movehmm An R package for the analysis of animal movement data
|
|
|
- Meghan Wheeler
- 5 years ago
- Views:
Transcription
1 movehmm An R package for the analysis of animal movement data Michelot T., Langrock R., and Patterson T. June 3, 2018 Contents 1 Introduction 2 2 A quick overview of the HMM approach to modelling animal movement 3 3 Example application Movement data Load and format the tracking data Use prepdata Use plot.movedata Fitting the model fithmm Dealing with numerical instability Confidence intervals Further inference tools and visualization of the model Plot the model State decoding Stationary state probabilities Model selection with AIC Model checking Use plotsat Dealing with one-dimensional data Package features Model options Distributions Zero-inflation Covariates Stationarity Main functions prepdata fithmm Generic methods Other operations on movehmm simdata plotsat
2 1 Introduction The analysis of animal movement data has become increasingly important in terrestrial and marine ecology. Improvements in telemetry technology have resulted in an explosion in the volume of high precision data being collected. As a result, there are two challenges which researchers collecting these data regularly face: (1) data volume and (2) employing statistical methods which can accommodate some of the specific features of movement data (Patterson et al., 2009). A substantial part of the literature on statistical modelling of animal movement data has focused on the intuitive approach of decomposing movement time series into distinct behavioural modes (a.k.a. bouts, states), via the use of so-called state-switching models. These approaches typically involve assuming movements of animals to be driven by stochastically evolving states, such as a slow moving state, which may be indicative of resting or foraging, versus faster movement states which might indicate transits between foraging patches. Associated with changes in movement speeds are changes to the distribution of directional changes in the movement (known as the turning angle see further description below). Bayesian methods which employ MCMC approaches have become very popular tools for the analysis of movement data using state-switching models (e.g. Jonsen et al., 2005; Morales et al., 2004). Typically, these have been implemented using WinBUGS(although see McClintock et al., 2012). While these models are relatively straightforward to build and hence fit in WinBUGS, the estimation can be painfully slow due to slow mixing of the MCMC samplers. However, for an important subset of movement data, namely highly accurate position data (e.g. from GPS) and more generally all time series of locations where the measurement error is negligible relative to the scale of the movement the task of statistical classification of behaviour can be done much more efficiently using hidden Markov models(hmms) and associated frequentist inferential tools. HMMs are increasingly popular in this field, due to their flexibility and to the associated very efficient recursive algorithms available for conducting statistical inference (Patterson et al., 2009; Langrock et al., 2012). The crucial requirements on movement data in order for HMMs to be suitable are that measurement error in positions is negligible and that there is a regular sampling unit (e.g. one positional observation per hour, or per dive, or any other meaningful unit). movehmm is an R package which implements HMMs and associated tools for state decoding, model selection etc. specifically tailored to animal movement modelling. Particular attention was paid to computational efficiency with the fitting algorithm implemented in C++. The high computational speed makes it feasible to analyze very large data sets e.g. tens of thousands of positions collected for each of a dozen individual animals on standard desktop PCs. The package also allows users to incorporate covariate data into their models, which is particularly useful when inferring the drivers of changes in behaviour. Our hope is that the movehmm package will provide users who collect movement data with an interface to sophisticated and adequate methods for a statistical analysis of their data. The package is structured so as to allow the users to prepare their data for analysis, fit a variety of HMMs to their data and perform diagnostics on these fitted models. The package is presented in Michelot et al. (2016), where its use is illustrated on simulated movement data of wild haggises. In this vignette, we briefly introduce HMMs in the context of animal movement. We then provide a detailed example of a typical use of the package (preprocessing movement data, fitting an HMM to the data, and analyzing the fitted model). Finally, we describe more technically the structure of the package and its main functions. 2
3 2 A quick overview of the HMM approach to modelling animal movement For full details of the HMM approach in the context of animal movement modelling, the user should refer to the relevant primary publications (see, e.g., Patterson et al., 2009, Langrock et al., 2012, Zucchini et al., 2016). Here, we briefly highlight the essential features of the HMM approach. The standard HMM approach to model an individual animal s movement considers bivariate time series comprising the step length and the turning angle at each time point (see illustration in Figure 1). The associated locations need to be sampled at equally spaced points in time (though missing data on an otherwise regular grid can easily be handled) and are assumed to be observed with no or only negligible error. The movehmm package is restricted to such discrete time data and involves the assumption that the locations are observed with zero, or at least negligible error. (x t+2,y t+2 ) (x t,y t ) (x t,y t ) φ t (x t+2,y t+2 ) l t 1 l t l t+1 (x t+1,y t+1 ) (x t+1,y t+1 ) φ t+1 (x t 1,y t 1 ) (x t 1,y t 1 ) Figure 1: Illustration of step lengths and turning angles At each time point, the parameters of the step length distribution (e.g., a gamma distribution) and the parameters of the turning angle distribution (e.g., a von Mises distribution) are determined by an underlying unobserved state. There are finitely many states which provide rough classifications of the movement (e.g. more active vs. less active), often interpreted as proxies for the animal s behavioural states (e.g. transiting vs. foraging). The sequence of states is assumed to be generated by a Markov chain, usually with a tendency of remaining in a state for some time before switching to another state. The corresponding state transition probabilities, as well as the parameters characterising the statedependent distributions, are model parameters to be estimated. The number of states (movement modes) is unknown and has to be specified by the user. Typically with movement data one assumes a low number of states (say 4). For example, an animal may be foraging (low speed, high rates of turning) and transiting (high speeds, low rates of turning). Biologically interesting inference often involves modelling the state transition probabilities as functions of environmental covariates. For a given set of model parameters, the likelihood of the data can be calculated using a recursive algorithm (the forward algorithm, cf. Patterson et al., 2016), which in a very effective way considers all possible state sequences that might have given rise to the observed time series. This makes numerical maximization of the (log-)likelihood, and hence maximum likelihood estimation, feasible in most cases. Having estimated the model parameters (and examined useful diagnostics of model fit), the user can estimate the most likely sequence of behavioural states. We encourage the user to consult a good primary text such as Zucchini et al. (2016) to familiarize themselves with the technical details of hidden Markov models. 3 Example application Before we provide a detailed description of the various features of the movehmm package in the subsequent section, we illustrate a typical HMM-based analysis of movement data using the main functions of the package, via an example. We use the data from Morales et al. (2004), collected on four elk in Canada. 3
4 3.1 Movement data The input data need to have the correct format for subsequent processing and analysis. The data need to be provided as a data.frame, with two mandatory columns: Easting or longitude (default name: x) Northing or latitude (default name: y) It is possible to have a column ID, which contains the identifiers of the observed animals. If no column named ID is provided, all the observations will be considered to belong to a single animal. Additional columns are considered as covariates. Note that, within this package, covariates need to have numerical values (rather than e.g. character values) Load and format the tracking data The elk data considered in Morales et al. (2004) are loaded with the package, as the data frame elk data. The data frame has four columns: ID, Easting, Northing, and dist water. The last one is the distance of the animal to water, which for illustration purposes we want to include in the model as a covariate. head(elk_data) ID Easting Northing dist_water 1 elk elk elk elk elk elk The easting and northing values are expressed in meters in the data, and we decide that we want to deal with distances in kilometers for the step lengths. To achieve this, we transform the coordinates into meters. elk_data$easting <- elk_data$easting/1000 elk_data$northing <- elk_data$northing/1000 As a result, this is what the data look like: head(elk_data) ID Easting Northing dist_water 1 elk elk elk elk elk elk Use prepdata The data are in the proper format, and can be processed using prepdata to compute step lengths and angles. We choose the arguments carefully: 4
5 type specifies whether the coordinates are easting/northing (type="utm") or longitude/latitude (type="ll") values. The latter is the default, so we need to call the function with the argument type="utm", to indicate that UTM coordinates are provided. coordnames are the names of the coordinates in the input data frame. The default is x and y, so we need to call the function with the argument coordnames=c("easting","northing"). The call to the function for these data is, data <- prepdata(elk_data,type="utm",coordnames=c("easting","northing")) The step lengths and turning angles are computed, and the returned object is a data frame. head(data) ID step angle x y dist_water 1 elk NA elk elk elk elk elk Note that the coordinates have been renamed x and y. This makes the processing of the data simpler. If the data set contains covariates which have missing values, then those are imputed using the closest non-missing value, by default the previous one if it is available. This is arbitrary and might not be appropriate in all situations. It is also possible to print summary information about the data, using the function summary, e.g. summary(data) Movement data for 4 tracks: elk observations elk observations elk observations elk observations Covariate(s): dist_water Min. 25% Median Mean 75% Max Use plot.movedata Once the data have been preprocessed, they can be plotted using the generic function plot. This displays maps of the animals tracks, times series of the steps and angles, and histograms of the steps and angles. A few plotting options are available. These are described in the documentation. To plot all animals tracks on a single map, we call: plot(data,compact=t) The resulting map, and the steps and angles graphs for the first animal, are displayed in Figure 2 (we omit the graphs for the three other animals, which are also displayed when using the above command). The time series of step lengths is one way to check the data for outliers. 5
6 Animal ID: elk 115 y step length Frequency t turning angle (radians) Frequency π π 2 0 π 2 π t π π 2 0 π 2 π x step length turning angle (radians) Figure 2: Map of the animals tracks (left) each color represents an animal. Time series and histograms of the step lengths and turning angles for one individual, elk-115 (right). 3.2 Fitting the model fithmm The function fithmm is used to fit an HMM to the data. Its arguments are described in the documentation. Here are a few choices we make: nbstates=2, i.e. we fit a 2-state HMM to the data; beta0=null and delta0=null, i.e. we use the default values for the initial values beta0 and delta0; formula= dist water, i.e. the transition probabilities are functions of the covariate dist water ; stepdist="gamma", to model the step lengths with the gamma distribution (note that it is the default, so we do not need to explicitely specify it); angledist="vm", to model the turning angles with the von Mises distribution (default); anglemean=null, because we want to estimate the mean of the angle distribution (default); stationary=false, as due to the covariates the process is not stationary (default). We also need to specify initial values for the parameters of the state-dependent distributions, to be used by the optimization function. Note that this choice is crucial, and that the algorithm might not find the global optimum of the likelihood function if the initial parameters are poorly chosen. The initial parameters should be specified in two vectors, steppar0 (for the step distribution) and anglepar0 (for the angle distribution). The necessary parameters of each distribution are detailed in Section Zero-inflation (as described in Section 4.1.2) must be included in the step length distribution if some steps are of length exactly zero (which is the case for the elk data). To do so, another parameter is added to the step distribution: its mass on zero. Here, the initial values are chosen such that they correspond to the commonly observed pattern in two-state HMMs for animal movement data, with state 1 involving relatively short steps and many 6
7 turnings (hence the choice of a small initial value for the mean of the gamma step length distribution and an initial value of pi for the mean turning angle) and state 2 involving longer steps and fewer turnings (hence the choice of a larger initial value for the mean of the gamma step length distribution and an initial value of 0 for the mean turning angle). For numerical stability, we decide to standardize the covariate values before fitting the model (explanations in Section 3.2.2). standardize covariate values data$dist_water <- (data$dist_water-mean(data$dist_water))/sd(data$dist_water) initial parameters for gamma and von Mises distributions mu0 <- c(0.1,1) # step mean (two parameters: one for each state) sigma0 <- c(0.1,1) # step SD zeromass0 <- c(0.1,0.05) # step zero-mass steppar0 <- c(mu0,sigma0,zeromass0) anglemean0 <- c(pi,0) # angle mean kappa0 <- c(1,1) # angle concentration anglepar0 <- c(anglemean0,kappa0) call to fitting function m <- fithmm(data=data,nbstates=2,steppar0=steppar0, anglepar0=anglepar0,formula=~dist_water) The returned object, m, is of the class movehmm. It can be printed in order to obtain the maximum likelihood estimates of all model parameters. m Value of the maximum log-likelihood: Step length parameters: state 1 state 2 mean e+00 sd e+00 zero-mass e-09 Turning angle parameters: state 1 state 2 mean concentration Regression coeffs for the transition probabilities: > 2 2 -> 1 intercept dist_water Initial distribution: [1] The argument knownstates of the function fithmm makes it possible to set some values of the state process to fixed values, prior to fitting the model. This can be useful e.g. when the animal s behaviour 7
8 is known for some time points, but we discourage users to take advantage of this option to make the states match their expectations (instead of letting the data speak for themselves) Dealing with numerical instability As mentioned above, the numerical maximization routine might not identify the global maximum of the likelihood function, or even fail to converge altogether, for poorly chosen initial values of the parameters. In such a case, the optimization routine nlm might produce an error such as: Error in nlm(nloglike, wpar, nbstates, bounds, parsize, data, stepdist, : non-finite value supplied by 'nlm' The best way to deal with such numerical problems is to test different sets of initial values, possibly chosen randomly. By comparing the resulting estimates for the different initial values used, one usually obtains a good feeling for any potential sensitivity of the numerical search to its chosen starting point. Note, however, that in any case there will usually be no certainty that the global maximum of the likelihood, i.e. the maximum likelihood estimate, has been identified. During preliminary tests on the elk data, we noticed that, in this example, the numerical search is highly sensitive to the choice of the initial parameters beta0. This is due to the high values of the covariate: a small change in the associated regression coefficients can make a big difference in the likelihood function. In such cases, it is advisable to standardize the covariate values before fitting the model, for example by calling: data$dist_water <- (data$dist_water-mean(data$dist_water))/sd(data$dist_water) This allows for greater numerical stability, with the convergence of the fitting function depending less on the choice of initial values. The value of the maximum log-likelihood is not affected by the standardization of the covariate values, only the maximum likelihood estimate of beta is Confidence intervals Confidence intervals for the model parameters can be computed with the function CI, passing as an argument the object created by fithmm. It is possible to give the significance level of the desired confidence interval as an argument, e.g for 99% confidence intervals. By default, 95% confidence intervals are returned. Below we show the 95% confidence intervals for the parameters of the 2-state model fitted to the elk data. CI(m)$stepPar corresponds to the bounds of the confidence intervals for the step parameters (CI(m)$stepPar$lower and CI(m)$stepPar$upper). CI(m)$anglePar and CI(m)$beta are respectively the bounds of the confidence intervals for the angle parameters and the regression coefficients of the transition probabilities. CI(m) $steppar $steppar$lower state 1 state 2 mean sd zero-mass NA $steppar$upper state 1 state 2 mean sd
9 zero-mass NA $anglepar $anglepar$lower state 1 state 2 mean concentration $anglepar$upper state 1 state 2 mean concentration $beta $beta$lower 1 -> 2 2 -> 1 intercept dist_water $beta$upper 1 -> 2 2 -> 1 intercept dist_water Note that a warning message is also output: Warning message: In CI(m) : Some of the parameter estimates seem to lie close to the boundaries of their parameter space. The associated CIs are probably unreliable (or might not be computable). It here refers to the zero-inflation parameter in the second state. Its estimate is very close to zero, the inferior boundary of its range (this parameter is in the interval [0,1]), and this causes the corresponding confidence interval to be unreliable. The function can sometimes fail to compute such a confidence interval, and returns NA instead, as in this example. 3.3 Further inference tools and visualization of the model Various options are available for the class movehmm, and here we explain how to use them in the elk example Plot the model The fitted model can be plotted, using the generic function plot. A few graphical options are available and listed in the documentation. Here, we call: plot(m, plotci=true) This outputs: an histogram of step lengths of all animals, with the fitted state-dependent densities, an histogram of turning angles of all animals, with the fitted state-dependent densities, plots of the transition probabilities as functions of the covariate considered, 9
10 a map of each animal s track, colored by states. Figure 3 displays those plots, but showing only one of the plotted maps, namely the one corresponding to the first animal, elk-115. The state-dependent densities are weighted by the relative frequency of each state in the most probable state sequence (decoded with the Viterbi algorithm, see Section 3.3.2). For example, if according to the most probable state sequence, one third of the observations is allocated to the first state, and two thirds to the second state, the plots of the densities in the first state are weighted with a factor 1/3, and in the second state with a factor 2/3. All animals All animals Density State 1 State 2 Total Density State 1 State 2 Total π π 2 0 π 2 π step length turning angle (radians) Transition probabilities Animal ID: elk > 1 2 > dist_water > 2 2 > dist_water y dist_water dist_water x Figure 3: Output of plot.movehmm. Histogram of step lengths with fitted distributions (top-left), histogram of turning angles with fitted distributions (top-right), transition probabilities as functions of dist water with 95% confidence intervals (bottom-left), and map of decoded track for the first animal (bottom-right). The first state (in orange on the plots) corresponds to short steps, and angles centered around π, and the second state (in blue on the plots) corresponds to longer steps, and angles centered around 0. 10
11 The plots of the transition probabilities as function of the covariate indicate that animals tend to switch from the second state to the first state when they are far from water, whereas they stay in the second state when closer to water State decoding Two functions can be used to decode the state process. Viterbi algorithm To globally decode the state process, the Viterbi algorithm is implemented in the function viterbi. This function outputs the most likely sequence of states to have generated the observation, under the fitted model. Below are the most probable states for the first 25 observations of the first individual: states <- viterbi(m) states[1:25] [1] State probabilities To get more accurate information on the state process, it is possible to compute the state probabilities for each observation, using stateprobs. This returns a matrix with as many columns as there are states in the model, and as many rows as there are observations (stacking all animals observations). The elements of the matrix are defined as where {S t } is the state process. For example: sp <- stateprobs(m) head(sp) [,1] [,2] [1,] e [2,] e [3,] e [4,] e [5,] e [6,] e stateprobs(m)[t,j] = Pr(S t = j) The state with highest probability according to stateprobs might not be the same as the state in the most probable sequence returned by the Viterbi algorithm. This is because the Viterbi algorithm performs global decoding, whereas the state probabilities are local decoding. For more details, see Zucchini et al. (2016). The function plotstates can be used to visualize the results of viterbi and stateprobs. Figure 4 shows the plots of the most likely state sequence decoded by the Viterbi algorithm, as well as both columns of the matrix of state probabilities, for one individual, elk-115. It was obtained with the following command: plotstates(m,animals="elk-115") 11
12 Animal ID: elk 115 State Pr(State=1) Observation index Pr(State=2) Observation index Figure 4: Decoded states sequence (top row), and state probabilities of observations (middle and bottom rows) for elk Stationary state probabilities For a transition probability matrix Γ, the stationary distribution is the vector δ that solves the equation δ = δγ, subject to N i=1 δ i = 1 (see Section for more information). It reflects the long-term proportion of time the model spends in each state. When the transition probabilities are time-varying (i.e. functions of covariates), the stationary distribution does not exist. However, for fixed values of the covariates, we can obtain one transition probability matrix, and thus one stationary distribution. The function plotstationary does this over a grid of values of each covariate, and plots the resulting stationary state probabilites. They can be interpreted as the long-term probabilities of being in each state at different values of the covariate. plotstationary(m, plotci=true) Model selection with AIC The generic method AIC is available to compare movehmm models. For example, we now fit a 3-state HMM to the data, and want to compare the AICs of the 2-state and 3-state models. # initial parameters mu0 <- c(0.1,0.5,3) sigma0 <- c(0.05,0.5,1) zeromass0 <- c(0.05,0.0001,0.0001) steppar0 <- c(mu0,sigma0,zeromass0) anglemean0 <- c(pi,pi,0) 12
13 Stationary state probabilities State 1 State dist_water Figure 5: Output of plotstationary. Stationary state probabilities, as functions of the distance to water, with 95% confidence intervals. 13
14 kappa0 <- c(1,1,1) anglepar0 <- c(anglemean0,kappa0) # fit the 3-state model m3 <- fithmm(data=data,nbstates=3,steppar0=steppar0, anglepar0=anglepar0,formula=~dist_water) And, to compare them: AIC(m,m3) Model AIC 1 m m In terms of AIC, the 3-state model is favoured over the 2-state model in this example Model checking The pseudo-residuals (a.k.a. quantile residuals) of the model can be computed with pseudores. These follow a standard normal distribution if the fitted model is the true data-generating process. In other words, a deviation from normality indicates a lack of fit. For more theoretical background on pseudoresiduals, see Zucchini et al. (2016). The pseudo-residuals of the 2-state model fitted to the elk data are displayed in Figure 6. They can be computed and plotted with the following commands. # compute the pseudo-residuals pr <- pseudores(m) # time series, qq-plots, and ACF of the pseudo-residuals plotpr(m) If some steps are of length zero (i.e. if the step distribution is zero-inflated), the corresponding pseudo-residuals are plotted as segments on the qq-plot. It is the case for the smallest step pseudoresidual located in the bottom-left corner of the step qq-plot in Figure 6. The pseudo-residuals of discrete data are defined as segments, and in this case, the segments start in. 3.4 Use plotsat The function plotsat plots the tracking data on a satellite image, with the help of the package ggmap (Kahle and Wickham, 2013). Il only works with longitude and latitude values, so we need to convert the UTM coordinates first. We can do this e.g. with the help of the packages sp (Pebesma and Bivand, 2005) and rgdal (Bivand et al., 2016) (more information can be found in the documentations of those packages). library(rgdal) utmcoord <- SpatialPoints(cbind(data$x*1000,data$y*1000), proj4string=crs("+proj=utm +zone=17")) llcoord <- sptransform(utmcoord,crs("+proj=longlat")) lldata <- data.frame(id=data$id,x=attr(llcoord,"coords")[,1], y=attr(llcoord,"coords")[,2]) In the code above, we need to multiply the UTM coordinates by 1000, as we had divided them by 1000 earlier to work with distances in kilometres. In the function SpatialPoints, we indicate +zone=17, because the data come from the UTM zone
15 Steps pseudo residuals Observation index Steps pseudo residuals Angles pseudo residuals Observation index Angles pseudo residuals Theoretical Quantiles Sample Quantiles Theoretical Quantiles Sample Quantiles Lag ACF Lag ACF Figure 6: Time series, qq-plots, and autocorrelation functions of the pseudo-residuals of the 2-state model. The data frame lldata contains the converted longitude and latitude coordinates of the observations. We can plot it with plotsat; the result is shown in Figure 7. As the satellite image is fetched from Google, an Internet connection is required to use plotsat. plotsat(lldata,zoom=8) 3.5 Dealing with one-dimensional data It is sometimes of interest to model one-dimensional movement data, like e.g. dive data. This can be done in movehmm by setting all values of the second coordinate to zero in the data, and use the option angledist="none". In the case where one-dimensional data are provided, the plotting functions for the data and for the model will output plots of the first coordinate as a function of time, instead of a map of the track. 15
16 45.5 lat ID elk 115 elk 163 elk 287 elk lon Figure 7: Plot of the elk data, obtained with plotsat. 4 Package features In this section, we describe the global structure of the package, and then describe in more detail the main functions required to fit an HMM to movement data. The package is articulated in terms of two S3 classes: movedata and movehmm. The first extends the native R data frame, essentially gathering time series of the movement metrics of interest, namely the step lengths and turning angles, as well as the covariate values. A movehmm object is a fitted model, which stores in particular the values of the MLE of the parameters. To create a movedata object, the function prepdata is called on the tracking data (track points coordinates). Then, the function fithmm is called on the movedata, and returns a movehmm. Both classes can be used through their methods (e.g. plot.movedata, AIC.moveHMM), and a variety of other functions can be called on movehmm objects. All functions are described in more detail in Section 4.2 and their use is explained on an example in Section 3. Figure 8 illustrates the links between the main components of the package. Note we will occasionally omit the class to which a method belongs, if the context makes it clear. 16
17 Figure 8: Structure of the main components of the package. The blue boxes are S3 classes, and the green boxes are functions. The arrows indicate input and output of data. Besides, as illustrated in the example in Section 3, it is not necessary to specify the class when calling the R function (e.g. calling plot on a movehmm object automatically refers to plot.movehmm). 4.1 Model options Distributions Here is the list of distributions included, with the names they have in the package. Step length: gamma( gamma ), Weibull( weibull ), exponential( exp ) and log-normal( lnorm ). Turning angle: von Mises ( vm ) and wrapped-cauchy ( wrpcauchy ). It is also possible to specify angledist="none", if the angles are not modelled. The parameters depend on the distribution used. The gamma distribution expects the mean and standard deviation, and all other distributions expect the same parameters as the corresponding R density function, i.e. Distribution Parameters gamma mean (0, + ) standard deviation (0, + ) Weibull shape (0, + ) scale (0, + ) log-normal location R scale (0, + ) exponential rate (0, + ) von Mises mean ( π, π] concentration (0, + ) wrapped Cauchy mean ( π, π] concentration (0, 1) For the gamma distribution, the link between the mean/standard deviation (expected by fithmm) and shape/rate (expected by dgamma) is given by: shape = mean2 SD 2, mean rate = SD 2 17
18 4.1.2 Zero-inflation If some steps are exactly equal to zero, then strictly positive distributions such as the gamma are inadequate. In such cases, zero-inflated distributions can be considered. A zero-inflated step length distribution simply assumes that there is a probability z of observing a 0 and a probability of 1 z of observing a positive value distributed according to a standard positive distribution (e.g. a gamma). Within the package movehmm, zero-inflation will automatically be included if there are zero steps. In that case the (state-dependent) values z will be estimated, with the remaining positive distribution, weighted by 1 z, specified as one of the available standard step length distributions, listed in Section Covariates In practice it is often of interest to model the state transition probabilities as functions of time-varying covariates. This can( be done ) by assuming the Markov chain to be time-varying, with transition probability matrix Γ (t) = γ (t) ij, linking the transition probabilities to the covariate(s) via the multinomial logit link. In the general case of N states, γ (t) ij = Pr ( S t = j S t 1 = i ) exp(η ij ) = N k=1 exp(η ik), where η ij = { β (ij) 0 + p l=1 β(ij) l w lt if i j, 0 otherwise, for i,j = 1,...,N. Here {S t } is the state process, w lt is the l-th covariate at time t and p is the number of covariates considered. The β parameters directly affect the off-diagonal elements in Γ (t) with an increase in the linear predictor η ij resulting in an increase in γ (t) ij and hence also the diagonal entries due to the row constraints (with the entries in each row summing to one). Note in particular that we have to fix η ii = 0 for all i since otherwise the model would be overparameterized (not identifiable). Within movehmm, the β coefficients for the off-diagonal transition probabilities are stored in an (p+1) (N (N 1)) matrix. For example, for a 3-state HMM with two covariates, the matrix beta is β (12) 0 β (13) 0 β (21) 0 β (23) 0 β (31) 0 β (32) 0 β (12) 1 β (13) 1 β (21) 1 β (23) 1 β (31) 1 β (32) 1 β (12) 2 β (13) 2 β (21) 2 β (23) 2 β (31) 2 β (32) 2 Here the first row corresponds to the intercept terms and the other two rows to the slope coefficients associated with the two covariates. There are as many columns as there are off-diagonal entries in the 3 3 transition probability matrix, and that matrix is filled row-wise (i.e. column 1 in beta is linked to γ (t) 12, column 2 is linked to γ(t) 13, column 3 is linked to γ(t) 21, etc.). In practice, many movement models involve only two states, in which case the above equations boil 18
19 down to ( 1+exp Γ (t) = ( exp 1+exp 1 β (12) 0 + ) p l=1 β(12) l w lt β (21) 0 + ) p l=1 β(21) l w lt ( β (21) 0 + ) p l=1 β(21) l w lt ( exp β (12) 0 + ) p l=1 β(12) l w lt ( 1+exp β (12) 0 + ) p l=1 β(12) l w lt 1 ( 1+exp β (21) 0 + ) p l=1 β(21) l w lt 1 logit 1( β (12) 0 + ) p l=1 β(12) l w lt = logit 1( β (21) 0 + ) p l=1 β(21) l w lt logit 1( β (12) 0 + ) p l=1 β(12) l w lt 1 logit 1( β (21) 0 + ) p l=1 β(21) l w lt The inverse logit link function is applied in order to map the real-valued predictor onto the interval [0, 1] (with the above multinomial logit link representing a generalization of this approach to the case of N > 2 states). In the case of two states, the matrix β in movehmm is structured as follows: Stationarity β (12) 0 β (21) 0 β (12) 1 β (21) 1.. β p (12) β p (21) The function fithmm includes the option of fitting a stationary model(using the option stationary=true, with the default being stationary=false). This is only possible if no covariates are incorporated into the model. (Otherwise the transition probabilities will be time-dependent, such that the Markov chain is non-homogeneous and in particular cannot be stationary.) When no covariates are considered and the option stationary=true is selected, then the initial state distribution of the Markov chain will automatically be chosen as the stationary distribution (a.k.a. steady-state distribution) implied by the estimated transition probability matrix (as opposed to being estimated when stationary=false). This stationary distribution is the vector δ that solves the equation δ = δγ subject to N i=1 δ i = 1. In practice, this solution almost always exists. 4.2 Main functions prepdata Tracking data usually consist of time series of either easting-northing coordinates or longitude-latitude values. However, with the HMM approach the derived quantities step lengths and turning angles are modelled. The function prepdata computes the steps and angles from the coordinates. As input, this function takes an R data frame with columns x (either easting or longitude) and y (either northing or latitude). If the names of the coordinates columns are not x and y, then the argument coordnames should specify them. If several animals were observed, there should also be a column ID which identifies the animal being observed. If there is no ID column, all observations will be considered to be associated with a single animal. All additional columns are considered as covariates. In addition to the data frame, prepdata takes an argument type, which can either be LL (longitude-latitude, the default) or UTM. The former indicates that the coordinates are longitude-latitude values, and the latter that they are easting-northing values.. 19
20 To compute the step lengths, prepdata calls the function spdistn1 from the package sp. The step lengths are in the unit of the input if easting/northing are provided, and in kilometres if longitude/latitude are provided. prepdata outputs a data frame, with the same columns as the input, plus columns step and angle. This object is of the class movedata, and can be plotted using the generic function plot fithmm Using the function fithmm, an HMM can be fitted to an object of class movedata, via numerical maximum likelihood. The list of the arguments of fithmm is detailed in the documentation. The maximum likelihood estimation is carried out using the R function nlm. This function outputs a list of information about the model. Most elements of that list are only meant to be used by the movehmm functions (see Sections and 4.2.4), but a few can be informative per se: mle contains the estimates of the parameters of the model; mod contains the output of the optimization function nlm, including mod$minimum (minimum of the negative log-likelihood) and mod$hessian, the Hessian of the negative log-likelihood function at its minimum Generic methods Methods (i.e. class functions) are available for both movedata and movehmm objects, to operate on them. For details on the options see the documentation, and for an example of their use, see Section 3. plot.movedata plots a few graphs to illustrate the data: a map of each animal s track, time series of the steps and angles, histograms of the steps and angles. summary.movedata outputs some summary information about a movedata object: the number of animals, the number of observations for each animal, and quantiles of the covariate values. plot.movehmm plots a few graphs to illustrate the fitted model: a map of each animal s track, colored by states, plots of the estimated density functions, plots of the transition probabilities as functions of the covariates. print.movehmm prints the value of the maximum log-likelihood, and the maximum likelihood estimates of the parameters of the model. AIC.moveHMM returns the AIC of one or several fitted models Other operations on movehmm Other functions can be called on a movehmm object, for further analysis. CI computes confidence intervals for the step length distribution parameters, for the turning angle distribution parameters, and for the regression coefficients of the transition probabilities. pseudores computes the pseudo-residuals of the model. These can be used to assess the goodness of fit. If the model is the true data-generating process, then the pseudo-residuals follow a standard normal distribution (Zucchini et al., 2016). stateprobs computes the probabilities of the underlying Markov chain being in the different states, at each observation, under the fitted model. 20
21 viterbi computes the sequence of most probable states, under the fitted model, using the Viterbi algorithm (Zucchini et al., 2016). plotstates plots the most probable state sequence (as decoded with viterbi), and the state probabilities (as computed with stateprobs). plotpr plots time series, qq-plots, and the sample autocorrelation(acf) functions of the pseudoresiduals of the fitted model (Zucchini et al., 2016). The qq-plots can be used to visually assess whether or not the pseudo-residuals are standard normally distributed. The points in the qq-plot will be close to the straight line if the model fits the data well. If the sample ACFs display a residual autocorrelation, then this is an indication that the model might not have captured all relevant correlation structure in the data simdata The function simdata simulates movement data from an HMM, given its parameters. The returned object is of the class movedata, and can be visualized using plot, or fitted using fithmm. The arguments of simdata are detailed in the documentation. It is possible to call simdata on a fitted model directly, to simulate data from it. This can be used to assess the fit, by checking that the simulated data display the same features as the real data plotsat Based on functions provided by the package ggmap(kahle and Wickham, 2013), plotsat plots tracking data on a satellite image retrieved from Google. Note that it only works with longitude and latitude values, and not with easting and northing coordinates. It can be used on any data frame with fields x and y, e.g. typically a movedata object. It is necessary to specify manually the zoom level wanted, with the argument zoom, which can take values in {3,4,5,...,20,21}. Other arguments are optional, and are described in the documentation. 21
22 References Bivand, R., Keitt, T., and Rowlingson, B. (2016) rgdal: Bindings for the Geospatial Data Abstraction Library, R package version 1.1-8, URL: Jonsen, I.D., Flemming, J.M., Myers, R.A. (2005), Robust state-space modeling of animal movement data, Ecology, 86 (11), Kahle, D., Wickham, H. (2013) ggmap: Spatial Visualization with ggplot2, The R Journal, 5 (1), URL: Langrock R., King R., Matthiopoulos J., Thomas L., Fortin D., Morales J.M. (2012), Flexible and practical modeling of animal telemetry data: hidden Markov models and extensions Ecology, 93 (11), McClintock, B.T., King, R., Thomas, L., Matthiopoulos, J., McConnell, B.J., Morales, J.M. (2012), A general discrete-time modeling framework for animal movement using multistate random walks, Ecological Monographs, 82 (3), Michelot, T., Langrock, R., Patterson, T.A. (2016), movehmm: an R package for analysing animal movement data using hidden Markov models, Methods in Ecology and Evolution, 7 (11), Morales, J.M., Haydon, D.T., Frair, J., Holsinger, K.E., Fryxell, J.M. (2004), Extracting more out of relocation data: building movement models as mixtures of random walks, Ecology, 85 (9), Patterson T.A., Basson M., Bravington M.V., Gunn J.S. (2009), Classifying movement behaviour in relation to environmental conditions using hidden Markov models, Journal of Animal Ecology, 78 (6), Patterson T.A., Parton, A., Langrock, R., Blackwell, P.G., Thomas, L., King, R. (2016), Statistical modelling of animal movement: a myopic review and a discussion of good practice, arxiv preprint, arxiv: Pebesma, E.J., Bivand, R.S. (2005) Classes and methods for spatial data in R, R News, 5 (2), URL: Zucchini, W. MacDonald, I.L., Langrock, R. (2016). Hidden Markov Models for Time Series: An Introduction Using R, Second Edition. Chapman & Hall/CRC Press, Boca Raton, FL. 22
Ecological applications of hidden Markov models and related doubly stochastic processes
 . Ecological applications of hidden Markov models and related doubly stochastic processes Roland Langrock School of Mathematics and Statistics & CREEM Motivating example HMM machinery Some ecological applications
. Ecological applications of hidden Markov models and related doubly stochastic processes Roland Langrock School of Mathematics and Statistics & CREEM Motivating example HMM machinery Some ecological applications
HMM Workshop. Rocío Joo. Rocío Joo HMM Workshop 1 / 43
 HMM Workshop Rocío Joo Rocío Joo HMM Workshop 1 / 43 Structure of the workshop Introduction to HMMs Data for application 3 HMM applications Simple HMM with 3 states HMM with 4 states with constraints in
HMM Workshop Rocío Joo Rocío Joo HMM Workshop 1 / 43 Structure of the workshop Introduction to HMMs Data for application 3 HMM applications Simple HMM with 3 states HMM with 4 states with constraints in
Nonparametric inference in hidden Markov and related models
 Nonparametric inference in hidden Markov and related models Roland Langrock, Bielefeld University Roland Langrock Bielefeld University 1 / 47 Introduction and motivation Roland Langrock Bielefeld University
Nonparametric inference in hidden Markov and related models Roland Langrock, Bielefeld University Roland Langrock Bielefeld University 1 / 47 Introduction and motivation Roland Langrock Bielefeld University
momentuhmm: R package for generalized hidden Markov models of animal movement
 Received: 26 January 2018 Accepted: 16 February 2018 DOI: 10.1111/2041-210X.12995 APPLICATION momentuhmm: R package for generalized hidden Markov models of animal movement Brett T. McClintock 1 Théo Michelot
Received: 26 January 2018 Accepted: 16 February 2018 DOI: 10.1111/2041-210X.12995 APPLICATION momentuhmm: R package for generalized hidden Markov models of animal movement Brett T. McClintock 1 Théo Michelot
momentuhmm: R package for analysis of telemetry data using generalized multivariate hidden Markov models of animal movement
 momentuhmm: R package for analysis of telemetry data using generalized multivariate hidden Markov models of animal movement Brett T. McClintock 1 and Théo Michelot 2 1 Marine Mammal Laboratory Alaska Fisheries
momentuhmm: R package for analysis of telemetry data using generalized multivariate hidden Markov models of animal movement Brett T. McClintock 1 and Théo Michelot 2 1 Marine Mammal Laboratory Alaska Fisheries
Estimation by direct maximization of the likelihood
 CHAPTER 3 Estimation by direct maximization of the likelihood 3.1 Introduction We saw in Equation (2.12) that the likelihood of an HMM is given by ( L T =Pr X (T ) = x (T )) = δp(x 1 )ΓP(x 2 ) ΓP(x T )1,
CHAPTER 3 Estimation by direct maximization of the likelihood 3.1 Introduction We saw in Equation (2.12) that the likelihood of an HMM is given by ( L T =Pr X (T ) = x (T )) = δp(x 1 )ΓP(x 2 ) ΓP(x T )1,
401 Review. 6. Power analysis for one/two-sample hypothesis tests and for correlation analysis.
 401 Review Major topics of the course 1. Univariate analysis 2. Bivariate analysis 3. Simple linear regression 4. Linear algebra 5. Multiple regression analysis Major analysis methods 1. Graphical analysis
401 Review Major topics of the course 1. Univariate analysis 2. Bivariate analysis 3. Simple linear regression 4. Linear algebra 5. Multiple regression analysis Major analysis methods 1. Graphical analysis
Model selection and checking
 CHAPTER 6 Model selection and checking In the basic HMM with m states, increasing m always improves the fit of the model (as judged by the likelihood). But along with the improvement comes a quadratic
CHAPTER 6 Model selection and checking In the basic HMM with m states, increasing m always improves the fit of the model (as judged by the likelihood). But along with the improvement comes a quadratic
Bayesian Methods for Machine Learning
 Bayesian Methods for Machine Learning CS 584: Big Data Analytics Material adapted from Radford Neal s tutorial (http://ftp.cs.utoronto.ca/pub/radford/bayes-tut.pdf), Zoubin Ghahramni (http://hunch.net/~coms-4771/zoubin_ghahramani_bayesian_learning.pdf),
Bayesian Methods for Machine Learning CS 584: Big Data Analytics Material adapted from Radford Neal s tutorial (http://ftp.cs.utoronto.ca/pub/radford/bayes-tut.pdf), Zoubin Ghahramni (http://hunch.net/~coms-4771/zoubin_ghahramani_bayesian_learning.pdf),
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 11 Project
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 11 Project
STA 414/2104: Machine Learning
 STA 414/2104: Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistics! rsalakhu@cs.toronto.edu! http://www.cs.toronto.edu/~rsalakhu/ Lecture 9 Sequential Data So far
STA 414/2104: Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistics! rsalakhu@cs.toronto.edu! http://www.cs.toronto.edu/~rsalakhu/ Lecture 9 Sequential Data So far
PACKAGE LMest FOR LATENT MARKOV ANALYSIS
 PACKAGE LMest FOR LATENT MARKOV ANALYSIS OF LONGITUDINAL CATEGORICAL DATA Francesco Bartolucci 1, Silvia Pandofi 1, and Fulvia Pennoni 2 1 Department of Economics, University of Perugia (e-mail: francesco.bartolucci@unipg.it,
PACKAGE LMest FOR LATENT MARKOV ANALYSIS OF LONGITUDINAL CATEGORICAL DATA Francesco Bartolucci 1, Silvia Pandofi 1, and Fulvia Pennoni 2 1 Department of Economics, University of Perugia (e-mail: francesco.bartolucci@unipg.it,
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 3 Linear
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 3 Linear
Markov Chain Monte Carlo methods
 Markov Chain Monte Carlo methods By Oleg Makhnin 1 Introduction a b c M = d e f g h i 0 f(x)dx 1.1 Motivation 1.1.1 Just here Supresses numbering 1.1.2 After this 1.2 Literature 2 Method 2.1 New math As
Markov Chain Monte Carlo methods By Oleg Makhnin 1 Introduction a b c M = d e f g h i 0 f(x)dx 1.1 Motivation 1.1.1 Just here Supresses numbering 1.1.2 After this 1.2 Literature 2 Method 2.1 New math As
COMPUTER SESSION: ARMA PROCESSES
 UPPSALA UNIVERSITY Department of Mathematics Jesper Rydén Stationary Stochastic Processes 1MS025 Autumn 2010 COMPUTER SESSION: ARMA PROCESSES 1 Introduction In this computer session, we work within the
UPPSALA UNIVERSITY Department of Mathematics Jesper Rydén Stationary Stochastic Processes 1MS025 Autumn 2010 COMPUTER SESSION: ARMA PROCESSES 1 Introduction In this computer session, we work within the
Estimating the distribution of demersal fishing effort from VMS data using hidden Markov models
 Estimating the distribution of demersal fishing effort from VMS data using hidden Markov models D.L. Borchers Centre for Research into Ecological and Environmental Modelling, The Observatory, Buchanan
Estimating the distribution of demersal fishing effort from VMS data using hidden Markov models D.L. Borchers Centre for Research into Ecological and Environmental Modelling, The Observatory, Buchanan
Part 6: Multivariate Normal and Linear Models
 Part 6: Multivariate Normal and Linear Models 1 Multiple measurements Up until now all of our statistical models have been univariate models models for a single measurement on each member of a sample of
Part 6: Multivariate Normal and Linear Models 1 Multiple measurements Up until now all of our statistical models have been univariate models models for a single measurement on each member of a sample of
M. H. Gonçalves 1 M. S. Cabral 2 A. Azzalini 3
 M. H. Gonçalves 1 M. S. Cabral 2 A. Azzalini 3 1 University of Algarve, Portugal 2 University of Lisbon, Portugal 3 Università di Padova, Italy user!2010, July 21-23 1 2 3 4 5 What is bild? an R. parametric
M. H. Gonçalves 1 M. S. Cabral 2 A. Azzalini 3 1 University of Algarve, Portugal 2 University of Lisbon, Portugal 3 Università di Padova, Italy user!2010, July 21-23 1 2 3 4 5 What is bild? an R. parametric
Introduction to Bayesian Statistics and Markov Chain Monte Carlo Estimation. EPSY 905: Multivariate Analysis Spring 2016 Lecture #10: April 6, 2016
 Introduction to Bayesian Statistics and Markov Chain Monte Carlo Estimation EPSY 905: Multivariate Analysis Spring 2016 Lecture #10: April 6, 2016 EPSY 905: Intro to Bayesian and MCMC Today s Class An
Introduction to Bayesian Statistics and Markov Chain Monte Carlo Estimation EPSY 905: Multivariate Analysis Spring 2016 Lecture #10: April 6, 2016 EPSY 905: Intro to Bayesian and MCMC Today s Class An
Fitting Narrow Emission Lines in X-ray Spectra
 Outline Fitting Narrow Emission Lines in X-ray Spectra Taeyoung Park Department of Statistics, University of Pittsburgh October 11, 2007 Outline of Presentation Outline This talk has three components:
Outline Fitting Narrow Emission Lines in X-ray Spectra Taeyoung Park Department of Statistics, University of Pittsburgh October 11, 2007 Outline of Presentation Outline This talk has three components:
Modelling Non-linear and Non-stationary Time Series
 Modelling Non-linear and Non-stationary Time Series Chapter 7(extra): (Generalized) Hidden Markov Models Henrik Madsen Lecture Notes September 2016 Henrik Madsen (02427 Adv. TS Analysis) Lecture Notes
Modelling Non-linear and Non-stationary Time Series Chapter 7(extra): (Generalized) Hidden Markov Models Henrik Madsen Lecture Notes September 2016 Henrik Madsen (02427 Adv. TS Analysis) Lecture Notes
Joint work with Nottingham colleagues Simon Preston and Michail Tsagris.
 /pgf/stepx/.initial=1cm, /pgf/stepy/.initial=1cm, /pgf/step/.code=1/pgf/stepx/.expanded=- 10.95415pt,/pgf/stepy/.expanded=- 10.95415pt, /pgf/step/.value required /pgf/images/width/.estore in= /pgf/images/height/.estore
/pgf/stepx/.initial=1cm, /pgf/stepy/.initial=1cm, /pgf/step/.code=1/pgf/stepx/.expanded=- 10.95415pt,/pgf/stepy/.expanded=- 10.95415pt, /pgf/step/.value required /pgf/images/width/.estore in= /pgf/images/height/.estore
4 : Exact Inference: Variable Elimination
 10-708: Probabilistic Graphical Models 10-708, Spring 2014 4 : Exact Inference: Variable Elimination Lecturer: Eric P. ing Scribes: Soumya Batra, Pradeep Dasigi, Manzil Zaheer 1 Probabilistic Inference
10-708: Probabilistic Graphical Models 10-708, Spring 2014 4 : Exact Inference: Variable Elimination Lecturer: Eric P. ing Scribes: Soumya Batra, Pradeep Dasigi, Manzil Zaheer 1 Probabilistic Inference
Introduction to mtm: An R Package for Marginalized Transition Models
 Introduction to mtm: An R Package for Marginalized Transition Models Bryan A. Comstock and Patrick J. Heagerty Department of Biostatistics University of Washington 1 Introduction Marginalized transition
Introduction to mtm: An R Package for Marginalized Transition Models Bryan A. Comstock and Patrick J. Heagerty Department of Biostatistics University of Washington 1 Introduction Marginalized transition
Bayesian Inference for Regression Parameters
 Bayesian Inference for Regression Parameters 1 Bayesian inference for simple linear regression parameters follows the usual pattern for all Bayesian analyses: 1. Form a prior distribution over all unknown
Bayesian Inference for Regression Parameters 1 Bayesian inference for simple linear regression parameters follows the usual pattern for all Bayesian analyses: 1. Form a prior distribution over all unknown
Introduction to cthmm (Continuous-time hidden Markov models) package
 Introduction to cthmm (Continuous-time hidden Markov models) package Abstract A disease process refers to a patient s traversal over time through a disease with multiple discrete states. Multistate models
Introduction to cthmm (Continuous-time hidden Markov models) package Abstract A disease process refers to a patient s traversal over time through a disease with multiple discrete states. Multistate models
COMS 4771 Probabilistic Reasoning via Graphical Models. Nakul Verma
 COMS 4771 Probabilistic Reasoning via Graphical Models Nakul Verma Last time Dimensionality Reduction Linear vs non-linear Dimensionality Reduction Principal Component Analysis (PCA) Non-linear methods
COMS 4771 Probabilistic Reasoning via Graphical Models Nakul Verma Last time Dimensionality Reduction Linear vs non-linear Dimensionality Reduction Principal Component Analysis (PCA) Non-linear methods
Math 350: An exploration of HMMs through doodles.
 Math 350: An exploration of HMMs through doodles. Joshua Little (407673) 19 December 2012 1 Background 1.1 Hidden Markov models. Markov chains (MCs) work well for modelling discrete-time processes, or
Math 350: An exploration of HMMs through doodles. Joshua Little (407673) 19 December 2012 1 Background 1.1 Hidden Markov models. Markov chains (MCs) work well for modelling discrete-time processes, or
CS Homework 3. October 15, 2009
 CS 294 - Homework 3 October 15, 2009 If you have questions, contact Alexandre Bouchard (bouchard@cs.berkeley.edu) for part 1 and Alex Simma (asimma@eecs.berkeley.edu) for part 2. Also check the class website
CS 294 - Homework 3 October 15, 2009 If you have questions, contact Alexandre Bouchard (bouchard@cs.berkeley.edu) for part 1 and Alex Simma (asimma@eecs.berkeley.edu) for part 2. Also check the class website
Machine Learning Overview
 Machine Learning Overview Sargur N. Srihari University at Buffalo, State University of New York USA 1 Outline 1. What is Machine Learning (ML)? 2. Types of Information Processing Problems Solved 1. Regression
Machine Learning Overview Sargur N. Srihari University at Buffalo, State University of New York USA 1 Outline 1. What is Machine Learning (ML)? 2. Types of Information Processing Problems Solved 1. Regression
A REVIEW AND APPLICATION OF HIDDEN MARKOV MODELS AND DOUBLE CHAIN MARKOV MODELS
 A REVIEW AND APPLICATION OF HIDDEN MARKOV MODELS AND DOUBLE CHAIN MARKOV MODELS Michael Ryan Hoff A Dissertation submitted to the Faculty of Science, University of the Witwatersrand, Johannesburg, in fulfilment
A REVIEW AND APPLICATION OF HIDDEN MARKOV MODELS AND DOUBLE CHAIN MARKOV MODELS Michael Ryan Hoff A Dissertation submitted to the Faculty of Science, University of the Witwatersrand, Johannesburg, in fulfilment
Chapter 1 Statistical Inference
 Chapter 1 Statistical Inference causal inference To infer causality, you need a randomized experiment (or a huge observational study and lots of outside information). inference to populations Generalizations
Chapter 1 Statistical Inference causal inference To infer causality, you need a randomized experiment (or a huge observational study and lots of outside information). inference to populations Generalizations
Computer Vision Group Prof. Daniel Cremers. 10a. Markov Chain Monte Carlo
 Group Prof. Daniel Cremers 10a. Markov Chain Monte Carlo Markov Chain Monte Carlo In high-dimensional spaces, rejection sampling and importance sampling are very inefficient An alternative is Markov Chain
Group Prof. Daniel Cremers 10a. Markov Chain Monte Carlo Markov Chain Monte Carlo In high-dimensional spaces, rejection sampling and importance sampling are very inefficient An alternative is Markov Chain
Package OUwie. August 29, 2013
 Package OUwie August 29, 2013 Version 1.34 Date 2013-5-21 Title Analysis of evolutionary rates in an OU framework Author Jeremy M. Beaulieu , Brian O Meara Maintainer
Package OUwie August 29, 2013 Version 1.34 Date 2013-5-21 Title Analysis of evolutionary rates in an OU framework Author Jeremy M. Beaulieu , Brian O Meara Maintainer
Statistics Toolbox 6. Apply statistical algorithms and probability models
 Statistics Toolbox 6 Apply statistical algorithms and probability models Statistics Toolbox provides engineers, scientists, researchers, financial analysts, and statisticians with a comprehensive set of
Statistics Toolbox 6 Apply statistical algorithms and probability models Statistics Toolbox provides engineers, scientists, researchers, financial analysts, and statisticians with a comprehensive set of
A hidden semi-markov model for the occurrences of water pipe bursts
 A hidden semi-markov model for the occurrences of water pipe bursts T. Economou 1, T.C. Bailey 1 and Z. Kapelan 1 1 School of Engineering, Computer Science and Mathematics, University of Exeter, Harrison
A hidden semi-markov model for the occurrences of water pipe bursts T. Economou 1, T.C. Bailey 1 and Z. Kapelan 1 1 School of Engineering, Computer Science and Mathematics, University of Exeter, Harrison
University of Cambridge. MPhil in Computer Speech Text & Internet Technology. Module: Speech Processing II. Lecture 2: Hidden Markov Models I
 University of Cambridge MPhil in Computer Speech Text & Internet Technology Module: Speech Processing II Lecture 2: Hidden Markov Models I o o o o o 1 2 3 4 T 1 b 2 () a 12 2 a 3 a 4 5 34 a 23 b () b ()
University of Cambridge MPhil in Computer Speech Text & Internet Technology Module: Speech Processing II Lecture 2: Hidden Markov Models I o o o o o 1 2 3 4 T 1 b 2 () a 12 2 a 3 a 4 5 34 a 23 b () b ()
Hidden Markov Models. Hal Daumé III. Computer Science University of Maryland CS 421: Introduction to Artificial Intelligence 19 Apr 2012
 Hidden Markov Models Hal Daumé III Computer Science University of Maryland me@hal3.name CS 421: Introduction to Artificial Intelligence 19 Apr 2012 Many slides courtesy of Dan Klein, Stuart Russell, or
Hidden Markov Models Hal Daumé III Computer Science University of Maryland me@hal3.name CS 421: Introduction to Artificial Intelligence 19 Apr 2012 Many slides courtesy of Dan Klein, Stuart Russell, or
Model checking overview. Checking & Selecting GAMs. Residual checking. Distribution checking
 Model checking overview Checking & Selecting GAMs Simon Wood Mathematical Sciences, University of Bath, U.K. Since a GAM is just a penalized GLM, residual plots should be checked exactly as for a GLM.
Model checking overview Checking & Selecting GAMs Simon Wood Mathematical Sciences, University of Bath, U.K. Since a GAM is just a penalized GLM, residual plots should be checked exactly as for a GLM.
Why Bayesian approaches? The average height of a rare plant
 Why Bayesian approaches? The average height of a rare plant Estimation and comparison of averages is an important step in many ecological analyses and demographic models. In this demonstration you will
Why Bayesian approaches? The average height of a rare plant Estimation and comparison of averages is an important step in many ecological analyses and demographic models. In this demonstration you will
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistical Sciences! rsalakhu@cs.toronto.edu! h0p://www.cs.utoronto.ca/~rsalakhu/ Lecture 7 Approximate
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistical Sciences! rsalakhu@cs.toronto.edu! h0p://www.cs.utoronto.ca/~rsalakhu/ Lecture 7 Approximate
36-463/663Multilevel and Hierarchical Models
 36-463/663Multilevel and Hierarchical Models From Bayes to MCMC to MLMs Brian Junker 132E Baker Hall brian@stat.cmu.edu 1 Outline Bayesian Statistics and MCMC Distribution of Skill Mastery in a Population
36-463/663Multilevel and Hierarchical Models From Bayes to MCMC to MLMs Brian Junker 132E Baker Hall brian@stat.cmu.edu 1 Outline Bayesian Statistics and MCMC Distribution of Skill Mastery in a Population
Lecture 21: Spectral Learning for Graphical Models
 10-708: Probabilistic Graphical Models 10-708, Spring 2016 Lecture 21: Spectral Learning for Graphical Models Lecturer: Eric P. Xing Scribes: Maruan Al-Shedivat, Wei-Cheng Chang, Frederick Liu 1 Motivation
10-708: Probabilistic Graphical Models 10-708, Spring 2016 Lecture 21: Spectral Learning for Graphical Models Lecturer: Eric P. Xing Scribes: Maruan Al-Shedivat, Wei-Cheng Chang, Frederick Liu 1 Motivation
UNIVERSITY of PENNSYLVANIA CIS 520: Machine Learning Final, Fall 2013
 UNIVERSITY of PENNSYLVANIA CIS 520: Machine Learning Final, Fall 2013 Exam policy: This exam allows two one-page, two-sided cheat sheets; No other materials. Time: 2 hours. Be sure to write your name and
UNIVERSITY of PENNSYLVANIA CIS 520: Machine Learning Final, Fall 2013 Exam policy: This exam allows two one-page, two-sided cheat sheets; No other materials. Time: 2 hours. Be sure to write your name and
Chapter 3 - Estimation by direct maximization of the likelihood
 Chapter 3 - Estimation by direct maximization of the likelihood 02433 - Hidden Markov Models Martin Wæver Pedersen, Henrik Madsen Course week 3 MWP, compiled June 7, 2011 Recall: Recursive scheme for the
Chapter 3 - Estimation by direct maximization of the likelihood 02433 - Hidden Markov Models Martin Wæver Pedersen, Henrik Madsen Course week 3 MWP, compiled June 7, 2011 Recall: Recursive scheme for the
Sequence Modelling with Features: Linear-Chain Conditional Random Fields. COMP-599 Oct 6, 2015
 Sequence Modelling with Features: Linear-Chain Conditional Random Fields COMP-599 Oct 6, 2015 Announcement A2 is out. Due Oct 20 at 1pm. 2 Outline Hidden Markov models: shortcomings Generative vs. discriminative
Sequence Modelling with Features: Linear-Chain Conditional Random Fields COMP-599 Oct 6, 2015 Announcement A2 is out. Due Oct 20 at 1pm. 2 Outline Hidden Markov models: shortcomings Generative vs. discriminative
Bayesian Mixture Modeling of Significant P Values: A Meta-Analytic Method to Estimate the Degree of Contamination from H 0 : Supplemental Material
 Bayesian Mixture Modeling of Significant P Values: A Meta-Analytic Method to Estimate the Degree of Contamination from H 0 : Supplemental Material Quentin Frederik Gronau 1, Monique Duizer 1, Marjan Bakker
Bayesian Mixture Modeling of Significant P Values: A Meta-Analytic Method to Estimate the Degree of Contamination from H 0 : Supplemental Material Quentin Frederik Gronau 1, Monique Duizer 1, Marjan Bakker
Some general observations.
 Modeling and analyzing data from computer experiments. Some general observations. 1. For simplicity, I assume that all factors (inputs) x1, x2,, xd are quantitative. 2. Because the code always produces
Modeling and analyzing data from computer experiments. Some general observations. 1. For simplicity, I assume that all factors (inputs) x1, x2,, xd are quantitative. 2. Because the code always produces
STAT 425: Introduction to Bayesian Analysis
 STAT 425: Introduction to Bayesian Analysis Marina Vannucci Rice University, USA Fall 2017 Marina Vannucci (Rice University, USA) Bayesian Analysis (Part 2) Fall 2017 1 / 19 Part 2: Markov chain Monte
STAT 425: Introduction to Bayesian Analysis Marina Vannucci Rice University, USA Fall 2017 Marina Vannucci (Rice University, USA) Bayesian Analysis (Part 2) Fall 2017 1 / 19 Part 2: Markov chain Monte
Stat 5101 Lecture Notes
 Stat 5101 Lecture Notes Charles J. Geyer Copyright 1998, 1999, 2000, 2001 by Charles J. Geyer May 7, 2001 ii Stat 5101 (Geyer) Course Notes Contents 1 Random Variables and Change of Variables 1 1.1 Random
Stat 5101 Lecture Notes Charles J. Geyer Copyright 1998, 1999, 2000, 2001 by Charles J. Geyer May 7, 2001 ii Stat 5101 (Geyer) Course Notes Contents 1 Random Variables and Change of Variables 1 1.1 Random
Community Health Needs Assessment through Spatial Regression Modeling
 Community Health Needs Assessment through Spatial Regression Modeling Glen D. Johnson, PhD CUNY School of Public Health glen.johnson@lehman.cuny.edu Objectives: Assess community needs with respect to particular
Community Health Needs Assessment through Spatial Regression Modeling Glen D. Johnson, PhD CUNY School of Public Health glen.johnson@lehman.cuny.edu Objectives: Assess community needs with respect to particular
Linear Regression. Data Model. β, σ 2. Process Model. ,V β. ,s 2. s 1. Parameter Model
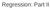 Regression: Part II Linear Regression y~n X, 2 X Y Data Model β, σ 2 Process Model Β 0,V β s 1,s 2 Parameter Model Assumptions of Linear Model Homoskedasticity No error in X variables Error in Y variables
Regression: Part II Linear Regression y~n X, 2 X Y Data Model β, σ 2 Process Model Β 0,V β s 1,s 2 Parameter Model Assumptions of Linear Model Homoskedasticity No error in X variables Error in Y variables
Maximum likelihood estimation of mark-recapture-recovery models in the presence of continuous covariates
 Maximum likelihood estimation of mark-recapture-recovery models in the presence of continuous covariates & Ruth King University of St Andrews........ 1 Outline of the problem 2 MRR data in the absence
Maximum likelihood estimation of mark-recapture-recovery models in the presence of continuous covariates & Ruth King University of St Andrews........ 1 Outline of the problem 2 MRR data in the absence
CHAPTER 9 EXAMPLES: MULTILEVEL MODELING WITH COMPLEX SURVEY DATA
 Examples: Multilevel Modeling With Complex Survey Data CHAPTER 9 EXAMPLES: MULTILEVEL MODELING WITH COMPLEX SURVEY DATA Complex survey data refers to data obtained by stratification, cluster sampling and/or
Examples: Multilevel Modeling With Complex Survey Data CHAPTER 9 EXAMPLES: MULTILEVEL MODELING WITH COMPLEX SURVEY DATA Complex survey data refers to data obtained by stratification, cluster sampling and/or
STAT 518 Intro Student Presentation
 STAT 518 Intro Student Presentation Wen Wei Loh April 11, 2013 Title of paper Radford M. Neal [1999] Bayesian Statistics, 6: 475-501, 1999 What the paper is about Regression and Classification Flexible
STAT 518 Intro Student Presentation Wen Wei Loh April 11, 2013 Title of paper Radford M. Neal [1999] Bayesian Statistics, 6: 475-501, 1999 What the paper is about Regression and Classification Flexible
LAB 2 - ONE DIMENSIONAL MOTION
 Name Date Partners L02-1 LAB 2 - ONE DIMENSIONAL MOTION OBJECTIVES Slow and steady wins the race. Aesop s fable: The Hare and the Tortoise To learn how to use a motion detector and gain more familiarity
Name Date Partners L02-1 LAB 2 - ONE DIMENSIONAL MOTION OBJECTIVES Slow and steady wins the race. Aesop s fable: The Hare and the Tortoise To learn how to use a motion detector and gain more familiarity
Hidden Markov models: definition and properties
 CHAPTER Hidden Markov models: definition and properties. A simple hidden Markov model Consider again the observed earthquake series displayed in Figure. on p.. The observations are unbounded counts, making
CHAPTER Hidden Markov models: definition and properties. A simple hidden Markov model Consider again the observed earthquake series displayed in Figure. on p.. The observations are unbounded counts, making
Ch3. TRENDS. Time Series Analysis
 3.1 Deterministic Versus Stochastic Trends The simulated random walk in Exhibit 2.1 shows a upward trend. However, it is caused by a strong correlation between the series at nearby time points. The true
3.1 Deterministic Versus Stochastic Trends The simulated random walk in Exhibit 2.1 shows a upward trend. However, it is caused by a strong correlation between the series at nearby time points. The true
Unobserved Heterogeneity and the Statistical Analysis of Highway Accident Data. Fred Mannering University of South Florida
 Unobserved Heterogeneity and the Statistical Analysis of Highway Accident Data Fred Mannering University of South Florida Highway Accidents Cost the lives of 1.25 million people per year Leading cause
Unobserved Heterogeneity and the Statistical Analysis of Highway Accident Data Fred Mannering University of South Florida Highway Accidents Cost the lives of 1.25 million people per year Leading cause
Ronald Christensen. University of New Mexico. Albuquerque, New Mexico. Wesley Johnson. University of California, Irvine. Irvine, California
 Texts in Statistical Science Bayesian Ideas and Data Analysis An Introduction for Scientists and Statisticians Ronald Christensen University of New Mexico Albuquerque, New Mexico Wesley Johnson University
Texts in Statistical Science Bayesian Ideas and Data Analysis An Introduction for Scientists and Statisticians Ronald Christensen University of New Mexico Albuquerque, New Mexico Wesley Johnson University
Approximate Inference
 Approximate Inference Simulation has a name: sampling Sampling is a hot topic in machine learning, and it s really simple Basic idea: Draw N samples from a sampling distribution S Compute an approximate
Approximate Inference Simulation has a name: sampling Sampling is a hot topic in machine learning, and it s really simple Basic idea: Draw N samples from a sampling distribution S Compute an approximate
Hidden Markov Models Part 1: Introduction
 Hidden Markov Models Part 1: Introduction CSE 6363 Machine Learning Vassilis Athitsos Computer Science and Engineering Department University of Texas at Arlington 1 Modeling Sequential Data Suppose that
Hidden Markov Models Part 1: Introduction CSE 6363 Machine Learning Vassilis Athitsos Computer Science and Engineering Department University of Texas at Arlington 1 Modeling Sequential Data Suppose that
BUGS Bayesian inference Using Gibbs Sampling
 BUGS Bayesian inference Using Gibbs Sampling Glen DePalma Department of Statistics May 30, 2013 www.stat.purdue.edu/~gdepalma 1 / 20 Bayesian Philosophy I [Pearl] turned Bayesian in 1971, as soon as I
BUGS Bayesian inference Using Gibbs Sampling Glen DePalma Department of Statistics May 30, 2013 www.stat.purdue.edu/~gdepalma 1 / 20 Bayesian Philosophy I [Pearl] turned Bayesian in 1971, as soon as I
CSEP 573: Artificial Intelligence
 CSEP 573: Artificial Intelligence Hidden Markov Models Luke Zettlemoyer Many slides over the course adapted from either Dan Klein, Stuart Russell, Andrew Moore, Ali Farhadi, or Dan Weld 1 Outline Probabilistic
CSEP 573: Artificial Intelligence Hidden Markov Models Luke Zettlemoyer Many slides over the course adapted from either Dan Klein, Stuart Russell, Andrew Moore, Ali Farhadi, or Dan Weld 1 Outline Probabilistic
Glossary. The ISI glossary of statistical terms provides definitions in a number of different languages:
 Glossary The ISI glossary of statistical terms provides definitions in a number of different languages: http://isi.cbs.nl/glossary/index.htm Adjusted r 2 Adjusted R squared measures the proportion of the
Glossary The ISI glossary of statistical terms provides definitions in a number of different languages: http://isi.cbs.nl/glossary/index.htm Adjusted r 2 Adjusted R squared measures the proportion of the
Dynamic resource sharing
 J. Virtamo 38.34 Teletraffic Theory / Dynamic resource sharing and balanced fairness Dynamic resource sharing In previous lectures we have studied different notions of fair resource sharing. Our focus
J. Virtamo 38.34 Teletraffic Theory / Dynamic resource sharing and balanced fairness Dynamic resource sharing In previous lectures we have studied different notions of fair resource sharing. Our focus
Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features. Yangxin Huang
 Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features Yangxin Huang Department of Epidemiology and Biostatistics, COPH, USF, Tampa, FL yhuang@health.usf.edu January
Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features Yangxin Huang Department of Epidemiology and Biostatistics, COPH, USF, Tampa, FL yhuang@health.usf.edu January
13: Variational inference II
 10-708: Probabilistic Graphical Models, Spring 2015 13: Variational inference II Lecturer: Eric P. Xing Scribes: Ronghuo Zheng, Zhiting Hu, Yuntian Deng 1 Introduction We started to talk about variational
10-708: Probabilistic Graphical Models, Spring 2015 13: Variational inference II Lecturer: Eric P. Xing Scribes: Ronghuo Zheng, Zhiting Hu, Yuntian Deng 1 Introduction We started to talk about variational
Bagging During Markov Chain Monte Carlo for Smoother Predictions
 Bagging During Markov Chain Monte Carlo for Smoother Predictions Herbert K. H. Lee University of California, Santa Cruz Abstract: Making good predictions from noisy data is a challenging problem. Methods
Bagging During Markov Chain Monte Carlo for Smoother Predictions Herbert K. H. Lee University of California, Santa Cruz Abstract: Making good predictions from noisy data is a challenging problem. Methods
STOCHASTIC MODELING OF ENVIRONMENTAL TIME SERIES. Richard W. Katz LECTURE 5
 STOCHASTIC MODELING OF ENVIRONMENTAL TIME SERIES Richard W Katz LECTURE 5 (1) Hidden Markov Models: Applications (2) Hidden Markov Models: Viterbi Algorithm (3) Non-Homogeneous Hidden Markov Model (1)
STOCHASTIC MODELING OF ENVIRONMENTAL TIME SERIES Richard W Katz LECTURE 5 (1) Hidden Markov Models: Applications (2) Hidden Markov Models: Viterbi Algorithm (3) Non-Homogeneous Hidden Markov Model (1)
1. Introduction. Hang Qian 1 Iowa State University
 Users Guide to the VARDAS Package Hang Qian 1 Iowa State University 1. Introduction The Vector Autoregression (VAR) model is widely used in macroeconomics. However, macroeconomic data are not always observed
Users Guide to the VARDAS Package Hang Qian 1 Iowa State University 1. Introduction The Vector Autoregression (VAR) model is widely used in macroeconomics. However, macroeconomic data are not always observed
Lecture 2: Univariate Time Series
 Lecture 2: Univariate Time Series Analysis: Conditional and Unconditional Densities, Stationarity, ARMA Processes Prof. Massimo Guidolin 20192 Financial Econometrics Spring/Winter 2017 Overview Motivation:
Lecture 2: Univariate Time Series Analysis: Conditional and Unconditional Densities, Stationarity, ARMA Processes Prof. Massimo Guidolin 20192 Financial Econometrics Spring/Winter 2017 Overview Motivation:
CS839: Probabilistic Graphical Models. Lecture 7: Learning Fully Observed BNs. Theo Rekatsinas
 CS839: Probabilistic Graphical Models Lecture 7: Learning Fully Observed BNs Theo Rekatsinas 1 Exponential family: a basic building block For a numeric random variable X p(x ) =h(x)exp T T (x) A( ) = 1
CS839: Probabilistic Graphical Models Lecture 7: Learning Fully Observed BNs Theo Rekatsinas 1 Exponential family: a basic building block For a numeric random variable X p(x ) =h(x)exp T T (x) A( ) = 1
Statistics Boot Camp. Dr. Stephanie Lane Institute for Defense Analyses DATAWorks 2018
 Statistics Boot Camp Dr. Stephanie Lane Institute for Defense Analyses DATAWorks 2018 March 21, 2018 Outline of boot camp Summarizing and simplifying data Point and interval estimation Foundations of statistical
Statistics Boot Camp Dr. Stephanie Lane Institute for Defense Analyses DATAWorks 2018 March 21, 2018 Outline of boot camp Summarizing and simplifying data Point and interval estimation Foundations of statistical
Generative Models for Discrete Data
 Generative Models for Discrete Data ddebarr@uw.edu 2016-04-21 Agenda Bayesian Concept Learning Beta-Binomial Model Dirichlet-Multinomial Model Naïve Bayes Classifiers Bayesian Concept Learning Numbers
Generative Models for Discrete Data ddebarr@uw.edu 2016-04-21 Agenda Bayesian Concept Learning Beta-Binomial Model Dirichlet-Multinomial Model Naïve Bayes Classifiers Bayesian Concept Learning Numbers
The Model Building Process Part I: Checking Model Assumptions Best Practice
 The Model Building Process Part I: Checking Model Assumptions Best Practice Authored by: Sarah Burke, PhD 31 July 2017 The goal of the STAT T&E COE is to assist in developing rigorous, defensible test
The Model Building Process Part I: Checking Model Assumptions Best Practice Authored by: Sarah Burke, PhD 31 July 2017 The goal of the STAT T&E COE is to assist in developing rigorous, defensible test
Metropolis-Hastings Algorithm
 Strength of the Gibbs sampler Metropolis-Hastings Algorithm Easy algorithm to think about. Exploits the factorization properties of the joint probability distribution. No difficult choices to be made to
Strength of the Gibbs sampler Metropolis-Hastings Algorithm Easy algorithm to think about. Exploits the factorization properties of the joint probability distribution. No difficult choices to be made to
The main algorithms used in the seqhmm package
 The main algorithms used in the seqhmm package Jouni Helske University of Jyväskylä, Finland May 9, 2018 1 Introduction This vignette contains the descriptions of the main algorithms used in the seqhmm
The main algorithms used in the seqhmm package Jouni Helske University of Jyväskylä, Finland May 9, 2018 1 Introduction This vignette contains the descriptions of the main algorithms used in the seqhmm
Linear Dynamical Systems
 Linear Dynamical Systems Sargur N. srihari@cedar.buffalo.edu Machine Learning Course: http://www.cedar.buffalo.edu/~srihari/cse574/index.html Two Models Described by Same Graph Latent variables Observations
Linear Dynamical Systems Sargur N. srihari@cedar.buffalo.edu Machine Learning Course: http://www.cedar.buffalo.edu/~srihari/cse574/index.html Two Models Described by Same Graph Latent variables Observations
mlegp: an R package for Gaussian process modeling and sensitivity analysis
 mlegp: an R package for Gaussian process modeling and sensitivity analysis Garrett Dancik January 30, 2018 1 mlegp: an overview Gaussian processes (GPs) are commonly used as surrogate statistical models
mlegp: an R package for Gaussian process modeling and sensitivity analysis Garrett Dancik January 30, 2018 1 mlegp: an overview Gaussian processes (GPs) are commonly used as surrogate statistical models
Bayesian Networks in Educational Assessment
 Bayesian Networks in Educational Assessment Estimating Parameters with MCMC Bayesian Inference: Expanding Our Context Roy Levy Arizona State University Roy.Levy@asu.edu 2017 Roy Levy MCMC 1 MCMC 2 Posterior
Bayesian Networks in Educational Assessment Estimating Parameters with MCMC Bayesian Inference: Expanding Our Context Roy Levy Arizona State University Roy.Levy@asu.edu 2017 Roy Levy MCMC 1 MCMC 2 Posterior
BIOL 458 BIOMETRY Lab 9 - Correlation and Bivariate Regression
 BIOL 458 BIOMETRY Lab 9 - Correlation and Bivariate Regression Introduction to Correlation and Regression The procedures discussed in the previous ANOVA labs are most useful in cases where we are interested
BIOL 458 BIOMETRY Lab 9 - Correlation and Bivariate Regression Introduction to Correlation and Regression The procedures discussed in the previous ANOVA labs are most useful in cases where we are interested
Luke B Smith and Brian J Reich North Carolina State University May 21, 2013
 BSquare: An R package for Bayesian simultaneous quantile regression Luke B Smith and Brian J Reich North Carolina State University May 21, 2013 BSquare in an R package to conduct Bayesian quantile regression
BSquare: An R package for Bayesian simultaneous quantile regression Luke B Smith and Brian J Reich North Carolina State University May 21, 2013 BSquare in an R package to conduct Bayesian quantile regression
The Model Building Process Part I: Checking Model Assumptions Best Practice (Version 1.1)
 The Model Building Process Part I: Checking Model Assumptions Best Practice (Version 1.1) Authored by: Sarah Burke, PhD Version 1: 31 July 2017 Version 1.1: 24 October 2017 The goal of the STAT T&E COE
The Model Building Process Part I: Checking Model Assumptions Best Practice (Version 1.1) Authored by: Sarah Burke, PhD Version 1: 31 July 2017 Version 1.1: 24 October 2017 The goal of the STAT T&E COE
Visualizing Energy Usage and Consumption of the World
 Visualizing Energy Usage and Consumption of the World William Liew Abstract As the world becomes more and more environmentally aware, a simple layman s visualization of worldwide energy use is needed to
Visualizing Energy Usage and Consumption of the World William Liew Abstract As the world becomes more and more environmentally aware, a simple layman s visualization of worldwide energy use is needed to
Fast Algorithms for Large-State-Space HMMs with Applications to Web Usage Analysis
 Fast Algorithms for Large-State-Space HMMs with Applications to Web Usage Analysis Pedro F. Felzenszwalb 1, Daniel P. Huttenlocher 2, Jon M. Kleinberg 2 1 AI Lab, MIT, Cambridge MA 02139 2 Computer Science
Fast Algorithms for Large-State-Space HMMs with Applications to Web Usage Analysis Pedro F. Felzenszwalb 1, Daniel P. Huttenlocher 2, Jon M. Kleinberg 2 1 AI Lab, MIT, Cambridge MA 02139 2 Computer Science
Exponential Families
 Exponential Families David M. Blei 1 Introduction We discuss the exponential family, a very flexible family of distributions. Most distributions that you have heard of are in the exponential family. Bernoulli,
Exponential Families David M. Blei 1 Introduction We discuss the exponential family, a very flexible family of distributions. Most distributions that you have heard of are in the exponential family. Bernoulli,
STATISTICS 174: APPLIED STATISTICS FINAL EXAM DECEMBER 10, 2002
 Time allowed: 3 HOURS. STATISTICS 174: APPLIED STATISTICS FINAL EXAM DECEMBER 10, 2002 This is an open book exam: all course notes and the text are allowed, and you are expected to use your own calculator.
Time allowed: 3 HOURS. STATISTICS 174: APPLIED STATISTICS FINAL EXAM DECEMBER 10, 2002 This is an open book exam: all course notes and the text are allowed, and you are expected to use your own calculator.
Module 6: Model Diagnostics
 St@tmaster 02429/MIXED LINEAR MODELS PREPARED BY THE STATISTICS GROUPS AT IMM, DTU AND KU-LIFE Module 6: Model Diagnostics 6.1 Introduction............................... 1 6.2 Linear model diagnostics........................
St@tmaster 02429/MIXED LINEAR MODELS PREPARED BY THE STATISTICS GROUPS AT IMM, DTU AND KU-LIFE Module 6: Model Diagnostics 6.1 Introduction............................... 1 6.2 Linear model diagnostics........................
Computational statistics
 Computational statistics Markov Chain Monte Carlo methods Thierry Denœux March 2017 Thierry Denœux Computational statistics March 2017 1 / 71 Contents of this chapter When a target density f can be evaluated
Computational statistics Markov Chain Monte Carlo methods Thierry Denœux March 2017 Thierry Denœux Computational statistics March 2017 1 / 71 Contents of this chapter When a target density f can be evaluated
Multiple Sequence Alignment using Profile HMM
 Multiple Sequence Alignment using Profile HMM. based on Chapter 5 and Section 6.5 from Biological Sequence Analysis by R. Durbin et al., 1998 Acknowledgements: M.Sc. students Beatrice Miron, Oana Răţoi,
Multiple Sequence Alignment using Profile HMM. based on Chapter 5 and Section 6.5 from Biological Sequence Analysis by R. Durbin et al., 1998 Acknowledgements: M.Sc. students Beatrice Miron, Oana Răţoi,
TESTING FOR CO-INTEGRATION
 Bo Sjö 2010-12-05 TESTING FOR CO-INTEGRATION To be used in combination with Sjö (2008) Testing for Unit Roots and Cointegration A Guide. Instructions: Use the Johansen method to test for Purchasing Power
Bo Sjö 2010-12-05 TESTING FOR CO-INTEGRATION To be used in combination with Sjö (2008) Testing for Unit Roots and Cointegration A Guide. Instructions: Use the Johansen method to test for Purchasing Power
Generalized Linear Models for Non-Normal Data
 Generalized Linear Models for Non-Normal Data Today s Class: 3 parts of a generalized model Models for binary outcomes Complications for generalized multivariate or multilevel models SPLH 861: Lecture
Generalized Linear Models for Non-Normal Data Today s Class: 3 parts of a generalized model Models for binary outcomes Complications for generalized multivariate or multilevel models SPLH 861: Lecture
Undirected Graphical Models
 Outline Hong Chang Institute of Computing Technology, Chinese Academy of Sciences Machine Learning Methods (Fall 2012) Outline Outline I 1 Introduction 2 Properties Properties 3 Generative vs. Conditional
Outline Hong Chang Institute of Computing Technology, Chinese Academy of Sciences Machine Learning Methods (Fall 2012) Outline Outline I 1 Introduction 2 Properties Properties 3 Generative vs. Conditional
Dynamic Approaches: The Hidden Markov Model
 Dynamic Approaches: The Hidden Markov Model Davide Bacciu Dipartimento di Informatica Università di Pisa bacciu@di.unipi.it Machine Learning: Neural Networks and Advanced Models (AA2) Inference as Message
Dynamic Approaches: The Hidden Markov Model Davide Bacciu Dipartimento di Informatica Università di Pisa bacciu@di.unipi.it Machine Learning: Neural Networks and Advanced Models (AA2) Inference as Message
AUTOMATED TEMPLATE MATCHING METHOD FOR NMIS AT THE Y-12 NATIONAL SECURITY COMPLEX
 AUTOMATED TEMPLATE MATCHING METHOD FOR NMIS AT THE Y-1 NATIONAL SECURITY COMPLEX J. A. Mullens, J. K. Mattingly, L. G. Chiang, R. B. Oberer, J. T. Mihalczo ABSTRACT This paper describes a template matching
AUTOMATED TEMPLATE MATCHING METHOD FOR NMIS AT THE Y-1 NATIONAL SECURITY COMPLEX J. A. Mullens, J. K. Mattingly, L. G. Chiang, R. B. Oberer, J. T. Mihalczo ABSTRACT This paper describes a template matching
HMM part 1. Dr Philip Jackson
 Centre for Vision Speech & Signal Processing University of Surrey, Guildford GU2 7XH. HMM part 1 Dr Philip Jackson Probability fundamentals Markov models State topology diagrams Hidden Markov models -
Centre for Vision Speech & Signal Processing University of Surrey, Guildford GU2 7XH. HMM part 1 Dr Philip Jackson Probability fundamentals Markov models State topology diagrams Hidden Markov models -
Package breakaway. R topics documented: March 30, 2016
 Title Species Richness Estimation and Modeling Version 3.0 Date 2016-03-29 Author and John Bunge Maintainer Package breakaway March 30, 2016 Species richness estimation is an important
Title Species Richness Estimation and Modeling Version 3.0 Date 2016-03-29 Author and John Bunge Maintainer Package breakaway March 30, 2016 Species richness estimation is an important
Hidden Markov Models. Vibhav Gogate The University of Texas at Dallas
 Hidden Markov Models Vibhav Gogate The University of Texas at Dallas Intro to AI (CS 4365) Many slides over the course adapted from either Dan Klein, Luke Zettlemoyer, Stuart Russell or Andrew Moore 1
Hidden Markov Models Vibhav Gogate The University of Texas at Dallas Intro to AI (CS 4365) Many slides over the course adapted from either Dan Klein, Luke Zettlemoyer, Stuart Russell or Andrew Moore 1
Bayesian Analysis of Latent Variable Models using Mplus
 Bayesian Analysis of Latent Variable Models using Mplus Tihomir Asparouhov and Bengt Muthén Version 2 June 29, 2010 1 1 Introduction In this paper we describe some of the modeling possibilities that are
Bayesian Analysis of Latent Variable Models using Mplus Tihomir Asparouhov and Bengt Muthén Version 2 June 29, 2010 1 1 Introduction In this paper we describe some of the modeling possibilities that are
