Myosin V Passing over Arp2/3 Junctions: Branching Ratio Calculated from the Elastic Lever Arm Model
|
|
|
- Willis Marshall
- 5 years ago
- Views:
Transcription
1 Biophysical Journal Volume 94 May Myosin V Passing over Arp2/3 Junctions: Branching Ratio Calculated from the Elastic Lever Arm Model Andrej Vilfan J. Stefan Institute, Ljubljana, Slovenia ABSTRACT Myosin V is a two-headed processive motor protein that walks in a hand-over-hand fashion along actin filaments. When it encounters a filament branch, formed by the Arp2/3 complex, it can either stay on the straight mother filament, or switch to the daughter filament. We study both probabilities using the elastic lever arm model for myosin V. We calculate the shapes and bending energies of all relevant configurations in which the trail head is bound to the actin filament before Arp2/3 and the lead head is bound either to the mother or to the daughter filament. Based on the assumption that the probability for a head to bind to a certain actin subunit is proportional to the Boltzmann factor obtained from the elastic energy, we calculate the mother/ daughter filament branching ratio. Our model predicts a value of 27% for the daughter and 73% for the mother filament. This result is in good agreement with recent experimental data. INTRODUCTION Myosin V is a two-headed processive motor protein from the myosin superfamily, involved in different forms of intracellular transport (,2). It has drawn a lot of attention in recent years and is now one of the best studied motor proteins. The experiments have characterized it mechanically (3 ), biochemically ( 4), optically (5 7), and structurally (8 2). These studies have shown that myosin V walks along actin filaments in a hand-over-hand fashion (5,22) with an average step size of ;35 nm, roughly corresponding to the periodicity of actin filaments (3,4,6,6,8), a stall force of ;2 pn (3), and a run length of a few microns (3,23,24). Under physiological conditions, ADP release has been identified as the time-limiting step in the duty cycle (3,). The Arp2/3 complex (25) initiates actin filament branching in the vicinity of a protruding edge of a cell. The complex consists of seven subunits (Arp2, Arp3, and ARPC through ARPC5) and is activated by WASp/Scar proteins (26). It binds to the side of one ( mother ) actin filament and initiates the nucleation of a second ( daughter filament), which starts growing with the fast growing end ( end) away from the Arp2/3 complex. The mother and the daughter filament enclose an angle of 7. The question of what happens to a myosin V motor when it arrives at an Arp2/3-mediated actin filament junction is of interest for several reasons. First, the branching behavior is important for understanding vesicle transport in the actin cortex. And second, it is of high interest when studying the fundamental mechanism of myosin V stepping, because it represents a well defined situation in which predictions from different theoretical models can be tested against the experimental result. Our aim in this article is to use the elastic lever arm model for myosin V, which is described in detail in a Submitted August 27, 27, and accepted for publication December, 27. Address reprint requests to Andrej Vilfan, Tel.: ; andrej.vilfan@ijs.si. Editor: E. Michael Ostap. Ó 28 by the Biophysical Society /8/5/345/8 $2. previous article (27), to predict the dynamics of a myosin V motor that passes over an Arp2/3 junction. In particular, we will calculate the probabilities that a motor continues along the mother or the daughter filament. MODEL The idea behind the elastic lever arm model for myosin V is to describe the dimeric motor as an assembly of two identical heads, connected to each other and to the cargo-binding tail with elastic lever arms. The model allows us to derive the properties of a dimeric molecule, such as step size distribution, force-velocity relation, and processivity, from the properties of an individual head, such as geometry, chemical kinetics, and elasticity. In this respect, the approach is different from the class of discrete stochastic models, which describe the motor as a single unit (28,29). We describe each head with a five-state mechano-chemical model, similar to that for muscle myosin (e.g., (3)), where each state (with bound ADP.Pi; ADP, pre-powerstroke; ADP, post-powerstroke; without a nucleotide; detached) is connected with a certain orientation of the lever arm, as determined with electron microscopy (8,9). The lever arms are modeled as elastic beams, connected with a flexible joint. A recent study measuring fluctuations in the position of the free head (3) has demonstrated that the lead head diffuses around the joint freely before binding to the next actin site, meaning that there is no detectable elastic energy cost connected with variation in the angle between the two lever arms. The very nature of protein flexibility, which mainly originates from the twisting of bonds between carbon atoms in the backbone, leads us to the conclusion that the joint is also fully flexible with regard to rotation of each lever arm along its axis. Similar flexibility has also been observed in myosin II (32 34). The calculation of the branching probability is greatly simplified if we make the following assumptions. First, we doi:.529/biophysj
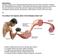
2 346 Vilfan assume that the binding of the lead head always advances to a step, which means that its unbinding is significantly slower than the step that follows in the regular cycle (Pi release). Second, we assume that the probability for the lead head to bind to site j if the trail head is bound to site i is given by the Boltzmann factor Ui;j exp P jji ¼ k B T + j9 exp U i;j9 k B T : () U i, j denotes the elastic energy of deformed lever arms when the trail head is in the post-powerstroke state, bound to site i, and the lead head in the pre-powerstroke state to site j. In the following, we will use the notation where the sites on the mother filament are marked with (M,i) and those on the daughter filament with (D,i). For example, P D,2jM, 9 denotes the conditional probability for the lead head to bind to site 2 on the daughter filament if the trail head is bound to the site 9 on the mother filament. We enumerate the actin subunits so that the central subunit under the Arp2/3 complex on the mother filament has the index. Positive indices denote subunits toward the plus () end and negative toward the minus ( ) end. Subunits of the daughter filament are enumerated from onwards. Note that sites numbered 2,, and 2 on the mother filament are not accessible for a myosin V head, because of steric hindrance with the Arp2/3 complex. The structure of the Arp2/3 complex and both actin filaments (Fig. ) has been determined from electron microscopy studies (35,36) and its parameters are summarized in Table. While we approximated actin with the commonly assumed 3/6 helix in the original article (27), we use a more accurate 28/3 helix here, with the angle u ¼ 67.4 between adjacent subunits. A detailed discussion on different helix models and their consequence for the calculated step size distribution can be found in Vilfan (37). When a head is bound to the site i, the starting point and the unit vector giving the initial direction of the lever arm are given by x ¼ R x ð iu Þ@ ia d R A ˆt ¼ R x ð iu Þ@ cosf i sinf i A; (2) where R x denotes the rotation matrix around the x axis, R x ðuþ ¼@ cosu sinu A: (3) sinu cosu For a head bound to the daughter filament, the two vectors read r ia d B C x ¼ R y ð bþr x ðg iu Þ@ A R cosf i B C ˆt ¼ R y ð bþr x ðg iu Þ@ A; (4) sinf i with cosb sinb R y ðbþ ¼@ A: (5) sinb cosb Here b ¼ 7 denotes the angle between the mother and the daughter filament and g ¼ 39 the rotation of the daughter filament around its axis (see Fig. ). We calculate the shapes of both lever arms as described in Vilfan (27) by minimizing the bending energy U ¼ R ds EIðCðsÞÞ 2 =2; where C(s) denotes local curvature. For the bending modulus of the lever arm we use the value EI ¼ 5 FIGURE Arp2/3 junction and a dimeric myosin V motor. (a)thearp2/3 complex is attached to the side of the mother filament occupying subunits 2,, and 2. It nucleates the growth of a daughter filament, whose position is determined by the angles b (branching angle), g (twist), and r (distance from the first subunit to the center of the mother filament). (b) The myosin V motor consists of two heads, connected with lever arms, which we describe as elastic rods. The proximal end of each lever arm always leaves the head at a fixed angle f, which depends on the nucleotide state of that head. The distal ends of both lever arms are connected with a fully flexible joint. TABLE Geometric parameters of the Arp2/3 junction and a myosin V head Parameter Symbol Value Distance actin subunits a 2.75 nm Angle actin subunits u 67.4 Daughter filament angle b 7 Daughter filament offset r 2 nm Daughter filament rotation g 39 Lever arm start: radial position R 8nm Lever arm start: displacement d pre PS d post PS 3.5 nm Lever arm angle f pre PS 5 f post PS 5 Lever arm length L 26 nm
3 Myosin V Passing over Arp2/3 Junctions 347 pn nm 2, which corresponds to a spring constant of k ¼ 3EI/ L 3 ¼.25 pn/nm, measured at the tip of the lever arm. This value was originally estimated from the stall force, but it shows good agreement with direct optical tweezer measurements (38). We neglect any additional compliance resulting from the head or converter domain. Because most of the bending takes place in the proximal part of the lever arm, we expect that its effect would not be significantly different. For the temperature we use the value T ¼ 27 C. RESULTS Stepping of myosin V in the absence of the Arp2/3 complex When a myosin V motor is sufficiently far away from the Arp2/3 complex, it exhibits the stepping pattern that has already been discussed (27,37). We restrict the step lengths of an unperturbed motor to, 3, and 5 subunits, with probabilities P ¼ P M; ijm; i :5; (6) P 3 ¼ P M; i3jm; i :895; (7) P 5 ¼ P M; i5jm; i :; (8) calculated from Eq.. Probabilities for other step sizes, such as P 9 and P 7 turn out to be very small. Note that, in this calculation, P and P 5 are likely to be somewhat underestimated; their values are somewhat higher if we take into account torsional fluctuations in the actin helix (37). The average step size can be calculated from these probabilities as l ¼ P 3P 3 5P 5 3:2: (9) In the absence of the Arp2/3 junction, the fraction of sites that get accessed by a passing myosin V motor is = l: This is also FIGURE 2 Different pathways on which a myosin V motor can approach the Arp2/3 junction. The probability distributions for the first accessed site in the two intervals marked with dashed lines are denoted as P I,i and P J,i. P I,i represents the probability that i is the first accessed site with 7 # i # 3 (between dashed lines in the first or second column). For values between 7 and 7, this state can be reached with three different step sizes. For i ¼ 6and i ¼ 5, it can be reached with two different step sizes (3 and 5). If it is reached by a shorter step ( subunits), it means that the preceding binding site was already inside the interval , so i is not the first accessed site within it. For i ¼ 4and i ¼ 3, the site can only be reached with a 5-subunit step. P J,i denotes the probability that i is the first accessed subunit in the interval 3 # i # (dashed lines in the right column).
4 348 Vilfan the probability that a site before the junction (i # 7, as will be shown later) ever gets accessed by the motor: P M;i ¼ l for i# 7: () Initial state: first accessed site in the interval 7 # i # 3 We start our analysis at the point where a head of an approaching myosin V first passes the subunit 7 or binds to it. This way the initial state is definitely not influenced by the presence of the Arp2/3 complex. By counting only the first accessed site in this interval, we avoid double-counting of events where the motor binds, for example, first to site 7 and then to 4. We denote with P I,i the probability that once the first head has bound to any subunit in the interval 7 # i # 3, it has happened at subunit i. P I,i can be calculated in a way that is illustrated in Fig. 2. The sites between 7 and 7 can only be reached from outside the interval. Therefore, whenever the motor binds to one of them, it becomes the first accessed site in this interval. The probability P I,i then equals the probability that the site i ever gets accessed by the motor, which is =l; according to Eq.. For i ¼ 6and higher the situation becomes different. Site 6 counts as the first site in this interval if the preceding step size is 3 or 5 subunits, but not if it is. Therefore, the corresponding probability is P I; 6 ¼ ðp 3 P 5 Þ= l: The probability that the first site accessed in this interval is 5, P I, 5, has the same value. Finally, the sites 4 and 3 will be the first accessed sites in the interval if they are following a 5-subunit step. Therefore, their probabilities are P I; 4 ¼ P I; 3 ¼ P 5 =l: The probabilities P I,i are given in the second column of Table 2 and shown in the top graph of Fig. 3. As a next step, we will derive the probability distribution P J,i for the first accessed site in the interval 3 # i #. We choose this interval because (M, 3) is the first site from which the motor can reach the daughter filament. This distribution can be obtained from P I,i by redistributing probabilities for sites between 7 and 4 according to the conditional probability for the next step P J;j ¼ P I;j 4 + P M;jjM;i P I;i ; () i¼ 7 as shown in Fig. 3. The values of P J,j are given in the third and fourth column of Table 2. This distribution defines the state from which we will calculate the branching ratio at the Arp2/3 junction in the following section. Conditional branching ratio Now we can calculate the conditional probabilities that the lead head binds to the daughter filament, if the trail head is TABLE 2 Probability distribution P I,i for the first accessed binding site in the interval 7 # i # 3 i P I,i P J,i P J,i 7 =l 6 =l 5 =l 4 =l 3 =l =l =l. =l =l =l ðp 3 P 5 Þ=l ðp 3 P 5 ðp =P P 3 ÞÞ=l ðp 3 P 5 Þ=l =l P 5 =l ðp 5 ðp 3 =P P 3 ÞÞ=l P 5 =l =l ðp 5 P 3 Þ=l.754 P 5 =l.75 P J,i denotes the probability distribution for the first accessed site in the interval 3 # i #. P J,i is calculated according to Eq.. bound to a mother filament subunit i fulfilling 3 # i #. We denote this probability as P djm,i. Fig. 4 shows the most relevant dimer configurations with the trail head on mother filament sites between 5 and 6 and the lead head either on mother or on daughter filament. For each trail head position, the conditional probability that the lead head binds to a certain site is given by Eq. with the index j9 running over all accessible mother- as well as daughter-filament sites corresponding to one row in Fig. 4. Generally speaking, the daughter filament can either be reached directly, in a one-step process such as (M, 6) / (D,3), or in a two-step process such as (M, 9) / (M,4) / FIGURE 3 Probability distribution P I,i (upper diagram) for the first accessed binding site on the mother filament in the interval 7 # i # 3. The lower diagram shows the probability distribution P J,i for the first accessed site with 3 # i #. P J,i is calculated according to Eq., by redistributing the probabilities for 7 to 4 to other sites, as indicated by arrows. For example, if 7 is the first accessed site with 7 # i # 3, the first accessed site with 3 # i # can either be 6or 4. Note that the total probability that the motor binds to site 4 is higher than for any other site, which is due to the inaccessibility of site 2.
5 Myosin V Passing over Arp2/3 Junctions 349 FIGURE 4 Most probable lead head binding sites for different trail head positions. For selected trail head binding sites i, a collection of lead head binding sites and their bending energies U i,j (in units of k B T) is shown. Some trail head positions which do not have significant branching probabilities (e.g., 2) are omitted. The shape of both lever arms in each configuration is calculated numerically, by minimizing the bending energy. (D,8). Two examples are shown in Fig. 5. The probability that a motor with the trail head bound to site (M,i) eventually binds to the daughter filament is P djm;i ¼ + j P D;jjM;i + P D;jjM;k P M;kjM;i k : (2) Here the first term denotes one-step processes and the second term two-step processes, where the motor first binds to site k on the mother filament and subsequently to site j on daughter filament. Our numerical calculation shows that the only significant terms are those with k ¼ 4. The branching ratio P djm,i for each trail head position is shown in Fig. 6. The graph shows separately the contributions of one- and twostep processes. Total branching ratio With these probabilities and weights P J, i we finally obtain the branching ratio for the daughter filament: P d ¼ + P J;i P djm;i :27: (3) i¼ 3 If one head binds to the daughter filament, there is still some probability that the next head will bind back to the mother filament. One such example, with the trail head on the site (D,), is shown in the last row in Fig. 4. However, the contribution of such events to the total branching ratio is not significant. DISCUSSION While the exact result calculated above does require us to take into account all the individual configurations, its order of magnitude can also be understood with a simple handwaving argument, which goes as follows. Roughly speaking, the approaching myosin V motor can either reach the Arp2/3 complex on the opposite side of the actin filament, in which case it cannot switch to the daughter filament, or on the same side in which case the probability to switch to the
6 34 Vilfan FIGURE 5 An example of a one-step (upper path) and two-step (lower path) process through which the myosin V motor can reach the daughter filament. The probability that the trail head is bound to subunit (M, 9) in the initial state is P J, 9. In the next step, the lead head can bind to site (D,2) with conditional probability P D,2jM, 9. We denote such processes as one-step. Alternatively, the lead head can bind to the site (M,4) with conditional probability P M,4jM, 9 and, subsequently, the other head can bind to site (D,8) with conditional probability P D,8jM,4. This is an example of a two-step process. Note that this scheme shows just two examples of possible pathways and omits alternatives that are indicated by dashed arrows. daughter filament is ;/2. Together, this gives a branching ratio of /4, not far from the exact result of.27. An experiment measuring (among other quantities) the branching ratio at Arp2/3-mediated actin filament junctions was recently carried out by Ali et al. (39). The results are not directly comparable in the experiment, the actin filaments were attached to a glass surface so that some binding sites were not accessible for myosin V. However, because the blocked sites are different depending whether the side filament branches to the left or to the right, we expect that in statistical average, the calculated branching ratio still gives a good approximation. In the experiment 8% of the molecules dissociated, 2% continued on the daughter filament, and 62% on the mother filament. If we discount dissociation events, this means that the fraction that switched to the daughter filament was 24%. The statistical error of this figure is ;65% (the total number of events observed was 76). Therefore, the result can be considered in excellent agreement with the model calculation. FIGURE 6 Probability that the motor will step on and continue its walk along the daughter filament if the trail head is initially bound to site i on the mother filament, P djm,i (solid and hatched bars), as calculated from Eq. 2. The solid bars show the contribution of one-step processes, in which the lead head binds to a site on the daughter filament immediately. The hatched bars show the contribution of two-step processes in which the motor first binds to another site on the mother filament (usually site 4) and then in the second step to the daughter filament. The shaded bars show the probability that the motor continues along the mother filament. Note that our calculation only concerns stepping patterns and does not take into account kinetics. This has an important advantage that the result only depends on geometric and elastic parameters, but not on kinetic constants, some of which are still less well known (27). Combining this model with the full kinetic scheme could influence the predicted branching ratio as follows. First, the binding rate of the lead head can be different in the vicinity of the Arp2/3 complex. And second, the ADP release rate of the rear head can be influenced by intramolecular strain, possibly also by its lateral (off-axis) component, as observed by Purcell and coworkers (4). Both these effects do not have a direct influence on the branching ratio, but they might have an indirect one by influencing the dissociation probability. In this calculation, events where the whole myosin V molecule dissociates from the actin filament are not taken into account. There is, however, evidence that the predominant dissociation path leads through detachment of a head in the ADP state (23,4) these processes are denoted as Pathway 2 in Vilfan (27). It is therefore plausible that the termination rate increases if either the lead head is hindered in its search for a binding site, or the ADP release in the trail head is slowed down. In our model (see elastic energies in Fig. 4), the lead head binding rate is strongly reduced if the trail head is bound to site 3; it is also reduced somewhat if the trail head is bound to, while it is accelerated for trail head positions 9 and 4. In total, dissociation can be accelerated by the presence of the Arp2/3 junction in 2 out of l cases. Without going into quantitative details, one can conclude that the dissociation probability could theoretically increase by up to 2=l :5: The branching ratio for the daughter filament could then be somewhat smaller, because those trail head positions that have the higher branching ratio are also more likely to lead to dissociation. The effect caused by the strain-dependent ADP release rate is more difficult to estimate, mainly because the exact influence of off-axis strain on the ADP release is not yet known quantitatively. To check the robustness of our result against uncertainties in model parameters, we calculated the variation of the branching ratio with several model parameters. Note that
7 Myosin V Passing over Arp2/3 Junctions 34 these calculations were carried out numerically and took into account all relevant processes, including longer and shorter steps, as well as transitions not shown in Fig. 4. Data in Fig. 7 shows a variation of ;6% if either the lead head or the trail head angle is modified by 65. The allowed variation of these two parameters that keeps the model consistent with the experimental result is therefore restricted, although parameter sets where both angles are increased or decreased simultaneously cannot be excluded. The result is more robust against variations in the lever arm stiffness EI, where deviations do not exceed a few percent if the value of EI is changed by a factor of 3 in either direction. We can therefore conclude that the calculated branching ratio adds support to the elastic lever arm model presented in Vilfan (27) and the geometric parameters used there. However, we cannot use it as a criterion to determine the lever arm stiffness, which is still not known precisely. Another open question is to what extent the result can be reproduced with alternative models, such as Lan and Sun (42), which uses a more complex model for the elasticity of the lever arm, with a soft longitudinal (;/3 of the value used there), but very stiff azimuthal component. The completely different class of hotspot models, which proposes that the position of the next binding site is determined by a propagating conformational change in the actin filament (43), on the other hand, seems less compatible with the finding, unless the conformational change could propagate through the Arp2/3 complex as well. I thank Mojca Vilfan for comments on the manuscript and Stan Burgess for helpful discussions. This work was supported by the Slovenian Research Agency (grant No. P-99). REFERENCES FIGURE 7 Dependence of the calculated branching ratio P d on different model parameters. (a) Dependence on the lead head angle f pre PS (solid line, lower scale) and trail head angle f post PS (dashed line, upper scale). (b) Dependence on the lever arm stiffness (bending modulus) EI. In both diagrams, the dot shows the value used in all other calculations throughout the article.. Vale, R. D. 23. Myosin V motor proteins: marching stepwise towards a mechanism. J. Cell Biol. 63: Sellers, J. R., and C. Veigel. 26. Walking with myosin V. Curr. Opin. Cell Biol. 8: Rief, M., R. S. Rock, A. D. Mehta, M. S. Mooseker, R. E. Cheney, and J. A. Spudich. 2. Myosin-V stepping kinetics: a molecular model for processivity. Proc. Natl. Acad. Sci. USA. 97: Mehta, A. D., R. S. Rock, M. Rief, J. A. Spudich, M. S. Mooseker, and R. E. Cheney Myosin-V is a processive actin-based motor. Nature. 4: Rock, R. S., M. Rief, A. D. Mehta, and J. A. Spudich. 2. In vitro assays of processive myosin motors. Methods. 22: Veigel, C., F. Wang, M. L. Bartoo, J. R. Sellers, and J. E. Molloy. 22. The gated gait of the processive molecular motor, myosin V. Nat. Cell Biol. 4: Purcell, T. J., C. Morris, J. A. Spudich, and H. L. Sweeney. 22. Role of the lever arm in the processive stepping of myosin V. Proc. Natl. Acad. Sci. USA. 99: Gebhardt, J. C., A. E. Clemen, J. Jaud, and M. Rief. 26. Myosin-V is a mechanical ratchet. Proc. Natl. Acad. Sci. USA. 3: Clemen, A. E., M. Vilfan, J. Jaud, J. Zhang, M. Barmann, and M. Rief. 25. Force-dependent stepping kinetics of myosin-v. Biophys. J. 88: Uemura, S., H. Higuchi, A. O. Olivares, E. M. De La Cruz, and S. Ishiwata. 24. Mechanochemical coupling of two substeps in a single myosin V motor. Nat. Struct. Mol. Biol. : De La Cruz, E. M., A. L. Wells, S. S. Rosenfeld, E. M. Ostap, and H. L. Sweeney The kinetic mechanism of myosin V. Proc. Natl. Acad. Sci. USA. 96: De La Cruz, E. M., A. L. Wells, H. L. Sweeney, and E. M. Ostap. 2. Actin and light chain isoform dependence of myosin V kinetics. Biochemistry. 39: De La Cruz, E. M., H. L. Sweeney, and E. M. Ostap. 2. ADP inhibition of myosin V ATPase activity. Biophys. J. 79: Yengo, C. M., E. M. De la Cruz, D. Safer, E. M. Ostap, and H. L. Sweeney. 22. Kinetic characterization of the weak binding states of myosin V. Biochemistry. 4: Yildiz, A., J. N. Forkey, S. A. McKinney, T. Ha, Y. E. Goldman, and P. R. Selvin. 23. Myosin V walks hand-over-hand: single fluorophore imaging with.5-nm localization. Science. 3:
8 342 Vilfan 6. Ali, M. Y., S. Uemura, K. Adachi, H. Itoh, K. Kinosita, Jr., and S. Ishiwata. 22. Myosin V is a left-handed spiral motor on the righthanded actin helix. Nat. Struct. Biol. 9: Forkey, J. N., M. E. Quinlan, M. A. Shaw, J. E. Corrie, and Y. E. Goldman. 23. Three-dimensional structural dynamics of myosin V by single-molecule fluorescence polarization. Nature. 422: Walker, M. L., S. A. Burgess, J. R. Sellers, F. Wang, J. A. Hammer, J. Trinick, and P. J. Knight. 2. Two-headed binding of a processive myosin to F-actin. Nature. 45: Burgess, S., M. Walker, F. Wang, J. R. Sellers, H. D. White, P. J. Knight, and J. Trinick. 22. The prepower stroke conformation of myosin V. J. Cell Biol. 59: Wang, F., K. Thirumurugan, W. F. Stafford, J. A. Hammer, P. J. Knight, and J. R. Sellers. 23. Regulated conformation of myosin V. J. Biol. Chem. 279: Coureux, P. D., A. L. Wells, J. Menetrey, C. M. Yengo, C. A. Morris, H. L. Sweeney, and A. Houdusse. 23. A structural state of the myosin V motor without bound nucleotide. Nature. 425: Warshaw, D. M., G. G. Kennedy, S. S. Work, E. B. Krementsova, S. Beck, and K. M. Trybus. 25. Differential labeling of myosin V heads with quantum dots allows direct visualization of hand-over-hand processivity. Biophys. J. 88:L3 L Baker, J. E., E. B. Krementsova, G. G. Kennedy, A. Armstrong, K. M. Trybus, and D. M. Warshaw. 24. Myosin V processivity: multiple kinetic pathways for head-to-head coordination. Proc. Natl. Acad. Sci. USA. : Sakamoto, T., F. Wang, S. Schmitz, Y. Xu, Q. Xu, J. E. Molloy, C. Veigel, and J. R. Sellers. 23. Neck length and processivity of myosin V. J. Biol. Chem. 278: Pantaloni, D., R. Boujemaa, D. Didry, P. Gounon, and M. F. Carlier. 2. The Arp2/3 complex branches filament barbed ends: functional antagonism with capping proteins. Nat. Cell Biol. 2: Machesky, L. M., R. D. Mullins, H. N. Higgs, D. A. Kaiser, L. Blanchoin, R. C. May, M. E. Hall, and T. D. Pollard Scar, a WASp-related protein, activates nucleation of actin filaments by the Arp2/3 complex. Proc. Natl. Acad. Sci. USA. 96: Vilfan, A. 25. Elastic lever-arm model for myosin V. Biophys. J. 88: Kolomeisky, A. B., and M. E. Fisher. 23. A simple kinetic model describes the processivity of myosin-v. Biophys. J. 84: Skau, K. I., R. B. Hoyle, and M. S. Turner. 26. A kinetic model describing the processivity of myosin-v. Biophys. J. 9: Vilfan, A., and T. Duke. 23. Instabilities in the transient response of muscle. Biophys. J. 85: Dunn, A. R., and J. A. Spudich. 27. Dynamics of the unbound head during myosin V processive translocation. Nat. Struct. Mol. Biol. 4: Knight, P., and J. Trinick Structure of the myosin projections on native thick filaments from vertebrate skeletal muscle. J. Mol. Biol. 77: Offer, G., and P. Knight The structure of the head-tail junction of the myosin molecule. J. Mol. Biol. 256: Knight, P. J Dynamic behavior of the head-tail junction of myosin. J. Mol. Biol. 255: Egile, C., I. Rouiller, X. P. Xu, N. Volkmann, R. Li, and D. Hanein. 25. Mechanism of filament nucleation and branch stability revealed by the structure of the Arp2/3 complex at actin branch junctions. PLoS Biol. 3:e Volkmann, N., K. J. Amann, S. Stoilova-McPhie, C. Egile, D. C. Winter, L. Hazelwood, J. E. Heuser, R. Li, T. D. Pollard, and D. Hanein. 2. Structure of Arp2/3 complex in its activated state and in actin filament branch junctions. Science. 293: Vilfan, A. 25. Influence of fluctuations in actin structure on myosin V step size. J. Chem. Inf. Model. 45: Veigel, C., S. Schmitz, F. Wang, and J. R. Sellers. 25. Loaddependent kinetics of myosin-v can explain its high processivity. Nat. Cell Biol. 7: Ali, M. Y., E. B. Krementsova, G. G. Kennedy, R. Mahaffy, T. D. Pollard, K. M. Trybus, and D. M. Warshaw. 27. Myosin Va maneuvers through actin intersections and diffuses along microtubules. Proc. Natl. Acad. Sci. USA. 4: Purcell, T. J., H. L. Sweeney, and J. A. Spudich. 25. A forcedependent state controls the coordination of processive myosin V. Proc. Natl. Acad. Sci. USA. 2: Wu, Y., Y. Q. Gao, and M. Karplus. 27. A kinetic model of coordinated myosin V. Biochemistry. 46: Lan, G., and S. X. Sun. 25. Dynamics of myosin-v processivity. Biophys. J. 88: Watanabe, T. M., H. Tanaka, A. H. Iwane, S. Maki-Yonekura, K. Homma, A. Inoue, R. Ikebe, T. Yanagida, and M. Ikebe. 24. A oneheaded class V myosin molecule develops multiple large (approximately 32-nm) steps successively. Proc. Natl. Acad. Sci. USA. :
Five models for myosin V
 Five models for myosin V Andrej Vilfan J. Stefan Institute, Jamova 39, 1000 Ljubljana, Slovenia Myosin V was the first discovered processive motor from the myosin family. It has therefore been subject
Five models for myosin V Andrej Vilfan J. Stefan Institute, Jamova 39, 1000 Ljubljana, Slovenia Myosin V was the first discovered processive motor from the myosin family. It has therefore been subject
Elastic Lever-Arm Model for Myosin V
 3792 Biophysical Journal Volume 88 June 2005 3792 3805 Elastic Lever-Arm Model for Myosin V Andrej Vilfan J. Stefan Institute, Ljubljana, Slovenia ABSTRACT We present a mechanochemical model for myosin
3792 Biophysical Journal Volume 88 June 2005 3792 3805 Elastic Lever-Arm Model for Myosin V Andrej Vilfan J. Stefan Institute, Ljubljana, Slovenia ABSTRACT We present a mechanochemical model for myosin
A new model for myosin dimeric motors incorporating Brownian ratchet and powerstroke mechanisms
 Invited Paper A new model for myosin dimeric motors incorporating Brownian ratchet and powerstroke mechanisms Brian Geislinger and Ryoichi Kawai Department of Physics, University of Alabama at Birmingham,
Invited Paper A new model for myosin dimeric motors incorporating Brownian ratchet and powerstroke mechanisms Brian Geislinger and Ryoichi Kawai Department of Physics, University of Alabama at Birmingham,
arxiv: v1 [physics.bio-ph] 9 Aug 2011
![arxiv: v1 [physics.bio-ph] 9 Aug 2011 arxiv: v1 [physics.bio-ph] 9 Aug 2011](/thumbs/95/124074436.jpg) Phenomenological analysis of ATP dependence of motor protein Yunxin Zhang Laboratory of Mathematics for Nonlinear Science, Centre for Computational System Biology, School of Mathematical Sciences, Fudan
Phenomenological analysis of ATP dependence of motor protein Yunxin Zhang Laboratory of Mathematics for Nonlinear Science, Centre for Computational System Biology, School of Mathematical Sciences, Fudan
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09450 Supplementary Table 1 Summary of kinetic parameters. Kinetic parameters were V = V / 1 K / ATP and obtained using the relationships max ( + m [ ]) V d s /( 1/ k [ ATP] + 1 k ) =,
doi:10.1038/nature09450 Supplementary Table 1 Summary of kinetic parameters. Kinetic parameters were V = V / 1 K / ATP and obtained using the relationships max ( + m [ ]) V d s /( 1/ k [ ATP] + 1 k ) =,
Dynamics of Myosin-V Processivity
 Biophysical Journal Volume 88 February 2005 999 1008 999 Dynamics of Myosin-V Processivity Ganhui Lan* and Sean X. Sun* y *Department of Mechanical Engineering, and y Whitaker Institute of Biomedical Engineering,
Biophysical Journal Volume 88 February 2005 999 1008 999 Dynamics of Myosin-V Processivity Ganhui Lan* and Sean X. Sun* y *Department of Mechanical Engineering, and y Whitaker Institute of Biomedical Engineering,
Coarse-Grained Structural Modeling of Molecular Motors Using Multibody Dynamics
 Cellular and Molecular Bioengineering, Vol. 2, No. 3, September 2009 (Ó 2009) pp. 366 374 DOI: 10.1007/s12195-009-0084-4 Coarse-Grained Structural Modeling of Molecular Motors Using Multibody Dynamics
Cellular and Molecular Bioengineering, Vol. 2, No. 3, September 2009 (Ó 2009) pp. 366 374 DOI: 10.1007/s12195-009-0084-4 Coarse-Grained Structural Modeling of Molecular Motors Using Multibody Dynamics
Supplementary Information
 Supplementary Information Switching of myosin-v motion between the lever-arm swing and Brownian search-and-catch Keisuke Fujita 1*, Mitsuhiro Iwaki 2,3*, Atsuko H. Iwane 1, Lorenzo Marcucci 1 & Toshio
Supplementary Information Switching of myosin-v motion between the lever-arm swing and Brownian search-and-catch Keisuke Fujita 1*, Mitsuhiro Iwaki 2,3*, Atsuko H. Iwane 1, Lorenzo Marcucci 1 & Toshio
Introduction: actin and myosin
 Introduction: actin and myosin Actin Myosin Myosin V and actin 375 residues Found in all eukaryotes Polymeric Forms track for myosin Many other cellular functions 36 nm pseudo-helical repeat Catalytic
Introduction: actin and myosin Actin Myosin Myosin V and actin 375 residues Found in all eukaryotes Polymeric Forms track for myosin Many other cellular functions 36 nm pseudo-helical repeat Catalytic
Interhead distance measurements in myosin VI via SHRImP support a simplified
 Biophys J BioFAST, published on April 29, 2005 as doi:10.1529/biophysj.105.060608 Interhead distance measurements in myosin VI via SHRImP support a simplified hand-over-hand model Hamza Balci 1, Taekjip
Biophys J BioFAST, published on April 29, 2005 as doi:10.1529/biophysj.105.060608 Interhead distance measurements in myosin VI via SHRImP support a simplified hand-over-hand model Hamza Balci 1, Taekjip
NIH Public Access Author Manuscript Cell Mol Bioeng. Author manuscript; available in PMC 2010 April 27.
 NIH Public Access Author Manuscript Published in final edited form as: Cell Mol Bioeng. 2009 September 1; 2(3): 366 374. doi:10.1007/s12195-009-0084-4. Coarse-Grained Structural Modeling of Molecular Motors
NIH Public Access Author Manuscript Published in final edited form as: Cell Mol Bioeng. 2009 September 1; 2(3): 366 374. doi:10.1007/s12195-009-0084-4. Coarse-Grained Structural Modeling of Molecular Motors
Mechanochemical coupling of two substeps in a single myosin V motor
 Mechanochemical coupling of two substeps in a single myosin V motor Sotaro Uemura 1, Hideo Higuchi 2,3, Adrian O Olivares 4, Enrique M De La Cruz 4 & Shin ichi Ishiwata 1,5 Myosin V is a double-headed
Mechanochemical coupling of two substeps in a single myosin V motor Sotaro Uemura 1, Hideo Higuchi 2,3, Adrian O Olivares 4, Enrique M De La Cruz 4 & Shin ichi Ishiwata 1,5 Myosin V is a double-headed
SINGLE-MOLECULE PHYSIOLOGY
 SINGLE-MOLECULE PHYSIOLOGY Kazuhiko Kinosita, Jr. Center for Integrative Bioscience, Okazaki National Research Institutes Higashiyama 5-1, Myodaiji, Okazaki 444-8585, Japan Single-Molecule Physiology under
SINGLE-MOLECULE PHYSIOLOGY Kazuhiko Kinosita, Jr. Center for Integrative Bioscience, Okazaki National Research Institutes Higashiyama 5-1, Myodaiji, Okazaki 444-8585, Japan Single-Molecule Physiology under
A Simple Kinetic Model Describes the Processivity of Myosin-V
 1642 Biophysical Journal Volume 84 March 2003 1642 1650 A Simple Kinetic Model Describes the Processivity of Myosin-V Anatoly B. Kolomeisky* and Michael E. Fisher y *Department of Chemistry, Rice University,
1642 Biophysical Journal Volume 84 March 2003 1642 1650 A Simple Kinetic Model Describes the Processivity of Myosin-V Anatoly B. Kolomeisky* and Michael E. Fisher y *Department of Chemistry, Rice University,
Anatoly B. Kolomeisky. Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS?
 Anatoly B. Kolomeisky Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS? Motor Proteins Enzymes that convert the chemical energy into mechanical
Anatoly B. Kolomeisky Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS? Motor Proteins Enzymes that convert the chemical energy into mechanical
Detailed Tuning of Structure and Intramolecular Communication Are Dispensable for Processive Motion of Myosin VI
 430 Biophysical Journal Volume 100 January 2011 430 439 Detailed Tuning of Structure and Intramolecular Communication Are Dispensable for Processive Motion of Myosin VI Mary Williard Elting, Zev Bryant,
430 Biophysical Journal Volume 100 January 2011 430 439 Detailed Tuning of Structure and Intramolecular Communication Are Dispensable for Processive Motion of Myosin VI Mary Williard Elting, Zev Bryant,
Acto-myosin: from muscles to single molecules. Justin Molloy MRC National Institute for Medical Research LONDON
 Acto-myosin: from muscles to single molecules. Justin Molloy MRC National Institute for Medical Research LONDON Energy in Biological systems: 1 Photon = 400 pn.nm 1 ATP = 100 pn.nm 1 Ion moving across
Acto-myosin: from muscles to single molecules. Justin Molloy MRC National Institute for Medical Research LONDON Energy in Biological systems: 1 Photon = 400 pn.nm 1 ATP = 100 pn.nm 1 Ion moving across
Motor Proteins as Nanomachines: The Roles of Thermal Fluctuations in Generating Force and Motion
 Séminaire Poincaré XII (2009) 33 44 Séminaire Poincaré Motor Proteins as Nanomachines: The Roles of Thermal Fluctuations in Generating Force and Motion Jonathon Howard Max Planck Institute of Molecular
Séminaire Poincaré XII (2009) 33 44 Séminaire Poincaré Motor Proteins as Nanomachines: The Roles of Thermal Fluctuations in Generating Force and Motion Jonathon Howard Max Planck Institute of Molecular
Effects of Nonequilibrium Processes on Actin Polymerization and Force Generation
 Effects of Nonequilibrium Processes on Actin Polymerization and Force Generation Anders Carlsson Washington University in St Louis Biological cells constantly utilize the influx of energy mediated by ATP
Effects of Nonequilibrium Processes on Actin Polymerization and Force Generation Anders Carlsson Washington University in St Louis Biological cells constantly utilize the influx of energy mediated by ATP
Are motor proteins power strokers, Brownian motors or both?
 Invited Paper Are motor proteins power strokers, Brownian motors or both? Brian Geislinger a,erindarnell b, Kimberly Farris c,andryoichikawai a a Dept. of Physics, Univ. of Alabama at Birmingham, 153 3rd
Invited Paper Are motor proteins power strokers, Brownian motors or both? Brian Geislinger a,erindarnell b, Kimberly Farris c,andryoichikawai a a Dept. of Physics, Univ. of Alabama at Birmingham, 153 3rd
The biological motors
 Motor proteins The definition of motor proteins Miklós Nyitrai, November 30, 2016 Molecular machines key to understand biological processes machines in the micro/nano-world (unidirectional steps): nm,
Motor proteins The definition of motor proteins Miklós Nyitrai, November 30, 2016 Molecular machines key to understand biological processes machines in the micro/nano-world (unidirectional steps): nm,
arxiv:cond-mat/ v2 [cond-mat.stat-mech] 8 Jun 2006
![arxiv:cond-mat/ v2 [cond-mat.stat-mech] 8 Jun 2006 arxiv:cond-mat/ v2 [cond-mat.stat-mech] 8 Jun 2006](/thumbs/93/113637182.jpg) Brownian molecular motors driven by rotation-translation coupling Brian Geislinger and Ryoichi Kawai Department of Physics, University of Alabama at Birmingham, Birmingham, AL 35294 arxiv:cond-mat/6487v2
Brownian molecular motors driven by rotation-translation coupling Brian Geislinger and Ryoichi Kawai Department of Physics, University of Alabama at Birmingham, Birmingham, AL 35294 arxiv:cond-mat/6487v2
Transport of single molecules along the periodic parallel lattices with coupling
 THE JOURNAL OF CHEMICAL PHYSICS 124 204901 2006 Transport of single molecules along the periodic parallel lattices with coupling Evgeny B. Stukalin The James Franck Institute The University of Chicago
THE JOURNAL OF CHEMICAL PHYSICS 124 204901 2006 Transport of single molecules along the periodic parallel lattices with coupling Evgeny B. Stukalin The James Franck Institute The University of Chicago
Intracellular transport
 Transport in cells Intracellular transport introduction: transport in cells, molecular players etc. cooperation of motors, forces good and bad transport regulation, traffic issues, Stefan Klumpp image
Transport in cells Intracellular transport introduction: transport in cells, molecular players etc. cooperation of motors, forces good and bad transport regulation, traffic issues, Stefan Klumpp image
Muscle regulation and Actin Topics: Tropomyosin and Troponin, Actin Assembly, Actin-dependent Movement
 1 Muscle regulation and Actin Topics: Tropomyosin and Troponin, Actin Assembly, Actin-dependent Movement In the last lecture, we saw that a repeating alternation between chemical (ATP hydrolysis) and vectorial
1 Muscle regulation and Actin Topics: Tropomyosin and Troponin, Actin Assembly, Actin-dependent Movement In the last lecture, we saw that a repeating alternation between chemical (ATP hydrolysis) and vectorial
Department of Physics, University at Buffalo, Buffalo, NY INTRODUCTION
 proteins STRUCTURE O FUNCTION O BIOINFORMATICS Coarse-grained modeling of conformational transitions underlying the processive stepping of myosin V dimer along filamentous actin Wenjun Zheng* Department
proteins STRUCTURE O FUNCTION O BIOINFORMATICS Coarse-grained modeling of conformational transitions underlying the processive stepping of myosin V dimer along filamentous actin Wenjun Zheng* Department
Dynamics of an inchworm nano walker
 Dynamics of an inchworm nano walker A. Ciudad a, J.M. Sancho a A.M. Lacasta b a Departament d Estructura i Constituents de la Matèria, Facultat de Física, Universitat de Barcelona, Diagonal 647, E-08028
Dynamics of an inchworm nano walker A. Ciudad a, J.M. Sancho a A.M. Lacasta b a Departament d Estructura i Constituents de la Matèria, Facultat de Física, Universitat de Barcelona, Diagonal 647, E-08028
According to the diagram, which of the following is NOT true?
 Instructions: Review Chapter 44 on muscular-skeletal systems and locomotion, and then complete the following Blackboard activity. This activity will introduce topics that will be covered in the next few
Instructions: Review Chapter 44 on muscular-skeletal systems and locomotion, and then complete the following Blackboard activity. This activity will introduce topics that will be covered in the next few
Enhancement of cargo processivity by cooperating molecular motors
 Enhancement of cargo processivity by cooperating molecular motors Filippo Posta, a Maria R. D Orsogna b and Tom Chou c a Dept. Biomathematics, UCLA, Los Angeles CA. fposta@ucla.edu b Dept. Mathematics,
Enhancement of cargo processivity by cooperating molecular motors Filippo Posta, a Maria R. D Orsogna b and Tom Chou c a Dept. Biomathematics, UCLA, Los Angeles CA. fposta@ucla.edu b Dept. Mathematics,
Actin and myosin constitute thin and thick filaments, respectively,
 Conformational change of the actomyosin complex drives the multiple stepping movement Tomoki P. Terada, Masaki Sasai, and Tetsuya Yomo Graduate School of Human Informatics, Nagoya University, Nagoya 464-860,
Conformational change of the actomyosin complex drives the multiple stepping movement Tomoki P. Terada, Masaki Sasai, and Tetsuya Yomo Graduate School of Human Informatics, Nagoya University, Nagoya 464-860,
BE/APh161 Physical Biology of the Cell. Rob Phillips Applied Physics and Bioengineering California Institute of Technology
 BE/APh161 Physical Biology of the Cell Rob Phillips Applied Physics and Bioengineering California Institute of Technology Cells Decide: Where to Go The Hunters of the Immune Response (Berman et al.) There
BE/APh161 Physical Biology of the Cell Rob Phillips Applied Physics and Bioengineering California Institute of Technology Cells Decide: Where to Go The Hunters of the Immune Response (Berman et al.) There
662 Biophysical Journal Volume 107 August The Winch Model Can Explain both Coordinated and Uncoordinated Stepping of Cytoplasmic Dynein
 662 Biophysical Journal Volume 107 August 2014 662 671 Article The Winch Model Can Explain both Coordinated and Uncoordinated Stepping of Cytoplasmic Dynein Andreja Sarlah 1 and Andrej Vilfan 1,2, * 1
662 Biophysical Journal Volume 107 August 2014 662 671 Article The Winch Model Can Explain both Coordinated and Uncoordinated Stepping of Cytoplasmic Dynein Andreja Sarlah 1 and Andrej Vilfan 1,2, * 1
arxiv: v1 [physics.bio-ph] 27 Sep 2014
![arxiv: v1 [physics.bio-ph] 27 Sep 2014 arxiv: v1 [physics.bio-ph] 27 Sep 2014](/thumbs/95/122806024.jpg) Ensemble velocity of non-processive molecular motors with multiple chemical states Andrej Vilfan J. Stefan Institute, Jamova 39, Ljubljana, Slovenia and Faculty of Mathematics and Physics, University of
Ensemble velocity of non-processive molecular motors with multiple chemical states Andrej Vilfan J. Stefan Institute, Jamova 39, Ljubljana, Slovenia and Faculty of Mathematics and Physics, University of
Efficiency of molecular motors at maximum power
 epl draft arxiv:81.3743v2 [cond-mat.stat-mech] 23 Jul 8 II. Institut für Theoretische Physik, Universität Stuttgart, 755 Stuttgart, Germany PACS 5.7.LN Nonequilibrium and irreversible thermodynamics PACS
epl draft arxiv:81.3743v2 [cond-mat.stat-mech] 23 Jul 8 II. Institut für Theoretische Physik, Universität Stuttgart, 755 Stuttgart, Germany PACS 5.7.LN Nonequilibrium and irreversible thermodynamics PACS
Conventional kinesin is a processive motor protein that walks
 Kinesin crouches to sprint but resists pushing Michael E. Fisher* and Young C. Kim Institute for Physical Science and Technology, University of Maryland, College Park, MD 20742 Contributed by Michael E.
Kinesin crouches to sprint but resists pushing Michael E. Fisher* and Young C. Kim Institute for Physical Science and Technology, University of Maryland, College Park, MD 20742 Contributed by Michael E.
EXPERIMENTAL PROCEDURES
 THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 284, NO. 4, pp. 2138 2149, January 23, 2009 Printed in the U.S.A. Switch 1 Mutation S217A Converts Myosin V into a Low Duty Ratio Motor * Received for publication,
THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 284, NO. 4, pp. 2138 2149, January 23, 2009 Printed in the U.S.A. Switch 1 Mutation S217A Converts Myosin V into a Low Duty Ratio Motor * Received for publication,
Biophysik der Moleküle!
 Biophysik der Moleküle!!"#$%&'()*+,-$./0()'$12$34!4! Molecular Motors:! - linear motors" 6. Dec. 2010! Muscle Motors and Cargo Transporting Motors! There are striking structural similarities but functional
Biophysik der Moleküle!!"#$%&'()*+,-$./0()'$12$34!4! Molecular Motors:! - linear motors" 6. Dec. 2010! Muscle Motors and Cargo Transporting Motors! There are striking structural similarities but functional
c 2006 by Prasanth Sankar. All rights reserved.
 c 2006 by Prasanth Sankar. All rights reserved. PHENOMENOLOGICAL MODELS OF MOTOR PROTEINS BY PRASANTH SANKAR M. S., University of Illinois at Urbana-Champaign, 2000 DISSERTATION Submitted in partial fulfillment
c 2006 by Prasanth Sankar. All rights reserved. PHENOMENOLOGICAL MODELS OF MOTOR PROTEINS BY PRASANTH SANKAR M. S., University of Illinois at Urbana-Champaign, 2000 DISSERTATION Submitted in partial fulfillment
Chapter 16. Cellular Movement: Motility and Contractility. Lectures by Kathleen Fitzpatrick Simon Fraser University Pearson Education, Inc.
 Chapter 16 Cellular Movement: Motility and Contractility Lectures by Kathleen Fitzpatrick Simon Fraser University Two eukaryotic motility systems 1. Interactions between motor proteins and microtubules
Chapter 16 Cellular Movement: Motility and Contractility Lectures by Kathleen Fitzpatrick Simon Fraser University Two eukaryotic motility systems 1. Interactions between motor proteins and microtubules
Neurite formation & neuronal polarization
 Neurite formation & neuronal polarization Paul Letourneau letou001@umn.edu Chapter 16; The Cytoskeleton; Molecular Biology of the Cell, Alberts et al. 1 An immature neuron in cell culture first sprouts
Neurite formation & neuronal polarization Paul Letourneau letou001@umn.edu Chapter 16; The Cytoskeleton; Molecular Biology of the Cell, Alberts et al. 1 An immature neuron in cell culture first sprouts
BMB Class 17, November 30, Single Molecule Biophysics (II)
 BMB 178 2018 Class 17, November 30, 2018 15. Single Molecule Biophysics (II) New Advances in Single Molecule Techniques Atomic Force Microscopy Single Molecule Manipulation - optical traps and tweezers
BMB 178 2018 Class 17, November 30, 2018 15. Single Molecule Biophysics (II) New Advances in Single Molecule Techniques Atomic Force Microscopy Single Molecule Manipulation - optical traps and tweezers
Neurite formation & neuronal polarization. The cytoskeletal components of neurons have characteristic distributions and associations
 Mechanisms of neuronal migration & Neurite formation & neuronal polarization Paul Letourneau letou001@umn.edu Chapter 16; The Cytoskeleton; Molecular Biology of the Cell, Alberts et al. 1 The cytoskeletal
Mechanisms of neuronal migration & Neurite formation & neuronal polarization Paul Letourneau letou001@umn.edu Chapter 16; The Cytoskeleton; Molecular Biology of the Cell, Alberts et al. 1 The cytoskeletal
SUPPLEMENTARY FIGURE 1. Force dependence of the unbinding rate: (a) Force-dependence
 (b) BSA-coated beads double exponential low force exponential high force exponential 1 unbinding time tb [sec] (a) unbinding time tb [sec] SUPPLEMENTARY FIGURES BSA-coated beads without BSA.2.1 5 1 load
(b) BSA-coated beads double exponential low force exponential high force exponential 1 unbinding time tb [sec] (a) unbinding time tb [sec] SUPPLEMENTARY FIGURES BSA-coated beads without BSA.2.1 5 1 load
MYOSIN IX: A SINGLE-HEADED PROCESSIVE MOTOR
 MYOSIN IX: A SINGLE-HEADED PROCESSIVE MOTOR BY TAKETOSHI KAMBARA A Dissertation Submitted to the Faculty of the WORCESTER POLYTECHNIC INSTITUTE in partial fulfillment of the requirements for the Degree
MYOSIN IX: A SINGLE-HEADED PROCESSIVE MOTOR BY TAKETOSHI KAMBARA A Dissertation Submitted to the Faculty of the WORCESTER POLYTECHNIC INSTITUTE in partial fulfillment of the requirements for the Degree
Polymerization/depolymerization motors
 Polymerization/depolymerization motors Movement formation Kuo Lab, J.H.U. http://www.nature.com/nature/journal/v407/n6807/extref/40 71026a0_S3.mov http://www.bme.jhu.edu/~skuo/movies/macrophchase.mov http://www.bme.jhu.edu/~skuo/movies/gc_filo.mov
Polymerization/depolymerization motors Movement formation Kuo Lab, J.H.U. http://www.nature.com/nature/journal/v407/n6807/extref/40 71026a0_S3.mov http://www.bme.jhu.edu/~skuo/movies/macrophchase.mov http://www.bme.jhu.edu/~skuo/movies/gc_filo.mov
Thermodynamics and Kinetics of Molecular Motors
 Biophysical Journal Volume 98 June 2010 2401 2409 2401 Thermodynamics and Kinetics of Molecular Motors R. Dean Astumian* Department of Physics and Astronomy, University of Maine, Orono, Maine; and Department
Biophysical Journal Volume 98 June 2010 2401 2409 2401 Thermodynamics and Kinetics of Molecular Motors R. Dean Astumian* Department of Physics and Astronomy, University of Maine, Orono, Maine; and Department
3 Biopolymers Uncorrelated chains - Freely jointed chain model
 3.4 Entropy When we talk about biopolymers, it is important to realize that the free energy of a biopolymer in thermal equilibrium is not constant. Unlike solids, biopolymers are characterized through
3.4 Entropy When we talk about biopolymers, it is important to realize that the free energy of a biopolymer in thermal equilibrium is not constant. Unlike solids, biopolymers are characterized through
Fitting Force-dependent Kinetic Models to Single-Molecule Dwell Time Records
 Fitting Force-dependent Kinetic Models to Single-Molecule Dwell Time Records Abstract Micromanipulation techniques such as optical tweezers atomic force microscopy allow us to identify characterize force-dependent
Fitting Force-dependent Kinetic Models to Single-Molecule Dwell Time Records Abstract Micromanipulation techniques such as optical tweezers atomic force microscopy allow us to identify characterize force-dependent
Research Article Flexural Stiffness of Myosin Va Subdomains as Measured from Tethered Particle Motion
 Journal of Biophysics Volume 2015, Article ID 465693, 9 pages http://dx.doi.org/10.1155/2015/465693 Research Article Flexural Stiffness of Myosin Va Subdomains as Measured from Tethered Particle Motion
Journal of Biophysics Volume 2015, Article ID 465693, 9 pages http://dx.doi.org/10.1155/2015/465693 Research Article Flexural Stiffness of Myosin Va Subdomains as Measured from Tethered Particle Motion
Papers and Reference Numbers 1. Bustamante, C., Chemla, Y. R., Forde, N. R., & Izhaky, D. (2004). Mechanical processes in biochemistry.
 Papers and Reference Numbers 1. Bustamante, C., Chemla, Y. R., Forde, N. R., & Izhaky, D. (2004). Mechanical processes in biochemistry. Annual review of biochemistry, 73, 705-48. 1.1. not his data. This
Papers and Reference Numbers 1. Bustamante, C., Chemla, Y. R., Forde, N. R., & Izhaky, D. (2004). Mechanical processes in biochemistry. Annual review of biochemistry, 73, 705-48. 1.1. not his data. This
BME Engineering Molecular Cell Biology. Basics of the Diffusion Theory. The Cytoskeleton (I)
 BME 42-620 Engineering Molecular Cell Biology Lecture 07: Basics of the Diffusion Theory The Cytoskeleton (I) BME42-620 Lecture 07, September 20, 2011 1 Outline Diffusion: microscopic theory Diffusion:
BME 42-620 Engineering Molecular Cell Biology Lecture 07: Basics of the Diffusion Theory The Cytoskeleton (I) BME42-620 Lecture 07, September 20, 2011 1 Outline Diffusion: microscopic theory Diffusion:
SECOND PUBLIC EXAMINATION. Honour School of Physics Part C: 4 Year Course. Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS
 2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2011 Monday, 27 June, 9.30 am 12.30 pm Answer
2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2011 Monday, 27 June, 9.30 am 12.30 pm Answer
BME Engineering Molecular Cell Biology. Review: Basics of the Diffusion Theory. The Cytoskeleton (I)
 BME 42-620 Engineering Molecular Cell Biology Lecture 08: Review: Basics of the Diffusion Theory The Cytoskeleton (I) BME42-620 Lecture 08, September 22, 2011 1 Outline Background: FRAP & SPT Review: microscopic
BME 42-620 Engineering Molecular Cell Biology Lecture 08: Review: Basics of the Diffusion Theory The Cytoskeleton (I) BME42-620 Lecture 08, September 22, 2011 1 Outline Background: FRAP & SPT Review: microscopic
Single molecule force spectroscopy reveals a highly compliant helical
 Supplementary Information Single molecule force spectroscopy reveals a highly compliant helical folding for the 30 nm chromatin fiber Maarten Kruithof, Fan-Tso Chien, Andrew Routh, Colin Logie, Daniela
Supplementary Information Single molecule force spectroscopy reveals a highly compliant helical folding for the 30 nm chromatin fiber Maarten Kruithof, Fan-Tso Chien, Andrew Routh, Colin Logie, Daniela
Bifurcation of Velocity Distributions in Cooperative Transport of Filaments by Fast and Slow Motors
 666 Biophysical Journal Volume 104 February 2013 666 676 Bifurcation of Velocity Distributions in Cooperative Transport of Filaments by Fast and Slow Motors Xin Li, Reinhard Lipowsky, and Jan Kierfeld
666 Biophysical Journal Volume 104 February 2013 666 676 Bifurcation of Velocity Distributions in Cooperative Transport of Filaments by Fast and Slow Motors Xin Li, Reinhard Lipowsky, and Jan Kierfeld
Macromolecular Crowding
 Macromolecular Crowding Keng-Hwee Chiam Mathematical and Theoretical Biology Group Goodsell (1994) Macromolecular Crowding, Oct. 15, 2003 p.1/33 Outline What: introduction, definition Why: implications
Macromolecular Crowding Keng-Hwee Chiam Mathematical and Theoretical Biology Group Goodsell (1994) Macromolecular Crowding, Oct. 15, 2003 p.1/33 Outline What: introduction, definition Why: implications
The Riboswitch is functionally separated into the ligand binding APTAMER and the decision-making EXPRESSION PLATFORM
 The Riboswitch is functionally separated into the ligand binding APTAMER and the decision-making EXPRESSION PLATFORM Purine riboswitch TPP riboswitch SAM riboswitch glms ribozyme In-line probing is used
The Riboswitch is functionally separated into the ligand binding APTAMER and the decision-making EXPRESSION PLATFORM Purine riboswitch TPP riboswitch SAM riboswitch glms ribozyme In-line probing is used
Lecture 13, 05 October 2004 Chapter 10, Muscle. Vertebrate Physiology ECOL 437 University of Arizona Fall instr: Kevin Bonine t.a.
 Lecture 13, 05 October 2004 Chapter 10, Muscle Vertebrate Physiology ECOL 437 University of Arizona Fall 2004 instr: Kevin Bonine t.a.: Nate Swenson Vertebrate Physiology 437 18 1. Muscle A. Sarcomere
Lecture 13, 05 October 2004 Chapter 10, Muscle Vertebrate Physiology ECOL 437 University of Arizona Fall 2004 instr: Kevin Bonine t.a.: Nate Swenson Vertebrate Physiology 437 18 1. Muscle A. Sarcomere
For slowly varying probabilities, the continuum form of these equations is. = (r + d)p T (x) (u + l)p D (x) ar x p T(x, t) + a2 r
 3.2 Molecular Motors A variety of cellular processes requiring mechanical work, such as movement, transport and packaging material, are performed with the aid of protein motors. These molecules consume
3.2 Molecular Motors A variety of cellular processes requiring mechanical work, such as movement, transport and packaging material, are performed with the aid of protein motors. These molecules consume
In biomolecular systems, free energy frequently needs to be
 Kinetic equilibrium of forces and molecular events in muscle contraction Erwin W. Becker* Institut für Mikrostrukturtechnik, Forschungszentrum, and Universität Karlsruhe, 76021 Karlsruhe, P.O. Box 3640,
Kinetic equilibrium of forces and molecular events in muscle contraction Erwin W. Becker* Institut für Mikrostrukturtechnik, Forschungszentrum, and Universität Karlsruhe, 76021 Karlsruhe, P.O. Box 3640,
The mechanical events responsible for muscle contraction are
 Repriming the actomyosin crossbridge cycle Walter Steffen and John Sleep* Randall Centre, King s College, London SE1 1UL, United Kingdom Edited by Edwin W. Taylor, University of Chicago, Chicago, IL, and
Repriming the actomyosin crossbridge cycle Walter Steffen and John Sleep* Randall Centre, King s College, London SE1 1UL, United Kingdom Edited by Edwin W. Taylor, University of Chicago, Chicago, IL, and
MOLECULAR MOTORS. The miracle of motion. Primož Dolenc Adviser: Dr. Andrej Vilfan
 MOLECULAR MOTORS The miracle of motion Primož Dolenc Adviser: Dr. Andrej Vilfan University of Ljubljana Faculty of Mathematics and Physics February 006 Abstract Biological motion from the contraction of
MOLECULAR MOTORS The miracle of motion Primož Dolenc Adviser: Dr. Andrej Vilfan University of Ljubljana Faculty of Mathematics and Physics February 006 Abstract Biological motion from the contraction of
Supporting Information Converter domain mutations in myosin alter structural kinetics and motor function. Hershey, PA, MN 55455, USA
 Supporting Information Converter domain mutations in myosin alter structural kinetics and motor function Laura K. Gunther 1, John A. Rohde 2, Wanjian Tang 1, Shane D. Walton 1, William C. Unrath 1, Darshan
Supporting Information Converter domain mutations in myosin alter structural kinetics and motor function Laura K. Gunther 1, John A. Rohde 2, Wanjian Tang 1, Shane D. Walton 1, William C. Unrath 1, Darshan
1. The plasma membrane of eukaryotic cells is supported by a. actin filaments. b. microtubules. c. lamins. d. intermediate filaments.
 ANALYSIS AND MODELING OF CELL MECHANICS Homework #2 (due 1/30/13) This homework involves comprehension of key biomechanical concepts of the cytoskeleton, cell-matrix adhesions, and cellcell adhesions.
ANALYSIS AND MODELING OF CELL MECHANICS Homework #2 (due 1/30/13) This homework involves comprehension of key biomechanical concepts of the cytoskeleton, cell-matrix adhesions, and cellcell adhesions.
ATP hydrolysis 1 1 1
 ATP hydrolysis 1 1 1 ATP hydrolysis 2 2 2 The binding zipper 1 3 3 ATP hydrolysis/synthesis is coupled to a torque Yasuda, R., et al (1998). Cell 93:1117 1124. Abrahams, et al (1994). Nature 370:621-628.
ATP hydrolysis 1 1 1 ATP hydrolysis 2 2 2 The binding zipper 1 3 3 ATP hydrolysis/synthesis is coupled to a torque Yasuda, R., et al (1998). Cell 93:1117 1124. Abrahams, et al (1994). Nature 370:621-628.
Molecular Motors. Dave Wee 24 Sept Mathematical & Theoretical Biology Seminar
 Molecular Motors Dave Wee 24 Sept 2003 Mathematical & Theoretical Biology Seminar Overview Types of motors and their working mechanisms. Illustrate the importance of motors using the example of : ATP-Synthase
Molecular Motors Dave Wee 24 Sept 2003 Mathematical & Theoretical Biology Seminar Overview Types of motors and their working mechanisms. Illustrate the importance of motors using the example of : ATP-Synthase
Flexing Protein muscles: How to Pull with a "Burning Rope"
 Flexing Protein muscles: How to Pull with a "Burning Rope" J. P. Keener 215 Joint International Conference on via Guangzhou University and Huaihua University Huaihua 8/15 p.1/28 Eukaryotic Chromosomal
Flexing Protein muscles: How to Pull with a "Burning Rope" J. P. Keener 215 Joint International Conference on via Guangzhou University and Huaihua University Huaihua 8/15 p.1/28 Eukaryotic Chromosomal
Linear Motors. Nanostrukturphysik II, Manuel Bastuck
 Molecular l Motors I: Linear Motors Nanostrukturphysik II, Manuel Bastuck Why can he run so fast? Usain Bolt 100 m / 9,58 s Usain Bolt: http://www.wallpaperdev.com/stock/fantesty-usain-bolt.jpg muscle:
Molecular l Motors I: Linear Motors Nanostrukturphysik II, Manuel Bastuck Why can he run so fast? Usain Bolt 100 m / 9,58 s Usain Bolt: http://www.wallpaperdev.com/stock/fantesty-usain-bolt.jpg muscle:
Muscle tissue. Types. Functions. Cardiac, Smooth, and Skeletal
 Types Cardiac, Smooth, and Skeletal Functions movements posture and body position Support soft tissues Guard openings body temperature nutrient reserves Muscle tissue Special Characteristics of Muscle
Types Cardiac, Smooth, and Skeletal Functions movements posture and body position Support soft tissues Guard openings body temperature nutrient reserves Muscle tissue Special Characteristics of Muscle
PHYSIOLOGY CHAPTER 9 MUSCLE TISSUE Fall 2016
 PHYSIOLOGY CHAPTER 9 MUSCLE TISSUE Fall 2016 2 Chapter 9 Muscles and Muscle Tissue Overview of Muscle Tissue types of muscle: are all prefixes for muscle Contractility all muscles cells can Smooth & skeletal
PHYSIOLOGY CHAPTER 9 MUSCLE TISSUE Fall 2016 2 Chapter 9 Muscles and Muscle Tissue Overview of Muscle Tissue types of muscle: are all prefixes for muscle Contractility all muscles cells can Smooth & skeletal
Tilting and Wobble of Myosin V by High-Speed Single-Molecule Polarized Fluorescence Microscopy
 University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 3-2013 Tilting and Wobble of Myosin V by High-Speed Single-Molecule Polarized Fluorescence Microscopy John
University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 3-2013 Tilting and Wobble of Myosin V by High-Speed Single-Molecule Polarized Fluorescence Microscopy John
Direct Modeling of X-Ray Diffraction Pattern from Skeletal Muscle in Rigor
 1082 Biophysical Journal Volume 83 August 2002 1082 1097 Direct Modeling of X-Ray Diffraction Pattern from Skeletal Muscle in Rigor Natalia A. Koubassova and A. K. Tsaturyan Institute of Mechanics, Lomonosov
1082 Biophysical Journal Volume 83 August 2002 1082 1097 Direct Modeling of X-Ray Diffraction Pattern from Skeletal Muscle in Rigor Natalia A. Koubassova and A. K. Tsaturyan Institute of Mechanics, Lomonosov
Cytokinesis in fission yeast: Modeling the assembly of the contractile ring
 Cytokinesis in fission yeast: Modeling the assembly of the contractile ring Nikola Ojkic, Dimitrios Vavylonis Department of Physics, Lehigh University Damien Laporte, Jian-Qiu Wu Department of Molecular
Cytokinesis in fission yeast: Modeling the assembly of the contractile ring Nikola Ojkic, Dimitrios Vavylonis Department of Physics, Lehigh University Damien Laporte, Jian-Qiu Wu Department of Molecular
Myosin V Movement: Lessons from Molecular Dynamics Studies of IQ Peptides in the Lever Arm
 14524 Biochemistry 2007, 46, 14524-14536 Myosin V Movement: Lessons from Molecular Dynamics Studies of IQ Peptides in the Lever Arm Assaf Ganoth, Esther Nachliel, Ran Friedman, and Menachem Gutman* Laser
14524 Biochemistry 2007, 46, 14524-14536 Myosin V Movement: Lessons from Molecular Dynamics Studies of IQ Peptides in the Lever Arm Assaf Ganoth, Esther Nachliel, Ran Friedman, and Menachem Gutman* Laser
Supporting Text - Jégou et al.
 Supporting Text - Jégou et al. I. PAUSES DURING DEPOLYMERIZATION Pauses were observed during the ymerization of individual filaments grown from ADP-, CrATP- or MgATPactin, in the presence or in the absence
Supporting Text - Jégou et al. I. PAUSES DURING DEPOLYMERIZATION Pauses were observed during the ymerization of individual filaments grown from ADP-, CrATP- or MgATPactin, in the presence or in the absence
Single molecule biochemistry using optical tweezers
 FEBS 20359 FEBS Letters 430 (1998) 23^27 Minireview Single molecule biochemistry using optical tweezers Amit D. Mehta*, Katherine A. Pullen, James A. Spudich Department of Biochemistry, Stanford University
FEBS 20359 FEBS Letters 430 (1998) 23^27 Minireview Single molecule biochemistry using optical tweezers Amit D. Mehta*, Katherine A. Pullen, James A. Spudich Department of Biochemistry, Stanford University
Supporting Information. for An atomistic modeling of F-actin mechanical responses and determination of. mechanical properties
 Supporting Information for An atomistic modeling of F-actin mechanical responses and determination of mechanical properties Authors: Si Li 1, Jin Zhang 2, Chengyuan Wang 1, Perumal Nithiarasu 1 1. Zienkiewicz
Supporting Information for An atomistic modeling of F-actin mechanical responses and determination of mechanical properties Authors: Si Li 1, Jin Zhang 2, Chengyuan Wang 1, Perumal Nithiarasu 1 1. Zienkiewicz
Actin-Binding Proteins in Nerve Cell Growth Cones
 J Pharmacol Sci 105, 6 11 (2007) Journal of Pharmacological Sciences 2007 The Japanese Pharmacological Society Current Perspective Actin-Binding Proteins in Nerve Cell Growth Cones Ryoki Ishikawa 1, *
J Pharmacol Sci 105, 6 11 (2007) Journal of Pharmacological Sciences 2007 The Japanese Pharmacological Society Current Perspective Actin-Binding Proteins in Nerve Cell Growth Cones Ryoki Ishikawa 1, *
Quantitative Analysis of Forces in Cells
 Quantitative Analysis of Forces in Cells Anders Carlsson Washington University in St Louis Basic properties of forces in cells Measurement methods and magnitudes of particular types of forces: Polymerization
Quantitative Analysis of Forces in Cells Anders Carlsson Washington University in St Louis Basic properties of forces in cells Measurement methods and magnitudes of particular types of forces: Polymerization
Announcements. Homework 3 (Klaus Schulten s Lecture): Due Wednesday at noon. Next homework assigned. Due Wednesday March 1.
 Announcements Homework 3 (Klaus Schulten s Lecture): Due Wednesday at noon. Next homework assigned. Due Wednesday March 1. No lecture next Monday, Feb. 27 th! (Homework is a bit longer.) Marco will have
Announcements Homework 3 (Klaus Schulten s Lecture): Due Wednesday at noon. Next homework assigned. Due Wednesday March 1. No lecture next Monday, Feb. 27 th! (Homework is a bit longer.) Marco will have
Neurite initiation. Neurite formation begins with a bud that sprouts from the cell body. One or several neurites can sprout at a time.
 Neurite initiation. Neuronal maturation initiation f-actin polarization and maturation tubulin stage 1: "spherical" neuron stage 2: neurons extend several neurites stage 3: one neurite accelerates its
Neurite initiation. Neuronal maturation initiation f-actin polarization and maturation tubulin stage 1: "spherical" neuron stage 2: neurons extend several neurites stage 3: one neurite accelerates its
Compliant Realignment of Binding Sites in Muscle: Transient Behavior and Mechanical Tuning
 Biophysical Journal Volume 74 April 1998 1611 1621 1611 Compliant Realignment of Binding Sites in Muscle: Transient Behavior and Mechanical Tuning Thomas L. Daniel,* Alan C. Trimble,* and P. Bryant Chase
Biophysical Journal Volume 74 April 1998 1611 1621 1611 Compliant Realignment of Binding Sites in Muscle: Transient Behavior and Mechanical Tuning Thomas L. Daniel,* Alan C. Trimble,* and P. Bryant Chase
Introduction to Polymerization Kinetics
 Introduction to Polymerization Kinetics Perspecties The cytoskeletal biopolymers are largely semi-rigid rods on typical size scale of cells. We here examine their assembly kinetics in free polymerization
Introduction to Polymerization Kinetics Perspecties The cytoskeletal biopolymers are largely semi-rigid rods on typical size scale of cells. We here examine their assembly kinetics in free polymerization
The tendency of actin protein to spontaneously polymerize into
 Actin polymerization kinetics, cap structure, and fluctuations Dimitrios Vavylonis*, Qingbo Yang, and Ben O Shaughnessy* Departments of *Chemical Engineering and Physics, Columbia University, New York,
Actin polymerization kinetics, cap structure, and fluctuations Dimitrios Vavylonis*, Qingbo Yang, and Ben O Shaughnessy* Departments of *Chemical Engineering and Physics, Columbia University, New York,
Does lamellipod slip under cantilever? AFM measurements show that height of lamellipod~ nm
 SMR 1746-7 WORKSHOP ON DRIVEN STATES IN SOFT AND BIOLOGICAL MATTER 18-28 April 2006 New Methods to Study Cell Migration Part II Kenneth JACOBSON University of North Carolina at Chapel Hill, School of Medicine,
SMR 1746-7 WORKSHOP ON DRIVEN STATES IN SOFT AND BIOLOGICAL MATTER 18-28 April 2006 New Methods to Study Cell Migration Part II Kenneth JACOBSON University of North Carolina at Chapel Hill, School of Medicine,
Multimedia : Fibronectin and Titin unfolding simulation movies.
 I LECTURE 21: SINGLE CHAIN ELASTICITY OF BIOMACROMOLECULES: THE GIANT PROTEIN TITIN AND DNA Outline : REVIEW LECTURE #2 : EXTENSIBLE FJC AND WLC... 2 STRUCTURE OF MUSCLE AND TITIN... 3 SINGLE MOLECULE
I LECTURE 21: SINGLE CHAIN ELASTICITY OF BIOMACROMOLECULES: THE GIANT PROTEIN TITIN AND DNA Outline : REVIEW LECTURE #2 : EXTENSIBLE FJC AND WLC... 2 STRUCTURE OF MUSCLE AND TITIN... 3 SINGLE MOLECULE
Lecture 7 : Molecular Motors. Dr Eileen Nugent
 Lecture 7 : Molecular Motors Dr Eileen Nugent Molecular Motors Energy Sources: Protonmotive Force, ATP Single Molecule Biophysical Techniques : Optical Tweezers, Atomic Force Microscopy, Single Molecule
Lecture 7 : Molecular Motors Dr Eileen Nugent Molecular Motors Energy Sources: Protonmotive Force, ATP Single Molecule Biophysical Techniques : Optical Tweezers, Atomic Force Microscopy, Single Molecule
Regularity and synchrony in motor proteins
 Regularity and synchrony in motor proteins R. E. Lee DeVille and Eric Vanden-Eijnden Courant Institute of Mathematical Sciences, New York University, New York, NY 10012 1 Corresponding author: R. E. Lee
Regularity and synchrony in motor proteins R. E. Lee DeVille and Eric Vanden-Eijnden Courant Institute of Mathematical Sciences, New York University, New York, NY 10012 1 Corresponding author: R. E. Lee
Significant Impact on Muscle Mechanics of Small Nonlinearities in Myofilament Elasticity
 Biophysical Journal Volume 99 September 21 1869 1875 1869 Significant Impact on Muscle Mechanics of Small Nonlinearities in Myofilament Elasticity Alf Månsson* School of Natural Sciences, Linnaeus University,
Biophysical Journal Volume 99 September 21 1869 1875 1869 Significant Impact on Muscle Mechanics of Small Nonlinearities in Myofilament Elasticity Alf Månsson* School of Natural Sciences, Linnaeus University,
Supplementary Figure 1: Spn27A-RNAi embryos are partially ventralized, whereas
 Figure S Fat2- eve hkb sim sog wild-type Supplementary Figure : embryos are partially ventralized, whereas patterning is not affected in Fat2- embryos. RNA in situ hybridization of wild-type, and Fat2-
Figure S Fat2- eve hkb sim sog wild-type Supplementary Figure : embryos are partially ventralized, whereas patterning is not affected in Fat2- embryos. RNA in situ hybridization of wild-type, and Fat2-
Nuclear import of DNA: genetic modification of plants
 Nuclear import of DNA: genetic modification of plants gene delivery by Agrobacterium tumefaciens T. Tzfira & V. Citovsky. 2001. Trends in Cell Biol. 12: 121-129 VirE2 binds ssdna in vitro forms helical
Nuclear import of DNA: genetic modification of plants gene delivery by Agrobacterium tumefaciens T. Tzfira & V. Citovsky. 2001. Trends in Cell Biol. 12: 121-129 VirE2 binds ssdna in vitro forms helical
Molecular Dynamics Simulation and Coarse-Grained Analysis of the Arp2/3 Complex
 5324 Biophysical Journal Volume 95 December 2008 5324 5333 Molecular Dynamics Simulation and Coarse-Grained Analysis of the Arp2/3 Complex Jim Pfaendtner* y and Gregory A. Voth* y *Center for Biophysical
5324 Biophysical Journal Volume 95 December 2008 5324 5333 Molecular Dynamics Simulation and Coarse-Grained Analysis of the Arp2/3 Complex Jim Pfaendtner* y and Gregory A. Voth* y *Center for Biophysical
Sarcomere Lattice Geometry Influences Cooperative Myosin Binding in Muscle
 Sarcomere Lattice Geometry Influences Cooperative Myosin Binding in Muscle Bertrand C. W. Tanner 1, Thomas L. Daniel 2, Michael Regnier 1* 1 Department of Bioengineering, University of Washington, Seattle,
Sarcomere Lattice Geometry Influences Cooperative Myosin Binding in Muscle Bertrand C. W. Tanner 1, Thomas L. Daniel 2, Michael Regnier 1* 1 Department of Bioengineering, University of Washington, Seattle,
Nanomotors: Nanoscale machines
 Nanomotors: Nanoscale machines October 31, 2016 1 Introduction to nanomotors In this part of the course we will study nanomotors. First we will define what we mean by nanomotor. A motor (of any size) is
Nanomotors: Nanoscale machines October 31, 2016 1 Introduction to nanomotors In this part of the course we will study nanomotors. First we will define what we mean by nanomotor. A motor (of any size) is
BIOMECHANICS 3 Origins and consequences of forces in biological systems
 BIOMECHANICS 3 Origins and consequences of forces in biological systems MOLECULAR MECHANISMS OF BIOLOGICAL MOVEMENT AT THE LEVELOF ORGANISMS MOLECULAR BASIS OF MUSCLE CONTRACTION DR. BEÁTA BUGYI - BIOPHYSICS
BIOMECHANICS 3 Origins and consequences of forces in biological systems MOLECULAR MECHANISMS OF BIOLOGICAL MOVEMENT AT THE LEVELOF ORGANISMS MOLECULAR BASIS OF MUSCLE CONTRACTION DR. BEÁTA BUGYI - BIOPHYSICS
Outline. The ensemble folding kinetics of protein G from an all-atom Monte Carlo simulation. Unfolded Folded. What is protein folding?
 The ensemble folding kinetics of protein G from an all-atom Monte Carlo simulation By Jun Shimada and Eugine Shaknovich Bill Hawse Dr. Bahar Elisa Sandvik and Mehrdad Safavian Outline Background on protein
The ensemble folding kinetics of protein G from an all-atom Monte Carlo simulation By Jun Shimada and Eugine Shaknovich Bill Hawse Dr. Bahar Elisa Sandvik and Mehrdad Safavian Outline Background on protein
Improved Resolution of Tertiary Structure Elasticity in Muscle Protein
 Improved Resolution of Tertiary Structure Elasticity in Muscle Protein Jen Hsin and Klaus Schulten* Department of Physics and Beckman Institute, University of Illinois at Urbana-Champaign, Urbana, Illinois
Improved Resolution of Tertiary Structure Elasticity in Muscle Protein Jen Hsin and Klaus Schulten* Department of Physics and Beckman Institute, University of Illinois at Urbana-Champaign, Urbana, Illinois
Use of Stable Analogs of Myosin ATPase Intermediates for Kinetic Studies of the Weak Binding of Myosin Heads to F-Actin
 Biochemistry (Moscow), Vol. 64, No. 8, 1999, pp. 875-882. Translated from Biokhimiya, Vol. 64, No. 8, 1999, pp. 1043-1051. Original Russian Text Copyright 1999 by Rostkova, Moiseeva, Teplova, Nikolaeva,
Biochemistry (Moscow), Vol. 64, No. 8, 1999, pp. 875-882. Translated from Biokhimiya, Vol. 64, No. 8, 1999, pp. 1043-1051. Original Russian Text Copyright 1999 by Rostkova, Moiseeva, Teplova, Nikolaeva,
BMB Lecture 7. Allostery and Cooperativity
 BMB 178 2017 Lecture 7 October 18, 2017 Allostery and Cooperativity A means for exquisite control Allostery: the basis of enzymatic control From the Greek: allos = other stereos = solid or space Action
BMB 178 2017 Lecture 7 October 18, 2017 Allostery and Cooperativity A means for exquisite control Allostery: the basis of enzymatic control From the Greek: allos = other stereos = solid or space Action
Chapter 4 Bidirectional membrane tubes driven by nonprocessive motors
 Chapter 4 Bidirectional membrane tubes driven by nonprocessive motors In cells, membrane tubes are extracted by molecular motors. Although individual motors cannot provide enough force to pull a tube,
Chapter 4 Bidirectional membrane tubes driven by nonprocessive motors In cells, membrane tubes are extracted by molecular motors. Although individual motors cannot provide enough force to pull a tube,
