant k +α1 (s -1 ) n.d n.d ATP-binding to actin 10 o C, Scheme 2 ATP induced actin ss1dissociation rate constant K' 1k' +2 (µm -1 s -1 ) 2.5 ± ± 0.07 Equilibrium dissociation constant 1/K' 1 (µm) 276 ± 12 * ± 15 Rate constant of strong-to-weak binding transition of actin ss1 k' +2 (s -1 ) 677 ± 12 *** 991 ± 17 Nucleotide pocket isomerization equilibrium constant K α1 3.4 ± ± 0.1 Nucleotide pocket isomerization forward rate constant k +α1 (s -1 ) 53.3 ± 1.8 *** 78.9 ± 3 ADP-binding for actin 20 o C, Scheme 2,3 Equilibrium dissociation constant K AD =K' 6K' 7(µM) 7.6 ± ± 0.3 ADP release rate constant k' +6 (s -1 ) 60.0 ± ± 1.7 ADP release rate 23 o C k' +6 (s -1 ) 80.0 ± ± 10 Rate constant of ADP binding K' 7k' -6 (µm -1 s -1 ) Nucleotide pocket isomerization equilibrium constant in the presence of ADP K αd 16.4 ± 5 No second phase Nucleotide pocket isomerization forward rate constant in the presence of ADP k +αd (s -1 ) 4.2 ± 0.5 No second phase Coupling constant K' 6K' 7 / K 6K ss1-affinity for 20 o C, Scheme 3 Dissociation constant (nm) K A (nm) 19.7 ± 6.2 * 10 ± 1.8 ss1-affinity for actin at 100 mm 20 o C, Scheme 3 Dissociation constant (nm) K A (nm) 63.2 ± 16.7 * 23.4 ± 2.5 ss1-affinity for actin in the presence of 20 o C, Scheme 3 Dissociation constant (nm) K DA (nm) ± ± 24.1 Table S1: Rate constants and equilibrium constants for the various kinetic steps of wild-type and R403Q human β-cardiac ss1. All kinetic measurements for wild-type human β-cardiac ss1 (n = 3-4 exp, 3 preps) and R403Q ss1 (n = 3-4 exp, 2 preps) were done according to methodologies published in Deacon et al. (34) and Bloemink et al. (36). The buffer used contained 20 mm MOPS, 25 mm KCl, 5 mm MgCl2 and 1 mm DTT at 20 C, unless otherwise stated in the table. Errors reported are SEM. The values for R403Q that are shaded in orange are the ones which had statistical differences from wild-type
6 ss1. * P<0.05 and *** P< were determined by Student's t tests. For some of the measurements under 'ATP-binding to actin 20 C ' the rates were too fast and therefore unreliable, and hence are not determined (n.d). These measurements were repeated at 10 C (see 'ATP-binding to actin 10 C '). ki denotes a forward rate constant and k-i denotes a backward rate constants of steps (i) shown in scheme 1 and scheme 2 below. Ki (= ki/k-i) denotes equilibrium rate constants for a particular step. All rate constants and equilibrium constants without a prime sign denote kinetic steps involving only the ss1 and nucleotide (scheme 1), whereas those with the prime sign (') are for steps involving the ss1, actin and nucleotide (scheme 2). KA is the equilibrium dissociation constant of actin interaction with ss1 in the rigor state (scheme 3), KAD is the equilibrium dissociation constant of ADP affinity with the actin.ss1 complex (scheme 3), KDA is the equilibrium dissociation constant of actin affinity with the ADP.sS1 complex (scheme 3) and KD is the equilibrium dissociation constant of ss1 affinity with ADP (scheme 3).
7 Reaction Schemes Kinetic reaction scheme 1 shows the reaction mechanism of ss1 (M) with either ATP (T) or ADP (D) (33, 35). Phosphate is denoted as P. * and ** indicates different level of tryptophan fluorescence and represents different conformational states of ss1. Kinetic reaction scheme 2 shows the interaction of ss1 (M) with actin (A), ATP (T) and ADP (D). Actin complexed with ss1 exists in two conformations (A.M and A.M') in equilibrium. A M' (closed configuration) is unable to bind to nucleotide and must isomerize to A M (open configuration) before ATP can bind. A similar pair of conformations exist in the presence of bound ADP, where A M' D must isomerize to A M D before ADP can dissociate. * indicates different level of tryptophan fluorescence and represents different conformational states of ss1. Scheme 3 is a simplified combined scheme showing the affinity of ss1 with actin in the rigor state (the equilibrium dissociation constant denoted by KA), the affinity of actin ss1 complex to ADP (the equilibrium dissociation constant denoted by KAD=K'6K'7), the affinity of ss1 ADP complex to actin (the equilibrium dissociation constant denoted by KDA) and the affinity of ss1 to ADP (the equilibrium dissociation constant denoted by KD=K6K7). Scheme 1: Kinetic schemes for interaction of myosin only with nucleotide k 1 k 2 * k 3 ** k 4 * k 5 * k 6 k 7 M T M T M T M D P M D P P M D M D M D k 1 k 2 k 3 k 4 k 5 k 6 k 7 Scheme 2: Kinetic schemes for interaction of myosin with actin and nucleotide k D k 6 * k 7 T k 1 k 2 A M ' D A M D A M D D A M A M T A M T k D k 6 k 7 k 1 k 2 closed open open open open closed k 1 k 1 Scheme 3: A combined equilibrium scheme for myosin, actin and ADP binding to each other A M closed K AD K A K DA KD
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09450 Supplementary Table 1 Summary of kinetic parameters. Kinetic parameters were V = V / 1 K / ATP and obtained using the relationships max ( + m [ ]) V d s /( 1/ k [ ATP] + 1 k ) =,
doi:10.1038/nature09450 Supplementary Table 1 Summary of kinetic parameters. Kinetic parameters were V = V / 1 K / ATP and obtained using the relationships max ( + m [ ]) V d s /( 1/ k [ ATP] + 1 k ) =,
Supporting Information Converter domain mutations in myosin alter structural kinetics and motor function. Hershey, PA, MN 55455, USA
 Supporting Information Converter domain mutations in myosin alter structural kinetics and motor function Laura K. Gunther 1, John A. Rohde 2, Wanjian Tang 1, Shane D. Walton 1, William C. Unrath 1, Darshan
Supporting Information Converter domain mutations in myosin alter structural kinetics and motor function Laura K. Gunther 1, John A. Rohde 2, Wanjian Tang 1, Shane D. Walton 1, William C. Unrath 1, Darshan
Supplementary Information
 Supplementary Information Switching of myosin-v motion between the lever-arm swing and Brownian search-and-catch Keisuke Fujita 1*, Mitsuhiro Iwaki 2,3*, Atsuko H. Iwane 1, Lorenzo Marcucci 1 & Toshio
Supplementary Information Switching of myosin-v motion between the lever-arm swing and Brownian search-and-catch Keisuke Fujita 1*, Mitsuhiro Iwaki 2,3*, Atsuko H. Iwane 1, Lorenzo Marcucci 1 & Toshio
Analysis of nucleotide binding to p97 reveals the properties of a tandem AAA hexameric ATPase
 SUPPLEMENTARY INFORMATION Analysis of nucleotide binding to p97 reveals the properties of a tandem AAA hexameric ATPase Louise C Briggs, Geoff S Baldwin, Non Miyata, Hisao Kondo, Xiaodong Zhang, Paul S
SUPPLEMENTARY INFORMATION Analysis of nucleotide binding to p97 reveals the properties of a tandem AAA hexameric ATPase Louise C Briggs, Geoff S Baldwin, Non Miyata, Hisao Kondo, Xiaodong Zhang, Paul S
Kinetic Analysis of the Slow Skeletal Myosin MHC-1 Isoform from Bovine Masseter Muscle
 doi:10.1016/j.jmb.2007.08.050 J. Mol. Biol. (2007) 373, 1184 1197 Kinetic Analysis of the Slow Skeletal Myosin MHC-1 Isoform from Bovine Masseter Muscle M. J. Bloemink 1, N. Adamek 1, C. Reggiani 2 and
doi:10.1016/j.jmb.2007.08.050 J. Mol. Biol. (2007) 373, 1184 1197 Kinetic Analysis of the Slow Skeletal Myosin MHC-1 Isoform from Bovine Masseter Muscle M. J. Bloemink 1, N. Adamek 1, C. Reggiani 2 and
Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition
 Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Use of Stable Analogs of Myosin ATPase Intermediates for Kinetic Studies of the Weak Binding of Myosin Heads to F-Actin
 Biochemistry (Moscow), Vol. 64, No. 8, 1999, pp. 875-882. Translated from Biokhimiya, Vol. 64, No. 8, 1999, pp. 1043-1051. Original Russian Text Copyright 1999 by Rostkova, Moiseeva, Teplova, Nikolaeva,
Biochemistry (Moscow), Vol. 64, No. 8, 1999, pp. 875-882. Translated from Biokhimiya, Vol. 64, No. 8, 1999, pp. 1043-1051. Original Russian Text Copyright 1999 by Rostkova, Moiseeva, Teplova, Nikolaeva,
Anatoly B. Kolomeisky. Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS?
 Anatoly B. Kolomeisky Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS? Motor Proteins Enzymes that convert the chemical energy into mechanical
Anatoly B. Kolomeisky Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS? Motor Proteins Enzymes that convert the chemical energy into mechanical
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/10/eaat8797/dc1 Supplementary Materials for Single-molecule observation of nucleotide induced conformational changes in basal SecA-ATP hydrolysis Nagaraju Chada,
advances.sciencemag.org/cgi/content/full/4/10/eaat8797/dc1 Supplementary Materials for Single-molecule observation of nucleotide induced conformational changes in basal SecA-ATP hydrolysis Nagaraju Chada,
Introduction: actin and myosin
 Introduction: actin and myosin Actin Myosin Myosin V and actin 375 residues Found in all eukaryotes Polymeric Forms track for myosin Many other cellular functions 36 nm pseudo-helical repeat Catalytic
Introduction: actin and myosin Actin Myosin Myosin V and actin 375 residues Found in all eukaryotes Polymeric Forms track for myosin Many other cellular functions 36 nm pseudo-helical repeat Catalytic
Supporting information for
 Supporting information for Rewiring multi-domain protein switches: transforming a fluorescent Zn 2+ -sensor into a light-responsive Zn 2+ binding protein Stijn J.A. Aper and Maarten Merkx Laboratory of
Supporting information for Rewiring multi-domain protein switches: transforming a fluorescent Zn 2+ -sensor into a light-responsive Zn 2+ binding protein Stijn J.A. Aper and Maarten Merkx Laboratory of
The Fic protein Doc uses an inverted substrate to phosphorylate and. inactivate EF-Tu
 The Fic protein Doc uses an inverted substrate to phosphorylate and inactivate EF-Tu Daniel Castro-Roa 1, Abel Garcia-Pino 2,3 *, Steven De Gieter 2,3, Nico A.J. van Nuland 2,3, Remy Loris 2,3, Nikolay
The Fic protein Doc uses an inverted substrate to phosphorylate and inactivate EF-Tu Daniel Castro-Roa 1, Abel Garcia-Pino 2,3 *, Steven De Gieter 2,3, Nico A.J. van Nuland 2,3, Remy Loris 2,3, Nikolay
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1.
 Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/6/e1700147/dc1 Supplementary Materials for Ribosome rearrangements at the onset of translational bypassing Xabier Agirrezabala, Ekaterina Samatova, Mariia Klimova,
advances.sciencemag.org/cgi/content/full/3/6/e1700147/dc1 Supplementary Materials for Ribosome rearrangements at the onset of translational bypassing Xabier Agirrezabala, Ekaterina Samatova, Mariia Klimova,
Muscle regulation and Actin Topics: Tropomyosin and Troponin, Actin Assembly, Actin-dependent Movement
 1 Muscle regulation and Actin Topics: Tropomyosin and Troponin, Actin Assembly, Actin-dependent Movement In the last lecture, we saw that a repeating alternation between chemical (ATP hydrolysis) and vectorial
1 Muscle regulation and Actin Topics: Tropomyosin and Troponin, Actin Assembly, Actin-dependent Movement In the last lecture, we saw that a repeating alternation between chemical (ATP hydrolysis) and vectorial
Supplemental Data for: Direct Observation of Translocation in Individual DNA Polymerase Complexes
 Supplemental Data for: Direct Observation of Translocation in Individual DNA Polymerase Complexes Joseph M. Dahl 1, Ai H. Mai 1, Gerald M. Cherf 1, Nahid N. Jetha 4, Daniel R. Garalde 3, Andre Marziali
Supplemental Data for: Direct Observation of Translocation in Individual DNA Polymerase Complexes Joseph M. Dahl 1, Ai H. Mai 1, Gerald M. Cherf 1, Nahid N. Jetha 4, Daniel R. Garalde 3, Andre Marziali
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10955 Supplementary Figures Supplementary Figure 1. Electron-density maps and crystallographic dimer structures of the motor domain. (a f) Stereo views of the final electron-density maps
doi:10.1038/nature10955 Supplementary Figures Supplementary Figure 1. Electron-density maps and crystallographic dimer structures of the motor domain. (a f) Stereo views of the final electron-density maps
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a
 Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
SUPPLEMENTARY INFORMATION
 Supplementary Table 1: Amplitudes of three current levels. Level 0 (pa) Level 1 (pa) Level 2 (pa) TrkA- TrkH WT 200 K 0.01 ± 0.01 9.5 ± 0.01 18.7 ± 0.03 200 Na * 0.001 ± 0.01 3.9 ± 0.01 12.5 ± 0.03 200
Supplementary Table 1: Amplitudes of three current levels. Level 0 (pa) Level 1 (pa) Level 2 (pa) TrkA- TrkH WT 200 K 0.01 ± 0.01 9.5 ± 0.01 18.7 ± 0.03 200 Na * 0.001 ± 0.01 3.9 ± 0.01 12.5 ± 0.03 200
Fig. 1. Stereo images showing (A) the best fit of the atomic model for F actin and the F actin map obtained by cryo-em and image analysis, and (B) goo
 Fig. 1. Stereo images showing (A) the best fit of the atomic model for F actin and the F actin map obtained by cryo-em and image analysis, and (B) good correspondence between the location of Cys374 and
Fig. 1. Stereo images showing (A) the best fit of the atomic model for F actin and the F actin map obtained by cryo-em and image analysis, and (B) good correspondence between the location of Cys374 and
SINGLE-MOLECULE PHYSIOLOGY
 SINGLE-MOLECULE PHYSIOLOGY Kazuhiko Kinosita, Jr. Center for Integrative Bioscience, Okazaki National Research Institutes Higashiyama 5-1, Myodaiji, Okazaki 444-8585, Japan Single-Molecule Physiology under
SINGLE-MOLECULE PHYSIOLOGY Kazuhiko Kinosita, Jr. Center for Integrative Bioscience, Okazaki National Research Institutes Higashiyama 5-1, Myodaiji, Okazaki 444-8585, Japan Single-Molecule Physiology under
MARCH 28, 2008 VOLUME 283 NUMBER 13 JOURNAL OF BIOLOGICAL CHEMISTRY 8527
 THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 283, NO. 13, pp. 8527 8537, March 28, 2008 Printed in the U.S.A. Human Myosin Vc Is a Low Duty Ratio, Nonprocessive Molecular Motor * Received for publication,
THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 283, NO. 13, pp. 8527 8537, March 28, 2008 Printed in the U.S.A. Human Myosin Vc Is a Low Duty Ratio, Nonprocessive Molecular Motor * Received for publication,
National de la Recherche Scientifique and Université Paris Descartes, Paris, France.
 FAST-RESPONSE CALMODULIN-BASED FLUORESCENT INDICATORS REVEAL RAPID INTRACELLULAR CALCIUM DYNAMICS Nordine Helassa a, Xiao-hua Zhang b, Ianina Conte a,c, John Scaringi b, Elric Esposito d, Jonathan Bradley
FAST-RESPONSE CALMODULIN-BASED FLUORESCENT INDICATORS REVEAL RAPID INTRACELLULAR CALCIUM DYNAMICS Nordine Helassa a, Xiao-hua Zhang b, Ianina Conte a,c, John Scaringi b, Elric Esposito d, Jonathan Bradley
Supporting Information for: Kinetic Mechanisms Governing Stable Ribonucleotide Incorporation in Individual DNA Polymerase Complexes
 Supporting Information for: Kinetic Mechanisms Governing Stable Ribonucleotide Incorporation in Individual DNA Polymerase Complexes Joseph M. Dahl, Hongyun Wang, José M. Lázaro, Margarita Salas, and Kate
Supporting Information for: Kinetic Mechanisms Governing Stable Ribonucleotide Incorporation in Individual DNA Polymerase Complexes Joseph M. Dahl, Hongyun Wang, José M. Lázaro, Margarita Salas, and Kate
Dictyostelium discoideum Myosin II: Characterization of Functional Myosin Motor Fragments
 Biochemistry 1997, 36, 317-323 317 Dictyostelium discoideum Myosin II: Characterization of Functional Myosin Motor Fragments S. E. Kurzawa, D. J. Manstein, and M. A. Geeves*, Max-Planck-Institut für Molekulare
Biochemistry 1997, 36, 317-323 317 Dictyostelium discoideum Myosin II: Characterization of Functional Myosin Motor Fragments S. E. Kurzawa, D. J. Manstein, and M. A. Geeves*, Max-Planck-Institut für Molekulare
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Identification of the ScDcp2 minimal region interacting with both ScDcp1 and the ScEdc3 LSm domain. Pull-down experiment of untagged ScEdc3 LSm with various ScDcp1-Dcp2-His 6 fragments.
Supplementary Figure 1 Identification of the ScDcp2 minimal region interacting with both ScDcp1 and the ScEdc3 LSm domain. Pull-down experiment of untagged ScEdc3 LSm with various ScDcp1-Dcp2-His 6 fragments.
Functional Characterisation of Dictyostelium Myosin II with Conserved Tryptophanyl Residue 501 Mutated to Tyrosine
 Biol. Chem., Vol. 380, pp. 1017 1023, July/August 1999 Copyright by Walter de Gruyter Berlin New York Short Communication Functional Characterisation of Dictyostelium Myosin II with Conserved Tryptophanyl
Biol. Chem., Vol. 380, pp. 1017 1023, July/August 1999 Copyright by Walter de Gruyter Berlin New York Short Communication Functional Characterisation of Dictyostelium Myosin II with Conserved Tryptophanyl
Chapter 6. The interaction of Src SH2 with the focal adhesion kinase catalytic domain studied by NMR
 The interaction of Src SH2 with the focal adhesion kinase catalytic domain studied by NMR 103 Abstract The interaction of the Src SH2 domain with the catalytic domain of FAK, including the Y397 SH2 domain
The interaction of Src SH2 with the focal adhesion kinase catalytic domain studied by NMR 103 Abstract The interaction of the Src SH2 domain with the catalytic domain of FAK, including the Y397 SH2 domain
Description: Supplementary Figures, Supplementary Methods, and Supplementary References
 File Name: Supplementary Information Description: Supplementary Figures, Supplementary Methods, and Supplementary References File Name: Supplementary Movie 1 Description: Footage of time trace of seeds
File Name: Supplementary Information Description: Supplementary Figures, Supplementary Methods, and Supplementary References File Name: Supplementary Movie 1 Description: Footage of time trace of seeds
Biophysik der Moleküle!
 Biophysik der Moleküle!!"#$%&'()*+,-$./0()'$12$34!4! Molecular Motors:! - linear motors" 6. Dec. 2010! Muscle Motors and Cargo Transporting Motors! There are striking structural similarities but functional
Biophysik der Moleküle!!"#$%&'()*+,-$./0()'$12$34!4! Molecular Motors:! - linear motors" 6. Dec. 2010! Muscle Motors and Cargo Transporting Motors! There are striking structural similarities but functional
ATP binding controls distinct structural transitions. of Escherichia coli DNA gyrase in complex with DNA
 Supplementary Information ATP binding controls distinct structural transitions of Escherichia coli DNA gyrase in complex with DNA Aakash Basu, Allyn J. Schoeffler, James M. Berger, and Zev Bryant Table
Supplementary Information ATP binding controls distinct structural transitions of Escherichia coli DNA gyrase in complex with DNA Aakash Basu, Allyn J. Schoeffler, James M. Berger, and Zev Bryant Table
Lecture 13, 05 October 2004 Chapter 10, Muscle. Vertebrate Physiology ECOL 437 University of Arizona Fall instr: Kevin Bonine t.a.
 Lecture 13, 05 October 2004 Chapter 10, Muscle Vertebrate Physiology ECOL 437 University of Arizona Fall 2004 instr: Kevin Bonine t.a.: Nate Swenson Vertebrate Physiology 437 18 1. Muscle A. Sarcomere
Lecture 13, 05 October 2004 Chapter 10, Muscle Vertebrate Physiology ECOL 437 University of Arizona Fall 2004 instr: Kevin Bonine t.a.: Nate Swenson Vertebrate Physiology 437 18 1. Muscle A. Sarcomere
Chapter 16. Cellular Movement: Motility and Contractility. Lectures by Kathleen Fitzpatrick Simon Fraser University Pearson Education, Inc.
 Chapter 16 Cellular Movement: Motility and Contractility Lectures by Kathleen Fitzpatrick Simon Fraser University Two eukaryotic motility systems 1. Interactions between motor proteins and microtubules
Chapter 16 Cellular Movement: Motility and Contractility Lectures by Kathleen Fitzpatrick Simon Fraser University Two eukaryotic motility systems 1. Interactions between motor proteins and microtubules
type GroEL-GroES complex. Crystals were grown in buffer D (100 mm HEPES, ph 7.5,
 Supplementary Material Supplementary Materials and Methods Structure Determination of SR1-GroES-ADP AlF x SR1-GroES-ADP AlF x was purified as described in Materials and Methods for the wild type GroEL-GroES
Supplementary Material Supplementary Materials and Methods Structure Determination of SR1-GroES-ADP AlF x SR1-GroES-ADP AlF x was purified as described in Materials and Methods for the wild type GroEL-GroES
Supporting Text - Jégou et al.
 Supporting Text - Jégou et al. I. PAUSES DURING DEPOLYMERIZATION Pauses were observed during the ymerization of individual filaments grown from ADP-, CrATP- or MgATPactin, in the presence or in the absence
Supporting Text - Jégou et al. I. PAUSES DURING DEPOLYMERIZATION Pauses were observed during the ymerization of individual filaments grown from ADP-, CrATP- or MgATPactin, in the presence or in the absence
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing
 Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Structure and RNA-binding properties. of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex
 Structure and RNA-binding properties of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex Varun Bhaskar 1, Vladimir Roudko 2,3, Jerome Basquin 1, Kundan Sharma 4, Henning Urlaub 4, Bertrand Seraphin
Structure and RNA-binding properties of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex Varun Bhaskar 1, Vladimir Roudko 2,3, Jerome Basquin 1, Kundan Sharma 4, Henning Urlaub 4, Bertrand Seraphin
Acto-myosin: from muscles to single molecules. Justin Molloy MRC National Institute for Medical Research LONDON
 Acto-myosin: from muscles to single molecules. Justin Molloy MRC National Institute for Medical Research LONDON Energy in Biological systems: 1 Photon = 400 pn.nm 1 ATP = 100 pn.nm 1 Ion moving across
Acto-myosin: from muscles to single molecules. Justin Molloy MRC National Institute for Medical Research LONDON Energy in Biological systems: 1 Photon = 400 pn.nm 1 ATP = 100 pn.nm 1 Ion moving across
Supplementary Information. The protease GtgE from Salmonella exclusively targets. inactive Rab GTPases
 Supplementary Information The protease GtgE from Salmonella exclusively targets inactive Rab GTPases Table of Contents Supplementary Figures... 2 Supplementary Figure 1... 2 Supplementary Figure 2... 3
Supplementary Information The protease GtgE from Salmonella exclusively targets inactive Rab GTPases Table of Contents Supplementary Figures... 2 Supplementary Figure 1... 2 Supplementary Figure 2... 3
Supplementary Information to
 Supplementary Information to Wiesner et al.: A change in conformational dynamics underlies the activation of Eph receptor tyrosine kinases Supplementary Material and Methods Cloning and Mutagenesis Site-directed
Supplementary Information to Wiesner et al.: A change in conformational dynamics underlies the activation of Eph receptor tyrosine kinases Supplementary Material and Methods Cloning and Mutagenesis Site-directed
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/5/e1601684/dc1 Supplementary Materials for Directional mechanical stability of Bacteriophage ϕ29 motor s 3WJ-pRNA: Extraordinary robustness along portal axis
advances.sciencemag.org/cgi/content/full/3/5/e1601684/dc1 Supplementary Materials for Directional mechanical stability of Bacteriophage ϕ29 motor s 3WJ-pRNA: Extraordinary robustness along portal axis
Supplementary Information. Overlap between folding and functional energy landscapes for. adenylate kinase conformational change
 Supplementary Information Overlap between folding and functional energy landscapes for adenylate kinase conformational change by Ulrika Olsson & Magnus Wolf-Watz Contents: 1. Supplementary Note 2. Supplementary
Supplementary Information Overlap between folding and functional energy landscapes for adenylate kinase conformational change by Ulrika Olsson & Magnus Wolf-Watz Contents: 1. Supplementary Note 2. Supplementary
Table 1. Kinetic data obtained from SPR analysis of domain 11 mutants interacting with IGF-II. Kinetic parameters K D 1.
 Kinetics and Thermodynamics of the Insulin-like Growth Factor II (IGF-II) Interaction with IGF-II/Mannose 6-phosphate Receptor and the function of CD and AB Loop Solvent-exposed Residues. Research Team:
Kinetics and Thermodynamics of the Insulin-like Growth Factor II (IGF-II) Interaction with IGF-II/Mannose 6-phosphate Receptor and the function of CD and AB Loop Solvent-exposed Residues. Research Team:
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles
 Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/4/e1600663/dc1 Supplementary Materials for A dynamic hydrophobic core orchestrates allostery in protein kinases Jonggul Kim, Lalima G. Ahuja, Fa-An Chao, Youlin
advances.sciencemag.org/cgi/content/full/3/4/e1600663/dc1 Supplementary Materials for A dynamic hydrophobic core orchestrates allostery in protein kinases Jonggul Kim, Lalima G. Ahuja, Fa-An Chao, Youlin
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1. Definition and assessment of ciap1 constructs.
 Supplementary Figure 1 Definition and assessment of ciap1 constructs. (a) ciap1 constructs used in this study are shown as primary structure schematics with domains colored as in the main text. Mutations
Supplementary Figure 1 Definition and assessment of ciap1 constructs. (a) ciap1 constructs used in this study are shown as primary structure schematics with domains colored as in the main text. Mutations
SECOND PUBLIC EXAMINATION. Honour School of Physics Part C: 4 Year Course. Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS
 2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2011 Monday, 27 June, 9.30 am 12.30 pm Answer
2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2011 Monday, 27 June, 9.30 am 12.30 pm Answer
Supplementary Information for The depolymerizing kinesin MCAK uses lattice diffusion to rapidly target microtubule ends
 doi:10.1038/nature04736 nature Supplementary Information for The depolymerizing kinesin MCAK uses lattice diffusion to rapidly target microtubule ends Jonne H. Helenius 1, Gary J. Brouhard 1, Yannis Kalaidzidis
doi:10.1038/nature04736 nature Supplementary Information for The depolymerizing kinesin MCAK uses lattice diffusion to rapidly target microtubule ends Jonne H. Helenius 1, Gary J. Brouhard 1, Yannis Kalaidzidis
Thermal Denaturation and Aggregation of Myosin Subfragment 1 Isoforms with Different Essential Light Chains
 Int. J. Mol. Sci. 21, 11, 4194-4226; doi:1.339/ijms11114194 OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-67 www.mdpi.com/journal/ijms Article Thermal Denaturation and Aggregation of
Int. J. Mol. Sci. 21, 11, 4194-4226; doi:1.339/ijms11114194 OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-67 www.mdpi.com/journal/ijms Article Thermal Denaturation and Aggregation of
SUPPLEMENTARY FIGURE 1. Force dependence of the unbinding rate: (a) Force-dependence
 (b) BSA-coated beads double exponential low force exponential high force exponential 1 unbinding time tb [sec] (a) unbinding time tb [sec] SUPPLEMENTARY FIGURES BSA-coated beads without BSA.2.1 5 1 load
(b) BSA-coated beads double exponential low force exponential high force exponential 1 unbinding time tb [sec] (a) unbinding time tb [sec] SUPPLEMENTARY FIGURES BSA-coated beads without BSA.2.1 5 1 load
Supplemental Materials and Methods
 Supplemental Materials and Methods Time-resolved FRET (trfret) to probe for changes in the Box A/A stem upon complex assembly U3 MINI was folded and the decay of Fl fluorescence was measured at 20 ºC (see
Supplemental Materials and Methods Time-resolved FRET (trfret) to probe for changes in the Box A/A stem upon complex assembly U3 MINI was folded and the decay of Fl fluorescence was measured at 20 ºC (see
Supporting Information
 Supporting Information Mullins et al. 10.1073/pnas.0906781106 SI Text Detection of Calcium Binding by 45 Ca 2 Overlay. The 45 CaCl 2 (1 mci, 37 MBq) was obtained from NEN. The general method of 45 Ca 2
Supporting Information Mullins et al. 10.1073/pnas.0906781106 SI Text Detection of Calcium Binding by 45 Ca 2 Overlay. The 45 CaCl 2 (1 mci, 37 MBq) was obtained from NEN. The general method of 45 Ca 2
James B. Munro, Roger B. Altman, Nathan O Connor, and Scott C. Blanchard
 Molecular Cell, Volume 25 Supplemental Data Identification of Two Distinct Hybrid State Intermediates on the Ribosome James B. Munro, Roger B. Altman, Nathan O Connor, and Scott C. Blanchard Wild-type
Molecular Cell, Volume 25 Supplemental Data Identification of Two Distinct Hybrid State Intermediates on the Ribosome James B. Munro, Roger B. Altman, Nathan O Connor, and Scott C. Blanchard Wild-type
SUPPLEMENTARY INFORMATION
 doi:1.138/nature1737 Supplementary Table 1 variant Description FSEC - 2B12 a FSEC - 6A1 a K d (leucine) c Leucine uptake e K (wild-type like) K (Y18F) K (TS) K (TSY) K288A mutant, lipid facing side chain
doi:1.138/nature1737 Supplementary Table 1 variant Description FSEC - 2B12 a FSEC - 6A1 a K d (leucine) c Leucine uptake e K (wild-type like) K (Y18F) K (TS) K (TSY) K288A mutant, lipid facing side chain
Binding Theory Equations for Affinity and Kinetics Analysis
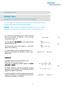 Technology Note #101 Binding Theory Equations for Affinity and Kinetics Analysis This technology note summarizes important equations underlying the theory of binding of solute analytes to surface-tethered
Technology Note #101 Binding Theory Equations for Affinity and Kinetics Analysis This technology note summarizes important equations underlying the theory of binding of solute analytes to surface-tethered
Supplementary Information. Structural basis for precursor protein-directed ribosomal peptide macrocyclization
 Supplementary Information Structural basis for precursor protein-directed ribosomal peptide macrocyclization Kunhua Li 1,3, Heather L. Condurso 1,3, Gengnan Li 1, Yousong Ding 2 and Steven D. Bruner 1*
Supplementary Information Structural basis for precursor protein-directed ribosomal peptide macrocyclization Kunhua Li 1,3, Heather L. Condurso 1,3, Gengnan Li 1, Yousong Ding 2 and Steven D. Bruner 1*
BME Engineering Molecular Cell Biology. Basics of the Diffusion Theory. The Cytoskeleton (I)
 BME 42-620 Engineering Molecular Cell Biology Lecture 07: Basics of the Diffusion Theory The Cytoskeleton (I) BME42-620 Lecture 07, September 20, 2011 1 Outline Diffusion: microscopic theory Diffusion:
BME 42-620 Engineering Molecular Cell Biology Lecture 07: Basics of the Diffusion Theory The Cytoskeleton (I) BME42-620 Lecture 07, September 20, 2011 1 Outline Diffusion: microscopic theory Diffusion:
BIOMECHANICS 3 Origins and consequences of forces in biological systems
 BIOMECHANICS 3 Origins and consequences of forces in biological systems MOLECULAR MECHANISMS OF BIOLOGICAL MOVEMENT AT THE LEVELOF ORGANISMS MOLECULAR BASIS OF MUSCLE CONTRACTION DR. BEÁTA BUGYI - BIOPHYSICS
BIOMECHANICS 3 Origins and consequences of forces in biological systems MOLECULAR MECHANISMS OF BIOLOGICAL MOVEMENT AT THE LEVELOF ORGANISMS MOLECULAR BASIS OF MUSCLE CONTRACTION DR. BEÁTA BUGYI - BIOPHYSICS
Supplemental Information. The Mitochondrial Fission Receptor MiD51. Requires ADP as a Cofactor
 Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/5/e1600519/dc1 Supplementary Materials for Encapsulation-free controlled release: Electrostatic adsorption eliminates the need for protein encapsulation in PLGA
advances.sciencemag.org/cgi/content/full/2/5/e1600519/dc1 Supplementary Materials for Encapsulation-free controlled release: Electrostatic adsorption eliminates the need for protein encapsulation in PLGA
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2215 Figure S1 Number of egfp-vps4a bursts versus cellular expression levels. The total number of egfp-vps4a bursts, counted at the end of each movie (frame 2000, after 1h 28 min) are plotted
DOI: 10.1038/ncb2215 Figure S1 Number of egfp-vps4a bursts versus cellular expression levels. The total number of egfp-vps4a bursts, counted at the end of each movie (frame 2000, after 1h 28 min) are plotted
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Crystallization. a, Crystallization constructs of the ET B receptor are shown, with all of the modifications to the human wild-type the ET B receptor indicated. Residues interacting
Supplementary Figure 1 Crystallization. a, Crystallization constructs of the ET B receptor are shown, with all of the modifications to the human wild-type the ET B receptor indicated. Residues interacting
In order to confirm the contribution of diffusion to the FRAP recovery curves of
 Fonseca et al., Supplementary Information FRAP data analysis ) Contribution of Diffusion to the recovery curves In order to confirm the contribution of diffusion to the FRAP recovery curves of PH::GFP
Fonseca et al., Supplementary Information FRAP data analysis ) Contribution of Diffusion to the recovery curves In order to confirm the contribution of diffusion to the FRAP recovery curves of PH::GFP
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11054 Supplementary Fig. 1 Sequence alignment of Na v Rh with NaChBac, Na v Ab, and eukaryotic Na v and Ca v homologs. Secondary structural elements of Na v Rh are indicated above the
doi:10.1038/nature11054 Supplementary Fig. 1 Sequence alignment of Na v Rh with NaChBac, Na v Ab, and eukaryotic Na v and Ca v homologs. Secondary structural elements of Na v Rh are indicated above the
A conserved P-loop anchor limits the structural dynamics that mediate. nucleotide dissociation in EF-Tu.
 Supplemental Material for A conserved P-loop anchor limits the structural dynamics that mediate nucleotide dissociation in EF-Tu. Evan Mercier 1,2, Dylan Girodat 1, and Hans-Joachim Wieden 1 * 1 Alberta
Supplemental Material for A conserved P-loop anchor limits the structural dynamics that mediate nucleotide dissociation in EF-Tu. Evan Mercier 1,2, Dylan Girodat 1, and Hans-Joachim Wieden 1 * 1 Alberta
Supplementary Information
 Supplementary Information Adenosyltransferase Tailors and Delivers Coenzyme B 12 Dominique Padovani 1,2, Tetyana Labunska 2, Bruce A. Palfey 1, David P. Ballou 1 and Ruma Banerjee 1,2 * 1 Biological Chemistry
Supplementary Information Adenosyltransferase Tailors and Delivers Coenzyme B 12 Dominique Padovani 1,2, Tetyana Labunska 2, Bruce A. Palfey 1, David P. Ballou 1 and Ruma Banerjee 1,2 * 1 Biological Chemistry
Interactions and Dynamics within the Troponin Complex
 Interactions and Dynamics within the Troponin Complex Tharin Blumenschein Steve Matthews Lab - Imperial College London (formerly Brian Sykes Lab, Canada) Striated muscle Thin filament proteins - regulation
Interactions and Dynamics within the Troponin Complex Tharin Blumenschein Steve Matthews Lab - Imperial College London (formerly Brian Sykes Lab, Canada) Striated muscle Thin filament proteins - regulation
1. The plasma membrane of eukaryotic cells is supported by a. actin filaments. b. microtubules. c. lamins. d. intermediate filaments.
 ANALYSIS AND MODELING OF CELL MECHANICS Homework #2 (due 1/30/13) This homework involves comprehension of key biomechanical concepts of the cytoskeleton, cell-matrix adhesions, and cellcell adhesions.
ANALYSIS AND MODELING OF CELL MECHANICS Homework #2 (due 1/30/13) This homework involves comprehension of key biomechanical concepts of the cytoskeleton, cell-matrix adhesions, and cellcell adhesions.
Mechanochemical coupling of two substeps in a single myosin V motor
 Mechanochemical coupling of two substeps in a single myosin V motor Sotaro Uemura 1, Hideo Higuchi 2,3, Adrian O Olivares 4, Enrique M De La Cruz 4 & Shin ichi Ishiwata 1,5 Myosin V is a double-headed
Mechanochemical coupling of two substeps in a single myosin V motor Sotaro Uemura 1, Hideo Higuchi 2,3, Adrian O Olivares 4, Enrique M De La Cruz 4 & Shin ichi Ishiwata 1,5 Myosin V is a double-headed
downstream (0.8 kb) homologous sequences to the genomic locus of DIC. A DIC mutant strain (ro- 6
 A B C D ts Figure S1 Generation of DIC- mcherry expressing N.crassa strain. A. N. crassa colony morphology. When a cot1 (top, left panel) strain is grown at permissive temperature (25 C), it exhibits straight
A B C D ts Figure S1 Generation of DIC- mcherry expressing N.crassa strain. A. N. crassa colony morphology. When a cot1 (top, left panel) strain is grown at permissive temperature (25 C), it exhibits straight
Processivity of dimeric kinesin-1 molecular motors
 Processivity of dimeric kinesin-1 molecular motors Si-Kao Guo 1,2, Xiao-Xuan Shi 1,2, Peng-Ye Wang 1,2 and Ping Xie 1,2 1 Key Laboratory of Soft Matter Physics, Beijing National Laboratory for Condensed
Processivity of dimeric kinesin-1 molecular motors Si-Kao Guo 1,2, Xiao-Xuan Shi 1,2, Peng-Ye Wang 1,2 and Ping Xie 1,2 1 Key Laboratory of Soft Matter Physics, Beijing National Laboratory for Condensed
Supporting Information for: Mechanism of Reversible Peptide-Bilayer. Attachment: Combined Simulation and Experimental Single-Molecule Study
 Supporting Information for: Mechanism of Reversible Peptide-Bilayer Attachment: Combined Simulation and Experimental Single-Molecule Study Nadine Schwierz a,, Stefanie Krysiak c, Thorsten Hugel c,d, Martin
Supporting Information for: Mechanism of Reversible Peptide-Bilayer Attachment: Combined Simulation and Experimental Single-Molecule Study Nadine Schwierz a,, Stefanie Krysiak c, Thorsten Hugel c,d, Martin
Nature Methods: doi: /nmeth Supplementary Figure 1. In vitro screening of recombinant R-CaMP2 variants.
 Supplementary Figure 1 In vitro screening of recombinant R-CaMP2 variants. Baseline fluorescence compared to R-CaMP1.07 at nominally zero calcium plotted versus dynamic range ( F/F) for 150 recombinant
Supplementary Figure 1 In vitro screening of recombinant R-CaMP2 variants. Baseline fluorescence compared to R-CaMP1.07 at nominally zero calcium plotted versus dynamic range ( F/F) for 150 recombinant
Cardiac troponin T mutations: correlation between the type of mutation and the nature of myofilament dysfunction in transgenic mice
 12644 Journal of Physiology (2001), 536.2, pp.583 592 583 Cardiac troponin T mutations: correlation between the type of mutation and the nature of myofilament dysfunction in transgenic mice David E. Montgomery,
12644 Journal of Physiology (2001), 536.2, pp.583 592 583 Cardiac troponin T mutations: correlation between the type of mutation and the nature of myofilament dysfunction in transgenic mice David E. Montgomery,
Supplementary Information
 Supplementary Information Structural analysis of leader peptide binding enables leaderfree cyanobactin processing Jesko Koehnke 1,2, Greg Mann 1,2, Andrew F Bent 1,2, Hannes Ludewig 1, Sally Shirran 1,
Supplementary Information Structural analysis of leader peptide binding enables leaderfree cyanobactin processing Jesko Koehnke 1,2, Greg Mann 1,2, Andrew F Bent 1,2, Hannes Ludewig 1, Sally Shirran 1,
Changes in microtubule overlap length regulate kinesin-14-driven microtubule sliding
 Supplementary information Changes in microtubule overlap length regulate kinesin-14-driven microtubule sliding Marcus Braun* 1,2,3, Zdenek Lansky* 1,2,3, Agata Szuba 1,2, Friedrich W. Schwarz 1,2, Aniruddha
Supplementary information Changes in microtubule overlap length regulate kinesin-14-driven microtubule sliding Marcus Braun* 1,2,3, Zdenek Lansky* 1,2,3, Agata Szuba 1,2, Friedrich W. Schwarz 1,2, Aniruddha
FW 1 CDR 1 FW 2 CDR 2
 Supplementary Figure 1 Supplementary Figure 1: Interface of the E9:Fas structure. The two interfaces formed by V H and V L of E9 with Fas are shown in stereo. The Fas receptor is represented as a surface
Supplementary Figure 1 Supplementary Figure 1: Interface of the E9:Fas structure. The two interfaces formed by V H and V L of E9 with Fas are shown in stereo. The Fas receptor is represented as a surface
Lecture 11: Enzymes: Kinetics [PDF] Reading: Berg, Tymoczko & Stryer, Chapter 8, pp
![Lecture 11: Enzymes: Kinetics [PDF] Reading: Berg, Tymoczko & Stryer, Chapter 8, pp Lecture 11: Enzymes: Kinetics [PDF] Reading: Berg, Tymoczko & Stryer, Chapter 8, pp](/thumbs/74/70848503.jpg) Lecture 11: Enzymes: Kinetics [PDF] Reading: Berg, Tymoczko & Stryer, Chapter 8, pp. 216-225 Updated on: 2/4/07 at 9:00 pm Key Concepts Kinetics is the study of reaction rates. Study of enzyme kinetics
Lecture 11: Enzymes: Kinetics [PDF] Reading: Berg, Tymoczko & Stryer, Chapter 8, pp. 216-225 Updated on: 2/4/07 at 9:00 pm Key Concepts Kinetics is the study of reaction rates. Study of enzyme kinetics
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/5/243/ra68/dc1 Supplementary Materials for Superbinder SH2 Domains Act as Antagonists of Cell Signaling Tomonori Kaneko, Haiming Huang, Xuan Cao, Xing Li, Chengjun
www.sciencesignaling.org/cgi/content/full/5/243/ra68/dc1 Supplementary Materials for Superbinder SH2 Domains Act as Antagonists of Cell Signaling Tomonori Kaneko, Haiming Huang, Xuan Cao, Xing Li, Chengjun
Simultaneous intracellular chloride and ph measurements using a GFPbased
 nature methods Simultaneous intracellular chloride and ph measurements using a GFPbased sensor Daniele Arosio, Fernanda Ricci, Laura Marchetti, Roberta Gualdani, Lorenzo Albertazzi & Fabio Beltram Supplementary
nature methods Simultaneous intracellular chloride and ph measurements using a GFPbased sensor Daniele Arosio, Fernanda Ricci, Laura Marchetti, Roberta Gualdani, Lorenzo Albertazzi & Fabio Beltram Supplementary
Supporting information for: Mechanism of lignin inhibition of enzymatic. biomass deconstruction
 Supporting information for: Mechanism of lignin inhibition of enzymatic biomass deconstruction Josh V. Vermaas,, Loukas Petridis, Xianghong Qi,, Roland Schulz,, Benjamin Lindner, and Jeremy C. Smith,,
Supporting information for: Mechanism of lignin inhibition of enzymatic biomass deconstruction Josh V. Vermaas,, Loukas Petridis, Xianghong Qi,, Roland Schulz,, Benjamin Lindner, and Jeremy C. Smith,,
Does the S2 Rod of Myosin II Uncoil upon Two-Headed Binding to Actin? A Leucine-Zippered HMM Study
 12886 Biochemistry 2003, 42, 12886-12892 Does the S2 Rod of Myosin II Uncoil upon Two-Headed Binding to Actin? A Leucine-Zippered HMM Study Tania Chakrabarty,, Chris Yengo, Corry Baldacchino, Li-Qiong
12886 Biochemistry 2003, 42, 12886-12892 Does the S2 Rod of Myosin II Uncoil upon Two-Headed Binding to Actin? A Leucine-Zippered HMM Study Tania Chakrabarty,, Chris Yengo, Corry Baldacchino, Li-Qiong
In biomolecular systems, free energy frequently needs to be
 Kinetic equilibrium of forces and molecular events in muscle contraction Erwin W. Becker* Institut für Mikrostrukturtechnik, Forschungszentrum, and Universität Karlsruhe, 76021 Karlsruhe, P.O. Box 3640,
Kinetic equilibrium of forces and molecular events in muscle contraction Erwin W. Becker* Institut für Mikrostrukturtechnik, Forschungszentrum, and Universität Karlsruhe, 76021 Karlsruhe, P.O. Box 3640,
Dynamics of an inchworm nano walker
 Dynamics of an inchworm nano walker A. Ciudad a, J.M. Sancho a A.M. Lacasta b a Departament d Estructura i Constituents de la Matèria, Facultat de Física, Universitat de Barcelona, Diagonal 647, E-08028
Dynamics of an inchworm nano walker A. Ciudad a, J.M. Sancho a A.M. Lacasta b a Departament d Estructura i Constituents de la Matèria, Facultat de Física, Universitat de Barcelona, Diagonal 647, E-08028
ATP Synthase. Proteins as nanomachines. ATP Synthase. Protein physics, Lecture 11. Create proton gradient across. Thermal motion and small objects
 Proteins as nanomachines Protein physics, Lecture 11 ATP Synthase ATP synthase, a molecular motor Thermal motion and small objects Brownian motors and ratchets Actin and Myosin using random motion to go
Proteins as nanomachines Protein physics, Lecture 11 ATP Synthase ATP synthase, a molecular motor Thermal motion and small objects Brownian motors and ratchets Actin and Myosin using random motion to go
NB-DNJ/GCase-pH 7.4 NB-DNJ+/GCase-pH 7.4 NB-DNJ+/GCase-pH 4.5
 SUPPLEMENTARY TABLES Suppl. Table 1. Protonation states at ph 7.4 and 4.5. Protonation states of titratable residues in GCase at ph 7.4 and 4.5. Histidine: HID, H at δ-nitrogen; HIE, H at ε-nitrogen; HIP,
SUPPLEMENTARY TABLES Suppl. Table 1. Protonation states at ph 7.4 and 4.5. Protonation states of titratable residues in GCase at ph 7.4 and 4.5. Histidine: HID, H at δ-nitrogen; HIE, H at ε-nitrogen; HIP,
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/1/e1500989/dc1 Supplementary Materials for An epidermis-driven mechanism positions and scales stem cell niches in plants Jérémy Gruel, Benoit Landrein, Paul Tarr,
advances.sciencemag.org/cgi/content/full/2/1/e1500989/dc1 Supplementary Materials for An epidermis-driven mechanism positions and scales stem cell niches in plants Jérémy Gruel, Benoit Landrein, Paul Tarr,
Department of Physics, University at Buffalo, Buffalo, NY INTRODUCTION
 proteins STRUCTURE O FUNCTION O BIOINFORMATICS Coarse-grained modeling of conformational transitions underlying the processive stepping of myosin V dimer along filamentous actin Wenjun Zheng* Department
proteins STRUCTURE O FUNCTION O BIOINFORMATICS Coarse-grained modeling of conformational transitions underlying the processive stepping of myosin V dimer along filamentous actin Wenjun Zheng* Department
4 Examples of enzymes
 Catalysis 1 4 Examples of enzymes Adding water to a substrate: Serine proteases. Carbonic anhydrase. Restrictions Endonuclease. Transfer of a Phosphoryl group from ATP to a nucleotide. Nucleoside monophosphate
Catalysis 1 4 Examples of enzymes Adding water to a substrate: Serine proteases. Carbonic anhydrase. Restrictions Endonuclease. Transfer of a Phosphoryl group from ATP to a nucleotide. Nucleoside monophosphate
The biological motors
 Motor proteins The definition of motor proteins Miklós Nyitrai, November 30, 2016 Molecular machines key to understand biological processes machines in the micro/nano-world (unidirectional steps): nm,
Motor proteins The definition of motor proteins Miklós Nyitrai, November 30, 2016 Molecular machines key to understand biological processes machines in the micro/nano-world (unidirectional steps): nm,
Structural basis for catalytically restrictive dynamics of a high-energy enzyme state
 Supplementary Material Structural basis for catalytically restrictive dynamics of a high-energy enzyme state Michael Kovermann, Jörgen Ådén, Christin Grundström, A. Elisabeth Sauer-Eriksson, Uwe H. Sauer
Supplementary Material Structural basis for catalytically restrictive dynamics of a high-energy enzyme state Michael Kovermann, Jörgen Ådén, Christin Grundström, A. Elisabeth Sauer-Eriksson, Uwe H. Sauer
Effects of substituting uridine triphosphate for ATP on the crossbridge cycle of rabbit muscle
 12248 Journal of Physiology (2001), 537.3, pp.907 921 907 Effects of substituting uridine triphosphate for ATP on the crossbridge cycle of rabbit muscle Chun Y. Seow, Howard D. White * and Lincoln E. Ford
12248 Journal of Physiology (2001), 537.3, pp.907 921 907 Effects of substituting uridine triphosphate for ATP on the crossbridge cycle of rabbit muscle Chun Y. Seow, Howard D. White * and Lincoln E. Ford
Online Supplementary Material. Messenger RNA Interactions in the Decoding Center Control the Rate of Translocation
 Online Supplementary Material Messenger RNA Interactions in the Decoding Center Control the Rate of Translocation Prashant K. Khade and Simpson Joseph Supplementary Figure 1 Dissociation of the f[ 35 S]Met-Phe-tRNA
Online Supplementary Material Messenger RNA Interactions in the Decoding Center Control the Rate of Translocation Prashant K. Khade and Simpson Joseph Supplementary Figure 1 Dissociation of the f[ 35 S]Met-Phe-tRNA
Structure of the α-helix
 Structure of the α-helix Structure of the β Sheet Protein Dynamics Basics of Quenching HDX Hydrogen exchange of amide protons is catalyzed by H 2 O, OH -, and H 3 O +, but it s most dominated by base
Structure of the α-helix Structure of the β Sheet Protein Dynamics Basics of Quenching HDX Hydrogen exchange of amide protons is catalyzed by H 2 O, OH -, and H 3 O +, but it s most dominated by base
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/5/e1602426/dc1 Supplementary Materials for In vivo mapping of tissue- and subcellular-specific proteomes in Caenorhabditis elegans The PDF file includes: Aaron
advances.sciencemag.org/cgi/content/full/3/5/e1602426/dc1 Supplementary Materials for In vivo mapping of tissue- and subcellular-specific proteomes in Caenorhabditis elegans The PDF file includes: Aaron
Supplementary Information
 Supplementary Information Supplementary Figure 1. Schematic pipeline for single-cell genome assembly, cleaning and annotation. a. The assembly process was optimized to account for multiple cells putatively
Supplementary Information Supplementary Figure 1. Schematic pipeline for single-cell genome assembly, cleaning and annotation. a. The assembly process was optimized to account for multiple cells putatively
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Structure of human carbamoyl phosphate synthetase: deciphering the on/off switch of human ureagenesis Sergio de Cima, Luis M. Polo, Carmen Díez-Fernández, Ana I. Martínez, Javier
SUPPLEMENTARY INFORMATION Structure of human carbamoyl phosphate synthetase: deciphering the on/off switch of human ureagenesis Sergio de Cima, Luis M. Polo, Carmen Díez-Fernández, Ana I. Martínez, Javier
Supplementary material
 Supplementary material Phosphorylation of the mitochondrial autophagy receptor Nix enhances its interaction with LC3 proteins Vladimir V. Rogov 1,*, Hironori Suzuki 2,3,*, Mija Marinković 4, Verena Lang
Supplementary material Phosphorylation of the mitochondrial autophagy receptor Nix enhances its interaction with LC3 proteins Vladimir V. Rogov 1,*, Hironori Suzuki 2,3,*, Mija Marinković 4, Verena Lang
Supplementary Figures
 1 Supplementary Figures Supplementary Figure 1 Type I FGFR1 inhibitors (a) Chemical structures of a pyrazolylaminopyrimidine inhibitor (henceforth referred to as PAPI; PDB-code of the FGFR1-PAPI complex:
1 Supplementary Figures Supplementary Figure 1 Type I FGFR1 inhibitors (a) Chemical structures of a pyrazolylaminopyrimidine inhibitor (henceforth referred to as PAPI; PDB-code of the FGFR1-PAPI complex:
