Structural investigation of single biomolecules
|
|
|
- Nicholas Harmon
- 5 years ago
- Views:
Transcription
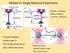
1 Structural investigation of single biomolecules NMR spectroscopy and X-ray crystallography are currently the most common techniques capable of determining the structures of biological macromolecules like proteins and nucleic acids at an atomic level of resolution. The atomic force microscope (AFM) is mostly known for its imaging capabilities, but it is also a very sensitive tool for quantitatively measuring forces on a single molecule level. This capability allows the use of AFM as an alternative technique for the investigation of the structural configuration of macromolecules under native conditions. Measuring forces within and between single molecules provides another approach to the understanding of molecular energy landscapes and chemical reaction kinetics. The AFM can measure individual bonds forming and breaking, rather than being limited to statistical averages over a population. Molecular stretching eperiments Force spectroscopy is an AFM mode where the tip is moved up and down above a single point on the surface, while the deflection or other cantilever properties are measured. This gives a profile of the forces at different heights above the surface. In molecular stretching eperiments, the tip is brought into contact with the surface, the molecules on the substrate or the tip are allowed to interact with each other, and then the cantilever is retracted. This is shown schematically in the image series in Figure 1, for the case of the tip binding to a molecule from the surface and stretching it. As the cantilever is moved away from the surface, the deflection eerts a pulling force. If a molecule is bound between the tip and the surface, then the deflection of the cantilever reflects the force eerted on the molecule as it is stretched. The details of the molecular stretching depend on the structure of the molecule. It may have a defined threedimensional folded structure, as for proteins, for eample, or it may be a disordered chain, such as many other long polymers. The adhesive interaction between the tip and the molecule usually takes place in the repulsive contact part of the force curve, but the features of interest for the molecular stretching are found in the retract curve. Long chain molecules, such as DNA or detran, can be stretched between the tip and the sample. The stiffness and persistence length can be seen from the initial stretching of the molecule. Internal molecular transitions can also be studied at higher forces, such as the melting transition in DNA as the backbone rearranges under higher tension. Molecules with a comple 3-dimensional structure, such as many proteins, can be unfolded in a controlled way so that the structural units can be investigated. Membrane proteins can be pulled out of the membrane, and the popping out of individual alpha-helices has been seen. Titin and bacteriorhodopsin are eamples of proteins that have been intensively studied. Approach Retract Molecule on surface Fig. 1 Schematic diagram of a force spectroscopy eperiment where the AFM tip is used to stretch a single molecule page 1/6 USA China Japan Europe & other regions
2 Characteristics of macromolecules in solution Long molecules in solution are rarely rod-like, and are usually highly fleible along the backbone. Even doublestranded DNA, a relatively "stiff" molecule because of the double-heli structure, is a random tangle for lengths over a few hundred nanometers. There is a large difference between the length of the molecule measured along the chain or backbone (contour length) and the typical dimensions in solution (e.g. the radius of gyration or the end-to-end length), which are much smaller [1]. A Low entropy conformation unlikely in solution Contour length B High entropy conformation typical in solution Typical dimensions << contour length Fig. 2 Schematic diagram of two conformations of a long molecule. Figure 2 shows a schematic representation of a long molecule in two different conformations. In A, the molecule is stretched out to a very etended conformation, where the end-to-end length is approimately the contour length. In B, the molecule is in a relaed random tangle that occupies a much smaller lateral dimension than the contour length. The number of similar conformations is much higher than in A, and this high level of disorder or entropy gives these states a lower free energy. worm-like chain (WLC) model is a continuous filament with chain bending stiffness built in, and is a better model for molecules such as double-stranded DNA. In the WLC model, the persistence length represents the length over which the initial orientation is randomised. Both models can be etended to higher force regimes by allowing for some chain elasticity, so that the length can increase beyond the initial contour length as the bonds are stretched. A brief overview of the models is given in the Appendi, more discussion can be found in reference [2]. Force-etension curves from AFM measurements with the JPK systems can be fitted automatically with these models in the data processing software. The main results are the contour length of the molecule being stretched, and the bending length, depending on the model Proteins generally have a well defined (folded) threedimensional structure in solution. This structure is the key to their biological function as enzymes or structural units, for eample. The polypeptide chain is folded into a relatively stiff structure and held together by many bonds. If the protein is unfolded (by force, chemical or thermal changes), then the free polypeptide chain behaves like a simple linear molecule. Certain subunits of the protein structure, such as alpha-helices or globular domains, can be unfolded sequentially, and the measured "free" length of the molecule based on the stretching curves will increase with each unfolding event. The protein unfolding steps provide information about the normal folded structure and its failure mechanisms. This free energy difference translates to a force that resists the straightening of a molecule from B to A, and acts like an entropic "spring". Force is required to etend the molecule towards a straight conformation even if the bonds in the backbone are not being stretched. Two basic models are commonly used for this entropic elasticity. In the freely jointed chain (FJC) model, the backbone is modelled as small units connected by completely "free" joints, so that adjacent units can have any angle between them without etra energy cost. The Xanthan simple molecular stretching Xanthan is a bacterial polysaccharide, with a molecular backbone identical to cellulose. The chemical properties of the side groups give the polymer many industrial applications, from food stabilization to oil recovery. When a Xanthan molecule is stretched between the tip and the surface, the behaviour is similar to a single (carboymethylated) cellulose chain [3]. High molecular weight anthan polymers can have lengths of several microns, and the stretching curves can be used as an eample for simple molecular etension. For low forces, page 2/6 USA China Japan Europe & other regions
3 the chain is stretched against the entropic spring. At some point, as the end-to-end length approaches the contour length, the elasticity of the backbone becomes important. such as proteins, a series of these stretching events are generally seen as different parts of the structure unfold and are etended, over shorter ranges. Figure 3 shows the retract curve of a single Xanthan molecule being stretched in a phosphate buffer solution, using the NanoWizard AFM. The cantilever deflection has been calibrated using the thermal noise method [4]. The data is plotted such that a larger negative force corresponds to the tip being deflected more towards the surface. For fitting with WLC or FJC models, the tipsample separation must be calculated first, to take account of the deflection of the cantilever. This is also included in the standard JPK data processing software, and the data here is shown as force against tip-sample separation. Titin sequential unfolding of subunits Titin is a protein from muscle tissue, which consists of several repeated globular domains. Within the muscle tissue it is responsible for the mechanical properties. The AFM can be used to study the nano-mechanical properties of individual titin molecules. Fig. 4 Schematic diagram of titin stretching, as successive globular domains unfold. Fig. 3 Force spectroscopy retract curve of a single Xanthan molecule being stretched. A fit curve for the etended FJC model is also shown in green. An etended FJC model fit curve is shown in Figure 3, fitted using a contour length of 1.24 microns. The fit for the simple FJC or WLC model was reasonable for small etension forces (around 50 pn in this case), but the curves diverged at higher forces. Hence the etended FJC model with the additional segment elasticity was used (see Appendi), to take account of the backbone stretching at higher forces. As the molecule is stretched, at some point the bonds within a particular domain will fail. This globular domain unfolds, increasing the contour length of the molecule, and temporarily reducing the force. With further etension, one of the other domains will reach its failure point. This is illustrated schematically in Figure 4. As the protein is pulled, the domains "pop" open sequentially, producing a characteristic sequence of peaks in the deflection curves. In the case of Xanthan, a single long chain is stretched between the tip and the surface. Since the molecule is so long, the force to etend it is very small for much of the stretching curve, and the behaviour is dominated by the higher-force stretching where the stretched length is near the contour length. For more comple folded molecules, Fig. 5 Force curve of titin stretching, showing sequential unfolding of the globular domains, with fitted WLC etension curves page 3/6 USA China Japan Europe & other regions
4 A typical titin etension force spectroscopy retract curve is shown in Figure 5. The characteristic sequence of unfolding events can be clearly seen. The curve of each successive etension part of the curve can be fitted with the FJC or WLC model to find the corresponding free amino acid length. The etended length of one domain (increase in contour length between successive peaks) is 28nm [5]. The JPK data processing software automatically recognises the jumps between different unfolding events, and a multi-fit routine automatically fits all the etension regions simultaneously with the selected fit model. In Figure 5, seven WLC fits are shown in green, for the seven Ig domains that have been unfolded. A schematic diagram of the unfolding process is shown in Figure 7. The seven alpha-helices are shown with the AFM tip picking up one end (in the actual protein, the alpha-helices are grouped together in a bundle, not a line). There are many possible unfolding pathways, but the most likely ones involve the alpha-helices being pulled out in pairs. A progression of three partially unfolded states is shown Figure 7, with pairs of alpha-helices unfolding as they are etracted from the membrane. Bacteriorhodopsin etraction and unfolding Bacteriorhodopsin is an integral membrane protein from bacterial cells, which generates energy for the cells from light photons. There are seven transmembrane alphahelices, with more disorganized peptide loops between them. When one end of the protein is grasped and pulled, then the alpha-helices are pulled out of the membrane, and they unfold as they are pulled out. Fig. 6 Bacteriorhodopsin topography image 30 nm 50 nm, 0.4 nm height range. Fig. 7 Schematic diagram of bacteriorhodopsin unfolding, with intermediate states as the alpha-helices are pulled out in pairs. Figure 8 shows a bacteriorhodopsin unfolding curve, with the force plotted against tip-sample separation. There is some short-range adhesion at the surface, followed by three main stretching and unfolding events, corresponding to the three states shown in Figure 7. The main peaks are around 20, 40 and 60 nm, as epected from the literature [6]. The cantilever was calibrated using the thermal noise method [4], and the eperiments carried out under buffer (10mM TRIS, 150mM KCl, ph 7.6). Purple membrane purified from bacteria consists of 25% lipids and 75% bacteriorhodopsin. The protein is arranged as trimers in a 2-dimensional crystal structure, which can be imaged using the AFM, as shown in Figure 6. The protein can be imaged and then individual proteins selected and pulled using the tip. The tip can be coated specially, and the molecules modified to pick up a particular part of the structure, or the tip can be pressed into the surface to pick up a molecule non-specifically [6]. Fig. 6 Unfolding curve of bacteriorhodopsin being pulled out of purple membrane. Three worm-like chain (WLC) fits are shown in green superimposed on the data. page 4/6 USA China Japan Europe & other regions
5 The eact position of the bond failure events varies from curve to curve, since the bonds have an energy in the range of the thermal energy at room temperature. For a bond with energy near the thermal energy, there is some probability of it failing within a certain time, even with no applied force. This corresponds to the normal "off-rate" seen for the binding reaction free in solution. For bonds with energies further above the thermal energy, which are normally stable, the probability of them failing spontaneously increases as a force is applied [7]. One way to think of a stretching event, therefore, is that the bonds are strained until the energy barrier to unfolding is within the thermal range. The actual moment when the bonds suddenly fail depends on a random thermal fluctuation, which takes the molecule over the unfolding energy barrier. The time dependence means that the measured unfolding rate will depend on the loading rate, or the speed with which the tip is pulling the free end of the molecule [7,8]. the tertiary structure of membrane proteins. Specific coating of the probe, and modification of the protein to have specific "pick-up" points can lead to pulling the protein at different locations, and hence further information about how the structural units are connected together in the 3-D structure [6,9,10]. Thus AFM is able to give insight into the configuration of a single molecule and allow an independent measurement of molecular properties to refine molecular modelling techniques. AFM can also be combined with singlemolecule fluorescence techniques such as TIRF or FRET. This report concentrates on the use of the AFM to manipulate single molecules to etract information about the molecular structure or conformation, and intramolecular binding forces. It is also possible to study time-dependent phenomena, such as intramolecular dynamics in macromolecules, molecular recognition or protein folding. The unfolding curves also show different pathways for the molecule to unfold and pull out of the membrane. From a knowledge of the protein structure and amino acid sequence, the etensional curves can be fitted to show clearly which unfolding path is taken in a particular force curve [9,10]. Applications A wide variety of proteins can be stretched and unfolded using the AFM, to gain information about both the normal protein structure and its failure modes. In the case of bacteriorhodopsin, the protein forms very highly packed structures in the bacterial cell wall, and so is one of the few membrane proteins that can be crystallized for highresolution structural determination. This method can be more generally applied, however. For most membrane proteins, the amino acid sequence is much easier to determine than the folded structure, since most membrane proteins are not suitable for crystallization. Unfolding events measured with the AFM, which show the change in amino acid length as different structural units unfold, can be used with the known sequence to interpret Literature [1] Israelachvili J. Intermolecular and surface forces, Academic Press, London ISBN [2] Janshoff A., Neitzert M., Obersörfer Y., and Fuchs H. "Force Spectroscopy of molecular systems single molecule spectroscopy of polymers and biomolecules". Angew. Chem. Int. Ed. 39: (2000). [3] Li H., Rief M., Oesterhelt F. and Gaub H.E. "Single-molecule force spectroscopy on Xanthan by AFM". Advanced Materials 3(4): (1998). [4] Hutter J.L. and Bechhoefer J. "Calibration of atomic-force microscope tips". Rev. Sci. Instrum. 64: (1993). [5] Rief M., Gautel M., Österhelt F., Fernandez J.M. and Gaub H.E. "Reversible unfolding of individual titin immunoglobulin domains by AFM" Science 276: (1997). [6] Oesterhelt F., Oesterhelt D., Pfeiffer M., Engel A, Gaub H.E. and Müller J. "Unfolding pathways of individual bacteriorhodopsins". Science 288: (2000). [7] Evans E. "Probing the relationship between force lifetime and chemistry in single molecular bonds" Annu. Rev. Biophys. Biomol. Struct. 30: (2001). [8] Evans E. and Ritchie K. "Dynamic strength of molecular adhesion bonds" Biophys. J. 72(4): (1997). [9] Kedrov A., Ziegler C. and Müller D.J. "Differentiating ligand and inhibitor interactions of a single antiporter" J. Mol. Biol. 362: (2006). [10] Kedrov A., Appel M., Baumann H., Ziegler C. and Müller D.J. "Eamining the dynamic energy landscape of an antiporter upon inhibitor binding J. Mol. Biol. 375: (2008). page 5/6 USA China Japan Europe & other regions
6 Appendi: Theory For more details, see the review by Janshoff et al. [1]. Freely jointed chain (FJC): Chain made from rigid links, with free joints between them. Chain link size (Kuhn length) l K Contour length L = n l K k T ( ) B F l p L 2 Fit parameter: Persistence length, l p L 1 4 l K End-to-end length This model can also be etended to include the elasticity of the polymer backbone. F l K ( F ) Lcoth k BT k BT F l K Fit parameter: Kuhn length, l K The model can be etended to cover elastic deformation of the backbone, so that it is possible to have an end-to-end length > L. The equation is modified with a segment elasticity, κ, to become: F l K ( F ) Lcoth k BT k BT F 1 F l K L Wormlike chain (WLC): l p Continuous filament, with resistance to bending. Average length over which the direction becomes random, (persistence length) l p Contour length L End-to-end length page 6/6 USA China Japan Europe & other regions
Multimedia : Fibronectin and Titin unfolding simulation movies.
 I LECTURE 21: SINGLE CHAIN ELASTICITY OF BIOMACROMOLECULES: THE GIANT PROTEIN TITIN AND DNA Outline : REVIEW LECTURE #2 : EXTENSIBLE FJC AND WLC... 2 STRUCTURE OF MUSCLE AND TITIN... 3 SINGLE MOLECULE
I LECTURE 21: SINGLE CHAIN ELASTICITY OF BIOMACROMOLECULES: THE GIANT PROTEIN TITIN AND DNA Outline : REVIEW LECTURE #2 : EXTENSIBLE FJC AND WLC... 2 STRUCTURE OF MUSCLE AND TITIN... 3 SINGLE MOLECULE
Announcements. Homework 3 (Klaus Schulten s Lecture): Due Wednesday at noon. Next homework assigned. Due Wednesday March 1.
 Announcements Homework 3 (Klaus Schulten s Lecture): Due Wednesday at noon. Next homework assigned. Due Wednesday March 1. No lecture next Monday, Feb. 27 th! (Homework is a bit longer.) Marco will have
Announcements Homework 3 (Klaus Schulten s Lecture): Due Wednesday at noon. Next homework assigned. Due Wednesday March 1. No lecture next Monday, Feb. 27 th! (Homework is a bit longer.) Marco will have
Single-Molecule Force Spectroscopy on Poly(acrylic acid) by AFM
 2120 Langmuir 1999, 15, 2120-2124 Single-Molecule Force Spectroscopy on Poly(acrylic acid) by AFM Hongbin Li, Bingbing Liu, Xi Zhang,*, Chunxiao Gao, Jiacong Shen, and Guangtian Zou Key Lab for Supramolecular
2120 Langmuir 1999, 15, 2120-2124 Single-Molecule Force Spectroscopy on Poly(acrylic acid) by AFM Hongbin Li, Bingbing Liu, Xi Zhang,*, Chunxiao Gao, Jiacong Shen, and Guangtian Zou Key Lab for Supramolecular
Sec. 2.1 Filaments in the cell 21 PART I - RODS AND ROPES
 Sec. 2.1 Filaments in the cell 21 PART I - RODS AND ROPES Sec. 2.1 Filaments in the cell 22 CHAPTER 2 - POLYMERS The structural elements of the cell can be broadly classified as filaments or sheets, where
Sec. 2.1 Filaments in the cell 21 PART I - RODS AND ROPES Sec. 2.1 Filaments in the cell 22 CHAPTER 2 - POLYMERS The structural elements of the cell can be broadly classified as filaments or sheets, where
Magnetic tweezers and its application to DNA mechanics
 Biotechnological Center Research group DNA motors (Seidel group) Handout for Practical Course Magnetic tweezers and its application to DNA mechanics When: 9.00 am Where: Biotec, 3 rd Level, Room 317 Tutors:
Biotechnological Center Research group DNA motors (Seidel group) Handout for Practical Course Magnetic tweezers and its application to DNA mechanics When: 9.00 am Where: Biotec, 3 rd Level, Room 317 Tutors:
Bringing optics into the nanoscale a double-scanner AFM brings advanced optical experiments within reach
 Bringing optics into the nanoscale a double-scanner AFM brings advanced optical experiments within reach Beyond the diffraction limit The resolution of optical microscopy is generally limited by the diffraction
Bringing optics into the nanoscale a double-scanner AFM brings advanced optical experiments within reach Beyond the diffraction limit The resolution of optical microscopy is generally limited by the diffraction
Models for Single Molecule Experiments
 Models for Single Molecule Experiments Binding Unbinding Folding - Unfolding -> Kramers Bell theory 1 Polymer elasticity: - Hooke s law (?) - FJC (Freely joined chain) - WLC (Worm-like chain) 3 Molecular
Models for Single Molecule Experiments Binding Unbinding Folding - Unfolding -> Kramers Bell theory 1 Polymer elasticity: - Hooke s law (?) - FJC (Freely joined chain) - WLC (Worm-like chain) 3 Molecular
SECOND PUBLIC EXAMINATION. Honour School of Physics Part C: 4 Year Course. Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS
 2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2013 Monday, 17 June, 2.30 pm 5.45 pm 15
2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2013 Monday, 17 June, 2.30 pm 5.45 pm 15
3 Biopolymers Uncorrelated chains - Freely jointed chain model
 3.4 Entropy When we talk about biopolymers, it is important to realize that the free energy of a biopolymer in thermal equilibrium is not constant. Unlike solids, biopolymers are characterized through
3.4 Entropy When we talk about biopolymers, it is important to realize that the free energy of a biopolymer in thermal equilibrium is not constant. Unlike solids, biopolymers are characterized through
Nature Methods: doi: /nmeth Supplementary Figure 1. Principle of force-distance (FD) curve based AFM.
 Supplementary Figure 1 Principle of force-distance (FD) curve based AFM. (a) FD-based AFM contours the sample surface while oscillating the AFM tip with a sine wave at a frequency of 0.25 khz. Pixel-by-pixel
Supplementary Figure 1 Principle of force-distance (FD) curve based AFM. (a) FD-based AFM contours the sample surface while oscillating the AFM tip with a sine wave at a frequency of 0.25 khz. Pixel-by-pixel
Scanning Force Microscopy. i.e. Atomic Force Microscopy. Josef A. Käs Institute for Soft Matter Physics Physics Department. Atomic Force Microscopy
 Scanning Force Microscopy i.e. Atomic Force Microscopy Josef A. Käs Institute for Soft Matter Physics Physics Department Atomic Force Microscopy Single Molecule Force Spectroscopy Zanjan 4 Hermann E. Gaub
Scanning Force Microscopy i.e. Atomic Force Microscopy Josef A. Käs Institute for Soft Matter Physics Physics Department Atomic Force Microscopy Single Molecule Force Spectroscopy Zanjan 4 Hermann E. Gaub
Mechanical Proteins. Stretching imunoglobulin and fibronectin. domains of the muscle protein titin. Adhesion Proteins of the Immune System
 Mechanical Proteins F C D B A domains of the muscle protein titin E Stretching imunoglobulin and fibronectin G NIH Resource for Macromolecular Modeling and Bioinformatics Theoretical Biophysics Group,
Mechanical Proteins F C D B A domains of the muscle protein titin E Stretching imunoglobulin and fibronectin G NIH Resource for Macromolecular Modeling and Bioinformatics Theoretical Biophysics Group,
Measurements of interaction forces in (biological) model systems
 Measurements of interaction forces in (biological) model systems Marina Ruths Department of Chemistry, UMass Lowell What can force measurements tell us about a system? Depending on the technique, we might
Measurements of interaction forces in (biological) model systems Marina Ruths Department of Chemistry, UMass Lowell What can force measurements tell us about a system? Depending on the technique, we might
Lecture 34 Protein Unfolding Thermodynamics
 Physical Principles in Biology Biology 3550 Fall 2018 Lecture 34 Protein Unfolding Thermodynamics Wednesday, 21 November c David P. Goldenberg University of Utah goldenberg@biology.utah.edu Clicker Question
Physical Principles in Biology Biology 3550 Fall 2018 Lecture 34 Protein Unfolding Thermodynamics Wednesday, 21 November c David P. Goldenberg University of Utah goldenberg@biology.utah.edu Clicker Question
Faculty of math and natural science. Universty of Split. Kratky-Porod model. Nataša Vučemilovid-Alagid. February, 2013.
 Faculty of math and natural science Universty of Split Kratky-Porod model Nataša Vučemilovid-Alagid February, 2013. Table of contents 1.Introduction:... 3 1. DNA:... 4 2. Models of polymer elasticity:...
Faculty of math and natural science Universty of Split Kratky-Porod model Nataša Vučemilovid-Alagid February, 2013. Table of contents 1.Introduction:... 3 1. DNA:... 4 2. Models of polymer elasticity:...
the spatial arrangement of atoms in a molecule and the chemical bonds that hold the atoms together Chemical structure Covalent bond Ionic bond
 Chemical structure the spatial arrangement of atoms in a molecule and the chemical bonds that hold the atoms together Covalent bond bond formed by the sharing of valence electrons between atoms Ionic bond
Chemical structure the spatial arrangement of atoms in a molecule and the chemical bonds that hold the atoms together Covalent bond bond formed by the sharing of valence electrons between atoms Ionic bond
MULTIPLE CHOICE. Circle the one alternative that best completes the statement or answers the question.
 Summer Work Quiz - Molecules and Chemistry Name MULTIPLE CHOICE. Circle the one alternative that best completes the statement or answers the question. 1) The four most common elements in living organisms
Summer Work Quiz - Molecules and Chemistry Name MULTIPLE CHOICE. Circle the one alternative that best completes the statement or answers the question. 1) The four most common elements in living organisms
Introduction to" Protein Structure
 Introduction to" Protein Structure Function, evolution & experimental methods Thomas Blicher, Center for Biological Sequence Analysis Learning Objectives Outline the basic levels of protein structure.
Introduction to" Protein Structure Function, evolution & experimental methods Thomas Blicher, Center for Biological Sequence Analysis Learning Objectives Outline the basic levels of protein structure.
Homework #4 Physics 498Bio Spring 2012 Prof. Paul Selvin
 Assigned Wednesday Feb. 22, 2012: Due Wednesday February 29, 10:30am. Hand in at start of class. Late homework is not accepted. (Solution sets will be posted shortly after deadline.) Note: Marco will give
Assigned Wednesday Feb. 22, 2012: Due Wednesday February 29, 10:30am. Hand in at start of class. Late homework is not accepted. (Solution sets will be posted shortly after deadline.) Note: Marco will give
THE UNIVERSITY OF MANITOBA. PAPER NO: 409 LOCATION: Fr. Kennedy Gold Gym PAGE NO: 1 of 6 DEPARTMENT & COURSE NO: CHEM 4630 TIME: 3 HOURS
 PAPER NO: 409 LOCATION: Fr. Kennedy Gold Gym PAGE NO: 1 of 6 DEPARTMENT & COURSE NO: CHEM 4630 TIME: 3 HOURS EXAMINATION: Biochemistry of Proteins EXAMINER: J. O'Neil Section 1: You must answer all of
PAPER NO: 409 LOCATION: Fr. Kennedy Gold Gym PAGE NO: 1 of 6 DEPARTMENT & COURSE NO: CHEM 4630 TIME: 3 HOURS EXAMINATION: Biochemistry of Proteins EXAMINER: J. O'Neil Section 1: You must answer all of
Supplementary Information
 Supplementary Information Mechanical unzipping and rezipping of a single SNARE complex reveals hysteresis as a force-generating mechanism Duyoung Min, Kipom Kim, Changbong Hyeon, Yong Hoon Cho, Yeon-Kyun
Supplementary Information Mechanical unzipping and rezipping of a single SNARE complex reveals hysteresis as a force-generating mechanism Duyoung Min, Kipom Kim, Changbong Hyeon, Yong Hoon Cho, Yeon-Kyun
Nature of matter. Chemical bond is a force that joins atoms
 Nature of matter Atom the smallest unit of matter that cannot be broken down by chemical means The subatomic particles of an atom consist of protons, neutrons and electrons Element is a pure substance
Nature of matter Atom the smallest unit of matter that cannot be broken down by chemical means The subatomic particles of an atom consist of protons, neutrons and electrons Element is a pure substance
Matter and Substances Section 3-1
 Matter and Substances Section 3-1 Key Idea: All matter is made up of atoms. An atom has a positively charges core surrounded by a negatively charged region. An atom is the smallest unit of matter that
Matter and Substances Section 3-1 Key Idea: All matter is made up of atoms. An atom has a positively charges core surrounded by a negatively charged region. An atom is the smallest unit of matter that
Protein Dynamics. The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron.
 Protein Dynamics The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron. Below is myoglobin hydrated with 350 water molecules. Only a small
Protein Dynamics The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron. Below is myoglobin hydrated with 350 water molecules. Only a small
2D Visualisation of SMFS data on membrane proteins
 2D Visualisation of SMFS data on membrane proteins Annalisa Marsico 1, Kerstin Vanselow 1, Jing Wang 2, and Dirk Labudde 1 1 Department of Bioinformatics, Center of Biotechnology, Dresden University of
2D Visualisation of SMFS data on membrane proteins Annalisa Marsico 1, Kerstin Vanselow 1, Jing Wang 2, and Dirk Labudde 1 1 Department of Bioinformatics, Center of Biotechnology, Dresden University of
Soft Matter and Biological Physics
 Dr. Ulrich F. Keyser - ufk20 (at) cam.ac.uk Soft Matter and Biological Physics Question Sheet Michaelmas 2011 Version: November 2, 2011 Question 0: Sedimentation Initially consider identical small particles
Dr. Ulrich F. Keyser - ufk20 (at) cam.ac.uk Soft Matter and Biological Physics Question Sheet Michaelmas 2011 Version: November 2, 2011 Question 0: Sedimentation Initially consider identical small particles
Proteins polymer molecules, folded in complex structures. Konstantin Popov Department of Biochemistry and Biophysics
 Proteins polymer molecules, folded in complex structures Konstantin Popov Department of Biochemistry and Biophysics Outline General aspects of polymer theory Size and persistent length of ideal linear
Proteins polymer molecules, folded in complex structures Konstantin Popov Department of Biochemistry and Biophysics Outline General aspects of polymer theory Size and persistent length of ideal linear
Ch 3: Chemistry of Life. Chemistry Water Macromolecules Enzymes
 Ch 3: Chemistry of Life Chemistry Water Macromolecules Enzymes Chemistry Atom = smallest unit of matter that cannot be broken down by chemical means Element = substances that have similar properties and
Ch 3: Chemistry of Life Chemistry Water Macromolecules Enzymes Chemistry Atom = smallest unit of matter that cannot be broken down by chemical means Element = substances that have similar properties and
Announcements & Lecture Points
 Announcements & Lecture Points Homework 3 (Klaus Schulten s Lecture): Due Wednesday. Quiz returned, next homework on Wednesday. Today s Lecture: Protein Folding, Misfolding, Aggregation. Experimental Approach
Announcements & Lecture Points Homework 3 (Klaus Schulten s Lecture): Due Wednesday. Quiz returned, next homework on Wednesday. Today s Lecture: Protein Folding, Misfolding, Aggregation. Experimental Approach
Introduction to Computational Structural Biology
 Introduction to Computational Structural Biology Part I 1. Introduction The disciplinary character of Computational Structural Biology The mathematical background required and the topics covered Bibliography
Introduction to Computational Structural Biology Part I 1. Introduction The disciplinary character of Computational Structural Biology The mathematical background required and the topics covered Bibliography
End-Tethered Elastin-Like Polypeptides
 Supporting Information Hydration and Conformational Mechanics of Single End-Tethered Elastin-Like Polypeptides Alexei Valiaev, Dong Woo Lim, Scott Schmidler,Robert L. Clark, Ashutosh Chilkoti, and Stefan
Supporting Information Hydration and Conformational Mechanics of Single End-Tethered Elastin-Like Polypeptides Alexei Valiaev, Dong Woo Lim, Scott Schmidler,Robert L. Clark, Ashutosh Chilkoti, and Stefan
BMB November 17, Single Molecule Biophysics (I)
 BMB 178 2017 November 17, 2017 14. Single Molecule Biophysics (I) Goals 1. Understand the information SM experiments can provide 2. Be acquainted with different SM approaches 3. Learn to interpret SM results
BMB 178 2017 November 17, 2017 14. Single Molecule Biophysics (I) Goals 1. Understand the information SM experiments can provide 2. Be acquainted with different SM approaches 3. Learn to interpret SM results
2.1 Atoms, Ions, and Molecules
 2.1 Atoms, Ions, and Molecules Living things consist of atoms of different elements. An atom is the smallest basic unit of matter. An element is one type of atom. 6 elements make up 99% of all living things
2.1 Atoms, Ions, and Molecules Living things consist of atoms of different elements. An atom is the smallest basic unit of matter. An element is one type of atom. 6 elements make up 99% of all living things
Biomolecules. Energetics in biology. Biomolecules inside the cell
 Biomolecules Energetics in biology Biomolecules inside the cell Energetics in biology The production of energy, its storage, and its use are central to the economy of the cell. Energy may be defined as
Biomolecules Energetics in biology Biomolecules inside the cell Energetics in biology The production of energy, its storage, and its use are central to the economy of the cell. Energy may be defined as
Mechanical Proteins. Stretching imunoglobulin and fibronectin. domains of the muscle protein titin. Adhesion Proteins of the Immune System
 Mechanical Proteins F C D B A domains of the muscle protein titin E Stretching imunoglobulin and fibronectin G NIH Resource for Macromolecular Modeling and Bioinformatics Theoretical Biophysics Group,
Mechanical Proteins F C D B A domains of the muscle protein titin E Stretching imunoglobulin and fibronectin G NIH Resource for Macromolecular Modeling and Bioinformatics Theoretical Biophysics Group,
Single molecule force spectroscopy reveals a highly compliant helical
 Supplementary Information Single molecule force spectroscopy reveals a highly compliant helical folding for the 30 nm chromatin fiber Maarten Kruithof, Fan-Tso Chien, Andrew Routh, Colin Logie, Daniela
Supplementary Information Single molecule force spectroscopy reveals a highly compliant helical folding for the 30 nm chromatin fiber Maarten Kruithof, Fan-Tso Chien, Andrew Routh, Colin Logie, Daniela
Basic Laboratory. Materials Science and Engineering. Atomic Force Microscopy (AFM)
 Basic Laboratory Materials Science and Engineering Atomic Force Microscopy (AFM) M108 Stand: 20.10.2015 Aim: Presentation of an application of the AFM for studying surface morphology. Inhalt 1.Introduction...
Basic Laboratory Materials Science and Engineering Atomic Force Microscopy (AFM) M108 Stand: 20.10.2015 Aim: Presentation of an application of the AFM for studying surface morphology. Inhalt 1.Introduction...
Quiz 2 Morphology of Complex Materials
 071003 Quiz 2 Morphology of Complex Materials 1) Explain the following terms: (for states comment on biological activity and relative size of the structure) a) Native State b) Unfolded State c) Denatured
071003 Quiz 2 Morphology of Complex Materials 1) Explain the following terms: (for states comment on biological activity and relative size of the structure) a) Native State b) Unfolded State c) Denatured
Lab Week 2 Module α 2. Denaturing of Proteins by Thermal and Mechanical Force
 3.014 Materials Laboratory October 13-19, Fall 2006 Lab Week 2 Module α 2 Denaturing of Proteins by hermal and Mechanical Force Instructor: Benjamin Wunsch Laboratory Instructions Prepared by: Gretchen
3.014 Materials Laboratory October 13-19, Fall 2006 Lab Week 2 Module α 2 Denaturing of Proteins by hermal and Mechanical Force Instructor: Benjamin Wunsch Laboratory Instructions Prepared by: Gretchen
UNIT 1: BIOCHEMISTRY
 UNIT 1: BIOCHEMISTRY UNIT 1: Biochemistry Chapter 6.1: Chemistry of Life I. Atoms, Ions, and Molecules A. Living things consist of atoms of different elements 1. An atom is the smallest basic unit of matter
UNIT 1: BIOCHEMISTRY UNIT 1: Biochemistry Chapter 6.1: Chemistry of Life I. Atoms, Ions, and Molecules A. Living things consist of atoms of different elements 1. An atom is the smallest basic unit of matter
Ch. 3 Metabolism and Enzymes
 Ch. 3 Metabolism and Enzymes Originally prepared by Kim B. Foglia. Revised and adapted by Nhan A. Pham Flow of energy through life Life is built on chemical reactions that enable energy to flow through
Ch. 3 Metabolism and Enzymes Originally prepared by Kim B. Foglia. Revised and adapted by Nhan A. Pham Flow of energy through life Life is built on chemical reactions that enable energy to flow through
Chemistry in Biology. Section 1. Atoms, Elements, and Compounds
 Section 1 Atoms, Elements, and Compounds Atoms! Chemistry is the study of matter.! Atoms are the building blocks of matter.! Neutrons and protons are located at the center of the atom.! Protons are positively
Section 1 Atoms, Elements, and Compounds Atoms! Chemistry is the study of matter.! Atoms are the building blocks of matter.! Neutrons and protons are located at the center of the atom.! Protons are positively
Chapter 6 Chemistry in Biology
 Section 1: Atoms, Elements, and Compounds Section 2: Chemical Reactions Section 3: Water and Solutions Section 4: The Building Blocks of Life Click on a lesson name to select. 6.1 Atoms, Elements, and
Section 1: Atoms, Elements, and Compounds Section 2: Chemical Reactions Section 3: Water and Solutions Section 4: The Building Blocks of Life Click on a lesson name to select. 6.1 Atoms, Elements, and
Biophysics of Macromolecules
 Biophysics of Macromolecules Lecture 11: Dynamic Force Spectroscopy Rädler/Lipfert SS 2014 - Forced Ligand-Receptor Unbinding - Bell-Evans Theory 22. Mai. 2014 AFM experiments with single molecules custom-built
Biophysics of Macromolecules Lecture 11: Dynamic Force Spectroscopy Rädler/Lipfert SS 2014 - Forced Ligand-Receptor Unbinding - Bell-Evans Theory 22. Mai. 2014 AFM experiments with single molecules custom-built
The protein folding problem consists of two parts:
 Energetics and kinetics of protein folding The protein folding problem consists of two parts: 1)Creating a stable, well-defined structure that is significantly more stable than all other possible structures.
Energetics and kinetics of protein folding The protein folding problem consists of two parts: 1)Creating a stable, well-defined structure that is significantly more stable than all other possible structures.
BBS501 Section 1 9:00 am 10:00 am Monday thru Friday LRC 105 A & B
 BBS501 Section 1 9:00 am 10:00 am Monday thru Friday LRC 105 A & B Lecturers: Dr. Yie-Hwa Chang Room M130 Phone: #79263 E-mail:changy@slu.edu Dr. Tomasz Heyduk Room M99 Phone: #79238 E-mail: heydukt@slu.edu
BBS501 Section 1 9:00 am 10:00 am Monday thru Friday LRC 105 A & B Lecturers: Dr. Yie-Hwa Chang Room M130 Phone: #79263 E-mail:changy@slu.edu Dr. Tomasz Heyduk Room M99 Phone: #79238 E-mail: heydukt@slu.edu
Denaturation and renaturation of proteins
 Denaturation and renaturation of proteins Higher levels of protein structure are formed without covalent bonds. Therefore, they are not as stable as peptide covalent bonds which make protein primary structure
Denaturation and renaturation of proteins Higher levels of protein structure are formed without covalent bonds. Therefore, they are not as stable as peptide covalent bonds which make protein primary structure
F. Piazza Center for Molecular Biophysics and University of Orléans, France. Selected topic in Physical Biology. Lecture 1
 Zhou Pei-Yuan Centre for Applied Mathematics, Tsinghua University November 2013 F. Piazza Center for Molecular Biophysics and University of Orléans, France Selected topic in Physical Biology Lecture 1
Zhou Pei-Yuan Centre for Applied Mathematics, Tsinghua University November 2013 F. Piazza Center for Molecular Biophysics and University of Orléans, France Selected topic in Physical Biology Lecture 1
Scanning Force Microscopy
 Scanning Force Microscopy Roland Bennewitz Rutherford Physics Building 405 Phone 398-3058 roland.bennewitz@mcgill.ca Scanning Probe is moved along scan lines over a sample surface 1 Force Microscopy Data
Scanning Force Microscopy Roland Bennewitz Rutherford Physics Building 405 Phone 398-3058 roland.bennewitz@mcgill.ca Scanning Probe is moved along scan lines over a sample surface 1 Force Microscopy Data
Protein Folding & Stability. Lecture 11: Margaret A. Daugherty. Fall How do we go from an unfolded polypeptide chain to a
 Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2004 How do we go from an unfolded polypeptide chain to a compact folded protein? (Folding of thioredoxin, F. Richards) Structure - Function
Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2004 How do we go from an unfolded polypeptide chain to a compact folded protein? (Folding of thioredoxin, F. Richards) Structure - Function
BIOCHEMISTRY GUIDED NOTES - AP BIOLOGY-
 BIOCHEMISTRY GUIDED NOTES - AP BIOLOGY- ELEMENTS AND COMPOUNDS - anything that has mass and takes up space. - cannot be broken down to other substances. - substance containing two or more different elements
BIOCHEMISTRY GUIDED NOTES - AP BIOLOGY- ELEMENTS AND COMPOUNDS - anything that has mass and takes up space. - cannot be broken down to other substances. - substance containing two or more different elements
The Chemistry of Life
 The Chemistry of Life Things you should be able to do 1. Describe how the unique properties of water support life on Earth. 2. Explain how carbon is uniquely suited to form biological macromolecules. 3.
The Chemistry of Life Things you should be able to do 1. Describe how the unique properties of water support life on Earth. 2. Explain how carbon is uniquely suited to form biological macromolecules. 3.
, to obtain a way to calculate stress from the energy function U(r).
 BIOEN 36 014 LECTURE : MOLECULAR BASIS OF ELASTICITY Estimating Young s Modulus from Bond Energies and Structures First we consider solids, which include mostly nonbiological materials, such as metals,
BIOEN 36 014 LECTURE : MOLECULAR BASIS OF ELASTICITY Estimating Young s Modulus from Bond Energies and Structures First we consider solids, which include mostly nonbiological materials, such as metals,
Biophysics II. Key points to be covered. Molecule and chemical bonding. Life: based on materials. Molecule and chemical bonding
 Biophysics II Life: based on materials By A/Prof. Xiang Yang Liu Biophysics & Micro/nanostructures Lab Department of Physics, NUS Organelle, formed by a variety of molecules: protein, DNA, RNA, polysaccharides
Biophysics II Life: based on materials By A/Prof. Xiang Yang Liu Biophysics & Micro/nanostructures Lab Department of Physics, NUS Organelle, formed by a variety of molecules: protein, DNA, RNA, polysaccharides
Guided Notes Unit 1: Biochemistry
 Name: Date: Block: Chapter 2: The Chemistry of Life I. Concept 2.1: Atoms, Ions, and Molecules a. Atoms Guided Notes Unit 1: Biochemistry i. Atom: _ ii. (They are SUPER small! It would take 3 million carbon
Name: Date: Block: Chapter 2: The Chemistry of Life I. Concept 2.1: Atoms, Ions, and Molecules a. Atoms Guided Notes Unit 1: Biochemistry i. Atom: _ ii. (They are SUPER small! It would take 3 million carbon
Chapter 1. Topic: Overview of basic principles
 Chapter 1 Topic: Overview of basic principles Four major themes of biochemistry I. What are living organism made from? II. How do organism acquire and use energy? III. How does an organism maintain its
Chapter 1 Topic: Overview of basic principles Four major themes of biochemistry I. What are living organism made from? II. How do organism acquire and use energy? III. How does an organism maintain its
Principles of Physical Biochemistry
 Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
EXAM I COURSE TFY4310 MOLECULAR BIOPHYSICS December Suggested resolution
 page 1 of 7 EXAM I COURSE TFY4310 MOLECULAR BIOPHYSICS December 2013 Suggested resolution Exercise 1. [total: 25 p] a) [t: 5 p] Describe the bonding [1.5 p] and the molecular orbitals [1.5 p] of the ethylene
page 1 of 7 EXAM I COURSE TFY4310 MOLECULAR BIOPHYSICS December 2013 Suggested resolution Exercise 1. [total: 25 p] a) [t: 5 p] Describe the bonding [1.5 p] and the molecular orbitals [1.5 p] of the ethylene
Contents. xiii. Preface v
 Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
Final exam. Please write your name on the exam and keep an ID card ready. You may use a calculator (but no computer or smart phone) and a dictionary.
 Biophysics of Macromolecules Prof. D. Braun and Prof. J. Lipfert SS 2015 Final exam Final exam Name: Student number ( Matrikelnummer ): Please write your name on the exam and keep an ID card ready. You
Biophysics of Macromolecules Prof. D. Braun and Prof. J. Lipfert SS 2015 Final exam Final exam Name: Student number ( Matrikelnummer ): Please write your name on the exam and keep an ID card ready. You
Dana Alsulaibi. Jaleel G.Sweis. Mamoon Ahram
 15 Dana Alsulaibi Jaleel G.Sweis Mamoon Ahram Revision of last lectures: Proteins have four levels of structures. Primary,secondary, tertiary and quaternary. Primary structure is the order of amino acids
15 Dana Alsulaibi Jaleel G.Sweis Mamoon Ahram Revision of last lectures: Proteins have four levels of structures. Primary,secondary, tertiary and quaternary. Primary structure is the order of amino acids
Lab Week 2 Module α 2. AFM/DSC Study of Protein Denaturation. Instructor: Gretchen DeVries
 3.014 Materials Laboratory October 24-28, 2005 Lab Week 2 Module α 2 AFM/DSC Study of Protein Denaturation Instructor: Gretchen DeVries Objectives Understand the theory and instrumentation in Differential
3.014 Materials Laboratory October 24-28, 2005 Lab Week 2 Module α 2 AFM/DSC Study of Protein Denaturation Instructor: Gretchen DeVries Objectives Understand the theory and instrumentation in Differential
Secondary Structure. Bioch/BIMS 503 Lecture 2. Structure and Function of Proteins. Further Reading. Φ, Ψ angles alone determine protein structure
 Bioch/BIMS 503 Lecture 2 Structure and Function of Proteins August 28, 2008 Robert Nakamoto rkn3c@virginia.edu 2-0279 Secondary Structure Φ Ψ angles determine protein structure Φ Ψ angles are restricted
Bioch/BIMS 503 Lecture 2 Structure and Function of Proteins August 28, 2008 Robert Nakamoto rkn3c@virginia.edu 2-0279 Secondary Structure Φ Ψ angles determine protein structure Φ Ψ angles are restricted
Biomolecules: lecture 10
 Biomolecules: lecture 10 - understanding in detail how protein 3D structures form - realize that protein molecules are not static wire models but instead dynamic, where in principle every atom moves (yet
Biomolecules: lecture 10 - understanding in detail how protein 3D structures form - realize that protein molecules are not static wire models but instead dynamic, where in principle every atom moves (yet
CAP 5510 Lecture 3 Protein Structures
 CAP 5510 Lecture 3 Protein Structures Su-Shing Chen Bioinformatics CISE 8/19/2005 Su-Shing Chen, CISE 1 Protein Conformation 8/19/2005 Su-Shing Chen, CISE 2 Protein Conformational Structures Hydrophobicity
CAP 5510 Lecture 3 Protein Structures Su-Shing Chen Bioinformatics CISE 8/19/2005 Su-Shing Chen, CISE 1 Protein Conformation 8/19/2005 Su-Shing Chen, CISE 2 Protein Conformational Structures Hydrophobicity
The Chemistry of Life.
 The Chemistry of Life http://www.chem.ufl.edu/~itl/2045_s00/matter/fg01_011.gif Atom: the smallest unit of matter Subatomic particles 1. neutron a. In nucleus b. No charge c. Weight 1dalton 2. proton a.
The Chemistry of Life http://www.chem.ufl.edu/~itl/2045_s00/matter/fg01_011.gif Atom: the smallest unit of matter Subatomic particles 1. neutron a. In nucleus b. No charge c. Weight 1dalton 2. proton a.
The biomolecules of terrestrial life
 Functional groups in biomolecules Groups of atoms that are responsible for the chemical properties of biomolecules The biomolecules of terrestrial life Planets and Astrobiology (2017-2018) G. Vladilo 1
Functional groups in biomolecules Groups of atoms that are responsible for the chemical properties of biomolecules The biomolecules of terrestrial life Planets and Astrobiology (2017-2018) G. Vladilo 1
Recently developed single-molecule atomic force microscope
 Stepwise unfolding of titin under force-clamp atomic force microscopy Andres F. Oberhauser*, Paul K. Hansma, Mariano Carrion-Vazquez*, and Julio M. Fernandez* *Department of Physiology and Biophysics,
Stepwise unfolding of titin under force-clamp atomic force microscopy Andres F. Oberhauser*, Paul K. Hansma, Mariano Carrion-Vazquez*, and Julio M. Fernandez* *Department of Physiology and Biophysics,
2.1 Atoms, Ions, and Molecules. 2.1 Atoms, Ions, and Molecules. 2.1 Atoms, Ions, and Molecules. 2.1 Atoms, Ions, and Molecules
 All living things are based on atoms and their interactions. Living things consist of atoms of different elements. An atom is the smallest basic unit of matter. An element is one type of atom. ydrogen
All living things are based on atoms and their interactions. Living things consist of atoms of different elements. An atom is the smallest basic unit of matter. An element is one type of atom. ydrogen
Lecture 11: Protein Folding & Stability
 Structure - Function Protein Folding: What we know Lecture 11: Protein Folding & Stability 1). Amino acid sequence dictates structure. 2). The native structure represents the lowest energy state for a
Structure - Function Protein Folding: What we know Lecture 11: Protein Folding & Stability 1). Amino acid sequence dictates structure. 2). The native structure represents the lowest energy state for a
Protein Folding & Stability. Lecture 11: Margaret A. Daugherty. Fall Protein Folding: What we know. Protein Folding
 Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2003 Structure - Function Protein Folding: What we know 1). Amino acid sequence dictates structure. 2). The native structure represents
Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2003 Structure - Function Protein Folding: What we know 1). Amino acid sequence dictates structure. 2). The native structure represents
Motif Prediction in Amino Acid Interaction Networks
 Motif Prediction in Amino Acid Interaction Networks Omar GACI and Stefan BALEV Abstract In this paper we represent a protein as a graph where the vertices are amino acids and the edges are interactions
Motif Prediction in Amino Acid Interaction Networks Omar GACI and Stefan BALEV Abstract In this paper we represent a protein as a graph where the vertices are amino acids and the edges are interactions
Supplementary Information
 1 Supplementary Information Figure S1 The V=0.5 Harker section of an anomalous difference Patterson map calculated using diffraction data from the NNQQNY crystal at 1.3 Å resolution. The position of the
1 Supplementary Information Figure S1 The V=0.5 Harker section of an anomalous difference Patterson map calculated using diffraction data from the NNQQNY crystal at 1.3 Å resolution. The position of the
EVPP 110 Lecture Exam #1 Study Questions Fall 2003 Dr. Largen
 EVPP 110 Lecture Exam #1 Study Questions Fall 2003 Dr. Largen These study questions are meant to focus your study of the material for the first exam. The absence here of a topic or point covered in lecture
EVPP 110 Lecture Exam #1 Study Questions Fall 2003 Dr. Largen These study questions are meant to focus your study of the material for the first exam. The absence here of a topic or point covered in lecture
Biophysics 490M Project
 Biophysics 490M Project Dan Han Department of Biochemistry Structure Exploration of aa 3 -type Cytochrome c Oxidase from Rhodobacter sphaeroides I. Introduction: All organisms need energy to live. They
Biophysics 490M Project Dan Han Department of Biochemistry Structure Exploration of aa 3 -type Cytochrome c Oxidase from Rhodobacter sphaeroides I. Introduction: All organisms need energy to live. They
Lecture 12: Biomaterials Characterization in Aqueous Environments
 3.051J/20.340J 1 Lecture 12: Biomaterials Characterization in Aqueous Environments High vacuum techniques are important tools for characterizing surface composition, but do not yield information on surface
3.051J/20.340J 1 Lecture 12: Biomaterials Characterization in Aqueous Environments High vacuum techniques are important tools for characterizing surface composition, but do not yield information on surface
CHEMISTRY. 2 Types of Properties Associated with Matter. Composition of Matter. Physical: properties that do not change the identity of the substance
 CHEMISTRY Composition of Matter Matter Mass Anything that occupies space and has mass Quantity of matter an object has Weight Pull of gravity on an object 2 Types of Properties Associated with Matter Physical:
CHEMISTRY Composition of Matter Matter Mass Anything that occupies space and has mass Quantity of matter an object has Weight Pull of gravity on an object 2 Types of Properties Associated with Matter Physical:
Elasticity of the human red blood cell skeleton
 Biorheology 40 (2003) 247 251 247 IOS Press Elasticity of the human red blood cell skeleton G. Lenormand, S. Hénon, A. Richert, J. Siméon and F. Gallet Laboratoire de Biorhéologie et d Hydrodynamique Physico-Chimique,
Biorheology 40 (2003) 247 251 247 IOS Press Elasticity of the human red blood cell skeleton G. Lenormand, S. Hénon, A. Richert, J. Siméon and F. Gallet Laboratoire de Biorhéologie et d Hydrodynamique Physico-Chimique,
Weight and contact forces: Young's modulus, Hooke's law and material properties
 Weight and contact forces: Young's modulus, Hooke's law and material properties Many objects deform according to Hooke's law; many materials behave elastically and have a Young's modulus. In this section,
Weight and contact forces: Young's modulus, Hooke's law and material properties Many objects deform according to Hooke's law; many materials behave elastically and have a Young's modulus. In this section,
2: CHEMICAL COMPOSITION OF THE BODY
 1 2: CHEMICAL COMPOSITION OF THE BODY Although most students of human physiology have had at least some chemistry, this chapter serves very well as a review and as a glossary of chemical terms. In particular,
1 2: CHEMICAL COMPOSITION OF THE BODY Although most students of human physiology have had at least some chemistry, this chapter serves very well as a review and as a glossary of chemical terms. In particular,
Biochemistry Prof. S. DasGupta Department of Chemistry Indian Institute of Technology Kharagpur. Lecture - 06 Protein Structure IV
 Biochemistry Prof. S. DasGupta Department of Chemistry Indian Institute of Technology Kharagpur Lecture - 06 Protein Structure IV We complete our discussion on Protein Structures today. And just to recap
Biochemistry Prof. S. DasGupta Department of Chemistry Indian Institute of Technology Kharagpur Lecture - 06 Protein Structure IV We complete our discussion on Protein Structures today. And just to recap
Unit 1: Chemistry - Guided Notes
 Scientific Method Notes: Unit 1: Chemistry - Guided Notes 1 Common Elements in Biology: Atoms are made up of: 1. 2. 3. In order to be stable, an atom of an element needs a full valence shell of electrons.
Scientific Method Notes: Unit 1: Chemistry - Guided Notes 1 Common Elements in Biology: Atoms are made up of: 1. 2. 3. In order to be stable, an atom of an element needs a full valence shell of electrons.
The Molecules of Life Chapter 2
 The Molecules of Life Chapter 2 Core concepts 1.The atom is the fundamental unit of matter. 2.Atoms can combine to form molecules linked by chemical bonds. 3.Water is essential for life. 4.Carbon is the
The Molecules of Life Chapter 2 Core concepts 1.The atom is the fundamental unit of matter. 2.Atoms can combine to form molecules linked by chemical bonds. 3.Water is essential for life. 4.Carbon is the
2/18/2013 CHEMISTRY OF CELLS. Carbon Structural Formations. 4 Classes of Organic Compounds (biomolecules)
 CHEMISTRY OF CELLS 11 elements make up all organisms C, O, N, H: 96% weight of human body ORGANIC CHEMISTRY Organic compounds: contain C Inorganic compounds: no C Bonding and Structural Formulas H and
CHEMISTRY OF CELLS 11 elements make up all organisms C, O, N, H: 96% weight of human body ORGANIC CHEMISTRY Organic compounds: contain C Inorganic compounds: no C Bonding and Structural Formulas H and
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability
 Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
Protein Structure. W. M. Grogan, Ph.D. OBJECTIVES
 Protein Structure W. M. Grogan, Ph.D. OBJECTIVES 1. Describe the structure and characteristic properties of typical proteins. 2. List and describe the four levels of structure found in proteins. 3. Relate
Protein Structure W. M. Grogan, Ph.D. OBJECTIVES 1. Describe the structure and characteristic properties of typical proteins. 2. List and describe the four levels of structure found in proteins. 3. Relate
BIRKBECK COLLEGE (University of London)
 BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
The Riboswitch is functionally separated into the ligand binding APTAMER and the decision-making EXPRESSION PLATFORM
 The Riboswitch is functionally separated into the ligand binding APTAMER and the decision-making EXPRESSION PLATFORM Purine riboswitch TPP riboswitch SAM riboswitch glms ribozyme In-line probing is used
The Riboswitch is functionally separated into the ligand binding APTAMER and the decision-making EXPRESSION PLATFORM Purine riboswitch TPP riboswitch SAM riboswitch glms ribozyme In-line probing is used
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09450 Supplementary Table 1 Summary of kinetic parameters. Kinetic parameters were V = V / 1 K / ATP and obtained using the relationships max ( + m [ ]) V d s /( 1/ k [ ATP] + 1 k ) =,
doi:10.1038/nature09450 Supplementary Table 1 Summary of kinetic parameters. Kinetic parameters were V = V / 1 K / ATP and obtained using the relationships max ( + m [ ]) V d s /( 1/ k [ ATP] + 1 k ) =,
Lecture 26: Polymers: DNA Packing and Protein folding 26.1 Problem Set 4 due today. Reading for Lectures 22 24: PKT Chapter 8 [ ].
![Lecture 26: Polymers: DNA Packing and Protein folding 26.1 Problem Set 4 due today. Reading for Lectures 22 24: PKT Chapter 8 [ ]. Lecture 26: Polymers: DNA Packing and Protein folding 26.1 Problem Set 4 due today. Reading for Lectures 22 24: PKT Chapter 8 [ ].](/thumbs/84/90810734.jpg) Lecture 26: Polymers: DA Packing and Protein folding 26.1 Problem Set 4 due today. eading for Lectures 22 24: PKT hapter 8 DA Packing for Eukaryotes: The packing problem for the larger eukaryotic genomes
Lecture 26: Polymers: DA Packing and Protein folding 26.1 Problem Set 4 due today. eading for Lectures 22 24: PKT hapter 8 DA Packing for Eukaryotes: The packing problem for the larger eukaryotic genomes
Multimedia : Podcast : Sacrificial Bonds in Biological Materials; Fantner, et al. Biophys. J , 1411
 3.52 Nanomechanics o Materials and Biomaterials Tuesday 5/1/7 Pro. C. Ortiz, MIT-DMSE I LECTURE 2: THEORETICAL ASPECTS O SINGLE MOLECULE ORCE SPECTROSCOPY 2 : EXTENSIBILITY AND THE WORM LIKE CHAIN (WLC)
3.52 Nanomechanics o Materials and Biomaterials Tuesday 5/1/7 Pro. C. Ortiz, MIT-DMSE I LECTURE 2: THEORETICAL ASPECTS O SINGLE MOLECULE ORCE SPECTROSCOPY 2 : EXTENSIBILITY AND THE WORM LIKE CHAIN (WLC)
Phys 450 Spring 2011 Solution set 6. A bimolecular reaction in which A and B combine to form the product P may be written as:
 Problem Phys 45 Spring Solution set 6 A bimolecular reaction in which A and combine to form the product P may be written as: k d A + A P k d k a where k d is a diffusion-limited, bimolecular rate constant
Problem Phys 45 Spring Solution set 6 A bimolecular reaction in which A and combine to form the product P may be written as: k d A + A P k d k a where k d is a diffusion-limited, bimolecular rate constant
Protein structure (and biomolecular structure more generally) CS/CME/BioE/Biophys/BMI 279 Sept. 28 and Oct. 3, 2017 Ron Dror
 Protein structure (and biomolecular structure more generally) CS/CME/BioE/Biophys/BMI 279 Sept. 28 and Oct. 3, 2017 Ron Dror Please interrupt if you have questions, and especially if you re confused! Assignment
Protein structure (and biomolecular structure more generally) CS/CME/BioE/Biophys/BMI 279 Sept. 28 and Oct. 3, 2017 Ron Dror Please interrupt if you have questions, and especially if you re confused! Assignment
3.052 Nanomechanics of Materials and Biomaterials Tuesday 04/03/07 Prof. C. Ortiz, MIT-DMSE I LECTURE 13: MIDTERM #1 SOLUTIONS REVIEW
 I LECTURE 13: MIDTERM #1 SOLUTIONS REVIEW Outline : HIGH RESOLUTION FORCE SPECTROSCOPY...2-10 General Experiment Description... 2 Verification of Surface Functionalization:Imaging of Planar Substrates...
I LECTURE 13: MIDTERM #1 SOLUTIONS REVIEW Outline : HIGH RESOLUTION FORCE SPECTROSCOPY...2-10 General Experiment Description... 2 Verification of Surface Functionalization:Imaging of Planar Substrates...
Single-Molecule Force Measurements by Nano-Handling of Individual Dendronized Polymers
 This is an open access article published under a Creative Commons Attribution (CC-BY) License, which permits unrestricted use, distribution and reproduction in any medium, provided the author and source
This is an open access article published under a Creative Commons Attribution (CC-BY) License, which permits unrestricted use, distribution and reproduction in any medium, provided the author and source
Quantitative Analysis of Forces in Cells
 Quantitative Analysis of Forces in Cells Anders Carlsson Washington University in St Louis Basic properties of forces in cells Measurement methods and magnitudes of particular types of forces: Polymerization
Quantitative Analysis of Forces in Cells Anders Carlsson Washington University in St Louis Basic properties of forces in cells Measurement methods and magnitudes of particular types of forces: Polymerization
The four forces of nature. Intermolecular forces, surface forces & the Atomic Force Microscope (AFM) Force- and potential curves
 Intermolecular forces, surface forces & the Atomic Force Microscope (AFM) The four forces of nature Strong interaction Holds neutrons and protons together in atomic nuclei. Weak interaction β and elementary
Intermolecular forces, surface forces & the Atomic Force Microscope (AFM) The four forces of nature Strong interaction Holds neutrons and protons together in atomic nuclei. Weak interaction β and elementary
Nanobiotechnology. Place: IOP 1 st Meeting Room Time: 9:30-12:00. Reference: Review Papers. Grade: 40% midterm, 60% final report (oral + written)
 Nanobiotechnology Place: IOP 1 st Meeting Room Time: 9:30-12:00 Reference: Review Papers Grade: 40% midterm, 60% final report (oral + written) Midterm: 5/18 Oral Presentation 1. 20 minutes each person
Nanobiotechnology Place: IOP 1 st Meeting Room Time: 9:30-12:00 Reference: Review Papers Grade: 40% midterm, 60% final report (oral + written) Midterm: 5/18 Oral Presentation 1. 20 minutes each person
Untangling the Mechanics of Entangled Biopolymers
 Untangling the Mechanics of Entangled Biopolymers Rae M. Robertson-Anderson Physics Department University of San Diego students/postdocs: Cole Chapman, PhD Tobias Falzone, PhD Stephanie Gorczyca, USD 16
Untangling the Mechanics of Entangled Biopolymers Rae M. Robertson-Anderson Physics Department University of San Diego students/postdocs: Cole Chapman, PhD Tobias Falzone, PhD Stephanie Gorczyca, USD 16
Viscoelasticity of a Single Polymer Chain
 Netsu Sokutei 33 4 183-190 2006 7 10 2006 7 22 Viscoelasticity of a Single Polymer Chain Ken Nakajima and Toshio Nishi (Received July 10, 2006; Accepted July 22, 2006) Atomic force microscopy (AFM) enabled
Netsu Sokutei 33 4 183-190 2006 7 10 2006 7 22 Viscoelasticity of a Single Polymer Chain Ken Nakajima and Toshio Nishi (Received July 10, 2006; Accepted July 22, 2006) Atomic force microscopy (AFM) enabled
