Supplementary Information
|
|
|
- Scot Bond
- 5 years ago
- Views:
Transcription
1 Supplementary Information Adenosyltransferase Tailors and Delivers Coenzyme B 12 Dominique Padovani 1,2, Tetyana Labunska 2, Bruce A. Palfey 1, David P. Ballou 1 and Ruma Banerjee 1,2 * 1 Biological Chemistry Department, University of Michigan, Ann Arbor, MI and 2 Redox Biology Center and Department of Biochemistry, University of Nebraska, Lincoln, NE
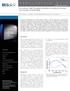
2 Figure S1. Binding of AdoCbl to MCM. Representative kinetic traces (black) for the binding of AdoCbl (11, 23 and 40 µm, bottom to top respectively) to apo-mcm (7 µm after mixing) in 50 mm KPi buffer, ph 7.5 at 20ºC. The kinetic simulations (in red) were generated using the model shown in Fig. S5a (upper) and the parameters k 1 -k 6 for the dissociative mechanism shown in Table SI. Inset: Dependence of the k obs for AdoCbl binding to MCM ( ) on cofactor concentration. The red line represents the simulated dependence of k obs on AdoCbl and yields values for the slope and intercept of µm -1 s -1 and 0.49 s -1 respectively, similar to the experimentally determined values of 0.10±0.02 µm -1 s -1 and 0.26±0.05 s -1. Based on the observed ΔA 525nm (Δε of 1.06 mm -1 cm -1 ), one AdoCbl is bound per heterodimer of MCM.
3 Figure S2. Properties of ATR from M. extorquens AM1. (a) The oligomeric state of ATR in solution was determined by size exclusion chromatography performed as described under Supplementary Methods. The molecular weights of the standards (solid line) are: thyroglobulin (670 kd), γ-globulin (158 kd), ovalbumin (44 kd), myoglobin (17 kda) and vitamin B 12 (1.35 kd). The retention time of 52.0 ± 0.2 min for ATR (dashed line) yields a native molecular mass of 55.2 ± 1.1 kd, corresponding to a homotrimer. (b) Scatchard analysis of AdoCbl binding to apo-atr reveals the presence of two nonequivalent AdoCbl binding sites per ATR trimer, with n 1 = 1.3 ± 0.3, K D1 = 0.12 ± 0.03 µm and n 2 = 1.99 ± 0.10,
4 K D2 = 1.0 ± 0.16 µm (n=3). Inset: Titration of AdoCbl (34.7 µm) with increasing concentrations of apo-atr ( µm) as described under Supplementary Methods. Representative traces are shown for clarity. (c) ITC titration of apo-atr (30.6 µm) in 50 mm KPi buffer, ph 7.5, 300 mm KCl with aliquots of a 1.0 mm stock solution of AdoCbl. The top panel shows the raw data in power versus time. The area under each spike is proportional to the heat produced with each injection. The lower panel shows the integrated areas normalized to the number of moles of AdoCbl added with each injection. Data were well-fitted to a two-site binding model and yielded values of K D1 = 0.6 ± 0.1 µm and K D2 = 1.5 ± 0.4 µm respectively. (d) Representative kinetic traces (in black) for the binding of AdoCbl (10-30 µm top to bottom) to apo-atr (5 µm). The simulations (in red) were performed using the model described in Fig. S5a (lower) and using the kinetic parameters (k 7 -k 14 ) for the dissociative model shown in Table SI. Inset: Dependence of k obs1 ( ) and k obs2 ( ) for binding of AdoCbl on cofactor concentration. A linear fit to the experimental data yields the following values: slope 1 =3.4±0.4 µm -1 s -1, intercept 1 =44±9 s -1, slope 2 =1.3±0.4 µm -1 s -1 and intercept 2 =13±3 s -1. The simulated data (in red) yield comparable values: slope 1 ~3.75 µm -1 s -1, intercept 1 ~30.2 s -1, slope 2 ~1.18 µm -1 s -1 and intercept 2 ~24.9 s -1. The data are represented as mean ± S.D.
5 Figure S3. Kinetics of interprotein AdoCbl transfer. (a) A representative set of stopped-flow scans for the transfer of AdoCbl between holo-atr (9.5 µm AdoCbl after mixing) and wildtype apo-mcm (90 µm, after mixing) in 50 mm KPi buffer, ph 7.5, at 20ºC. Traces were recorded every 0.5 sec. The arrows denote the direction of absorption changes. (b) Representative stopped-flow traces observed for AdoCbl transfer between holo-atr (9.6 µm bound AdoCbl, after mixing) and wild-type apo-mcm (67.5 µm, after mixing) (trace 1) or between holo-mcm (15 µm, after mixing) and apo-atr (38.5 µm, after mixing) (trace 2). The red lines represent biphasic fits (R 2 ~0.998) to the experimental data. In the range of concentrations studied, transfer of AdoCbl occurs with changes in amplitude ΔA 1 ~0.9-1ΔA 2 and ΔA 1 ~ ΔA 2, in the forward and reverse directions, respectively. The k obs1 and k obs2 values obtained from the kinetic traces at each concentration of apo-[e2] were then used to generate Figs. S4a and b.
6 Figure S4. Dependence of AdoCbl transfer kinetics on the concentration of the acceptor protein. (a) Hyperbolic dependence of k obs1 and k obs2 (inset) for AdoCbl transfer from holo- ATR on the concentration of apo-mcm. The values for forward transfer k trans+1 and k trans+2, were obtained as described under Supplementary Methods. We note that the k obs values are not true rate constants but eigenvalues that are composites of individual rate constants. The eigenvalues appear to decrease because the amplitudes become larger at increasing concentrations of E2 and it therefore takes longer to make the complete transfers. (b) Hyperbolic dependence of k obs1 and k obs2 (inset) for AdoCbl transfer from holo-mcm on the concentration of apo-atr. The values for reverse transfer (k trans-1, k trans-2 ) were obtained from these plots as described under Supplementary Methods. The data are represented as mean ± S.D.
7 Figure S5. Alternative mechanisms for AdoCbl transfer. Equations describing the dissociative (a) and associative (b) models for cofactor binding to and transfer between ATR and MCM.
8 Figure S6. Temperature dependence of AdoCbl binding to MCM. Comparison of the temperature dependence of the observed rates of transfer k trans+1 ( ) and k trans+2 ( ) from holo-atr (10 µm) to apo-mcm (100 µm) and of the binding of AdoCbl (50 µm) from solution to apo-mcm (10 µm) ( ) in 50 mm KPi buffer, ph 7.5. The data are represented as mean ± S.D.
9 Table S1 Kinetic parameters obtained from simulations of the dissociative and associative mechanisms. Dissociative mechanism Associative mechanism Kinetic parameters K D Kinetic parameters K D k 1, µm -1 s } 43.5 µm k 1, µm -1 s k 2, s k 2, s } 7.05 µm k 3, s k 3, s k 4, s k 4, s k 5, s k 5, s k 6, s k 6, µm -1 s k 7, µm -1 s k 7, µm -1 s } 69.4 µm k 8, s k 8, s } 4.35 µm } 0.36 µm k 9, s k 9, s k 10, s k 10, s k 11, µm -1 s k 11, s } 0.37 µm k 12, s k 12, µm 1 s } 2.19 µm k 13, s k 14, s -1 20
10 Table S2 Comparison of the activation parameters for transfer of AdoCbl from holo-atr to apo-mcm and for the binding of AdoCbl to apo-mcm from solution. Parameters Site 1 a Site 2 a solution E a, kcal/mol 10.3± ± ±0.9 ΔH, kcal/mol 9.7± ± ±0.9 ΔS, cal/(mol.k) -24.4± ± ±3.2 ΔG, kcal/mol 16.8± ± ±0.1 a The sites refer to the two binding sites for AdoCbl on ATR.
11 Table S3 Kinetic and thermodynamic parameters for the H596N/A mutants of MCM. a Parameters Wild-type b H596A H596N k cat, s ± ± ±0.006 K M-[R]MCoA, µm 86±13 107±32 64±21 k cat /K M, M -1 s -1 (1.53±0.19) ±6 322±75 K act-adocbl, µm 1.6±0.1 c 4.5± ±0.3 K D, µm d 0.40 ± ± ± 1.7 a The kinetic parameters for MCM were determined using the radiolabeled assay at 37 C as described under Methods and represent the average (± S.D.) of three independent experiments. b From Padovani, D., and Banerjee, R. (2006) Biochemistry 45, except for (c), which was determined as described under Methods. d Determined by ITC (in 50 mm KPi, ph 7.5, containing 300 mm KCl, at 20 C) as described under Supplementary Methods.
12 Supplementary Methods Materials. AdoCbl and other reagent grade chemicals were purchased from Sigma. [ 14 C]- CH 3 -malonyl-coa (56 Ci/mol) was purchased from New England Nuclear. Construction of Site-Specific Mutants. The plasmid containing the M. extorquens AM1 MCM and ATR were a generous gift from Mary E. Lidstrom (University of Washington, Seattle). The site-directed mutants H596N and H596A were created using the QuickChange kit (Stratagene) and the following sense primers: 5 - CAAGATGGGCCAGGACGGGAACGACCGCGGCCAGAAGGTG-3 for H596N and 5 - CAAGATGGGCCAGGACGGGGCCGACCGCGGCCAGAAGGTG-3 for H596A. The antisense mutagenic primers had the complementary sequences. Following PCR amplification, the mutations were confirmed by nucleotide sequence determination at the Genomics Core Facility (University of Nebraska-Lincoln). Enzymes Expression and Purification. The wild-type and mutant MCMs 1 and ATR 2 were purified as previously described. Enzyme assays. The specific activity of MCM was determined in the radiolabeled assay as described previously. 3 Each experiment was performed in triplicate. (i) Kinetic parameters for wild-type and mutant MCM. The kinetic parameters for the mutants, H596A/N, were determined at 37 C by increasing the duration of the assay from 3 to 10 min and the amount of enzyme 200- to 1,000-fold. K M and V max were determined in the presence of varying concentrations of (R,S)-[ 14 C]-methylmalonyl-CoA ( µm) while keeping constant the concentration of AdoCbl at 100 µm in the assay. The K act for AdoCbl was obtained by varying the concentration of the cofactor (1-50 µm) in the presence of a saturating concentration of the substrate (4 mm (R,S)-[ 14 C]-methylmalonyl- CoA).
13 (ii) Modulation of MCM activity by ATR. To assess MCM activity under conditions where the percent cofactor transfer was monitored spectroscopically, the concentration of enzymes in the assay had to be increased ~1,000-fold. As a consequence, after the AdoCbl transfer experiments, the activity measurements were performed on ice to be under initial velocity conditions in the radiolabeled MCM assay. AdoCbl transfer was accomplished at 20ºC for 10 min by incubating in the forward direction, 5 µm holo-atr (10 µm bound AdoCbl) with various concentrations of apo-mcm ( µm) and in the reverse direction, 10 µm holo-mcm (containing a stoichiometric amount of AdoCbl) with varying concentrations of apo-atr (0-50 µm final concentration), in 50 mm potassium phosphate (KPi) buffer, ph 7.5. Then, the solutions were incubated for 20 min on ice before starting the assay by addition of 5 mm ice-cold (R,S)-[ 14 C]-methylmalonyl-CoA. After 1 min incubation, the reaction mixtures were quenched and the samples treated as previously described. 3 Oligomeric State of ATR. To determine the oligomeric state of apo-atr in solution, ~0.2-1 mg of the enzyme was loaded on a 2 x 70 cm Sephacryl 200 column in 50 mm KPi, ph 7.5, containing 100 mm KCl at a flow rate of 2 ml min -1. Prior to loading, the protein was filtered through a 0.2-µm Anotop 10 filter (Whatman). The column was calibrated using gel filtration standards from Bio-Rad. AdoCbl binding to ATR: UV visible spectroscopy. The large blue shift in the UV-visible spectrum (from 525 to 458 nm) accompanying binding of the free cofactor to ATR was monitored to determine the binding affinity of ATR for AdoCbl. Briefly, successive aliquots of apo-atr (1-45 µm) were added to a fixed concentration of AdoCbl (20-35 µm). Following each addition of apo-atr, the solution was incubated at 20 C for 10 min prior to recording of the UV-visible spectrum. The concentrations of bound (L b ) and unbound (L u ) AdoCbl after each addition of apo-atr were determined using equations 2 and 3.
14 L b =(ΔA 525nm /ΔA 525nm max)*[adocbl] total [2] L u =[AdoCbl] total -L b [3] The values from equations 2 and 3 were used to obtain the parameters, K D and n (the number of binding sites) using the Scatchard plot described by equation 4. The experiment was performed in triplicate. ([L b ]/[E]) = ( n " [L b ]/[E]) [L u ] 1 K D [4] Since the Scatchard plot was clearly biphasic, each phase was analyzed independently (Fig. S2b). Isothermal titration calorimetry (ITC). All calorimetric binding experiments were performed as described previously. 1 Each experiment was performed at least in triplicate and the data were analyzed using Microcal ORIGIN software. (i) Binding of AdoCbl to apo-atr. Apo-ATR (10-45 µm in different experiments) was titrated with fifty nine 5 µl aliquots of a µm solution of AdoCbl in 50 mm KPi buffer, ph 7.5 at 20.0 ± 0.1 C. When the same experiments were performed with buffer containing 300 mm KCl, no significant difference was observed. The calorimetric signals were integrated, and the data fit well to a two-site binding model to estimate the equilibrium association constants, K A, and the binding enthalpies, ΔH. The Gibbs free energy of binding, ΔG, and the entropic contribution to the binding free energy, -TΔS, were calculated using equations 5 and 6. ΔG = -RT ln K A [5] ΔG = ΔH - TΔS [6]
15 (ii) Binding of AdoCbl to MCM mutants. Enzyme (10-50 µm of H596N/A mutants of MCM) in 50 mm KPi buffer, ph 7.5, was titrated with twenty nine 10 µl aliquots of a µm solution of AdoCbl at 20.0 ± 0.1 C. When the same experiments were carried out in buffer containing 300 mm KCl, very small changes ( 6%) in the binding parameters were observed. The data were analyzed as described for ATR but using a single-site binding model. AdoCbl transfer between ATR and MCM. The transfer of AdoCbl between the two enzyme active sites (in 50 mm KPi buffer, ph 7.5) was monitored by absorption spectroscopy at 20ºC. In the forward direction, 7 µm holo-atr (14 µm bound AdoCbl) was added to a final concentration of µm apo-mcm (wild-type or mutant). In the reverse direction, apo- ATR (3-60 µm final concentration) was added to 15 µm wild-type holo-mcm (containing 15 µm bound AdoCbl). The amount of AdoCbl transferred was estimated from the increase (forward) or decrease (reverse) in absorbance at 525 nm (Δε 525 nm = 7.75 mm -1 cm -1 for wild-type MCM and Δε 525 nm = 5.46 mm -1 cm -1 for the mutants). Stopped-flow spectroscopy. Stopped-flow UV-visible spectroscopy experiments were performed on an Applied Photophysics spectrophotometer (ISX.MV18) under red light illumination. Rapid-scanning stopped-flow fluorescence kinetic measurements were made using an OLIS stopped-flow fluorescence spectrophotometer. Fluorescence emission spectra were recorded between 325 and 474 nm using an emission slit width of 0.6 µm and acquired at 62 scans/s. The excitation wavelength was 282 nm and two consecutive slits of 0.6 µm width were used, as reported previously. 4 All solutions were prepared in 50 mm KPi buffer, ph 7.5, filtered through a 0.02 µm syringe filter (Whatman), transferred to the loading syringes and allowed to equilibrate for at least 20 min before initiating the experiments. An external water bath was used to maintain the loading syringes and the mixing chamber at 20 ± 0.5 ºC.
16 (i) Kinetic analysis of AdoCbl binding to apo-mcm. The kinetic parameters for binding of AdoCbl to apo-mcm were determined by mixing AdoCbl ( µm before mixing) and a fixed concentration of apo-mcm (10-15 µm before mixing in different experiments) in 50 mm KPi buffer, ph 7.5, and monitoring the increase in A 525nm (Δε 525 nm = 1.06 mm -1 cm -1 ). Kinetic traces were well-fitted to a single-exponential function to obtain k obs and the change in amplitude, ΔA, according to equation 7, where A is the absorbance at time t and A 0 is the offset for the exponential increase. A = A o + "A(1# e (#k obs t ) ) [7] The k obs value was then plotted as a function of AdoCbl concentration. The kinetic parameters for binding of AdoCbl to apo-mcm were also determined by mixing AdoCbl (2-15 µm after mixing) and a fixed concentration of apo-mcm (0.5-1 µm after mixing in different experiments) in 50 mm KPi buffer, ph 7.5, and monitoring the quenching of fluorescence at 340 nm. Kinetic traces were well-fitted to a single-exponential function to obtain k obs and the change in fluorescence, ΔF, according to equation 8, where F is the absorbance at time t and F 0 is the offset for the exponential decrease and c is the rate of background fluorescence decay. F ( = F + " F(1! e o! kobs t) )! ct [8] The rate of background fluorescence decay (c) was recorded by mixing µm apo- MCM (after mixing) against buffer. The k obs value was then plotted as a function of AdoCbl concentration. (ii) Kinetic analysis of AdoCbl binding to apo-atr. Binding of AdoCbl to apo-atr was carried out as described above for MCM by rapidly mixing varying concentrations of AdoCbl ( µm before mixing) with a fixed concentration of apo-atr (5-16 µm before
17 mixing) in 50 mm KPi buffer, ph 7.5, and monitoring the decrease in A 525nm (Δε 525 nm = -6.7 mm -1 cm -1 ). Kinetic traces were best fitted to a double exponential function to obtain k obs1 and k obs2 and the changes in amplitude associated with each phase, ΔA 1 and ΔA 2, as described in equation 9, where A is the absorbance at time t and A 0 is the offset for the exponential decrease. A = A o + "A 1 e (#k obs1t ) + "A 2 e (#k obs 2t ) [9] Then, the k obs (k obs1 and k obs2 ) values were each plotted as a function of AdoCbl concentration and the plots were fitted to a linear function to obtain the slope and intercept for each binding site. (iii) AdoCbl transfer experiments. Cofactor transfer in the forward direction (from holo-atr to apo-mcm) was investigated by rapidly mixing a fixed concentration of holo-atr (10-20 µm before mixing) with increasing concentrations of apo-mcm (3-180 µm before mixing). The concentration of AdoCbl bound to ATR was determined spectrophotometrically (ε 458 nm = 8 mm -1 cm -1 ). Transfer was monitored at 525 nm (Δε 525nm ~ 7.75 mm -1 cm -1 ) or 458 nm (Δε 428nm ~ mm -1 cm -1 ). Kinetic traces were best fitted to a double exponential function using equation 9, when monitoring the transfer at 458 nm, or equation 10, when monitoring the transfer at 525 nm, to obtain both the observed rates of transfer (k obs1 and k obs2 ) and the change in amplitude (ΔA 1 and ΔA 2 ) associated with each phase. A = A o + "A 1 (1# e (#k obs1t ) ) + "A 2 (1# e (#k obs 2t ) ) [10] In this equation, A represents the absorbance at time t and A 0 the offset for the exponential increase. The dependence on apo-mcm concentration associated with the transfer of AdoCbl from each binding site on holo-atr was then obtained by plotting the observed rates of transfer (k obs1 and k obs2 ) as a function of apo-mcm concentration and by fitting the
18 data to a hyperbolic decay function, yielding the values for k trans+1 and k trans+2 at saturating concentrations of apo-mcm. At least 6 independent traces were recorded at each apo-mcm concentration and the experiment was performed in duplicate. The reactions were also monitored in a scanning mode with a photodiode array detector. A total of scans were collected over a range of sec using the program XScan (Applied Photophysics) and analyzed using Sigma Plot. We note that in the scanning mode, photolysis of cofactor occurs after ~6 sec. To monitor the transfer of AdoCbl in the reverse direction, varying concentrations of apo- ATR (4-160 µm before mixing) and a fixed concentration of holo-mcm (20-30 µm before mixing, based on the bound AdoCbl concentration using ε 525 nm = 9.06 mm -1 cm -1 ) were rapidly mixed and absorbance changes were monitored at 458 nm or 525 nm. The kinetic traces were well-fitted to a double exponential function using equations 9 and 10. The k trans- 1 and k trans-2 values for the reverse transfer of AdoCbl from holo-mcm to each binding site of apo-atr were estimated at saturating concentrations of apo-atr, as described above. (iv) Thermodynamic parameters for AdoCbl transfer- The activation parameters for the transfer of AdoCbl from holo-atr were obtained from the temperature dependence of the k trans+1 and k trans+2 between 4-28ºC. Holo-ATR (20 µm before mixing) and apo-mcm (200 µm before mixing) were rapidly mixed and absorption change at 525 nm was monitored. The enthalpy (ΔH ) and the entropy of activation (ΔS ) were obtained from the slope and intercept, respectively, of the Eyring plot (equation 11), where where k B is the Boltzmann ln(k transfer /T) = ln (k B /ħ) - ΔH /RT + ΔS /R [11] constant, ħ is Planck's constant and R is the molar gas constant. The Gibbs energy of activation was then obtained at a given temperature, T, using equation 12. ΔG = ΔH - TΔS [12]
19 The activation parameters for binding of AdoCbl (100 µm before mixing) to apo-mcm (20 µm before mixing) were determined between 7 and 32 C as analyzed as described above. Kinetic Simulations- The kinetic simulation program, Berkeley Madonna ( was used to fit the cofactor binding data for the individual enzymes and to distinguish between the dissociative versus associative mechanisms for cofactor transfer between ATR and MCM (described in Fig. S5). All curve fittings were performed using the Runge Kutta 4 method. To simulate the data for cofactor binding to the individual enzymes, 3-5 representative experimental traces were employed (i) Cofactor Binding to MCM. Cofactor binding to MCM was simulated using the equations described in Fig. 5a (k 1 -k 6 ), which describes a model for B 12 binding that has been reported previously ( Model B described in Chowdhury, S., and Banerjee, R. Biochemistry 38, (1999)). The salient steps in the model include: formation of a pre-docking complex of enzyme and cofactor (MCM B 12 )*, followed by a ph-sensitive step (with a pk a ~7.3) to give the protonated complex, (MCM B 12 H + )*. A subsequent conformational change results in docking of the cofactor and enzyme (MCM B 12 H + ). For the simulations, the following assumptions were made and the kinetic parameters were allowed to float: (i) the macroscopic K D (k 2 /k 1 ), for the pre-docking complex, is high ( µm range); (ii) the ph sensitive step is slow (k 4 >k 3 ) and (iii) cofactor docking after the phsensitive step is fast (k 5 >>k 6 ). The simulated traces were fitted to equation 7 and the dependence of the rate constant for AdoCbl binding on cofactor concentration was compared to the experimental data (Fig. S1, inset). An excellent correspondence was observed between the simulated and experimental data (Fig. S1), supporting the validity of the model for B 12 binding to MCM shown in Fig. S5a.
20 (ii) Cofactor Binding to ATR. Cofactor binding to ATR was simulated using equations k 7 -k 14 in Fig. S5a. The salient feature of the model is that it describes negative cooperativity between the two binding sites for B 12 in ATR. A conformational change is shown to follow the binding of each equivalent of B 12. The simulations were subjected to the following assumptions and the kinetic parameters were allowed to float: (i) the first conformational change is very fast (k 9 high); (ii) the first conformational change affects the second conformational change that will then become the rate-limiting step for the binding of the second ligand (k 13 <k 14 ); (iii) the macroscopic K D (k 8 /k 7 and k 12 /k 11 ), for the pre-docking complexes are in the µm range. The simulated kinetic traces were fitted to equation 9 above and the k obs (k obs1 and k obs2 ) values were plotted as a function of AdoCbl concentration to determine the slope and intercept values for the binding of the first and second equivalent of B 12 (Fig. S2d, inset). The correspondence between the experimental and simulated kinetic parameters (Fig. S2d) provides support for the validity of the model describing cofactor binding to ATR. (iii) Dissociative Mechanism. The excellent correspondence between the experimental and simulated data for cofactor binding to the individual enzymes, ATR and MCM described above, provided confidence in the simulated kinetic parameters that were then employed to predict the behavior of the system when the cofactor was transferred from one enzyme to the other via a dissociative mechanism. The values for the first and second order rate constants for binding of B 12 to MCM and ATR (k 1 -k 14 ) described in Table SI were used to fit the dissociative model described in Fig. S5a. Five kinetic traces (three in the forward direction and two in the reverse direction) were fitted to the model (Fig. 2a). (iv) Associative Mechanism. For the associative mechanism, the same five experimental traces used for the dissociative mechanism, were fitted to the model described by equations k 1 -k 12 in Fig. S5b. The salient features of the associative model
21 are: (i) unlike the dissociative model, a rate-limiting ph-sensitive step is not involved, (ii) cofactor transfer occurs in a step-wise manner and thus, a protein-protein complex forms between one ATR and one MCM at a time, (iii) the K D values for complex formation are high, in the micromolar range, consistent with the complex not being detectable by gel filtration chromatography or in a native gel; (iv) the on-rates are higher in the reverse direction than in the forward direction (k 1, k 7 < k 6, k 12 ), since under our experimental conditions, the reverse transfer (from holo-mcm to apo-atr) is favored; (v) the initial estimates for the kinetic parameters were: k 5 =k 11 =100 s -1 and k 6 =k 12 =10 µm -1 s -1, k 2 =k 8 =10 s -1 and k 1 =k 7 =2 µm -1 s -1 ; (vi) the rates of cofactor transfer are faster in the reverse than in the forward direction (k 10 >k 9 and k 4 >k 3 ). The parameters obtained from the initial simulation run were then adjusted manually using the sliders in the software package to obtain values that yielded better overall fits to the experimental data (Fig. 2b) and are reported in Table SI. References for supporting methods 1. Padovani, D., Labunska, T. & Banerjee, R. J. Biol. Chem. 281, (2006). 2. Yamanishi, M., Labunska, T. & Banerjee, R. J Am Chem Soc 127, (2005). 3. Taoka, S., Padmakumar, R., Lai, M.-t., Liu, H.-w. & Banerjee, R. J. Biol. Chem. 269, (1994). 4. Chowdhury, S. & Banerjee, R. Biochemistry 38, (1999).
ISoTherMal TITraTIon Calorimetry
 ISoTherMal TITraTIon Calorimetry With the Nano ITC, heat effects as small as 1 nanojoules are detectable using one nanomole or less of biopolymer. The Nano ITC uses a solid-state thermoelectric heating
ISoTherMal TITraTIon Calorimetry With the Nano ITC, heat effects as small as 1 nanojoules are detectable using one nanomole or less of biopolymer. The Nano ITC uses a solid-state thermoelectric heating
1. Use the Data for RNAse to estimate:
 Chem 78 - - Spr 1 03/14/01 Assignment 4 - Answers Thermodynamic Analysis of RNAseA Denaturation by UV- Vis Difference Absorption Spectroscopy (and Differential Scanning Calorimetry). The accompanying excel
Chem 78 - - Spr 1 03/14/01 Assignment 4 - Answers Thermodynamic Analysis of RNAseA Denaturation by UV- Vis Difference Absorption Spectroscopy (and Differential Scanning Calorimetry). The accompanying excel
Crystal Violet as a Fluorescent Switch-On Probe for I-Motif: Label-Free DNA-Based Logic Gate
 Crystal Violet as a Fluorescent Switch-On Probe for I-Motif: Label-Free DNA-Based Logic Gate Dik-Lung Ma,* a Maria Hiu-Tung Kwan, a Daniel Shiu-Hin Chan, a Paul Lee, a Hui Yang, a Victor Pui-Yan Ma, a
Crystal Violet as a Fluorescent Switch-On Probe for I-Motif: Label-Free DNA-Based Logic Gate Dik-Lung Ma,* a Maria Hiu-Tung Kwan, a Daniel Shiu-Hin Chan, a Paul Lee, a Hui Yang, a Victor Pui-Yan Ma, a
Microcalorimetry for the Life Sciences
 Microcalorimetry for the Life Sciences Why Microcalorimetry? Microcalorimetry is universal detector Heat is generated or absorbed in every chemical process In-solution No molecular weight limitations Label-free
Microcalorimetry for the Life Sciences Why Microcalorimetry? Microcalorimetry is universal detector Heat is generated or absorbed in every chemical process In-solution No molecular weight limitations Label-free
Problem solving steps
 Problem solving steps Determine the reaction Write the (balanced) equation ΔG K v Write the equilibrium constant v Find the equilibrium constant using v If necessary, solve for components K K = [ p ] ν
Problem solving steps Determine the reaction Write the (balanced) equation ΔG K v Write the equilibrium constant v Find the equilibrium constant using v If necessary, solve for components K K = [ p ] ν
Presentation Microcalorimetry for Life Science Research
 Presentation Microcalorimetry for Life Science Research MicroCalorimetry The Universal Detector Heat is either generated or absorbed in every chemical process Capable of thermal measurements over a wide
Presentation Microcalorimetry for Life Science Research MicroCalorimetry The Universal Detector Heat is either generated or absorbed in every chemical process Capable of thermal measurements over a wide
LABORATORY OF ELEMENTARY BIOPHYSICS. Isothermal Titration Calorimetry as a tool for determining thermodynamic parameters of chemical reactions
 LABORATORY OF ELEMENTARY BIOPHYSICS Experimental exercises for III year of the First cycle studies Field: Applications of physics in biology and medicine Specialization: Molecular Biophysics Isothermal
LABORATORY OF ELEMENTARY BIOPHYSICS Experimental exercises for III year of the First cycle studies Field: Applications of physics in biology and medicine Specialization: Molecular Biophysics Isothermal
Lecture 13: Data Analysis for the V versus [S] Experiment and Interpretation of the Michaelis-Menten Parameters
![Lecture 13: Data Analysis for the V versus [S] Experiment and Interpretation of the Michaelis-Menten Parameters Lecture 13: Data Analysis for the V versus [S] Experiment and Interpretation of the Michaelis-Menten Parameters](/thumbs/89/100978514.jpg) Biological Chemistry Laboratory Biology 3515/Chemistry 3515 Spring 2018 Lecture 13: Data Analysis for the V versus [S] Experiment and Interpretation of the Michaelis-Menten Parameters 20 February 2018
Biological Chemistry Laboratory Biology 3515/Chemistry 3515 Spring 2018 Lecture 13: Data Analysis for the V versus [S] Experiment and Interpretation of the Michaelis-Menten Parameters 20 February 2018
Isothermal titration calorimetry (ITC)
 Isothermal titration calorimetry (ITC) Peter.gimeson@malvern.com Why microcalorimetry? Label-free Broad dynamic range Information rich Ease-of-use Direct measurement of heat change (ITC) Direct measurement
Isothermal titration calorimetry (ITC) Peter.gimeson@malvern.com Why microcalorimetry? Label-free Broad dynamic range Information rich Ease-of-use Direct measurement of heat change (ITC) Direct measurement
Table 1. Kinetic data obtained from SPR analysis of domain 11 mutants interacting with IGF-II. Kinetic parameters K D 1.
 Kinetics and Thermodynamics of the Insulin-like Growth Factor II (IGF-II) Interaction with IGF-II/Mannose 6-phosphate Receptor and the function of CD and AB Loop Solvent-exposed Residues. Research Team:
Kinetics and Thermodynamics of the Insulin-like Growth Factor II (IGF-II) Interaction with IGF-II/Mannose 6-phosphate Receptor and the function of CD and AB Loop Solvent-exposed Residues. Research Team:
Supplementary Information. Overlap between folding and functional energy landscapes for. adenylate kinase conformational change
 Supplementary Information Overlap between folding and functional energy landscapes for adenylate kinase conformational change by Ulrika Olsson & Magnus Wolf-Watz Contents: 1. Supplementary Note 2. Supplementary
Supplementary Information Overlap between folding and functional energy landscapes for adenylate kinase conformational change by Ulrika Olsson & Magnus Wolf-Watz Contents: 1. Supplementary Note 2. Supplementary
Analysis of nucleotide binding to p97 reveals the properties of a tandem AAA hexameric ATPase
 SUPPLEMENTARY INFORMATION Analysis of nucleotide binding to p97 reveals the properties of a tandem AAA hexameric ATPase Louise C Briggs, Geoff S Baldwin, Non Miyata, Hisao Kondo, Xiaodong Zhang, Paul S
SUPPLEMENTARY INFORMATION Analysis of nucleotide binding to p97 reveals the properties of a tandem AAA hexameric ATPase Louise C Briggs, Geoff S Baldwin, Non Miyata, Hisao Kondo, Xiaodong Zhang, Paul S
CHAPTER - 3 ANALYTICAL PROFILE. 3.1 Estimation of Drug in Pharmaceutical Formulation Estimation of Drugs
 CHAPTER - 3 ANALYTICAL PROFILE 3.1 Estimation of Drug in Pharmaceutical Formulation 3.1.1 Estimation of Drugs ANALYTICAL PROFILE 84 3.1 ESTIMATION OF DRUG IN PHARMACEUTICAL FORMULATION. Agrawal A et al
CHAPTER - 3 ANALYTICAL PROFILE 3.1 Estimation of Drug in Pharmaceutical Formulation 3.1.1 Estimation of Drugs ANALYTICAL PROFILE 84 3.1 ESTIMATION OF DRUG IN PHARMACEUTICAL FORMULATION. Agrawal A et al
S2004 Methods for characterization of biomolecular interactions - classical versus modern
 S2004 Methods for characterization of biomolecular interactions - classical versus modern Isothermal Titration Calorimetry (ITC) Eva Dubská email: eva.dubska@ceitec.cz Outline Calorimetry - history + a
S2004 Methods for characterization of biomolecular interactions - classical versus modern Isothermal Titration Calorimetry (ITC) Eva Dubská email: eva.dubska@ceitec.cz Outline Calorimetry - history + a
Supplemental Materials and Methods
 Supplemental Materials and Methods Time-resolved FRET (trfret) to probe for changes in the Box A/A stem upon complex assembly U3 MINI was folded and the decay of Fl fluorescence was measured at 20 ºC (see
Supplemental Materials and Methods Time-resolved FRET (trfret) to probe for changes in the Box A/A stem upon complex assembly U3 MINI was folded and the decay of Fl fluorescence was measured at 20 ºC (see
Biological Thermodynamics
 Biological Thermodynamics Classical thermodynamics is the only physical theory of universal content concerning which I am convinced that, within the framework of applicability of its basic contents, will
Biological Thermodynamics Classical thermodynamics is the only physical theory of universal content concerning which I am convinced that, within the framework of applicability of its basic contents, will
Cholera Toxin Invasion
 Protein-carbohydrate interactions: Isothermal Titration Calorimetry Dr Bruce Turnbull School of Chemistry and Astbury Centre for Structural Molecular Biology University of Leeds Cholera Toxin Invasion
Protein-carbohydrate interactions: Isothermal Titration Calorimetry Dr Bruce Turnbull School of Chemistry and Astbury Centre for Structural Molecular Biology University of Leeds Cholera Toxin Invasion
TA Instruments Application Note
 Q ( µj) TA Instruments Application Note Isothermal Titration Calorimetry (ITC) with Reduced Cell Volumes: A Comparison of the TA Instruments Nano ITC-Low Volume with the GE Healthcare Auto-iTC 200. Colette
Q ( µj) TA Instruments Application Note Isothermal Titration Calorimetry (ITC) with Reduced Cell Volumes: A Comparison of the TA Instruments Nano ITC-Low Volume with the GE Healthcare Auto-iTC 200. Colette
Effect of the Single and Double Chain Surfactant Cobalt(III) Complexes on Their Hydrophobicity, Micelle Formation,
 Electronic Supplementary Material (ESI) for Inorganic Chemistry Frontiers. This journal is the Partner Organisations 2014 Supplementary Information Effect of the Single and Double Chain Surfactant Cobalt(III)
Electronic Supplementary Material (ESI) for Inorganic Chemistry Frontiers. This journal is the Partner Organisations 2014 Supplementary Information Effect of the Single and Double Chain Surfactant Cobalt(III)
A BODIPY-based fluorescent probe for the differential
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 15 Supporting information A BODIPY-based fluorescent probe for the differential recognition of Hg(II)
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 15 Supporting information A BODIPY-based fluorescent probe for the differential recognition of Hg(II)
A Single Outer Sphere Mutation Stabilizes apo- Mn Superoxide Dismutase by 35 C and. Disfavors Mn Binding.
 Supporting information for A Single Outer Sphere Mutation Stabilizes apo- Mn Superoxide Dismutase by 35 C and Disfavors Mn Binding. Anne-Frances Miller* and Ting Wang Department of Chemistry, University
Supporting information for A Single Outer Sphere Mutation Stabilizes apo- Mn Superoxide Dismutase by 35 C and Disfavors Mn Binding. Anne-Frances Miller* and Ting Wang Department of Chemistry, University
Substrate-dependent switching of the allosteric binding mechanism of a dimeric enzyme
 Supplementary Information: Substrate-dependent switching of the allosteric binding mechanism of a dimeric enzyme Lee Freiburger, 1 Teresa Miletti, 1 Siqi Zhu, 1 Oliver Baettig, Albert Berghuis, Karine
Supplementary Information: Substrate-dependent switching of the allosteric binding mechanism of a dimeric enzyme Lee Freiburger, 1 Teresa Miletti, 1 Siqi Zhu, 1 Oliver Baettig, Albert Berghuis, Karine
A Determination of DNA-DAPI Binding using Fluorescence Spectroscopy
 CHEM 311L Revision 1.2 A Determination of DNA-DAPI Binding using Fluorescence Spectroscopy In this Laboratory Exercise, we will determine the binding constant K f for complex formation between 4'-6-diamidino-2-phenylindole
CHEM 311L Revision 1.2 A Determination of DNA-DAPI Binding using Fluorescence Spectroscopy In this Laboratory Exercise, we will determine the binding constant K f for complex formation between 4'-6-diamidino-2-phenylindole
Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition
 Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Bioengineering Laboratory I. Enzyme Assays. Part II: Determination of Kinetic Parameters Fall Semester
 Bioengineering Laboratory I Enzyme Assays Part II: Determination of Kinetic Parameters 2016-2017 Fall Semester 1. Theoretical background There are several mathematical models to determine the kinetic constants
Bioengineering Laboratory I Enzyme Assays Part II: Determination of Kinetic Parameters 2016-2017 Fall Semester 1. Theoretical background There are several mathematical models to determine the kinetic constants
Biology Chemistry & Physics of Biomolecules. Examination #1. Proteins Module. September 29, Answer Key
 Biology 5357 Chemistry & Physics of Biomolecules Examination #1 Proteins Module September 29, 2017 Answer Key Question 1 (A) (5 points) Structure (b) is more common, as it contains the shorter connection
Biology 5357 Chemistry & Physics of Biomolecules Examination #1 Proteins Module September 29, 2017 Answer Key Question 1 (A) (5 points) Structure (b) is more common, as it contains the shorter connection
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a
 Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for New Journal of Chemistry. This journal is The Royal Society of Chemistry and the Centre National de la Recherche Scientifique 2018 Electronic Supplementary Information
Electronic Supplementary Material (ESI) for New Journal of Chemistry. This journal is The Royal Society of Chemistry and the Centre National de la Recherche Scientifique 2018 Electronic Supplementary Information
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Supplementary Figure 1 Protein sequence alignment of Vibrionaceae with either a 40-residue insertion or a 44-residue insertion. Identical residues are indicated by red background.
SUPPLEMENTARY FIGURES Supplementary Figure 1 Protein sequence alignment of Vibrionaceae with either a 40-residue insertion or a 44-residue insertion. Identical residues are indicated by red background.
MicroCal itc 200. System MicroCal Auto-iTC 200. System. GE Healthcare Life Sciences. System design and description. provide:
 GE Healthcare Life Sciences Data file 28-97822 AC MicroCal label-free interaction analysis MicroCal itc 2 System MicroCal Auto-iTC 2 System MicroCal itc 2 and MicroCal Auto-iTC 2 isothermal titration calorimetry
GE Healthcare Life Sciences Data file 28-97822 AC MicroCal label-free interaction analysis MicroCal itc 2 System MicroCal Auto-iTC 2 System MicroCal itc 2 and MicroCal Auto-iTC 2 isothermal titration calorimetry
ITC Expert User s Manual
 ITC Expert User s Manual 1 Section 1: ITC Expert Background... 3 Minimal Heats and Injections... 3 C Parameter... 3 C Limitations... 4 High C... 4 Low C... 6 Concentrations Ratio... 6 Section 2: ITC Expert
ITC Expert User s Manual 1 Section 1: ITC Expert Background... 3 Minimal Heats and Injections... 3 C Parameter... 3 C Limitations... 4 High C... 4 Low C... 6 Concentrations Ratio... 6 Section 2: ITC Expert
A New Solvatochromic Fluorophore for Exploring Nonpolar Environments Created by Biopolymers
 Electronic Supplementary Information A New Solvatochromic Fluorophore for Exploring Nonpolar Environments Created by Biopolymers Abulfazl Fakhari M. and Steven E. Rokita Contribution from the Department
Electronic Supplementary Information A New Solvatochromic Fluorophore for Exploring Nonpolar Environments Created by Biopolymers Abulfazl Fakhari M. and Steven E. Rokita Contribution from the Department
Problem Set 5 Question 1
 2.32 Problem Set 5 Question As discussed in class, drug discovery often involves screening large libraries of small molecules to identify those that have favorable interactions with a certain druggable
2.32 Problem Set 5 Question As discussed in class, drug discovery often involves screening large libraries of small molecules to identify those that have favorable interactions with a certain druggable
Big Idea 1: Structure of Matter Learning Objective Check List
 Big Idea 1: Structure of Matter Learning Objective Check List Structure of Matter Mole Concept: Empirical Formula, Percent Composition, Stoichiometry Learning objective 1.1 The student can justify the
Big Idea 1: Structure of Matter Learning Objective Check List Structure of Matter Mole Concept: Empirical Formula, Percent Composition, Stoichiometry Learning objective 1.1 The student can justify the
Structural basis for catalytically restrictive dynamics of a high-energy enzyme state
 Supplementary Material Structural basis for catalytically restrictive dynamics of a high-energy enzyme state Michael Kovermann, Jörgen Ådén, Christin Grundström, A. Elisabeth Sauer-Eriksson, Uwe H. Sauer
Supplementary Material Structural basis for catalytically restrictive dynamics of a high-energy enzyme state Michael Kovermann, Jörgen Ådén, Christin Grundström, A. Elisabeth Sauer-Eriksson, Uwe H. Sauer
Supporting Information
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 214 Supporting Information Rapid and sensitive detection of acrylic acid using a novel fluorescence
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 214 Supporting Information Rapid and sensitive detection of acrylic acid using a novel fluorescence
Supplementary Figures
 1 Supplementary Figures Supplementary Figure 1 Type I FGFR1 inhibitors (a) Chemical structures of a pyrazolylaminopyrimidine inhibitor (henceforth referred to as PAPI; PDB-code of the FGFR1-PAPI complex:
1 Supplementary Figures Supplementary Figure 1 Type I FGFR1 inhibitors (a) Chemical structures of a pyrazolylaminopyrimidine inhibitor (henceforth referred to as PAPI; PDB-code of the FGFR1-PAPI complex:
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Supporting Information Kinetic Resolution of Constitutional Isomers Controlled by Selective Protection inside a Supramolecular Nanocapsule Simin Liu, Haiying Gan, Andrew T. Hermann,
SUPPLEMENTARY INFORMATION Supporting Information Kinetic Resolution of Constitutional Isomers Controlled by Selective Protection inside a Supramolecular Nanocapsule Simin Liu, Haiying Gan, Andrew T. Hermann,
Thermodynamic Stability of Hoogsteen and Watson-Crick. Base Pairs in the Presence of Histone H3-Mimicking Peptide
 Supporting information Thermodynamic Stability of Hoogsteen and Watson-Crick Base Pairs in the Presence of Histone H3-Mimicking Peptide Smritimoy Pramanik a, Kaori Nakamura a, Kenji Usui a,b, Shu-ichi
Supporting information Thermodynamic Stability of Hoogsteen and Watson-Crick Base Pairs in the Presence of Histone H3-Mimicking Peptide Smritimoy Pramanik a, Kaori Nakamura a, Kenji Usui a,b, Shu-ichi
Protein-Ligand Interactions: hydrodynamics and calorimetry
 Protein-Ligand Interactions: hydrodynamics and calorimetry Approach Stephen E. Harding Babur Z. Chowdhry OXFORD UNIVERSITY PRESS , New York Oxford University Press, 2001 978-0-19-963746-1 List of protocols
Protein-Ligand Interactions: hydrodynamics and calorimetry Approach Stephen E. Harding Babur Z. Chowdhry OXFORD UNIVERSITY PRESS , New York Oxford University Press, 2001 978-0-19-963746-1 List of protocols
shown by line A. The net effect of temperature on an enzyme-catalyzed reaction is given by line C.
 Experiment 8 EFFECT OF TEMPERATURE ON ENZYME ACTIVITY Temperature affects the stability of an enzyme as well as the binding of substrate and its transformation to product. Line B of Fig. 8-l shows the
Experiment 8 EFFECT OF TEMPERATURE ON ENZYME ACTIVITY Temperature affects the stability of an enzyme as well as the binding of substrate and its transformation to product. Line B of Fig. 8-l shows the
GeNei TM Enzyme Kinetics Teaching Kit Manual
 Teaching Kit Manual Cat No. New Cat No. KT89 106209 Revision No.: 00140806 CONTENTS Page No. Objective 3 Principle 3 Kit Description 4 Materials Provided 5 Procedure 6 Result 12 Interpretation 17 ORDERING
Teaching Kit Manual Cat No. New Cat No. KT89 106209 Revision No.: 00140806 CONTENTS Page No. Objective 3 Principle 3 Kit Description 4 Materials Provided 5 Procedure 6 Result 12 Interpretation 17 ORDERING
Supplementary Figure 1. Stability constants of metal monohydroxides. The log K values are summarized according to the atomic number of each element
 Supplementary Figure 1. Stability constants of metal monohydroxides. The log K values are summarized according to the atomic number of each element as determined in a previous study 1. The log K value
Supplementary Figure 1. Stability constants of metal monohydroxides. The log K values are summarized according to the atomic number of each element as determined in a previous study 1. The log K value
specified quantity of a solvent at a given temperature. To deconvolute the value from the
 S.1 Calculations of Dilution Enthalpy and Enthalpic Interaction Coefficients. When a solute is dissolved in a solvent a solution is formed. During dissolution of a solute in any solvent, heat is either
S.1 Calculations of Dilution Enthalpy and Enthalpic Interaction Coefficients. When a solute is dissolved in a solvent a solution is formed. During dissolution of a solute in any solvent, heat is either
Chapter 19 Chemical Thermodynamics Entropy and free energy
 Chapter 19 Chemical Thermodynamics Entropy and free energy Learning goals and key skills: Explain and apply the terms spontaneous process, reversible process, irreversible process, and isothermal process.
Chapter 19 Chemical Thermodynamics Entropy and free energy Learning goals and key skills: Explain and apply the terms spontaneous process, reversible process, irreversible process, and isothermal process.
National de la Recherche Scientifique and Université Paris Descartes, Paris, France.
 FAST-RESPONSE CALMODULIN-BASED FLUORESCENT INDICATORS REVEAL RAPID INTRACELLULAR CALCIUM DYNAMICS Nordine Helassa a, Xiao-hua Zhang b, Ianina Conte a,c, John Scaringi b, Elric Esposito d, Jonathan Bradley
FAST-RESPONSE CALMODULIN-BASED FLUORESCENT INDICATORS REVEAL RAPID INTRACELLULAR CALCIUM DYNAMICS Nordine Helassa a, Xiao-hua Zhang b, Ianina Conte a,c, John Scaringi b, Elric Esposito d, Jonathan Bradley
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/NCHEM.2633 Mechanically controlled radical polymerization initiated by ultrasound Hemakesh Mohapatra, Maya Kleiman, Aaron P. Esser-Kahn Contents 1. Materials and methods 2 2. Procedure for
DOI: 10.1038/NCHEM.2633 Mechanically controlled radical polymerization initiated by ultrasound Hemakesh Mohapatra, Maya Kleiman, Aaron P. Esser-Kahn Contents 1. Materials and methods 2 2. Procedure for
B L U E V A L L E Y D I S T R I C T C U R R I C U L U M Science AP Chemistry
 B L U E V A L L E Y D I S T R I C T C U R R I C U L U M Science AP Chemistry ORGANIZING THEME/TOPIC UNIT 1: ATOMIC STRUCTURE Atomic Theory Electron configuration Periodic Trends Big Idea 1: The chemical
B L U E V A L L E Y D I S T R I C T C U R R I C U L U M Science AP Chemistry ORGANIZING THEME/TOPIC UNIT 1: ATOMIC STRUCTURE Atomic Theory Electron configuration Periodic Trends Big Idea 1: The chemical
Dual Use of a Chemical Auxiliary: Molecularly Imprinted Polymers for the Selective Recovery of Products from Biocatalytic Reaction Mixtures
 SUPPORTING INFORMATION Dual Use of a Chemical Auxiliary: Molecularly Imprinted Polymers for the Selective Recovery of Products from Biocatalytic Reaction Mixtures Aaron T. Larsen, Tiffany Lai, Vanja Polic,
SUPPORTING INFORMATION Dual Use of a Chemical Auxiliary: Molecularly Imprinted Polymers for the Selective Recovery of Products from Biocatalytic Reaction Mixtures Aaron T. Larsen, Tiffany Lai, Vanja Polic,
Instantaneous and Quantitative Functionalization of Gold Nanoparticles with Thiolated DNA Using a ph-assisted and Surfactant-Free Route
 Supporting Information Instantaneous and Quantitative Functionalization of Gold Nanoparticles with Thiolated DNA Using a ph-assisted and Surfactant-Free Route Xu Zhang,, Mark R. Servos and Juewen Liu *
Supporting Information Instantaneous and Quantitative Functionalization of Gold Nanoparticles with Thiolated DNA Using a ph-assisted and Surfactant-Free Route Xu Zhang,, Mark R. Servos and Juewen Liu *
Concept review: Binding equilibria
 Concept review: Binding equilibria 1 Binding equilibria and association/dissociation constants 2 The binding of a protein to a ligand at equilibrium can be written as: P + L PL And so the equilibrium constant
Concept review: Binding equilibria 1 Binding equilibria and association/dissociation constants 2 The binding of a protein to a ligand at equilibrium can be written as: P + L PL And so the equilibrium constant
Ligand Binding A. Binding to a Single Site:
 A. Binding to a Single Site: The uilibrium constant (also known as association constant or affinity constant) for the binding of a ligand to a protein is described by the following uation (note: A ): [
A. Binding to a Single Site: The uilibrium constant (also known as association constant or affinity constant) for the binding of a ligand to a protein is described by the following uation (note: A ): [
Chirascan 6-Cell Peltier Cell Holder: Rapid Optimisation of Buffer Conditions for Stabilising Protein Therapeutics
 CHIRASCAN SERIES APPLICATION NOTE Chirascan 6-Cell Peltier Cell Holder: Rapid Optimisation of Buffer Conditions for Stabilising Protein Therapeutics Abstract: The selection of buffer conditions that maximise
CHIRASCAN SERIES APPLICATION NOTE Chirascan 6-Cell Peltier Cell Holder: Rapid Optimisation of Buffer Conditions for Stabilising Protein Therapeutics Abstract: The selection of buffer conditions that maximise
Supporting Information
 Supporting Information Materials and Methods: Tris-hydroxymethylaminomethane (Tris) was purchased from USB. All the other reagents used in the experiments were purchased from Sigma. All the DNA oligonucleotides
Supporting Information Materials and Methods: Tris-hydroxymethylaminomethane (Tris) was purchased from USB. All the other reagents used in the experiments were purchased from Sigma. All the DNA oligonucleotides
BIOCHEMISTRY. František Vácha. JKU, Linz.
 BIOCHEMISTRY František Vácha http://www.prf.jcu.cz/~vacha/ JKU, Linz Recommended reading: D.L. Nelson, M.M. Cox Lehninger Principles of Biochemistry D.J. Voet, J.G. Voet, C.W. Pratt Principles of Biochemistry
BIOCHEMISTRY František Vácha http://www.prf.jcu.cz/~vacha/ JKU, Linz Recommended reading: D.L. Nelson, M.M. Cox Lehninger Principles of Biochemistry D.J. Voet, J.G. Voet, C.W. Pratt Principles of Biochemistry
Supporting Information. Time-Resolved Botulinum Neurotoxin A Activity Monitored using. Peptide-Functionalized Au Nanoparticle Energy Transfer Sensors
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2014 Supporting Information Time-Resolved Botulinum Neurotoxin A Activity Monitored using Peptide-Functionalized
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2014 Supporting Information Time-Resolved Botulinum Neurotoxin A Activity Monitored using Peptide-Functionalized
Yves J. M. Bollen, Sanne M. Nabuurs, Willem J. H. van Berkel, and Carlo P. M. van Mierlo
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 280, No. 9, Issue of March 4, pp. 7836 7844, 2005 2005 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Last In, First Out
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 280, No. 9, Issue of March 4, pp. 7836 7844, 2005 2005 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Last In, First Out
Experimental procedures and characterization of the supramolecular complexes
 Supporting Information for Impact of cyclodextrins on the behavior of amphiphilic ligands in aqueous organometallic catalysis Hervé Bricout 1, Estelle Léonard 2, Christophe Len 2, David Landy 3, Frédéric
Supporting Information for Impact of cyclodextrins on the behavior of amphiphilic ligands in aqueous organometallic catalysis Hervé Bricout 1, Estelle Léonard 2, Christophe Len 2, David Landy 3, Frédéric
Awanish Kumar, Anjeeta Rani and Pannuru Venkatesu* Department of Chemistry, University of Delhi, Delhi
 Electronic Supplementary Material (ESI) for New Journal of Chemistry. This journal is The Royal Society of Chemistry and the Centre National de la Recherche Scientifique 2014 Supplimentary Informations
Electronic Supplementary Material (ESI) for New Journal of Chemistry. This journal is The Royal Society of Chemistry and the Centre National de la Recherche Scientifique 2014 Supplimentary Informations
Isothermal experiments characterize time-dependent aggregation and unfolding
 1 Energy Isothermal experiments characterize time-dependent aggregation and unfolding Technical ote Introduction Kinetic measurements have, for decades, given protein scientists insight into the mechanisms
1 Energy Isothermal experiments characterize time-dependent aggregation and unfolding Technical ote Introduction Kinetic measurements have, for decades, given protein scientists insight into the mechanisms
SUPPORTING INFORMATION
 SUPPORTING INFORMATION How Does Competition between Anionic Pollutants Affect Adsorption onto Mg-Al Layered Double Hydroxide? Three Competition Schemes Ganna DARMOGRAI, Benedicte PRELOT,* Amine GENESTE,
SUPPORTING INFORMATION How Does Competition between Anionic Pollutants Affect Adsorption onto Mg-Al Layered Double Hydroxide? Three Competition Schemes Ganna DARMOGRAI, Benedicte PRELOT,* Amine GENESTE,
Energetics of chitooligosaccharide binding to pumpkin (Cucurbita maxima) phloem exudate Lectin
 Chapter 3 Energetics of chitooligosaccharide binding to pumpkin (Cucurbita maxima) phloem exudate Lectin 50 Chapter 3 Energetics of 51 Summary Binding of chitooligosaccharides [(GlcNAc) 2-6 ] to pumpkin
Chapter 3 Energetics of chitooligosaccharide binding to pumpkin (Cucurbita maxima) phloem exudate Lectin 50 Chapter 3 Energetics of 51 Summary Binding of chitooligosaccharides [(GlcNAc) 2-6 ] to pumpkin
FLUORESCENCE STUDY ON TRYPTOPHAN POTASSIUM IODIDE INTERACTION
 PROCEEDINGS OF THE YEREVAN STATE UNIVERSITY C h e m i s t r y a n d B i o l o g y 08, 5(), p. 75 79 FLUORESCENCE STUDY ON TRYPTOPHAN POTASSIUM IODIDE INTERACTION C h emistr y H. A. SHILAJYAN, K. R. GRIGORYAN
PROCEEDINGS OF THE YEREVAN STATE UNIVERSITY C h e m i s t r y a n d B i o l o g y 08, 5(), p. 75 79 FLUORESCENCE STUDY ON TRYPTOPHAN POTASSIUM IODIDE INTERACTION C h emistr y H. A. SHILAJYAN, K. R. GRIGORYAN
Supramolecular Free Radicals: Near-infrared Organic Materials with Enhanced Photothermal Conversion. Supporting Information
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2015 Supramolecular Free Radicals: Near-infrared Organic Materials with Enhanced Photothermal
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2015 Supramolecular Free Radicals: Near-infrared Organic Materials with Enhanced Photothermal
Stopped-Flow Studies of the Kinetics of Single-Stranded DNA Binding and Wrapping around the Escherichia coli SSB Tetramer
 6032 Biochemistry 2002, 41, 6032-6044 Stopped-Flow Studies of the Kinetics of Single-Stranded DNA Binding and Wrapping around the Escherichia coli SSB Tetramer Alexander G. Kozlov and Timothy M. Lohman*
6032 Biochemistry 2002, 41, 6032-6044 Stopped-Flow Studies of the Kinetics of Single-Stranded DNA Binding and Wrapping around the Escherichia coli SSB Tetramer Alexander G. Kozlov and Timothy M. Lohman*
Data Sheet. Azide Cy5 RNA T7 Transcription Kit
 Cat. No. Size 1. Description PP-501-Cy5 10 reactions à 40 µl For in vitro use only Quality guaranteed for 12 months Store all components at -20 C. Avoid freeze and thaw cycles. DBCO-Sulfo-Cy5 must be stored
Cat. No. Size 1. Description PP-501-Cy5 10 reactions à 40 µl For in vitro use only Quality guaranteed for 12 months Store all components at -20 C. Avoid freeze and thaw cycles. DBCO-Sulfo-Cy5 must be stored
Supplementary Information for. Direct nitration and azidation of aliphatic carbons by an iron-dependent halogenase
 Supplementary Information for Direct nitration and azidation of aliphatic carbons by an iron-dependent halogenase Megan L Matthews, Wei-chen Chang, Andrew P Layne, Linde A Miles, Carsten Krebs, J Martin
Supplementary Information for Direct nitration and azidation of aliphatic carbons by an iron-dependent halogenase Megan L Matthews, Wei-chen Chang, Andrew P Layne, Linde A Miles, Carsten Krebs, J Martin
EXPERIMENT 14. ACID DISSOCIATION CONSTANT OF METHYL RED 1
 EXPERIMET 14. ACID DISSOCIATIO COSTAT OF METHYL RED 1 The acid dissociation constant, Ka, of a dye is determined using spectrophotometry. Introduction In aqueous solution, methyl red is a zwitterion and
EXPERIMET 14. ACID DISSOCIATIO COSTAT OF METHYL RED 1 The acid dissociation constant, Ka, of a dye is determined using spectrophotometry. Introduction In aqueous solution, methyl red is a zwitterion and
Redox-Responsive Complexation between a. Pillar[5]arene with Mono ethylene oxide Substituents. and Paraquat
![Redox-Responsive Complexation between a. Pillar[5]arene with Mono ethylene oxide Substituents. and Paraquat Redox-Responsive Complexation between a. Pillar[5]arene with Mono ethylene oxide Substituents. and Paraquat](/thumbs/93/111729438.jpg) Redox-Responsive Complexation between a Pillar[5]arene with Mono ethylene oxide Substituents and Paraquat Xiaodong Chi, Min Xue, Yong Yao and Feihe Huang* MOE Key Laboratory of Macromolecular Synthesis
Redox-Responsive Complexation between a Pillar[5]arene with Mono ethylene oxide Substituents and Paraquat Xiaodong Chi, Min Xue, Yong Yao and Feihe Huang* MOE Key Laboratory of Macromolecular Synthesis
Electronic Supplementary Information for:
 Electronic Supplementary Material (ESI) for Energy & Environmental Science. This journal is The Royal Society of Chemistry 216 Electronic Supplementary Information for: Nitrogenase bioelectrocatalysis:
Electronic Supplementary Material (ESI) for Energy & Environmental Science. This journal is The Royal Society of Chemistry 216 Electronic Supplementary Information for: Nitrogenase bioelectrocatalysis:
Supporting Information
 Supporting Information Self-Assembly of Glutathione S-transferases into Nanowires Wei Zhang, a Quan Luo,* a Lu Miao, a Yushi Bai, a Zeyuan Dong, a Jiayun Xu, a and Junqiu Liu* a a State Key Laboratory
Supporting Information Self-Assembly of Glutathione S-transferases into Nanowires Wei Zhang, a Quan Luo,* a Lu Miao, a Yushi Bai, a Zeyuan Dong, a Jiayun Xu, a and Junqiu Liu* a a State Key Laboratory
Precision and accuracy of protein size determination using the ActiPix TDA200 Nano-Sizing System
 Precision and accuracy of protein size determination using the ActiPix TDA200 Nano-Sizing System Keywords: Hydrodynamic radius, diffusion coefficient, sizing, Taylor dispersion analysis, protein, antibodies,
Precision and accuracy of protein size determination using the ActiPix TDA200 Nano-Sizing System Keywords: Hydrodynamic radius, diffusion coefficient, sizing, Taylor dispersion analysis, protein, antibodies,
Ratiometric and intensity-based zinc sensors built on rhodol and rhodamine platforms
 Supporting Information Ratiometric and intensity-based zinc sensors built on rhodol and rhodamine platforms Elisa Tomat and Stephen J. Lippard* Department of Chemistry, Massachusetts Institute of Technology,
Supporting Information Ratiometric and intensity-based zinc sensors built on rhodol and rhodamine platforms Elisa Tomat and Stephen J. Lippard* Department of Chemistry, Massachusetts Institute of Technology,
pyridoxal phosphate synthase
 Supplementary Information 13 C-NMR snapshots of the complex reaction coordinate of pyridoxal phosphate synthase Jeremiah W. Hanes, Ivan Keresztes, and Tadhg P. Begley * Department of Chemistry and Chemical
Supplementary Information 13 C-NMR snapshots of the complex reaction coordinate of pyridoxal phosphate synthase Jeremiah W. Hanes, Ivan Keresztes, and Tadhg P. Begley * Department of Chemistry and Chemical
A. One-Substrate Reactions (1) Kinetic concepts
 A. One-Substrate Reactions (1) Kinetic concepts (2) Kinetic analysis (a) Briggs-Haldane steady-state treatment (b) Michaelis constant (K m ) (c) Specificity constant (3) Graphical analysis (4) Practical
A. One-Substrate Reactions (1) Kinetic concepts (2) Kinetic analysis (a) Briggs-Haldane steady-state treatment (b) Michaelis constant (K m ) (c) Specificity constant (3) Graphical analysis (4) Practical
type GroEL-GroES complex. Crystals were grown in buffer D (100 mm HEPES, ph 7.5,
 Supplementary Material Supplementary Materials and Methods Structure Determination of SR1-GroES-ADP AlF x SR1-GroES-ADP AlF x was purified as described in Materials and Methods for the wild type GroEL-GroES
Supplementary Material Supplementary Materials and Methods Structure Determination of SR1-GroES-ADP AlF x SR1-GroES-ADP AlF x was purified as described in Materials and Methods for the wild type GroEL-GroES
Supporting Information
 Supporting Information Boehr et al. 10.1073/pnas.0914163107 SI Text Materials and Methods. R 2 relaxation dispersion experiments. 15 NR 2 relaxation dispersion data measured at 1 H Larmor frequencies of
Supporting Information Boehr et al. 10.1073/pnas.0914163107 SI Text Materials and Methods. R 2 relaxation dispersion experiments. 15 NR 2 relaxation dispersion data measured at 1 H Larmor frequencies of
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing
 Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi: 1.138/nchem.892 Mutual modulation Modulation between membrane-embedded Membrane Embedded receptor Receptors clustering lustering and ligand binding in lipid membranes Ligand
SUPPLEMENTARY INFORMATION doi: 1.138/nchem.892 Mutual modulation Modulation between membrane-embedded Membrane Embedded receptor Receptors clustering lustering and ligand binding in lipid membranes Ligand
Microcalorimetric techniques
 Microcalorimetric techniques Isothermal titration calorimetry (ITC) Differential scanning calorimetry (DSC) Filip Šupljika Filip.Supljika@irb.hr Laboratory for the study of interactions of biomacromolecules
Microcalorimetric techniques Isothermal titration calorimetry (ITC) Differential scanning calorimetry (DSC) Filip Šupljika Filip.Supljika@irb.hr Laboratory for the study of interactions of biomacromolecules
Low-volume, High Throughput Workflow for Analysis of Nucleic Acid Samples for Biobanking
 A p p l i c a t i o n N o t e Low-volume, High Throughput Workflow for Analysis of Nucleic Acid Samples for Biobanking Peter J. Brescia, Jr., Chris Wilson, and Peter Banks, BioTek Instruments, Inc., Winooski,
A p p l i c a t i o n N o t e Low-volume, High Throughput Workflow for Analysis of Nucleic Acid Samples for Biobanking Peter J. Brescia, Jr., Chris Wilson, and Peter Banks, BioTek Instruments, Inc., Winooski,
Macromolecular Interactions the equilibrium element
 Macromolecular Interactions the equilibrium element Physical Reality Quantitative P + L PL K d,overall K d,overall = [P][L] [PL] Driving force is difference in ground state free energies ΔG f o ΔG f o
Macromolecular Interactions the equilibrium element Physical Reality Quantitative P + L PL K d,overall K d,overall = [P][L] [PL] Driving force is difference in ground state free energies ΔG f o ΔG f o
Spanish Fork High School Unit Topics and I Can Statements AP Chemistry
 Spanish Fork High School 2014-15 Unit Topics and I Can Statements AP Chemistry Properties of Elements I can describe how mass spectroscopy works and use analysis of elements to calculate the atomic mass
Spanish Fork High School 2014-15 Unit Topics and I Can Statements AP Chemistry Properties of Elements I can describe how mass spectroscopy works and use analysis of elements to calculate the atomic mass
Isothermal Titration Calorimetry in Drug Discovery. Geoff Holdgate Structure & Biophysics, Discovery Sciences, AstraZeneca October 2017
 Isothermal Titration Calorimetry in Drug Discovery Geoff Holdgate Structure & Biophysics, Discovery Sciences, AstraZeneca October 217 Introduction Introduction to ITC Strengths / weaknesses & what is required
Isothermal Titration Calorimetry in Drug Discovery Geoff Holdgate Structure & Biophysics, Discovery Sciences, AstraZeneca October 217 Introduction Introduction to ITC Strengths / weaknesses & what is required
Supplementary information for. Observation of photovoltaic action from photoacid-modified Nafion due to light-driven ion transport
 Supplementary information for Observation of photovoltaic action from photoacid-modified Nafion due to light-driven ion transport William White, a# Christopher D. Sanborn, a# Ronald S. Reiter, a David
Supplementary information for Observation of photovoltaic action from photoacid-modified Nafion due to light-driven ion transport William White, a# Christopher D. Sanborn, a# Ronald S. Reiter, a David
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1.
 Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Supporting Text Z = 2Γ 2+ + Γ + Γ [1]
![Supporting Text Z = 2Γ 2+ + Γ + Γ [1] Supporting Text Z = 2Γ 2+ + Γ + Γ [1]](/thumbs/91/105230015.jpg) Supporting Text RNA folding experiments are typically carried out in a solution containing a mixture of monovalent and divalent ions, usually MgCl 2 and NaCl or KCl. All three species of ions, Mg, M +
Supporting Text RNA folding experiments are typically carried out in a solution containing a mixture of monovalent and divalent ions, usually MgCl 2 and NaCl or KCl. All three species of ions, Mg, M +
A BODIPY aldoxime-based chemodosimeter for highly selective and rapid detection of hypochlorous acid
 Supporting Information A BODIPY aldoxime-based chemodosimeter for highly selective and rapid detection of hypochlorous acid Mustafa Emrullahoğlu,* Muhammed Üçüncü and Erman Karakuş Department of Chemistry,
Supporting Information A BODIPY aldoxime-based chemodosimeter for highly selective and rapid detection of hypochlorous acid Mustafa Emrullahoğlu,* Muhammed Üçüncü and Erman Karakuş Department of Chemistry,
Chapter 13 Lecture Lecture Presentation. Chapter 13. Chemical Kinetics. Sherril Soman Grand Valley State University Pearson Education, Inc.
 Chapter 13 Lecture Lecture Presentation Chapter 13 Chemical Kinetics Sherril Soman Grand Valley State University Ectotherms Lizards, and other cold-blooded creatures, are ectotherms animals whose body
Chapter 13 Lecture Lecture Presentation Chapter 13 Chemical Kinetics Sherril Soman Grand Valley State University Ectotherms Lizards, and other cold-blooded creatures, are ectotherms animals whose body
Online Supplementary Material. Messenger RNA Interactions in the Decoding Center Control the Rate of Translocation
 Online Supplementary Material Messenger RNA Interactions in the Decoding Center Control the Rate of Translocation Prashant K. Khade and Simpson Joseph Supplementary Figure 1 Dissociation of the f[ 35 S]Met-Phe-tRNA
Online Supplementary Material Messenger RNA Interactions in the Decoding Center Control the Rate of Translocation Prashant K. Khade and Simpson Joseph Supplementary Figure 1 Dissociation of the f[ 35 S]Met-Phe-tRNA
UW Department of Chemistry Lab Lectures Online
 Lab 4: Effect of Temperature on Solubility and Fractional Crystallization Part I: Fractional Crystallization of Potassium Nitrate (KNO 3 ) Part II: Determining the Solubility Curve of Potassium Nitrate
Lab 4: Effect of Temperature on Solubility and Fractional Crystallization Part I: Fractional Crystallization of Potassium Nitrate (KNO 3 ) Part II: Determining the Solubility Curve of Potassium Nitrate
Chem 460 Laboratory Fall 2008 Experiment 3: Investigating Fumarase: ph Profile, Stereospecificity and Thermodynamics of Reaction
 1 Chem 460 Laboratory Fall 2008 Experiment 3: Investigating Fumarase: ph Profile, Stereospecificity and Thermodynamics of Reaction Before Lab Week 1 -- ph Profile for Fumarase Read Box 11-1 (page 323)
1 Chem 460 Laboratory Fall 2008 Experiment 3: Investigating Fumarase: ph Profile, Stereospecificity and Thermodynamics of Reaction Before Lab Week 1 -- ph Profile for Fumarase Read Box 11-1 (page 323)
A. K. R. Junker et al 1 Investigating subtle 4f vs. 5f...
 Electronic Supplementary Material (ESI) for Dalton Transactions. This journal is The Royal Society of Chemistry 2018 A. K. R. Junker et al 1 Investigating subtle 4f vs. 5f... Electronic Supplementary Information
Electronic Supplementary Material (ESI) for Dalton Transactions. This journal is The Royal Society of Chemistry 2018 A. K. R. Junker et al 1 Investigating subtle 4f vs. 5f... Electronic Supplementary Information
Multivalent interactions in human biology
 Cooperativity Multivalent interactions in human biology Multivalent interactions in supramolecular chemistry Additivity (?) Multivalent interactions in supramolecular chemistry In order to obtain a
Cooperativity Multivalent interactions in human biology Multivalent interactions in supramolecular chemistry Additivity (?) Multivalent interactions in supramolecular chemistry In order to obtain a
Supplementary materials. Crystal structure of the carboxyltransferase domain. of acetyl coenzyme A carboxylase. Department of Biological Sciences
 Supplementary materials Crystal structure of the carboxyltransferase domain of acetyl coenzyme A carboxylase Hailong Zhang, Zhiru Yang, 1 Yang Shen, 1 Liang Tong Department of Biological Sciences Columbia
Supplementary materials Crystal structure of the carboxyltransferase domain of acetyl coenzyme A carboxylase Hailong Zhang, Zhiru Yang, 1 Yang Shen, 1 Liang Tong Department of Biological Sciences Columbia
Thermodynamics of Borax Dissolution
 Thermodynamics of Borax Dissolution Introduction In this experiment, you will determine the values of H, G and S for the reaction which occurs when borax (sodium tetraborate octahydrate) dissolves in water.
Thermodynamics of Borax Dissolution Introduction In this experiment, you will determine the values of H, G and S for the reaction which occurs when borax (sodium tetraborate octahydrate) dissolves in water.
Effects of Temperature and Concentration on the Rate of Photo-bleaching of Erythrosine in Water
 Supporting Information for: Effects of Temperature and Concentration on the Rate of Photo-bleaching of Erythrosine in Water Joshua K. G. Karlsson, Owen J. Woodford, Roza Al-Aqar and Anthony Harriman* Molecular
Supporting Information for: Effects of Temperature and Concentration on the Rate of Photo-bleaching of Erythrosine in Water Joshua K. G. Karlsson, Owen J. Woodford, Roza Al-Aqar and Anthony Harriman* Molecular
Supporting Information for: Kinetic Mechanisms Governing Stable Ribonucleotide Incorporation in Individual DNA Polymerase Complexes
 Supporting Information for: Kinetic Mechanisms Governing Stable Ribonucleotide Incorporation in Individual DNA Polymerase Complexes Joseph M. Dahl, Hongyun Wang, José M. Lázaro, Margarita Salas, and Kate
Supporting Information for: Kinetic Mechanisms Governing Stable Ribonucleotide Incorporation in Individual DNA Polymerase Complexes Joseph M. Dahl, Hongyun Wang, José M. Lázaro, Margarita Salas, and Kate
The Effect of the Stoichiometry in the Supramolecular Chirality Transfer to Zinc Bisporphyrins Systems
 Supporting Information The Effect of the Stoichiometry in the Supramolecular Chirality Transfer to Zinc Bisporphyrins Systems Juan Etxebarria, Anton Vidal-Ferran* and Pablo Ballester* Catalan Institution
Supporting Information The Effect of the Stoichiometry in the Supramolecular Chirality Transfer to Zinc Bisporphyrins Systems Juan Etxebarria, Anton Vidal-Ferran* and Pablo Ballester* Catalan Institution
Previous Class. Reasons for analyzing pre-steady state conditions Methods for pre-steady state measurements. Today
 Previous Class Reasons for analyzing pre-steady state conditions Methods for pre-steady state measurements Today Spectrophotometry Spectrofluorimetry Radioactive Procedures ph dependency Spectrophotometry
Previous Class Reasons for analyzing pre-steady state conditions Methods for pre-steady state measurements Today Spectrophotometry Spectrofluorimetry Radioactive Procedures ph dependency Spectrophotometry
