Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1. Different crystal forms obtained for Sky
|
|
|
- Randell Harper
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1 Different crystal forms obtained for Sky (a) Crystal form 1 obtained in the presence of 20% PEG 3350 and 0.2 M ammonium citrate tribasic ph 7.0. (b) Crystal form 1 of the selenomethionine-labelled Sky obtained in the presence of 20% PEG 3350 and 0.2 M ammonium citrate tribasic ph 7.0 (c) Crystal form 2 obtained in the presence of 25% PEG 1500 and 0.1 M succinate/phosphate/glycine ph 7.0. This crystal form was used to obtain the Sky IP 3 crystal structure after soaking with 5 mm IP 3.
2
3 Supplementary Figure 2 Sequence alignment of Sky with the TBC domain of human TBC1D24 and representative RabGAP proteins for which the crystal structure has been reported. The residues corresponding to the arginine and glutamine fingers in conventional TBC Rab-GAP proteins are highlighted in green. The residues corresponding to the cationic pocket residues in Sky are highlighted in blue. Residues corresponding to patient mutations are red, with those located in the cationic pocket additionally indicated by a red star. The α-helices determined from the Sky structure are indicated above the alignment, with the numbering corresponding to the numbering used in the Gyp1 structure. The sequences used are human TBC1D24 (UniProt: Q9ULP9), Gyp1 from Saccharomyces cerevisiae (UniProt: Q08484), human TBC1D1 (UniProt: Q86TI0), human TBC1D4 (Uniprot: O60343), human TBC1D7 (UniProt: Q9P0N9), human TBC1D11 (UniProt: Q9Y3P9), human TBC1D14 (UniProt: Q9P2M4), human TBC1D18 (UniProt: Q5R372), human TBC1D20 (UniProt: Q96BZ9), human TBC1D22A (UniProt: Q8WUA7), human TBC1D22B (UniProt: Q9NU19), CrfRabGAP from Chlamydomonas reinhardtii (UniProt: A8JCA4).
4 Supplementary Figure 3 Surface electrostatics of Sky in comparison to other TBC domains. The electrostatic potential mapped on the solvent accessible surface of Sky in the first panel shows the cationic pocket located on the opposite side of the GTPase binding region. The electrostatic potential surfaces of the 11 other TBC domains deposited in the PDB are shown in exactly the same orientation. None of these proteins seems to harbor a well-defined cationic pocket similar to Sky The structures of the TBC domains shown in this figure are: Gyp1 (pdb 1FKM; Rak, A. et al., EMBO J. 19, , 2000), TBC1D1 (pdb 3QYE; Park, S.-Y. et al., J. Biol. Chem. 286, , 2011), TBC1D4 (pdb 3QYB; Park, S.-Y. et al., J. Biol. Chem. 286, , 2011), TBC1D7 (pdb 3QWL; unpublished), TBC1D11 (pdb 4NC6; unpublished), TBC1D14 (pdb 2QQ8; unpublished), TBC1D18 (pdb 3HZJ; unpublished), TBC1D20 (pdb 4HL4; Gavriljuk, K. et al., Proc. Natl. Acad. Sci. U.S.A. 109, , 2012), TBC1D22A (pdb 2QFZ; unpublished), TBC1D22B (pdb 3DZX; unpublished), CrfRabGAP (pdb 4P17; Bhogaraju, S. & Lorentzen, E. Proteins 82, , 2014).
5 Supplementary Figure 4 Influence of PI(4,5)P 2 and salt concentrations on the binding of Sky to liposomes. (a) Western blots of liposome flotation assays using wild type Sky and liposomes (PC:PS) enriched with different concentrations of PI(4,5)P 2 (0%, 0.5%, 2% and 5%). n = 2 independent experiments. (b) Western blots of liposome flotation assays using wild type Sky and liposomes (PC:PS) enriched with 2% of PI(4,5)P 2 in the presence of three different NaCl concentrations (30 mm, 150 mm and 500 mm). n = 2 independent experiments. Original blots can be found in Supplementary Data Set 1.
6 Supplementary Figure 5 Comparison of the phosphoinositide-binding pocket of Sky with other typical phosphoinositide-binding domains. Structures of representatives of well-established phosphoinositide-binding domains (ENTH, PX, FYVE and PH) in complex with a phosphoinositide head group are shown in electrostatic surface representation and are compared to the Sky structure in complex with IP 3. The boxes show a close-up of the phosphoinositide binding pocket in cartoon representation with the phosphoinositide head group and the interacting residues shown as sticks. The structures that are shown are: the TBC domain of Sky bound to IP 3 (this study), the ENTH domain of epsin bound to IP 3 (pdb 1H0A; Ford, M. G. J. et al., Nature 419, , 2002), the PX domain of P40 phox bound to dibutanoyl IP 2 (pdb 1H6H; Bravo, J. et al., Mol. Cell 8, , 2001), the FYVE domain of EEA1 bound to IP 2 (pdb 1JOC; Dumas, J. J. et al., Mol. Cell 8, , 2001), the PH domain of PLC-δ1 bound to IP 3 (pdb 1MAI; Ferguson, K. M. et al., Cell 83, , 1995) and the PH domain of spectrin bound to IP 3 (pdb 1BTN; Hyvönen, M. et al., EMBO J. 14, , 1995).
7 Supplementary Figure 6 Wild-type and mutant Skywalker-GFP traffic to and are present at synaptic terminals (a) Confocal images of the ventral nerve cord in third instar Drosophila larvae expressing wild type GFP-Sky (Sky WT ) or mutant GFP- Sky (Sky R79C, Sky R281C or Sky 3Glu ). Scale bar for all top panels: 20 m. (b) Confocal images of larval neuromuscular junction endplates of animals with genotypes as in (a). (n = 6 animals), scale bar for all panels in (b): 20 m. GFP intensities between the GFP-Sky mutants is not significantly different. (c) Quantification of third-instar NMJ GFP intensity at the membrane in single confocal sections. Mean Fluorescence Intensity between the genotypes GFP-Sky WT, GFP-Sky R79C and GFP-Sky 3Glu is similar. Error bars: mean ± s.e.m. (n = 6) P = 0.446, ns: not significant by
8 ANOVA, Dunnett s. (d) Quantification of Western blot band intensity of GFP-Sky and GFP-Syt in protein isolations from adult fly heads. Intensities are normalized to Syntaxin control levels. GFP-Sky WT and mutant levels are similar. Error bars: mean ± s.e.m. (n = 3) P = 0.681, ns: not significant by ANOVA, Dunnett s. The levels of Synaptotagmin-GFP (Syt) are shown as a control.
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Crystallization. a, Crystallization constructs of the ET B receptor are shown, with all of the modifications to the human wild-type the ET B receptor indicated. Residues interacting
Supplementary Figure 1 Crystallization. a, Crystallization constructs of the ET B receptor are shown, with all of the modifications to the human wild-type the ET B receptor indicated. Residues interacting
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3267 Supplementary Figure 1 A group of genes required for formation or orientation of annular F-actin bundles and aecm ridges: RNAi phenotypes and their validation by standard mutations.
DOI: 10.1038/ncb3267 Supplementary Figure 1 A group of genes required for formation or orientation of annular F-actin bundles and aecm ridges: RNAi phenotypes and their validation by standard mutations.
Rho1 binding site PtdIns(4,5)P2 binding site Both sites
 localization Mutation site DMSO LatB WT F77A I115A I131A K134A Rho1 binding site PtdIns(4,5)P2 binding site Both sites E186A E199A N201A R84A-E186A-E199A L131A-K136A-E186A L131A-E186A-E199A K136A-E186A-E199A
localization Mutation site DMSO LatB WT F77A I115A I131A K134A Rho1 binding site PtdIns(4,5)P2 binding site Both sites E186A E199A N201A R84A-E186A-E199A L131A-K136A-E186A L131A-E186A-E199A K136A-E186A-E199A
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a
 Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
Supplementary figure 1 Application of tmfret in LeuT. (a) To assess the feasibility of using tmfret for distance-dependent measurements in LeuT, a series of tmfret-pairs comprised of single cysteine mutants
Structure and RNA-binding properties. of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex
 Structure and RNA-binding properties of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex Varun Bhaskar 1, Vladimir Roudko 2,3, Jerome Basquin 1, Kundan Sharma 4, Henning Urlaub 4, Bertrand Seraphin
Structure and RNA-binding properties of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex Varun Bhaskar 1, Vladimir Roudko 2,3, Jerome Basquin 1, Kundan Sharma 4, Henning Urlaub 4, Bertrand Seraphin
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10923 Supplementary Figure 1 Ten-a and Ten-m antibody and cell type specificities. a c, Representative single confocal sections of a Drosophila NMJ stained with antibodies to Ten-a (red),
doi:10.1038/nature10923 Supplementary Figure 1 Ten-a and Ten-m antibody and cell type specificities. a c, Representative single confocal sections of a Drosophila NMJ stained with antibodies to Ten-a (red),
Supplementary Figure 1. Biochemical and sequence alignment analyses the
 Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
Supplementary Figure 1. Biochemical and sequence alignment analyses the interaction of OPTN and TBK1. (a) Analytical gel filtration chromatography analysis of the interaction between TBK1 CTD and OPTN(1-119).
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11085 Supplementary Tables: Supplementary Table 1. Summary of crystallographic and structure refinement data Structure BRIL-NOP receptor Data collection Number of crystals 23 Space group
doi:10.1038/nature11085 Supplementary Tables: Supplementary Table 1. Summary of crystallographic and structure refinement data Structure BRIL-NOP receptor Data collection Number of crystals 23 Space group
Supplementary Figure 1 Crystal packing of ClR and electron density maps. Crystal packing of type A crystal (a) and type B crystal (b).
 Supplementary Figure 1 Crystal packing of ClR and electron density maps. Crystal packing of type A crystal (a) and type B crystal (b). Crystal contacts at B-C loop are magnified and stereo view of A-weighted
Supplementary Figure 1 Crystal packing of ClR and electron density maps. Crystal packing of type A crystal (a) and type B crystal (b). Crystal contacts at B-C loop are magnified and stereo view of A-weighted
Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and
 Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and Lifeact-Ruby (red) were imaged in vivo to visualize secretion
Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and Lifeact-Ruby (red) were imaged in vivo to visualize secretion
SUPPLEMENTARY INFORMATION. doi: /nature07461
 Figure S1 Electrophysiology. a ph-activation of. Two-electrode voltage clamp recordings of Xenopus oocytes expressing in comparison to waterinjected oocytes. Currents were recorded at 40 mv. The ph of
Figure S1 Electrophysiology. a ph-activation of. Two-electrode voltage clamp recordings of Xenopus oocytes expressing in comparison to waterinjected oocytes. Currents were recorded at 40 mv. The ph of
Nature Neuroscience: doi: /nn.2717
 Supplementary Fig. 1. Dendrite length is not secondary to body length. Dendrite growth proceeds independently of the rate of body growth and decreases in rate in adults. n 20 on dendrite measurement, n
Supplementary Fig. 1. Dendrite length is not secondary to body length. Dendrite growth proceeds independently of the rate of body growth and decreases in rate in adults. n 20 on dendrite measurement, n
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
SUPPLEMENTARY INFORMATION doi:10.1038/nature11524 Supplementary discussion Functional analysis of the sugar porter family (SP) signature motifs. As seen in Fig. 5c, single point mutation of the conserved
NB-DNJ/GCase-pH 7.4 NB-DNJ+/GCase-pH 7.4 NB-DNJ+/GCase-pH 4.5
 SUPPLEMENTARY TABLES Suppl. Table 1. Protonation states at ph 7.4 and 4.5. Protonation states of titratable residues in GCase at ph 7.4 and 4.5. Histidine: HID, H at δ-nitrogen; HIE, H at ε-nitrogen; HIP,
SUPPLEMENTARY TABLES Suppl. Table 1. Protonation states at ph 7.4 and 4.5. Protonation states of titratable residues in GCase at ph 7.4 and 4.5. Histidine: HID, H at δ-nitrogen; HIE, H at ε-nitrogen; HIP,
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/5/243/ra68/dc1 Supplementary Materials for Superbinder SH2 Domains Act as Antagonists of Cell Signaling Tomonori Kaneko, Haiming Huang, Xuan Cao, Xing Li, Chengjun
www.sciencesignaling.org/cgi/content/full/5/243/ra68/dc1 Supplementary Materials for Superbinder SH2 Domains Act as Antagonists of Cell Signaling Tomonori Kaneko, Haiming Huang, Xuan Cao, Xing Li, Chengjun
Expanded View Figures
 The EMBO Journal Structure of a Dm peptide bound to the OT module Tobias Raisch et al Expanded View Figures A Hs Dm 262 297 685 8 HEAT HEAT MIF4G 9BD 1SHD 761 91 193 169 1152 1317 16 1376 1467 HEAT HEAT
The EMBO Journal Structure of a Dm peptide bound to the OT module Tobias Raisch et al Expanded View Figures A Hs Dm 262 297 685 8 HEAT HEAT MIF4G 9BD 1SHD 761 91 193 169 1152 1317 16 1376 1467 HEAT HEAT
SUPPLEMENTARY INFORMATION
 Supplementary Figure 1 Sns and Duf co-localise in embryonic nephrocytes a-c, Wild-type stage 11 (a),14 (b) and 16 (c) embryos stained with anti-duf (green) and anti-sns (red). Higher magnification images
Supplementary Figure 1 Sns and Duf co-localise in embryonic nephrocytes a-c, Wild-type stage 11 (a),14 (b) and 16 (c) embryos stained with anti-duf (green) and anti-sns (red). Higher magnification images
SUPPLEMENTARY INFORMATION
 Supplementary Table 1: Amplitudes of three current levels. Level 0 (pa) Level 1 (pa) Level 2 (pa) TrkA- TrkH WT 200 K 0.01 ± 0.01 9.5 ± 0.01 18.7 ± 0.03 200 Na * 0.001 ± 0.01 3.9 ± 0.01 12.5 ± 0.03 200
Supplementary Table 1: Amplitudes of three current levels. Level 0 (pa) Level 1 (pa) Level 2 (pa) TrkA- TrkH WT 200 K 0.01 ± 0.01 9.5 ± 0.01 18.7 ± 0.03 200 Na * 0.001 ± 0.01 3.9 ± 0.01 12.5 ± 0.03 200
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature17991 Supplementary Discussion Structural comparison with E. coli EmrE The DMT superfamily includes a wide variety of transporters with 4-10 TM segments 1. Since the subfamilies of the
doi:10.1038/nature17991 Supplementary Discussion Structural comparison with E. coli EmrE The DMT superfamily includes a wide variety of transporters with 4-10 TM segments 1. Since the subfamilies of the
THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN
 THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN HSP90 AND BOTH SCF E3 UBIQUITIN LIGASES AND KINETOCHORES Oliver Willhoft, Richard Kerr, Dipali Patel, Wenjuan Zhang, Caezar Al-Jassar, Tina
THE CRYSTAL STRUCTURE OF THE SGT1-SKP1 COMPLEX: THE LINK BETWEEN HSP90 AND BOTH SCF E3 UBIQUITIN LIGASES AND KINETOCHORES Oliver Willhoft, Richard Kerr, Dipali Patel, Wenjuan Zhang, Caezar Al-Jassar, Tina
Supplementary Information. The protease GtgE from Salmonella exclusively targets. inactive Rab GTPases
 Supplementary Information The protease GtgE from Salmonella exclusively targets inactive Rab GTPases Table of Contents Supplementary Figures... 2 Supplementary Figure 1... 2 Supplementary Figure 2... 3
Supplementary Information The protease GtgE from Salmonella exclusively targets inactive Rab GTPases Table of Contents Supplementary Figures... 2 Supplementary Figure 1... 2 Supplementary Figure 2... 3
TBC1D9B functions as a GTPase-activating protein for Rab11a in polarized MDCK cells
 MBoC ARTICLE TBC1D9B functions as a GTPase-activating protein for Rab11a in polarized MDCK cells Luciana I. Gallo a, Yong Liao b, Wily G. Ruiz a, Dennis R. Clayton a, Min Li b,c, Yong-Jian Liu d, Yu Jiang
MBoC ARTICLE TBC1D9B functions as a GTPase-activating protein for Rab11a in polarized MDCK cells Luciana I. Gallo a, Yong Liao b, Wily G. Ruiz a, Dennis R. Clayton a, Min Li b,c, Yong-Jian Liu d, Yu Jiang
Enhanced zinc-finger-nuclease activity with improved obligate heterodimeric architectures
 Nature Methods Enhanced zinc-finger-nuclease activity with improved obligate heterodimeric architectures Yannick Doyon, Thuy D Vo, Matthew C Mendel, Shon G Greenberg, Jianbin Wang, Danny F Xia, Jeffrey
Nature Methods Enhanced zinc-finger-nuclease activity with improved obligate heterodimeric architectures Yannick Doyon, Thuy D Vo, Matthew C Mendel, Shon G Greenberg, Jianbin Wang, Danny F Xia, Jeffrey
Ab1 (57-68) Ab2 ( ) Ab3 ( ) Ab4 ( ) GBP-N domain GBP-C domain CARD domain
 MDKPVCLIDTGSDGKLCVQQAALQVLQQIQQPVVVVAVVGLYRTGKSFLMNRLAG 55 KRTGFALSSNIKPKTEGIWMWCVPHPTKAGTSLVLLDTKGLGDVEKGDSKRDTYI 110 FSLTVLLSSTLVYNSRGVIDNKAMEELQYVTELIEHIKVTPDEDADDCTAFAKFF 165 PHFIWCLRDFTLELKLDGKDLTEDEYLEFALKLRPGTLKKVMMYNLPRECIQKFF
MDKPVCLIDTGSDGKLCVQQAALQVLQQIQQPVVVVAVVGLYRTGKSFLMNRLAG 55 KRTGFALSSNIKPKTEGIWMWCVPHPTKAGTSLVLLDTKGLGDVEKGDSKRDTYI 110 FSLTVLLSSTLVYNSRGVIDNKAMEELQYVTELIEHIKVTPDEDADDCTAFAKFF 165 PHFIWCLRDFTLELKLDGKDLTEDEYLEFALKLRPGTLKKVMMYNLPRECIQKFF
Supplementary Figure 1. Markedly decreased numbers of marginal zone B cells in DOCK8 mutant mice Supplementary Figure 2.
 Supplementary Figure 1. Markedly decreased numbers of marginal zone B cells in DOCK8 mutant mice. Percentage of marginal zone B cells in the spleen of wild-type mice (+/+), mice homozygous for cpm or pri
Supplementary Figure 1. Markedly decreased numbers of marginal zone B cells in DOCK8 mutant mice. Percentage of marginal zone B cells in the spleen of wild-type mice (+/+), mice homozygous for cpm or pri
Supporting Information
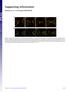 Supporting Information Fleissner et al. 10.1073/pnas.0907039106 Fig. S1. (A) MAK-2-GFP localized to CATs tips is not bound by membrane. his-3::pccg1 mak-2-gfp; mak-2 strain labeled with membrane dye FM4
Supporting Information Fleissner et al. 10.1073/pnas.0907039106 Fig. S1. (A) MAK-2-GFP localized to CATs tips is not bound by membrane. his-3::pccg1 mak-2-gfp; mak-2 strain labeled with membrane dye FM4
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2647 Figure S1 Other Rab GTPases do not co-localize with the ER. a, Cos-7 cells cotransfected with an ER luminal marker (either KDEL-venus or mch-kdel) and mch-tagged human Rab5 (mch-rab5,
DOI: 10.1038/ncb2647 Figure S1 Other Rab GTPases do not co-localize with the ER. a, Cos-7 cells cotransfected with an ER luminal marker (either KDEL-venus or mch-kdel) and mch-tagged human Rab5 (mch-rab5,
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature12791 Supplementary Figure 1 (1/3) WWW.NATURE.COM/NATURE 1 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 1 (2/3) 2 WWW.NATURE.COM/NATURE SUPPLEMENTARY
SUPPLEMENTARY INFORMATION doi:10.1038/nature12791 Supplementary Figure 1 (1/3) WWW.NATURE.COM/NATURE 1 RESEARCH SUPPLEMENTARY INFORMATION Supplementary Figure 1 (2/3) 2 WWW.NATURE.COM/NATURE SUPPLEMENTARY
Supplementary Figure 1 Schematic overview of ASTNs in neuronal migration. (a) Schematic of roles played by ASTNs 1 and 2. ASTN-1-mediated adhesions
 Supplementary Figure 1 Schematic overview of ASTNs in neuronal migration. (a) Schematic of roles played by ASTNs 1 and 2. ASTN-1-mediated adhesions undergo endocytosis into clathrin-coated vesicles dependent
Supplementary Figure 1 Schematic overview of ASTNs in neuronal migration. (a) Schematic of roles played by ASTNs 1 and 2. ASTN-1-mediated adhesions undergo endocytosis into clathrin-coated vesicles dependent
Nature Neuroscience: doi: /nn.2662
 Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Identification of the ScDcp2 minimal region interacting with both ScDcp1 and the ScEdc3 LSm domain. Pull-down experiment of untagged ScEdc3 LSm with various ScDcp1-Dcp2-His 6 fragments.
Supplementary Figure 1 Identification of the ScDcp2 minimal region interacting with both ScDcp1 and the ScEdc3 LSm domain. Pull-down experiment of untagged ScEdc3 LSm with various ScDcp1-Dcp2-His 6 fragments.
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Chemical structure of LPS and LPS biogenesis in Gram-negative bacteria. a. Chemical structure of LPS. LPS molecule consists of Lipid A, core oligosaccharide and O-antigen. The polar
Supplementary Figure 1 Chemical structure of LPS and LPS biogenesis in Gram-negative bacteria. a. Chemical structure of LPS. LPS molecule consists of Lipid A, core oligosaccharide and O-antigen. The polar
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.1244624/dc1 Supplementary Materials for Cytoneme-Mediated Contact-Dependent Transport of the Drosophila Decapentaplegic Signaling Protein Sougata Roy, Hai Huang,
www.sciencemag.org/cgi/content/full/science.1244624/dc1 Supplementary Materials for Cytoneme-Mediated Contact-Dependent Transport of the Drosophila Decapentaplegic Signaling Protein Sougata Roy, Hai Huang,
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12045 Supplementary Table 1 Data collection and refinement statistics. Native Pt-SAD X-ray source SSRF BL17U SPring-8 BL41XU Wavelength (Å) 0.97947 1.07171 Space group P2 1 2 1 2 1 P2
doi:10.1038/nature12045 Supplementary Table 1 Data collection and refinement statistics. Native Pt-SAD X-ray source SSRF BL17U SPring-8 BL41XU Wavelength (Å) 0.97947 1.07171 Space group P2 1 2 1 2 1 P2
Supplementary materials. Crystal structure of the carboxyltransferase domain. of acetyl coenzyme A carboxylase. Department of Biological Sciences
 Supplementary materials Crystal structure of the carboxyltransferase domain of acetyl coenzyme A carboxylase Hailong Zhang, Zhiru Yang, 1 Yang Shen, 1 Liang Tong Department of Biological Sciences Columbia
Supplementary materials Crystal structure of the carboxyltransferase domain of acetyl coenzyme A carboxylase Hailong Zhang, Zhiru Yang, 1 Yang Shen, 1 Liang Tong Department of Biological Sciences Columbia
SUPPLEMENTARY INFORMATION
 Fig. 1 Influences of crystal lattice contacts on Pol η structures. a. The dominant lattice contact between two hpol η molecules (silver and gold) in the type 1 crystals. b. A close-up view of the hydrophobic
Fig. 1 Influences of crystal lattice contacts on Pol η structures. a. The dominant lattice contact between two hpol η molecules (silver and gold) in the type 1 crystals. b. A close-up view of the hydrophobic
SUPPLEMENTARY INFORMATION
 Table of Contents Page Supplementary Table 1. Diffraction data collection statistics 2 Supplementary Table 2. Crystallographic refinement statistics 3 Supplementary Fig. 1. casic1mfc packing in the R3
Table of Contents Page Supplementary Table 1. Diffraction data collection statistics 2 Supplementary Table 2. Crystallographic refinement statistics 3 Supplementary Fig. 1. casic1mfc packing in the R3
Supplemental Information. The Mitochondrial Fission Receptor MiD51. Requires ADP as a Cofactor
 Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Structure, Volume 22 Supplemental Information The Mitochondrial Fission Receptor MiD51 Requires ADP as a Cofactor Oliver C. Losón, Raymond Liu, Michael E. Rome, Shuxia Meng, Jens T. Kaiser, Shu-ou Shan,
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/10/eaat8797/dc1 Supplementary Materials for Single-molecule observation of nucleotide induced conformational changes in basal SecA-ATP hydrolysis Nagaraju Chada,
advances.sciencemag.org/cgi/content/full/4/10/eaat8797/dc1 Supplementary Materials for Single-molecule observation of nucleotide induced conformational changes in basal SecA-ATP hydrolysis Nagaraju Chada,
downstream (0.8 kb) homologous sequences to the genomic locus of DIC. A DIC mutant strain (ro- 6
 A B C D ts Figure S1 Generation of DIC- mcherry expressing N.crassa strain. A. N. crassa colony morphology. When a cot1 (top, left panel) strain is grown at permissive temperature (25 C), it exhibits straight
A B C D ts Figure S1 Generation of DIC- mcherry expressing N.crassa strain. A. N. crassa colony morphology. When a cot1 (top, left panel) strain is grown at permissive temperature (25 C), it exhibits straight
Supplementary Figure 1. Voltage clamp speed. Capacity membrane current in response to a 4- mv voltage step (black). Solid red line corresponds to a
 Supplementary Figure 1. Voltage clamp speed. Capacity membrane current in response to a 4- mv voltage step (black). Solid red line corresponds to a mono-exponential fit with a time constant of 5.5 µs.
Supplementary Figure 1. Voltage clamp speed. Capacity membrane current in response to a 4- mv voltage step (black). Solid red line corresponds to a mono-exponential fit with a time constant of 5.5 µs.
Supplementary Figure 1 Structure of the Orai channel. (a) The hexameric Drosophila Orai channel structure derived from crystallography 1 comprises
 Supplementary Figure 1 Structure of the Orai channel. (a) The hexameric Drosophila Orai channel structure derived from crystallography 1 comprises six Orai subunits, each with identical amino acid sequences
Supplementary Figure 1 Structure of the Orai channel. (a) The hexameric Drosophila Orai channel structure derived from crystallography 1 comprises six Orai subunits, each with identical amino acid sequences
Sensitive NMR Approach for Determining the Binding Mode of Tightly Binding Ligand Molecules to Protein Targets
 Supporting information Sensitive NMR Approach for Determining the Binding Mode of Tightly Binding Ligand Molecules to Protein Targets Wan-Na Chen, Christoph Nitsche, Kala Bharath Pilla, Bim Graham, Thomas
Supporting information Sensitive NMR Approach for Determining the Binding Mode of Tightly Binding Ligand Molecules to Protein Targets Wan-Na Chen, Christoph Nitsche, Kala Bharath Pilla, Bim Graham, Thomas
Supporting online material
 Supporting online material Materials and Methods Target proteins All predicted ORFs in the E. coli genome (1) were downloaded from the Colibri data base (2) (http://genolist.pasteur.fr/colibri/). 737 proteins
Supporting online material Materials and Methods Target proteins All predicted ORFs in the E. coli genome (1) were downloaded from the Colibri data base (2) (http://genolist.pasteur.fr/colibri/). 737 proteins
Structural characterization of NiV N 0 P in solution and in crystal.
 Supplementary Figure 1 Structural characterization of NiV N 0 P in solution and in crystal. (a) SAXS analysis of the N 32-383 0 -P 50 complex. The Guinier plot for complex concentrations of 0.55, 1.1,
Supplementary Figure 1 Structural characterization of NiV N 0 P in solution and in crystal. (a) SAXS analysis of the N 32-383 0 -P 50 complex. The Guinier plot for complex concentrations of 0.55, 1.1,
 DOI: 10.1038/ncb2819 Gαi3 / Actin / Acetylated Tubulin Gαi3 / Actin / Acetylated Tubulin a a Gαi3 a Actin Gαi3 WT Gαi3 WT Gαi3 WT b b Gαi3 b Actin Gαi3 KO Gαi3 KO Gαi3 KO # # Figure S1 Loss of protein
DOI: 10.1038/ncb2819 Gαi3 / Actin / Acetylated Tubulin Gαi3 / Actin / Acetylated Tubulin a a Gαi3 a Actin Gαi3 WT Gαi3 WT Gαi3 WT b b Gαi3 b Actin Gαi3 KO Gαi3 KO Gαi3 KO # # Figure S1 Loss of protein
Supplemental material
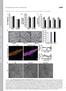 Supplemental material THE JOURNAL OF CELL BIOLOGY Mourier et al., http://www.jcb.org/cgi/content/full/jcb.201411100/dc1 Figure S1. Size and mitochondrial content in Mfn1 and Mfn2 knockout hearts. (A) Body
Supplemental material THE JOURNAL OF CELL BIOLOGY Mourier et al., http://www.jcb.org/cgi/content/full/jcb.201411100/dc1 Figure S1. Size and mitochondrial content in Mfn1 and Mfn2 knockout hearts. (A) Body
Supplementary material
 Supplementary material Phosphorylation of the mitochondrial autophagy receptor Nix enhances its interaction with LC3 proteins Vladimir V. Rogov 1,*, Hironori Suzuki 2,3,*, Mija Marinković 4, Verena Lang
Supplementary material Phosphorylation of the mitochondrial autophagy receptor Nix enhances its interaction with LC3 proteins Vladimir V. Rogov 1,*, Hironori Suzuki 2,3,*, Mija Marinković 4, Verena Lang
SUPPLEMENTARY INFORMATION
 Supplementary Notes Downregulation of atlastin does not affect secretory traffic We investigated whether Atlastin might play a role in secretory traffic. Traffic impairment results in disruption of Golgi
Supplementary Notes Downregulation of atlastin does not affect secretory traffic We investigated whether Atlastin might play a role in secretory traffic. Traffic impairment results in disruption of Golgi
Structure and evolution of the spliceosomal peptidyl-prolyl cistrans isomerase Cwc27
 Acta Cryst. (2014). D70, doi:10.1107/s1399004714021695 Supporting information Volume 70 (2014) Supporting information for article: Structure and evolution of the spliceosomal peptidyl-prolyl cistrans isomerase
Acta Cryst. (2014). D70, doi:10.1107/s1399004714021695 Supporting information Volume 70 (2014) Supporting information for article: Structure and evolution of the spliceosomal peptidyl-prolyl cistrans isomerase
Chapter 4 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays.
 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Structure and mechanism of an intramembrane liponucleotide synthetase central for phospholipid biosynthesis
 Structure and mechanism of an intramembrane liponucleotide synthetase central for phospholipid biosynthesis Xiuying Liu 1,3, Yan Yin 1,2,3, Jinjun Wu 1 and Zhenfeng Liu 1 1 National Laboratory of Biomacromolecules,
Structure and mechanism of an intramembrane liponucleotide synthetase central for phospholipid biosynthesis Xiuying Liu 1,3, Yan Yin 1,2,3, Jinjun Wu 1 and Zhenfeng Liu 1 1 National Laboratory of Biomacromolecules,
Bacterial Outer Membrane Porins as Electrostatic Nanosieves: Exploring Transport Rules of Small Polar Molecules
 Bacterial Outer Membrane Porins as Electrostatic Nanosieves: Exploring Transport Rules of Small Polar Molecules Harsha Bajaj, Silvia Acosta Gutiérrez, Igor Bodrenko, Giuliano Malloci, Mariano Andrea Scorciapino,
Bacterial Outer Membrane Porins as Electrostatic Nanosieves: Exploring Transport Rules of Small Polar Molecules Harsha Bajaj, Silvia Acosta Gutiérrez, Igor Bodrenko, Giuliano Malloci, Mariano Andrea Scorciapino,
13-3. Synthesis-Secretory pathway: Sort lumenal proteins, Secrete proteins, Sort membrane proteins
 13-3. Synthesis-Secretory pathway: Sort lumenal proteins, Secrete proteins, Sort membrane proteins Molecular sorting: specific budding, vesicular transport, fusion 1. Why is this important? A. Form and
13-3. Synthesis-Secretory pathway: Sort lumenal proteins, Secrete proteins, Sort membrane proteins Molecular sorting: specific budding, vesicular transport, fusion 1. Why is this important? A. Form and
Supplementary Figure 1.
 Supplementary Figure 1. Characterisation of IHG-1 overexpressing and knockdown cell lines. (A) Total cellular RNA was prepared from HeLa cells stably overexpressing IHG-1 or mts-ihg-1. IHG-1 mrna was quantified
Supplementary Figure 1. Characterisation of IHG-1 overexpressing and knockdown cell lines. (A) Total cellular RNA was prepared from HeLa cells stably overexpressing IHG-1 or mts-ihg-1. IHG-1 mrna was quantified
Open Syntaxin Docks Synaptic Vesicles
 Marc Hammarlund 1,2[, Mark T. Palfreyman 1,2[, Shigeki Watanabe 1,2[, Shawn Olsen 1,2, Erik M. Jorgensen 1,2* PLoS BIOLOGY 1 Department of Biology, University of Utah, Salt Lake City, Utah, United States
Marc Hammarlund 1,2[, Mark T. Palfreyman 1,2[, Shigeki Watanabe 1,2[, Shawn Olsen 1,2, Erik M. Jorgensen 1,2* PLoS BIOLOGY 1 Department of Biology, University of Utah, Salt Lake City, Utah, United States
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10955 Supplementary Figures Supplementary Figure 1. Electron-density maps and crystallographic dimer structures of the motor domain. (a f) Stereo views of the final electron-density maps
doi:10.1038/nature10955 Supplementary Figures Supplementary Figure 1. Electron-density maps and crystallographic dimer structures of the motor domain. (a f) Stereo views of the final electron-density maps
Supplementary figure 1. Comparison of unbound ogm-csf and ogm-csf as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine
 Supplementary figure 1. Comparison of unbound and as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine GM-CSF (slate) with bound GM-CSF in the GIF:GM-CSF complex (GIF: green,
Supplementary figure 1. Comparison of unbound and as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine GM-CSF (slate) with bound GM-CSF in the GIF:GM-CSF complex (GIF: green,
Waithe et al Supplementary Figures
 Waithe et al Supplementary Figures Supplementary Figure 1 Expression and properties of WT and W391A mutant YFP- Ca V 2.2. A Immunoblot using Ca V 2.2 Ab for untransfected cells (UT, lane 1), YFP-Ca V 2.2
Waithe et al Supplementary Figures Supplementary Figure 1 Expression and properties of WT and W391A mutant YFP- Ca V 2.2. A Immunoblot using Ca V 2.2 Ab for untransfected cells (UT, lane 1), YFP-Ca V 2.2
Supplementary Figure S1. MscS orientation in spheroplasts and liposomes (a) Current-voltage relationship for wild-type MscS expressed in E.
 a b c Supplementary Figure S1. MscS orientation in spheroplasts and liposomes (a) Current-voltage relationship for wild-type MscS expressed in E. coli giant spheroplasts (MJF465) and reconstituted into
a b c Supplementary Figure S1. MscS orientation in spheroplasts and liposomes (a) Current-voltage relationship for wild-type MscS expressed in E. coli giant spheroplasts (MJF465) and reconstituted into
SUPPLEMENTARY INFORMATION
 Figure S1. Secondary structure of CAP (in the camp 2 -bound state) 10. α-helices are shown as cylinders and β- strands as arrows. Labeling of secondary structure is indicated. CDB, DBD and the hinge are
Figure S1. Secondary structure of CAP (in the camp 2 -bound state) 10. α-helices are shown as cylinders and β- strands as arrows. Labeling of secondary structure is indicated. CDB, DBD and the hinge are
Anterograde Activin Signaling Regulates Postsynaptic Membrane Potential and GluRIIA/B Abundance at the Drosophila Neuromuscular Junction
 Anterograde Activin Signaling Regulates Postsynaptic Membrane Potential and GluRIIA/B Abundance at the Drosophila Neuromuscular Junction Myung-Jun Kim, Michael B. O Connor* Department of Genetics, Cell
Anterograde Activin Signaling Regulates Postsynaptic Membrane Potential and GluRIIA/B Abundance at the Drosophila Neuromuscular Junction Myung-Jun Kim, Michael B. O Connor* Department of Genetics, Cell
Role of Mitochondrial Remodeling in Programmed Cell Death in
 Developmental Cell, Vol. 12 Supplementary Data Role of Mitochondrial Remodeling in Programmed Cell Death in Drosophila melanogaster Gaurav Goyal, Brennan Fell, Apurva Sarin, Richard J. Youle, V. Sriram.
Developmental Cell, Vol. 12 Supplementary Data Role of Mitochondrial Remodeling in Programmed Cell Death in Drosophila melanogaster Gaurav Goyal, Brennan Fell, Apurva Sarin, Richard J. Youle, V. Sriram.
Supplementary Figure 3 a. Structural comparison between the two determined structures for the IL 23:MA12 complex. The overall RMSD between the two
 Supplementary Figure 1. Biopanningg and clone enrichment of Alphabody binders against human IL 23. Positive clones in i phage ELISA with optical density (OD) 3 times higher than background are shown for
Supplementary Figure 1. Biopanningg and clone enrichment of Alphabody binders against human IL 23. Positive clones in i phage ELISA with optical density (OD) 3 times higher than background are shown for
Examples of Protein Modeling. Protein Modeling. Primary Structure. Protein Structure Description. Protein Sequence Sources. Importing Sequences to MOE
 Examples of Protein Modeling Protein Modeling Visualization Examination of an experimental structure to gain insight about a research question Dynamics To examine the dynamics of protein structures To
Examples of Protein Modeling Protein Modeling Visualization Examination of an experimental structure to gain insight about a research question Dynamics To examine the dynamics of protein structures To
SUPPLEMENTARY INFORMATION
 Supplementary materials Figure S1 Fusion protein of Sulfolobus solfataricus SRP54 and a signal peptide. a, Expression vector for the fusion protein. The signal peptide of yeast dipeptidyl aminopeptidase
Supplementary materials Figure S1 Fusion protein of Sulfolobus solfataricus SRP54 and a signal peptide. a, Expression vector for the fusion protein. The signal peptide of yeast dipeptidyl aminopeptidase
7.06 Spring 2004 PS 6 KEY 1 of 14
 7.06 Spring 2004 PS 6 KEY 1 of 14 Problem Set 6. Question 1. You are working in a lab that studies hormones and hormone receptors. You are tasked with the job of characterizing a potentially new hormone
7.06 Spring 2004 PS 6 KEY 1 of 14 Problem Set 6. Question 1. You are working in a lab that studies hormones and hormone receptors. You are tasked with the job of characterizing a potentially new hormone
RNA Polymerase I Contains a TFIIF-Related DNA-Binding Subcomplex
 Molecular Cell, Volume 39 Supplemental Information RNA Polymerase I Contains a TFIIFRelated DNABinding Subcomplex Sebastian R. Geiger, Kristina Lorenzen, Amelie Schreieck, Patrizia Hanecker, Dirk Kostrewa,
Molecular Cell, Volume 39 Supplemental Information RNA Polymerase I Contains a TFIIFRelated DNABinding Subcomplex Sebastian R. Geiger, Kristina Lorenzen, Amelie Schreieck, Patrizia Hanecker, Dirk Kostrewa,
EST1 Homology Domain. 100 aa. hest1a / SMG6 PIN TPR TPR. Est1-like DBD? hest1b / SMG5. TPR-like TPR. a helical. hest1c / SMG7.
 hest1a / SMG6 EST1 Homology Domain 100 aa 853 695 761 780 1206 hest1 / SMG5 -like? -like 109 145 214 237 497 165 239 1016 114 207 212 381 583 hest1c / SMG7 a helical 1091 Sc 57 185 267 284 699 Figure S1:
hest1a / SMG6 EST1 Homology Domain 100 aa 853 695 761 780 1206 hest1 / SMG5 -like? -like 109 145 214 237 497 165 239 1016 114 207 212 381 583 hest1c / SMG7 a helical 1091 Sc 57 185 267 284 699 Figure S1:
7.06 Cell Biology EXAM #3 KEY
 7.06 Cell Biology EXAM #3 KEY May 2, 2006 This is an OPEN BOOK exam, and you are allowed access to books, a calculator, and notes BUT NOT computers or any other types of electronic devices. Please write
7.06 Cell Biology EXAM #3 KEY May 2, 2006 This is an OPEN BOOK exam, and you are allowed access to books, a calculator, and notes BUT NOT computers or any other types of electronic devices. Please write
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Structure of human carbamoyl phosphate synthetase: deciphering the on/off switch of human ureagenesis Sergio de Cima, Luis M. Polo, Carmen Díez-Fernández, Ana I. Martínez, Javier
SUPPLEMENTARY INFORMATION Structure of human carbamoyl phosphate synthetase: deciphering the on/off switch of human ureagenesis Sergio de Cima, Luis M. Polo, Carmen Díez-Fernández, Ana I. Martínez, Javier
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11744 Supplementary Table 1. Crystallographic data collection and refinement statistics. Wild-type Se-Met-BcsA-B SmCl 3 -soaked EMTS-soaked Data collection Space
SUPPLEMENTARY INFORMATION doi:10.1038/nature11744 Supplementary Table 1. Crystallographic data collection and refinement statistics. Wild-type Se-Met-BcsA-B SmCl 3 -soaked EMTS-soaked Data collection Space
Nature Genetics: doi: /ng Supplementary Figure 1. The phenotypes of PI , BR121, and Harosoy under short-day conditions.
 Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Modulation of central pattern generator output by peripheral sensory cells in Drosophila larvae. BioNB4910 Cornell University.
 Modulation of central pattern generator output by peripheral sensory cells in Drosophila larvae BioNB4910 Cornell University Goals 1) Observe the behavioral effects of remotely activating different populations
Modulation of central pattern generator output by peripheral sensory cells in Drosophila larvae BioNB4910 Cornell University Goals 1) Observe the behavioral effects of remotely activating different populations
Structural basis of PROTAC cooperative recognition for selective protein degradation
 SUPPLEMENTARY INFORMATION Structural basis of PROTAC cooperative recognition for selective protein degradation Morgan S. Gadd 1, Andrea Testa 1, Xavier Lucas 1, Kwok-Ho Chan, Wenzhang Chen, Douglas J.
SUPPLEMENTARY INFORMATION Structural basis of PROTAC cooperative recognition for selective protein degradation Morgan S. Gadd 1, Andrea Testa 1, Xavier Lucas 1, Kwok-Ho Chan, Wenzhang Chen, Douglas J.
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Overexpression of YFP::GPR-1 in the germline.
 Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
TNFα 18hr. Control. CHX 18hr. TNFα+ CHX 18hr. TNFα: 18 18hr (KDa) PARP. Cleaved. Cleaved. Cleaved. Caspase3. Pellino3 shrna. Control shrna.
 Survival ( %) a. TNFα 18hr b. Control sirna Pellino3 sirna TNFα: 18 18hr c. Control shrna Pellino3 shrna Caspase3 Actin Control d. Control shrna Pellino3 shrna *** 100 80 60 CHX 18hr 40 TNFα+ CHX 18hr
Survival ( %) a. TNFα 18hr b. Control sirna Pellino3 sirna TNFα: 18 18hr c. Control shrna Pellino3 shrna Caspase3 Actin Control d. Control shrna Pellino3 shrna *** 100 80 60 CHX 18hr 40 TNFα+ CHX 18hr
7.06 Problem Set #4, Spring 2005
 7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
Impact of the crystallization condition on importin-β conformation
 Supporting information Volume 72 (2016) Supporting information for article: Impact of the crystallization condition on importin-β conformation Marcel J. Tauchert, Clément Hémonnot, Piotr Neumann, Sarah
Supporting information Volume 72 (2016) Supporting information for article: Impact of the crystallization condition on importin-β conformation Marcel J. Tauchert, Clément Hémonnot, Piotr Neumann, Sarah
Supplementary Information. Overlap between folding and functional energy landscapes for. adenylate kinase conformational change
 Supplementary Information Overlap between folding and functional energy landscapes for adenylate kinase conformational change by Ulrika Olsson & Magnus Wolf-Watz Contents: 1. Supplementary Note 2. Supplementary
Supplementary Information Overlap between folding and functional energy landscapes for adenylate kinase conformational change by Ulrika Olsson & Magnus Wolf-Watz Contents: 1. Supplementary Note 2. Supplementary
T H E J O U R N A L O F C E L L B I O L O G Y
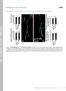 Supplemental material Kasprowicz et al., http://www.jcb.org/cgi/content/full/jcb.201310090/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. NMJ morphology of shi 12-12B ; shi-4c not treated
Supplemental material Kasprowicz et al., http://www.jcb.org/cgi/content/full/jcb.201310090/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. NMJ morphology of shi 12-12B ; shi-4c not treated
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Purification of yeast CKM. (a) Silver-stained SDS-PAGE analysis of CKM purified through a TAP-tag engineered into the Cdk8 C-terminus. (b) Kinase activity
Supplementary Figures Supplementary Figure 1. Purification of yeast CKM. (a) Silver-stained SDS-PAGE analysis of CKM purified through a TAP-tag engineered into the Cdk8 C-terminus. (b) Kinase activity
Supplementary Information
 Supplementary Information Resveratrol Serves as a Protein-Substrate Interaction Stabilizer in Human SIRT1 Activation Xuben Hou,, David Rooklin, Hao Fang *,,, Yingkai Zhang Department of Medicinal Chemistry
Supplementary Information Resveratrol Serves as a Protein-Substrate Interaction Stabilizer in Human SIRT1 Activation Xuben Hou,, David Rooklin, Hao Fang *,,, Yingkai Zhang Department of Medicinal Chemistry
SUPPLEMENTARY INFORMATION
 doi:1.138/nature1737 Supplementary Table 1 variant Description FSEC - 2B12 a FSEC - 6A1 a K d (leucine) c Leucine uptake e K (wild-type like) K (Y18F) K (TS) K (TSY) K288A mutant, lipid facing side chain
doi:1.138/nature1737 Supplementary Table 1 variant Description FSEC - 2B12 a FSEC - 6A1 a K d (leucine) c Leucine uptake e K (wild-type like) K (Y18F) K (TS) K (TSY) K288A mutant, lipid facing side chain
Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A)
 Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the
Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the
Binding Studies on Trafficking Proteins Using Microcalorimetry McMahon lab
 Binding Studies on Trafficking Proteins Using Microcalorimetry McMahon lab Neurobiology Division Laboratory of Molecular Biology Cambridge Clathrin Mediated Endocytosis Binding Recruitment Coating Budding
Binding Studies on Trafficking Proteins Using Microcalorimetry McMahon lab Neurobiology Division Laboratory of Molecular Biology Cambridge Clathrin Mediated Endocytosis Binding Recruitment Coating Budding
Supplemental Data. Gao et al. (2012). Plant Cell /tpc
 Supplemental Figure 1. Plant EMP Proteins. (A) The Accession numbers of the 12 EMP members from Arabidopsis. (B) Phylogenetic analysis of EMP proteins from Arabidopsis, human and yeast using the Mac Vector
Supplemental Figure 1. Plant EMP Proteins. (A) The Accession numbers of the 12 EMP members from Arabidopsis. (B) Phylogenetic analysis of EMP proteins from Arabidopsis, human and yeast using the Mac Vector
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11054 Supplementary Fig. 1 Sequence alignment of Na v Rh with NaChBac, Na v Ab, and eukaryotic Na v and Ca v homologs. Secondary structural elements of Na v Rh are indicated above the
doi:10.1038/nature11054 Supplementary Fig. 1 Sequence alignment of Na v Rh with NaChBac, Na v Ab, and eukaryotic Na v and Ca v homologs. Secondary structural elements of Na v Rh are indicated above the
Cks1 CDK1 CDK1 CDK1 CKS1. are ice- lobe. conserved. conserved
 Cks1 d CKS1 Supplementary Figure 1 The -Cks1 crystal lattice. (a) Schematic of the - Cks1 crystal lattice. -Cks1 crystallizes in a lattice that contains c 4 copies of the t - Cks1 dimer in the crystallographic
Cks1 d CKS1 Supplementary Figure 1 The -Cks1 crystal lattice. (a) Schematic of the - Cks1 crystal lattice. -Cks1 crystallizes in a lattice that contains c 4 copies of the t - Cks1 dimer in the crystallographic
Peptide-derived Inhibitors of Protein-Protein Interactions
 Peptide-derived Inhibitors of Protein-Protein Interactions Sven Hennig Department of Chemistry and Pharmaceutical Sciences Vrije Universiteit Amsterdam 1 Biomolecular recognitions Classification via interaction
Peptide-derived Inhibitors of Protein-Protein Interactions Sven Hennig Department of Chemistry and Pharmaceutical Sciences Vrije Universiteit Amsterdam 1 Biomolecular recognitions Classification via interaction
Analysis of nucleotide binding to p97 reveals the properties of a tandem AAA hexameric ATPase
 SUPPLEMENTARY INFORMATION Analysis of nucleotide binding to p97 reveals the properties of a tandem AAA hexameric ATPase Louise C Briggs, Geoff S Baldwin, Non Miyata, Hisao Kondo, Xiaodong Zhang, Paul S
SUPPLEMENTARY INFORMATION Analysis of nucleotide binding to p97 reveals the properties of a tandem AAA hexameric ATPase Louise C Briggs, Geoff S Baldwin, Non Miyata, Hisao Kondo, Xiaodong Zhang, Paul S
Supplementary Information. The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes
 Supplementary Information The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes Xuesong Shi, a Lin Huang, b David M. J. Lilley, b Pehr B. Harbury a,c and Daniel Herschlag
Supplementary Information The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes Xuesong Shi, a Lin Huang, b David M. J. Lilley, b Pehr B. Harbury a,c and Daniel Herschlag
Tex 25mer ssrna Binding Stoichiometry
 Figure S. Determination of Tex:2nt ssrna binding stoichiometry using fluorescence polarization. Fluorescein labeled RNA was held at a constant concentration 2-fold above the K d. Tex protein was titrated
Figure S. Determination of Tex:2nt ssrna binding stoichiometry using fluorescence polarization. Fluorescein labeled RNA was held at a constant concentration 2-fold above the K d. Tex protein was titrated
We used the PSI-BLAST program (http://www.ncbi.nlm.nih.gov/blast/) to search the
 SUPPLEMENTARY METHODS - in silico protein analysis We used the PSI-BLAST program (http://www.ncbi.nlm.nih.gov/blast/) to search the Protein Data Bank (PDB, http://www.rcsb.org/pdb/) and the NCBI non-redundant
SUPPLEMENTARY METHODS - in silico protein analysis We used the PSI-BLAST program (http://www.ncbi.nlm.nih.gov/blast/) to search the Protein Data Bank (PDB, http://www.rcsb.org/pdb/) and the NCBI non-redundant
Supplementary Figures for Tong et al.: Structure and function of the intracellular region of the plexin-b1 transmembrane receptor
 Supplementary Figures for Tong et al.: Structure and function of the intracellular region of the plexin-b1 transmembrane receptor Figure S1. Plexin-B1 GAP segments are homologous to RasGAPs. Sequence alignment
Supplementary Figures for Tong et al.: Structure and function of the intracellular region of the plexin-b1 transmembrane receptor Figure S1. Plexin-B1 GAP segments are homologous to RasGAPs. Sequence alignment
Supporting Information
 Supporting Information Mullins et al. 10.1073/pnas.0906781106 SI Text Detection of Calcium Binding by 45 Ca 2 Overlay. The 45 CaCl 2 (1 mci, 37 MBq) was obtained from NEN. The general method of 45 Ca 2
Supporting Information Mullins et al. 10.1073/pnas.0906781106 SI Text Detection of Calcium Binding by 45 Ca 2 Overlay. The 45 CaCl 2 (1 mci, 37 MBq) was obtained from NEN. The general method of 45 Ca 2
Supplementary Information
 Supplementary Information Supplementary Figure 1. JAK/STAT in early wing development (a-f) Wing primordia of second instar larvae of the indicated genotypes labeled to visualize expression of upd mrna
Supplementary Information Supplementary Figure 1. JAK/STAT in early wing development (a-f) Wing primordia of second instar larvae of the indicated genotypes labeled to visualize expression of upd mrna
Supporting Information
 Supporting Information Reaction Mechanism of Adenylyltransferase DrrA from Legionella pneumophila Elucidated by Time-Resolved Fourier Transform Infrared Spectroscopy Konstantin Gavriljuk, Jonas Schartner,
Supporting Information Reaction Mechanism of Adenylyltransferase DrrA from Legionella pneumophila Elucidated by Time-Resolved Fourier Transform Infrared Spectroscopy Konstantin Gavriljuk, Jonas Schartner,
Molecular dynamics simulations of anti-aggregation effect of ibuprofen. Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov
 Biophysical Journal, Volume 98 Supporting Material Molecular dynamics simulations of anti-aggregation effect of ibuprofen Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov Supplemental
Biophysical Journal, Volume 98 Supporting Material Molecular dynamics simulations of anti-aggregation effect of ibuprofen Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov Supplemental
SUPPLEMENTARY INFORMATION
 www.nature.com/nature 1 Figure S1 Sequence alignment. a Structure based alignment of the plgic of E. chrysanthemi (ELIC), the acetylcholine binding protein from the snail Lymnea stagnalis (AchBP, PDB code
www.nature.com/nature 1 Figure S1 Sequence alignment. a Structure based alignment of the plgic of E. chrysanthemi (ELIC), the acetylcholine binding protein from the snail Lymnea stagnalis (AchBP, PDB code
