robustness: revisting the significance of mirna-mediated regulation
|
|
|
- Georgiana Haynes
- 5 years ago
- Views:
Transcription
1 : revisting the significance of mirna-mediated regulation Hervé Seitz IGH (CNRS), Montpellier, France October 13, 2012
2 microrna target identification
3 .. microrna target identification mir: target: N NNNNNNNNNNNNNN NNNNNN 3 the seed
4 Identification of mirna targets Computational programs for target prediction: look for seed matches in 3 UTRs, select the ones that were conserved in evolution.
5 Identification of mirna targets Computational programs for target prediction: look for seed matches in 3 UTRs, select the ones that were conserved in evolution. Such short matches are very frequent (60 % of human coding genes seem to be targeted: Friedman et al., 2009).
6 Identification of mirna targets Computational programs for target prediction: look for seed matches in 3 UTRs, select the ones that were conserved in evolution. Such short matches are very frequent (60 % of human coding genes seem to be targeted: Friedman et al., 2009). = mirnas are implicated in every physiological process in animals.
7 Identification of mirna targets Computational programs for target prediction: look for seed matches in 3 UTRs, select the ones that were conserved in evolution. Such short matches are very frequent (60 % of human coding genes seem to be targeted: Friedman et al., 2009). = mirnas are implicated in every physiological process in animals. mirna-mediated repression is very modest (usually < 2-fold): lower than well tolerated fluctuations in gene expression (e.g., haplosufficiency). Why have these sites been conserved if they are not functional?
8
9 Most computationally predicted targets are not functionally targeted (not repressed enough).
10 Most computationally predicted targets are not functionally targeted (not repressed enough). Their phylogenetic conservation means that these binding sites have a function.
11 Most computationally predicted targets are not functionally targeted (not repressed enough). Their phylogenetic conservation means that these binding sites have a function. That function could be to repress the mirna by titrating it.
12 Most computationally predicted targets are not functionally targeted (not repressed enough). Their phylogenetic conservation means that these binding sites have a function. That function could be to repress the mirna by titrating it..
13 Most computationally predicted targets are not functionally targeted (not repressed enough). Their phylogenetic conservation means that these binding sites have a function. That function could be to repress the mirna by titrating it..
14 Most computationally predicted targets are not functionally targeted (not repressed enough). Their phylogenetic conservation means that these binding sites have a function. That function could be to repress the mirna by titrating it.. pseudo target (insensitive) real target (sensitive)
15 A new interpretation The molecular event is the same (interaction between an mrna and a mirna), but the interpretation of that interaction is different: Current theory: the mirna represses the mrna. New : the mrna represses the mirna (except for a few real targets).
16 Discriminative predictions According to the new : According to the current theory:
17 Discriminative predictions According to the new : Abundantly expressed pseudo-targets have a stronger mirna-modulating effect, so their interaction with the mirna should be better conserved According to the current theory:
18 Discriminative predictions According to the new : Abundantly expressed pseudo-targets have a stronger mirna-modulating effect, so their interaction with the mirna should be better conserved = mrna abundance should be positively correlated with mirna binding site conservation. According to the current theory:
19 Discriminative predictions According to the new : Abundantly expressed pseudo-targets have a stronger mirna-modulating effect, so their interaction with the mirna should be better conserved = mrna abundance should be positively correlated with mirna binding site conservation. According to the current theory: Each target is a real target, its interaction is more or less conserved depending on the functional importance of the regulation (it is not expected to correlate with gene expression).
20 mrna abundance and seed match conservation Does mrna abundance correlate positively with mirna binding site conservation?
21 mrna abundance and seed match conservation Does mrna abundance correlate positively with mirna binding site conservation? mirna binding site conservation: measured by TargetScan s P CT score. mrna abundance in published microarray experiments.
22 mrna abundance and seed match conservation Kendall's τ = (p-value = ) Conservation score (P CT ) of binding sites to mir Gene expression in hypothalamus
23 mrna abundance and seed match conservation Adjusted p value 1e 16 1e 12 1e 08 1e 04 1 hypothalamus Adjusted p value 1e 16 1e 12 1e 08 1e 04 1 kidney Kendall s τ Kendall s τ
24 mrna abundance and seed match conservation Adjusted p value 1e 16 1e 12 1e 08 1e 04 1 hypothalamus Adjusted p value 1e 16 1e 12 1e 08 1e 04 1 kidney Kendall s τ Kendall s τ +34 other mouse tissues, same results: +33 fly tissues, same results:
25 Discriminative predictions According to the new : According to the current theory:
26 Discriminative predictions According to the new : Pseudo-target expression levels can fluctuate between individuals in a natural population (phenotype is robust). According to the current theory:
27 Discriminative predictions According to the new : Pseudo-target expression levels can fluctuate between individuals in a natural population (phenotype is robust). According to the current theory: mirna targets are tightly regulated (inter-individual fluctuation should not exceed repression).
28 Natural variability vs. repression
29 Natural variability vs. repression Baek et al., 2008: quantification of mir-223-mediated repression in mouse neutrophils.
30 Natural variability vs. repression Baek et al., 2008: quantification of mir-223-mediated repression in mouse neutrophils. Blood collection Neutrophil isolation RNA extraction cdna labeling, array hybridization
31 Natural variability vs. repression Baek et al., 2008: quantification of mir-223-mediated repression in mouse neutrophils. Blood collection Neutrophil isolation Blood collection Pooled blood Split in 5 replicates Neutrophil isolation RNA extraction RNA extraction cdna labeling, array hybridization cdna labeling, array hybridization
32 Natural variability vs. repression
33 Natural variability vs. repression
34 Natural variability vs. repression
35 Natural variability vs. repression p: probability that the difference between two individual mice is smaller than repression
36 Natural variability vs. repression
37 Natural variability vs. repression For 168 predicted targets out of 189: inter-individual fluctuations across 5 wild-type mice exceeds mirna-mediated regulation (p-value < 0.05).
38 Natural variability vs. repression For 168 predicted targets out of 189: inter-individual fluctuations across 5 wild-type mice exceeds mirna-mediated regulation (p-value < 0.05).
39 : revisiting mirna target definition
40 : revisiting mirna target definition Every measurable change in gene expression does not translate into a macroscopic, evolutionarily selectable phenotype.
41 : revisiting gene regulation definition High-throughput identification of transcription factor or RNA-binding protein targets: thousands of genes are bound, many of them are not under selective pressure to keep these binding sites.
42 : revisiting gene regulation definition High-throughput identification of transcription factor or RNA-binding protein targets: thousands of genes are bound, many of them are not under selective pressure to keep these binding sites. Microscopic events which are neutral in evolutionary terms.
43 : revisiting gene regulation definition High-throughput identification of transcription factor or RNA-binding protein targets: thousands of genes are bound, many of them are not under selective pressure to keep these binding sites. Microscopic events which are neutral in evolutionary terms. Are there pseudo-targets for these other regulators?
44 Acknowledgements Anna Sergeeva Laura Martinez Natalia Pinzo n Restrepo
45 Acknowledgements Anna Sergeeva Laura Martinez Natalia Pinzo n Restrepo Jessy Presumey and Florence Apparailly (INM, Montpellier, France)
46 Acknowledgements Anna Sergeeva Laura Martinez Natalia Pinzo n Restrepo Jessy Presumey and Florence Apparailly (INM, Montpellier, France)
47 Robustness of pathways Conservative approximations in the assessment of abundance/conservation correlation Positive correlation between gene expression and conservation of mirna binding sites
48 Second paradox enzyme 1 enzyme 2 enzyme 3 Substrate Product 1 Product 2 Product 3
49 mrna abundance and seed match conservation Most predicted targets are expected to be pseudo-targets (conservative approximation: we will consider every predicted target). mirna binding site should correlate with that mrna s abundance in the cells where mirna titration is beneficial (conservative approximation: we will consider whole tissues and organs). Poorly abundant mrnas (without a real titration effect on the mirna) should not exhibit such correlation (conservative approximation: we will consider every mrna). Return
50 mrna abundance and seed match conservation Adjusted p value 1e 16 1e 12 1e 08 1e 04 1 hypothalamus Adjusted p value 1e 16 1e 12 1e 08 1e 04 1 spleen Adjusted p value 1e 16 1e 12 1e 08 1e 04 1 naive B cells Kendall s τ Kendall s τ Kendall s τ kidney ES cells ovary Adjusted p value 1e 16 1e 12 1e 08 1e 04 1 Adjusted p value 1e 16 1e 12 1e 08 1e 04 1 Adjusted p value 1e 16 1e 12 1e 08 1e Kendall s τ Kendall s τ Kendall s τ Return
51 mrna abundance and seed match conservation Adjusted p value 1e 16 1e 12 1e 08 1e 04 1 adult ovary Adjusted p value 1e 16 1e 12 1e 08 1e 04 1 adult eye Adjusted p value 1e 16 1e 12 1e 08 1e 04 1 adult heart Expression of deeply conserved/expression of poorly conserved Expression of deeply conserved/expression of poorly conserved Expression of deeply conserved/expression of poorly conserved adult hind gut adult salivary gland larval feeding trachea Adjusted p value 1e 16 1e 12 1e 08 1e 04 1 Adjusted p value 1e 16 1e 12 1e 08 1e 04 1 Adjusted p value 1e 16 1e 12 1e 08 1e Expression of deeply conserved/expression of poorly conserved Expression of deeply conserved/expression of poorly conserved Expression of deeply conserved/expression of poorly conserved Return
microrna pseudo-targets
 microrna pseudo-targets Natalia Pinzón Restrepo Institut de Génétique Humaine (CNRS), Montpellier, France August, 2012 microrna target identification mir: target: 2 7 5 N NNNNNNNNNNNNNN NNNNNN 3 the seed
microrna pseudo-targets Natalia Pinzón Restrepo Institut de Génétique Humaine (CNRS), Montpellier, France August, 2012 microrna target identification mir: target: 2 7 5 N NNNNNNNNNNNNNN NNNNNN 3 the seed
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday
 Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
10-810: Advanced Algorithms and Models for Computational Biology. microrna and Whole Genome Comparison
 10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
Lesson Overview. Gene Regulation and Expression. Lesson Overview Gene Regulation and Expression
 13.4 Gene Regulation and Expression THINK ABOUT IT Think of a library filled with how-to books. Would you ever need to use all of those books at the same time? Of course not. Now picture a tiny bacterium
13.4 Gene Regulation and Expression THINK ABOUT IT Think of a library filled with how-to books. Would you ever need to use all of those books at the same time? Of course not. Now picture a tiny bacterium
Gene Autoregulation via Intronic micrornas and its Functions
 Gene Autoregulation via Intronic micrornas and its Functions Carla Bosia Department of Theoretical Physics University of Torino and INFN, Italy cbosia@to.infn.it Annecy-le-Vieux 20-22/10/2010 OUTLINE microrna
Gene Autoregulation via Intronic micrornas and its Functions Carla Bosia Department of Theoretical Physics University of Torino and INFN, Italy cbosia@to.infn.it Annecy-le-Vieux 20-22/10/2010 OUTLINE microrna
13.4 Gene Regulation and Expression
 13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
TRANSCRIPTOMICS. (or the analysis of the transcriptome) Mario Cáceres. Main objectives of genomics. Determine the entire DNA sequence of an organism
 TRANSCRIPTOMICS (or the analysis of the transcriptome) Mario Cáceres Main objectives of genomics Determine the entire DNA sequence of an organism Identify and annotate the complete set of genes encoded
TRANSCRIPTOMICS (or the analysis of the transcriptome) Mario Cáceres Main objectives of genomics Determine the entire DNA sequence of an organism Identify and annotate the complete set of genes encoded
Supplementary Figure 1: To test the role of mir-17~92 in orthologous genetic model of ADPKD, we generated Ksp/Cre;Pkd1 F/F (Pkd1-KO) and Ksp/Cre;Pkd1
 Supplementary Figure 1: To test the role of mir-17~92 in orthologous genetic model of ADPKD, we generated Ksp/Cre;Pkd1 F/F (Pkd1-KO) and Ksp/Cre;Pkd1 F/F ;mir-17~92 F/F (Pkd1-miR-17~92KO) mice. (A) Q-PCR
Supplementary Figure 1: To test the role of mir-17~92 in orthologous genetic model of ADPKD, we generated Ksp/Cre;Pkd1 F/F (Pkd1-KO) and Ksp/Cre;Pkd1 F/F ;mir-17~92 F/F (Pkd1-miR-17~92KO) mice. (A) Q-PCR
Hairpin Database: Why and How?
 Hairpin Database: Why and How? Clark Jeffries Research Professor Renaissance Computing Institute and School of Pharmacy University of North Carolina at Chapel Hill, United States Why should a database
Hairpin Database: Why and How? Clark Jeffries Research Professor Renaissance Computing Institute and School of Pharmacy University of North Carolina at Chapel Hill, United States Why should a database
Gene Regulation and Expression
 THINK ABOUT IT Think of a library filled with how-to books. Would you ever need to use all of those books at the same time? Of course not. Now picture a tiny bacterium that contains more than 4000 genes.
THINK ABOUT IT Think of a library filled with how-to books. Would you ever need to use all of those books at the same time? Of course not. Now picture a tiny bacterium that contains more than 4000 genes.
Biology. Biology. Slide 1 of 26. End Show. Copyright Pearson Prentice Hall
 Biology Biology 1 of 26 Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 2 of 26 Gene Regulation: An Example Gene Regulation: An Example
Biology Biology 1 of 26 Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 2 of 26 Gene Regulation: An Example Gene Regulation: An Example
the noisy gene Biology of the Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB)
 Biology of the the noisy gene Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB) day III: noisy bacteria - Regulation of noise (B. subtilis) - Intrinsic/Extrinsic
Biology of the the noisy gene Universidad Autónoma de Madrid Jan 2008 Juan F. Poyatos Spanish National Biotechnology Centre (CNB) day III: noisy bacteria - Regulation of noise (B. subtilis) - Intrinsic/Extrinsic
12-5 Gene Regulation
 12-5 Gene Regulation Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 1 of 26 12-5 Gene Regulation Gene Regulation: An Example Gene
12-5 Gene Regulation Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 1 of 26 12-5 Gene Regulation Gene Regulation: An Example Gene
Upstream Elements Regulating mir-241 and mir-48 Abstract Introduction
 Upstream Elements Regulating mir-241 and mir-48 Hanna Vollbrecht, Tamar Resnick, and Ann Rougvie University of Minnesota: Twin Cities Undergraduate Research Scholarship 2012-2013 Abstract Caenorhabditis
Upstream Elements Regulating mir-241 and mir-48 Hanna Vollbrecht, Tamar Resnick, and Ann Rougvie University of Minnesota: Twin Cities Undergraduate Research Scholarship 2012-2013 Abstract Caenorhabditis
UNIT 6 PART 3 *REGULATION USING OPERONS* Hillis Textbook, CH 11
 UNIT 6 PART 3 *REGULATION USING OPERONS* Hillis Textbook, CH 11 REVIEW: Signals that Start and Stop Transcription and Translation BUT, HOW DO CELLS CONTROL WHICH GENES ARE EXPRESSED AND WHEN? First of
UNIT 6 PART 3 *REGULATION USING OPERONS* Hillis Textbook, CH 11 REVIEW: Signals that Start and Stop Transcription and Translation BUT, HOW DO CELLS CONTROL WHICH GENES ARE EXPRESSED AND WHEN? First of
CONJOINT 541. Translating a Transcriptome at Specific Times and Places. David Morris. Department of Biochemistry
 CONJOINT 541 Translating a Transcriptome at Specific Times and Places David Morris Department of Biochemistry http://faculty.washington.edu/dmorris/ Lecture 1 The Biology and Experimental Analysis of mrna
CONJOINT 541 Translating a Transcriptome at Specific Times and Places David Morris Department of Biochemistry http://faculty.washington.edu/dmorris/ Lecture 1 The Biology and Experimental Analysis of mrna
Predicting Protein Functions and Domain Interactions from Protein Interactions
 Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
MIR-237 is Likely a Developmental Timing Gene that Regulates the L2-to-L3 Transition in C. Elegans
 Marquette University e-publications@marquette Master's Theses (2009 -) Dissertations, Theses, and Professional Projects MIR-237 is Likely a Developmental Timing Gene that Regulates the L2-to-L3 Transition
Marquette University e-publications@marquette Master's Theses (2009 -) Dissertations, Theses, and Professional Projects MIR-237 is Likely a Developmental Timing Gene that Regulates the L2-to-L3 Transition
Lecture 18 June 2 nd, Gene Expression Regulation Mutations
 Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
Chapter 15 Active Reading Guide Regulation of Gene Expression
 Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
SUPPLEMENTARY INFORMATION
 Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
Lecture 7. Development of the Fruit Fly Drosophila
 BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
BIOMED Programming & Modeling for BME Final Exam, , Instructor: Ahmet Sacan
 BIOMED 201 - Programming & Modeling for BME Final Exam, 2011.12.01, Instructor: Ahmet Sacan Sign the honor code below. No credit will be given for the exam without a signed pledge. I have neither given
BIOMED 201 - Programming & Modeling for BME Final Exam, 2011.12.01, Instructor: Ahmet Sacan Sign the honor code below. No credit will be given for the exam without a signed pledge. I have neither given
GCD3033:Cell Biology. Transcription
 Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
GLOBEX Bioinformatics (Summer 2015) Genetic networks and gene expression data
 GLOBEX Bioinformatics (Summer 2015) Genetic networks and gene expression data 1 Gene Networks Definition: A gene network is a set of molecular components, such as genes and proteins, and interactions between
GLOBEX Bioinformatics (Summer 2015) Genetic networks and gene expression data 1 Gene Networks Definition: A gene network is a set of molecular components, such as genes and proteins, and interactions between
Proteomics. Areas of Interest
 Introduction to BioMEMS & Medical Microdevices Proteomics and Protein Microarrays Companion lecture to the textbook: Fundamentals of BioMEMS and Medical Microdevices, by Prof., http://saliterman.umn.edu/
Introduction to BioMEMS & Medical Microdevices Proteomics and Protein Microarrays Companion lecture to the textbook: Fundamentals of BioMEMS and Medical Microdevices, by Prof., http://saliterman.umn.edu/
Matteo Figliuzzi
 Supervisors: Prof. Andrea De Martino, Prof. Enzo Marinari Dottorato in sica, XXVI ciclo (N,1) model (N,M) model 25-10-2013 Systems Biology & Networks (N,1) model (N,M) model Arabidopsis regulatory network
Supervisors: Prof. Andrea De Martino, Prof. Enzo Marinari Dottorato in sica, XXVI ciclo (N,1) model (N,M) model 25-10-2013 Systems Biology & Networks (N,1) model (N,M) model Arabidopsis regulatory network
BME 5742 Biosystems Modeling and Control
 BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
VCE BIOLOGY Relationship between the key knowledge and key skills of the Study Design and the Study Design
 VCE BIOLOGY 2006 2014 Relationship between the key knowledge and key skills of the 2000 2005 Study Design and the 2006 2014 Study Design The following table provides a comparison of the key knowledge (and
VCE BIOLOGY 2006 2014 Relationship between the key knowledge and key skills of the 2000 2005 Study Design and the 2006 2014 Study Design The following table provides a comparison of the key knowledge (and
1 GO: regulation of cell size E-04 2 GO: negative regulation of cell growth GO:
 Table S2: The biological modulated by mir-5701 Sr. No Term Id 1 Term Name 2 Hit Gene Number 3 P-Value 4 1 GO:0008361 regulation of cell size 9 4.37E-04 2 GO:0030308 negative regulation of cell growth 8
Table S2: The biological modulated by mir-5701 Sr. No Term Id 1 Term Name 2 Hit Gene Number 3 P-Value 4 1 GO:0008361 regulation of cell size 9 4.37E-04 2 GO:0030308 negative regulation of cell growth 8
Translation Part 2 of Protein Synthesis
 Translation Part 2 of Protein Synthesis IN: How is transcription like making a jello mold? (be specific) What process does this diagram represent? A. Mutation B. Replication C.Transcription D.Translation
Translation Part 2 of Protein Synthesis IN: How is transcription like making a jello mold? (be specific) What process does this diagram represent? A. Mutation B. Replication C.Transcription D.Translation
3.B.1 Gene Regulation. Gene regulation results in differential gene expression, leading to cell specialization.
 3.B.1 Gene Regulation Gene regulation results in differential gene expression, leading to cell specialization. We will focus on gene regulation in prokaryotes first. Gene regulation accounts for some of
3.B.1 Gene Regulation Gene regulation results in differential gene expression, leading to cell specialization. We will focus on gene regulation in prokaryotes first. Gene regulation accounts for some of
Identifying Signaling Pathways
 These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony Gitter, Mark Craven, Colin Dewey Identifying Signaling Pathways BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018
These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by Anthony Gitter, Mark Craven, Colin Dewey Identifying Signaling Pathways BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2018
Biology I Level - 2nd Semester Final Review
 Biology I Level - 2nd Semester Final Review The 2 nd Semester Final encompasses all material that was discussed during second semester. It s important that you review ALL notes and worksheets from the
Biology I Level - 2nd Semester Final Review The 2 nd Semester Final encompasses all material that was discussed during second semester. It s important that you review ALL notes and worksheets from the
AP Biology Essential Knowledge Cards BIG IDEA 1
 AP Biology Essential Knowledge Cards BIG IDEA 1 Essential knowledge 1.A.1: Natural selection is a major mechanism of evolution. Essential knowledge 1.A.4: Biological evolution is supported by scientific
AP Biology Essential Knowledge Cards BIG IDEA 1 Essential knowledge 1.A.1: Natural selection is a major mechanism of evolution. Essential knowledge 1.A.4: Biological evolution is supported by scientific
Computational Genomics. Reconstructing dynamic regulatory networks in multiple species
 02-710 Computational Genomics Reconstructing dynamic regulatory networks in multiple species Methods for reconstructing networks in cells CRH1 SLT2 SLR3 YPS3 YPS1 Amit et al Science 2009 Pe er et al Recomb
02-710 Computational Genomics Reconstructing dynamic regulatory networks in multiple species Methods for reconstructing networks in cells CRH1 SLT2 SLR3 YPS3 YPS1 Amit et al Science 2009 Pe er et al Recomb
Genome wide analysis of protein and mrna half lives reveals dynamic properties of mammalian gene expression
 Genome wide analysis of protein and mrna half lives reveals dynamic properties of mammalian gene expression Matthias Selbach Cell Signaling and Mass Spectrometry Max Delbrück Center for Molecular Medicine
Genome wide analysis of protein and mrna half lives reveals dynamic properties of mammalian gene expression Matthias Selbach Cell Signaling and Mass Spectrometry Max Delbrück Center for Molecular Medicine
Controlling Gene Expression
 Controlling Gene Expression Control Mechanisms Gene regulation involves turning on or off specific genes as required by the cell Determine when to make more proteins and when to stop making more Housekeeping
Controlling Gene Expression Control Mechanisms Gene regulation involves turning on or off specific genes as required by the cell Determine when to make more proteins and when to stop making more Housekeeping
RANK. Alternative names. Discovery. Structure. William J. Boyle* SUMMARY BACKGROUND
 RANK William J. Boyle* Department of Cell Biology, Amgen, Inc., One Amgen Center Drive, Thousand Oaks, CA 91320-1799, USA * corresponding author tel: 805-447-4304, fax: 805-447-1982, e-mail: bboyle@amgen.com
RANK William J. Boyle* Department of Cell Biology, Amgen, Inc., One Amgen Center Drive, Thousand Oaks, CA 91320-1799, USA * corresponding author tel: 805-447-4304, fax: 805-447-1982, e-mail: bboyle@amgen.com
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1. HSP21 expression in 35S:HSP21 and hsp21 knockdown plants. (a) Since no T- DNA insertion line for HSP21 is available in the publicly available T-DNA collections,
Supplementary Figure 1 Supplementary Figure 1. HSP21 expression in 35S:HSP21 and hsp21 knockdown plants. (a) Since no T- DNA insertion line for HSP21 is available in the publicly available T-DNA collections,
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Edward M. Golenberg Wayne State University Detroit, MI
 Edward M. Golenberg Wayne State University Detroit, MI Targeting Phragmites Success As Invasive Species Phragmites displays multiple life history parameters High seed output and small seed size Rapid growth
Edward M. Golenberg Wayne State University Detroit, MI Targeting Phragmites Success As Invasive Species Phragmites displays multiple life history parameters High seed output and small seed size Rapid growth
SCOTCAT Credits: 20 SCQF Level 7 Semester 1 Academic year: 2018/ am, Practical classes one per week pm Mon, Tue, or Wed
 Biology (BL) modules BL1101 Biology 1 SCOTCAT Credits: 20 SCQF Level 7 Semester 1 10.00 am; Practical classes one per week 2.00-5.00 pm Mon, Tue, or Wed This module is an introduction to molecular and
Biology (BL) modules BL1101 Biology 1 SCOTCAT Credits: 20 SCQF Level 7 Semester 1 10.00 am; Practical classes one per week 2.00-5.00 pm Mon, Tue, or Wed This module is an introduction to molecular and
Written Exam 15 December Course name: Introduction to Systems Biology Course no
 Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Utilizing Illumina high-throughput sequencing technology to gain insights into small RNA biogenesis and function
 Utilizing Illumina high-throughput sequencing technology to gain insights into small RNA biogenesis and function Brian D. Gregory Department of Biology Penn Genome Frontiers Institute University of Pennsylvania
Utilizing Illumina high-throughput sequencing technology to gain insights into small RNA biogenesis and function Brian D. Gregory Department of Biology Penn Genome Frontiers Institute University of Pennsylvania
Name Period The Control of Gene Expression in Prokaryotes Notes
 Bacterial DNA contains genes that encode for many different proteins (enzymes) so that many processes have the ability to occur -not all processes are carried out at any one time -what allows expression
Bacterial DNA contains genes that encode for many different proteins (enzymes) so that many processes have the ability to occur -not all processes are carried out at any one time -what allows expression
Comparative Network Analysis
 Comparative Network Analysis BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by
Comparative Network Analysis BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by
Prokaryotic Regulation
 Prokaryotic Regulation Control of transcription initiation can be: Positive control increases transcription when activators bind DNA Negative control reduces transcription when repressors bind to DNA regulatory
Prokaryotic Regulation Control of transcription initiation can be: Positive control increases transcription when activators bind DNA Negative control reduces transcription when repressors bind to DNA regulatory
Understanding Science Through the Lens of Computation. Richard M. Karp Nov. 3, 2007
 Understanding Science Through the Lens of Computation Richard M. Karp Nov. 3, 2007 The Computational Lens Exposes the computational nature of natural processes and provides a language for their description.
Understanding Science Through the Lens of Computation Richard M. Karp Nov. 3, 2007 The Computational Lens Exposes the computational nature of natural processes and provides a language for their description.
Basic Synthetic Biology circuits
 Basic Synthetic Biology circuits Note: these practices were obtained from the Computer Modelling Practicals lecture by Vincent Rouilly and Geoff Baldwin at Imperial College s course of Introduction to
Basic Synthetic Biology circuits Note: these practices were obtained from the Computer Modelling Practicals lecture by Vincent Rouilly and Geoff Baldwin at Imperial College s course of Introduction to
Clustering and Network
 Clustering and Network Jing-Dong Jackie Han jdhan@picb.ac.cn http://www.picb.ac.cn/~jdhan Copy Right: Jing-Dong Jackie Han What is clustering? A way of grouping together data samples that are similar in
Clustering and Network Jing-Dong Jackie Han jdhan@picb.ac.cn http://www.picb.ac.cn/~jdhan Copy Right: Jing-Dong Jackie Han What is clustering? A way of grouping together data samples that are similar in
Proteomics. Yeast two hybrid. Proteomics - PAGE techniques. Data obtained. What is it?
 Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
Introduction to Bioinformatics
 CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
Computational Genomics. Systems biology. Putting it together: Data integration using graphical models
 02-710 Computational Genomics Systems biology Putting it together: Data integration using graphical models High throughput data So far in this class we discussed several different types of high throughput
02-710 Computational Genomics Systems biology Putting it together: Data integration using graphical models High throughput data So far in this class we discussed several different types of high throughput
Prokaryotic Gene Expression (Learning Objectives)
 Prokaryotic Gene Expression (Learning Objectives) 1. Learn how bacteria respond to changes of metabolites in their environment: short-term and longer-term. 2. Compare and contrast transcriptional control
Prokaryotic Gene Expression (Learning Objectives) 1. Learn how bacteria respond to changes of metabolites in their environment: short-term and longer-term. 2. Compare and contrast transcriptional control
Network Biology-part II
 Network Biology-part II Jun Zhu, Ph. D. Professor of Genomics and Genetic Sciences Icahn Institute of Genomics and Multi-scale Biology The Tisch Cancer Institute Icahn Medical School at Mount Sinai New
Network Biology-part II Jun Zhu, Ph. D. Professor of Genomics and Genetic Sciences Icahn Institute of Genomics and Multi-scale Biology The Tisch Cancer Institute Icahn Medical School at Mount Sinai New
16 CONTROL OF GENE EXPRESSION
 16 CONTROL OF GENE EXPRESSION Chapter Outline 16.1 REGULATION OF GENE EXPRESSION IN PROKARYOTES The operon is the unit of transcription in prokaryotes The lac operon for lactose metabolism is transcribed
16 CONTROL OF GENE EXPRESSION Chapter Outline 16.1 REGULATION OF GENE EXPRESSION IN PROKARYOTES The operon is the unit of transcription in prokaryotes The lac operon for lactose metabolism is transcribed
APGRU6L2. Control of Prokaryotic (Bacterial) Genes
 APGRU6L2 Control of Prokaryotic (Bacterial) Genes 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP u if they have enough of a product, need to stop production
APGRU6L2 Control of Prokaryotic (Bacterial) Genes 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP u if they have enough of a product, need to stop production
Inferring disease-associated lncrnas using expression data and disease-associated protein coding genes
 Inferring disease-associated lncrnas using expression data and disease-associated protein coding genes Xiaoyong Pan Center for non-coding RNA in Technology and Health, Department of Clinical Veterinary
Inferring disease-associated lncrnas using expression data and disease-associated protein coding genes Xiaoyong Pan Center for non-coding RNA in Technology and Health, Department of Clinical Veterinary
Transcription Regulation and Gene Expression in Eukaryotes FS08 Pharmacenter/Biocenter Auditorium 1 Wednesdays 16h15-18h00.
 Transcription Regulation and Gene Expression in Eukaryotes FS08 Pharmacenter/Biocenter Auditorium 1 Wednesdays 16h15-18h00. Promoters and Enhancers Systematic discovery of transcriptional regulatory motifs
Transcription Regulation and Gene Expression in Eukaryotes FS08 Pharmacenter/Biocenter Auditorium 1 Wednesdays 16h15-18h00. Promoters and Enhancers Systematic discovery of transcriptional regulatory motifs
microrna Studies Chen-Hanson Ting SVFIG June 23, 2018
 microrna Studies Chen-Hanson Ting SVFIG June 23, 2018 Summary MicroRNA (mirna) Species and organisms studied mirna in mitocondria Huge genome files mirna in human Chromosome 1 mirna in bacteria Tools used
microrna Studies Chen-Hanson Ting SVFIG June 23, 2018 Summary MicroRNA (mirna) Species and organisms studied mirna in mitocondria Huge genome files mirna in human Chromosome 1 mirna in bacteria Tools used
Control of Gene Expression
 Control of Gene Expression Mechanisms of Gene Control Gene Control in Eukaryotes Master Genes Gene Control In Prokaryotes Epigenetics Gene Expression The overall process by which information flows from
Control of Gene Expression Mechanisms of Gene Control Gene Control in Eukaryotes Master Genes Gene Control In Prokaryotes Epigenetics Gene Expression The overall process by which information flows from
Unit 3: Control and regulation Higher Biology
 Unit 3: Control and regulation Higher Biology To study the roles that genes play in the control of growth and development of organisms To be able to Give some examples of features which are controlled
Unit 3: Control and regulation Higher Biology To study the roles that genes play in the control of growth and development of organisms To be able to Give some examples of features which are controlled
Unit 4 Evaluation Question 1:
 Name: Unit 4 Evaluation Question 1: /7 points A naturally occurring dominant mutant in mice is the Doublefoot (Dbf) mutant. Below is an image of the bones from a wildtype (wt) and Doublefoot mutant mouse.
Name: Unit 4 Evaluation Question 1: /7 points A naturally occurring dominant mutant in mice is the Doublefoot (Dbf) mutant. Below is an image of the bones from a wildtype (wt) and Doublefoot mutant mouse.
Inferring Transcriptional Regulatory Networks from High-throughput Data
 Inferring Transcriptional Regulatory Networks from High-throughput Data Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20
Inferring Transcriptional Regulatory Networks from High-throughput Data Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20
Exploring the mirna Regulatory Network Using Evolutionary Correlations
 Exploring the mirna Regulatory Network Using Evolutionary Correlations The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters. Citation
Exploring the mirna Regulatory Network Using Evolutionary Correlations The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters. Citation
BIS &003 Answers to Assigned Problems May 23, Week /18.6 How would you distinguish between an enhancer and a promoter?
 Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
Inferring Transcriptional Regulatory Networks from Gene Expression Data II
 Inferring Transcriptional Regulatory Networks from Gene Expression Data II Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday
Inferring Transcriptional Regulatory Networks from Gene Expression Data II Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday
Biology 112 Practice Midterm Questions
 Biology 112 Practice Midterm Questions 1. Identify which statement is true or false I. Bacterial cell walls prevent osmotic lysis II. All bacterial cell walls contain an LPS layer III. In a Gram stain,
Biology 112 Practice Midterm Questions 1. Identify which statement is true or false I. Bacterial cell walls prevent osmotic lysis II. All bacterial cell walls contain an LPS layer III. In a Gram stain,
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION
 MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
Correspondence of D. melanogaster and C. elegans developmental stages revealed by alternative splicing characteristics of conserved exons
 Gao and Li BMC Genomics (2017) 18:234 DOI 10.1186/s12864-017-3600-2 RESEARCH ARTICLE Open Access Correspondence of D. melanogaster and C. elegans developmental stages revealed by alternative splicing characteristics
Gao and Li BMC Genomics (2017) 18:234 DOI 10.1186/s12864-017-3600-2 RESEARCH ARTICLE Open Access Correspondence of D. melanogaster and C. elegans developmental stages revealed by alternative splicing characteristics
The geneticist s questions. Deleting yeast genes. Functional genomics. From Wikipedia, the free encyclopedia
 From Wikipedia, the free encyclopedia Functional genomics..is a field of molecular biology that attempts to make use of the vast wealth of data produced by genomic projects (such as genome sequencing projects)
From Wikipedia, the free encyclopedia Functional genomics..is a field of molecular biology that attempts to make use of the vast wealth of data produced by genomic projects (such as genome sequencing projects)
Regulation of Transcription in Eukaryotes. Nelson Saibo
 Regulation of Transcription in Eukaryotes Nelson Saibo saibo@itqb.unl.pt In eukaryotes gene expression is regulated at different levels 1 - Transcription 2 Post-transcriptional modifications 3 RNA transport
Regulation of Transcription in Eukaryotes Nelson Saibo saibo@itqb.unl.pt In eukaryotes gene expression is regulated at different levels 1 - Transcription 2 Post-transcriptional modifications 3 RNA transport
Bi 1x Spring 2014: LacI Titration
 Bi 1x Spring 2014: LacI Titration 1 Overview In this experiment, you will measure the effect of various mutated LacI repressor ribosome binding sites in an E. coli cell by measuring the expression of a
Bi 1x Spring 2014: LacI Titration 1 Overview In this experiment, you will measure the effect of various mutated LacI repressor ribosome binding sites in an E. coli cell by measuring the expression of a
56:198:582 Biological Networks Lecture 8
 56:198:582 Biological Networks Lecture 8 Course organization Two complementary approaches to modeling and understanding biological networks Constraint-based modeling (Palsson) System-wide Metabolism Steady-state
56:198:582 Biological Networks Lecture 8 Course organization Two complementary approaches to modeling and understanding biological networks Constraint-based modeling (Palsson) System-wide Metabolism Steady-state
L3.1: Circuits: Introduction to Transcription Networks. Cellular Design Principles Prof. Jenna Rickus
 L3.1: Circuits: Introduction to Transcription Networks Cellular Design Principles Prof. Jenna Rickus In this lecture Cognitive problem of the Cell Introduce transcription networks Key processing network
L3.1: Circuits: Introduction to Transcription Networks Cellular Design Principles Prof. Jenna Rickus In this lecture Cognitive problem of the Cell Introduce transcription networks Key processing network
Mole_Oce Lecture # 24: Introduction to genomics
 Mole_Oce Lecture # 24: Introduction to genomics DEFINITION: Genomics: the study of genomes or he study of genes and their function. Genomics (1980s):The systematic generation of information about genes
Mole_Oce Lecture # 24: Introduction to genomics DEFINITION: Genomics: the study of genomes or he study of genes and their function. Genomics (1980s):The systematic generation of information about genes
Entrance Exam. BIOLOGY (30 questions, 90 minutes) Name:
 Sarajevo School of Science and Technology Sarajevo, April 2014 Entrance Exam BIOLOGY (30 questions, 90 minutes) Name: Questions 1-22, circle the correct answer. 1. The first evolutionary molecule that
Sarajevo School of Science and Technology Sarajevo, April 2014 Entrance Exam BIOLOGY (30 questions, 90 minutes) Name: Questions 1-22, circle the correct answer. 1. The first evolutionary molecule that
Biomolecular Feedback Systems
 Biomolecular Feedback Systems Domitilla Del Vecchio MIT Richard M. Murray Caltech Version 1.0b, September 14, 2014 c 2014 by Princeton University Press All rights reserved. This is the electronic edition
Biomolecular Feedback Systems Domitilla Del Vecchio MIT Richard M. Murray Caltech Version 1.0b, September 14, 2014 c 2014 by Princeton University Press All rights reserved. This is the electronic edition
Multiple Choice Review- Eukaryotic Gene Expression
 Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Molecular Developmental Physiology and Signal Transduction
 Prof. Dr. J. Vanden Broeck (Animal Physiology and Neurobiology - Dept. of Biology - KU Leuven) Molecular Developmental Physiology and Signal Transduction My Research Team Insect species under study +
Prof. Dr. J. Vanden Broeck (Animal Physiology and Neurobiology - Dept. of Biology - KU Leuven) Molecular Developmental Physiology and Signal Transduction My Research Team Insect species under study +
Bio/Life: Cell Biology
 Bio/Life: Cell Biology 1a The fundamental life processes of plants and animals depend on a variety of chemical reactions that occur in specialized areas of the organism's cells. As a basis for understanding
Bio/Life: Cell Biology 1a The fundamental life processes of plants and animals depend on a variety of chemical reactions that occur in specialized areas of the organism's cells. As a basis for understanding
3/8/ Complex adaptations. 2. often a novel trait
 Chapter 10 Adaptation: from genes to traits p. 302 10.1 Cascades of Genes (p. 304) 1. Complex adaptations A. Coexpressed traits selected for a common function, 2. often a novel trait A. not inherited from
Chapter 10 Adaptation: from genes to traits p. 302 10.1 Cascades of Genes (p. 304) 1. Complex adaptations A. Coexpressed traits selected for a common function, 2. often a novel trait A. not inherited from
A complementation test would be done by crossing the haploid strains and scoring the phenotype in the diploids.
 Problem set H answers 1. To study DNA repair mechanisms, geneticists isolated yeast mutants that were sensitive to various types of radiation; for example, mutants that were more sensitive to UV light.
Problem set H answers 1. To study DNA repair mechanisms, geneticists isolated yeast mutants that were sensitive to various types of radiation; for example, mutants that were more sensitive to UV light.
Physiology. Organization of the Body. Assumptions in Physiology. Chapter 1. Physiology is the study of how living organisms function
 Introduction to Physiology and Homeostasis Chapter 1 Physiology Physiology is the study of how living organisms function On the street explanations are in terms of meeting a bodily need Physiologic explanations
Introduction to Physiology and Homeostasis Chapter 1 Physiology Physiology is the study of how living organisms function On the street explanations are in terms of meeting a bodily need Physiologic explanations
Regulation and signaling. Overview. Control of gene expression. Cells need to regulate the amounts of different proteins they express, depending on
 Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
What is the central dogma of biology?
 Bellringer What is the central dogma of biology? A. RNA DNA Protein B. DNA Protein Gene C. DNA Gene RNA D. DNA RNA Protein Review of DNA processes Replication (7.1) Transcription(7.2) Translation(7.3)
Bellringer What is the central dogma of biology? A. RNA DNA Protein B. DNA Protein Gene C. DNA Gene RNA D. DNA RNA Protein Review of DNA processes Replication (7.1) Transcription(7.2) Translation(7.3)
CAPE Biology Unit 1 Scheme of Work
 CAPE Biology Unit 1 Scheme of Work 2011-2012 Term 1 DATE SYLLABUS OBJECTIVES TEXT PAGES ASSIGNMENTS COMMENTS Orientation Introduction to CAPE Biology syllabus content and structure of the exam Week 05-09
CAPE Biology Unit 1 Scheme of Work 2011-2012 Term 1 DATE SYLLABUS OBJECTIVES TEXT PAGES ASSIGNMENTS COMMENTS Orientation Introduction to CAPE Biology syllabus content and structure of the exam Week 05-09
GENE REGULATION AND PROBLEMS OF DEVELOPMENT
 GENE REGULATION AND PROBLEMS OF DEVELOPMENT By Surinder Kaur DIET Ropar Surinder_1998@ yahoo.in Mob No 9988530775 GENE REGULATION Gene is a segment of DNA that codes for a unit of function (polypeptide,
GENE REGULATION AND PROBLEMS OF DEVELOPMENT By Surinder Kaur DIET Ropar Surinder_1998@ yahoo.in Mob No 9988530775 GENE REGULATION Gene is a segment of DNA that codes for a unit of function (polypeptide,
Supplemental material
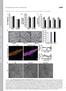 Supplemental material THE JOURNAL OF CELL BIOLOGY Mourier et al., http://www.jcb.org/cgi/content/full/jcb.201411100/dc1 Figure S1. Size and mitochondrial content in Mfn1 and Mfn2 knockout hearts. (A) Body
Supplemental material THE JOURNAL OF CELL BIOLOGY Mourier et al., http://www.jcb.org/cgi/content/full/jcb.201411100/dc1 Figure S1. Size and mitochondrial content in Mfn1 and Mfn2 knockout hearts. (A) Body
Frequently Asked Questions (FAQs)
 Frequently Asked Questions (FAQs) Q1. What is meant by Satellite and Repetitive DNA? Ans: Satellite and repetitive DNA generally refers to DNA whose base sequence is repeated many times throughout the
Frequently Asked Questions (FAQs) Q1. What is meant by Satellite and Repetitive DNA? Ans: Satellite and repetitive DNA generally refers to DNA whose base sequence is repeated many times throughout the
Genetic transcription and regulation
 Genetic transcription and regulation Central dogma of biology DNA codes for DNA DNA codes for RNA RNA codes for proteins not surprisingly, many points for regulation of the process DNA codes for DNA replication
Genetic transcription and regulation Central dogma of biology DNA codes for DNA DNA codes for RNA RNA codes for proteins not surprisingly, many points for regulation of the process DNA codes for DNA replication
Bioinformatics 2. Yeast two hybrid. Proteomics. Proteomics
 GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
RNA Infrastructure and Networks
 RNA Infrastructure and Networks Edited by Lesley J. Collins Institute of Fundamental Sciences, Massey University, Palmerston North, New Zealand volume e contact Springer Science+Business Media, LLC Landes
RNA Infrastructure and Networks Edited by Lesley J. Collins Institute of Fundamental Sciences, Massey University, Palmerston North, New Zealand volume e contact Springer Science+Business Media, LLC Landes
JEFFERSON COLLEGE VERTEBRATE ANATOMY
 JEFFERSON COLLEGE COURSE SYLLABUS BIO207 VERTEBRATE ANATOMY 4 Credit Hours Prepared by: Mr. Jim McCain Revised Date: November 2005 by Dr. Ken Balak Division of Arts & Science Education Dr. Mindy Selsor,
JEFFERSON COLLEGE COURSE SYLLABUS BIO207 VERTEBRATE ANATOMY 4 Credit Hours Prepared by: Mr. Jim McCain Revised Date: November 2005 by Dr. Ken Balak Division of Arts & Science Education Dr. Mindy Selsor,
Measuring TF-DNA interactions
 Measuring TF-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Regulation of Gene Expression by Transcription Factors TF trans-acting factors TF TF TF TF
Measuring TF-DNA interactions How is Biological Complexity Achieved? Mediated by Transcription Factors (TFs) 2 Regulation of Gene Expression by Transcription Factors TF trans-acting factors TF TF TF TF
Inferring Protein-Signaling Networks
 Inferring Protein-Signaling Networks Lectures 14 Nov 14, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20 Johnson Hall (JHN) 022 1
Inferring Protein-Signaling Networks Lectures 14 Nov 14, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20 Johnson Hall (JHN) 022 1
Co-ordination occurs in multiple layers Intracellular regulation: self-regulation Intercellular regulation: coordinated cell signalling e.g.
 Gene Expression- Overview Differentiating cells Achieved through changes in gene expression All cells contain the same whole genome A typical differentiated cell only expresses ~50% of its total gene Overview
Gene Expression- Overview Differentiating cells Achieved through changes in gene expression All cells contain the same whole genome A typical differentiated cell only expresses ~50% of its total gene Overview
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/345/ra91/dc1 Supplementary Materials for TGF-β induced epithelial-to-mesenchymal transition proceeds through stepwise activation of multiple feedback loops Jingyu
www.sciencesignaling.org/cgi/content/full/7/345/ra91/dc1 Supplementary Materials for TGF-β induced epithelial-to-mesenchymal transition proceeds through stepwise activation of multiple feedback loops Jingyu
Variation in the genetic response to high temperature in Montastraea faveolata from the Florida Keys & Mexico
 Variation in the genetic response to high temperature in Montastraea faveolata from the Florida Keys & Mexico Nicholas R. Polato 1, Christian R. Voolstra 2, Julia Schnetzer 3, Michael K. DeSalvo 4, Carly
Variation in the genetic response to high temperature in Montastraea faveolata from the Florida Keys & Mexico Nicholas R. Polato 1, Christian R. Voolstra 2, Julia Schnetzer 3, Michael K. DeSalvo 4, Carly
