Anion-π and π-π cooperative interactions
|
|
|
- Dominic Jennings
- 6 years ago
- Views:
Transcription
1 Chapter 5 Anion-π and π-π cooperative interactions 5.1 Introduction The design of selective receptors of anionic species is a very active area of research within supramolecular chemistry due to the potential applications to catalysis, separation processes, and biomolecular systems [87]. Common neutral receptors bind the anion by hydrogen bonding or coordinate the anion at the Lewis acidic center of an organometallic ligand. As compared to cationic hosts, neutral receptors avoid the presence of competing negative counterions and are characterized by higher selectivity due to the directionality of the interactions. In recent years, the alternative route of anion complexation by neutral hosts via anion-π interactions has received considerable interest. The favorable binding interactions between an anionic species and an electron-deficient π-electron compound has been demonstrated in several theoretical studies [88 104] and experimental evidence of these attractive interactions is now cumulating from both X-ray structures [ ] and solution data [114, 115]. Aromatic systems which have been investigated as potential anion-host candidates are either substituted benzene or electron-poor heteroaromatics such as triazines. Even though π-systems are expected to interact repulsively with anions, the presence of electron-withdrawing substituting atoms in the aromatic compounds modulates the reactivity inverting their natural electron-donor character. Hexafluorobenzene is the extreme example with a permanent quadrupole moment of similar magnitude as benzene but opposite sign, leading to attractive electrostatic interactions with electron-donor species [116]. In general, the interactions between the anion and the π-system 117
2 5.1. Introduction 5. Anion-π and π-π cooperative interactions are predominantly of electrostatic and polarization nature, but dispersion forces and charge transfer [93,94,102] also contribute to the stability of these complexes. Recently, experimental evidence of anion-π-π interactions has emerged from crystallographic studies [ ] on synthesized coordination compounds based on the electron-deficient 1,3,5-triazine moieties, suggesting the possibility to enhance anion-π binding by π-π stacking. Particularly intriguing are the structural features of the nitrate-triazine-triazine complex of Ref. [109], as the two aromatic rings are staggered and not perfectly faced, and the nitrate ion is not parallel to the closest ring. The unusual asymmetrical configuration of the closely stacked triazines could be induced by the particular coordination within the compound, or governed by a subtle interplay between anion-π and π-π interactions [101, 104]. In the original paper, the compound was also investigated theoretically but, due to the poor description of dispersive interactions by the standard density functional theory approach employed, no conclusions could be drawn on the stabilization effect of π-π interactions on the whole complex. In the present theoretical study, we investigate and rationalize the structural features of this anion-π-π complex, and quantitatively address the issue of cooperativity of anion-π and π-π interactions using a combination of dispersion-corrected density functional theory (DFT) and quantum Monte Carlo (QMC) calculations. The calculated structure is remarkably close to the one observed experimentally even though the anion-π-π complex was not additionally coordinated as in the crystal structure. Therefore, the unusual stacking is an intrinsic feature which stabilizes the anion-π binding, indicating that the principle of anion-π-π cooperativity is regulating the self-assembly in this coordination compound. Energetically, the cooperative effect of anion-π and π-π interactions in the triazine-triazine-nitrate complex is not negligible but amounts to roughly 6% of the total binding energy. We want to emphasize that the theoretical investigation of anion-π-π interactions is particularly demanding. In anion-π systems, correlation significantly contributes to the interaction energy [93, 94] and must therefore be accurately treated. In the presence of aromatic stacking, the need to also address π-π interactions further complicates matters. Finally, even though most previous studies of anion-π systems within the MP2 approach were limited to highly symmetrical configurations, it is important to be able to explore lower-symmetry complexes for a realistic representation of the anionπ-π systems observed experimentally. Therefore, we choose here to employ the efficient DFT approach in combination with the recently proposed dispersion-corrected atom-centered pseudopotentials (DCACPs) [117, 118], which we validate against accurate highly-correlated quantum Monte Carlo 118
3 5. Anion-π and π-π cooperative interactions5.2. Computational approaches calculations. The DFT-DCACP method is found to reliably predict equilibrium structures as well as the relative stability of different complexes, and is therefore a very promising tool for the investigation of even larger anion-π systems. 5.2 Computational approaches We briefly review below the two theoretical methods employed in this work, that is, the recently developed semi-empirical DFT scheme augmented with dispersion-corrected atom-centered potentials (DCACPs) and the quantum Monte Carlo (QMC) approach. We also give all relevant computational details Semi-empirical dispersion corrected DFT A simple semi-empirical approach has been recently proposed to correct the deficiency of approximate density functionals in describing London dispersion forces [117, 118]. The non-local electron-nucleus pseudopotentials used in the Kohn-Sham DFT scheme are augmented with DCACPs whose parameters are fitted against references data obtained in ab-initio highly-correlated approaches. By construction, these potentials do not affect valence electronic properties but appear to significantly improve the description within DFT of weakly bound systems at no additional computational cost [119, 120]. While the original DCACPs were calibrated against MP2 reference properties, we use here the latest library of potentials constructed from more accurate coupled-cluster singles and doubles with a perturbative treatment of the triples [CCSD(T)] and configuration interaction data [121]. We employ the generalized gradient approximation (GGA) functional of Becke, Lee, Yang and Parr (BLYP) [16] and the corresponding DCACPs non-local potentials given in the Troullier-Martins [122] form. We use the DCACPs for all the atomic species except for F where we use the original Troullier-Martins BLYP pseudopotential as the corresponding DCACP is not yet available. We expect that the absence of dispersion corrections for F is not important in the complexes studied in Section as the F-F interaction is dominated by electrostatic repulsion. All the DFT calculations are performed using the plane-wave basis set program CPMD [46] with a plane-wave cutoff of 80 Ry. We employ the isolated system module in CPMD which allows studying an isolated molecule or complex within periodic boundary conditions. The Poisson equations are solved with the Hockney method [123]. We use a box size of Å 3 which is sufficiently large for all the complexes 119
4 5.2. Computational approaches5. Anion-π and π-π cooperative interactions considered in this work. All the geometry optimizations are performed without imposing any symmetry constraints and with a threshold for the residual force of a.u. The binding energies of complexes are computed by subtracting the energies of the optimized fragments from the total energy of the complex. We note that all computed binding energies do not include zero point energy corrections Quantum Monte Carlo methods QMC methods [124,125] offer an efficient alternative to conventional highlycorrelated ab-initio methods as they can be applied to sufficiently large systems and still provide an accurate description of both dynamical and static electronic correlation. The key ingredient which determines the quality of a QMC calculation is the many-body trial wave function which, in the present work, is chosen of the Jastrow-Slater type with the particular form, Ψ=D D A,i,j J (r ij,r ia,r ja ), (5.1) where D and D are Slater determinants of single-particle orbitals for the up- and down-spin electrons, respectively, and the orbitals are represented using atomic Gaussian basis. The Jastrow correlation factor J depends on the distance r ij between electrons i and j, and on the distance r ia and r ja of electrons i and j from nucleus A. The Jastrow factor is here expressed as the exponential of the sum of three fifth-order polynomials of electronnuclear, of electron-electron, and of pure three-body mixed electron-electron and electron-nucleus distances, respectively [126]. Different Jastrow factors are used to describe the correlation with different atom types. In variational Monte Carlo (VMC), the square of the wave function is sampled using the Metropolis algorithm and the expectation value of the Hamiltonian on the wave function is computed by statistically averaging over a large number of electronic configurations sampled from Ψ 2.Thewave function is then used in diffusion Monte Carlo (DMC), which produces the best energy within the fixed-node approximation, i.e. the lowest-energy state with the same zeros (nodes) as the trial wave function Ψ. All QMC results presented below are from DMC calculations. All QMC calculations are performed with the program package CHAMP [127]. We employ scalar-relativistic energy-consistent Hartree-Fock pseudopotentials [67] for all the elements, and the hydrogen potential is softened by removing the Coulomb divergence. To represent the orbitals in the determinantal component, we employ the Gaussian basis sets [67] constructed 120
5 5. Anion-π and π-π cooperative interactions 5.3. Results for these pseudopotentials and augment them with diffuse functions. All calculations are performed with the cc-pvdz basis augmented with two additional diffuse s and p functions with exponents and for carbon, and for nitrogen, and for oxygen, and and for chlorine [68]. Only in the computation of the binding energy of the far-triazine-no 3 fragment (see Table 5.3), the basis is further augmented with two diffuse s and p functions with exponents and for carbon, and for nitrogen, and and for oxygen [68]. The use of these additional diffuse functions allows a stable QMC simulation in this compound where the triazine and NO 3 molecules are at very large distances (about 7 Å). Further augmentation of the basis with two diffuse d functions for all heavier atoms does not change the binding energy of the compound. The parameters in the Jastrow factor are always optimized within VMC by energy minimization [66] and, when stated, the coefficients of the orbital expansions over the atomic Gaussian basis are simultaneously optimized with the Jastrow component. Otherwise, orbitals from a B3LYP density functional theory [16, 19] calculation are employed, which are obtained using the same pseudopotentials and basis set with the program GAMESS(US) [57]. An imaginary time step of a.u. is used in the DMC calculations. As side test, we compute the DMC binding energy and equilibrium distance of the prototypical triazine-chloride complex with the ion along the C 3 axes of the ring, which has been the subject of several MP2 studies [88, 92, 93,95,96,99]. We perform a correlated sampling run [128] using as reference the MP2/aug-cc-pVDZ geometry with a chloride-centroid distance of 3.13 Å [93], and the corresponding fully optimized QMC wave function. We find a DMC equilibrium distance of 3.24 Å, which is in the range of the MP2 values obtained with different basis sets [93]. The DMC binding energy of 6.0(3) kcal/mol is slightly smaller than the MP2/aug-cc-pVDZ value of 6.93 kcal/mol [93]. We note that using optimized or B3LYP orbitals in the determinantal component of the wave functions yields statistically equivalent results even though the B3LYP approach underestimates the binding energy by about 2 kcal/mol [95]. 5.3 Results To investigate cooperative effects of anion-π and π-π interactions in the unusual triazine-triazine-nitrate complex observed experimentally, we first need to characterize how the anion-π and π-π fragments are separately stabilized. Studying these smaller components also allows us to access the performance 121
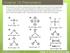
6 5.3. Results 5. Anion-π and π-π cooperative interactions of QMC and in particular of the semi-empirical DCACP approach, by comparing to MP2 or CCSD(T) calculations when available Triazine and NO 3 Figure 5.1: Side (A) and top (B) view of the triazine-nitrate complex in the parallel geometry (left) and in the T-like form (right). The triazine-no 3 complex represents one of the first examples of anionπ interactions observed experimentally [109, 110], and has already been the subject of several computational studies both at the DFT and MP2 level [93, 109,110]. Thus, it is an ideal system to assess the accuracy and transferability 122
7 5. Anion-π and π-π cooperative interactions 5.3. Results of the DCACPs in describing this novel weak interactions as all the atomic elements in the complex are available in the current library of dispersion corrected pseudopotentials [121]. Table 5.1: DFT/BLYP-DCACP and DMC binding energies in kcal/mol of the triazine-no 3 complex. The parallel and T-like geometries corresponding to the MP2 and DFT/BLYP-DCACP equilibrium distances are shown. The MP2 binding energies are also given when available. R 0 and R 0 are the equilibrium distances in Å between the ring centroid and the nitrogen atom of NO 3, and between the centroid and the closest oxygen atom of NO 3, respectively. The statistical error on the last figure of the DMC binding energy is given in parenthesis. Geometry R 0 R 0 DFT DMC MP2 DFT parallel (3) MP2 parallel 2.90 a -4.9(3) c -6.8 a DFT T-like (3) MP2 T-like 2.75 b -8.4 b a Ref. [93] MP2/aug-cc-pVDZ; b Ref. [110] MP2/ G(3df,p). c Wave function fully optimized with VMC [129]. In Table 5.1, we show the binding energy and the equilibrium distance between the ring centroid and the nitrogen of NO 3, calculated with different approaches. We focus first on the symmetrical geometry where the plane of the nitrate is parallel to the triazine plane with the nitrogen of the anion located on top of the ring centroid and the oxygens facing the carbon atoms (Fig. 5.1). We find that DFT/BLYP-DCACP gives a binding energy which is very close to the one obtained within the MP2 approach [93] but with a slightly larger (about 10%) equilibrium distance. DMC gives a binding energy 1.7 kcal/mol smaller than the DFT and MP2 values. A DMC equilibrium distance of about 2.9 Å is estimated in a correlated sampling run using as reference geometry the MP2 geometry and the corresponding fully optimized QMC wave function [128]. In Fig. 5.2, we show the DFT binding energies obtained with the standard BLYP and the BLYP-DCACP pseudopotentials for the parallel geometry at different distances between the planes. The structure of the fragments is kept fixed at the optimal geometry of the complex while changing the triazine-nitrate distance. The computed BLYP binding energy of 2.4 kcal/mol is in line with the value previously obtained with a Slater-type orbital ET-pVQZ basis [109]. It clearly appears that the dispersion corrections have a very large effect on the description of the anion-π interactions. With the inclusion of the DCACP pseudopoten- 123
8 5.3. Results 5. Anion-π and π-π cooperative interactions tials, the binding energy increases from 2.4 kcal/mol to 6.7 kcal/mol and the equilibrium distance decreases by about 6%. The triazine-nitrate complexes observed experimentally [109, 110] show however a very different structure from the more intuitive face-to-face configuration. This observation prompted further calculations both at the DFT and MP2 level [109, 110], which yield an energetically lower T-like arrangement of the triazine-nitrate complex. As shown in Table 5.1, the structure and the binding energy of the T-like complex computed within DFT/BLYP- DCACP compare well with the MP2 values [110]. In particular, the dispersion corrected potentials predict this configuration to be more stable than the parallel arrangement by about 2 kcal/mol. Similarly to the MP2 results, we find that the two oxygens of the nitrate point towards the triazine plane, one facing the ring centroid and the other facing one hydrogen atom at a distance d O...H =2.52Å (see also Fig. 5.1). The interaction with the hydrogen appears to further stabilize this geometrical arrangement. The shortest oxygen-centroid distance R 0 predicted by DFT/BLYP-DCACP is again about 10% larger than the MP2 value. The DMC calculations performed using the DFT optimized complexes give the same trend for the binding energy with the T-like geometry being the most stable. DMC results however indicate that both DFT and MP2 tend to overestimate the binding energy by 2 kcal/mol The triazine dimer The DCACPs have been developed with the main aim of improving the generally poor description of π-π interactions within DFT. The triazine dimer makes no exception in this respect, with the BLYP functional giving an unbound state. At the contrary, ab-initio correlated methods such as MP2 and CCSD(T) find a bound state with the optimal conformation being a stacked complex with a 60 relative orientation and a vertical distance of 3.4 Å [130, 131]. As shown in Table 5.2, the MP2 method tends to overestimates the binding energy in comparison with the more accurate CCSD(T) approach, a feature which appears to be general in the MP2 description of van der Waals complexes [ ]. In Table 5.2, we show the results for the triazine dimer obtained with the DFT/BLYP-DCACP approach. We consider the face-to-face conformation with relative orientations, 0,30,and60, between the two triazine molecules (Fig. 5.3). All structures are bound with the binding energy increasing as we move from 0 to 60. The DFT-DCACP global minimum at 60 is consistent with the MP2 finding, but has a slightly smaller binding energy and a centroid-to-centroid distance 6% larger. DMC calculations per- 124
9 5. Anion-π and π-π cooperative interactions 5.3. Results Binding energy (kcal/mol) Triazine-NO 3 - DFT/BLYP-DCACPs DFT/BLYP Distance (Å) Figure 5.2: Binding energy of the triazine-no 3 system in the parallel geometry as a function of the distance between the centroid of the triazine ring and the nitrogen atom of NO 3. The curves are computed within DFT/BLYP with and without the DCACPs. formed on the DFT optimized geometries display the same trend as DFT but give a systematically smaller binding energy by about 1 kcal/mol. We note that, at the MP2 geometry, the DMC binding energy is equal to the CCSD(T) prediction. Once more, these results indicate that the BLYP- DCACPs are able to predict the proper trend and global minimum of the weakly-bound triazine dimer although they slightly overestimate the binding energy similarly to the MP2 approach. In the crystalline structure [109] containing the triazine-triazine-nitrate complex, an unusual triazine-triazine moiety is observed: The planes of the triazine molecules are slightly off-centered with a centroid-to-centroid distance of 3.45 Å, and have a relative orientation of about 30 with a dihedral angle of about 15. If we optimize the triazine dimer within DFT-BLYP- DCACP starting from this experimental conformation, we find a local mini- 125
10 5.3. Results 5. Anion-π and π-π cooperative interactions Figure 5.3: Side (A) and top (B) view of four different conformations of the triazine-triazine dimer. From left to right, the two molecules are rotated with respect to each other by 0,30,and60. The bended geometry (right) is rotated by roughly 30. mum very similar to the experiment, that is, a distance between the centroids of 3.79 Å and a dihedral angle between the two rings of 11 (see Fig. 5.3). As shown in Table 5.2, the predicted binding energy for this local minimum is only 0.6 kcal/mol lower than the global minimum. However, the bending slightly stabilizes the dimer with respect to the parallel conformation at the same angle of 30. For the bended conformation, DMC gives the same energy within statistical error as for the DFT global minimum geometry Cooperativity of anion-π-π interactions We now focus on the experimental triazine-triazine-nitrate complex observed in Ref. [109] to understand the unusual structural features and quantify the stabilization induced on the anion-π system by π-π stacking. The relevant anion-π-π unit of the crystalline structure represents the starting point for our calculations and is shown in Fig. 5.4 together with the DFT/BLYP-DCACP optimal geometry. The similarity between the experimental and theoretical complexes is remarkable: The two triazine are staggered by about 30 and bended, and slip with respect to each other with the nitrate bound in a T-like configuration. Moreover, the nitrogen of the anion is displaced with respect to the normal through the centroid of the ring, consistently with the experiment. More specifically, we find a centroid-to-centroid distance of 3.74 Å and a distance from the centroid to the N of NO 3 of 3.50 Å as compared to the experimental values of 3.45 Å and 3.71 Å, respectively. The two aro- 126
11 5. Anion-π and π-π cooperative interactions 5.3. Results Table 5.2: DFT/BLYP-DCACP and DMC binding energies in kcal/mol of different conformations of the triazine-triazine dimer. The equilibrium distance R 0 in Å is between the centroids of the two rings and the angle indicates their relative rotation. Geometry R 0 DFT DMC MP2 CCSD(T) DFT (3) DFT (3) DFT (3) MP2 a (3) -3.8 a -2.8 a DFT 30 bended (3) DFT 60 bended a Ref. [130, 131] with a diffuse cc-pvdz basis set. matic rings form a dihedral angle of 22, close to the experimental value of 18. Finally, two oxygen atoms of the nitrate are interacting with the triazine ring, similarly to the observed structure. A more detailed comparison with the experimental structure is not appropriate as the experimental complex is embedded in the crystalline environment which may induce further distortions. We note that, with respect to the geometries of the subunits separately optimized, the presence of the nitrate does not significantly change the centroid-centroid distance but yields a larger tilt angle between the rings. As for triazine-nitrate system, the main geometrical effect is that the nitrate is more parallel and closer to the ring with a centroid-oxygen distance of 2.98 Å. The total computed binding energy of this anion-π-π complex is kcal/mol. To establish the stabilization effect induced by π-π stacking on this supramolecular complex, we separately compute the π-π and anion-π contributions to the binding energy. We extract from the optimized structure the three fragments corresponding to the triazine dimer and the two possible triazine-nitrate units, and compute their binding energies which are listed in Table 5.3. By subtracting these contributions from the total binding energy of the anion-π-π system, we derive a cooperative contribution to the binding energy of 0.8 kcal/mol, which corresponds to a stabilization enhancement of about 6%. The cooperative energy predicted by DMC using the DFT geometries is compatible with the DFT value within two standard deviations, while the DMC binding energies of the individual fragments are always lower as already pointed out in the previous sections. In particular, the compound given by the nitrate and the distant triazine ring is unbound within DMC due to geometrical deformations of the molecules within the triazine-triazinenitrate complex. 127
12 5.3. Results 5. Anion-π and π-π cooperative interactions Figure 5.4: Side (A) and top (B) view of the triazine-triazine-no 3 complex as obtained within DFT/BLYP-DCACP (left) and experimentally [109] (right). In the top view, the bottom ring is parallel to the page. We note that the triazine rings are not in the optimal staggered configuration of 60 (see Table 5.2) but have a relative orientation of about 30.This is certainly due to the geometrical constraints within the crystalline frame- 128
13 5. Anion-π and π-π cooperative interactions 5.3. Results work as the triazine rings are strongly coordinated via a complex network to the metal centers of the compound. To generate a triazine-triazine-nitrate complex arrangement consistent with the optimal geometries for the separate fragments, we thus start from a structure in which the triazine rings are parallel in the optimal 60 orientation, with the nitrate in the T-like form and two oxygens interacting with the ring (Fig. 5.6). The geometry optimization shows that the relative orientation remains at 60, and the centroid-centroid distance is unchanged with respect to the isolated dimer. However, the presence of the anion induces a tilt of about 14 of the triazine close to it, which is likely induced by the interaction between one oxygen of the nitrate and one hydrogen of the ring. As expected, the binding energy of this complex is slightly larger than in the structure derived from the experiment (see Table 5.3) and the cooperative effect is of comparable magnitude. Table 5.3: DFT/BLYP-DCACP and DMC binding energies in kcal/mol of five π-π complexes interacting with NO 3, i.e. two triazine (TAZ) rings at 30, two TAZ rings at 60, two trifluorotriazine (TFT) at 60,andthetwo mixed systems with one TAZ and one TFT ring at 60. The mixed systems denoted TAZ-TFT and the TFT-TAZ have the TFT and TAZ ring closer to the NO 3, respectively. The geometry of the ternary compound (ring-ring-no 3 ) is optimized within DFT/BLYP-DCACP and the geometries of the ring-ring, close-ring-no 3, and far-ring-no 3 fragments are here kept fixed when computing their binding energies. The binding energies are obtained as the difference between the total energy and the energies of the isolated ring and NO 3 optimized separately. For example, the binding energy of the ternary compound is given by E b (ring-ring-no 3 )=E(ring-ring- NO 3 ) 2 E(ring) E(NO 3 ). The cooperative energy is defined as E b (ringring-no 3 ) E b(close-ring-no 3 ) E b(far-ring-no 3 ) E b(ring-ring). Compound TAZ 30 TAZ 60 TFT TAZ-TFT TFT-TAZ DFT DMC DFT DFT DFT DFT Ring-ring-NO (2) Ring-ring (2) Close-ring-NO (2) Far-ring-NO (2) Cooperativity (4) Finally, we check the effect of enhancing the π-anion interaction by substituting the hydrogens in the triazine rings at 60 with the strongly electronegative fluorine [88,92,96]. With this procedure, we generate three complexes, i.e. (1) two trifluorotriazine rings with a nitrate, (2) one triazine and one tri- 129
14 5.3. Results 5. Anion-π and π-π cooperative interactions Figure 5.5: Side (A) and top (B) view of four different π-π complexes interacting with NO 3. From left to right, we show two triazine (TAZ) rings at 60, two trifluorotriazine (TFT) at 60, and the two mixed systems, TFT- TAZ and the TAZ-TFT. The TFT-TAZ and TAZ-TFT complexes have the TFT and TAZ ring closer to the NO 3, respectively. fluorotriazine coordinated with the nitrate, and (3) one trifluorotriazine and one triazine coordinated with the nitrate. The three models are also shown in Fig As seen in Table 5.3, the total binding is increased by the fluorine substitution due to the stronger attractive interaction between the nitrate and the trifluorotriazine ring(s). However, this increase does not always correlate with a cooperative enhancement of the π-anion interaction by π-π stacking. In particular, in the complex with two trifluorotriazine rings, the cooperative effect is smaller than in the original system with two triazine rings as the π-π system is now very weakly bound: The binding energy of the ring-ring fragment is only 0.2 kcal/mol, and significantly reduced from the 2.0 kcal/mol of the trifluorotriazine dimer optimized in the absence of the nitrate. This small binding can be explained with the strong deformation of the trifluorotriazine ring close to the nitrate, with one of the fluorine s bending out of the ring plane away from one of the nitrate oxygens. In the mixed triazine-trifluorotriazine compounds, we observe instead a more favorable balance of π-π and anion-π interactions, leading to an increased cooperativity. In both complexes, the ring-ring fragment is still significantly bound even though its binding energy is smaller than the optimal 130
15 5. Anion-π and π-π cooperative interactions 5.3. Results value of 3.2 kcal/mol for the isolated dimer. The ring-ring binding energy of the complex with the triazine coordinated to the nitrate is larger by about 1 kcal/mol than the energy obtained in the case of the trifluorotriazine coordinated to the nitrate. The reduced binding of the latter is due to a strong deformation of the trifluorotriazine ring vicinal to the nitrate, similarly to what observed in the compound with two trifluorotriazine rings. Correspondingly, the largest cooperative effect is observed in the mixed complex with the triazine coordinated to the nitrate where the non-additive contribution amounts to roughly 10% of the total binding energy. Finally, we note that the cooperative effect is always present in all the studied complexes while this does not appear to be a general feature [104] of anion-π-π complexes, and points to the very versatile nature of the triazine moiety in supramolecular chemistry [133]. Figure 5.6: Triazine-triazine-NO 3 complex with relative orientation of the rings of 60 optimized within DFT/BLYP-DCACP (grey). In blue, we show the starting symmetrical geometry. 131
16 5.4. Conclusions 5. Anion-π and π-π cooperative interactions 5.4 Conclusions Using state-of-the-art dispersion corrected DFT and QMC calculations, we have investigated the geometrical and energetic effects induced by π-π stacking on the anion-π system of the unusual triazine-triazine-nitrate complex recently observed experimentally. We have reproduced and rationalized the highly asymmetrical features of the structure, which are not imposed by the coordination of the anion-π-π subunit within the particular synthesized compound. We show that the two triazines are staggered and bended, and slip with respect to each other with the nitrate bound off-center in a T-like configuration. The stabilization induced by π-π stacking amounts energetically to about 6% of the total binding energy. An increased cooperative effect of 10% is obtained if the hydrogens in one of the triazine rings are substituted with the strongly electronegative fluorine atoms and the dimer is further staggered in a 60 orientation. We want to emphasize that the theoretical investigation of a realistic anion-π-π system as the one treated in this paper is particularly demanding as correlation plays an important role, while the system is not small and must be treated without symmetry constraints. We find that the use of the recently proposed dispersion corrected DFT approach represents a good compromise between accuracy and the ability to study complex and realistic systems involving weak interactions. 132
QMC dissociation energy of the water dimer: Time step errors and backflow calculations
 QMC dissociation energy of the water dimer: Time step errors and backflow calculations Idoia G. de Gurtubay and Richard J. Needs TCM group. Cavendish Laboratory University of Cambridge Idoia G. de Gurtubay.
QMC dissociation energy of the water dimer: Time step errors and backflow calculations Idoia G. de Gurtubay and Richard J. Needs TCM group. Cavendish Laboratory University of Cambridge Idoia G. de Gurtubay.
Quantum Monte Carlo backflow calculations of benzene dimers
 Quantum Monte Carlo backflow calculations of benzene dimers Kathleen Schwarz*, Cyrus Umrigar**, Richard Hennig*** *Cornell University Department of Chemistry, **Cornell University Department of Physics,
Quantum Monte Carlo backflow calculations of benzene dimers Kathleen Schwarz*, Cyrus Umrigar**, Richard Hennig*** *Cornell University Department of Chemistry, **Cornell University Department of Physics,
DFT calculations of NMR indirect spin spin coupling constants
 DFT calculations of NMR indirect spin spin coupling constants Dalton program system Program capabilities Density functional theory Kohn Sham theory LDA, GGA and hybrid theories Indirect NMR spin spin coupling
DFT calculations of NMR indirect spin spin coupling constants Dalton program system Program capabilities Density functional theory Kohn Sham theory LDA, GGA and hybrid theories Indirect NMR spin spin coupling
Oslo node. Highly accurate calculations benchmarking and extrapolations
 Oslo node Highly accurate calculations benchmarking and extrapolations Torgeir Ruden, with A. Halkier, P. Jørgensen, J. Olsen, W. Klopper, J. Gauss, P. Taylor Explicitly correlated methods Pål Dahle, collaboration
Oslo node Highly accurate calculations benchmarking and extrapolations Torgeir Ruden, with A. Halkier, P. Jørgensen, J. Olsen, W. Klopper, J. Gauss, P. Taylor Explicitly correlated methods Pål Dahle, collaboration
Paper No. 1: ORGANIC CHEMISTRY- I (Nature of Bonding and Stereochemistry)
 Subject Chemistry Paper No and Title Paper 1: ORGANIC - I (Nature of Bonding Module No and Title Module Tag CHE_P1_M10 TABLE OF CONTENTS 1. Learning Outcomes 2. Introduction 3. Non-Covalent Interactions
Subject Chemistry Paper No and Title Paper 1: ORGANIC - I (Nature of Bonding Module No and Title Module Tag CHE_P1_M10 TABLE OF CONTENTS 1. Learning Outcomes 2. Introduction 3. Non-Covalent Interactions
Computational Chemistry. An Introduction to Molecular Dynamic Simulations
 Computational Chemistry An Introduction to Molecular Dynamic Simulations Computational chemistry simulates chemical structures and reactions numerically, based in full or in part on the fundamental laws
Computational Chemistry An Introduction to Molecular Dynamic Simulations Computational chemistry simulates chemical structures and reactions numerically, based in full or in part on the fundamental laws
Electronic structure theory: Fundamentals to frontiers. 2. Density functional theory
 Electronic structure theory: Fundamentals to frontiers. 2. Density functional theory MARTIN HEAD-GORDON, Department of Chemistry, University of California, and Chemical Sciences Division, Lawrence Berkeley
Electronic structure theory: Fundamentals to frontiers. 2. Density functional theory MARTIN HEAD-GORDON, Department of Chemistry, University of California, and Chemical Sciences Division, Lawrence Berkeley
Quantum Chemical Simulations and Descriptors. Dr. Antonio Chana, Dr. Mosè Casalegno
 Quantum Chemical Simulations and Descriptors Dr. Antonio Chana, Dr. Mosè Casalegno Classical Mechanics: basics It models real-world objects as point particles, objects with negligible size. The motion
Quantum Chemical Simulations and Descriptors Dr. Antonio Chana, Dr. Mosè Casalegno Classical Mechanics: basics It models real-world objects as point particles, objects with negligible size. The motion
Example questions for Molecular modelling (Level 4) Dr. Adrian Mulholland
 Example questions for Molecular modelling (Level 4) Dr. Adrian Mulholland 1) Question. Two methods which are widely used for the optimization of molecular geometies are the Steepest descents and Newton-Raphson
Example questions for Molecular modelling (Level 4) Dr. Adrian Mulholland 1) Question. Two methods which are widely used for the optimization of molecular geometies are the Steepest descents and Newton-Raphson
Intermolecular Forces in Density Functional Theory
 Intermolecular Forces in Density Functional Theory Problems of DFT Peter Pulay at WATOC2005: There are 3 problems with DFT 1. Accuracy does not converge 2. Spin states of open shell systems often incorrect
Intermolecular Forces in Density Functional Theory Problems of DFT Peter Pulay at WATOC2005: There are 3 problems with DFT 1. Accuracy does not converge 2. Spin states of open shell systems often incorrect
Ionic Bonds. H He: ... Li Be B C :N :O :F: :Ne:
 Ionic Bonds Valence electrons - the electrons in the highest occupied energy level - always electrons in the s and p orbitals - maximum of 8 valence electrons - elements in the same group have the same
Ionic Bonds Valence electrons - the electrons in the highest occupied energy level - always electrons in the s and p orbitals - maximum of 8 valence electrons - elements in the same group have the same
Exercise 1: Structure and dipole moment of a small molecule
 Introduction to computational chemistry Exercise 1: Structure and dipole moment of a small molecule Vesa Hänninen 1 Introduction In this exercise the equilibrium structure and the dipole moment of a small
Introduction to computational chemistry Exercise 1: Structure and dipole moment of a small molecule Vesa Hänninen 1 Introduction In this exercise the equilibrium structure and the dipole moment of a small
MO Calculation for a Diatomic Molecule. /4 0 ) i=1 j>i (1/r ij )
 MO Calculation for a Diatomic Molecule Introduction The properties of any molecular system can in principle be found by looking at the solutions to the corresponding time independent Schrodinger equation
MO Calculation for a Diatomic Molecule Introduction The properties of any molecular system can in principle be found by looking at the solutions to the corresponding time independent Schrodinger equation
with the larger dimerization energy also exhibits the larger structural changes.
 A7. Looking at the image and table provided below, it is apparent that the monomer and dimer are structurally almost identical. Although angular and dihedral data were not included, these data are also
A7. Looking at the image and table provided below, it is apparent that the monomer and dimer are structurally almost identical. Although angular and dihedral data were not included, these data are also
Chapter 11 Chemical Bonds: The Formation of Compounds from Atoms Advanced Chemistry Periodic Trends in Atomic Properties Learning Objective
 Chapter 11 Chemical Bonds: The Formation of Compounds from Atoms Advanced Chemistry 11.1 Periodic Trends in Atomic Properties Discuss the atomic trends Metals are located on the left side of the periodic
Chapter 11 Chemical Bonds: The Formation of Compounds from Atoms Advanced Chemistry 11.1 Periodic Trends in Atomic Properties Discuss the atomic trends Metals are located on the left side of the periodic
Size-extensive wave functions for QMC A linear-scaling GVB approach
 Size-extensive wave functions for QMC A linear-scaling GVB approach Claudia Filippi, University of Twente, The Netherlands Francesco Fracchia, University of Pisa, Italy Claudio Amovilli, University of
Size-extensive wave functions for QMC A linear-scaling GVB approach Claudia Filippi, University of Twente, The Netherlands Francesco Fracchia, University of Pisa, Italy Claudio Amovilli, University of
Atomic and molecular interaction forces in biology
 Atomic and molecular interaction forces in biology 1 Outline Types of interactions relevant to biology Van der Waals interactions H-bond interactions Some properties of water Hydrophobic effect 2 Types
Atomic and molecular interaction forces in biology 1 Outline Types of interactions relevant to biology Van der Waals interactions H-bond interactions Some properties of water Hydrophobic effect 2 Types
Session 1. Introduction to Computational Chemistry. Computational (chemistry education) and/or (Computational chemistry) education
 Session 1 Introduction to Computational Chemistry 1 Introduction to Computational Chemistry Computational (chemistry education) and/or (Computational chemistry) education First one: Use computational tools
Session 1 Introduction to Computational Chemistry 1 Introduction to Computational Chemistry Computational (chemistry education) and/or (Computational chemistry) education First one: Use computational tools
Teoría del Funcional de la Densidad (Density Functional Theory)
 Teoría del Funcional de la Densidad (Density Functional Theory) Motivation: limitations of the standard approach based on the wave function. The electronic density n(r) as the key variable: Functionals
Teoría del Funcional de la Densidad (Density Functional Theory) Motivation: limitations of the standard approach based on the wave function. The electronic density n(r) as the key variable: Functionals
Computational Chemistry I
 Computational Chemistry I Text book Cramer: Essentials of Quantum Chemistry, Wiley (2 ed.) Chapter 3. Post Hartree-Fock methods (Cramer: chapter 7) There are many ways to improve the HF method. Most of
Computational Chemistry I Text book Cramer: Essentials of Quantum Chemistry, Wiley (2 ed.) Chapter 3. Post Hartree-Fock methods (Cramer: chapter 7) There are many ways to improve the HF method. Most of
Computational Methods. Chem 561
 Computational Methods Chem 561 Lecture Outline 1. Ab initio methods a) HF SCF b) Post-HF methods 2. Density Functional Theory 3. Semiempirical methods 4. Molecular Mechanics Computational Chemistry " Computational
Computational Methods Chem 561 Lecture Outline 1. Ab initio methods a) HF SCF b) Post-HF methods 2. Density Functional Theory 3. Semiempirical methods 4. Molecular Mechanics Computational Chemistry " Computational
QMC dissociation energies of three-electron hemibonded radical cation dimers... and water clusters
 QMC dissociation energies of three-electron hemibonded radical cation dimers... and water clusters Idoia G. de Gurtubay TCM group. Cavendish Laboratory University of Cambridge Idoia G. de Gurtubay. Quantum
QMC dissociation energies of three-electron hemibonded radical cation dimers... and water clusters Idoia G. de Gurtubay TCM group. Cavendish Laboratory University of Cambridge Idoia G. de Gurtubay. Quantum
Quantum Monte Carlo wave functions and their optimization for quantum chemistry
 Quantum Monte Carlo wave functions and their optimization for quantum chemistry Julien Toulouse Université Pierre & Marie Curie and CNRS, Paris, France CEA Saclay, SPhN Orme des Merisiers April 2015 Outline
Quantum Monte Carlo wave functions and their optimization for quantum chemistry Julien Toulouse Université Pierre & Marie Curie and CNRS, Paris, France CEA Saclay, SPhN Orme des Merisiers April 2015 Outline
3/30/2015. Third energy level. Second energy level. Energy absorbed. First energy level. Atomic nucleus. Energy released (as light)
 Chapter 2 An Introduction Chemistry Lecture 2: Energy Levels and Chemical Bonding Electrons are always moving Outside the nucleus in atomic orbitals Maybe usually Average distance from nucleus (size of
Chapter 2 An Introduction Chemistry Lecture 2: Energy Levels and Chemical Bonding Electrons are always moving Outside the nucleus in atomic orbitals Maybe usually Average distance from nucleus (size of
Earth Solid Earth Rocks Minerals Atoms. How to make a mineral from the start of atoms?
 Earth Solid Earth Rocks Minerals Atoms How to make a mineral from the start of atoms? Formation of ions Ions excess or deficit of electrons relative to protons Anions net negative charge Cations net
Earth Solid Earth Rocks Minerals Atoms How to make a mineral from the start of atoms? Formation of ions Ions excess or deficit of electrons relative to protons Anions net negative charge Cations net
CHAPTER 2 INTERATOMIC FORCES. atoms together in a solid?
 CHAPTER 2 INTERATOMIC FORCES What kind of force holds the atoms together in a solid? Interatomic Binding All of the mechanisms which cause bonding between the atoms derive from electrostatic interaction
CHAPTER 2 INTERATOMIC FORCES What kind of force holds the atoms together in a solid? Interatomic Binding All of the mechanisms which cause bonding between the atoms derive from electrostatic interaction
Covalent Bonding. a. O b. Mg c. Ar d. C. a. K b. N c. Cl d. B
 Covalent Bonding 1. Obtain the number of valence electrons for each of the following atoms from its group number and draw the correct Electron Dot Notation (a.k.a. Lewis Dot Structures). a. K b. N c. Cl
Covalent Bonding 1. Obtain the number of valence electrons for each of the following atoms from its group number and draw the correct Electron Dot Notation (a.k.a. Lewis Dot Structures). a. K b. N c. Cl
CCSD(T) benchmarks of non-equilibrium water clusters: the importance of monomer deformation
 CCSD(T) benchmarks of non-equilibrium water clusters: the importance of monomer deformation Biswajit Santra 1, Angelos Michaelides 1,2, and Matthias Scheffler 1 1 Fritz-Haber-Institut der MPG, Berlin,
CCSD(T) benchmarks of non-equilibrium water clusters: the importance of monomer deformation Biswajit Santra 1, Angelos Michaelides 1,2, and Matthias Scheffler 1 1 Fritz-Haber-Institut der MPG, Berlin,
Solutions and Non-Covalent Binding Forces
 Chapter 3 Solutions and Non-Covalent Binding Forces 3.1 Solvent and solution properties Molecules stick together using the following forces: dipole-dipole, dipole-induced dipole, hydrogen bond, van der
Chapter 3 Solutions and Non-Covalent Binding Forces 3.1 Solvent and solution properties Molecules stick together using the following forces: dipole-dipole, dipole-induced dipole, hydrogen bond, van der
CHAPTER-IV. FT-IR and FT-Raman investigation on m-xylol using ab-initio HF and DFT calculations
 4.1. Introduction CHAPTER-IV FT-IR and FT-Raman investigation on m-xylol using ab-initio HF and DFT calculations m-xylol is a material for thermally stable aramid fibers or alkyd resins [1]. In recent
4.1. Introduction CHAPTER-IV FT-IR and FT-Raman investigation on m-xylol using ab-initio HF and DFT calculations m-xylol is a material for thermally stable aramid fibers or alkyd resins [1]. In recent
Quantum Mechanical Simulations
 Quantum Mechanical Simulations Prof. Yan Wang Woodruff School of Mechanical Engineering Georgia Institute of Technology Atlanta, GA 30332, U.S.A. yan.wang@me.gatech.edu Topics Quantum Monte Carlo Hartree-Fock
Quantum Mechanical Simulations Prof. Yan Wang Woodruff School of Mechanical Engineering Georgia Institute of Technology Atlanta, GA 30332, U.S.A. yan.wang@me.gatech.edu Topics Quantum Monte Carlo Hartree-Fock
1 Density functional theory (DFT)
 1 Density functional theory (DFT) 1.1 Introduction Density functional theory is an alternative to ab initio methods for solving the nonrelativistic, time-independent Schrödinger equation H Φ = E Φ. The
1 Density functional theory (DFT) 1.1 Introduction Density functional theory is an alternative to ab initio methods for solving the nonrelativistic, time-independent Schrödinger equation H Φ = E Φ. The
Intermolecular Forces
 Intermolecular Forces Molecular Compounds The simplest molecule is H 2 : Increased electron density draws nuclei together The pair of shared electrons constitutes a covalent bond. Intermolecular Forces
Intermolecular Forces Molecular Compounds The simplest molecule is H 2 : Increased electron density draws nuclei together The pair of shared electrons constitutes a covalent bond. Intermolecular Forces
AP Chemistry Chapter 7: Bonding
 AP Chemistry Chapter 7: Bonding Types of Bonding I. holds everything together! I All bonding occurs because of! Electronegativity difference and bond character A. A difference in electronegativity between
AP Chemistry Chapter 7: Bonding Types of Bonding I. holds everything together! I All bonding occurs because of! Electronegativity difference and bond character A. A difference in electronegativity between
AN INTRODUCTION TO QUANTUM CHEMISTRY. Mark S. Gordon Iowa State University
 AN INTRODUCTION TO QUANTUM CHEMISTRY Mark S. Gordon Iowa State University 1 OUTLINE Theoretical Background in Quantum Chemistry Overview of GAMESS Program Applications 2 QUANTUM CHEMISTRY In principle,
AN INTRODUCTION TO QUANTUM CHEMISTRY Mark S. Gordon Iowa State University 1 OUTLINE Theoretical Background in Quantum Chemistry Overview of GAMESS Program Applications 2 QUANTUM CHEMISTRY In principle,
OVERVIEW OF QUANTUM CHEMISTRY METHODS
 OVERVIEW OF QUANTUM CHEMISTRY METHODS Outline I Generalities Correlation, basis sets Spin II Wavefunction methods Hartree-Fock Configuration interaction Coupled cluster Perturbative methods III Density
OVERVIEW OF QUANTUM CHEMISTRY METHODS Outline I Generalities Correlation, basis sets Spin II Wavefunction methods Hartree-Fock Configuration interaction Coupled cluster Perturbative methods III Density
CHEMICAL BONDS. Determining Percentage Composition, Empirical, and Molecular Formulas for Compounds:
 CHEMICAL BONDS Chemical Bonds: The strong electrostatic forces of attraction holding atoms together in a unit are called chemical bonds (EU 2.C). Reflect a balance in the attractive and repulsive forces
CHEMICAL BONDS Chemical Bonds: The strong electrostatic forces of attraction holding atoms together in a unit are called chemical bonds (EU 2.C). Reflect a balance in the attractive and repulsive forces
Beyond the Hartree-Fock Approximation: Configuration Interaction
 Beyond the Hartree-Fock Approximation: Configuration Interaction The Hartree-Fock (HF) method uses a single determinant (single electronic configuration) description of the electronic wavefunction. For
Beyond the Hartree-Fock Approximation: Configuration Interaction The Hartree-Fock (HF) method uses a single determinant (single electronic configuration) description of the electronic wavefunction. For
Unit 5: Bonding. Place a checkmark next to each item that you can do. If a sample problem is given, complete it as evidence.
 Unit 5: Bonding Place a checkmark next to each item that you can do. If a sample problem is given, complete it as evidence. Intramolecular Forces: forces of attraction within the same molecule. Examples:
Unit 5: Bonding Place a checkmark next to each item that you can do. If a sample problem is given, complete it as evidence. Intramolecular Forces: forces of attraction within the same molecule. Examples:
Ch 6 Chemical Bonding
 Ch 6 Chemical Bonding What you should learn in this section (objectives): Define chemical bond Explain why most atoms form chemical bonds Describe ionic and covalent bonding Explain why most chemical bonding
Ch 6 Chemical Bonding What you should learn in this section (objectives): Define chemical bond Explain why most atoms form chemical bonds Describe ionic and covalent bonding Explain why most chemical bonding
Fixed-Node quantum Monte Carlo for Chemistry
 Fixed-Node quantum Monte Carlo for Chemistry Michel Caffarel Lab. Physique et Chimie Quantiques, CNRS-IRSAMC, Université de Toulouse e-mail : caffarel@irsamc.ups-tlse.fr. p.1/29 The N-body problem of Chemistry
Fixed-Node quantum Monte Carlo for Chemistry Michel Caffarel Lab. Physique et Chimie Quantiques, CNRS-IRSAMC, Université de Toulouse e-mail : caffarel@irsamc.ups-tlse.fr. p.1/29 The N-body problem of Chemistry
Aqueous solutions. Solubility of different compounds in water
 Aqueous solutions Solubility of different compounds in water The dissolution of molecules into water (in any solvent actually) causes a volume change of the solution; the size of this volume change is
Aqueous solutions Solubility of different compounds in water The dissolution of molecules into water (in any solvent actually) causes a volume change of the solution; the size of this volume change is
Australian Journal of Basic and Applied Sciences
 AENSI Journals Australian Journal of Basic and Applied Sciences ISSN:1991-8178 Journal home page: www.ajbasweb.com Theoretical Study for the Effect of Hydroxyl Radical on the Electronic Properties of Cyclobutadiene
AENSI Journals Australian Journal of Basic and Applied Sciences ISSN:1991-8178 Journal home page: www.ajbasweb.com Theoretical Study for the Effect of Hydroxyl Radical on the Electronic Properties of Cyclobutadiene
DENSITY FUNCTIONAL THEORY FOR NON-THEORISTS JOHN P. PERDEW DEPARTMENTS OF PHYSICS AND CHEMISTRY TEMPLE UNIVERSITY
 DENSITY FUNCTIONAL THEORY FOR NON-THEORISTS JOHN P. PERDEW DEPARTMENTS OF PHYSICS AND CHEMISTRY TEMPLE UNIVERSITY A TUTORIAL FOR PHYSICAL SCIENTISTS WHO MAY OR MAY NOT HATE EQUATIONS AND PROOFS REFERENCES
DENSITY FUNCTIONAL THEORY FOR NON-THEORISTS JOHN P. PERDEW DEPARTMENTS OF PHYSICS AND CHEMISTRY TEMPLE UNIVERSITY A TUTORIAL FOR PHYSICAL SCIENTISTS WHO MAY OR MAY NOT HATE EQUATIONS AND PROOFS REFERENCES
Concepts of Chemical Bonding and Molecular Geometry Part 1: Ionic and Covalent Bonds. David A. Katz Pima Community College Tucson, AZ
 Concepts of Chemical Bonding and Molecular Geometry Part 1: Ionic and Covalent Bonds David A. Katz Pima Community College Tucson, AZ Chemical Bonds Three basic types of bonds: Ionic Electrostatic attraction
Concepts of Chemical Bonding and Molecular Geometry Part 1: Ionic and Covalent Bonds David A. Katz Pima Community College Tucson, AZ Chemical Bonds Three basic types of bonds: Ionic Electrostatic attraction
Big Idea: Ionic Bonds: Ionic Bonds: Metals: Nonmetals: Covalent Bonds: Ionic Solids: What ions do atoms form? Electron Electron
 Chapter 13: Phenomena Phenomena: Scientists measured the bond angles of some common molecules. In the pictures below each line represents a bond that contains 2 electrons. If multiple lines are drawn together
Chapter 13: Phenomena Phenomena: Scientists measured the bond angles of some common molecules. In the pictures below each line represents a bond that contains 2 electrons. If multiple lines are drawn together
Molecular mechanics. classical description of molecules. Marcus Elstner and Tomáš Kubař. April 29, 2016
 classical description of molecules April 29, 2016 Chemical bond Conceptual and chemical basis quantum effect solution of the SR numerically expensive (only small molecules can be treated) approximations
classical description of molecules April 29, 2016 Chemical bond Conceptual and chemical basis quantum effect solution of the SR numerically expensive (only small molecules can be treated) approximations
Why Is Molecular Interaction Important in Our Life
 Why Is Molecular Interaction Important in ur Life QuLiS and Graduate School of Science iroshima University http://www.nabit.hiroshima-u.ac.jp/iwatasue/indexe.htm Suehiro Iwata Sept. 29, 2007 Department
Why Is Molecular Interaction Important in ur Life QuLiS and Graduate School of Science iroshima University http://www.nabit.hiroshima-u.ac.jp/iwatasue/indexe.htm Suehiro Iwata Sept. 29, 2007 Department
T6.2 Molecular Mechanics
 T6.2 Molecular Mechanics We have seen that Benson group additivities are capable of giving heats of formation of molecules with accuracies comparable to those of the best ab initio procedures. However,
T6.2 Molecular Mechanics We have seen that Benson group additivities are capable of giving heats of formation of molecules with accuracies comparable to those of the best ab initio procedures. However,
Jack Simons, Henry Eyring Scientist and Professor Chemistry Department University of Utah
 1. Born-Oppenheimer approx.- energy surfaces 2. Mean-field (Hartree-Fock) theory- orbitals 3. Pros and cons of HF- RHF, UHF 4. Beyond HF- why? 5. First, one usually does HF-how? 6. Basis sets and notations
1. Born-Oppenheimer approx.- energy surfaces 2. Mean-field (Hartree-Fock) theory- orbitals 3. Pros and cons of HF- RHF, UHF 4. Beyond HF- why? 5. First, one usually does HF-how? 6. Basis sets and notations
Molecular Mechanics: The Ab Initio Foundation
 Molecular Mechanics: The Ab Initio Foundation Ju Li GEM4 Summer School 2006 Cell and Molecular Mechanics in BioMedicine August 7 18, 2006, MIT, Cambridge, MA, USA 2 Outline Why are electrons quantum? Born-Oppenheimer
Molecular Mechanics: The Ab Initio Foundation Ju Li GEM4 Summer School 2006 Cell and Molecular Mechanics in BioMedicine August 7 18, 2006, MIT, Cambridge, MA, USA 2 Outline Why are electrons quantum? Born-Oppenheimer
Chimica Farmaceutica
 Chimica Farmaceutica Drug Targets Why should chemicals, some of which have remarkably simple structures, have such an important effect «in such a complicated and large structure as a human being? The answer
Chimica Farmaceutica Drug Targets Why should chemicals, some of which have remarkably simple structures, have such an important effect «in such a complicated and large structure as a human being? The answer
in Halogen-Bonded Complexes
 9 Resonance Assistance and Cooperativity in Halogen-Bonded Complexes Previously appeared as Covalency in Resonance-Assisted Halogen Bonds Demonstrated with Cooperativity in N-Halo-Guanine Quartets L. P.
9 Resonance Assistance and Cooperativity in Halogen-Bonded Complexes Previously appeared as Covalency in Resonance-Assisted Halogen Bonds Demonstrated with Cooperativity in N-Halo-Guanine Quartets L. P.
Introduction to computational chemistry Exercise I: Structure and electronic energy of a small molecule. Vesa Hänninen
 Introduction to computational chemistry Exercise I: Structure and electronic energy of a small molecule Vesa Hänninen 1 Introduction In this exercise the equilibrium structure and the electronic energy
Introduction to computational chemistry Exercise I: Structure and electronic energy of a small molecule Vesa Hänninen 1 Introduction In this exercise the equilibrium structure and the electronic energy
CHEMICAL BONDING IONIC BONDS COVALENT BONDS HYDROGEN BONDS METALLIC BONDS
 CHEMICAL BONDING IONIC BONDS COVALENT BONDS HYDROGEN BONDS METALLIC BONDS IONIC BONDING When an atom of a nonmetal takes one or more electrons from an atom of a metal so both atoms end up with eight valence
CHEMICAL BONDING IONIC BONDS COVALENT BONDS HYDROGEN BONDS METALLIC BONDS IONIC BONDING When an atom of a nonmetal takes one or more electrons from an atom of a metal so both atoms end up with eight valence
Unit 9: CHEMICAL BONDING
 Unit 9: CEMICAL BNDING Unit 9: Bonding: 1. Electronegativity 2. Intramolecular Bonding 3. Intermolecular Bonding 4. Drawing Lewis Structures 5. Lewis Structures for Polyatomic Ions 6. Exceptions to the
Unit 9: CEMICAL BNDING Unit 9: Bonding: 1. Electronegativity 2. Intramolecular Bonding 3. Intermolecular Bonding 4. Drawing Lewis Structures 5. Lewis Structures for Polyatomic Ions 6. Exceptions to the
Ionic and Covalent Bonding
 1. Define the following terms: a) valence electrons Ionic and Covalent Bonding the electrons in the highest occupied energy level always electrons in the s and p orbitals maximum of 8 valence electrons
1. Define the following terms: a) valence electrons Ionic and Covalent Bonding the electrons in the highest occupied energy level always electrons in the s and p orbitals maximum of 8 valence electrons
Lec20 Fri 3mar17
 564-17 Lec20 Fri 3mar17 [PDF]GAUSSIAN 09W TUTORIAL www.molcalx.com.cn/wp-content/uploads/2015/01/gaussian09w_tutorial.pdf by A Tomberg - Cited by 8 - Related articles GAUSSIAN 09W TUTORIAL. AN INTRODUCTION
564-17 Lec20 Fri 3mar17 [PDF]GAUSSIAN 09W TUTORIAL www.molcalx.com.cn/wp-content/uploads/2015/01/gaussian09w_tutorial.pdf by A Tomberg - Cited by 8 - Related articles GAUSSIAN 09W TUTORIAL. AN INTRODUCTION
1.1 The Fundamental Chemistry of life
 1.1 The Fundamental Chemistry of life Matter makes up everything in the universe, including all living organisms. Matter is composed of elements, a pure substance that cannot be broken down into simpler
1.1 The Fundamental Chemistry of life Matter makes up everything in the universe, including all living organisms. Matter is composed of elements, a pure substance that cannot be broken down into simpler
Characteristics of the interaction in azulene (H 2 X) n=1,2 (X=O,S) clusters.
 Characteristics of the interaction in azulene (H 2 X) n=1,2 (X=O,S) clusters. Enrique M. Cabaleiro-Lago (a), Ángeles Peña-Gallego (b), Jesús Rodríguez-Otero (b), M. Merced Montero-Campillo (b) (a) Departamento
Characteristics of the interaction in azulene (H 2 X) n=1,2 (X=O,S) clusters. Enrique M. Cabaleiro-Lago (a), Ángeles Peña-Gallego (b), Jesús Rodríguez-Otero (b), M. Merced Montero-Campillo (b) (a) Departamento
States of matter Part 1
 Physical pharmacy I 1. States of matter (2 Lectures) 2. Thermodynamics (2 Lectures) 3. Solution of non-electrolyte 4. Solution of electrolyte 5. Ionic equilibria 6. Buffered and isotonic solution Physical
Physical pharmacy I 1. States of matter (2 Lectures) 2. Thermodynamics (2 Lectures) 3. Solution of non-electrolyte 4. Solution of electrolyte 5. Ionic equilibria 6. Buffered and isotonic solution Physical
Unit 9: CHEMICAL BONDING
 Unit 9: CHEMICAL BONDING 1 Unit 9: Bonding: 1. Electronegativity 2. Intramolecular Bonding 3. Intermolecular Bonding 4. Drawing Lewis Structures 5. Lewis Structures for Polyatomic Ions 6. Exceptions to
Unit 9: CHEMICAL BONDING 1 Unit 9: Bonding: 1. Electronegativity 2. Intramolecular Bonding 3. Intermolecular Bonding 4. Drawing Lewis Structures 5. Lewis Structures for Polyatomic Ions 6. Exceptions to
This is a very succinct primer intended as supplementary material for an undergraduate course in physical chemistry.
 1 Computational Chemistry (Quantum Chemistry) Primer This is a very succinct primer intended as supplementary material for an undergraduate course in physical chemistry. TABLE OF CONTENTS Methods...1 Basis
1 Computational Chemistry (Quantum Chemistry) Primer This is a very succinct primer intended as supplementary material for an undergraduate course in physical chemistry. TABLE OF CONTENTS Methods...1 Basis
States of matter Part 1. Lecture 1. University of Kerbala. Hamid Alghurabi Assistant Lecturer in Pharmaceutics. Physical Pharmacy
 Physical pharmacy I 1. States of matter (2 Lectures) 2. Thermodynamics (2 Lectures) 3. Solution of non-electrolyte 4. Solution of electrolyte 5. Ionic equilibria 6. Buffered and isotonic solution Physical
Physical pharmacy I 1. States of matter (2 Lectures) 2. Thermodynamics (2 Lectures) 3. Solution of non-electrolyte 4. Solution of electrolyte 5. Ionic equilibria 6. Buffered and isotonic solution Physical
Atoms can form stable units called molecules by sharing electrons.
 Atoms can form stable units called molecules by sharing electrons. The formation of molecules is the result of intramolecular bonding (within the molecule) e.g. ionic, covalent. Forces that cause the aggregation
Atoms can form stable units called molecules by sharing electrons. The formation of molecules is the result of intramolecular bonding (within the molecule) e.g. ionic, covalent. Forces that cause the aggregation
Atomic structure & interatomic bonding. Chapter two
 Atomic structure & interatomic bonding Chapter two 1 Atomic Structure Mass Charge Proton 1.67 х 10-27 kg + 1.60 х 10-19 C Neutron 1.67 х 10-27 kg Neutral Electron 9.11 х 10-31 kg - 1.60 х 10-19 C Electron
Atomic structure & interatomic bonding Chapter two 1 Atomic Structure Mass Charge Proton 1.67 х 10-27 kg + 1.60 х 10-19 C Neutron 1.67 х 10-27 kg Neutral Electron 9.11 х 10-31 kg - 1.60 х 10-19 C Electron
CHEMISTRY XL-14A CHEMICAL BONDS
 CHEMISTRY XL-14A CHEMICAL BONDS July 16, 2011 Robert Iafe Office Hours 2 July 18-July 22 Monday: 2:00pm in Room MS-B 3114 Tuesday-Thursday: 3:00pm in Room MS-B 3114 Chapter 2 Overview 3 Ionic Bonds Covalent
CHEMISTRY XL-14A CHEMICAL BONDS July 16, 2011 Robert Iafe Office Hours 2 July 18-July 22 Monday: 2:00pm in Room MS-B 3114 Tuesday-Thursday: 3:00pm in Room MS-B 3114 Chapter 2 Overview 3 Ionic Bonds Covalent
8.1 Types of Chemical Bonds List and define three types of bonding. chapter 8 Bonding General Concepts.notebook. September 10, 2015
 chapter 8 Bonding General Concepts.notebook Chapter 8: Bonding: General Concepts Mar 13 11:15 AM 8.1 Types of Chemical Bonds List and define three types of bonding. Bonds are forces that hold groups of
chapter 8 Bonding General Concepts.notebook Chapter 8: Bonding: General Concepts Mar 13 11:15 AM 8.1 Types of Chemical Bonds List and define three types of bonding. Bonds are forces that hold groups of
Elements react to attain stable (doublet or octet) electronic configurations of the noble gases.
 digitalteachers.co.ug Chemical bonding This chapter teaches the different types and names of bonds that exist in substances that keep their constituent particles together. We will understand how these
digitalteachers.co.ug Chemical bonding This chapter teaches the different types and names of bonds that exist in substances that keep their constituent particles together. We will understand how these
Fragmentation methods
 Fragmentation methods Scaling of QM Methods HF, DFT scale as N 4 MP2 scales as N 5 CC methods scale as N 7 What if we could freeze the value of N regardless of the size of the system? Then each method
Fragmentation methods Scaling of QM Methods HF, DFT scale as N 4 MP2 scales as N 5 CC methods scale as N 7 What if we could freeze the value of N regardless of the size of the system? Then each method
Chapter 8 : Covalent Bonding. Section 8.1: Molecular Compounds
 Chapter 8 : Covalent Bonding Section 8.1: Molecular Compounds What is a molecule? A molecular compound? A molecule is a neutral group of atoms joined together by covalent bonds A molecular compound is
Chapter 8 : Covalent Bonding Section 8.1: Molecular Compounds What is a molecule? A molecular compound? A molecule is a neutral group of atoms joined together by covalent bonds A molecular compound is
ATOMIC BONDING Atomic Bonding
 ATOMIC BONDING Atomic Bonding Primary Bonds Secondary Bonds Ionic Covalent Metallic van der Waals 1. IONIC BONDING q 11 Na & 17 Cl These two ions are attracted to eachother by the electrostatic force developed
ATOMIC BONDING Atomic Bonding Primary Bonds Secondary Bonds Ionic Covalent Metallic van der Waals 1. IONIC BONDING q 11 Na & 17 Cl These two ions are attracted to eachother by the electrostatic force developed
Chemical bonding in solids from ab-initio Calculations
 Chemical bonding in solids from ab-initio Calculations 1 Prof.P. Ravindran, Department of Physics, Central University of Tamil Nadu, India & Center for Materials Science and Nanotechnology, University
Chemical bonding in solids from ab-initio Calculations 1 Prof.P. Ravindran, Department of Physics, Central University of Tamil Nadu, India & Center for Materials Science and Nanotechnology, University
Intermolecular and Intramolecular Forces. Introduction
 Intermolecular and Intramolecular Forces Introduction Atoms can form stable units called molecules by sharing electrons. The formation of molecules is the result of intramolecular bonding (within the molecule)
Intermolecular and Intramolecular Forces Introduction Atoms can form stable units called molecules by sharing electrons. The formation of molecules is the result of intramolecular bonding (within the molecule)
The Chemical Context of Life
 Elements and Compounds The Chemical Context of Life Sodium Chlorine! Sodium chloride! An element is a substance that cannot be broken down to other substances by chemical reactions A compound is a substance
Elements and Compounds The Chemical Context of Life Sodium Chlorine! Sodium chloride! An element is a substance that cannot be broken down to other substances by chemical reactions A compound is a substance
Name: Hr: 8 Basic Concepts of Chemical Bonding
 8.1-8.2 8.3-8.5 8.5-8.7 8.8 Name: Hr: 8 Basic Concepts of Chemical Bonding 8.1 Chemical Bonds, Lewis Symbols, and the Octet Rule State the type of bond (ionic, covalent, or metallic) formed between any
8.1-8.2 8.3-8.5 8.5-8.7 8.8 Name: Hr: 8 Basic Concepts of Chemical Bonding 8.1 Chemical Bonds, Lewis Symbols, and the Octet Rule State the type of bond (ionic, covalent, or metallic) formed between any
Chapter 8: Bonding. Section 8.1: Lewis Dot Symbols
 Chapter 8: Bonding Section 8.1: Lewis Dot Symbols The Lewis electron dot symbol is named after Gilbert Lewis. In the Lewis dot symbol, the element symbol represents the nucleus and the inner electrons.
Chapter 8: Bonding Section 8.1: Lewis Dot Symbols The Lewis electron dot symbol is named after Gilbert Lewis. In the Lewis dot symbol, the element symbol represents the nucleus and the inner electrons.
Ab-initio molecular dynamics for High pressure Hydrogen
 Ab-initio molecular dynamics for High pressure Hydrogen Claudio Attaccalite Institut d'electronique, Microélectronique et Nanotechnologie (IEMN), Lille Outline A brief introduction to Quantum Monte Carlo
Ab-initio molecular dynamics for High pressure Hydrogen Claudio Attaccalite Institut d'electronique, Microélectronique et Nanotechnologie (IEMN), Lille Outline A brief introduction to Quantum Monte Carlo
Chapter 6 Chemistry Review
 Chapter 6 Chemistry Review Multiple Choice Identify the choice that best completes the statement or answers the question. Put the LETTER of the correct answer in the blank. 1. The electrons involved in
Chapter 6 Chemistry Review Multiple Choice Identify the choice that best completes the statement or answers the question. Put the LETTER of the correct answer in the blank. 1. The electrons involved in
3: Many electrons. Orbital symmetries. l =2 1. m l
 3: Many electrons Orbital symmetries Atomic orbitals are labelled according to the principal quantum number, n, and the orbital angular momentum quantum number, l. Electrons in a diatomic molecule experience
3: Many electrons Orbital symmetries Atomic orbitals are labelled according to the principal quantum number, n, and the orbital angular momentum quantum number, l. Electrons in a diatomic molecule experience
CHEMISTRY 4021/8021 MIDTERM EXAM 1 SPRING 2014
 CHEMISTRY 4021/8021 Q1) Propose a simple, united-atom molecular mechanics force-field needed to generate a potential energy surface for an isolated molecule of acetone (Me 2 CO). I.e., provide an energy
CHEMISTRY 4021/8021 Q1) Propose a simple, united-atom molecular mechanics force-field needed to generate a potential energy surface for an isolated molecule of acetone (Me 2 CO). I.e., provide an energy
Chapter 13: Phenomena
 Chapter 13: Phenomena Phenomena: Scientists measured the bond angles of some common molecules. In the pictures below each line represents a bond that contains 2 electrons. If multiple lines are drawn together
Chapter 13: Phenomena Phenomena: Scientists measured the bond angles of some common molecules. In the pictures below each line represents a bond that contains 2 electrons. If multiple lines are drawn together
Interplay of hydrogen bonding and aryl-perfluoroaryl interactions in construction of supramolecular aggregates
 Interplay of hydrogen bonding and aryl-perfluoroaryl interactions in construction of supramolecular aggregates Katarzyna EICHSTAEDT Keywords: supramolecular chemistry, crystalengineering, Hydrogen bonding,
Interplay of hydrogen bonding and aryl-perfluoroaryl interactions in construction of supramolecular aggregates Katarzyna EICHSTAEDT Keywords: supramolecular chemistry, crystalengineering, Hydrogen bonding,
Chapter 4. Molecular Structure and Orbitals
 Chapter 4 Molecular Structure and Orbitals Chapter 4 Table of Contents (4.1) (4.2) (4.3) (4.4) (4.5) (4.6) (4.7) Molecular structure: The VSEPR model Bond polarity and dipole moments Hybridization and
Chapter 4 Molecular Structure and Orbitals Chapter 4 Table of Contents (4.1) (4.2) (4.3) (4.4) (4.5) (4.6) (4.7) Molecular structure: The VSEPR model Bond polarity and dipole moments Hybridization and
CHEMICAL BONDS. Electrical forces. Reflect a balance in the attractive and repulsive forces between electrically charged particles
 CHEMICAL BONDS Chemical Bonds: Electrical forces. Reflect a balance in the attractive and repulsive forces between electrically charged particles Lewis Theory of Bonding: Electrons play a fundamental role
CHEMICAL BONDS Chemical Bonds: Electrical forces. Reflect a balance in the attractive and repulsive forces between electrically charged particles Lewis Theory of Bonding: Electrons play a fundamental role
Uptake of OH radical to aqueous aerosol: a computational study
 Uptake of OH radical to aqueous aerosol: a computational study Grigory Andreev Karpov Institute of Physical Chemistry 10 Vorontsovo pole, Moscow, 105064, Russia Institute of Physical Chemistry and Electrochemistry
Uptake of OH radical to aqueous aerosol: a computational study Grigory Andreev Karpov Institute of Physical Chemistry 10 Vorontsovo pole, Moscow, 105064, Russia Institute of Physical Chemistry and Electrochemistry
Unit 9: CHEMICAL BONDING
 Unit 9: CHEMICAL BONDING 1 Unit 9: Bonding: 1. Electronegativity 2. Intramolecular Bonding 3. Intermolecular Bonding 4. Drawing Lewis Structures 5. Lewis Structures for Polyatomic Ions 6. Exceptions to
Unit 9: CHEMICAL BONDING 1 Unit 9: Bonding: 1. Electronegativity 2. Intramolecular Bonding 3. Intermolecular Bonding 4. Drawing Lewis Structures 5. Lewis Structures for Polyatomic Ions 6. Exceptions to
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015,
 Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Study of Ozone in Tribhuvan University, Kathmandu, Nepal. Prof. S. Gurung Central Department of Physics, Tribhuvan University, Kathmandu, Nepal
 Study of Ozone in Tribhuvan University, Kathmandu, Nepal Prof. S. Gurung Central Department of Physics, Tribhuvan University, Kathmandu, Nepal 1 Country of the Mt Everest 2 View of the Mt Everest 3 4 5
Study of Ozone in Tribhuvan University, Kathmandu, Nepal Prof. S. Gurung Central Department of Physics, Tribhuvan University, Kathmandu, Nepal 1 Country of the Mt Everest 2 View of the Mt Everest 3 4 5
States of Matter. Intermolecular Forces. The States of Matter. Intermolecular Forces. Intermolecular Forces
 Intermolecular Forces Have studied INTRAmolecular forces the forces holding atoms together to form compounds. Now turn to forces between molecules INTERmolecular forces. Forces between molecules, between
Intermolecular Forces Have studied INTRAmolecular forces the forces holding atoms together to form compounds. Now turn to forces between molecules INTERmolecular forces. Forces between molecules, between
Unit 6: Molecular Geometry
 Unit 6: Molecular Geometry Molecular Geometry [6-5] the polarity of each bond, along with the geometry of the molecule determines Molecular Polarity. To predict the geometries of more complicated molecules,
Unit 6: Molecular Geometry Molecular Geometry [6-5] the polarity of each bond, along with the geometry of the molecule determines Molecular Polarity. To predict the geometries of more complicated molecules,
The energy associated with electrostatic interactions is governed by Coulomb s law:
 Chapter 8 Concepts of Chemical Bonding Chemical Bonds Three basic types of bonds: Ionic Electrostatic attraction between ions Covalent Sharing of electrons Metallic Metal atoms bonded to several other
Chapter 8 Concepts of Chemical Bonding Chemical Bonds Three basic types of bonds: Ionic Electrostatic attraction between ions Covalent Sharing of electrons Metallic Metal atoms bonded to several other
Density Functional Theory
 Density Functional Theory March 26, 2009 ? DENSITY FUNCTIONAL THEORY is a method to successfully describe the behavior of atomic and molecular systems and is used for instance for: structural prediction
Density Functional Theory March 26, 2009 ? DENSITY FUNCTIONAL THEORY is a method to successfully describe the behavior of atomic and molecular systems and is used for instance for: structural prediction
GEM4 Summer School OpenCourseWare
 GEM4 Summer School OpenCourseWare http://gem4.educommons.net/ http://www.gem4.org/ Lecture: Molecular Mechanics by Ju Li. Given August 9, 2006 during the GEM4 session at MIT in Cambridge, MA. Please use
GEM4 Summer School OpenCourseWare http://gem4.educommons.net/ http://www.gem4.org/ Lecture: Molecular Mechanics by Ju Li. Given August 9, 2006 during the GEM4 session at MIT in Cambridge, MA. Please use
Correlation in correlated materials (mostly transition metal oxides) Lucas K. Wagner University of Illinois at Urbana-Champaign
 Correlation in correlated materials (mostly transition metal oxides) Lucas K. Wagner University of Illinois at Urbana-Champaign Understanding of correlated materials is mostly phenomenological FN- DMC
Correlation in correlated materials (mostly transition metal oxides) Lucas K. Wagner University of Illinois at Urbana-Champaign Understanding of correlated materials is mostly phenomenological FN- DMC
362 Lecture 6 and 7. Spring 2017 Monday, Jan 30
 362 Lecture 6 and 7 Spring 2017 Monday, Jan 30 Quantum Numbers n is the principal quantum number, indicates the size of the orbital, has all positive integer values of 1 to (infinity) l is the angular
362 Lecture 6 and 7 Spring 2017 Monday, Jan 30 Quantum Numbers n is the principal quantum number, indicates the size of the orbital, has all positive integer values of 1 to (infinity) l is the angular
Chapter 11 Intermolecular Forces, Liquids, and Solids. Intermolecular Forces
 Chapter 11, Liquids, and Solids States of Matter The fundamental difference between states of matter is the distance between particles. States of Matter Because in the solid and liquid states particles
Chapter 11, Liquids, and Solids States of Matter The fundamental difference between states of matter is the distance between particles. States of Matter Because in the solid and liquid states particles
Essential Organic Chemistry. Chapter 1
 Essential Organic Chemistry Paula Yurkanis Bruice Chapter 1 Electronic Structure and Covalent Bonding Periodic Table of the Elements 1.1 The Structure of an Atom Atoms have an internal structure consisting
Essential Organic Chemistry Paula Yurkanis Bruice Chapter 1 Electronic Structure and Covalent Bonding Periodic Table of the Elements 1.1 The Structure of an Atom Atoms have an internal structure consisting
The wavefunction that describes a bonding pair of electrons:
 4.2. Molecular Properties from VB Theory a) Bonding and Bond distances The wavefunction that describes a bonding pair of electrons: Ψ b = a(h 1 ) + b(h 2 ) where h 1 and h 2 are HAOs on adjacent atoms
4.2. Molecular Properties from VB Theory a) Bonding and Bond distances The wavefunction that describes a bonding pair of electrons: Ψ b = a(h 1 ) + b(h 2 ) where h 1 and h 2 are HAOs on adjacent atoms
There are two types of bonding that exist between particles interparticle and intraparticle bonding.
 There are two types of bonding that exist between particles interparticle and intraparticle bonding. Intraparticle bonding describes the forces that exist within a particle such as a molecule or ionic
There are two types of bonding that exist between particles interparticle and intraparticle bonding. Intraparticle bonding describes the forces that exist within a particle such as a molecule or ionic
