Characterization of the rice NLA family reveals a key role for OsNLA1 in phosphate homeostasis
|
|
|
- Harry Lawrence
- 6 years ago
- Views:
Transcription
1 Yang et al. Rice (2017) 10:52 DOI /s y SHORT COMMUNICATION Open Access Characterization of the rice NLA family reveals a key role for OsNLA1 in phosphate homeostasis Jian Yang 1, Lan Wang 1,2, Chuanzao Mao 3 and Honghui Lin 1* Abstract Background: Phosphate (Pi), an essential mineral nutrient for plant development and reproduction, is one of the main components of fertilizers in modern agriculture. Previous research demonstrated that AtNLA1 mediates ubiquitination of Pi transporters in the plasma membrane and triggers their endocytosis and degradation in Arabidopsis. In this study, we researched the function of NLA homologous proteins in Pi homeostasis in rice. Findings: Two OsNLA homologs from rice (Oryza sativa L.) were identified by bioinformatics and phylogenetic analysis and designated OsNLA1 and OsNLA2. TheOsNLA1 clustered with Arabidopsis AtNLA1, was expressed higher than OsNLA2 and was transcriptionally repressed under Pi-deficient condition. Loss-of-function of OsNLA1 caused P overaccumulation and growth inhibitions in both root and shoot under Pi-sufficient condition. Furthermore, mutation of OsNLA1 affected expression of Pi tranporters and root hair development under Pi-sufficient and/or Pi-deficient conditions. Conclusions: OsNLA1 plays a key role in maintaining phosphate homeostasis in rice. Keywords: Rice, Phosphate, OsNLA1, Pi-homeostasis Findings Phosphorus (P) is a mineral nutrient essential for plant development and reproduction, and is integral to several macromolecules such as phospholipids and nucleic acids. Despite the indispensable role of P for plants, levels of phosphate (orthophosphate; Pi), the only form of P that can be taken up by plants, are commonly limited because of chemical fixation and microbial activity (Raghothama, 1999). To cope with suboptimal Pi conditions, plants have developed a series of adaptive responses, such as induction of Pi transporters and modification of root system architecture (Raghothama, 1999; Lin et al., 2009; Wu et al., 2013). Plant uptake of Pi is largely mediated by plasma membrane -localized Pi transporters belonging to the PHOSPHATE TRANSPORTER1 (PT) symporter family. Thirteen PT genes have been identified in rice (Oryza sativa) and nine in Arabidopsis thaliana (Goff et al., 2002; Karthikeyan et al., 2002). OsPTs differ in tissue expression * Correspondence: hhlin@scu.edu.cn 1 Ministry of Education Key Laboratory for Bio-Resource and Eco-Environment, College of Life Science, Sichuan University, Chengdu , China Full list of author information is available at the end of the article patterns and affinities for Pi, resulting in diverse functions in plants. For instance, the high-affinity Pi transporter OsPT8 is universally expressed in rice, and is responsible for half of its Pi uptake (Chen et al., 2011; Jia et al., 2011). Although most OsPTs in rice are induced at the transcriptional level by Pi starvation or mycorrhizal symbiosis (Yang et al., 2012; Secco et al., 2013), post-transcriptional regulating of OsPT family proteins is also important to their activities (Gonzalez et al., 2005; Bayle et al., 2011; Chen et al., 2011; Chen et al., 2015). NITROGEN LIMITATION ADAPTATION (NLA), designated AtNLA1 in this study, was first identified as a positive regulator for the adaptability of Arabidopsis to nitrogen limitation (Peng et al., 2007), and later analysis of Pi concentration revealed that the early senescence phenotype of atnla mutant plants was due to Pi toxicity (Kant et al., 2011). In Arabidopsis, AtNLA1 can interact with AtPTs members via its SPX domain, and mediate ubiquitination of AtPTs in the plasma membrane and trigger their endocytosis and degradation (Lin et al., 2013; Park et al., 2014). Recently, two research groups separately reported roles of OsNLA1 in mantaining Pi homeostasis in rice (Yue et al., The Author(s) Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License ( which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
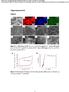
2 Yang et al. Rice (2017) 10:52 Page 2 of ; Zhong et al., 2017). Yue et al., (2017) additionally reported OsNLA1 functioned as a ubiquitin ligase to degrade Pi transporters in rice, with a similar function of AtNLA1 in Arabidopsis.. In this study, we were interested in the phylogenetic relationship of the NLA family and expression of OsPTs and root hair development in osnla1 mutant. An unrooted phylogenetic analysis of the NLA family proteins with four monocots (B. distachyon, S. bicolor, S. italica and rice) and five dicots (grapevine, soybean, apple, M. truncatula and Arabidopsis), revealed the presence of two distinct clades. Although all plants had proteins belonging to clade I, in which AtNLA1 involved in regulating Pi homeostasis was present (Kant et al., 2011), all monocots and only some dicots had NLA members belonging to clade II (Fig. 1a). This suggested that NLA members of clade I conservatively functioned in maintaining Pi homeostasis among different plant species. Quantitative reverse-transcription PCR (qrt-pcr) was performed on different tissues for rice plants grown in nutrient solutions under Pi- sufficient (300 μm) condition (Additional file 1). The amplification efficiencies of gene-specific primers for OsNLA1 and OsNLA2 were assessed and found that they were approximately equal (95.5% for OsNLA1 and 94.3% for OsNLA2) (Additional file 2: Figure S1 and Additional file 3: Table S1). Then, the transcript level of OsNLA1 and OsNLA2 in plants were compared, and found that the transcript level of OsNLA1 was higher than that of OsNLA2 in all tissues tested, with about 1.5-fold in shoot base, 4.5-fold in root and 80-fold in leaf sheath (Fig. 1b and c). Previous transcriptome analysis also shown that OsNLA1 abounace was higher than OsNLA2 in both root and shoot (Secco et al., 2013). Furthermore, the OsNLA1 transcript was differentially regulated by Pi availability, with higher expression in Pi- sufficient and lower expression in Pi -deficient conditions (Fig. 1d), as AtNLA1 in Arabidopsis (Lin et al., 2013); however, the OsNLA2 transcripts remained at relatively constant levels (Fig. 1e). The transcriptional change of OsNLA1 in response to Pi supply was also identified by RNA sequencing (Secco et al., 2013). Based on phylogenetic relationships and expression levels and responses to Pi starvation of the NLA family in rice, we suggest that OsNLA1 might plays a major role in regulating Pi homeostasis, as does AtNLA1 in Arabidopsis. To characterize the functions of the NLA gene family in rice, we searched for publicly available mutants in different rice genomic resources. One T-DNA null mutant in OsNLA1 gene (PFG_1B-12,301) was obtained from RISD DB (Rice T-DNA Insertion Sequence Database) (Fig. 2a and b). After growth for 30 d under Pi-sufficient condition (300 μm Pi; +P), the shoots and roots of osnla1 were inhibited compared with wild-type (WT) plants (Fig. 2c and d). In addition, osnla1 displayed leaf tip necrosis on old leaves, which was a typical Pi toxicity symptom in rice (Fig. 2c, Additional file 2: Figure S1). This symptom in osnla1 was not observed when grown in Pideficient condition (10 μm Pi; -P). Moreover, the inhibited root phenotype of Osnla1 was reversed when grown in Pi-deficient condition (Fig. 2c and d). Total P concentrations in all tissues of osnla1 were higher than of WT, with 1.24-fold in roots and fold in leaves under Pi-sufficient condition (Fig. 2e). This indicating that OsNLA1 played a key role in Pi uptake in rice, as previously reported (Yue et al., 2017; Zhong et al., 2017). However, under Pi-deficient condition, total P concentrations in old leaves (leaves 2 and 3) of osnla1 were decreased by 17 25%, while total P concentrations in youngest leaves (leaves 7) of osnla1 were increased by 21% compared with WT. The total P distribution rate in WT plants grown in Pi-deficient condition was 1.68-fold higher than that in plants grown in Pi-sufficient condition. However, the rate in osnla1 mutants grown in Pi-deficient condition was 3.19-fold higher than that in plants grown in Pi-sufficient condition (Fig. 2f). This significant increased total P distribution rate under Pi limiting condition sustained the newly leaves growth (Fig. 2c and d). Pi is the major form of P transported within the plants, and old leaves Pi pool would be the source of Pi in regard to young leaves under Pi-deficient condition, resulting in higher P concentration in young leaves (Li et al., 2015, b). Thus, these results indicated that OsNLA1 was also involved in Pi remobilization besides Pi uptake. This was expected because OsPT1 and OsPT8 also function in the redistribution of Pi from source to sink organs (Sun et al., 2012; Li et al., 2015, b). In a recent study, AtNLA1 was also involved in mediating degradation of NRT1.7 and further remobilizing nitrate from source to sink in Arabidopsis (Liu et al., 2016). Whether OsNLA1 also functions in nitrate remobilization need further studies. As plants may modify root system architecture when growth in suboptimal Pi condition, we analyzed root hairs when osnla1 and WT were grown in Pi-sufficient and -deficient conditions. After Pi-deficient growth for 10 d, WT developed many root hairs (Fig. 3a and b), as previously reported (Zhou et al., 2008; Sun et al., 2012). However, the length of root hairs on osnla1 mutant was 1.5-fold those of WT plants grown in Pi-deficient condition. Furthermore, osnla1 mutant also had increased root hairs length under Pi-sufficient condition compared with WT plants. Since OsNLA1 could mediate
3 Yang et al. Rice (2017) 10:52 Page 3 of 6 Fig. 1 Phylogenetic relationships of NLA proteins and OsNLA family expression patterns in rice. a Unrooted phylogenetic tree of the NLA family using MEGA 5.10 by the neighbor-joining method. Dicots: Vitis vinifera (GSVIV), Glycine max (Glyma), Malus domestica (MDP), Medicago truncatula (Medtr), Arabidopsis thaliana (At); monocots: Brachypodium distachyon (Bradi), Sorghum bicolor (Sobic) and Setaria italica (Si). OsNLA family expression patterns in rice. b, c Spatial expression of OsNLAs transcripts. Transcript levels for OsNLA1 (b) and OsNLA2 (c)in leaf sheath, leaf blade, roots and shoot base in 30-d-old seedlings grown in nutrient solutions containing 300 μm Pi and in panicle before flowering. Expression of OsNLA1 and OsNLA2 is relative to OsACTIN2. d, e Transcript levels of OsNLA1 (d) and OsNLA2 (e) in root and shoot of plants grown for 10 d in nutrient solutions with different Pi concentrations. Data represent mean ± SD of three replicates. Different letters represent significant differences according to Duncan s multiple range test (P < 0.05) the degradation of OsPT2 and OsPT8 (Yue et al., 2017), inhibition of root growth and induce of root hair in osnla1 was expected because OsPTs were involved in regulating root growth and root hair development (Jia et al., 2011; Sun et al., 2012). Since changing the expression of OsPT4 or OsPT8 affects the expression of other Pi transporters in rice (Jia et al., 2011; Sun et al., 2012; Li et al., 2015, b) and protein levels of OsPT2 and OsPT8 accumulated in osnla1 mutants (Yue et al., 2017), we then analyzed the transcriptional levels of OsPTs following WT and osnla1 mutant growth in Pi-sufficient and -deficient conditions for 10d. In the shoot of osnla1 mutant, transcripts of most of Pi transporters were induced (Fig. 3c). Compared with the WT, expressions of OsPT6 and OsPT8 were greatly induced under both Pi-sufficient and -deficient conditions. Although OsPT2, OsPT4 and OsPT10 were also upregulated under Pi-sufficient condition, their transcript levels did not change under Pi-deficient condition. Expression of OsPT1 was induced only when osnla1 mutant was
4 Yang et al. Rice (2017) 10:52 Page 4 of 6 Fig. 2 Phenotypical characteristics of osnla1 mutant. a Genomic structure of rice OsNLA1. Position of the T-DNA insertion in OsNLA1 is indicated by a triangle. Small arrows are the gene-specific primers for RT-PCR. b RT-PCR analysis of OsNLA1 expression in roots of the mutant and wild-type (Dongjin; WT). c Phenotype comparison between WT and osnla1 mutant. 30-d-old WT and osnla1 grown under Pi-sufficient (300 μm; +P; upper) and Pi-deficient (10 μm; -P; lower) conditions. Bars = 5 cm. d Biomass of 30-d-old WT and osnla1 in (c). Data represent mean ± SD of eight replicates. e Total P concentration in different leaves and roots of 30-d-old WT and osnla1 in (c). Data represent mean ± SD of three replicates. f Distribution ratio of total P between young leaves (leaf 7) and old leaves (leaf 2) in osnla1 and WT. Asterisks represent a significant difference with the corresponding WT (**, P < 0.01; ***, P < 0.001) grown under Pi-deficient condition. However, contrary to our finding, Yue et al. (2017) found that OsPT2 and OsPT8 were unchanged in leaf under Pi-sufficient conditions. This might be resulted from transcriptional levels of OsNLA1 and Pi transporters differed in various tissues (Fig. 1b; Remy et al., 2012). Unlike Pi transporters induced in the shoot, transcripts of Pi transporters were differentially regulated under Pi-sufficient and -deficient conditions in root of osnla1 mutant (Fig. 3c). The transcriptional levels of OsPT1 and OsPT4 were induced under Pi-sufficient condition, but unchanged under Pi-deficient condition. In contrast, OsPT6, OsPT8 and OsPT10 were downregulated under Pi-deficient condition, but unchanged under Pi-sufficient condition. The increased or repressed expression of these Pi transporters was caused, at least in part, by accumulated protein level of Pi transporters in osnla1 mutant, because changing the expression of OsPT4 or OsPT8 affects the expression of Pi transporters in rice (Jia et al., 2011; Zhang et al., 2015). Moreover, induced expression of OsPT1 and OsPT8 in shoot of osnla1 mutant under Pi-deficient condition would further remobilize Pi from old to young leaves (Sun et al., 2012; Li et al., 2015, b).
5 Yang et al. Rice (2017) 10:52 Page 5 of 6 Fig. 3 Root hair proliferation and expression of Pi transporter genes in osnla1 mutant and wild-type (WT). a Root hair proliferation of WT and osnla1 grown on Pi-sufficient (+P; left) and Pi-deficient ( P; right). Bars = 100 μm. b Root hair length in the maturation zone of roots. Data represent mean ± SD of eight replicates. Different letters represent significant differences according to Duncan s multiple range test (P < 0.05). c Expression of Pi transporter genes in osnla1 mutant and WT. Ten-day-old plants grown in Pi-sufficient (300 μm Pi) nutrient solution were transferred to Pi-sufficient (+P) and -deficient (-P) conditions for 10 d. RNA was extracted from shoots (upper) and roots (lower) for qrt-pcr. Data represent mean ± SD of three replicates. Asterisks represent significant difference with the corresponding WT (*, P < 0.05; **, P < 0.01; ***, P < 0.001) Since OsNLA1 mediates degradation of OsPTs and plays a key role in maintaining Pi homeostasis in rice. In this research, we identified OsNLA1 could regulate root system architecture, Pi transporters at the transcriptional levels and Pi redistribution from source to sink organs. These results presented here will provide a novel insight into the function of OsNLA1 in rice. Abbreviations NLA: Nitrogen Limitation Adaptation; Pi: Phosphate; PT: Phosphate transporter; RISD DB: Rice T-DNA Insertion Sequence Database; WT: Wild type Acknowledgements This work was supported by the National Natural Science Foundation of China ( ), Foundation of Sichuan University (2017SCU12007) and National Key Research and Development Program of China (2016YFD ) Additional files Additional file 1: Materials and methods. (DOCX 19 kb) Additional file 2: Figure S1. Calculation of PCR efficiencies. Figure S2. Leaf blades of 30-d-old WT and osnla1 grown under Pi-sufficient (300 μm; +P) and Pi-deficient (10 μm; -P) conditions. (PPTX 304 kb) Additional file 3: Table S1. Primers used in this study. (DOCX 14 kb) Authors Contributions HHL, CZM and JY designed the experiments. JY developed relevant research materials and performed the experiments together with LW. HHL, CZM and JY wrote the manuscript. All authors read and approved the final manuscript. Competing Interests The authors declare that they have no competing interests.
6 Yang et al. Rice (2017) 10:52 Page 6 of 6 Publisher s Note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations. Author details 1 Ministry of Education Key Laboratory for Bio-Resource and Eco-Environment, College of Life Science, Sichuan University, Chengdu , China. 2 Biogas Institute of Ministry of Agriculture, Chengdu , China. 3 State Key Laboratory of Plant Physiology and Biochemistry, College of Life Sciences, Zhejiang University, Hangzhou , China. Received: 15 July 2017 Accepted: 12 December 2017 References BayleV,ArrighiJF,CreffA,NespoulousC,VialaretJ,RossignolM,GonzalezE, Paz-Ares J, Nussaume L (2011) Arabidopsis thaliana high-affinity phosphate transporters exhibit multiple levels of posttranslational regulation. Plant Cell 23: Chen J, Liu Y, Ni J, Wang Y, Bai Y, Shi J, Gan J, Wu Z, Wu P (2011) OsPHF1 regulates the plasma membrane localization of low- and high-affinity inorganic phosphate transporters and determines inorganic phosphate uptake and translocation in rice. Plant Physiol 157: Chen J, Wang Y, Wang F, Yang J, Gao M, Li C, Liu Y, Yamaji N, Ma JF, Paz-Ares J, Nussaume L, Zhang S, Yi K, Wu Z, Wu P (2015) The rice CK2 kinase regulates trafficking of phosphate transporters in response to phosphate levels. Plant Cell 27: Goff SA, Ricke D, Lan T-H, Presting G, Wang R, Dunn M, Glazebrook J, Sessions A, Oeller P, Varma H, Hadley D, Hutchison D, Martin C, Katagiri F, Lange BM, Moughamer T, Xia Y, Budworth P, Zhong J, Miguel T, Paszkowski U, Zhang S, Colbert M, W-l S, Chen L, Cooper B, Park S, Wood TC, Mao L, Quail P, Wing R, Dean R, Yu Y, Zharkikh A, Shen R, Sahasrabudhe S, Thomas A, Cannings R, Gutin A, Pruss D, Reid J, Tavtigian S, Mitchell J, Eldredge G, Scholl T, Miller RM, Bhatnagar S, Adey N, Rubano T, Tusneem N, Robinson R, Feldhaus J, Macalma T, Oliphant A, Briggs S (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. japonica). Science 296: Gonzalez E, Solano R, Rubio V, Leyva A, Paz-Ares J (2005) PHOSPHATE TRANSPORTER TRAFFIC FACILITATOR1 is a plant-specific SEC12-related protein that enables the endoplasmic reticulum exit of a high-affinity phosphate transporter in Arabidopsis. Plant Cell 17: Jia H, Ren H, Gu M, Zhao J, Sun S, Zhang X, Chen J, Wu P, Xu G (2011) The phosphate transporter gene OsPht1;8 is involved in phosphate homeostasis in rice. Plant Physiol 156: Kant S, Peng M, Rothstein SJ (2011) Genetic regulation by NLA and microrna827 for maintaining nitrate-dependent phosphate homeostasis in Arabidopsis. PLoS Genet 7:e Karthikeyan AS, Varadarajan DK, Mukatira UT, D'Urzo MP, Damsz B, Raghothama KG (2002) Regulated expression of Arabidopsis phosphate transporters. Plant Physiol 130: Li XM, Chao DY, Wu Y, Huang X, Chen K, Cui LG, Su L, Ye WW, Chen H, Chen HC, Dong NQ, Guo T, Shi M, Feng Q, Zhang P, Han B, Shan JX, Gao JP, Lin HX (2015) Natural alleles of a proteasome alpha2 subunit gene contribute to thermotolerance and adaptation of African rice. Nat Genet 47: Li Y, Zhang J, Zhang X, Fan H, Gu M, Qu H, Xu G (2015) Phosphate transporter OsPht1;8 in rice plays an important role in phosphorus redistribution from source to sink organs and allocation between embryo and endosperm of seeds. Plant Sci 230:23 32 Lin WY, Huang TK, Chiou TJ (2013) NITROGEN LIMITATION ADAPTATION, a target of MicroRNA827, mediates degradation of plasma membrane-localized phosphate transporters to maintain phosphate homeostasis in Arabidopsis. Plant Cell 25: Lin WY, Lin SI, Chiou TJ (2009) Molecular regulators of phosphate homeostasis in plants. J Exp Bot 60: Liu W, Sun Q, Wang K, Du Q, Li WX (2016) Nitrogen Limitation Adaptation (NLA) is involved in source-to-sink remobilization of nitrate by mediating the degradation of NRT1.7 in Arabidopsis. New Phytol 214: Park BS, Seo JS, Chua NH (2014) NITROGEN LIMITATION ADAPTATION recruits PHOSPHATE2 to target the phosphate transporter PT2 for degradation during the regulation of Arabidopsis phosphate homeostasis. Plant Cell 26: Peng M, Hannam C, Gu H, Bi YM, Rothstein SJ (2007) A mutation in NLA, which encodes a RING-type ubiquitin ligase, disrupts the adaptability of Arabidopsis to nitrogen limitation. Plant J 50: Raghothama KG (1999) Phosphate acquisition. Annu Rev Plant Biol 50: Remy E, Cabrito TR, Batista RA, Teixeira MC, Sa-Correia I, Duque P (2012) The Pht1;9 and Pht1;8 transporters mediate inorganic phosphate acquisition by the Arabidopsis thaliana root during phosphorus starvation. New Phytol 195: Secco D, Jabnoune M, Walker H, Shou H, Wu P, Poirier Y, Whelan J (2013) Spatiotemporal transcript profiling of rice roots and shoots in response to phosphate starvation and recovery. Plant Cell 25: Sun S, Gu M, Cao Y, Huang X, Zhang X, Ai P, Zhao J, Fan X, Xu G (2012) A constitutive expressed phosphate transporter, OsPht1;1, modulates phosphate uptake and translocation in phosphate-replete rice. Plant Physiol 159: Wu P, Shou H, Xu G, Lian X (2013) Improvement of phosphorus efficiency in rice on the basis of understanding phosphate signaling and homeostasis. Curr Opin Plant Biol 16: Yang SY, Gronlund M, Jakobsen I, Grotemeyer MS, Rentsch D, Miyao A, Hirochika H, Kumar CS, Sundaresan V, Salamin N, Catausan S, Mattes N, Heuer S, Paszkowski U (2012) Nonredundant regulation of rice arbuscular mycorrhizal symbiosis by two members of the phosphate transporter1 gene family. Plant Cell 24: Yue W, Ying Y, Wang C, Zhao Y, Dong C, Whelan J, Shou H (2017) OsNLA1, a RING-type ubiquitin ligase, maintains phosphate homeostasis in Oryza sativa via degradation of phosphate transporters. Plant J 90: Zhang F, Sun Y, Pei W, Jain A, Sun R, Cao Y, Wu X, Jiang T, Zhang L, Fan X, Chen A, Xu G, Sun S (2015) Involvement of OsPht1;4 in phosphate acquisition and mobilization facilitates embryo development in rice. Plant J 82: Zhong S, Mahmood K, Bi YM, Rothstein SJ, Ranathunge K (2017) Altered expression of OsNLA1 modulates Pi accumulation in rice (Oryza sativa L.) plants. Front Plant Sci 8: 928 Zhou J, Jiao F, Wu Z, Li Y, Wang X, He X, Zhong W, Wu P (2008) OsPHR2 is involved in phosphate-starvation signaling and excessive phosphate accumulation in shoots of plants. Plant Physiol 146:
Regulation of Phosphate Homeostasis by microrna in Plants
 Regulation of Phosphate Homeostasis by microrna in Plants Tzyy-Jen Chiou 1 *, Kyaw Aung 1,2, Shu-I Lin 1,3, Chia-Chune Wu 1, Su-Fen Chiang 1, and Chun-Lin Su 1 Abstract Upon phosphate (Pi) starvation,
Regulation of Phosphate Homeostasis by microrna in Plants Tzyy-Jen Chiou 1 *, Kyaw Aung 1,2, Shu-I Lin 1,3, Chia-Chune Wu 1, Su-Fen Chiang 1, and Chun-Lin Su 1 Abstract Upon phosphate (Pi) starvation,
Small RNA in rice genome
 Vol. 45 No. 5 SCIENCE IN CHINA (Series C) October 2002 Small RNA in rice genome WANG Kai ( 1, ZHU Xiaopeng ( 2, ZHONG Lan ( 1,3 & CHEN Runsheng ( 1,2 1. Beijing Genomics Institute/Center of Genomics and
Vol. 45 No. 5 SCIENCE IN CHINA (Series C) October 2002 Small RNA in rice genome WANG Kai ( 1, ZHU Xiaopeng ( 2, ZHONG Lan ( 1,3 & CHEN Runsheng ( 1,2 1. Beijing Genomics Institute/Center of Genomics and
Supplemental Data. Yang et al. (2012). Plant Cell /tpc
 Supplemental Figure 1. Mature flowers of P. heterotricha. (A) An inflorescence of P. heterotricha showing the front view of a zygomorphic flower characterized by two small dorsal petals and only two fertile
Supplemental Figure 1. Mature flowers of P. heterotricha. (A) An inflorescence of P. heterotricha showing the front view of a zygomorphic flower characterized by two small dorsal petals and only two fertile
Nature Genetics: doi: /ng Supplementary Figure 1. The phenotypes of PI , BR121, and Harosoy under short-day conditions.
 Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
GENETIC ANALYSES OF ROOT SYSTEM DEVELOPMENT IN THE TOMATO CROP MODEL
 GENETIC ANALYSES OF ROOT SYSTEM DEVELOPMENT IN THE TOMATO CROP MODEL Kelsey Hoth 1 Dr. Maria Ivanchenko 2 Bioresourse Research 1, Department of Botany and Plant Physiology 2, Oregon State University, Corvallis,
GENETIC ANALYSES OF ROOT SYSTEM DEVELOPMENT IN THE TOMATO CROP MODEL Kelsey Hoth 1 Dr. Maria Ivanchenko 2 Bioresourse Research 1, Department of Botany and Plant Physiology 2, Oregon State University, Corvallis,
POTASSIUM IN PLANT GROWTH AND YIELD. by Ismail Cakmak Sabanci University Istanbul, Turkey
 POTASSIUM IN PLANT GROWTH AND YIELD by Ismail Cakmak Sabanci University Istanbul, Turkey Low K High K High K Low K Low K High K Low K High K Control K Deficiency Cakmak et al., 1994, J. Experimental Bot.
POTASSIUM IN PLANT GROWTH AND YIELD by Ismail Cakmak Sabanci University Istanbul, Turkey Low K High K High K Low K Low K High K Low K High K Control K Deficiency Cakmak et al., 1994, J. Experimental Bot.
23-. Shoot and root development depend on ratio of IAA/CK
 Balance of Hormones regulate growth and development Environmental factors regulate hormone levels light- e.g. phototropism gravity- e.g. gravitropism temperature Mode of action of each hormone 1. Signal
Balance of Hormones regulate growth and development Environmental factors regulate hormone levels light- e.g. phototropism gravity- e.g. gravitropism temperature Mode of action of each hormone 1. Signal
Utilizing Illumina high-throughput sequencing technology to gain insights into small RNA biogenesis and function
 Utilizing Illumina high-throughput sequencing technology to gain insights into small RNA biogenesis and function Brian D. Gregory Department of Biology Penn Genome Frontiers Institute University of Pennsylvania
Utilizing Illumina high-throughput sequencing technology to gain insights into small RNA biogenesis and function Brian D. Gregory Department of Biology Penn Genome Frontiers Institute University of Pennsylvania
Ethylene is critical to the maintenance of primary root growth and Fe. homeostasis under Fe stress in Arabidopsis
 Ethylene is critical to the maintenance of primary root growth and Fe homeostasis under Fe stress in Arabidopsis Guangjie Li, Weifeng Xu, Herbert J. Kronzucker, Weiming Shi * Supplementary Data Supplementary
Ethylene is critical to the maintenance of primary root growth and Fe homeostasis under Fe stress in Arabidopsis Guangjie Li, Weifeng Xu, Herbert J. Kronzucker, Weiming Shi * Supplementary Data Supplementary
The Role of Inorganic Carbon Transport and Accumulation in the CO 2 -Concentrating Mechanism and CO 2 Assimilation in Chlamydomonas
 The Role of Inorganic Carbon Transport and Accumulation in the CO 2 -Concentrating Mechanism and CO 2 Assimilation in Chlamydomonas Is there a Role for the CCM in Increasing Biological CO 2 Capture? Generalized
The Role of Inorganic Carbon Transport and Accumulation in the CO 2 -Concentrating Mechanism and CO 2 Assimilation in Chlamydomonas Is there a Role for the CCM in Increasing Biological CO 2 Capture? Generalized
The Science of Plants in Agriculture Pl.Sci 102. Getting to Know Plants
 The Science of Plants in Agriculture Pl.Sci 102 Getting to Know Plants Growth and Development of Plants Growth and Development of Plants Why it s important to have knowledge about plant development. What
The Science of Plants in Agriculture Pl.Sci 102 Getting to Know Plants Growth and Development of Plants Growth and Development of Plants Why it s important to have knowledge about plant development. What
Cytokinin Interaction to Cope with Phosphorous Starvation in Rice (Oryza sativa)
 INTERNATIONAL JOURNAL OF AGRICULTURE & BIOLOGY ISSN Print: 1560 8530; ISSN Online: 1814 9596 18 0233/201x/00 0 000 000 DOI: 10.17957/IJAB/15.0789 http://www.fspublishers.org Full Length Article Cytokinin
INTERNATIONAL JOURNAL OF AGRICULTURE & BIOLOGY ISSN Print: 1560 8530; ISSN Online: 1814 9596 18 0233/201x/00 0 000 000 DOI: 10.17957/IJAB/15.0789 http://www.fspublishers.org Full Length Article Cytokinin
GFP GAL bp 3964 bp
 Supplemental Data. Møller et al. (2009) Shoot Na + exclusion and increased salinity tolerance engineered by cell type-specific alteration of Na + transport in Arabidopsis Supplemental Figure 1. Salt-sensitive
Supplemental Data. Møller et al. (2009) Shoot Na + exclusion and increased salinity tolerance engineered by cell type-specific alteration of Na + transport in Arabidopsis Supplemental Figure 1. Salt-sensitive
Supplemental Data. Chen and Thelen (2010). Plant Cell /tpc
 Supplemental Data. Chen and Thelen (2010). Plant Cell 10.1105/tpc.109.071837 1 C Total 5 kg 20 kg 100 kg Transmission Image 100 kg soluble pdtpi-gfp Plastid (PDH-alpha) Mito (PDH-alpha) GFP Image vector
Supplemental Data. Chen and Thelen (2010). Plant Cell 10.1105/tpc.109.071837 1 C Total 5 kg 20 kg 100 kg Transmission Image 100 kg soluble pdtpi-gfp Plastid (PDH-alpha) Mito (PDH-alpha) GFP Image vector
Genome-wide Identification of Lineage Specific Genes in Arabidopsis, Oryza and Populus
 Genome-wide Identification of Lineage Specific Genes in Arabidopsis, Oryza and Populus Xiaohan Yang Sara Jawdy Timothy Tschaplinski Gerald Tuskan Environmental Sciences Division Oak Ridge National Laboratory
Genome-wide Identification of Lineage Specific Genes in Arabidopsis, Oryza and Populus Xiaohan Yang Sara Jawdy Timothy Tschaplinski Gerald Tuskan Environmental Sciences Division Oak Ridge National Laboratory
Comparative studies on sequence characteristics around translation initiation codon in four eukaryotes
 Indian Academy of Sciences RESEARCH NOTE Comparative studies on sequence characteristics around translation initiation codon in four eukaryotes QINGPO LIU and QINGZHONG XUE* Department of Agronomy, College
Indian Academy of Sciences RESEARCH NOTE Comparative studies on sequence characteristics around translation initiation codon in four eukaryotes QINGPO LIU and QINGZHONG XUE* Department of Agronomy, College
Penghui Li, Beibei Chen, Gaoyang Zhang, Longxiang Chen, Qiang Dong, Jiangqi Wen, Kirankumar S. Mysore and Jian Zhao
 New Phytologist Supporting Information Regulation of anthocyanin and proanthocyanidin biosynthesis by Medicago truncatula bhlh transcription factor MtTT8 Penghui Li, Beibei Chen, Gaoyang Zhang, Longxiang
New Phytologist Supporting Information Regulation of anthocyanin and proanthocyanidin biosynthesis by Medicago truncatula bhlh transcription factor MtTT8 Penghui Li, Beibei Chen, Gaoyang Zhang, Longxiang
. Supplementary Information
 . Supplementary Information Supplementary Figure S1. Mature embryo sac observations. Supplementary Figure S2. STT observations. Supplementary Figure S3. Comparison of the PTB1 cdna with that of the mutant.
. Supplementary Information Supplementary Figure S1. Mature embryo sac observations. Supplementary Figure S2. STT observations. Supplementary Figure S3. Comparison of the PTB1 cdna with that of the mutant.
Supplemental Data. Perea-Resa et al. Plant Cell. (2012) /tpc
 Supplemental Data. Perea-Resa et al. Plant Cell. (22)..5/tpc.2.3697 Sm Sm2 Supplemental Figure. Sequence alignment of Arabidopsis LSM proteins. Alignment of the eleven Arabidopsis LSM proteins. Sm and
Supplemental Data. Perea-Resa et al. Plant Cell. (22)..5/tpc.2.3697 Sm Sm2 Supplemental Figure. Sequence alignment of Arabidopsis LSM proteins. Alignment of the eleven Arabidopsis LSM proteins. Sm and
** * * * Col-0 cau1 CAU1. Actin2 CAS. Actin2. Supplemental Figure 1. CAU1 affects calcium accumulation.
 Ca 2+ ug g -1 DW Ca 2+ ug g -1 DW Ca 2+ ug g -1 DW Supplemental Data. Fu et al. Plant Cell. (213). 1.115/tpc.113.113886 A 5 4 3 * Col- cau1 B 4 3 2 Col- cau1 ** * * ** C 2 1 25 2 15 1 5 Shoots Roots *
Ca 2+ ug g -1 DW Ca 2+ ug g -1 DW Ca 2+ ug g -1 DW Supplemental Data. Fu et al. Plant Cell. (213). 1.115/tpc.113.113886 A 5 4 3 * Col- cau1 B 4 3 2 Col- cau1 ** * * ** C 2 1 25 2 15 1 5 Shoots Roots *
Supporting material. Figures
 Electronic Supplementary Material (ESI) for New Journal of Chemistry. This journal is The Royal Society of Chemistry and the Centre National de la Recherche Scientifique 2018 Supporting material Figures
Electronic Supplementary Material (ESI) for New Journal of Chemistry. This journal is The Royal Society of Chemistry and the Centre National de la Recherche Scientifique 2018 Supporting material Figures
Supplemental Figure 1. Comparison of Tiller Bud Formation between the Wild Type and d27. (A) and (B) Longitudinal sections of shoot apex in wild-type
 A B 2 3 3 2 1 1 Supplemental Figure 1. Comparison of Tiller Bud Formation between the Wild Type and d27. (A) and (B) Longitudinal sections of shoot apex in wild-type (A) and d27 (B) seedlings at the four
A B 2 3 3 2 1 1 Supplemental Figure 1. Comparison of Tiller Bud Formation between the Wild Type and d27. (A) and (B) Longitudinal sections of shoot apex in wild-type (A) and d27 (B) seedlings at the four
Other funding Sources Agency Name: MSU Agricultural Experiment Station /Project GREEEN Amount requested or awarded: 30,000
 FINAL PROJECT REPORT Project Title: Functional genomics of flowering in apple PI: Herb Aldwinckle Co-PI(2): Steve VanNocker Organization: Cornell University Organization: Michigan State University Telephone/email:
FINAL PROJECT REPORT Project Title: Functional genomics of flowering in apple PI: Herb Aldwinckle Co-PI(2): Steve VanNocker Organization: Cornell University Organization: Michigan State University Telephone/email:
USDA-DOE Plant Feedstock Genomics for Bioenergy
 USDA-DOE Plant Feedstock Genomics for Bioenergy BERAC Thursday, June 7, 2012 Cathy Ronning, DOE-BER Ed Kaleikau, USDA-NIFA Plant Feedstock Genomics for Bioenergy Joint competitive grants program initiated
USDA-DOE Plant Feedstock Genomics for Bioenergy BERAC Thursday, June 7, 2012 Cathy Ronning, DOE-BER Ed Kaleikau, USDA-NIFA Plant Feedstock Genomics for Bioenergy Joint competitive grants program initiated
Heterosis and inbreeding depression of epigenetic Arabidopsis hybrids
 Heterosis and inbreeding depression of epigenetic Arabidopsis hybrids Plant growth conditions The soil was a 1:1 v/v mixture of loamy soil and organic compost. Initial soil water content was determined
Heterosis and inbreeding depression of epigenetic Arabidopsis hybrids Plant growth conditions The soil was a 1:1 v/v mixture of loamy soil and organic compost. Initial soil water content was determined
Supporting Information
 Supporting Information Ultrasensitive Label-Free Resonance Rayleigh Scattering Aptasensor for Hg 2+ Using Hg 2+ -Triggered Exonuclease III-Assisted Target Recycling and Growth of G-Wires for Signal Amplification
Supporting Information Ultrasensitive Label-Free Resonance Rayleigh Scattering Aptasensor for Hg 2+ Using Hg 2+ -Triggered Exonuclease III-Assisted Target Recycling and Growth of G-Wires for Signal Amplification
As negative mycorrhizal growth responses (MGR) have received more experimental attention
 Supplemental Material: Annu. Rev. Plant Biol. 2011. 62:227-250 Supplementary A Negative mycorrhizal responses As negative mycorrhizal growth responses (MGR) have received more experimental attention it
Supplemental Material: Annu. Rev. Plant Biol. 2011. 62:227-250 Supplementary A Negative mycorrhizal responses As negative mycorrhizal growth responses (MGR) have received more experimental attention it
Keystone Exams: Biology Assessment Anchors and Eligible Content. Pennsylvania Department of Education
 Assessment Anchors and Pennsylvania Department of Education www.education.state.pa.us 2010 PENNSYLVANIA DEPARTMENT OF EDUCATION General Introduction to the Keystone Exam Assessment Anchors Introduction
Assessment Anchors and Pennsylvania Department of Education www.education.state.pa.us 2010 PENNSYLVANIA DEPARTMENT OF EDUCATION General Introduction to the Keystone Exam Assessment Anchors Introduction
Supplementary Figure 1 Characterization of wild type (WT) and abci8 mutant in the paddy field.
 Supplementary Figure 1 Characterization of wild type (WT) and abci8 mutant in the paddy field. A, Phenotypes of 30-day old wild-type (WT) and abci8 mutant plants grown in a paddy field under normal sunny
Supplementary Figure 1 Characterization of wild type (WT) and abci8 mutant in the paddy field. A, Phenotypes of 30-day old wild-type (WT) and abci8 mutant plants grown in a paddy field under normal sunny
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION Identification and validation of circrnas Ribo-minus RNA-seq data sets used in identification of circrnas in Oryza sativa and Arabidopsis thaliana were downloaded from NCBI (https://www.ncbi.nlm.nih.gov/).
SUPPLEMENTAL INFORMATION Identification and validation of circrnas Ribo-minus RNA-seq data sets used in identification of circrnas in Oryza sativa and Arabidopsis thaliana were downloaded from NCBI (https://www.ncbi.nlm.nih.gov/).
Supplementary Methods
 Supplementary Methods Microarray analysis Grains of 7 DAP of the wild-type and gif1 were harvested for RNA preparation. Microarray analysis was performed with the Affymetrix (Santa Clara, CA) GeneChip
Supplementary Methods Microarray analysis Grains of 7 DAP of the wild-type and gif1 were harvested for RNA preparation. Microarray analysis was performed with the Affymetrix (Santa Clara, CA) GeneChip
Cytokinin. Fig Cytokinin needed for growth of shoot apical meristem. F Cytokinin stimulates chloroplast development in the dark
 Cytokinin Abundant in young, dividing cells Shoot apical meristem Root apical meristem Synthesized in root tip, developing embryos, young leaves, fruits Transported passively via xylem into shoots from
Cytokinin Abundant in young, dividing cells Shoot apical meristem Root apical meristem Synthesized in root tip, developing embryos, young leaves, fruits Transported passively via xylem into shoots from
Plant Growth and Development
 Plant Growth and Development Concept 26.1 Plants Develop in Response to the Environment Factors involved in regulating plant growth and development: 1. Environmental cues (e.g., day length) 2. Receptors
Plant Growth and Development Concept 26.1 Plants Develop in Response to the Environment Factors involved in regulating plant growth and development: 1. Environmental cues (e.g., day length) 2. Receptors
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/345/ra91/dc1 Supplementary Materials for TGF-β induced epithelial-to-mesenchymal transition proceeds through stepwise activation of multiple feedback loops Jingyu
www.sciencesignaling.org/cgi/content/full/7/345/ra91/dc1 Supplementary Materials for TGF-β induced epithelial-to-mesenchymal transition proceeds through stepwise activation of multiple feedback loops Jingyu
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1. HSP21 expression in 35S:HSP21 and hsp21 knockdown plants. (a) Since no T- DNA insertion line for HSP21 is available in the publicly available T-DNA collections,
Supplementary Figure 1 Supplementary Figure 1. HSP21 expression in 35S:HSP21 and hsp21 knockdown plants. (a) Since no T- DNA insertion line for HSP21 is available in the publicly available T-DNA collections,
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature10534 Supplementary Fig. 1. Diagrammatic representation of the N-end rule pathway of targeted proteolysis (after Graciet and Wellmer 2010 9 ). Tertiary, secondary
SUPPLEMENTARY INFORMATION doi:10.1038/nature10534 Supplementary Fig. 1. Diagrammatic representation of the N-end rule pathway of targeted proteolysis (after Graciet and Wellmer 2010 9 ). Tertiary, secondary
The Plant Cell, November. 2017, American Society of Plant Biologists. All rights reserved
 The Genetics of Floral Development Teaching Guide Overview The development of flowers in angiosperm plants provided a critical evolutionary advantage, allowing more options for pollen dispersal and seed
The Genetics of Floral Development Teaching Guide Overview The development of flowers in angiosperm plants provided a critical evolutionary advantage, allowing more options for pollen dispersal and seed
Activation of MKK9-MPK3/MPK6 enhances phosphate acquisition in Arabidopsis thaliana
 Research Activation of MKK9-MPK3/MPK6 enhances phosphate acquisition in Arabidopsis thaliana Lei Lei, Yuan Li, Qian Wang, Juan Xu, Yifang Chen, Hailian Yang and Dongtao Ren State Key Laboratory of Plant
Research Activation of MKK9-MPK3/MPK6 enhances phosphate acquisition in Arabidopsis thaliana Lei Lei, Yuan Li, Qian Wang, Juan Xu, Yifang Chen, Hailian Yang and Dongtao Ren State Key Laboratory of Plant
PYR1 PYL1 PYL2 PYL3 PYL4 PYL5 PYL6 PYL7 PYL8 PYL9 PYL10 PYL11/12
 Supplemental Data. Gonzalez-Guzman et al. Plant Cell. (212). 1.115/tpc.112.98574 Supplemental Figure 1. Gene expression levels of the PYR/PYL/RCAR ABA receptors in the Arabidopsis transcriptome genomic
Supplemental Data. Gonzalez-Guzman et al. Plant Cell. (212). 1.115/tpc.112.98574 Supplemental Figure 1. Gene expression levels of the PYR/PYL/RCAR ABA receptors in the Arabidopsis transcriptome genomic
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Journal of Materials Chemistry A. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Cation exchange MOF-derived nitrogen-doped
Electronic Supplementary Material (ESI) for Journal of Materials Chemistry A. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Cation exchange MOF-derived nitrogen-doped
Supplemental Data. Perrella et al. (2013). Plant Cell /tpc
 Intensity Intensity Intensity Intensity Intensity Intensity 150 50 150 0 10 20 50 C 150 0 10 20 50 D 0 10 20 Distance (μm) 50 20 40 E 50 F 0 10 20 50 0 15 30 Distance (μm) Supplemental Figure 1: Co-localization
Intensity Intensity Intensity Intensity Intensity Intensity 150 50 150 0 10 20 50 C 150 0 10 20 50 D 0 10 20 Distance (μm) 50 20 40 E 50 F 0 10 20 50 0 15 30 Distance (μm) Supplemental Figure 1: Co-localization
Genome-wide discovery of G-quadruplex forming sequences and their functional
 *Correspondence and requests for materials should be addressed to R.G. (rohini@nipgr.ac.in) Genome-wide discovery of G-quadruplex forming sequences and their functional relevance in plants Rohini Garg*,
*Correspondence and requests for materials should be addressed to R.G. (rohini@nipgr.ac.in) Genome-wide discovery of G-quadruplex forming sequences and their functional relevance in plants Rohini Garg*,
Prospecting for Green Revolution Genes
 Learning Objectives: Prospecting for Green Revolution Genes 1) Discover how changes in individual genes produce phenotypic change 2) Learn to apply bioinformatics tools to identify groups of related genes
Learning Objectives: Prospecting for Green Revolution Genes 1) Discover how changes in individual genes produce phenotypic change 2) Learn to apply bioinformatics tools to identify groups of related genes
Supplementary Figure 3
 Supplementary Figure 3 7.0 Col Kas-1 Line FTH1A 8.4 F3PII3 8.9 F26H11 ATQ1 T9I22 PLS8 F26B6-B 9.6 F27L4 9.81 F27D4 9.92 9.96 10.12 10.14 10.2 11.1 0.5 Mb T1D16 Col % RGR 83.3 101 227 93.5 75.9 132 90 375
Supplementary Figure 3 7.0 Col Kas-1 Line FTH1A 8.4 F3PII3 8.9 F26H11 ATQ1 T9I22 PLS8 F26B6-B 9.6 F27L4 9.81 F27D4 9.92 9.96 10.12 10.14 10.2 11.1 0.5 Mb T1D16 Col % RGR 83.3 101 227 93.5 75.9 132 90 375
HRS1 Acts as a Negative Regulator of Abscisic Acid Signaling to Promote Timely Germination of Arabidopsis Seeds
 HRS1 Acts as a Negative Regulator of Abscisic Acid Signaling to Promote Timely Germination of Arabidopsis Seeds Chongming Wu 1,2., Juanjuan Feng 1,2., Ran Wang 1,2, Hong Liu 1,2, Huixia Yang 1,2, Pedro
HRS1 Acts as a Negative Regulator of Abscisic Acid Signaling to Promote Timely Germination of Arabidopsis Seeds Chongming Wu 1,2., Juanjuan Feng 1,2., Ran Wang 1,2, Hong Liu 1,2, Huixia Yang 1,2, Pedro
Supplemental Data. Wang et al. (2014). Plant Cell /tpc
 Supplemental Figure1: Mock and NPA-treated tomato plants. (A) NPA treated tomato (cv. Moneymaker) developed a pin-like inflorescence (arrowhead). (B) Comparison of first and second leaves from mock and
Supplemental Figure1: Mock and NPA-treated tomato plants. (A) NPA treated tomato (cv. Moneymaker) developed a pin-like inflorescence (arrowhead). (B) Comparison of first and second leaves from mock and
Figure 1. Identification of UGT74E2 as an IBA glycosyltransferase. (A) Relative conversion rates of different plant hormones to their glucosylated
 Figure 1. Identification of UGT74E2 as an IBA glycosyltransferase. (A) Relative conversion rates of different plant hormones to their glucosylated form by recombinant UGT74E2. The naturally occurring auxin
Figure 1. Identification of UGT74E2 as an IBA glycosyltransferase. (A) Relative conversion rates of different plant hormones to their glucosylated form by recombinant UGT74E2. The naturally occurring auxin
Table S1 List of primers used for genotyping and qrt-pcr.
 Table S1 List of primers used for genotyping and qrt-pcr. genotyping! allele! ligomer*! 5'-sequence-3'! rice! d10-2! F! TTGGCTTTGCCTCGTTTC!!! R! AGCCTCCACTTGTACTGTG! Arabidopsis! max2-3, max2-4! F! ACTCTCTCCGACCTCCCTGACG!!!
Table S1 List of primers used for genotyping and qrt-pcr. genotyping! allele! ligomer*! 5'-sequence-3'! rice! d10-2! F! TTGGCTTTGCCTCGTTTC!!! R! AGCCTCCACTTGTACTGTG! Arabidopsis! max2-3, max2-4! F! ACTCTCTCCGACCTCCCTGACG!!!
Supplementary Figure S1. Amino acid alignment of selected monocot FT-like and TFL-like sequences. Sequences were aligned using ClustalW and analyzed
 Supplementary Figure S1. Amino acid alignment of selected monocot FT-like and TFL-like sequences. Sequences were aligned using ClustalW and analyzed using the Geneious software. Accession numbers of the
Supplementary Figure S1. Amino acid alignment of selected monocot FT-like and TFL-like sequences. Sequences were aligned using ClustalW and analyzed using the Geneious software. Accession numbers of the
The Coch gene controls the subsequent differentiation of pea axial meristems into lateral structures
 The Coch gene controls the subsequent differentiation of pea axial meristems into lateral structures Rozov, S.M. 1, Institute of Cytology and Genetics SD RAS, Novosibirsk, Russia Voroshilova, V.A. 2, 2
The Coch gene controls the subsequent differentiation of pea axial meristems into lateral structures Rozov, S.M. 1, Institute of Cytology and Genetics SD RAS, Novosibirsk, Russia Voroshilova, V.A. 2, 2
Life Science Journal 2014;11(9) Cryptochrome 2 negatively regulates ABA-dependent seed germination in Arabidopsis
 Cryptochrome 2 negatively regulates ABA-dependent seed germination in Arabidopsis Sung-Il Kim 1, Sang Ik Song 3, Hak Soo Seo 1, 2, 4 * 1 Department of Plant Science and Research Institute of Agriculture
Cryptochrome 2 negatively regulates ABA-dependent seed germination in Arabidopsis Sung-Il Kim 1, Sang Ik Song 3, Hak Soo Seo 1, 2, 4 * 1 Department of Plant Science and Research Institute of Agriculture
Supporting Information
 Electronic Supplementary Material (ESI) for Journal of Materials Chemistry B. This journal is The Royal Society of Chemistry 2018 Supporting Information Highly Photoluminescent Carbon Dots Derived from
Electronic Supplementary Material (ESI) for Journal of Materials Chemistry B. This journal is The Royal Society of Chemistry 2018 Supporting Information Highly Photoluminescent Carbon Dots Derived from
Leber s Hereditary Optic Neuropathy
 Leber s Hereditary Optic Neuropathy Dear Editor: It is well known that the majority of Leber s hereditary optic neuropathy (LHON) cases was caused by 3 mtdna primary mutations (m.3460g A, m.11778g A, and
Leber s Hereditary Optic Neuropathy Dear Editor: It is well known that the majority of Leber s hereditary optic neuropathy (LHON) cases was caused by 3 mtdna primary mutations (m.3460g A, m.11778g A, and
Supplementary Information
 Supplementary Information Rice APC/C TE controls tillering through mediating the degradation of MONOCULM 1 Qibing Lin 1*, Dan Wang 1*, Hui Dong 2*, Suhai Gu 1, Zhijun Cheng 1, Jie Gong 2, Ruizhen Qin 1,
Supplementary Information Rice APC/C TE controls tillering through mediating the degradation of MONOCULM 1 Qibing Lin 1*, Dan Wang 1*, Hui Dong 2*, Suhai Gu 1, Zhijun Cheng 1, Jie Gong 2, Ruizhen Qin 1,
Study on form distribution of soil iron in western Jilin and its correlation with soil properties
 35 2 2016 6 GLOBAL GEOLOGY Vol. 35 No. 2 Jun. 2016 1004 5589 2016 02 0593 08 1 1 2 1 1 1. 130061 2. 130012 50 A > B > C > D > E > F > G A A CEC B ph C E A C D G B C D P595 S151. 9 A doi 10. 3969 /j. issn.
35 2 2016 6 GLOBAL GEOLOGY Vol. 35 No. 2 Jun. 2016 1004 5589 2016 02 0593 08 1 1 2 1 1 1. 130061 2. 130012 50 A > B > C > D > E > F > G A A CEC B ph C E A C D G B C D P595 S151. 9 A doi 10. 3969 /j. issn.
Supporting information
 a Supporting information Core-Shell Nanocomposites Based on Gold Nanoparticle@Zinc-Iron- Embedded Porous Carbons Derived from Metal Organic Frameworks as Efficient Dual Catalysts for Oxygen Reduction and
a Supporting information Core-Shell Nanocomposites Based on Gold Nanoparticle@Zinc-Iron- Embedded Porous Carbons Derived from Metal Organic Frameworks as Efficient Dual Catalysts for Oxygen Reduction and
Morphology and photosynthetic enzyme activity of maize phosphoenolpyruvate carboxylase transgenic rice
 Morphology and photosynthetic enzyme activity of maize phosphoenolpyruvate carboxylase transgenic rice W.C. Li 1, J. Wang 1, Y.L. Sun 1, S.D. Ji 1 and S.W. Guo 2 1 College of Life Sciences, Henan Normal
Morphology and photosynthetic enzyme activity of maize phosphoenolpyruvate carboxylase transgenic rice W.C. Li 1, J. Wang 1, Y.L. Sun 1, S.D. Ji 1 and S.W. Guo 2 1 College of Life Sciences, Henan Normal
Hydrothermally Activated Graphene Fiber Fabrics for Textile. Electrodes of Supercapacitors
 Supporting Information for Hydrothermally Activated Graphene Fiber Fabrics for Textile Electrodes of Supercapacitors Zheng Li, Tieqi Huang, Weiwei Gao*, Zhen Xu, Dan Chang, Chunxiao Zhang, and Chao Gao*
Supporting Information for Hydrothermally Activated Graphene Fiber Fabrics for Textile Electrodes of Supercapacitors Zheng Li, Tieqi Huang, Weiwei Gao*, Zhen Xu, Dan Chang, Chunxiao Zhang, and Chao Gao*
EFFECTS OF CROP LOAD ON VEGETATIVE GROWTH OF CITRUS
 EFFECTS OF CROP LOAD ON VEGETATIVE GROWTH OF CITRUS HOS 6545 ADVANCED CITRICULTURE I Regulation of Vegetative Growth L. GENE ALBRIGO Smith, P.F. 1976. Collapse of Murcott tangerine trees. J. Amer. Soc.
EFFECTS OF CROP LOAD ON VEGETATIVE GROWTH OF CITRUS HOS 6545 ADVANCED CITRICULTURE I Regulation of Vegetative Growth L. GENE ALBRIGO Smith, P.F. 1976. Collapse of Murcott tangerine trees. J. Amer. Soc.
California Subject Examinations for Teachers
 California Subject Examinations for Teachers TEST GUIDE SCIENCE SUBTEST II: LIFE SCIENCES Subtest Description This document contains the Life Sciences subject matter requirements arranged according to
California Subject Examinations for Teachers TEST GUIDE SCIENCE SUBTEST II: LIFE SCIENCES Subtest Description This document contains the Life Sciences subject matter requirements arranged according to
Miller & Levine Biology 2010
 A Correlation of 2010 to the Pennsylvania Assessment Anchors Grades 9-12 INTRODUCTION This document demonstrates how 2010 meets the Pennsylvania Assessment Anchors, grades 9-12. Correlation page references
A Correlation of 2010 to the Pennsylvania Assessment Anchors Grades 9-12 INTRODUCTION This document demonstrates how 2010 meets the Pennsylvania Assessment Anchors, grades 9-12. Correlation page references
Introduction. Gene expression is the combined process of :
 1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
THE ROLE OF THE PHYTOCHROME B PHOTORECEPTOR IN THE REGULATION OF PHOTOPERIODIC FLOWERING. AnitaHajdu. Thesis of the Ph.D.
 THE ROLE OF THE PHYTOCHROME B PHOTORECEPTOR IN THE REGULATION OF PHOTOPERIODIC FLOWERING AnitaHajdu Thesis of the Ph.D. dissertation Supervisor: Dr. LászlóKozma-Bognár - senior research associate Doctoral
THE ROLE OF THE PHYTOCHROME B PHOTORECEPTOR IN THE REGULATION OF PHOTOPERIODIC FLOWERING AnitaHajdu Thesis of the Ph.D. dissertation Supervisor: Dr. LászlóKozma-Bognár - senior research associate Doctoral
Transmembrane Domains (TMDs) of ABC transporters
 Transmembrane Domains (TMDs) of ABC transporters Most ABC transporters contain heterodimeric TMDs (e.g. HisMQ, MalFG) TMDs show only limited sequence homology (high diversity) High degree of conservation
Transmembrane Domains (TMDs) of ABC transporters Most ABC transporters contain heterodimeric TMDs (e.g. HisMQ, MalFG) TMDs show only limited sequence homology (high diversity) High degree of conservation
Scalable Preparation of Hierarchical Porous Activated Carbon/graphene composite for High-Performance Supercapacitors
 Electronic Supplementary Material (ESI) for Journal of Materials Chemistry A. This journal is The Royal Society of Chemistry 2019 Supplementary Information Scalable Preparation of Hierarchical Porous Activated
Electronic Supplementary Material (ESI) for Journal of Materials Chemistry A. This journal is The Royal Society of Chemistry 2019 Supplementary Information Scalable Preparation of Hierarchical Porous Activated
AUXIN BINDING PROTEIN 1 (ABP1): A matter of fact. Chun-Ming Liu, Professor
 Commentary AUXIN BINDING PROTEIN 1 (ABP1): A matter of fact Chun-Ming Liu, Professor Editor-in-Chief Journal of Integrative Plant Biology (JIPB) Correspondence: cmliu@ibcas.ac.cn This article has been
Commentary AUXIN BINDING PROTEIN 1 (ABP1): A matter of fact Chun-Ming Liu, Professor Editor-in-Chief Journal of Integrative Plant Biology (JIPB) Correspondence: cmliu@ibcas.ac.cn This article has been
Duplication of an upstream silencer of FZP increases grain yield in rice
 SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41477-017-0042-4 In the format provided by the authors and unedited. Duplication of an upstream silencer of FZP increases grain yield in rice
SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41477-017-0042-4 In the format provided by the authors and unedited. Duplication of an upstream silencer of FZP increases grain yield in rice
Potato Genome Analysis
 Potato Genome Analysis Xin Liu Deputy director BGI research 2016.1.21 WCRTC 2016 @ Nanning Reference genome construction???????????????????????????????????????? Sequencing HELL RIEND WELCOME BGI ZHEN LLOFRI
Potato Genome Analysis Xin Liu Deputy director BGI research 2016.1.21 WCRTC 2016 @ Nanning Reference genome construction???????????????????????????????????????? Sequencing HELL RIEND WELCOME BGI ZHEN LLOFRI
Effects of nitrogen application rate on flag leaf photosynthetic characteristics and grain growth and development of high2quality wheat
 2006,12(1) :49-55 Plant Nutrition and Fertilizer Science,,,, (, 271018) 3 :,, 8901 N 120 360 kg/ hm 2, SN1391 N 120 240 kg/ hm 2, 360 kg/ hm 2,,, ; ; : ; ; ; : S5121106 : A : 1008-505X(2006) 01-0049- 07
2006,12(1) :49-55 Plant Nutrition and Fertilizer Science,,,, (, 271018) 3 :,, 8901 N 120 360 kg/ hm 2, SN1391 N 120 240 kg/ hm 2, 360 kg/ hm 2,,, ; ; : ; ; ; : S5121106 : A : 1008-505X(2006) 01-0049- 07
Flowering Time Control in Plants -How plants know the time to flower?
 Advanced Molecular and Cell Biology II, 2015/12/04 Flowering Time Control in Plants -How plants know the time to flower? Masaki NIWA Grad. Sch. Biostudies, Kyoto Univ. Why can plants bloom every year in
Advanced Molecular and Cell Biology II, 2015/12/04 Flowering Time Control in Plants -How plants know the time to flower? Masaki NIWA Grad. Sch. Biostudies, Kyoto Univ. Why can plants bloom every year in
Optimal bounds for Neuman-Sándor mean in terms of the convex combination of the logarithmic and the second Seiffert means
 Chen et al. Journal of Inequalities and Applications (207) 207:25 DOI 0.86/s660-07-56-7 R E S E A R C H Open Access Optimal bounds for Neuman-Sándor mean in terms of the conve combination of the logarithmic
Chen et al. Journal of Inequalities and Applications (207) 207:25 DOI 0.86/s660-07-56-7 R E S E A R C H Open Access Optimal bounds for Neuman-Sándor mean in terms of the conve combination of the logarithmic
Course Descriptions Biology
 Course Descriptions Biology BIOL 1010 (F/S) Human Anatomy and Physiology I. An introductory study of the structure and function of the human organ systems including the nervous, sensory, muscular, skeletal,
Course Descriptions Biology BIOL 1010 (F/S) Human Anatomy and Physiology I. An introductory study of the structure and function of the human organ systems including the nervous, sensory, muscular, skeletal,
Properties for the Perron complement of three known subclasses of H-matrices
 Wang et al Journal of Inequalities and Applications 2015) 2015:9 DOI 101186/s13660-014-0531-1 R E S E A R C H Open Access Properties for the Perron complement of three known subclasses of H-matrices Leilei
Wang et al Journal of Inequalities and Applications 2015) 2015:9 DOI 101186/s13660-014-0531-1 R E S E A R C H Open Access Properties for the Perron complement of three known subclasses of H-matrices Leilei
RNA-seq to study rice defense responses upon parasitic nematode infections. Tina Kyndt
 RNA-seq to study rice defense responses upon parasitic nematode infections Tina Kyndt 1 Rice (Oryza sativa L.) Most important staple food for at least half of the human population Monocot model plant Paddy
RNA-seq to study rice defense responses upon parasitic nematode infections Tina Kyndt 1 Rice (Oryza sativa L.) Most important staple food for at least half of the human population Monocot model plant Paddy
I. Molecules and Cells: Cells are the structural and functional units of life; cellular processes are based on physical and chemical changes.
 I. Molecules and Cells: Cells are the structural and functional units of life; cellular processes are based on physical and chemical changes. A. Chemistry of Life B. Cells 1. Water How do the unique chemical
I. Molecules and Cells: Cells are the structural and functional units of life; cellular processes are based on physical and chemical changes. A. Chemistry of Life B. Cells 1. Water How do the unique chemical
Bioinformatics tools to analyze complex genomes. Yves Van de Peer Ghent University/VIB
 Bioinformatics tools to analyze complex genomes Yves Van de Peer Ghent University/VIB Detecting colinearity and large-scale gene duplications A 1 2 3 4 5 6 7 8 9 10 11 Speciation/Duplicatio n S1 S2 1
Bioinformatics tools to analyze complex genomes Yves Van de Peer Ghent University/VIB Detecting colinearity and large-scale gene duplications A 1 2 3 4 5 6 7 8 9 10 11 Speciation/Duplicatio n S1 S2 1
Photoreceptor Regulation of Constans Protein in Photoperiodic Flowering
 Photoreceptor Regulation of Constans Protein in Photoperiodic Flowering by Valverde et. Al Published in Science 2004 Presented by Boyana Grigorova CBMG 688R Feb. 12, 2007 Circadian Rhythms: The Clock Within
Photoreceptor Regulation of Constans Protein in Photoperiodic Flowering by Valverde et. Al Published in Science 2004 Presented by Boyana Grigorova CBMG 688R Feb. 12, 2007 Circadian Rhythms: The Clock Within
AUXIN CONJUGATE HYDROLYSIS DURING PLANT-MICROBE INTERACTION AND EVOLUTION
 AUXIN CONJUGATE HYDROLYSIS DURING PLANT-MICROBE INTERACTION AND EVOLUTION J. Ludwig-Müller 1, A. Schuller 1, A.F. Olajide 2, V. Bakllamaja 2, J.J. Campanella 2 ABSTRACT Plants regulate auxin balance through
AUXIN CONJUGATE HYDROLYSIS DURING PLANT-MICROBE INTERACTION AND EVOLUTION J. Ludwig-Müller 1, A. Schuller 1, A.F. Olajide 2, V. Bakllamaja 2, J.J. Campanella 2 ABSTRACT Plants regulate auxin balance through
A MicroRNA as a Translational Repressor of APETALA2 in Arabidopsis Flower Development
 A MicroRNA as a Translational Repressor of APETALA2 in Arabidopsis Flower Development Xuemei Chen Waksman Institute, Rutgers University, Piscataway, NJ 08854, USA. E-mail: xuemei@waksman.rutgers.edu Plant
A MicroRNA as a Translational Repressor of APETALA2 in Arabidopsis Flower Development Xuemei Chen Waksman Institute, Rutgers University, Piscataway, NJ 08854, USA. E-mail: xuemei@waksman.rutgers.edu Plant
purpose of this Chapter is to highlight some problems that will likely provide new
 119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
ADVANCED PLACEMENT BIOLOGY
 ADVANCED PLACEMENT BIOLOGY Description Advanced Placement Biology is designed to be the equivalent of a two-semester college introductory course for Biology majors. The course meets seven periods per week
ADVANCED PLACEMENT BIOLOGY Description Advanced Placement Biology is designed to be the equivalent of a two-semester college introductory course for Biology majors. The course meets seven periods per week
Supporting Information
 Supporting Information Nitrogen-doped coal tar pitch based microporous carbons with superior CO 2 capture performance Dai Yu, Jun Hu, Lihui Zhou *, Jinxia Li, Jing Tang, Changjun Peng, and Honglai Liu
Supporting Information Nitrogen-doped coal tar pitch based microporous carbons with superior CO 2 capture performance Dai Yu, Jun Hu, Lihui Zhou *, Jinxia Li, Jing Tang, Changjun Peng, and Honglai Liu
Supplemental Data. Fernández-Calvo et al. Plant Cell. (2011) /tpc
 Supplemental Data. Fernández-Calvo et al. Plant Cell. (2011). 10.1105/tpc.110.080788 Supplemental Figure S1. Phylogenetic tree of MYC2-related proteins from Arabidopsis and other plants. Phenogram representation
Supplemental Data. Fernández-Calvo et al. Plant Cell. (2011). 10.1105/tpc.110.080788 Supplemental Figure S1. Phylogenetic tree of MYC2-related proteins from Arabidopsis and other plants. Phenogram representation
Genome-wide multilevel spatial interactome model of rice
 Sino-German Workshop on Multiscale Spatial Computational Systems Biology, Beijing, Oct 8-12, 2015 Genome-wide multilevel spatial interactome model of rice Ming CHEN ( 陈铭 ) mchen@zju.edu.cn College of Life
Sino-German Workshop on Multiscale Spatial Computational Systems Biology, Beijing, Oct 8-12, 2015 Genome-wide multilevel spatial interactome model of rice Ming CHEN ( 陈铭 ) mchen@zju.edu.cn College of Life
Effect of nitrogen application on nitrogen absorption, distribution and yield of Stra wberry
 26,12(3) :4-45 Plant Nutrition and Fertilizer Science,,,, (, 27118) : 15 N 15 N, 15 N,, 5215 %, 3412 % 85 %, 15 % ; 7212 %, 2718 % ; 3113 % 15 N 15 N, SPAD, N 225 kg/ hm 2 : ; ; 15 N ; ; ; : S66814 ; S66
26,12(3) :4-45 Plant Nutrition and Fertilizer Science,,,, (, 27118) : 15 N 15 N, 15 N,, 5215 %, 3412 % 85 %, 15 % ; 7212 %, 2718 % ; 3113 % 15 N 15 N, SPAD, N 225 kg/ hm 2 : ; ; 15 N ; ; ; : S66814 ; S66
Curriculum Links. AQA GCE Biology. AS level
 Curriculum Links AQA GCE Biology Unit 2 BIOL2 The variety of living organisms 3.2.1 Living organisms vary and this variation is influenced by genetic and environmental factors Causes of variation 3.2.2
Curriculum Links AQA GCE Biology Unit 2 BIOL2 The variety of living organisms 3.2.1 Living organisms vary and this variation is influenced by genetic and environmental factors Causes of variation 3.2.2
CONTROL OF PLANT GROWTH AND DEVELOPMENT BI-2232 RIZKITA R E
 CONTROL OF PLANT GROWTH AND DEVELOPMENT BI-2232 RIZKITA R E The development of a plant the series of progressive changes that take place throughout its life is regulated in complex ways. Factors take part
CONTROL OF PLANT GROWTH AND DEVELOPMENT BI-2232 RIZKITA R E The development of a plant the series of progressive changes that take place throughout its life is regulated in complex ways. Factors take part
Supporting Information. Co 4 N Nanosheets Assembled Mesoporous Sphere as a Matrix for Ultrahigh Sulfur Content Lithium Sulfur Batteries
 Supporting Information Co 4 N Nanosheets Assembled Mesoporous Sphere as a Matrix for Ultrahigh Sulfur Content Lithium Sulfur Batteries Ding-Rong Deng, Fei Xue, Yue-Ju Jia, Jian-Chuan Ye, Cheng-Dong Bai,
Supporting Information Co 4 N Nanosheets Assembled Mesoporous Sphere as a Matrix for Ultrahigh Sulfur Content Lithium Sulfur Batteries Ding-Rong Deng, Fei Xue, Yue-Ju Jia, Jian-Chuan Ye, Cheng-Dong Bai,
Compare and contrast the cellular structures and degrees of complexity of prokaryotic and eukaryotic organisms.
 Subject Area - 3: Science and Technology and Engineering Education Standard Area - 3.1: Biological Sciences Organizing Category - 3.1.A: Organisms and Cells Course - 3.1.B.A: BIOLOGY Standard - 3.1.B.A1:
Subject Area - 3: Science and Technology and Engineering Education Standard Area - 3.1: Biological Sciences Organizing Category - 3.1.A: Organisms and Cells Course - 3.1.B.A: BIOLOGY Standard - 3.1.B.A1:
Experimental and numerical simulation studies of the squeezing dynamics of the UBVT system with a hole-plug device
 Experimental numerical simulation studies of the squeezing dynamics of the UBVT system with a hole-plug device Wen-bin Gu 1 Yun-hao Hu 2 Zhen-xiong Wang 3 Jian-qing Liu 4 Xiao-hua Yu 5 Jiang-hai Chen 6
Experimental numerical simulation studies of the squeezing dynamics of the UBVT system with a hole-plug device Wen-bin Gu 1 Yun-hao Hu 2 Zhen-xiong Wang 3 Jian-qing Liu 4 Xiao-hua Yu 5 Jiang-hai Chen 6
AP Biology Essential Knowledge Cards BIG IDEA 1
 AP Biology Essential Knowledge Cards BIG IDEA 1 Essential knowledge 1.A.1: Natural selection is a major mechanism of evolution. Essential knowledge 1.A.4: Biological evolution is supported by scientific
AP Biology Essential Knowledge Cards BIG IDEA 1 Essential knowledge 1.A.1: Natural selection is a major mechanism of evolution. Essential knowledge 1.A.4: Biological evolution is supported by scientific
Supporting Information
 Supporting Information Electrochemiluminescent Pb 2+ -Driven Circular Etching Sensor Coupled to a DNA Micronet-Carrier Wen-Bin Liang,, Ying Zhuo, Ying-Ning Zheng, Cheng-Yi Xiong, Ya-Qin Chai, *, and Ruo
Supporting Information Electrochemiluminescent Pb 2+ -Driven Circular Etching Sensor Coupled to a DNA Micronet-Carrier Wen-Bin Liang,, Ying Zhuo, Ying-Ning Zheng, Cheng-Yi Xiong, Ya-Qin Chai, *, and Ruo
The Pht1;9 and Pht1;8 transporters mediate inorganic phosphate acquisition by the Arabidopsis thaliana root during phosphorus starvation
 Research The Pht1;9 and Pht1;8 transporters mediate inorganic phosphate acquisition by the Arabidopsis thaliana root during phosphorus starvation E. Remy 1, T. R. Cabrito 2, R. A. Batista 1, M. C. Teixeira
Research The Pht1;9 and Pht1;8 transporters mediate inorganic phosphate acquisition by the Arabidopsis thaliana root during phosphorus starvation E. Remy 1, T. R. Cabrito 2, R. A. Batista 1, M. C. Teixeira
Increasing Processing Tomato Fruit Soluble Solids
 Increasing Processing Tomato Fruit Soluble Solids Diane M Beckles Department of Plant Sciences dmbeckles@ucdavis.edu Processing Tomato Conference @ UC Davis December 13 th 2018 Soil Micronutrients Cultivar
Increasing Processing Tomato Fruit Soluble Solids Diane M Beckles Department of Plant Sciences dmbeckles@ucdavis.edu Processing Tomato Conference @ UC Davis December 13 th 2018 Soil Micronutrients Cultivar
Classical Selection, Balancing Selection, and Neutral Mutations
 Classical Selection, Balancing Selection, and Neutral Mutations Classical Selection Perspective of the Fate of Mutations All mutations are EITHER beneficial or deleterious o Beneficial mutations are selected
Classical Selection, Balancing Selection, and Neutral Mutations Classical Selection Perspective of the Fate of Mutations All mutations are EITHER beneficial or deleterious o Beneficial mutations are selected
Model plants and their Role in genetic manipulation. Mitesh Shrestha
 Model plants and their Role in genetic manipulation Mitesh Shrestha Definition of Model Organism Specific species or organism Extensively studied in research laboratories Advance our understanding of Cellular
Model plants and their Role in genetic manipulation Mitesh Shrestha Definition of Model Organism Specific species or organism Extensively studied in research laboratories Advance our understanding of Cellular
High nitrogen conditions enhance the cooling damage to pollen in rice plants : proteome analysis of mature anthers
 High nitrogen conditions enhance the cooling damage to pollen in rice plants : proteome analysis of mature anthers Takami Hayashi 1, Tomoya Yamaguchi 1, Katsuhiro Nakayama 1, Setsuko Komatsu 2 and Setsuo
High nitrogen conditions enhance the cooling damage to pollen in rice plants : proteome analysis of mature anthers Takami Hayashi 1, Tomoya Yamaguchi 1, Katsuhiro Nakayama 1, Setsuko Komatsu 2 and Setsuo
Biology II : Embedded Inquiry
 Biology II : Embedded Inquiry Conceptual Strand Understandings about scientific inquiry and the ability to conduct inquiry are essential for living in the 21 st century. Guiding Question What tools, skills,
Biology II : Embedded Inquiry Conceptual Strand Understandings about scientific inquiry and the ability to conduct inquiry are essential for living in the 21 st century. Guiding Question What tools, skills,
1. (a) Why are the two kinds of self-incompatibiltiy (SI) mechanisms called sporophytic and gametophytic?
 Bio 328 -Spring 2005 NAME: Test #1 Please provide succinct answers in the space provided under each question. Unless otherwise noted in the margin the value of each question is 3 points. 1. (a) Why are
Bio 328 -Spring 2005 NAME: Test #1 Please provide succinct answers in the space provided under each question. Unless otherwise noted in the margin the value of each question is 3 points. 1. (a) Why are
Supplementary Information
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2017 Supplementary Information Supramolecular interactions via hydrogen bonding contributing to
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2017 Supplementary Information Supramolecular interactions via hydrogen bonding contributing to
