Multi-state models: a flexible approach for modelling complex event histories in epidemiology
|
|
|
- Paulina Horn
- 6 years ago
- Views:
Transcription
1 Multi-state models: a flexible approach for modelling complex event histories in epidemiology Ahmadou Alioum Centre de Recherche INSERM U897 "Epidémiologie et Biostatistique Institut de Santé Publique, d Epidémiologie et de Développement (ISPED) Université Bordeaux Segalen, France March 7, 2013 Short-Course "Statistique Appliquée pour le Développement en Afrique 2013" (SADA 13), 4-8 mars 2013, Cotonou, Bénin
2 Organization of the course 1 Two-states (survival) models 2 Multistate models - Examples 3 Multistate processes - Markov processes 4 Transition probabilities and transition intensities 5 Patterns of observation 6 Likelihood 7 Likelihood inference 8 Non-parametric, semi-parametric estimation (Package mstate) 9 Problem with interval-censored data (Package SmoothHazard)
3 Two-state (survival) model Survival analysis in epidemiology = observing individuals over time, focusing on the occurence of a event of interest (death, occurrence of a disease or a complication,...) and time of occurrence The occurrence of the event of interest is a transition from one state to another Alive Death Dead State 0 α 01 (t) State 1 Onset of disease Healthy Diseased
4 Two-state (survival) model Positive random variable T = time from a given origin (in state 0 at time 0) to the occurrence of the event death If X = (X t, t 0) is a stochastic process with X t the state occupied by an individual at time t 0, X t S = {0, 1}, then T is the smallest time at which the process is not in the initial state 0 anymore T := inf{t : X t 0} Hazard rate function transition intensity λ(t) = lim t 0 P(t T < t + t T t) t = α 01 (t)
5 Two-state (survival) model Cumulative hazard function cumulative transition intensity Λ(t) = t 0 λ(u)du = A 01 (0, t) Survival function or distribution function transition probabilities S(t) = P(T > t) = exp[ Λ(t)] = P(X t = 0 X 0 = 0) = P 00 (0, t) F(t) = P(T t) = 1 S(t) = P(X t = 1 X 0 = 0) = P 01 (0, t) Transition probability matrix ( ) P00 (0, t) P P(0, t) = 01 (0, t) 0 1 Inference for hazard rate and survival functions taking into account incomplete observation of event histories (right-censoring, left-truncation,...)
6 Multi-states model : examples Generalization of survival models (two-state models) : more than one transition between more than two states Multi-state model (MSM) = model of continuous-time stochastic process allowing individuals to move among a finite number of states Extensive litterature on MSMs Books : ABGK (1993), Hougaard (2000), Aalen, Borgan and Gjessing (2008), Beyersmann, Schumacher and Allignol (2012) Reviews : Commenges (1999), Hougaard (1999), Andersen and Keiding (2002) Special Issues of : Statistical Methods in Medical Research on MSMs (2002) ; Journal of Statistical Software (2011)
7 Multi-states model : examples Competing risks model : one transient state (0: alive) and several absorbing states representing different causes of deaths 0: Alive. Death cause 1 Death cause 2 Death cause k
8 Multi-states model : examples Splitting the initial state into two or more states gives a progressive multi-state model HIV+ asymptomatic HIV+ symptomatic non AIDS AIDS
9 Multi-states model : examples Illness-death model : very useful in epidemiology to study both incidence of a disease and the mortality rate α 01 (t) : incidence of dementia α 02 (t) : mortality rate for healthy α 12 (t) : mortality rate for demented 0 : Healthy α 01 (t) 1: Demented α 02 (t) 2 : Dead α 12 (t)
10 Multi-states model : examples Bi-directional illness-death model : possibility of recovery from diseased state α 01 (t) 0 1 Healthy Diseased α 10 (t) α 02 (t) α 12 (t) 2 Dead
11 Multi-states model : progression of HIV infection, HIV diagnosis, inclusion in a cohort and death Infection ν(t) VIH+ asymptomatic VIH+ symptomatic AIDS HIV diagnosis Inclusion in the cohort Death
12 Stochastic processes and filtration Continuous-time stochastic process X = (X t, t 0) We should think of a filtration as a flow of information. The σ-algebra F t contains the events that can happen "up to time t". The filtration defined by F X t = σ(x s : s t) is called the filtration generated by X or the natural filtration of X (keep tracks of the "history" of the process) A multi-state process is a process which can take a finite number of states; that is, for any t, the variable X t has values in S = {0, 1,..., K }. The law of a multi-state process is defined by its finite dimensional distribution: P(X t1 = j 1, X t2 = j 2,..., X tn = j n ), n N
13 Specification of multi-state process A multi-state model is fully characterized through Transition probabilities between state h and state j : P hj (s, t; F s ) = P(X t = j X s = h, F s ), for s < t Transition intensities α hj (t; F t ) = lim t 0 P hj (t, t + t; F t ) t Cumulative (integrated) transition intensities A hj (t; F t ) = t 0 α hj (u; F u )du
14 Markov processes Markov assumption : the future and the past of the process are independent given its present state P hj (s, t; F s ) = P hj (s, t) = P(X t = j X s = h) s < t, h, j S, h j and P hh (s, t) = 1 j h P hj(s, t) P(s, t) = {P hj (s, t)} matrix of transition probabilities Transition intensities are then defined as: α hj (t) = lim t 0 P hj (t, t + t)/ t, h j, and α hh (t) = j h α hj(t); j h α hj(t) is the hazard function associated with the distribution of the sojourn time h. a(t) = {α hj (t)} matrix of transition intensities A(t) = {A hj (t)} matrix of cumulative transition intensities, where A hj (t) = t 0 α hj(u)du
15 Relationship between transition probabilities and intensities Chapman-Kolmogorov equation: P(s, t) = P(s, u)p(u, t), s < u < t P(s, t) is the unique solution of the Kolmogorov forward differential equation: t P(s, t) = P(s, t)a(t); P(s, s) = I P(s, t) can be recovered from the transition intensities through product integration where is a product integral P(s, t) = (I + da(u)) u (s,t]
16 Relationship between transition probabilities and intensities For simple models, such as the illness-death model, the transition probabilities has explicit expression P 00 (s, t) = e [A 01(s,t)+A 02 (s,t)] P 11 (s, t) = e A 12(s,t) P 01 (s, t) = t s P 00(s, u)α 01 (u)p 11 (u, t)du Particular case : homogeneous Markov chain a(t) is constant; that is α hj (t) = α hj does not depend on time t, then P(s, t) = P(0, t s) = P(t s) The solution of Kolmogorov equation is : P(t) = e ta If the transition intensities matrix a can be diagonalized, then P(t) = V diag(e ρ 0t,..., e ρ K t ) V 1 Explicit formulas for the illness-death model
17 Aalen-Johansen estimator N hj (t): number of observed direct transitions from state h to state j up to time t; Y h (t): number of individuals under observation in state h just before time t da(t) matrix of elements d(a hj (t)) h,j = (α hj (t)) h,j dt Nelson-Aalen estimator of the cumulative transition intensities:  hj (t) = u t dâhj(t) = dn hj(t) Y h (t) dâhj(t), h j and Âhh(t) = h j  hj (t) Aalen-Johansen estimator of the transition probabilities : Â(t) is a matrix of step-functions with finite number of jumps in (s, t] ˆP(s, t) = (I + dâ(u)) s<u t
18 Special case : Illness-death model Aalen-Johansen estimators of P 00 (s, t) and P 11 (s, t) are equal to the corresponding Kaplan-Meier estimators ˆP 00 (s, t) = s<u t (1 dâ01(u) dâ02(u)) ˆP 11 (s, t) = s<u t (1 dâ12(u)) ˆP 01 (s, t) = s<u t ˆP 00 (s, u )dâ01(u)ˆp 11 (u+, t) Then since ˆP 02 (s, t) = 1 ˆP 00 (s, t) ˆP 01 (s, t) and ˆP 12 (s, t) = 1 ˆP 11 (s, t) ˆP 02 (s, t) = s<u t ˆP 00 (s, u )dâ01(u)ˆp 11 (u+, t) + s<u t ˆP 00 (s, u )dâ02(u) ˆP 12 (s, t) = s<u t ˆP 11 (s, u )dâ12(u)
19 Semi-Markov process The most used semi-markov process can be specified by: P hj (s, t, T h ) = P(X t = j X s = h, T h ), s < t α hj (t, T h ) = lim t 0 P hj (t, t + t, T h )/ t These quantities are random and we do not have the Kolmogorov equations. In general we may still be able to derive transition probabilities from transition intensities but the theory is more complex. Clock reset models : the relevant time is the time spent in the current state (the clock is "reset" to 0 each time a patient enters a new state)
20 Semi-Markov model : an example 0: HIV- α 01 (t) α 12 (τ) 1: HIV+ 2: AIDS Figure: Progressive three-state Model: if τ = t this is a Markov model; if τ is the time since the 0 1 transition this is a semi-markov model.
21 Multistate models for a population States and time Associate to each subject i of a population a multistate process X i = (Xt i) t 0 with law specified by a i (.). States of the X i s and time t have same meaning for all the subjects. However, several choices of t are possible: t may be the calendar time or time since a subject-specific event; if the event is birth, t is age. Heterogeneity of intensities The population may be homogeneous or heterogeneous. Heterogenity may be modelled as depending of observed covariates Z i (t): α i hj (t) = φ hj(α hj0 (t), Z i (t)) Proportional intensity assumption: α hj (t, Z i (t)) = α hj0 (t)e β hj Z i (t) Additive intensity α hj (t, Z i (t)) = α hj0 (t) + β hj (t)z i (t)
22 Patterns of observation Selection of the sample For estimation purpose we have to draw a sample of size n from a population. It often happens that subjects can enter the sample only if they are in a given state. For instance in an illness-death model the study sample may be a random sample of healthy subject. The condition is: X i t i0 = 0 This is a kind of left-truncation which must be taken into account for inference.
23 Patterns of observation Left- and right-censoring Subject i may be first observed at time L i until time C i. We observe: (X i t, L i t C i ) If L i = 0 this the extension of right-censoring to multistate processes.
24 Patterns of observation Discrete-time observations We may consider that the state of the process i is observed at only a finite number of times V i 0, V i 1,..., V i m. This typically happens in cohort studies where fixed visit times have been planned. In such cases the exact times of transitions are not known; it is only known that they occurred during a particular interval; these observations are said to be interval-censored. It is also possible that the state of the process is not exactly observed but it is known that it belongs to a subset of {0, 1,..., K }
25 Patterns of observation Mixed continuous and discrete-time observations The most common pattern of observation is in fact a mixing of discrete and continuous time observations. This is because most multistate models include states which represent clinical status and one state which represents death: most often clinical status is observed at discrete times (visits) while the (nearly) exact time of death can be retrieved. It may also happen that some events other than death are observed exactly.
26 Likelihood Using the likelihood approach for inference taking into account selection and censoring. The likelihood can be written easily in terms of the P hj (.,.) and the α hj (.) under the assumption of ignorability conditional on the state at the first observation time V 0 the processes X i s are independent the processes are Markov
27 Likelihood Likelihood: continuous-time observations The process X is continuously observed from V 0 to C. We observe that transitions have occured at (exactly) times T 1, T 2,..., T m. With the convention T 0 = V 0, the likelihood (conditional on X(V 0 )) is: m 1 L = [ P X(T r ),X(Tr )(T r,t r+1 )α X(T r ),X(T r+1 )(T r+1 )]P X(T m),x(tm)(t m,c). r=0 Likelihood: discrete-time observations Observations of X are taken at V 0, V 1,..., V m. We write the likelihood as if the V j were fixed (ignorability): L = m 1 r=0 P X(V r ),X(V r+1 )(V r,v r+1 ).
28 Likelihood Likelihood: Mixed discrete-continuous time observations For simplicity we give the likelihood when the process is observed at discrete times but time of transition towards one absorbing state, representing generally death, is exactly observed or right-censored. Denote by K this absorbing state. Observations of X are taken at V 0, V 1,..., V m and the vital status is observed until C ( C V m ). Let us call T the follow-up time that is T = min(t, C), where T is the time of death; we observe T and δ = I{T C}. The likelihood is: L = m 1 r=0 P X(V r ),X(V r+1 )(V r,v r+1 ) P X(V m),j (V m, T )α j,k (T ) δ. j K
29 Example: likelihood in the illness-death model If the subject starts in state health, has never been observed in the illness state and was last seen at visit L (at time V L ) the likelihood is: L = P 00 (V 0, V L )[P 00 (V L, T )α 02 ( T ) δ + P 01 (V L, T )α 12 ( T ) δ ] If the subject has been observed in the illness state for the first time at V J then the likelihood is: L = P 00 (V 0, V J 1 )P 01 (V J 1, V J )P 11 (V J, T )α 12 ( T ) δ.
30 Likelihood inference Once we have a likelihood L(a(.), X ), several approaches for getting estimators can be used Non-parametric: no restriction on a(.) Parametric: a(.) M Smooth non-parametric by smoothing a non-parametric estimate Smooth non-parametric by penalized likelihood Bayesian approach The non-parametric methods may be combined with a parametric form for effects of covariates leading to semi-parametric models. α hj (t, Z ) = α hj,0 (t) exp (β hj Z ) ˆP(s, t; Z ) = (I + dâ(u; Z )) s<u t
31 Case with continuous time observations If transition times are exactly observed or right-censored, the problem can be split in as many survival problems as there are transitions. For instance if subject i enters state h at Th i and makes a transition from state h to state j at Tj i his likelihood contribution to estimation of α hj (.) is e T i j T h i α hj (u)du αhj (T i j ) If a subject makes a transition to another state it is considered as censored. Estimation of α hj (.) can be done by survival methods allowing left-truncation (delayed entry) and right-censoring. Estimation may be parametric or semi-parametric (Cox model).
32 Illness-death model 0 : Healthy α 02 (t) α 01 (t) 2 : Dead 1: Diseased α 12 (t)
33 An example : Transplant treatment of patients with blood cancer From de Wreede, Fiocco and Putter (Journal of Statistics Software, 2011, vol 38, Issue 7)
34 An example : Transplant treatment of patients with blood cancer Pronostic factors for all patients Pronostic factor Categories n(%) (name in data) Donor recipient no gender mismatch 1734 (76) (match) gender mismatch 545 (24) Prophylaxis no 1730 (76) (proph) yes 549 (24) Year of transplant (28) (year) (39) (33) Age at transplant (years) (24) (agec1) (53) > (23)
35 Package mstate Estimation and prediction (non-parametric and semi-parametric) : cumulative transition intensities, transition probabilities, prediction Need to make a dataset with a line for each transition (data preparation)
36 Package mstate : semi-parametric models Incorporate covariate information in Cox PH models Separate Cox models for each of the transitions α hj (t; Z ) = α hj,0 (t) exp(β hj Z ) Build appropriate subsets of data and estimate covariate vector and baseline intensity for each transition (standard software) Transition-specific covariates Model all transitions in a single Cox regression model, using stratified regression and transition-specific covariates α hj (t; Z hj ) = α hj,0 (t) exp(βz hj ) Obtain different covariate effect estimates accross transition (transition-specific covariates) Obtain a single effect estimate assumed to be common for all transitions (stratified Cox regression using the global covariate)
37 Package mstate > library(mstate) > data("ebmt4") > ebmt <- ebmt4 > head(ebmt4) id rec rec.s ae ae.s recae recae.s rel rel.s srv srv.s year agecl > proph match 1 no no gender mismatch 2 no no gender mismatch 3 no no gender mismatch 4 no gender mismatch 5 no gender mismatch 6 no no gender mismatch > tmat <- transmat(x = list(c(2,3,5,6), c(4,5,6), c(4,5,6), c(5,6), c(), c()), names=c("tx", "Rec", "AE", "Rec+AE", "Rel", "Death")) > tmat to from Tx Rec AE Rec+AE Rel Death Tx NA 1 2 NA 3 4 Rec NA NA NA AE NA NA NA Rec+AE NA NA NA NA Rel NA NA NA NA NA NA Death NA NA NA NA NA NA
38 Package mstate > msebmt <- msprep(data=ebmt, trans=tmat, time=c(na, "rec", "ae", "recae", "rel", "srv"), status=c(na, "rec.s", "ae.s", "recae.s", "rel.s", "srv.s"), keep=c("match", "proph", "year", "agecl")) > msebmt[msebmt$id == 1, c(1:8, 10:12)] An object of class 'msdata' Data: id from to trans Tstart Tstop time status proph year agecl no no no no no no no > events(msebmt) $Frequencies to from Tx Rec AE Rec+AE Rel Death no event total entering Tx Rec AE Rec+AE Rel Death
39 Package mstate > covs <- c("match", "proph", "year", "agecl") > msebmt <- expand.covs(msebmt, covs, longnames=false) > msebmt[msebmt$id == 1, -c(9, 10, 12:48, 61:84)] An object of class 'msdata' Data: id from to trans Tstart Tstop time status year year2.1 year2.2 year year2.4 year2.5 year2.6 year2.7 year2.8 year2.9 year2.10 year2.11 year
40 Package mstate > msebmt[, c("tstart", "Tstop", "time")] <- msebmt[, c("tstart", "Tstop", "time")]/ > c0 <- coxph(surv(tstart, Tstop, status) ~ strata(trans), data=msebmt, method="breslow") > msf0 <- msfit(object=c0, vartype="greenwood", trans=tmat) > head(msf0$haz) time Haz trans > tail(msf0$haz) time Haz trans > plot (msf0, las=1, lty= rep(1:2, c(8,4)), xlab="years since transplantation")
41 Years since transplantation Package mstate Cumulative hazard
42 Package mstate > pt0 <- probtrans(msf0, predt=0, method="greenwood") > summary(pt0) An object of class 'probtrans' Prediction from state 1 (head and tail): time pstate1 pstate2 pstate3 pstate4 pstate5 pstate6 se se2 se3 se4 se5 se time pstate1 pstate2 pstate3 pstate4 pstate5 pstate6 se
43 Package mstate
44 The computational problem with interval censored observations Dealing with interval-censored data in multistate models
45 Two-state (survival) model with interval-censored data Non-parametric estimation of the survival function The equivalent of the Kaplan-Meier estimation for interval-censored data was first proposed by Peto (1973), and Turnbull (1976) then improved the numerical algorithm to estimate the survival function (EM algorithm) From the data {(L i, R i ], i = 1, 2,..., n} a set of non-overlapping intervals {(q 1, p 1 ], (q 2, p 2 ],..., (q m, p m ]} is generated over which S(t) is estimated : the NPMLE of S(t) can decrease only on intervals (q j, p j ]. PH model S(t; Z ) = S 0 (t) exp(βz ) : estimation (non-parametric) of the baseline survival function S 0 (t) and of the regression coefficients (Finkelstein, 1986; Alioum and Commenges, 1996; Goetghebeur and Ryan, 2000).
46 NPMLE of the survival curve
47 Illness-death model with interval-censored data Observations pattern : times of transition from 0 to 1 are interval-censored, but times to the absorbing state are assumed to be known exactly or to be right-censored If you re just interested in α 01, you can use survival methods for interval censored data, but α 01 will be underestimated (e.g. incidence of dementia ; Joly et al., 2002) Problem with survival analysis: using death as right-censoring leads to a biased estimates of age-specific incidence of dementia A more adapted model is an illness-death model (Commenges et al., 1998; Commenges et al., 2004)
48 Incidence of Dementia : survival and illness-death model
49 Parametric/Smooth non-parametric Parametric approach : easy to implement using likelihood methods Smooth non-parametric via penalized likelihood (Joly et al., 2002) α hj (t; Z ) = α hj,0 (t) exp (β hj Z ), hj {01, 02, 12} pl(α 01 (.), α 02 (.), α 12 (.); β 01, β 02, β 12 ) = + L(α hj (.); β hj ) κ 01 [α 01,0 (u)]2 du κ [α 0 02,0 (u)]2 du κ [α 12,0 (u)]2 du
50 Parametric/Smooth non-parametric NPMLE ˆα 01 (.), ˆα 02 (.), ˆα 12 (.) are approximated using M-splines α hj (t) = l θ l hj M l(t) Λ hj (t) = l θ l hj I l(t) κ 01, κ 02, κ 12 are estimated simultaneously by maximizing the standard crossvalidation score SmoothHazard : A new R package to fit illness-death model (Weibull or penalized likelihood) for left-truncated and/or interval-censored data (written by Tourraine, Joly and Gerds)
51 Package SmoothHazard
52 Package SmoothHazard t0: left truncation time t1: for diseased subjects, left endpoint of the censoring interval; for non diseased subjects, right censoring time for 0->1 transition t2: for diseased subjects, right endpoint of the censoring interval; for non diseased subjects, right censoring time for the disease event t3: for subjects who died, time of death; for alive subjects, censoring time for the death event > library(smoothhazard) > data(paq1000) > head(paq1000) dementia death t0 t1 t2 t3 certif gender
53 Package SmoothHazard > fit <- idm(formula02=hist(time,event=death,entry=entry)~certif, + formula01=hist(time=list(l,r),event=dementia)~certif, + data=d, hazard="splines") > print(fit) Call: idm(formula01 = Hist(time = list(l, R), event = dementia) ~ certif, formula02 = Hist(time, event = death, entry = entry) ~ certif, data = d, hazard = "Splines") Illness-death model using a penalized likelihood approach with splines approximation for the intensity functions. number of subjects: 1000 number of events '0-->1': 186 number of events '0-->2' or '0-->1-->2': 724 number of subjects: 1000 number of covariates: Smoothing parameters: transition01 transition02 transition12 knots 7e+00 7e+00 7 kappa 1e+06 5e coef SE.coef HR CI Wald p.value certif_ [0.41;0.93] certif_ [0.91;1.50] certif_ [0.59;1.47] Without cov With cov Penalized log likelihood > plot(fit)
54 Time Package SmoothHazard Splines transition intensities
55 Package SmoothHazard Other features of SmoothHazard for both Weibull model and penalized likelihood Plot baseline survival functions Predict transition probabilities Predict life expectancies (in state 0; for a non diseased subject; for a diseased subject) at time s for a subject with a given vector of covariates Fit survival model using either a penalized likelihood approach or a parametric (Weibull) approach from left-truncated and interval-censored data
56 More complex models : Dementia-Institution-Death 0: Healthy 1: Demented α 01 (t) α 13 (t) α 14 (t) α 04 (t) α 34 (t) 4: Dead 3: Dem+Inst α 02 (t) α 24 (t) α 23 (t) 2: Instit
57 More complex models : Dementia-Institution-Death Smooth non-parametric by penalized likelihood Need some additional assumptions (Joly et al., 2009) α 14 (t) = α 04 (t)e θ 14 α 23 (t) = α 01 (t)e θ 23 α 24 (t) = α 04 (t)e θ 24 α 13 (t) = α 02 (t)e θ 13 α 34 (t) = α 04 (t)e θ 34 pl(α 01 (.), α 02 (.), α 04 (.); θ) = L(α 01, α 02, α 04 ; θ) κ 01 + κ [α 0 [α 02 (u)]2 du κ (u)]2 du [α 04 (u)]2 du
58 Panel data States X i (t) of individual i(i = 1, 2,..., n) is only known at discrete consecutive follow-up times t = (t i,0, t i,1,..., t i,mi ), and the exact time of transitions from one state to another is not observed Likelihood conditionally on X(t i,0 ) n m i P X(ti,j 1 ),X(t i,j )(t i,j 1, t i,j ; Z i (t i,j 1 )) i=1 j=1 If X(t i,mi ) = K (single absorbing state) and t i,mi is the exact time of transition to K, then the expression of the last term of the product is given by P X(ti,mi 1),l(t i,mi 1, t i,mi ; Z i (t i,mi 1))α l,k (t i,mi ) l K
59 Example of more complex multi-state models Disablement process in the elderly (Paquid study)
60 HMM/NHMM with PCI NHMM with piecewise constant intensities: partitioning time into 2 or more intervals and assume constant intensity in each interval PIM : α hj (t; Z (t)) = α hj0 (t) exp (β Z (t)) where α hj0 (t) = α l hj0 for τ l 1 < t τ l, (l = 1, 2,..., r) The expressions of transition probabilities P hj (s, t) become more complicated (even if we keep the idea of fitting of homogeneous models to a series of r intervals) This method needs prior specification of the cutpoints; the choice of cutpoints should make sure that an appropriate number of observations fall in all intervals
61 Programs and Packages for fitting HMM/NHMM with PCI MKVPCI Fortran program (Alioum and Commenges, 2001) Fitting general k-state Markov models (k 1 transient states and the kth state can be optionally chosen as an absorbing state) with PCI and covariates The output includes estimates of the baseline transition intensities, regression coefficients, transition intensities in time intervals (τ l 1, τ l ], l = 1, 2,..., r, together with their estimated standard errors, and the results of the multivariate hypotheses tests.
62 Example 2: disablement process in the elderly File of input parameter for MKVPCI disab.dat gender educ
63 Example 2: disablement process in the elderly Multi-state Markov model with piecewise constant intensities Program developed by Ahmadou ALIOUM ISPED - University Victor Segalen Bordeaux 2 Markov model with piecewise constant intensities : two periods Data file : disab.dat Number of records : Number of subjects : 3461 Number of states : 5 Number of transitions : 10 Number of artificial covariates : 1 Cut points : Number of observed covariates : 0 Number of parameters : 20 Exact transition times to the absorbing state : 1 Baseline transition intensities Time intervals Tau(0) - Tau(1) Tau(1) - Tau(2) TRANSITION ESTIMATES STD ERROR ESTIMATES STD ERROR 1 > > > > > > > > > >
64 R package msm Package msm developped by Christopher H. Jackson (Journal of Statistical Software, 2011, vol. 38, issue 8) Markov model with any number of states and any pattern of transitions to panel data : HMM, NHMM with PCI Proportional intensities models : piecewise constant time-dependant covariates Exact death times, censored states (alive but in an unknown disease state at the end of the study) Hidden Markov models in which the states of the Markov chain are not observed : observed data X ij are governed by some probability distribution conditionally on the unobserved state S ij Tools for model assessment
65 Estimation of HIV incidence from a prevalent cohort Infection ν(t) VIH+ asymptomatic VIH+ symptomatic AIDS HIV diagnosis Inclusion in the cohort Death
66 Estimation of HIV incidence from a prevalent cohort Infection occurs according to a non-homogeneous Poisson process with intensity ν(t) Non-homogeneous Markov process with piecewise constant intensities Data from a prevalent cohort study : a total of n subjects were enrolled in the cohort by time T s i time of first HIV positive test X i (t i0 ), X i (t i1 ),..., X i (t imi ) Sampling criterion : a subject i is enrolled only if t i0 T The number n is the realization of a random variable N, and the observation of N = n brings information on ν(t)
67 Estimation of HIV incidence from a prevalent cohort Conditionally on the occurrence of a certain number of infections in the period [0, T ], the times of infection a iid with density function ν (t) = ν(t)/ T 0 ν(u)du Conditional on N = n, the likelihood of the trajectories of observed subjects is L 1 = n i=1 L i/p I where P I = T 0 ν (u)p (u)du, and L i = P(s i, t i0, x i0 ) m i j=1 P X i (t i,j 1 ),X i (t i,j )(t i,j 1, t i,j )
68 Estimation of HIV incidence from a prevalent cohort N follows a Poisson distribution with parameter A = T 0 ν(u)p (u)du : L 2 = P(N = n) Full likelihood : L = L 1 L 2 Smooth estimate of HIV is obtained by maximazing the penalized log-likelihood : pl(ν, α) = log L 1 + log L 2 λ ν (u) 2 du
69 Estimation of HIV incidence from a prevalent cohort Data from a hospital-based surveillance system of HIV infection implemented since 1987 in Aquitaine region, Southwestern France : homosexuals/bisexuals (λ = 100 ; GECSA, )
70 Concluding remarks Methods for inference in MSM from selected sample or/and interval-censored data are useful in epidemiology Some software are now available for fitting simple MSM models (mstate, msm, smoothhazard, among others) Some challenges for modelling in the MSM framework more complex processes unobserved heterogeneity, clustering dependant LTF or missing data model checking and validation
71 Some selected references PK Andersen, O Borgan, RD Gill and N Keiding (1993), Statistical Models Based on Counting Processes, Springer, NY P Hougaard (2000), Analysis of Multivariate Survival Data, Springer, NY Aalen, Borgan and Gjessing (2008), Event History Analysis, Springer, NY Beyersmann, Schumacher and Allignol (2012), Competing Risks and Multistate Models with R, Springer, NY A Alioum et al. (2005), Applied Statistics 54, PK Andersen and Keiding (2002), Statist. Methods Med. Res. 11, D Commenges (2002), Statist. Methods Med. Res. 11, D Commenges et al. (2004), Statist. Med. 23, P Joly et al. (2002), Biostatistics 3, de Wreede, Fiocco and Putter (2011), Journal of Statistical Software Volume 38, Issue 7,
Philosophy and Features of the mstate package
 Introduction Mathematical theory Practice Discussion Philosophy and Features of the mstate package Liesbeth de Wreede, Hein Putter Department of Medical Statistics and Bioinformatics Leiden University
Introduction Mathematical theory Practice Discussion Philosophy and Features of the mstate package Liesbeth de Wreede, Hein Putter Department of Medical Statistics and Bioinformatics Leiden University
Journal of Statistical Software
 JSS Journal of Statistical Software January 2011, Volume 38, Issue 7. http://www.jstatsoft.org/ mstate: An R Package for the Analysis of Competing Risks and Multi-State Models Liesbeth C. de Wreede Leiden
JSS Journal of Statistical Software January 2011, Volume 38, Issue 7. http://www.jstatsoft.org/ mstate: An R Package for the Analysis of Competing Risks and Multi-State Models Liesbeth C. de Wreede Leiden
Multi-state models: prediction
 Department of Medical Statistics and Bioinformatics Leiden University Medical Center Course on advanced survival analysis, Copenhagen Outline Prediction Theory Aalen-Johansen Computational aspects Applications
Department of Medical Statistics and Bioinformatics Leiden University Medical Center Course on advanced survival analysis, Copenhagen Outline Prediction Theory Aalen-Johansen Computational aspects Applications
A multi-state model for the prognosis of non-mild acute pancreatitis
 A multi-state model for the prognosis of non-mild acute pancreatitis Lore Zumeta Olaskoaga 1, Felix Zubia Olaskoaga 2, Guadalupe Gómez Melis 1 1 Universitat Politècnica de Catalunya 2 Intensive Care Unit,
A multi-state model for the prognosis of non-mild acute pancreatitis Lore Zumeta Olaskoaga 1, Felix Zubia Olaskoaga 2, Guadalupe Gómez Melis 1 1 Universitat Politècnica de Catalunya 2 Intensive Care Unit,
Multi-state Models: An Overview
 Multi-state Models: An Overview Andrew Titman Lancaster University 14 April 2016 Overview Introduction to multi-state modelling Examples of applications Continuously observed processes Intermittently observed
Multi-state Models: An Overview Andrew Titman Lancaster University 14 April 2016 Overview Introduction to multi-state modelling Examples of applications Continuously observed processes Intermittently observed
Robust estimates of state occupancy and transition probabilities for Non-Markov multi-state models
 Robust estimates of state occupancy and transition probabilities for Non-Markov multi-state models 26 March 2014 Overview Continuously observed data Three-state illness-death General robust estimator Interval
Robust estimates of state occupancy and transition probabilities for Non-Markov multi-state models 26 March 2014 Overview Continuously observed data Three-state illness-death General robust estimator Interval
A multi-state model for the prognosis of non-mild acute pancreatitis
 A multi-state model for the prognosis of non-mild acute pancreatitis Lore Zumeta Olaskoaga 1, Felix Zubia Olaskoaga 2, Marta Bofill Roig 1, Guadalupe Gómez Melis 1 1 Universitat Politècnica de Catalunya
A multi-state model for the prognosis of non-mild acute pancreatitis Lore Zumeta Olaskoaga 1, Felix Zubia Olaskoaga 2, Marta Bofill Roig 1, Guadalupe Gómez Melis 1 1 Universitat Politècnica de Catalunya
A multistate additive relative survival semi-markov model
 46 e Journées de Statistique de la SFdS A multistate additive semi-markov Florence Gillaizeau,, & Etienne Dantan & Magali Giral, & Yohann Foucher, E-mail: florence.gillaizeau@univ-nantes.fr EA 4275 - SPHERE
46 e Journées de Statistique de la SFdS A multistate additive semi-markov Florence Gillaizeau,, & Etienne Dantan & Magali Giral, & Yohann Foucher, E-mail: florence.gillaizeau@univ-nantes.fr EA 4275 - SPHERE
Lecture 7 Time-dependent Covariates in Cox Regression
 Lecture 7 Time-dependent Covariates in Cox Regression So far, we ve been considering the following Cox PH model: λ(t Z) = λ 0 (t) exp(β Z) = λ 0 (t) exp( β j Z j ) where β j is the parameter for the the
Lecture 7 Time-dependent Covariates in Cox Regression So far, we ve been considering the following Cox PH model: λ(t Z) = λ 0 (t) exp(β Z) = λ 0 (t) exp( β j Z j ) where β j is the parameter for the the
Multistate Modeling and Applications
 Multistate Modeling and Applications Yang Yang Department of Statistics University of Michigan, Ann Arbor IBM Research Graduate Student Workshop: Statistics for a Smarter Planet Yang Yang (UM, Ann Arbor)
Multistate Modeling and Applications Yang Yang Department of Statistics University of Michigan, Ann Arbor IBM Research Graduate Student Workshop: Statistics for a Smarter Planet Yang Yang (UM, Ann Arbor)
Modeling Prediction of the Nosocomial Pneumonia with a Multistate model
 Modeling Prediction of the Nosocomial Pneumonia with a Multistate model M.Nguile Makao 1 PHD student Director: J.F. Timsit 2 Co-Directors: B Liquet 3 & J.F. Coeurjolly 4 1 Team 11 Inserm U823-Joseph Fourier
Modeling Prediction of the Nosocomial Pneumonia with a Multistate model M.Nguile Makao 1 PHD student Director: J.F. Timsit 2 Co-Directors: B Liquet 3 & J.F. Coeurjolly 4 1 Team 11 Inserm U823-Joseph Fourier
Statistics 262: Intermediate Biostatistics Non-parametric Survival Analysis
 Statistics 262: Intermediate Biostatistics Non-parametric Survival Analysis Jonathan Taylor & Kristin Cobb Statistics 262: Intermediate Biostatistics p.1/?? Overview of today s class Kaplan-Meier Curve
Statistics 262: Intermediate Biostatistics Non-parametric Survival Analysis Jonathan Taylor & Kristin Cobb Statistics 262: Intermediate Biostatistics p.1/?? Overview of today s class Kaplan-Meier Curve
DAGStat Event History Analysis.
 DAGStat 2016 Event History Analysis Robin.Henderson@ncl.ac.uk 1 / 75 Schedule 9.00 Introduction 10.30 Break 11.00 Regression Models, Frailty and Multivariate Survival 12.30 Lunch 13.30 Time-Variation and
DAGStat 2016 Event History Analysis Robin.Henderson@ncl.ac.uk 1 / 75 Schedule 9.00 Introduction 10.30 Break 11.00 Regression Models, Frailty and Multivariate Survival 12.30 Lunch 13.30 Time-Variation and
Understanding product integration. A talk about teaching survival analysis.
 Understanding product integration. A talk about teaching survival analysis. Jan Beyersmann, Arthur Allignol, Martin Schumacher. Freiburg, Germany DFG Research Unit FOR 534 jan@fdm.uni-freiburg.de It is
Understanding product integration. A talk about teaching survival analysis. Jan Beyersmann, Arthur Allignol, Martin Schumacher. Freiburg, Germany DFG Research Unit FOR 534 jan@fdm.uni-freiburg.de It is
An Overview of Methods for Applying Semi-Markov Processes in Biostatistics.
 An Overview of Methods for Applying Semi-Markov Processes in Biostatistics. Charles J. Mode Department of Mathematics and Computer Science Drexel University Philadelphia, PA 19104 Overview of Topics. I.
An Overview of Methods for Applying Semi-Markov Processes in Biostatistics. Charles J. Mode Department of Mathematics and Computer Science Drexel University Philadelphia, PA 19104 Overview of Topics. I.
ADVANCED STATISTICAL ANALYSIS OF EPIDEMIOLOGICAL STUDIES. Cox s regression analysis Time dependent explanatory variables
 ADVANCED STATISTICAL ANALYSIS OF EPIDEMIOLOGICAL STUDIES Cox s regression analysis Time dependent explanatory variables Henrik Ravn Bandim Health Project, Statens Serum Institut 4 November 2011 1 / 53
ADVANCED STATISTICAL ANALYSIS OF EPIDEMIOLOGICAL STUDIES Cox s regression analysis Time dependent explanatory variables Henrik Ravn Bandim Health Project, Statens Serum Institut 4 November 2011 1 / 53
Survival Analysis I (CHL5209H)
 Survival Analysis Dalla Lana School of Public Health University of Toronto olli.saarela@utoronto.ca January 7, 2015 31-1 Literature Clayton D & Hills M (1993): Statistical Models in Epidemiology. Not really
Survival Analysis Dalla Lana School of Public Health University of Toronto olli.saarela@utoronto.ca January 7, 2015 31-1 Literature Clayton D & Hills M (1993): Statistical Models in Epidemiology. Not really
Multistate models and recurrent event models
 Multistate models Multistate models and recurrent event models Patrick Breheny December 10 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/22 Introduction Multistate models In this final lecture,
Multistate models Multistate models and recurrent event models Patrick Breheny December 10 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/22 Introduction Multistate models In this final lecture,
Definitions and examples Simple estimation and testing Regression models Goodness of fit for the Cox model. Recap of Part 1. Per Kragh Andersen
 Recap of Part 1 Per Kragh Andersen Section of Biostatistics, University of Copenhagen DSBS Course Survival Analysis in Clinical Trials January 2018 1 / 65 Overview Definitions and examples Simple estimation
Recap of Part 1 Per Kragh Andersen Section of Biostatistics, University of Copenhagen DSBS Course Survival Analysis in Clinical Trials January 2018 1 / 65 Overview Definitions and examples Simple estimation
Lecture 3. Truncation, length-bias and prevalence sampling
 Lecture 3. Truncation, length-bias and prevalence sampling 3.1 Prevalent sampling Statistical techniques for truncated data have been integrated into survival analysis in last two decades. Truncation in
Lecture 3. Truncation, length-bias and prevalence sampling 3.1 Prevalent sampling Statistical techniques for truncated data have been integrated into survival analysis in last two decades. Truncation in
Multistate models and recurrent event models
 and recurrent event models Patrick Breheny December 6 Patrick Breheny University of Iowa Survival Data Analysis (BIOS:7210) 1 / 22 Introduction In this final lecture, we will briefly look at two other
and recurrent event models Patrick Breheny December 6 Patrick Breheny University of Iowa Survival Data Analysis (BIOS:7210) 1 / 22 Introduction In this final lecture, we will briefly look at two other
Survival Regression Models
 Survival Regression Models David M. Rocke May 18, 2017 David M. Rocke Survival Regression Models May 18, 2017 1 / 32 Background on the Proportional Hazards Model The exponential distribution has constant
Survival Regression Models David M. Rocke May 18, 2017 David M. Rocke Survival Regression Models May 18, 2017 1 / 32 Background on the Proportional Hazards Model The exponential distribution has constant
Multivariate Survival Analysis
 Multivariate Survival Analysis Previously we have assumed that either (X i, δ i ) or (X i, δ i, Z i ), i = 1,..., n, are i.i.d.. This may not always be the case. Multivariate survival data can arise in
Multivariate Survival Analysis Previously we have assumed that either (X i, δ i ) or (X i, δ i, Z i ), i = 1,..., n, are i.i.d.. This may not always be the case. Multivariate survival data can arise in
Estimating transition probabilities for the illness-death model The Aalen-Johansen estimator under violation of the Markov assumption Torunn Heggland
 Estimating transition probabilities for the illness-death model The Aalen-Johansen estimator under violation of the Markov assumption Torunn Heggland Master s Thesis for the degree Modelling and Data Analysis
Estimating transition probabilities for the illness-death model The Aalen-Johansen estimator under violation of the Markov assumption Torunn Heggland Master s Thesis for the degree Modelling and Data Analysis
PhD course in Advanced survival analysis. One-sample tests. Properties. Idea: (ABGK, sect. V.1.1) Counting process N(t)
 PhD course in Advanced survival analysis. (ABGK, sect. V.1.1) One-sample tests. Counting process N(t) Non-parametric hypothesis tests. Parametric models. Intensity process λ(t) = α(t)y (t) satisfying Aalen
PhD course in Advanced survival analysis. (ABGK, sect. V.1.1) One-sample tests. Counting process N(t) Non-parametric hypothesis tests. Parametric models. Intensity process λ(t) = α(t)y (t) satisfying Aalen
Analysis of competing risks data and simulation of data following predened subdistribution hazards
 Analysis of competing risks data and simulation of data following predened subdistribution hazards Bernhard Haller Institut für Medizinische Statistik und Epidemiologie Technische Universität München 27.05.2013
Analysis of competing risks data and simulation of data following predened subdistribution hazards Bernhard Haller Institut für Medizinische Statistik und Epidemiologie Technische Universität München 27.05.2013
Lecture 5 Models and methods for recurrent event data
 Lecture 5 Models and methods for recurrent event data Recurrent and multiple events are commonly encountered in longitudinal studies. In this chapter we consider ordered recurrent and multiple events.
Lecture 5 Models and methods for recurrent event data Recurrent and multiple events are commonly encountered in longitudinal studies. In this chapter we consider ordered recurrent and multiple events.
Other Survival Models. (1) Non-PH models. We briefly discussed the non-proportional hazards (non-ph) model
 Other Survival Models (1) Non-PH models We briefly discussed the non-proportional hazards (non-ph) model λ(t Z) = λ 0 (t) exp{β(t) Z}, where β(t) can be estimated by: piecewise constants (recall how);
Other Survival Models (1) Non-PH models We briefly discussed the non-proportional hazards (non-ph) model λ(t Z) = λ 0 (t) exp{β(t) Z}, where β(t) can be estimated by: piecewise constants (recall how);
Multistate Modelling Vertical Transmission and Determination of R 0 Using Transition Intensities
 Applied Mathematical Sciences, Vol. 9, 2015, no. 79, 3941-3956 HIKARI Ltd, www.m-hikari.com http://dx.doi.org/10.12988/ams.2015.52130 Multistate Modelling Vertical Transmission and Determination of R 0
Applied Mathematical Sciences, Vol. 9, 2015, no. 79, 3941-3956 HIKARI Ltd, www.m-hikari.com http://dx.doi.org/10.12988/ams.2015.52130 Multistate Modelling Vertical Transmission and Determination of R 0
Longitudinal + Reliability = Joint Modeling
 Longitudinal + Reliability = Joint Modeling Carles Serrat Institute of Statistics and Mathematics Applied to Building CYTED-HAROSA International Workshop November 21-22, 2013 Barcelona Mainly from Rizopoulos,
Longitudinal + Reliability = Joint Modeling Carles Serrat Institute of Statistics and Mathematics Applied to Building CYTED-HAROSA International Workshop November 21-22, 2013 Barcelona Mainly from Rizopoulos,
Score test for random changepoint in a mixed model
 Score test for random changepoint in a mixed model Corentin Segalas and Hélène Jacqmin-Gadda INSERM U1219, Biostatistics team, Bordeaux GDR Statistiques et Santé October 6, 2017 Biostatistics 1 / 27 Introduction
Score test for random changepoint in a mixed model Corentin Segalas and Hélène Jacqmin-Gadda INSERM U1219, Biostatistics team, Bordeaux GDR Statistiques et Santé October 6, 2017 Biostatistics 1 / 27 Introduction
Cox s proportional hazards model and Cox s partial likelihood
 Cox s proportional hazards model and Cox s partial likelihood Rasmus Waagepetersen October 12, 2018 1 / 27 Non-parametric vs. parametric Suppose we want to estimate unknown function, e.g. survival function.
Cox s proportional hazards model and Cox s partial likelihood Rasmus Waagepetersen October 12, 2018 1 / 27 Non-parametric vs. parametric Suppose we want to estimate unknown function, e.g. survival function.
Parametric and Non Homogeneous semi-markov Process for HIV Control
 Parametric and Non Homogeneous semi-markov Process for HIV Control Eve Mathieu 1, Yohann Foucher 1, Pierre Dellamonica 2, and Jean-Pierre Daures 1 1 Clinical Research University Institute. Biostatistics
Parametric and Non Homogeneous semi-markov Process for HIV Control Eve Mathieu 1, Yohann Foucher 1, Pierre Dellamonica 2, and Jean-Pierre Daures 1 1 Clinical Research University Institute. Biostatistics
MAS3301 / MAS8311 Biostatistics Part II: Survival
 MAS3301 / MAS8311 Biostatistics Part II: Survival M. Farrow School of Mathematics and Statistics Newcastle University Semester 2, 2009-10 1 13 The Cox proportional hazards model 13.1 Introduction In the
MAS3301 / MAS8311 Biostatistics Part II: Survival M. Farrow School of Mathematics and Statistics Newcastle University Semester 2, 2009-10 1 13 The Cox proportional hazards model 13.1 Introduction In the
A COMPARISON OF POISSON AND BINOMIAL EMPIRICAL LIKELIHOOD Mai Zhou and Hui Fang University of Kentucky
 A COMPARISON OF POISSON AND BINOMIAL EMPIRICAL LIKELIHOOD Mai Zhou and Hui Fang University of Kentucky Empirical likelihood with right censored data were studied by Thomas and Grunkmier (1975), Li (1995),
A COMPARISON OF POISSON AND BINOMIAL EMPIRICAL LIKELIHOOD Mai Zhou and Hui Fang University of Kentucky Empirical likelihood with right censored data were studied by Thomas and Grunkmier (1975), Li (1995),
THESIS for the degree of MASTER OF SCIENCE. Modelling and Data Analysis
 PROPERTIES OF ESTIMATORS FOR RELATIVE RISKS FROM NESTED CASE-CONTROL STUDIES WITH MULTIPLE OUTCOMES (COMPETING RISKS) by NATHALIE C. STØER THESIS for the degree of MASTER OF SCIENCE Modelling and Data
PROPERTIES OF ESTIMATORS FOR RELATIVE RISKS FROM NESTED CASE-CONTROL STUDIES WITH MULTIPLE OUTCOMES (COMPETING RISKS) by NATHALIE C. STØER THESIS for the degree of MASTER OF SCIENCE Modelling and Data
Analysing geoadditive regression data: a mixed model approach
 Analysing geoadditive regression data: a mixed model approach Institut für Statistik, Ludwig-Maximilians-Universität München Joint work with Ludwig Fahrmeir & Stefan Lang 25.11.2005 Spatio-temporal regression
Analysing geoadditive regression data: a mixed model approach Institut für Statistik, Ludwig-Maximilians-Universität München Joint work with Ludwig Fahrmeir & Stefan Lang 25.11.2005 Spatio-temporal regression
Pricing and Risk Analysis of a Long-Term Care Insurance Contract in a non-markov Multi-State Model
 Pricing and Risk Analysis of a Long-Term Care Insurance Contract in a non-markov Multi-State Model Quentin Guibert Univ Lyon, Université Claude Bernard Lyon 1, ISFA, Laboratoire SAF EA2429, F-69366, Lyon,
Pricing and Risk Analysis of a Long-Term Care Insurance Contract in a non-markov Multi-State Model Quentin Guibert Univ Lyon, Université Claude Bernard Lyon 1, ISFA, Laboratoire SAF EA2429, F-69366, Lyon,
Session 9: Introduction to Sieve Analysis of Pathogen Sequences, for Assessing How VE Depends on Pathogen Genomics Part I
 Session 9: Introduction to Sieve Analysis of Pathogen Sequences, for Assessing How VE Depends on Pathogen Genomics Part I Peter B Gilbert Vaccine and Infectious Disease Division, Fred Hutchinson Cancer
Session 9: Introduction to Sieve Analysis of Pathogen Sequences, for Assessing How VE Depends on Pathogen Genomics Part I Peter B Gilbert Vaccine and Infectious Disease Division, Fred Hutchinson Cancer
Parametric multistate survival analysis: New developments
 Parametric multistate survival analysis: New developments Michael J. Crowther Biostatistics Research Group Department of Health Sciences University of Leicester, UK michael.crowther@le.ac.uk Victorian
Parametric multistate survival analysis: New developments Michael J. Crowther Biostatistics Research Group Department of Health Sciences University of Leicester, UK michael.crowther@le.ac.uk Victorian
Journal of Statistical Software
 JSS Journal of Statistical Software January 2011, Volume 38, Issue 2. http://www.jstatsoft.org/ Analyzing Competing Risk Data Using the R timereg Package Thomas H. Scheike University of Copenhagen Mei-Jie
JSS Journal of Statistical Software January 2011, Volume 38, Issue 2. http://www.jstatsoft.org/ Analyzing Competing Risk Data Using the R timereg Package Thomas H. Scheike University of Copenhagen Mei-Jie
Multivariate Survival Data With Censoring.
 1 Multivariate Survival Data With Censoring. Shulamith Gross and Catherine Huber-Carol Baruch College of the City University of New York, Dept of Statistics and CIS, Box 11-220, 1 Baruch way, 10010 NY.
1 Multivariate Survival Data With Censoring. Shulamith Gross and Catherine Huber-Carol Baruch College of the City University of New York, Dept of Statistics and CIS, Box 11-220, 1 Baruch way, 10010 NY.
arxiv: v3 [stat.ap] 29 Jun 2015
![arxiv: v3 [stat.ap] 29 Jun 2015 arxiv: v3 [stat.ap] 29 Jun 2015](/thumbs/79/79513447.jpg) Joint modelling of longitudinal and multi-state processes: application to clinical progressions in prostate cancer Loïc Ferrer 1,, Virginie Rondeau 1, James J. Dignam 2, Tom Pickles 3, Hélène Jacqmin-Gadda
Joint modelling of longitudinal and multi-state processes: application to clinical progressions in prostate cancer Loïc Ferrer 1,, Virginie Rondeau 1, James J. Dignam 2, Tom Pickles 3, Hélène Jacqmin-Gadda
Survival Analysis for Case-Cohort Studies
 Survival Analysis for ase-ohort Studies Petr Klášterecký Dept. of Probability and Mathematical Statistics, Faculty of Mathematics and Physics, harles University, Prague, zech Republic e-mail: petr.klasterecky@matfyz.cz
Survival Analysis for ase-ohort Studies Petr Klášterecký Dept. of Probability and Mathematical Statistics, Faculty of Mathematics and Physics, harles University, Prague, zech Republic e-mail: petr.klasterecky@matfyz.cz
Lecture 6 PREDICTING SURVIVAL UNDER THE PH MODEL
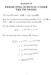 Lecture 6 PREDICTING SURVIVAL UNDER THE PH MODEL The Cox PH model: λ(t Z) = λ 0 (t) exp(β Z). How do we estimate the survival probability, S z (t) = S(t Z) = P (T > t Z), for an individual with covariates
Lecture 6 PREDICTING SURVIVAL UNDER THE PH MODEL The Cox PH model: λ(t Z) = λ 0 (t) exp(β Z). How do we estimate the survival probability, S z (t) = S(t Z) = P (T > t Z), for an individual with covariates
UNIVERSITY OF CALIFORNIA, SAN DIEGO
 UNIVERSITY OF CALIFORNIA, SAN DIEGO Estimation of the primary hazard ratio in the presence of a secondary covariate with non-proportional hazards An undergraduate honors thesis submitted to the Department
UNIVERSITY OF CALIFORNIA, SAN DIEGO Estimation of the primary hazard ratio in the presence of a secondary covariate with non-proportional hazards An undergraduate honors thesis submitted to the Department
Lecture 9. Statistics Survival Analysis. Presented February 23, Dan Gillen Department of Statistics University of California, Irvine
 Statistics 255 - Survival Analysis Presented February 23, 2016 Dan Gillen Department of Statistics University of California, Irvine 9.1 Survival analysis involves subjects moving through time Hazard may
Statistics 255 - Survival Analysis Presented February 23, 2016 Dan Gillen Department of Statistics University of California, Irvine 9.1 Survival analysis involves subjects moving through time Hazard may
REGRESSION ANALYSIS FOR TIME-TO-EVENT DATA THE PROPORTIONAL HAZARDS (COX) MODEL ST520
 REGRESSION ANALYSIS FOR TIME-TO-EVENT DATA THE PROPORTIONAL HAZARDS (COX) MODEL ST520 Department of Statistics North Carolina State University Presented by: Butch Tsiatis, Department of Statistics, NCSU
REGRESSION ANALYSIS FOR TIME-TO-EVENT DATA THE PROPORTIONAL HAZARDS (COX) MODEL ST520 Department of Statistics North Carolina State University Presented by: Butch Tsiatis, Department of Statistics, NCSU
for Time-to-event Data Mei-Ling Ting Lee University of Maryland, College Park
 Threshold Regression for Time-to-event Data Mei-Ling Ting Lee University of Maryland, College Park MLTLEE@UMD.EDU Outline The proportional hazards (PH) model is widely used in analyzing time-to-event data.
Threshold Regression for Time-to-event Data Mei-Ling Ting Lee University of Maryland, College Park MLTLEE@UMD.EDU Outline The proportional hazards (PH) model is widely used in analyzing time-to-event data.
[Part 2] Model Development for the Prediction of Survival Times using Longitudinal Measurements
![[Part 2] Model Development for the Prediction of Survival Times using Longitudinal Measurements [Part 2] Model Development for the Prediction of Survival Times using Longitudinal Measurements](/thumbs/89/99255856.jpg) [Part 2] Model Development for the Prediction of Survival Times using Longitudinal Measurements Aasthaa Bansal PhD Pharmaceutical Outcomes Research & Policy Program University of Washington 69 Biomarkers
[Part 2] Model Development for the Prediction of Survival Times using Longitudinal Measurements Aasthaa Bansal PhD Pharmaceutical Outcomes Research & Policy Program University of Washington 69 Biomarkers
Survival Analysis Math 434 Fall 2011
 Survival Analysis Math 434 Fall 2011 Part IV: Chap. 8,9.2,9.3,11: Semiparametric Proportional Hazards Regression Jimin Ding Math Dept. www.math.wustl.edu/ jmding/math434/fall09/index.html Basic Model Setup
Survival Analysis Math 434 Fall 2011 Part IV: Chap. 8,9.2,9.3,11: Semiparametric Proportional Hazards Regression Jimin Ding Math Dept. www.math.wustl.edu/ jmding/math434/fall09/index.html Basic Model Setup
Statistical Inference and Methods
 Department of Mathematics Imperial College London d.stephens@imperial.ac.uk http://stats.ma.ic.ac.uk/ das01/ 31st January 2006 Part VI Session 6: Filtering and Time to Event Data Session 6: Filtering and
Department of Mathematics Imperial College London d.stephens@imperial.ac.uk http://stats.ma.ic.ac.uk/ das01/ 31st January 2006 Part VI Session 6: Filtering and Time to Event Data Session 6: Filtering and
Statistical Analysis of Competing Risks With Missing Causes of Failure
 Proceedings 59th ISI World Statistics Congress, 25-3 August 213, Hong Kong (Session STS9) p.1223 Statistical Analysis of Competing Risks With Missing Causes of Failure Isha Dewan 1,3 and Uttara V. Naik-Nimbalkar
Proceedings 59th ISI World Statistics Congress, 25-3 August 213, Hong Kong (Session STS9) p.1223 Statistical Analysis of Competing Risks With Missing Causes of Failure Isha Dewan 1,3 and Uttara V. Naik-Nimbalkar
Lecture 7. Proportional Hazards Model - Handling Ties and Survival Estimation Statistics Survival Analysis. Presented February 4, 2016
 Proportional Hazards Model - Handling Ties and Survival Estimation Statistics 255 - Survival Analysis Presented February 4, 2016 likelihood - Discrete Dan Gillen Department of Statistics University of
Proportional Hazards Model - Handling Ties and Survival Estimation Statistics 255 - Survival Analysis Presented February 4, 2016 likelihood - Discrete Dan Gillen Department of Statistics University of
Approximation of Survival Function by Taylor Series for General Partly Interval Censored Data
 Malaysian Journal of Mathematical Sciences 11(3): 33 315 (217) MALAYSIAN JOURNAL OF MATHEMATICAL SCIENCES Journal homepage: http://einspem.upm.edu.my/journal Approximation of Survival Function by Taylor
Malaysian Journal of Mathematical Sciences 11(3): 33 315 (217) MALAYSIAN JOURNAL OF MATHEMATICAL SCIENCES Journal homepage: http://einspem.upm.edu.my/journal Approximation of Survival Function by Taylor
Survival Distributions, Hazard Functions, Cumulative Hazards
 BIO 244: Unit 1 Survival Distributions, Hazard Functions, Cumulative Hazards 1.1 Definitions: The goals of this unit are to introduce notation, discuss ways of probabilistically describing the distribution
BIO 244: Unit 1 Survival Distributions, Hazard Functions, Cumulative Hazards 1.1 Definitions: The goals of this unit are to introduce notation, discuss ways of probabilistically describing the distribution
Survival analysis in R
 Survival analysis in R Niels Richard Hansen This note describes a few elementary aspects of practical analysis of survival data in R. For further information we refer to the book Introductory Statistics
Survival analysis in R Niels Richard Hansen This note describes a few elementary aspects of practical analysis of survival data in R. For further information we refer to the book Introductory Statistics
Chapter 4 Fall Notations: t 1 < t 2 < < t D, D unique death times. d j = # deaths at t j = n. Y j = # at risk /alive at t j = n
 Bios 323: Applied Survival Analysis Qingxia (Cindy) Chen Chapter 4 Fall 2012 4.2 Estimators of the survival and cumulative hazard functions for RC data Suppose X is a continuous random failure time with
Bios 323: Applied Survival Analysis Qingxia (Cindy) Chen Chapter 4 Fall 2012 4.2 Estimators of the survival and cumulative hazard functions for RC data Suppose X is a continuous random failure time with
Modelling geoadditive survival data
 Modelling geoadditive survival data Thomas Kneib & Ludwig Fahrmeir Department of Statistics, Ludwig-Maximilians-University Munich 1. Leukemia survival data 2. Structured hazard regression 3. Mixed model
Modelling geoadditive survival data Thomas Kneib & Ludwig Fahrmeir Department of Statistics, Ludwig-Maximilians-University Munich 1. Leukemia survival data 2. Structured hazard regression 3. Mixed model
Instrumental variables estimation in the Cox Proportional Hazard regression model
 Instrumental variables estimation in the Cox Proportional Hazard regression model James O Malley, Ph.D. Department of Biomedical Data Science The Dartmouth Institute for Health Policy and Clinical Practice
Instrumental variables estimation in the Cox Proportional Hazard regression model James O Malley, Ph.D. Department of Biomedical Data Science The Dartmouth Institute for Health Policy and Clinical Practice
Part III Measures of Classification Accuracy for the Prediction of Survival Times
 Part III Measures of Classification Accuracy for the Prediction of Survival Times Patrick J Heagerty PhD Department of Biostatistics University of Washington 102 ISCB 2010 Session Three Outline Examples
Part III Measures of Classification Accuracy for the Prediction of Survival Times Patrick J Heagerty PhD Department of Biostatistics University of Washington 102 ISCB 2010 Session Three Outline Examples
Multistate models in survival and event history analysis
 Multistate models in survival and event history analysis Dorota M. Dabrowska UCLA November 8, 2011 Research supported by the grant R01 AI067943 from NIAID. The content is solely the responsibility of the
Multistate models in survival and event history analysis Dorota M. Dabrowska UCLA November 8, 2011 Research supported by the grant R01 AI067943 from NIAID. The content is solely the responsibility of the
STAT331. Cox s Proportional Hazards Model
 STAT331 Cox s Proportional Hazards Model In this unit we introduce Cox s proportional hazards (Cox s PH) model, give a heuristic development of the partial likelihood function, and discuss adaptations
STAT331 Cox s Proportional Hazards Model In this unit we introduce Cox s proportional hazards (Cox s PH) model, give a heuristic development of the partial likelihood function, and discuss adaptations
Lecture 22 Survival Analysis: An Introduction
 University of Illinois Department of Economics Spring 2017 Econ 574 Roger Koenker Lecture 22 Survival Analysis: An Introduction There is considerable interest among economists in models of durations, which
University of Illinois Department of Economics Spring 2017 Econ 574 Roger Koenker Lecture 22 Survival Analysis: An Introduction There is considerable interest among economists in models of durations, which
Log-linearity for Cox s regression model. Thesis for the Degree Master of Science
 Log-linearity for Cox s regression model Thesis for the Degree Master of Science Zaki Amini Master s Thesis, Spring 2015 i Abstract Cox s regression model is one of the most applied methods in medical
Log-linearity for Cox s regression model Thesis for the Degree Master of Science Zaki Amini Master s Thesis, Spring 2015 i Abstract Cox s regression model is one of the most applied methods in medical
I N S T I T U T D E S T A T I S T I Q U E B I O S T A T I S T I Q U E E T S C I E N C E S A C T U A R I E L L E S (I S B A)
 I N S T I T U T D E S T A T I S T I Q U E B I O S T A T I S T I Q U E E T S C I E N C E S A C T U A R I E L L E S (I S B A) UNIVERSITÉ CATHOLIQUE DE LOUVAIN D I S C U S S I O N P A P E R 153 ESTIMATION
I N S T I T U T D E S T A T I S T I Q U E B I O S T A T I S T I Q U E E T S C I E N C E S A C T U A R I E L L E S (I S B A) UNIVERSITÉ CATHOLIQUE DE LOUVAIN D I S C U S S I O N P A P E R 153 ESTIMATION
A Multistate Model for Bivariate Interval Censored Failure Time Data
 A Multistate Model for Bivariate Interval Censored Failure Time Data 1, Richard J. Cook, Leilei Zeng 2 and Ker-Ai Lee 1 1 Department of Statistics and Actuarial Science, University of Waterloo, Waterloo,
A Multistate Model for Bivariate Interval Censored Failure Time Data 1, Richard J. Cook, Leilei Zeng 2 and Ker-Ai Lee 1 1 Department of Statistics and Actuarial Science, University of Waterloo, Waterloo,
Estimation of Conditional Kendall s Tau for Bivariate Interval Censored Data
 Communications for Statistical Applications and Methods 2015, Vol. 22, No. 6, 599 604 DOI: http://dx.doi.org/10.5351/csam.2015.22.6.599 Print ISSN 2287-7843 / Online ISSN 2383-4757 Estimation of Conditional
Communications for Statistical Applications and Methods 2015, Vol. 22, No. 6, 599 604 DOI: http://dx.doi.org/10.5351/csam.2015.22.6.599 Print ISSN 2287-7843 / Online ISSN 2383-4757 Estimation of Conditional
CONTINUOUS TIME MULTI-STATE MODELS FOR SURVIVAL ANALYSIS. Erika Lynn Gibson
 CONTINUOUS TIME MULTI-STATE MODELS FOR SURVIVAL ANALYSIS By Erika Lynn Gibson A Thesis Submitted to the Graduate Faculty of WAKE FOREST UNIVERSITY in Partial Fulfillment of the Requirements for the Degree
CONTINUOUS TIME MULTI-STATE MODELS FOR SURVIVAL ANALYSIS By Erika Lynn Gibson A Thesis Submitted to the Graduate Faculty of WAKE FOREST UNIVERSITY in Partial Fulfillment of the Requirements for the Degree
Survival Analysis. Stat 526. April 13, 2018
 Survival Analysis Stat 526 April 13, 2018 1 Functions of Survival Time Let T be the survival time for a subject Then P [T < 0] = 0 and T is a continuous random variable The Survival function is defined
Survival Analysis Stat 526 April 13, 2018 1 Functions of Survival Time Let T be the survival time for a subject Then P [T < 0] = 0 and T is a continuous random variable The Survival function is defined
FULL LIKELIHOOD INFERENCES IN THE COX MODEL
 October 20, 2007 FULL LIKELIHOOD INFERENCES IN THE COX MODEL BY JIAN-JIAN REN 1 AND MAI ZHOU 2 University of Central Florida and University of Kentucky Abstract We use the empirical likelihood approach
October 20, 2007 FULL LIKELIHOOD INFERENCES IN THE COX MODEL BY JIAN-JIAN REN 1 AND MAI ZHOU 2 University of Central Florida and University of Kentucky Abstract We use the empirical likelihood approach
Stochastic Modelling Unit 1: Markov chain models
 Stochastic Modelling Unit 1: Markov chain models Russell Gerrard and Douglas Wright Cass Business School, City University, London June 2004 Contents of Unit 1 1 Stochastic Processes 2 Markov Chains 3 Poisson
Stochastic Modelling Unit 1: Markov chain models Russell Gerrard and Douglas Wright Cass Business School, City University, London June 2004 Contents of Unit 1 1 Stochastic Processes 2 Markov Chains 3 Poisson
STAT Sample Problem: General Asymptotic Results
 STAT331 1-Sample Problem: General Asymptotic Results In this unit we will consider the 1-sample problem and prove the consistency and asymptotic normality of the Nelson-Aalen estimator of the cumulative
STAT331 1-Sample Problem: General Asymptotic Results In this unit we will consider the 1-sample problem and prove the consistency and asymptotic normality of the Nelson-Aalen estimator of the cumulative
Direct likelihood inference on the cause-specific cumulative incidence function: a flexible parametric regression modelling approach
 Direct likelihood inference on the cause-specific cumulative incidence function: a flexible parametric regression modelling approach Sarwar I Mozumder 1, Mark J Rutherford 1, Paul C Lambert 1,2 1 Biostatistics
Direct likelihood inference on the cause-specific cumulative incidence function: a flexible parametric regression modelling approach Sarwar I Mozumder 1, Mark J Rutherford 1, Paul C Lambert 1,2 1 Biostatistics
Censoring and Truncation - Highlighting the Differences
 Censoring and Truncation - Highlighting the Differences Micha Mandel The Hebrew University of Jerusalem, Jerusalem, Israel, 91905 July 9, 2007 Micha Mandel is a Lecturer, Department of Statistics, The
Censoring and Truncation - Highlighting the Differences Micha Mandel The Hebrew University of Jerusalem, Jerusalem, Israel, 91905 July 9, 2007 Micha Mandel is a Lecturer, Department of Statistics, The
Logistic regression model for survival time analysis using time-varying coefficients
 Logistic regression model for survival time analysis using time-varying coefficients Accepted in American Journal of Mathematical and Management Sciences, 2016 Kenichi SATOH ksatoh@hiroshima-u.ac.jp Research
Logistic regression model for survival time analysis using time-varying coefficients Accepted in American Journal of Mathematical and Management Sciences, 2016 Kenichi SATOH ksatoh@hiroshima-u.ac.jp Research
CASE STUDY: Bayesian Incidence Analyses from Cross-Sectional Data with Multiple Markers of Disease Severity. Outline:
 CASE STUDY: Bayesian Incidence Analyses from Cross-Sectional Data with Multiple Markers of Disease Severity Outline: 1. NIEHS Uterine Fibroid Study Design of Study Scientific Questions Difficulties 2.
CASE STUDY: Bayesian Incidence Analyses from Cross-Sectional Data with Multiple Markers of Disease Severity Outline: 1. NIEHS Uterine Fibroid Study Design of Study Scientific Questions Difficulties 2.
Introduction to cthmm (Continuous-time hidden Markov models) package
 Introduction to cthmm (Continuous-time hidden Markov models) package Abstract A disease process refers to a patient s traversal over time through a disease with multiple discrete states. Multistate models
Introduction to cthmm (Continuous-time hidden Markov models) package Abstract A disease process refers to a patient s traversal over time through a disease with multiple discrete states. Multistate models
Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features. Yangxin Huang
 Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features Yangxin Huang Department of Epidemiology and Biostatistics, COPH, USF, Tampa, FL yhuang@health.usf.edu January
Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features Yangxin Huang Department of Epidemiology and Biostatistics, COPH, USF, Tampa, FL yhuang@health.usf.edu January
You know I m not goin diss you on the internet Cause my mama taught me better than that I m a survivor (What?) I m not goin give up (What?
 You know I m not goin diss you on the internet Cause my mama taught me better than that I m a survivor (What?) I m not goin give up (What?) I m not goin stop (What?) I m goin work harder (What?) Sir David
You know I m not goin diss you on the internet Cause my mama taught me better than that I m a survivor (What?) I m not goin give up (What?) I m not goin stop (What?) I m goin work harder (What?) Sir David
Frailty Modeling for clustered survival data: a simulation study
 Frailty Modeling for clustered survival data: a simulation study IAA Oslo 2015 Souad ROMDHANE LaREMFiQ - IHEC University of Sousse (Tunisia) souad_romdhane@yahoo.fr Lotfi BELKACEM LaREMFiQ - IHEC University
Frailty Modeling for clustered survival data: a simulation study IAA Oslo 2015 Souad ROMDHANE LaREMFiQ - IHEC University of Sousse (Tunisia) souad_romdhane@yahoo.fr Lotfi BELKACEM LaREMFiQ - IHEC University
Survival analysis in R
 Survival analysis in R Niels Richard Hansen This note describes a few elementary aspects of practical analysis of survival data in R. For further information we refer to the book Introductory Statistics
Survival analysis in R Niels Richard Hansen This note describes a few elementary aspects of practical analysis of survival data in R. For further information we refer to the book Introductory Statistics
Introduction to mtm: An R Package for Marginalized Transition Models
 Introduction to mtm: An R Package for Marginalized Transition Models Bryan A. Comstock and Patrick J. Heagerty Department of Biostatistics University of Washington 1 Introduction Marginalized transition
Introduction to mtm: An R Package for Marginalized Transition Models Bryan A. Comstock and Patrick J. Heagerty Department of Biostatistics University of Washington 1 Introduction Marginalized transition
University of California, Berkeley
 University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 24 Paper 153 A Note on Empirical Likelihood Inference of Residual Life Regression Ying Qing Chen Yichuan
University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 24 Paper 153 A Note on Empirical Likelihood Inference of Residual Life Regression Ying Qing Chen Yichuan
Survival Analysis. Lu Tian and Richard Olshen Stanford University
 1 Survival Analysis Lu Tian and Richard Olshen Stanford University 2 Survival Time/ Failure Time/Event Time We will introduce various statistical methods for analyzing survival outcomes What is the survival
1 Survival Analysis Lu Tian and Richard Olshen Stanford University 2 Survival Time/ Failure Time/Event Time We will introduce various statistical methods for analyzing survival outcomes What is the survival
Subject CT4 Models. October 2015 Examination INDICATIVE SOLUTION
 Institute of Actuaries of India Subject CT4 Models October 2015 Examination INDICATIVE SOLUTION Introduction The indicative solution has been written by the Examiners with the aim of helping candidates.
Institute of Actuaries of India Subject CT4 Models October 2015 Examination INDICATIVE SOLUTION Introduction The indicative solution has been written by the Examiners with the aim of helping candidates.
Statistical Methods for Alzheimer s Disease Studies
 Statistical Methods for Alzheimer s Disease Studies Rebecca A. Betensky, Ph.D. Department of Biostatistics, Harvard T.H. Chan School of Public Health July 19, 2016 1/37 OUTLINE 1 Statistical collaborations
Statistical Methods for Alzheimer s Disease Studies Rebecca A. Betensky, Ph.D. Department of Biostatistics, Harvard T.H. Chan School of Public Health July 19, 2016 1/37 OUTLINE 1 Statistical collaborations
Application of Time-to-Event Methods in the Assessment of Safety in Clinical Trials
 Application of Time-to-Event Methods in the Assessment of Safety in Clinical Trials Progress, Updates, Problems William Jen Hoe Koh May 9, 2013 Overview Marginal vs Conditional What is TMLE? Key Estimation
Application of Time-to-Event Methods in the Assessment of Safety in Clinical Trials Progress, Updates, Problems William Jen Hoe Koh May 9, 2013 Overview Marginal vs Conditional What is TMLE? Key Estimation
Typical Survival Data Arising From a Clinical Trial. Censoring. The Survivor Function. Mathematical Definitions Introduction
 Outline CHL 5225H Advanced Statistical Methods for Clinical Trials: Survival Analysis Prof. Kevin E. Thorpe Defining Survival Data Mathematical Definitions Non-parametric Estimates of Survival Comparing
Outline CHL 5225H Advanced Statistical Methods for Clinical Trials: Survival Analysis Prof. Kevin E. Thorpe Defining Survival Data Mathematical Definitions Non-parametric Estimates of Survival Comparing
A general mixed model approach for spatio-temporal regression data
 A general mixed model approach for spatio-temporal regression data Thomas Kneib, Ludwig Fahrmeir & Stefan Lang Department of Statistics, Ludwig-Maximilians-University Munich 1. Spatio-temporal regression
A general mixed model approach for spatio-temporal regression data Thomas Kneib, Ludwig Fahrmeir & Stefan Lang Department of Statistics, Ludwig-Maximilians-University Munich 1. Spatio-temporal regression
Lecture 11. Interval Censored and. Discrete-Time Data. Statistics Survival Analysis. Presented March 3, 2016
 Statistics 255 - Survival Analysis Presented March 3, 2016 Motivating Dan Gillen Department of Statistics University of California, Irvine 11.1 First question: Are the data truly discrete? : Number of
Statistics 255 - Survival Analysis Presented March 3, 2016 Motivating Dan Gillen Department of Statistics University of California, Irvine 11.1 First question: Are the data truly discrete? : Number of
Part IV Extensions: Competing Risks Endpoints and Non-Parametric AUC(t) Estimation
 Part IV Extensions: Competing Risks Endpoints and Non-Parametric AUC(t) Estimation Patrick J. Heagerty PhD Department of Biostatistics University of Washington 166 ISCB 2010 Session Four Outline Examples
Part IV Extensions: Competing Risks Endpoints and Non-Parametric AUC(t) Estimation Patrick J. Heagerty PhD Department of Biostatistics University of Washington 166 ISCB 2010 Session Four Outline Examples
Part III. Hypothesis Testing. III.1. Log-rank Test for Right-censored Failure Time Data
 1 Part III. Hypothesis Testing III.1. Log-rank Test for Right-censored Failure Time Data Consider a survival study consisting of n independent subjects from p different populations with survival functions
1 Part III. Hypothesis Testing III.1. Log-rank Test for Right-censored Failure Time Data Consider a survival study consisting of n independent subjects from p different populations with survival functions
Maximum likelihood estimation of a log-concave density based on censored data
 Maximum likelihood estimation of a log-concave density based on censored data Dominic Schuhmacher Institute of Mathematical Statistics and Actuarial Science University of Bern Joint work with Lutz Dümbgen
Maximum likelihood estimation of a log-concave density based on censored data Dominic Schuhmacher Institute of Mathematical Statistics and Actuarial Science University of Bern Joint work with Lutz Dümbgen
A GENERALIZED ADDITIVE REGRESSION MODEL FOR SURVIVAL TIMES 1. By Thomas H. Scheike University of Copenhagen
 The Annals of Statistics 21, Vol. 29, No. 5, 1344 136 A GENERALIZED ADDITIVE REGRESSION MODEL FOR SURVIVAL TIMES 1 By Thomas H. Scheike University of Copenhagen We present a non-parametric survival model
The Annals of Statistics 21, Vol. 29, No. 5, 1344 136 A GENERALIZED ADDITIVE REGRESSION MODEL FOR SURVIVAL TIMES 1 By Thomas H. Scheike University of Copenhagen We present a non-parametric survival model
Exercises. (a) Prove that m(t) =
 Exercises 1. Lack of memory. Verify that the exponential distribution has the lack of memory property, that is, if T is exponentially distributed with parameter λ > then so is T t given that T > t for
Exercises 1. Lack of memory. Verify that the exponential distribution has the lack of memory property, that is, if T is exponentially distributed with parameter λ > then so is T t given that T > t for
PENALIZED LIKELIHOOD PARAMETER ESTIMATION FOR ADDITIVE HAZARD MODELS WITH INTERVAL CENSORED DATA
 PENALIZED LIKELIHOOD PARAMETER ESTIMATION FOR ADDITIVE HAZARD MODELS WITH INTERVAL CENSORED DATA Kasun Rathnayake ; A/Prof Jun Ma Department of Statistics Faculty of Science and Engineering Macquarie University
PENALIZED LIKELIHOOD PARAMETER ESTIMATION FOR ADDITIVE HAZARD MODELS WITH INTERVAL CENSORED DATA Kasun Rathnayake ; A/Prof Jun Ma Department of Statistics Faculty of Science and Engineering Macquarie University
Frailty Models and Copulas: Similarities and Differences
 Frailty Models and Copulas: Similarities and Differences KLARA GOETHALS, PAUL JANSSEN & LUC DUCHATEAU Department of Physiology and Biometrics, Ghent University, Belgium; Center for Statistics, Hasselt
Frailty Models and Copulas: Similarities and Differences KLARA GOETHALS, PAUL JANSSEN & LUC DUCHATEAU Department of Physiology and Biometrics, Ghent University, Belgium; Center for Statistics, Hasselt
Stochastic modelling of epidemic spread
 Stochastic modelling of epidemic spread Julien Arino Centre for Research on Inner City Health St Michael s Hospital Toronto On leave from Department of Mathematics University of Manitoba Julien Arino@umanitoba.ca
Stochastic modelling of epidemic spread Julien Arino Centre for Research on Inner City Health St Michael s Hospital Toronto On leave from Department of Mathematics University of Manitoba Julien Arino@umanitoba.ca
Multistate models STK4080 H Competing risk setting 2. Illness-Death setting 3. General event histories and Aalen-Johansen estimator
 Multistate models p. 1/36 Multistate models STK4080 H16 1. Competing risk setting 2. Illness-Death setting 3. General event histories and Aalen-Johansen estimator Multistate models p. 2/36 Multistate models
Multistate models p. 1/36 Multistate models STK4080 H16 1. Competing risk setting 2. Illness-Death setting 3. General event histories and Aalen-Johansen estimator Multistate models p. 2/36 Multistate models
