Journal of Avian Biology
|
|
|
- Arnold Gordon
- 5 years ago
- Views:
Transcription
1 Journal of Avian Biology JAB5728 Brown, J. L. and Collopy, M. W Immigration stabilizes a population of threatened cavity-nesting raptors despite possibility of nest site imprinting. J. Avian Biol. 43: xxx xxx. Supplementary material
2 Appendix 1 Joint likelihood and prior probabilities of the integrated population models Joint likelihood function The joint likelihood of our female-based IPMs was the product of the individual likelihoods of each of our model components, which were informed by a population census, fecundity estimates, and Cormack-Jolly-Seber models of our marked adults and nestlings. We treated our data sets as independent although many of the same adult females were represented in multiple data sets, because a simulation study examining the impacts of violation of the assumption of independence suggested minor impacts on the accuracy of parameter estimates. Therefore, under the assumption of independence between the four data sources, the joint likelihood is L joint (m, J, R, y N,Φ juv, Φ ad, f, imm, p juv,p ad ) = L cr (m Φ juv, Φ ad,p juv,p ad ) L rp (J, R f) L ob (y N) L sy (N Φ juv, Φ ad, f, imm) where L cr is the likelihood for the Cormack-Jolly-Seber mark-recapture model, L rp is the likelihood for the fecundity model, and L ob and L sy are the likelihoods for the observation and state process components of the state-space model.
3 Prior probabilities All prior probabilities were selected to be vague and uninformative, except for initial population numbers in 2008, and are presented in WinBUGS format. We assumed our raw population count in 2008 was less than, but not by much, the latent number of individuals present. We adopted Normal(50, 0.001) priors for the initial population sizes per age class in 2008 for the model without immigration (normal distribution with a mean of 50 and precision, or 1/variance, of 0.001). For the model with immigration, we examined whether the final estimated distribution of kestrels amongst age-classes was dependent on the numbers specified in the priors for 2008 by varying these priors in separate model runs. Therefore, we ran a model with Normal(33, 0.01) for local yearlings, Normal(34, 0.01) for local adults, and Normal(33, 0.01) for immigrants, and another with Normal(10, 0.01), for yearlings, Normal(55, 0.01) for local adults, and Normal(35, 0.01) for immigrants; all truncated to 0. Model results were very similar regardless of priors chosen, so we performed the remaining analyses with the model with the roughly balanced priors, i.e. specifying the age-class distribution as 33, 34, and 33. The priors for the other model parameters were as follows (parameters of the uniform distribution specifies the upper and lower limits; parameters specified in the beta distribution are the α and β shape parameters): f t Uniform(0, 10) imm t Uniform(0, 20) Φ i,t Beta(1, 1) p i,t Beta(1, 1)
4 Appendix 2 Example WinBUGS code for integrated population model (IPM) with immigration ######################################################################## # integrated population assessment of American kestrels in Florida # uses CJS model of adult kestrels and nestling kestrels # observed nest box attendance as index of population size # and observed fecundity in nest boxes # code written to import and prepare data in R # and call WinBUGs using the package R2WinBUGS ####################################################################### library("r2winbugs") # making a stacked m-array of mark-recapture data with top two rows marked as # juveniles and bottom two rows mixture of previous juveniles (first enter # after first capture as adults) and marked as adults mfem <- matrix(c(2, 1, 180, 0, 6, 196, 32, 0, 35, 0, 14, 34), nrow=4, ncol=3, byrow=t) # temporal specs ni <- 2 # number of release occasions nj <- 2 # number of recapture occasions nyear<-3
5 T <- nyear # census data, which is observed females # making sure not to duplicate nests in territories yc3 <- c(88,83,85) # number of breeding females Nb <- c(80,77,74) # nestlings, half of observed nestlings nestlings3 <-c(113, 107, 122) ####################################################################### # integrated population model with immigration ####################################################################### sink("ipm_immcjs.txt") cat(" model { #defining recapture and survival parameters for (i in 1:nj) { pn[i] <- param[1]
6 p[i] <- param[2] Sn[i] <- param[2+i] S[i] <- param[4+i] # define priors on beta scale for (i in 1:6) { param[i] ~ dbeta(1,1) # define CJS model likelihood for (i in 1:2*ni) { mfem[i, 1:(nj+1)] ~ dmulti(pr[i, ], r[i]) # Calculate number of birds released each year, or row total for (i in 1:2*ni) { r[i] <- sum(mfem[i, ]) # Calculate multinomial cell probabilities # Calculate diagonal, above diagonal, and below for juveniles for (i in 1:ni) {
7 qn[i] <- 1-pn[i] pr[i,i] <- Sn[i]*pn[i] for (j in (i+1):nj) { pr[i,j] <- Sn[i]*prod(S[(i+1):j])*prod(qn[i:(j-1)])*p[j] for (j in 1:(i-1)) { pr[i,j] <- 0 # probability of juvenile bird not being seen again pr[i, nj+1] <- 1-sum(pr[i, 1:nj]) # Cell probs for adults # Calculate diagonal, above diagonal, and below for adults for (i in 1:ni) { q[i] <- 1-p[i] pr[i+ni,i] <- S[i]*p[i] for (j in (i+1):nj) { pr[i+ni,j] <- prod(s[i:j])*prod(q[i:(j-1)])*p[j] for (j in 1:(i-1)) { pr[i+ni,j] <- 0
8 # probability of adult bird not being seen again pr[i+ni, nj+1] <- 1-sum(pr[i+ni, 1:nj]) # define lambda, mean survival, recapture, and pop growth rate # could do means post hoc, but this method generates CI for(t in 1:T-1){ lambda[t] <- Ntot[t+1]/Ntot[t] logla[t] <- log(lambda[t]) logphij[t] <- log(sn[t]) logphia[t] <- log(s[t]) MELAM <- exp((1/(t-1))*sum(logla[1:(t-1)])) MEPHIJ <- exp((1/(t-1))*sum(logphij[1:(t-1)])) MEPHIA <- exp((1/(t-1))*sum(logphia[1:(t-1)])) # priors for population N1[1] ~ dnorm(10, 0.01)I(0,) Nad[1] ~ dnorm(55, 0.01)I(0,) Nadimm[1] ~ dnorm(35, 0.01)I(0,)
9 # system process using binomial and poisson for(tt in 2:T){ mean1[tt]<-fec[tt-1]*sn[tt-1]*ntot[tt-1] mpo[tt] <- Ntot[tt-1]*imm[tt-1] N1[tt]~dpois(mean1[tt]) Nad[tt]~dbin(S[tt-1],Ntot[tt-1]) Nadimm[tt] ~ dpois(mpo[tt]) # observation process for above for(tt in 1:T){ Ntot[tt]<-Nad[tt] + N1[tt] + Nadimm[tt] yc3[tt]~dpois(ntot[tt]) # immigration priors for(j in 1:nyear-1){ imm[j] ~ dunif(0,20) # fecundity priors for (t in 1:T){ fec[t] ~ dunif(0,10)
10 # likelihood for reproductive data for (t in 1:T){ nestlings3[t] ~ dpois(rho[t]) rho[t] <- Nb[t]*fec[t] ", fill=true) sink() # specifying data for WinBUGS data <- list ("ni", "nj", "T", "nyear", "Nb", "yc3", "nestlings3", "mfem") # specifying init generation functions inits <- function(){ list(n1=round(runif(3,5,30),0), Nad=round(runif(3,5,55),0), Nadimm=round(runif(3,5,55),0), fec=runif(3, 0.5, 4), param=runif(6, 0.01, 0.99), imm=runif(2,0,5) )
11 # specifying which parameters to monitor parameters <- c("param", "Ntot", "fec", "N1", "Nad", "rho", "lambda", "imm", "Nadimm", "MELAM", "MEPHIJ", "MEPHIA") # calling WinBUGS from within R IPM_ImmCJS.mod <- bugs (data, inits, parameters, "IPM_ImmCJS.txt", n.thin=12, n.chains=2, n.burnin=5000, n.iter=140000,debug=t, codapkg=t) # reading model results into CODA for further analysis IPM_ImmCJS.coda <- read.bugs(ipm_immcjs.mod) # exporting posterior probability summaries to csv files IPM_Imm.out <- summary(ipm_imm.coda, quantiles=c(0.025, 0.25, 0.5, 0.75, 0.975)) IPM_Imm.hpd <- HPDinterval(IPM_Imm.coda, prob=0.95) write.csv(ipm_imm.out$statistics, "IPM_ImmStats.csv") write.csv(ipm_imm.out$quantiles, "IPM_ImmQs.csv") write.csv(ipm_imm.hpd, "IPM_ImmHPD.csv")
Barn swallow post-fledging survival: Using Stan to fit a hierarchical ecological model. Fränzi Korner-Nievergelt Beat Naef-Daenzer Martin U.
 Barn swallow post-fledging survival: Using Stan to fit a hierarchical ecological model Fränzi Korner-Nievergelt Beat Naef-Daenzer Martin U. Grüebler Introduction Population dynamics Youngs Juvenile survival
Barn swallow post-fledging survival: Using Stan to fit a hierarchical ecological model Fränzi Korner-Nievergelt Beat Naef-Daenzer Martin U. Grüebler Introduction Population dynamics Youngs Juvenile survival
FW Laboratory Exercise. Program MARK: Joint Live Recapture and Dead Recovery Data and Pradel Model
 FW663 -- Laboratory Exercise Program MARK: Joint Live Recapture and Dead Recovery Data and Pradel Model Today s exercise explores parameter estimation using both live recaptures and dead recoveries. We
FW663 -- Laboratory Exercise Program MARK: Joint Live Recapture and Dead Recovery Data and Pradel Model Today s exercise explores parameter estimation using both live recaptures and dead recoveries. We
Cormack-Jolly-Seber Models
 Cormack-Jolly-Seber Models Estimating Apparent Survival from Mark-Resight Data & Open-Population Models Ch. 17 of WNC, especially sections 17.1 & 17.2 For these models, animals are captured on k occasions
Cormack-Jolly-Seber Models Estimating Apparent Survival from Mark-Resight Data & Open-Population Models Ch. 17 of WNC, especially sections 17.1 & 17.2 For these models, animals are captured on k occasions
Robust Bayesian Regression
 Readings: Hoff Chapter 9, West JRSSB 1984, Fúquene, Pérez & Pericchi 2015 Duke University November 17, 2016 Body Fat Data: Intervals w/ All Data Response % Body Fat and Predictor Waist Circumference 95%
Readings: Hoff Chapter 9, West JRSSB 1984, Fúquene, Pérez & Pericchi 2015 Duke University November 17, 2016 Body Fat Data: Intervals w/ All Data Response % Body Fat and Predictor Waist Circumference 95%
Advanced Statistical Modelling
 Markov chain Monte Carlo (MCMC) Methods and Their Applications in Bayesian Statistics School of Technology and Business Studies/Statistics Dalarna University Borlänge, Sweden. Feb. 05, 2014. Outlines 1
Markov chain Monte Carlo (MCMC) Methods and Their Applications in Bayesian Statistics School of Technology and Business Studies/Statistics Dalarna University Borlänge, Sweden. Feb. 05, 2014. Outlines 1
Introduction to capture-markrecapture
 E-iNET Workshop, University of Kent, December 2014 Introduction to capture-markrecapture models Rachel McCrea Overview Introduction Lincoln-Petersen estimate Maximum-likelihood theory* Capture-mark-recapture
E-iNET Workshop, University of Kent, December 2014 Introduction to capture-markrecapture models Rachel McCrea Overview Introduction Lincoln-Petersen estimate Maximum-likelihood theory* Capture-mark-recapture
WinBUGS : part 2. Bruno Boulanger Jonathan Jaeger Astrid Jullion Philippe Lambert. Gabriele, living with rheumatoid arthritis
 WinBUGS : part 2 Bruno Boulanger Jonathan Jaeger Astrid Jullion Philippe Lambert Gabriele, living with rheumatoid arthritis Agenda 2! Hierarchical model: linear regression example! R2WinBUGS Linear Regression
WinBUGS : part 2 Bruno Boulanger Jonathan Jaeger Astrid Jullion Philippe Lambert Gabriele, living with rheumatoid arthritis Agenda 2! Hierarchical model: linear regression example! R2WinBUGS Linear Regression
Lecture 7 Models for open populations: Tag recovery and CJS models, Goodness-of-fit
 WILD 7250 - Analysis of Wildlife Populations 1 of 16 Lecture 7 Models for open populations: Tag recovery and CJS models, Goodness-of-fit Resources Chapter 5 in Goodness of fit in E. Cooch and G.C. White
WILD 7250 - Analysis of Wildlife Populations 1 of 16 Lecture 7 Models for open populations: Tag recovery and CJS models, Goodness-of-fit Resources Chapter 5 in Goodness of fit in E. Cooch and G.C. White
Bayesian Inference in Camera Trapping Studies for a Class of Spatial Capture-Recapture Models. Appendix A: Analysis of the Poisson encounter model
 Bayesian Inference in Camera Trapping Studies for a Class of Spatial Capture-Recapture Models J. Andrew Royle, U.S. Geological Survey, Patuxent Wildlife Research Center, Laurel, Maryland, 20708, email:
Bayesian Inference in Camera Trapping Studies for a Class of Spatial Capture-Recapture Models J. Andrew Royle, U.S. Geological Survey, Patuxent Wildlife Research Center, Laurel, Maryland, 20708, email:
Joint live encounter & dead recovery data
 Joint live encounter & dead recovery data CHAPTER 8 The first chapters in this book focussed on typical open population mark-recapture models, where the probability of an individual being encountered (dead
Joint live encounter & dead recovery data CHAPTER 8 The first chapters in this book focussed on typical open population mark-recapture models, where the probability of an individual being encountered (dead
Trevor Davies, Dalhousie University, Halifax, Canada Steven J.D. Martell, International Pacific Halibut Commission, Seattle WA.
 Comparison of ADMB-RE and JAGS Bayesian State-space length disaggregated population model: Estimating total mortality by decade for winter skate (Leucoraja ocellata) Trevor Davies, Dalhousie University,
Comparison of ADMB-RE and JAGS Bayesian State-space length disaggregated population model: Estimating total mortality by decade for winter skate (Leucoraja ocellata) Trevor Davies, Dalhousie University,
7.2 EXPLORING ECOLOGICAL RELATIONSHIPS IN SURVIVAL AND ESTIMATING RATES OF POPULATION CHANGE USING PROGRAM MARK
 350 International Wildlife Management Congress 7.2 7.2 EXPLORING ECOLOGICAL RELATIONSHIPS IN SURVIVAL AND ESTIMATING RATES OF POPULATION CHANGE USING PROGRAM MARK ALAN B. FRANKLIN Colorado Cooperative
350 International Wildlife Management Congress 7.2 7.2 EXPLORING ECOLOGICAL RELATIONSHIPS IN SURVIVAL AND ESTIMATING RATES OF POPULATION CHANGE USING PROGRAM MARK ALAN B. FRANKLIN Colorado Cooperative
Bayesian Networks in Educational Assessment
 Bayesian Networks in Educational Assessment Estimating Parameters with MCMC Bayesian Inference: Expanding Our Context Roy Levy Arizona State University Roy.Levy@asu.edu 2017 Roy Levy MCMC 1 MCMC 2 Posterior
Bayesian Networks in Educational Assessment Estimating Parameters with MCMC Bayesian Inference: Expanding Our Context Roy Levy Arizona State University Roy.Levy@asu.edu 2017 Roy Levy MCMC 1 MCMC 2 Posterior
Bayesian Inference for Regression Parameters
 Bayesian Inference for Regression Parameters 1 Bayesian inference for simple linear regression parameters follows the usual pattern for all Bayesian analyses: 1. Form a prior distribution over all unknown
Bayesian Inference for Regression Parameters 1 Bayesian inference for simple linear regression parameters follows the usual pattern for all Bayesian analyses: 1. Form a prior distribution over all unknown
PIER HLM Course July 30, 2011 Howard Seltman. Discussion Guide for Bayes and BUGS
 PIER HLM Course July 30, 2011 Howard Seltman Discussion Guide for Bayes and BUGS 1. Classical Statistics is based on parameters as fixed unknown values. a. The standard approach is to try to discover,
PIER HLM Course July 30, 2011 Howard Seltman Discussion Guide for Bayes and BUGS 1. Classical Statistics is based on parameters as fixed unknown values. a. The standard approach is to try to discover,
Journal of Avian Biology
 Journal of Avian Biology JAV-01 McKnight, A., Blomberg, E. J., Golet, G. H., Irons, D. B., Loftin, C. S. and McKinney, S. T. 01. Experimental evidence of long-term reproductive costs in a colonial nesting
Journal of Avian Biology JAV-01 McKnight, A., Blomberg, E. J., Golet, G. H., Irons, D. B., Loftin, C. S. and McKinney, S. T. 01. Experimental evidence of long-term reproductive costs in a colonial nesting
FW Laboratory Exercise. Program MARK with Mark-Recapture Data
 FW663 -- Laboratory Exercise Program MARK with Mark-Recapture Data This exercise brings us to the land of the living! That is, instead of estimating survival from dead animal recoveries, we will now estimate
FW663 -- Laboratory Exercise Program MARK with Mark-Recapture Data This exercise brings us to the land of the living! That is, instead of estimating survival from dead animal recoveries, we will now estimate
Linear Regression. Data Model. β, σ 2. Process Model. ,V β. ,s 2. s 1. Parameter Model
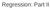 Regression: Part II Linear Regression y~n X, 2 X Y Data Model β, σ 2 Process Model Β 0,V β s 1,s 2 Parameter Model Assumptions of Linear Model Homoskedasticity No error in X variables Error in Y variables
Regression: Part II Linear Regression y~n X, 2 X Y Data Model β, σ 2 Process Model Β 0,V β s 1,s 2 Parameter Model Assumptions of Linear Model Homoskedasticity No error in X variables Error in Y variables
Environmental effects and individual body condition drive seasonal fecundity of rabbits: identifying acute and lagged processes
 Electronic supplementary material (ESM 2) for Environmental effects and individual body condition drive seasonal fecundity of rabbits: identifying acute and lagged processes Konstans Wells Robert B. O
Electronic supplementary material (ESM 2) for Environmental effects and individual body condition drive seasonal fecundity of rabbits: identifying acute and lagged processes Konstans Wells Robert B. O
DIC: Deviance Information Criterion
 (((( Welcome Page Latest News DIC: Deviance Information Criterion Contact us/bugs list WinBUGS New WinBUGS examples FAQs DIC GeoBUGS DIC (Deviance Information Criterion) is a Bayesian method for model
(((( Welcome Page Latest News DIC: Deviance Information Criterion Contact us/bugs list WinBUGS New WinBUGS examples FAQs DIC GeoBUGS DIC (Deviance Information Criterion) is a Bayesian method for model
WinBUGS for Population Ecologists: Bayesian Modeling Using Markov Chain Monte Carlo Methods
 WinBUGS for Population Ecologists: Bayesian Modeling Using Markov Chain Monte Carlo Methods Olivier Gimenez, Simon J. Bonner, Ruth King, Richard A. Parker, Stephen P. Brooks, Lara E. Jamieson, Vladimir
WinBUGS for Population Ecologists: Bayesian Modeling Using Markov Chain Monte Carlo Methods Olivier Gimenez, Simon J. Bonner, Ruth King, Richard A. Parker, Stephen P. Brooks, Lara E. Jamieson, Vladimir
Why Bayesian approaches? The average height of a rare plant
 Why Bayesian approaches? The average height of a rare plant Estimation and comparison of averages is an important step in many ecological analyses and demographic models. In this demonstration you will
Why Bayesian approaches? The average height of a rare plant Estimation and comparison of averages is an important step in many ecological analyses and demographic models. In this demonstration you will
Integrating mark-resight, count, and photograph data to more effectively monitor non-breeding American oystercatcher populations
 Integrating mark-resight, count, and photograph data to more effectively monitor non-breeding American oystercatcher populations Gibson, Daniel, Thomas V. Riecke, Tim Keyes, Chris Depkin, Jim Fraser, and
Integrating mark-resight, count, and photograph data to more effectively monitor non-breeding American oystercatcher populations Gibson, Daniel, Thomas V. Riecke, Tim Keyes, Chris Depkin, Jim Fraser, and
Input from capture mark recapture methods to the understanding of population biology
 Input from capture mark recapture methods to the understanding of population biology Roger Pradel, iostatistics and Population iology team CEFE, Montpellier, France 1 Why individual following? There are
Input from capture mark recapture methods to the understanding of population biology Roger Pradel, iostatistics and Population iology team CEFE, Montpellier, France 1 Why individual following? There are
THE JOLLY-SEBER MODEL AND MAXIMUM LIKELIHOOD ESTIMATION REVISITED. Walter K. Kremers. Biometrics Unit, Cornell, Ithaca, New York ABSTRACT
 THE JOLLY-SEBER MODEL AND MAXIMUM LIKELIHOOD ESTIMATION REVISITED BU-847-M by Walter K. Kremers August 1984 Biometrics Unit, Cornell, Ithaca, New York ABSTRACT Seber (1965, B~ometr~ka 52: 249-259) describes
THE JOLLY-SEBER MODEL AND MAXIMUM LIKELIHOOD ESTIMATION REVISITED BU-847-M by Walter K. Kremers August 1984 Biometrics Unit, Cornell, Ithaca, New York ABSTRACT Seber (1965, B~ometr~ka 52: 249-259) describes
Jolly-Seber models in MARK
 Chapter 12 Jolly-Seber models in MARK Carl James Schwarz, Simon Fraser University A. Neil Arnason, University of Manitoba The original Jolly-Seber (JS) model (Jolly, 1965; Seber, 1965) was primarily interested
Chapter 12 Jolly-Seber models in MARK Carl James Schwarz, Simon Fraser University A. Neil Arnason, University of Manitoba The original Jolly-Seber (JS) model (Jolly, 1965; Seber, 1965) was primarily interested
CHAPTER 12. Jolly-Seber models in MARK Protocol. Carl James Schwarz, Simon Fraser University A. Neil Arnason, University of Manitoba
 CHAPTER 12 Jolly-Seber models in MARK Carl James Schwarz, Simon Fraser University A. Neil Arnason, University of Manitoba The original Jolly-Seber (JS) model (Jolly, 1965; Seber, 1965) was primarily interested
CHAPTER 12 Jolly-Seber models in MARK Carl James Schwarz, Simon Fraser University A. Neil Arnason, University of Manitoba The original Jolly-Seber (JS) model (Jolly, 1965; Seber, 1965) was primarily interested
Parameter Redundancy with Applications in Statistical Ecology
 Parameter Redundancy with Applications in Statistical Ecology Ben Arthur Hubbard A Thesis submitted for the degree of Doctor of Philosophy in the subject of Statistics School of Mathematics, Statistics
Parameter Redundancy with Applications in Statistical Ecology Ben Arthur Hubbard A Thesis submitted for the degree of Doctor of Philosophy in the subject of Statistics School of Mathematics, Statistics
Mark-Recapture. Mark-Recapture. Useful in estimating: Lincoln-Petersen Estimate. Lincoln-Petersen Estimate. Lincoln-Petersen Estimate
 Mark-Recapture Mark-Recapture Modern use dates from work by C. G. J. Petersen (Danish fisheries biologist, 1896) and F. C. Lincoln (U. S. Fish and Wildlife Service, 1930) Useful in estimating: a. Size
Mark-Recapture Mark-Recapture Modern use dates from work by C. G. J. Petersen (Danish fisheries biologist, 1896) and F. C. Lincoln (U. S. Fish and Wildlife Service, 1930) Useful in estimating: a. Size
Parametric Modelling of Over-dispersed Count Data. Part III / MMath (Applied Statistics) 1
 Parametric Modelling of Over-dispersed Count Data Part III / MMath (Applied Statistics) 1 Introduction Poisson regression is the de facto approach for handling count data What happens then when Poisson
Parametric Modelling of Over-dispersed Count Data Part III / MMath (Applied Statistics) 1 Introduction Poisson regression is the de facto approach for handling count data What happens then when Poisson
Climate change drives microevolution in a wild bird
 Supplementary information for Climate change drives microevolution in a wild bird Patrik Karell, Kari Ahola, Teuvo Karstinen, Jari Valkama & Jon E. Brommer This document includes: -Supplementary Figures
Supplementary information for Climate change drives microevolution in a wild bird Patrik Karell, Kari Ahola, Teuvo Karstinen, Jari Valkama & Jon E. Brommer This document includes: -Supplementary Figures
R Demonstration ANCOVA
 R Demonstration ANCOVA Objective: The purpose of this week s session is to demonstrate how to perform an analysis of covariance (ANCOVA) in R, and how to plot the regression lines for each level of the
R Demonstration ANCOVA Objective: The purpose of this week s session is to demonstrate how to perform an analysis of covariance (ANCOVA) in R, and how to plot the regression lines for each level of the
Generalized Linear Model under the Extended Negative Multinomial Model and Cancer Incidence
 Generalized Linear Model under the Extended Negative Multinomial Model and Cancer Incidence Sunil Kumar Dhar Center for Applied Mathematics and Statistics, Department of Mathematical Sciences, New Jersey
Generalized Linear Model under the Extended Negative Multinomial Model and Cancer Incidence Sunil Kumar Dhar Center for Applied Mathematics and Statistics, Department of Mathematical Sciences, New Jersey
Webinar Session 1. Introduction to Modern Methods for Analyzing Capture- Recapture Data: Closed Populations 1
 Webinar Session 1. Introduction to Modern Methods for Analyzing Capture- Recapture Data: Closed Populations 1 b y Bryan F.J. Manly Western Ecosystems Technology Inc. Cheyenne, Wyoming bmanly@west-inc.com
Webinar Session 1. Introduction to Modern Methods for Analyzing Capture- Recapture Data: Closed Populations 1 b y Bryan F.J. Manly Western Ecosystems Technology Inc. Cheyenne, Wyoming bmanly@west-inc.com
36-463/663Multilevel and Hierarchical Models
 36-463/663Multilevel and Hierarchical Models From Bayes to MCMC to MLMs Brian Junker 132E Baker Hall brian@stat.cmu.edu 1 Outline Bayesian Statistics and MCMC Distribution of Skill Mastery in a Population
36-463/663Multilevel and Hierarchical Models From Bayes to MCMC to MLMs Brian Junker 132E Baker Hall brian@stat.cmu.edu 1 Outline Bayesian Statistics and MCMC Distribution of Skill Mastery in a Population
SUPPLEMENT TO EXTENDING THE LATENT MULTINOMIAL MODEL WITH COMPLEX ERROR PROCESSES AND DYNAMIC MARKOV BASES
 SUPPLEMENT TO EXTENDING THE LATENT MULTINOMIAL MODEL WITH COMPLEX ERROR PROCESSES AND DYNAMIC MARKOV BASES By Simon J Bonner, Matthew R Schofield, Patrik Noren Steven J Price University of Western Ontario,
SUPPLEMENT TO EXTENDING THE LATENT MULTINOMIAL MODEL WITH COMPLEX ERROR PROCESSES AND DYNAMIC MARKOV BASES By Simon J Bonner, Matthew R Schofield, Patrik Noren Steven J Price University of Western Ontario,
Climate, social factors and research disturbance influence population dynamics in a declining sociable weaver metapopulation
 DOI 10.1007/s00442-013-2768-7 Population ecology - Original research Climate, social factors and research disturbance influence population dynamics in a declining sociable weaver metapopulation Res Altwegg
DOI 10.1007/s00442-013-2768-7 Population ecology - Original research Climate, social factors and research disturbance influence population dynamics in a declining sociable weaver metapopulation Res Altwegg
MCMC 2: Lecture 2 Coding and output. Phil O Neill Theo Kypraios School of Mathematical Sciences University of Nottingham
 MCMC 2: Lecture 2 Coding and output Phil O Neill Theo Kypraios School of Mathematical Sciences University of Nottingham Contents 1. General (Markov) epidemic model 2. Non-Markov epidemic model 3. Debugging
MCMC 2: Lecture 2 Coding and output Phil O Neill Theo Kypraios School of Mathematical Sciences University of Nottingham Contents 1. General (Markov) epidemic model 2. Non-Markov epidemic model 3. Debugging
Review Article Putting Markov Chains Back into Markov Chain Monte Carlo
 Hindawi Publishing Corporation Journal of Applied Mathematics and Decision Sciences Volume 2007, Article ID 98086, 13 pages doi:10.1155/2007/98086 Review Article Putting Markov Chains Back into Markov
Hindawi Publishing Corporation Journal of Applied Mathematics and Decision Sciences Volume 2007, Article ID 98086, 13 pages doi:10.1155/2007/98086 Review Article Putting Markov Chains Back into Markov
EXERCISE 8: REPEATED COUNT MODEL (ROYLE) In collaboration with Heather McKenney
 EXERCISE 8: REPEATED COUNT MODEL (ROYLE) In collaboration with Heather McKenney University of Vermont, Rubenstein School of Environment and Natural Resources Please cite this work as: Donovan, T. M. and
EXERCISE 8: REPEATED COUNT MODEL (ROYLE) In collaboration with Heather McKenney University of Vermont, Rubenstein School of Environment and Natural Resources Please cite this work as: Donovan, T. M. and
Contents. 1 Introduction: what is overdispersion? 2 Recognising (and testing for) overdispersion. 1 Introduction: what is overdispersion?
 Overdispersion, and how to deal with it in R and JAGS (requires R-packages AER, coda, lme4, R2jags, DHARMa/devtools) Carsten F. Dormann 07 December, 2016 Contents 1 Introduction: what is overdispersion?
Overdispersion, and how to deal with it in R and JAGS (requires R-packages AER, coda, lme4, R2jags, DHARMa/devtools) Carsten F. Dormann 07 December, 2016 Contents 1 Introduction: what is overdispersion?
Continuous Covariates in Mark-Recapture-Recovery Analysis: A Comparison. of Methods
 Biometrics 000, 000 000 DOI: 000 000 0000 Continuous Covariates in Mark-Recapture-Recovery Analysis: A Comparison of Methods Simon J. Bonner 1, Byron J. T. Morgan 2, and Ruth King 3 1 Department of Statistics,
Biometrics 000, 000 000 DOI: 000 000 0000 Continuous Covariates in Mark-Recapture-Recovery Analysis: A Comparison of Methods Simon J. Bonner 1, Byron J. T. Morgan 2, and Ruth King 3 1 Department of Statistics,
BUGS Bayesian inference Using Gibbs Sampling
 BUGS Bayesian inference Using Gibbs Sampling Glen DePalma Department of Statistics May 30, 2013 www.stat.purdue.edu/~gdepalma 1 / 20 Bayesian Philosophy I [Pearl] turned Bayesian in 1971, as soon as I
BUGS Bayesian inference Using Gibbs Sampling Glen DePalma Department of Statistics May 30, 2013 www.stat.purdue.edu/~gdepalma 1 / 20 Bayesian Philosophy I [Pearl] turned Bayesian in 1971, as soon as I
Bayesian Mixture Modeling of Significant P Values: A Meta-Analytic Method to Estimate the Degree of Contamination from H 0 : Supplemental Material
 Bayesian Mixture Modeling of Significant P Values: A Meta-Analytic Method to Estimate the Degree of Contamination from H 0 : Supplemental Material Quentin Frederik Gronau 1, Monique Duizer 1, Marjan Bakker
Bayesian Mixture Modeling of Significant P Values: A Meta-Analytic Method to Estimate the Degree of Contamination from H 0 : Supplemental Material Quentin Frederik Gronau 1, Monique Duizer 1, Marjan Bakker
STAT 328 (Statistical Packages)
 Department of Statistics and Operations Research College of Science King Saud University Exercises STAT 328 (Statistical Packages) nashmiah r.alshammari ^-^ Excel and Minitab - 1 - Write the commands of
Department of Statistics and Operations Research College of Science King Saud University Exercises STAT 328 (Statistical Packages) nashmiah r.alshammari ^-^ Excel and Minitab - 1 - Write the commands of
EXAMINATIONS OF THE ROYAL STATISTICAL SOCIETY
 EXAMINATIONS OF THE ROYAL STATISTICAL SOCIETY GRADUATE DIPLOMA, 00 MODULE : Statistical Inference Time Allowed: Three Hours Candidates should answer FIVE questions. All questions carry equal marks. The
EXAMINATIONS OF THE ROYAL STATISTICAL SOCIETY GRADUATE DIPLOMA, 00 MODULE : Statistical Inference Time Allowed: Three Hours Candidates should answer FIVE questions. All questions carry equal marks. The
GROUPED DATA E.G. FOR SAMPLE OF RAW DATA (E.G. 4, 12, 7, 5, MEAN G x / n STANDARD DEVIATION MEDIAN AND QUARTILES STANDARD DEVIATION
 FOR SAMPLE OF RAW DATA (E.G. 4, 1, 7, 5, 11, 6, 9, 7, 11, 5, 4, 7) BE ABLE TO COMPUTE MEAN G / STANDARD DEVIATION MEDIAN AND QUARTILES Σ ( Σ) / 1 GROUPED DATA E.G. AGE FREQ. 0-9 53 10-19 4...... 80-89
FOR SAMPLE OF RAW DATA (E.G. 4, 1, 7, 5, 11, 6, 9, 7, 11, 5, 4, 7) BE ABLE TO COMPUTE MEAN G / STANDARD DEVIATION MEDIAN AND QUARTILES Σ ( Σ) / 1 GROUPED DATA E.G. AGE FREQ. 0-9 53 10-19 4...... 80-89
Comparing estimates of population change from occupancy and mark recapture models for a territorial species
 Comparing estimates of population change from occupancy and mark recapture models for a territorial species Mary M. Conner, 1,3, John J. Keane, 1 Claire V. Gallagher, 1 Thomas E. Munton, 2 and Paula A.
Comparing estimates of population change from occupancy and mark recapture models for a territorial species Mary M. Conner, 1,3, John J. Keane, 1 Claire V. Gallagher, 1 Thomas E. Munton, 2 and Paula A.
APPM 2360 Project 2: Exploring Stage-Structured Population Dynamics with Loggerhead Sea Turtles
 APPM 2360 Project 2: Exploring Stage-Structured Population Dynamics with Loggerhead Sea Turtles Due: March 22, 2018 by 11:59 PM Submit to the Dropbox on D2L as a PDF 1 Introduction In this lab, you will
APPM 2360 Project 2: Exploring Stage-Structured Population Dynamics with Loggerhead Sea Turtles Due: March 22, 2018 by 11:59 PM Submit to the Dropbox on D2L as a PDF 1 Introduction In this lab, you will
Stopover Models. Rachel McCrea. BES/DICE Workshop, Canterbury Collaborative work with
 BES/DICE Workshop, Canterbury 2014 Stopover Models Rachel McCrea Collaborative work with Hannah Worthington, Ruth King, Eleni Matechou Richard Griffiths and Thomas Bregnballe Overview Capture-recapture
BES/DICE Workshop, Canterbury 2014 Stopover Models Rachel McCrea Collaborative work with Hannah Worthington, Ruth King, Eleni Matechou Richard Griffiths and Thomas Bregnballe Overview Capture-recapture
Centre for Biodiversity Dynamics, Department of Biology, NTNU, NO-7491 Trondheim, Norway,
 Animal models and Integrated Nested Laplace Approximations Anna Marie Holand,1, Ingelin Steinsland, Sara Martino,, Henrik Jensen Centre for Biodiversity Dynamics, Department of Biology, NTNU, NO-7491 Trondheim,
Animal models and Integrated Nested Laplace Approximations Anna Marie Holand,1, Ingelin Steinsland, Sara Martino,, Henrik Jensen Centre for Biodiversity Dynamics, Department of Biology, NTNU, NO-7491 Trondheim,
Multilevel Statistical Models: 3 rd edition, 2003 Contents
 Multilevel Statistical Models: 3 rd edition, 2003 Contents Preface Acknowledgements Notation Two and three level models. A general classification notation and diagram Glossary Chapter 1 An introduction
Multilevel Statistical Models: 3 rd edition, 2003 Contents Preface Acknowledgements Notation Two and three level models. A general classification notation and diagram Glossary Chapter 1 An introduction
Mixed models in R using the lme4 package Part 4: Theory of linear mixed models
 Mixed models in R using the lme4 package Part 4: Theory of linear mixed models Douglas Bates Madison January 11, 11 Contents 1 Definition of linear mixed models Definition of linear mixed models As previously
Mixed models in R using the lme4 package Part 4: Theory of linear mixed models Douglas Bates Madison January 11, 11 Contents 1 Definition of linear mixed models Definition of linear mixed models As previously
Supplementary Materials for A Spatial Time-to-Event Approach for Estimating Associations between Air Pollution and Preterm Birth
 Supplementary Materials for A Spatial Time-to-Event Approach for Estimating Associations between Air Pollution and Preterm Birth Howard H. Chang Department of Biostatistics and Bioinformatics Emory University,
Supplementary Materials for A Spatial Time-to-Event Approach for Estimating Associations between Air Pollution and Preterm Birth Howard H. Chang Department of Biostatistics and Bioinformatics Emory University,
SuperMix2 features not available in HLM 7 Contents
 SuperMix2 features not available in HLM 7 Contents Spreadsheet display of.ss3 files... 2 Continuous Outcome Variables: Additional Distributions... 3 Additional Estimation Methods... 5 Count variables including
SuperMix2 features not available in HLM 7 Contents Spreadsheet display of.ss3 files... 2 Continuous Outcome Variables: Additional Distributions... 3 Additional Estimation Methods... 5 Count variables including
Bayes Tutorial using R and JAGS. James Brownlow AIR FORCE TEST CENTER EDWARDS AFB, CA May, 2015
 412TW-PA-15218 Bayes Tutorial using R and JAGS 4 1 2 T W James Brownlow AIR FORCE TEST CENTER EDWARDS AFB, CA 12-14 May, 2015 Approved for public release ; distribution is unlimited. 412TW-PA-15218 AIR
412TW-PA-15218 Bayes Tutorial using R and JAGS 4 1 2 T W James Brownlow AIR FORCE TEST CENTER EDWARDS AFB, CA 12-14 May, 2015 Approved for public release ; distribution is unlimited. 412TW-PA-15218 AIR
Lecture 6. Prior distributions
 Summary Lecture 6. Prior distributions 1. Introduction 2. Bivariate conjugate: normal 3. Non-informative / reference priors Jeffreys priors Location parameters Proportions Counts and rates Scale parameters
Summary Lecture 6. Prior distributions 1. Introduction 2. Bivariate conjugate: normal 3. Non-informative / reference priors Jeffreys priors Location parameters Proportions Counts and rates Scale parameters
Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features. Yangxin Huang
 Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features Yangxin Huang Department of Epidemiology and Biostatistics, COPH, USF, Tampa, FL yhuang@health.usf.edu January
Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features Yangxin Huang Department of Epidemiology and Biostatistics, COPH, USF, Tampa, FL yhuang@health.usf.edu January
Lecture 3. G. Cowan. Lecture 3 page 1. Lectures on Statistical Data Analysis
 Lecture 3 1 Probability (90 min.) Definition, Bayes theorem, probability densities and their properties, catalogue of pdfs, Monte Carlo 2 Statistical tests (90 min.) general concepts, test statistics,
Lecture 3 1 Probability (90 min.) Definition, Bayes theorem, probability densities and their properties, catalogue of pdfs, Monte Carlo 2 Statistical tests (90 min.) general concepts, test statistics,
Metric Predicted Variable With One Nominal Predictor Variable
 Metric Predicted Variable With One Nominal Predictor Variable Tim Frasier Copyright Tim Frasier This work is licensed under the Creative Commons Attribution 4.0 International license. Click here for more
Metric Predicted Variable With One Nominal Predictor Variable Tim Frasier Copyright Tim Frasier This work is licensed under the Creative Commons Attribution 4.0 International license. Click here for more
LONG-TERM CONTRASTED RESPONSES TO CLIMATE OF TWO ANTARCTIC SEABIRD SPECIES
 Ecology, 86(11), 2005, pp. 2889 2903 2005 by the Ecological Society of America LONG-TERM CONTRASTED RESPONSES TO CLIMATE OF TWO ANTARCTIC SEABIRD SPECIES STEPHANIE JENOUVRIER, 1 CHRISTOPHE BARBRAUD, AND
Ecology, 86(11), 2005, pp. 2889 2903 2005 by the Ecological Society of America LONG-TERM CONTRASTED RESPONSES TO CLIMATE OF TWO ANTARCTIC SEABIRD SPECIES STEPHANIE JENOUVRIER, 1 CHRISTOPHE BARBRAUD, AND
Joint Modeling of Longitudinal Item Response Data and Survival
 Joint Modeling of Longitudinal Item Response Data and Survival Jean-Paul Fox University of Twente Department of Research Methodology, Measurement and Data Analysis Faculty of Behavioural Sciences Enschede,
Joint Modeling of Longitudinal Item Response Data and Survival Jean-Paul Fox University of Twente Department of Research Methodology, Measurement and Data Analysis Faculty of Behavioural Sciences Enschede,
Chapter 4 Lecture. Populations with Age and Stage structures. Spring 2013
 Chapter 4 Lecture Populations with Age and Stage structures Spring 2013 4.1 Introduction Life Table- approach to quantify age specific fecundity and survivorship data Age (or Size Class) structured populations
Chapter 4 Lecture Populations with Age and Stage structures Spring 2013 4.1 Introduction Life Table- approach to quantify age specific fecundity and survivorship data Age (or Size Class) structured populations
Online appendix 1: Detailed statistical tests referred to in Wild goose dilemmas by Black, Prop & Larsson
 Online appendix 1: Detailed statistical tests referred to in Wild goose dilemmas by Black, Prop & Larsson Table 1. Variation in length of goslings association (days) with parents in relation to birth year
Online appendix 1: Detailed statistical tests referred to in Wild goose dilemmas by Black, Prop & Larsson Table 1. Variation in length of goslings association (days) with parents in relation to birth year
Eigenvalues and eigenvectors.
 Eigenvalues and eigenvectors. Example. A population of a certain animal can be broken into three classes: eggs, juveniles, and adults. Eggs take one year to become juveniles, and juveniles take one year
Eigenvalues and eigenvectors. Example. A population of a certain animal can be broken into three classes: eggs, juveniles, and adults. Eggs take one year to become juveniles, and juveniles take one year
A Parameter Expansion Approach to Bayesian SEM Estimation
 A Parameter Expansion Approach to Bayesian SEM Estimation Ed Merkle and Yves Rosseel Utrecht University 24 June 2016 Yves Rosseel A Parameter Expansion Approach to Bayesian SEM Estimation 1 / 51 overview
A Parameter Expansion Approach to Bayesian SEM Estimation Ed Merkle and Yves Rosseel Utrecht University 24 June 2016 Yves Rosseel A Parameter Expansion Approach to Bayesian SEM Estimation 1 / 51 overview
Ronald Christensen. University of New Mexico. Albuquerque, New Mexico. Wesley Johnson. University of California, Irvine. Irvine, California
 Texts in Statistical Science Bayesian Ideas and Data Analysis An Introduction for Scientists and Statisticians Ronald Christensen University of New Mexico Albuquerque, New Mexico Wesley Johnson University
Texts in Statistical Science Bayesian Ideas and Data Analysis An Introduction for Scientists and Statisticians Ronald Christensen University of New Mexico Albuquerque, New Mexico Wesley Johnson University
Chapter 4 Multi-factor Treatment Designs with Multiple Error Terms 93
 Contents Preface ix Chapter 1 Introduction 1 1.1 Types of Models That Produce Data 1 1.2 Statistical Models 2 1.3 Fixed and Random Effects 4 1.4 Mixed Models 6 1.5 Typical Studies and the Modeling Issues
Contents Preface ix Chapter 1 Introduction 1 1.1 Types of Models That Produce Data 1 1.2 Statistical Models 2 1.3 Fixed and Random Effects 4 1.4 Mixed Models 6 1.5 Typical Studies and the Modeling Issues
Exercises in Occupancy Estimation and Modeling; Donovan and Hines 2007 EXERCISE 7: ROYLE-NICHOLS ABUNDANCE INDUCED HETEROGENEITY
 EXERCISE 7: ROYLE-NICHOLS ABUNDANCE INDUCED HETEROGENEITY Estimating mean abundance from repeated presence-absence surveys In collaboration with Kurt Rinehart, University of Vermont, Rubenstein School
EXERCISE 7: ROYLE-NICHOLS ABUNDANCE INDUCED HETEROGENEITY Estimating mean abundance from repeated presence-absence surveys In collaboration with Kurt Rinehart, University of Vermont, Rubenstein School
Supplementary Materials for Bayesian Inference for the Causal Effect of Mediation
 Supplementary Materials for Bayesian Inference for the Causal Effect of Mediation M. J. Daniels, J. Roy, C. Kim, J.W. Hogan, and M.G. Perri Step 3. of sampling algorithm: Sampling (F M1,1, F M1,0, F M0,1,
Supplementary Materials for Bayesian Inference for the Causal Effect of Mediation M. J. Daniels, J. Roy, C. Kim, J.W. Hogan, and M.G. Perri Step 3. of sampling algorithm: Sampling (F M1,1, F M1,0, F M0,1,
JULIEN MARTIN, JAMES D. NICHOLS*, WILEY M. KITCHENS and JAMES E. HINES*
 Ecology 2006 75, Multiscale patterns of movement in fragmented landscapes Blackwell Publishing Ltd and consequences on demography of the snail kite in Florida JULIEN MARTIN, JAMES D. NICHOLS*, WILEY M.
Ecology 2006 75, Multiscale patterns of movement in fragmented landscapes Blackwell Publishing Ltd and consequences on demography of the snail kite in Florida JULIEN MARTIN, JAMES D. NICHOLS*, WILEY M.
Stat 587: Key points and formulae Week 15
 Odds ratios to compare two proportions: Difference, p 1 p 2, has issues when applied to many populations Vit. C: P[cold Placebo] = 0.82, P[cold Vit. C] = 0.74, Estimated diff. is 8% What if a year or place
Odds ratios to compare two proportions: Difference, p 1 p 2, has issues when applied to many populations Vit. C: P[cold Placebo] = 0.82, P[cold Vit. C] = 0.74, Estimated diff. is 8% What if a year or place
9 September N-mixture models. Emily Dennis, Byron Morgan and Martin Ridout. The N-mixture model. Data. Model. Multivariate Poisson model
 The Poisson 9 September 2015 Exeter 1 Stats Lab, Cambridge, 1973 The Poisson Exeter 2 The Poisson What the does can estimate animal abundance from a set of counts with both spatial and temporal replication
The Poisson 9 September 2015 Exeter 1 Stats Lab, Cambridge, 1973 The Poisson Exeter 2 The Poisson What the does can estimate animal abundance from a set of counts with both spatial and temporal replication
Bayesian Estimation of Expected Cell Counts by Using R
 Bayesian Estimation of Expected Cell Counts by Using R Haydar Demirhan 1 and Canan Hamurkaroglu 2 Department of Statistics, Hacettepe University, Beytepe, 06800, Ankara, Turkey Abstract In this article,
Bayesian Estimation of Expected Cell Counts by Using R Haydar Demirhan 1 and Canan Hamurkaroglu 2 Department of Statistics, Hacettepe University, Beytepe, 06800, Ankara, Turkey Abstract In this article,
36-463/663: Hierarchical Linear Models
 36-463/663: Hierarchical Linear Models Taste of MCMC / Bayes for 3 or more levels Brian Junker 132E Baker Hall brian@stat.cmu.edu 1 Outline Practical Bayes Mastery Learning Example A brief taste of JAGS
36-463/663: Hierarchical Linear Models Taste of MCMC / Bayes for 3 or more levels Brian Junker 132E Baker Hall brian@stat.cmu.edu 1 Outline Practical Bayes Mastery Learning Example A brief taste of JAGS
Analyzing the genetic structure of populations: a Bayesian approach
 Analyzing the genetic structure of populations: a Bayesian approach Introduction Our review of Nei s G st and Weir and Cockerham s θ illustrated two important principles: 1. It s essential to distinguish
Analyzing the genetic structure of populations: a Bayesian approach Introduction Our review of Nei s G st and Weir and Cockerham s θ illustrated two important principles: 1. It s essential to distinguish
Capture-Recapture Analyses of the Frog Leiopelma pakeka on Motuara Island
 Capture-Recapture Analyses of the Frog Leiopelma pakeka on Motuara Island Shirley Pledger School of Mathematical and Computing Sciences Victoria University of Wellington P.O.Box 600, Wellington, New Zealand
Capture-Recapture Analyses of the Frog Leiopelma pakeka on Motuara Island Shirley Pledger School of Mathematical and Computing Sciences Victoria University of Wellington P.O.Box 600, Wellington, New Zealand
Package SEMModComp. R topics documented: February 19, Type Package Title Model Comparisons for SEM Version 1.0 Date Author Roy Levy
 Type Package Title Model Comparisons for SEM Version 1.0 Date 2009-02-23 Author Roy Levy Package SEMModComp Maintainer Roy Levy February 19, 2015 Conduct tests of difference in fit for
Type Package Title Model Comparisons for SEM Version 1.0 Date 2009-02-23 Author Roy Levy Package SEMModComp Maintainer Roy Levy February 19, 2015 Conduct tests of difference in fit for
Number of Sites Where Spotted Owls Were Detected
 WILDLIFE ECOLOGY TEAM WILDLIFE HABITAT RELATIONSHIPS IN WASHINGTON AND OREGON FY2014 January 27, 2015 Title: Demographic characteristics of spotted owls in the Oregon Coast Ranges, 1990 2014. Principal
WILDLIFE ECOLOGY TEAM WILDLIFE HABITAT RELATIONSHIPS IN WASHINGTON AND OREGON FY2014 January 27, 2015 Title: Demographic characteristics of spotted owls in the Oregon Coast Ranges, 1990 2014. Principal
ECOLOGICAL STATISTICS: Analysis of Capture-Recapture Data
 ECOLOGICAL STATISTICS: Analysis of Capture-Recapture Data Rachel McCrea and Byron Morgan National Centre for Statistical Ecology, University of Kent, Canterbury. Outline 1. Introduction 2. Estimating abundance
ECOLOGICAL STATISTICS: Analysis of Capture-Recapture Data Rachel McCrea and Byron Morgan National Centre for Statistical Ecology, University of Kent, Canterbury. Outline 1. Introduction 2. Estimating abundance
Main Points. Note: Tutorial #1, help session, and due date have all been pushed back 1 class period. Will send Tutorial #1 by end of day.
 1) Demographic PVA -- age- and stage-structured matrices Main Points Pre-reading: Thursday 21 September: Marris Tuesday 26 September: NA Note: Tutorial #1, help session, and due date have all been pushed
1) Demographic PVA -- age- and stage-structured matrices Main Points Pre-reading: Thursday 21 September: Marris Tuesday 26 September: NA Note: Tutorial #1, help session, and due date have all been pushed
The lmm Package. May 9, Description Some improved procedures for linear mixed models
 The lmm Package May 9, 2005 Version 0.3-4 Date 2005-5-9 Title Linear mixed models Author Original by Joseph L. Schafer . Maintainer Jing hua Zhao Description Some improved
The lmm Package May 9, 2005 Version 0.3-4 Date 2005-5-9 Title Linear mixed models Author Original by Joseph L. Schafer . Maintainer Jing hua Zhao Description Some improved
Bayesian Model Diagnostics and Checking
 Earvin Balderama Quantitative Ecology Lab Department of Forestry and Environmental Resources North Carolina State University April 12, 2013 1 / 34 Introduction MCMCMC 2 / 34 Introduction MCMCMC Steps in
Earvin Balderama Quantitative Ecology Lab Department of Forestry and Environmental Resources North Carolina State University April 12, 2013 1 / 34 Introduction MCMCMC 2 / 34 Introduction MCMCMC Steps in
Measures of Agreement
 Measures of Agreement An interesting application is to measure how closely two individuals agree on a series of assessments. A common application for this is to compare the consistency of judgments of
Measures of Agreement An interesting application is to measure how closely two individuals agree on a series of assessments. A common application for this is to compare the consistency of judgments of
Package lmm. R topics documented: March 19, Version 0.4. Date Title Linear mixed models. Author Joseph L. Schafer
 Package lmm March 19, 2012 Version 0.4 Date 2012-3-19 Title Linear mixed models Author Joseph L. Schafer Maintainer Jing hua Zhao Depends R (>= 2.0.0) Description Some
Package lmm March 19, 2012 Version 0.4 Date 2012-3-19 Title Linear mixed models Author Joseph L. Schafer Maintainer Jing hua Zhao Depends R (>= 2.0.0) Description Some
Part 8: GLMs and Hierarchical LMs and GLMs
 Part 8: GLMs and Hierarchical LMs and GLMs 1 Example: Song sparrow reproductive success Arcese et al., (1992) provide data on a sample from a population of 52 female song sparrows studied over the course
Part 8: GLMs and Hierarchical LMs and GLMs 1 Example: Song sparrow reproductive success Arcese et al., (1992) provide data on a sample from a population of 52 female song sparrows studied over the course
Aedes egg laying behavior Erika Mudrak, CSCU November 7, 2018
 Aedes egg laying behavior Erika Mudrak, CSCU November 7, 2018 Introduction The current study investivates whether the mosquito species Aedes albopictus preferentially lays it s eggs in water in containers
Aedes egg laying behavior Erika Mudrak, CSCU November 7, 2018 Introduction The current study investivates whether the mosquito species Aedes albopictus preferentially lays it s eggs in water in containers
Markov Chain Monte Carlo (MCMC) and Model Evaluation. August 15, 2017
 Markov Chain Monte Carlo (MCMC) and Model Evaluation August 15, 2017 Frequentist Linking Frequentist and Bayesian Statistics How can we estimate model parameters and what does it imply? Want to find the
Markov Chain Monte Carlo (MCMC) and Model Evaluation August 15, 2017 Frequentist Linking Frequentist and Bayesian Statistics How can we estimate model parameters and what does it imply? Want to find the
GIS and conservation biology: A case study with animal movement data
 GIS and conservation biology: A case study with animal movement data BIOL 037: Conservation Genetics, week of April 22 2013 In this lab exercise we will explore bird migration data and how it can be used
GIS and conservation biology: A case study with animal movement data BIOL 037: Conservation Genetics, week of April 22 2013 In this lab exercise we will explore bird migration data and how it can be used
 Weakness of Beta priors (or conjugate priors in general) They can only represent a limited range of prior beliefs. For example... There are no bimodal beta distributions (except when the modes are at 0
Weakness of Beta priors (or conjugate priors in general) They can only represent a limited range of prior beliefs. For example... There are no bimodal beta distributions (except when the modes are at 0
CHAPTER 15. The robust design Decomposing the probability of subsequent encounter
 CHAPTER 15 The robust design William Kendall, USGS Colorado Cooperative Fish & Wildlife Research Unit Changes in population size through time are a function of births, deaths, immigration, and emigration.
CHAPTER 15 The robust design William Kendall, USGS Colorado Cooperative Fish & Wildlife Research Unit Changes in population size through time are a function of births, deaths, immigration, and emigration.
Measures of Association for I J tables based on Pearson's 2 Φ 2 = Note that I 2 = I where = n J i=1 j=1 J i=1 j=1 I i=1 j=1 (ß ij ß i+ ß +j ) 2 ß i+ ß
 Correlation Coefficient Y = 0 Y = 1 = 0 ß11 ß12 = 1 ß21 ß22 Product moment correlation coefficient: ρ = Corr(; Y ) E() = ß 2+ = ß 21 + ß 22 = E(Y ) E()E(Y ) q V ()V (Y ) E(Y ) = ß 2+ = ß 21 + ß 22 = ß
Correlation Coefficient Y = 0 Y = 1 = 0 ß11 ß12 = 1 ß21 ß22 Product moment correlation coefficient: ρ = Corr(; Y ) E() = ß 2+ = ß 21 + ß 22 = E(Y ) E()E(Y ) q V ()V (Y ) E(Y ) = ß 2+ = ß 21 + ß 22 = ß
Analysis of 2005 and 2006 Wolverine DNA Mark-Recapture Sampling at Daring Lake, Ekati, Diavik, and Kennady Lake, Northwest Territories
 Analysis of 2005 and 2006 Wolverine DNA Mark-Recapture Sampling at Daring Lake, Ekati, Diavik, and Kennady Lake, Northwest Territories John Boulanger, Integrated Ecological Research, 924 Innes St. Nelson
Analysis of 2005 and 2006 Wolverine DNA Mark-Recapture Sampling at Daring Lake, Ekati, Diavik, and Kennady Lake, Northwest Territories John Boulanger, Integrated Ecological Research, 924 Innes St. Nelson
A generalized integrated population model to estimate greater sage- grouse population dynamics
 A generalized integrated population model to estimate greater sage- grouse population dynamics Rebecca McCaffery 1, and Paul M. Lukacs Wildlife Biology Program, Department of Ecosystem and Conservation
A generalized integrated population model to estimate greater sage- grouse population dynamics Rebecca McCaffery 1, and Paul M. Lukacs Wildlife Biology Program, Department of Ecosystem and Conservation
Outline. Binomial, Multinomial, Normal, Beta, Dirichlet. Posterior mean, MAP, credible interval, posterior distribution
 Outline A short review on Bayesian analysis. Binomial, Multinomial, Normal, Beta, Dirichlet Posterior mean, MAP, credible interval, posterior distribution Gibbs sampling Revisit the Gaussian mixture model
Outline A short review on Bayesian analysis. Binomial, Multinomial, Normal, Beta, Dirichlet Posterior mean, MAP, credible interval, posterior distribution Gibbs sampling Revisit the Gaussian mixture model
CONSERVATION PLANNING FOR
 CHAPTER 26 Biotic Interactions and Global Change. 1993. Edited by Peter M. Kareiva Joel G. Kingsolver Raymond B. Huey Sinauer Assoc. Inc., Sunderland, Massachusetts CONSERVATION PLANNING FOR SPECIES OCCUPYING
CHAPTER 26 Biotic Interactions and Global Change. 1993. Edited by Peter M. Kareiva Joel G. Kingsolver Raymond B. Huey Sinauer Assoc. Inc., Sunderland, Massachusetts CONSERVATION PLANNING FOR SPECIES OCCUPYING
Lec 10: Bayesian Statistics for Genetics Imputation and software
 Lec 10: Bayesian Statistics for Genetics Imputation and software Ken Rice Swiss Institute in Statistical Genetics September, 2017 Overview In this final session: Overview of imputation where Bayesian (or
Lec 10: Bayesian Statistics for Genetics Imputation and software Ken Rice Swiss Institute in Statistical Genetics September, 2017 Overview In this final session: Overview of imputation where Bayesian (or
Exam C Solutions Spring 2005
 Exam C Solutions Spring 005 Question # The CDF is F( x) = 4 ( + x) Observation (x) F(x) compare to: Maximum difference 0. 0.58 0, 0. 0.58 0.7 0.880 0., 0.4 0.680 0.9 0.93 0.4, 0.6 0.53. 0.949 0.6, 0.8
Exam C Solutions Spring 005 Question # The CDF is F( x) = 4 ( + x) Observation (x) F(x) compare to: Maximum difference 0. 0.58 0, 0. 0.58 0.7 0.880 0., 0.4 0.680 0.9 0.93 0.4, 0.6 0.53. 0.949 0.6, 0.8
A CONDITIONALLY-UNBIASED ESTIMATOR OF POPULATION SIZE BASED ON PLANT-CAPTURE IN CONTINUOUS TIME. I.B.J. Goudie and J. Ashbridge
 This is an electronic version of an article published in Communications in Statistics Theory and Methods, 1532-415X, Volume 29, Issue 11, 2000, Pages 2605-2619. The published article is available online
This is an electronic version of an article published in Communications in Statistics Theory and Methods, 1532-415X, Volume 29, Issue 11, 2000, Pages 2605-2619. The published article is available online
Anne Trainor Matt Simon Geog 593 Final Project Report December 8, 2006
 Anne Trainor Matt Simon Geog 593 Final Project Report December 8, 2006 INTRODUCTION Habitat fragmentation has led to an increasingly patchy landscape. This has forced land managers to adopt a metapopulation
Anne Trainor Matt Simon Geog 593 Final Project Report December 8, 2006 INTRODUCTION Habitat fragmentation has led to an increasingly patchy landscape. This has forced land managers to adopt a metapopulation
