CE 590 Applied Bayesian Statistics. Mid-term Take-home Exam
|
|
|
- Grace Alyson Sutton
- 5 years ago
- Views:
Transcription
1 CE 590 Applied Bayesian Statistics Mid-term Take-home Exam Due: April 1, 2015
2 ST495/590 Mid-term take-home exam Due 4/1 This portion of the exam is take-home and must be dropped in my office by 5PM on Wednesday, April 1. THIS IS AN EXAM - YOU MAY NOT DISCUSS THE PROBLEMS WITH ANYONE (INCLUDING OTHER STUDENTS OR THE TA)! If you have questions please visit office hours or me. Data for this analysis were downloaded from For each National Basketball Association team and each season from , the data set includes several statistics describing the team s performance that season. Your objective is to build a predictive model for margin of victory in terms of the other variables. Variables are described on the back of the exam, and available for download at Use the data from to fit the model, and 2014 to test predictions. independence across teams and years. You may assume 1. Fit at least 2-3 different models to the data and select a final model. 2. Verify that the MCMC algorithm is producing reliable output for your final model. 3. Determine which variables in your final model are statistically significant. 4. Are the results sensitive to the prior? 5. Make predictions from your final model for each team in 2014, and plot the posterior predictive distributions versus actual 2014 data for the final model. Summarize prediction accuracy in terms of mean squared error and give the coverage of the prediction intervals. Turn in a report summarizing this analysis. The report should be no more than 4 pages (11 font, 1-inch margins) and should be in manuscript style with paragraphs of text and numbered figures and tables. A substantial portion of the grade will be based on clarity of presentation. You should describe in the text the methods you are using in enough detail that the analysis could be replicated by another student in the class. Attach commented code in a separate report. Staple all material for the exam together. HAVE FUN!
3 VARIABLE DESCRIPTIONS FOR THE NBA DATASET 1. Team: Team 2. Year: Year 3. Conference: East/West Conference 4. Division: Division within conference 5. MarginOfVictory: Average margin of victory on the season 6. AverageAge: Average age of players on the team 7. StrengthOfSchedule: Strength of schedule (positive means you played good teams) 8. Pace: Pace of play (possessions per game) 9. FreeThrowAttemptRate: Number of free throw attempts divided by number of field goal attempts 10. 3PointAttemptRate: Number of three point attempts divided by number of field goal attempts 11. TrueShootingPCT: Percentage of shots made (accounting for two verses three pointers) 12. TurnoverPCT: Turnovers per 100 plays 13. OffensiveReboundPCT: Percent of missed shots reclaimed on the rebound 14. ThreePointPCT: Three-point percentage 15. FreeThrowPCT: Free-throw percentage 16. Opp3pointPCT: Opponent s three-point percentage 17. oppfreethrowpct: Opponent s free-throw percentage 18. OffAveFGDist: Average distance of shot attempt 19. Off2PAssd: Percentage of made two-point shots that resulted from an assist 20. Dunks: Number of dunks 21. Off3PAssd: Percentage of made three-point shots that resulted from an assist 22. DefAveFGDist: Average distance of opponent s shots 23. Def2PAssd: Percentage of opponent s made two-point shots that resulted from an assist 24. Def3PAssd: Percentage of opponent s made three-point shots that resulted from an assist In this regression, use margin of victory as the response and variables 6-24 as predictors. For more information about these variables, see
4 M - 1 (ANSWER) 1. In this mid-term exam, the multiple linear regression analysis has been performed to fit the model based on NBA data and test the prediction using 2014 NBA data. The independence across teams and years is assumed. The margin of victory is defined as response (Y i ) and 19 variables from AverageAge (X i1 ) to Def3PAssd (X i19 ) of data are used as predictors. The basic model equation is represented in Equation (1) for i = 1~n observations. Since NBA teams are 30 and 4 seasons, n is 120 observations to fit the model. 2 Y i ~ Normal Xi 11 X i1919, (1) Here, all variables are centered and scaled. The priors σ 2 ~ InvGamma (0.01, 0.01) and α ~ Normal (0, ) are assumed for the error variance and intercept, respectively. And for the regression coefficients β j, four different prior models of Table 1 are selected, evaluated and compared in order to choose the best model. Specifically, for the best model selection, the mean squared error (MSE), Bias, average standard deviation (AVESD) and coverage of 95% prediction interval (COV) for the cross validation, and the deviance information criteria (DIC) for these models are calculated and represented in Table 2. As a result of cross validation, the Cauchy prior has the smallest prediction MSE and the best coverage out of four models. Regarding DIC result, the Gaussian 2 prior has the smallest DIC. Based on these results, the Cauchy prior or Gaussian 2 can be the best possible models. In this study, Gaussian 2 is set to be a final model. The detailed calculation is described in Appendix 1 and For checking whether the posterior results are sensitive to these four different priors, the mean, standard deviation (SD), 95% confidence interval (CI) are represented in Table 3. The distribution of the posterior of regression coefficients β 7 and β 13 as the representative examples (i.e. not sensitive and sensitive case) are illustrated in Figure 1. Overall, the posterior results seem not sensitive to 4 different priors. However, there are some changes about 8 regression coefficients of FreeThrowAttemptRate (β 4 ), 3PointAttemptRate (β 5 ), TurnoverPCT (β 6 ), ThreePointPCT (β 9 ), Opp3pointPCT (β 11 ), oppfreethrowpct (β 12 ), OffAveFGDist (β 13 ) and Dunks (β 15 ) in terms of 4 different priors. The detailed calculation is described in Appendix Since variables are deemed statistically significant if their 95% CI exclude zero, total 12 variables corresponding to regression coefficients such as β 1, β 2, β 6, β 7, β 8, β 9, β 11, β 12, β 13, β 15, β 17, and β 18 are to be statistically significant as shown in Table 4. Table 1. Four different priors of regression coefficients β j Gaussian 1 Gaussian 2 Cauchy Bayesian LASSO j ~ Normal 0, j ~ Normal 0, b j ~ t1 0, b j ~ DoubleEx 0, b 2 2 InvG InvG InvG b ~.,. b ~.,. Table 2. Cross validation and DIC Cross validation MSE BIAS AVESD COV b ~.,. DIC (penalized deviance) Gaussian Gaussian Cauchy Bayes LASSO
5 M - 2 Table 3. Posterior summary of regression coefficients β j with 4 different priors Gaussian 1 Gaussian 2 Cauchy Bayesian LASSO Mean SD 95% 95% 95% 95% Mean SD Mean SD Mean SD CI CI CI CI β β β β β β β β β β β β β β β β β β β Table 4. Posterior 95% confidence interval of regression coefficients β j with Gaussian 2 prior CL β 1 β 2 β 3 β 4 β 5 β 6 β 7 β 8 β 9 β % % CL β 11 β 12 β 13 β 14 β 15 β 16 β 17 β 18 β % % (a) β 7 (b) β 13 Figure 1. Posterior distributions of regression coefficients β 7 and β 13 with 4 different priors
6 M - 3 (ANSWER) 2. In order to verify that the MCMC algorithm is producing reliable output using the final model (Gaussian 2 prior as above mentioned), the convergence test for σ, σ b and β j is performed. Using MCMC algorithm of JAGS, samples have been drawn. 10,000 warm up samples are drawn by using update function. These are the burn-in samples. The 20,000 samples are more produced to approximate the posterior by using number of iteration in coda.samples. For the more thorough convergence test, the three chains are used by using n.chains of jags.model. The detailed code is represented in Appendix 4. Specifically, the trace of the parameters, the auto-correlation function (ACF), and the Gelman stat ( ˆR ) are produced, and ACF and ˆR of σ, σb, β 6 and β 13 are only represented in the Figure 2. The effective sample size (ESS) of all parameters is evaluated in Table 5. Table 5. Effective sample size (ESS) σ σ b β 1 β 2 β 3 β 4 β 5 β 6 β 7 β 8 β β 10 β 11 β 12 β 13 β 14 β 15 β 16 β 17 β 18 β As a result, if we see the ACF of β 6 (the worst cases based on ESS), the samples of a chain are being uncorrelated as lag is going on. The effective sample size of σ, σ b and β j are huge and greater than Based on Gelman stat graph, the Gelman stat of samples of β 6 is close to one after 14,000 samples. Therefore, the samplings for all parameters are converged, which leads to the reliable output. Figure 2. Auto-correlation function and Gelman stat of parameters of final model (ANSWER) 5. The predictions using the final model for each team in 2014 are summarized in Table 6. Since NBA teams are 30 and 1 seasons, n is 30 observations to test the model. The predictive posterior distributions (PPD) of Y 14 (Memphis) and Y 21 (Oklahoma City) are representatively illustrated with the actual or true 2014 data and plug-in distributions in Figure 3. The prediction accuracy of each team using the final model is described by using mean squared error (MSE), Bias, average standard deviation (AVESD), coverage of 95% prediction intervals (COV). Based on these results, it is observed that this model can predict the response (the margin of victory) of each team in 2014 season quite well. The detailed code is represented in Appendix 5.
7 M - 4 Table 6. Predictive posterior summary of each team Y i using final model Y i Mean SD 2.5% Q 97.5% Q 95% CI 1 Atlanta Boston Charlotte Chicago Cleveland Dallas Denver Detroit Golden State Houston Indiana LA Clippers LA Lakers Memphis Miami Milwaukee Minnesota New Jersey New Orleans New York Oklahoma City Orlando Philadelphia Phoenix Portland Sacramento San Antonio Toronto Utah Washington Table 7. Prediction accuracy using final model MSE BIAS AVESD COV (a) Y 14 : Memphis (b) Y 21 Figure 3. Predictive posterior distributions with the actual 2014 data for Y14 and Y21
8 A - 1 Appendix 1. Model Selection via Cross-validation # # # Model Selection via Cross-validation # # rm(list=ls()) ## Load and standardize NBA data dat <- read.csv(" Y <- dat[,6] Y <- (Y-mean(Y))/sd(Y) X <- dat[,7:25] X <- scale(x) # : observed data, 2014: test or prediction data obs <- dat[,3]!= 2014 prd <- dat[,3] == 2014 Yo <- Y[obs] Xo <- X[obs,] Yp <- Y[prd] Xp <- X[prd,] no <- length(yo) np <- length(yp) p <- ncol(xo) ## Fit the linear regression model # (1) Gaussian: beta_j ~ Normal(0,100^2) model_string1 <- "model{ for(i in 1:no){ Yo[i] ~ dnorm(muo[i],inv.var) muo[i] <- alpha + inprod(xo[i,],beta[]) # Prediction for(i in 1:np){ Yp[i] ~ dnorm(mup[i],inv.var) mup[i] <- alpha + inprod(xp[i,],beta[]) beta[j] ~ dnorm(0,0.0001) " # (2) Gaussian: beta_j ~ Normal(0,sigmab^2) & sigmab^2 ~ InvGamma(0.01,0.01) model_string2 <- "model{ for(i in 1:no){
9 A - 2 Yo[i] ~ dnorm(muo[i],inv.var) muo[i] <- alpha + inprod(xo[i,],beta[]) # Prediction for(i in 1:np){ Yp[i] ~ dnorm(mup[i],inv.var) mup[i] <- alpha + inprod(xp[i,],beta[]) beta[j] ~ dnorm(0,inv.var.b) " # (3) Cauchy: beta_j ~ t1(0,sigmab^2) & sigmab^2 ~ InvGamma(0.01,0.01) model_string3 <- "model{ for(i in 1:no){ Yo[i] ~ dnorm(muo[i],inv.var) muo[i] <- alpha + inprod(xo[i,],beta[]) # Prediction for(i in 1:np){ Yp[i] ~ dnorm(mup[i],inv.var) mup[i] <- alpha + inprod(xp[i,],beta[]) beta[j] ~ dt(0,inv.var.b,1) " # (4) Bayesian LASSO: beta_j ~ DoubleExpo(0,sigmab^2) & sigmab^2 ~ InvGamma(0.01,0.01) model_string4 <- "model{ for(i in 1:no){ Yo[i] ~ dnorm(muo[i],inv.var) muo[i] <- alpha + inprod(xo[i,],beta[]) # Prediction for(i in 1:np){ Yp[i] ~ dnorm(mup[i],inv.var) mup[i] <- alpha + inprod(xp[i,],beta[])
10 A - 3 beta[j] ~ ddexp(0,inv.var.b) " # Fit the model library(rjags) model1 <- jags.model(textconnection(model_string1), data = list(yo=yo,no=no,np=np,p=p,xo=xo,xp=xp)) update(model1, 10000, progress.bar="none") samps1 <- coda.samples(model1, variable.names=c("yp"), Yp1 <- samps1[[1]] model2 <- jags.model(textconnection(model_string2), data = list(yo=yo,no=no,np=np,p=p,xo=xo,xp=xp)) update(model2, 10000, progress.bar="none") samps2 <- coda.samples(model2, variable.names=c("yp"), Yp2 <- samps2[[1]] model3 <- jags.model(textconnection(model_string3), data = list(yo=yo,no=no,np=np,p=p,xo=xo,xp=xp)) update(model3, 10000, progress.bar="none") samps3 <- coda.samples(model3, variable.names=c("yp"), Yp3 <- samps3[[1]] model4 <- jags.model(textconnection(model_string4), data = list(yo=yo,no=no,np=np,p=p,xo=xo,xp=xp)) update(model4, 10000, progress.bar="none") samps4 <- coda.samples(model4, variable.names=c("yp"), Yp4 <- samps4[[1]] ## Compile the results post_mn1 <- apply(yp1,2,mean) post_sd1 <- apply(yp1,2,sd) post_low1 <- apply(yp1,2,quantile,0.025) post_high1 <- apply(yp1,2,quantile,0.975) post_mn2 <- apply(yp2,2,mean) post_sd2 <- apply(yp2,2,sd) post_low2 <- apply(yp2,2,quantile,0.025) post_high2 <- apply(yp2,2,quantile,0.975) post_mn3 post_sd3 <- apply(yp3,2,mean) <- apply(yp3,2,sd)
11 A - 4 post_low3 <- apply(yp3,2,quantile,0.025) post_high3 <- apply(yp3,2,quantile,0.975) post_mn4 <- apply(yp4,2,mean) post_sd4 <- apply(yp4,2,sd) post_low4 <- apply(yp4,2,quantile,0.025) post_high4 <- apply(yp4,2,quantile,0.975) MSE1 <- mean((post_mn1-yp)^2) BIAS1 <- mean(post_mn1-yp) AVESD1 <- mean(post_sd1) COV1 <- mean(yp>post_low1 & Yp<post_high1) MSE2 <- mean((post_mn2-yp)^2) BIAS2 <- mean(post_mn2-yp) AVESD2 <- mean(post_sd2) COV2 <- mean(yp>post_low2 & Yp<post_high2) MSE3 <- mean((post_mn3-yp)^2) BIAS3 <- mean(post_mn3-yp) AVESD3 <- mean(post_sd3) COV3 <- mean(yp>post_low3 & Yp<post_high3) MSE4 <- mean((post_mn4-yp)^2) BIAS4 <- mean(post_mn4-yp) AVESD4 <- mean(post_sd4) COV4 <- mean(yp>post_low4 & Yp<post_high4) MSE BIAS AVESD COV <- c(mse1,mse2,mse3,mse4) <- c(bias1,bias2,bias3,bias4) <- c(avesd1,avesd2,avesd3,avesd4) <- c(cov1,cov2,cov3,cov4) OUTPUT <- cbind(mse,bias,avesd,cov) rownames(output) <- c("gaussian1","gaussian2","cauchy","blasso") as.table(output,digits=2)
12 A - 5 Appendix 2. Model Selection via DIC # # # Model Selection via DIC # # rm(list=ls()) ## Load and standardize NBA data dat <- read.csv(" Y <- dat[,6] Y <- (Y-mean(Y))/sd(Y) X <- dat[,7:25] X <- scale(x) # only for NBA data obs <- dat[,3]!= 2014 Y <- Y[obs] X <- X[obs,] n <- length(y) p <- ncol(x) ## Fit the linear regression model # (1) Gaussian: beta_j ~ Normal(0,100^2) model_string1 <- "model{ for(i in 1:n){ Y[i] ~ dnorm(muo[i],inv.var) muo[i] <- alpha + inprod(x[i,],beta[]) beta[j] ~ dnorm(0,0.0001) # " # (2) Gaussian: beta_j ~ Normal(0,sigmab^2) & sigmab^2 ~ InvGamma(0.01,0.01) model_string2 <- "model{ for(i in 1:n){ Y[i] ~ dnorm(muo[i],inv.var) muo[i] <- alpha + inprod(x[i,],beta[]) beta[j] ~ dnorm(0,inv.var.b)
13 A - 6 " # (3) Cauchy: beta_j ~ t1(0,sigmab^2) & sigmab^2 ~ InvGamma(0.01,0.01) model_string3 <- "model{ for(i in 1:n){ Y[i] ~ dnorm(muo[i],inv.var) muo[i] <- alpha + inprod(x[i,],beta[]) beta[j] ~ dt(0,inv.var.b,1) " # (4) Bayesian LASSO: beta_j ~ DoubleExpo(0,sigmab^2) & sigmab^2 ~ InvGamma(0.01,0.01) model_string4 <- "model{ for(i in 1:n){ Y[i] ~ dnorm(muo[i],inv.var) muo[i] <- alpha + inprod(x[i,],beta[]) beta[j] ~ ddexp(0,inv.var.b) " # Fit the model library(rjags) model1 <- jags.model(textconnection(model_string1), data = list(y=y,n=n,x=x,p=p),n.chains=3) update(model1, 10000) dic1 <- dic.samples(model1, variable.names=c("beta"), model2 <- jags.model(textconnection(model_string2), data = list(y=y,n=n,x=x,p=p),n.chains=3) update(model2, 10000) dic2 <- dic.samples(model2, variable.names=c("beta"), model3 <- jags.model(textconnection(model_string3),
14 A - 7 data = list(y=y,n=n,x=x,p=p),n.chains=3) update(model3, 10000) dic3 <- dic.samples(model3, variable.names=c("beta"), model4 <- jags.model(textconnection(model_string4), data = list(y=y,n=n,x=x,p=p),n.chains=3) update(model4, 10000) dic4 <- dic.samples(model4, variable.names=c("beta"),
15 A - 8 Appendix 3. Multiple Linear Regression with Different Priors # # # Multiple linear regression using shrinkage priors # # rm(list=ls()) ## Load and standardize NBA data dat <- read.csv(" Y <- dat[,6] Y <- (Y-mean(Y))/sd(Y) X <- dat[,7:25] X <- scale(x) # only for NBA data obs <- dat[,3]!= 2014 Y <- Y[obs] X <- X[obs,] n <- length(y) p <- ncol(x) boxplot(x,las=3,main="standardized Covariates",cex.axis=0.75) image(1:p,1:p,abs(cor(x)), xlab="",ylab="",main="correlation between predictors", axes=false,col=gray(1-seq(0,1,.01))) axis(1,1:p,colnames(x),las=2) axis(2,1:p,colnames(x),las=2) ## Fit the linear regression model # (1) Gaussian: beta_j ~ Normal(0,100^2) model_string1 <- "model{ for(i in 1:n){ Y[i] ~ dnorm(muo[i],inv.var) muo[i] <- alpha + inprod(x[i,],beta[]) beta[j] ~ dnorm(0,0.0001) # " # (2) Gaussian: beta_j ~ Normal(0,sigmab^2) & sigmab^2 ~ InvGamma(0.01,0.01) model_string2 <- "model{ for(i in 1:n){ Y[i] ~ dnorm(muo[i],inv.var) muo[i] <- alpha + inprod(x[i,],beta[])
16 A - 9 beta[j] ~ dnorm(0,inv.var.b) " # (3) Cauchy: beta_j ~ t1(0,sigmab^2) & sigmab^2 ~ InvGamma(0.01,0.01) model_string3 <- "model{ for(i in 1:n){ Y[i] ~ dnorm(muo[i],inv.var) muo[i] <- alpha + inprod(x[i,],beta[]) beta[j] ~ dt(0,inv.var.b,1) " # (4) Bayesian LASSO: beta_j ~ DoubleExpo(0,sigmab^2) & sigmab^2 ~ InvGamma(0.01,0.01) model_string4 <- "model{ for(i in 1:n){ Y[i] ~ dnorm(muo[i],inv.var) muo[i] <- alpha + inprod(x[i,],beta[]) beta[j] ~ ddexp(0,inv.var.b) " # Fit the model library(rjags) model1 <- jags.model(textconnection(model_string1), data = list(y=y,n=n,x=x,p=p)) update(model1, 10000, progress.bar="none") samp1 <- coda.samples(model1, variable.names=c("beta"), model2 <- jags.model(textconnection(model_string2), data = list(y=y,n=n,x=x,p=p))
17 A - 10 update(model2, 10000, progress.bar="none") samp2 <- coda.samples(model2, variable.names=c("beta"), model3 <- jags.model(textconnection(model_string3), data = list(y=y,n=n,x=x,p=p)) update(model3, 10000, progress.bar="none") samp3 <- coda.samples(model3, variable.names=c("beta"), model4 <- jags.model(textconnection(model_string4), data = list(y=y,n=n,x=x,p=p)) update(model4, 10000, progress.bar="none") samp4 <- coda.samples(model4, variable.names=c("beta"), ## Compare the posteriors from the four fits # Extract the MCMC samples from each fit: s1 <- samp1[[1]] s2 <- samp2[[1]] s3 <- samp3[[1]] s4 <- samp4[[1]] # Plot the posterior for each covariance for all four models: for(index in 1:p){ d1 <- density(s1[,index]) d2 <- density(s2[,index]) d3 <- density(s3[,index]) d4 <- density(s4[,index]) mx <- max(d1$y,d2$y,d3$y,d4$y) plot(d1,ylim=c(0,mx),xlab="beta",ylab="posterior density",main=colnames(x)[index]) lines(d2,col=2) lines(d3,col=3) lines(d4,col=4) legend("topright",c("gaussian 1", "Gaussian 2", "Cauchy", "LASSO"),lty=1,col=1:4,inset=0.05)
18 A - 11 Appendix 4. Convergence Test # # # Convergence Test # # rm(list=ls()) ## Load and standardize NBA data dat <- read.csv(" Y <- dat[,6] Y <- (Y-mean(Y))/sd(Y) X <- dat[,7:25] X <- scale(x) # : observed data, 2014: test or prediction data obs <- dat[,3]!= 2014 prd <- dat[,3] == 2014 Yo <- Y[obs] Xo <- X[obs,] Yp <- Y[prd] Xp <- X[prd,] no <- length(yo) np <- length(yp) p <- ncol(xo) ## Fit the linear regression model # (2) Gaussian: beta_j ~ Normal(0,sigmab^2) & sigmab^2 ~ InvGamma(0.01,0.01) model_string1 <- "model{ for(i in 1:no){ Yo[i] ~ dnorm(muo[i],inv.var) muo[i] <- alpha + inprod(xo[i,],beta[]) # Prediction for(i in 1:np){ Yp[i] ~ dnorm(mup[i],inv.var) mup[i] <- alpha + inprod(xp[i,],beta[]) beta[j] ~ dnorm(0,inv.var.b) sigma <- 1/sqrt(inv.var) sigmab <- 1/sqrt(inv.var.b) " # Fit the model library(rjags) model1 <- jags.model(textconnection(model_string1),
19 A - 12 n.chains = 3, data = list(yo=yo,no=no,np=np,p=p,xo=xo,xp=xp)) update(model1, 10000) samp1 <- coda.samples(model1, variable.names=c("beta[1]","sigma","sigmab"), summary(samp1) plot(samp1) effectivesize(samp1) gelman.plot(samp1) autocorr.plot(samp1)
20 A - 13 Appendix 5. Prediction with Real Data # # # Multiple linear regression prediction # # rm(list=ls()) ## Load and standardize NBA data dat <- read.csv(" Y <- dat[,6] Y <- (Y-mean(Y))/sd(Y) X <- dat[,7:25] X <- scale(x) # : observed data, 2014: test or prediction data obs <- dat[,3]!= 2014 prd <- dat[,3] == 2014 Yo <- Y[obs] Xo <- X[obs,] Yp <- Y[prd] Xp <- X[prd,] no <- length(yo) np <- length(yp) p <- ncol(xo) ## Fit the linear regression model # (2) Gaussian: beta_j ~ Normal(0,sigmab^2) & sigmab^2 ~ InvGamma(0.01,0.01) model_string1 <- "model{ for(i in 1:no){ Yo[i] ~ dnorm(muo[i],inv.var) muo[i] <- alpha + inprod(xo[i,],beta[]) # Prediction for(i in 1:np){ Yp[i] ~ dnorm(mup[i],inv.var) mup[i] <- alpha + inprod(xp[i,],beta[]) beta[j] ~ dnorm(0,inv.var.b) sigma <- 1/sqrt(inv.var) " # Fit the model library(rjags) model1 <- jags.model(textconnection(model_string1), data = list(yo=yo,no=no,np=np,p=p,xo=xo,xp=xp))
21 A - 14 update(model1, 10000) samp1 <- coda.samples(model1, variable.names=c("alpha","beta","yp","sigma"), summary(samp1) #plot(samp1) ## Plot samples for each parameter #Extract the samples for each parameter samps1 <- samp1[[1]] Yp.samps1 <- samps1[,1:30] alpha.samps1 <- samps1[,31] beta.samps1 <- samps1[,32:50] sigma.samps1 <- samps1[,51] # Compute the posterior mean for the plug-in predictions beta.mn <- colmeans(beta.samps1) sigma.mn <- mean(sigma.samps1) alpha.mn <- mean(alpha.samps1) # Plot the PPD and plug-in for(j in 1:np){ # PPD plot(density(yp.samps1[,j]),xlab="y",main="ppd") # Plug-in mu <- alpha.mn+sum(xp[j,]*beta.mn) y <- rnorm(20000,mu,sigma.mn) lines(density(y),col=2) # Truth abline(v=yp[j],col=3,lwd=2) legend("topright",c("ppd","plug-in","truth"),col=1:3,lty=1,inset=0.05) ## Compile the results post_mn1 <- apply(yp.samps1,2,mean) post_sd1 <- apply(yp.samps1,2,sd) post_low1 <- apply(yp.samps1,2,quantile,0.025) post_high1 <- apply(yp.samps1,2,quantile,0.975) MSE1 <- mean((post_mn1-yp)^2) BIAS1 <- mean(post_mn1-yp) AVESD1 <- mean(post_sd1) COV1 <- mean(yp>post_low1 & Yp<post_high1) OUTPUT <- cbind(mse1,bias1,avesd1,cov1) as.table(output,digits=2)
Swarthmore Honors Exam 2012: Statistics
 Swarthmore Honors Exam 2012: Statistics 1 Swarthmore Honors Exam 2012: Statistics John W. Emerson, Yale University NAME: Instructions: This is a closed-book three-hour exam having six questions. You may
Swarthmore Honors Exam 2012: Statistics 1 Swarthmore Honors Exam 2012: Statistics John W. Emerson, Yale University NAME: Instructions: This is a closed-book three-hour exam having six questions. You may
Booklet of Code and Output for STAD29/STA 1007 Midterm Exam
 Booklet of Code and Output for STAD29/STA 1007 Midterm Exam List of Figures in this document by page: List of Figures 1 NBA attendance data........................ 2 2 Regression model for NBA attendances...............
Booklet of Code and Output for STAD29/STA 1007 Midterm Exam List of Figures in this document by page: List of Figures 1 NBA attendance data........................ 2 2 Regression model for NBA attendances...............
STAT Lecture 11: Bayesian Regression
 STAT 491 - Lecture 11: Bayesian Regression Generalized Linear Models Generalized linear models (GLMs) are a class of techniques that include linear regression, logistic regression, and Poisson regression.
STAT 491 - Lecture 11: Bayesian Regression Generalized Linear Models Generalized linear models (GLMs) are a class of techniques that include linear regression, logistic regression, and Poisson regression.
Solving Quadratic Equations by Graphing 6.1. ft /sec. The height of the arrow h(t) in terms
 Quadratic Function f ( x) ax bx c Solving Quadratic Equations by Graphing 6.1 Write each in quadratic form. Example 1 f ( x) 3( x + ) Example Graph f ( x) x + 6 x + 8 Example 3 An arrow is shot upward
Quadratic Function f ( x) ax bx c Solving Quadratic Equations by Graphing 6.1 Write each in quadratic form. Example 1 f ( x) 3( x + ) Example Graph f ( x) x + 6 x + 8 Example 3 An arrow is shot upward
ST440/540: Applied Bayesian Statistics. (9) Model selection and goodness-of-fit checks
 (9) Model selection and goodness-of-fit checks Objectives In this module we will study methods for model comparisons and checking for model adequacy For model comparisons there are a finite number of candidate
(9) Model selection and goodness-of-fit checks Objectives In this module we will study methods for model comparisons and checking for model adequacy For model comparisons there are a finite number of candidate
Bayesian linear regression
 Bayesian linear regression Linear regression is the basis of most statistical modeling. The model is Y i = X T i β + ε i, where Y i is the continuous response X i = (X i1,..., X ip ) T is the corresponding
Bayesian linear regression Linear regression is the basis of most statistical modeling. The model is Y i = X T i β + ε i, where Y i is the continuous response X i = (X i1,..., X ip ) T is the corresponding
WinBUGS : part 2. Bruno Boulanger Jonathan Jaeger Astrid Jullion Philippe Lambert. Gabriele, living with rheumatoid arthritis
 WinBUGS : part 2 Bruno Boulanger Jonathan Jaeger Astrid Jullion Philippe Lambert Gabriele, living with rheumatoid arthritis Agenda 2! Hierarchical model: linear regression example! R2WinBUGS Linear Regression
WinBUGS : part 2 Bruno Boulanger Jonathan Jaeger Astrid Jullion Philippe Lambert Gabriele, living with rheumatoid arthritis Agenda 2! Hierarchical model: linear regression example! R2WinBUGS Linear Regression
1. How can you tell if there is serial correlation? 2. AR to model serial correlation. 3. Ignoring serial correlation. 4. GLS. 5. Projects.
 1. How can you tell if there is serial correlation? 2. AR to model serial correlation. 3. Ignoring serial correlation. 4. GLS. 5. Projects. 1) Identifying serial correlation. Plot Y t versus Y t 1. See
1. How can you tell if there is serial correlation? 2. AR to model serial correlation. 3. Ignoring serial correlation. 4. GLS. 5. Projects. 1) Identifying serial correlation. Plot Y t versus Y t 1. See
Lecture 14: Shrinkage
 Lecture 14: Shrinkage Reading: Section 6.2 STATS 202: Data mining and analysis October 27, 2017 1 / 19 Shrinkage methods The idea is to perform a linear regression, while regularizing or shrinking the
Lecture 14: Shrinkage Reading: Section 6.2 STATS 202: Data mining and analysis October 27, 2017 1 / 19 Shrinkage methods The idea is to perform a linear regression, while regularizing or shrinking the
Authors: Antonella Zanobetti and Joel Schwartz
 Title: Mortality Displacement in the Association of Ozone with Mortality: An Analysis of 48 US Cities Authors: Antonella Zanobetti and Joel Schwartz ONLINE DATA SUPPLEMENT Additional Information on Materials
Title: Mortality Displacement in the Association of Ozone with Mortality: An Analysis of 48 US Cities Authors: Antonella Zanobetti and Joel Schwartz ONLINE DATA SUPPLEMENT Additional Information on Materials
Chapter 4: Displaying and Summarizing Quantitative Data
 Chapter 4: Displaying and Summarizing Quantitative Data This chapter discusses methods of displaying quantitative data. The objective is describe the distribution of the data. The figure below shows three
Chapter 4: Displaying and Summarizing Quantitative Data This chapter discusses methods of displaying quantitative data. The objective is describe the distribution of the data. The figure below shows three
Research Update: Race and Male Joblessness in Milwaukee: 2008
 Research Update: Race and Male Joblessness in Milwaukee: 2008 by: Marc V. Levine University of Wisconsin Milwaukee Center for Economic Development Briefing Paper September 2009 Overview Over the past decade,
Research Update: Race and Male Joblessness in Milwaukee: 2008 by: Marc V. Levine University of Wisconsin Milwaukee Center for Economic Development Briefing Paper September 2009 Overview Over the past decade,
36-463/663Multilevel and Hierarchical Models
 36-463/663Multilevel and Hierarchical Models From Bayes to MCMC to MLMs Brian Junker 132E Baker Hall brian@stat.cmu.edu 1 Outline Bayesian Statistics and MCMC Distribution of Skill Mastery in a Population
36-463/663Multilevel and Hierarchical Models From Bayes to MCMC to MLMs Brian Junker 132E Baker Hall brian@stat.cmu.edu 1 Outline Bayesian Statistics and MCMC Distribution of Skill Mastery in a Population
Vibrancy and Property Performance of Major U.S. Employment Centers. Appendix A
 Appendix A DOWNTOWN VIBRANCY SCORES Atlanta 103.3 Minneapolis 152.8 Austin 112.3 Nashville 83.5 Baltimore 151.3 New Orleans 124.3 Birmingham 59.3 New York Midtown 448.6 Charlotte 94.1 Oakland 157.7 Chicago
Appendix A DOWNTOWN VIBRANCY SCORES Atlanta 103.3 Minneapolis 152.8 Austin 112.3 Nashville 83.5 Baltimore 151.3 New Orleans 124.3 Birmingham 59.3 New York Midtown 448.6 Charlotte 94.1 Oakland 157.7 Chicago
1. Evaluation of maximum daily temperature
 1. Evaluation of maximum daily temperature The cumulative distribution of maximum daily temperature is shown in Figure S1. Overall, among all of the 23 states, the cumulative distributions of daily maximum
1. Evaluation of maximum daily temperature The cumulative distribution of maximum daily temperature is shown in Figure S1. Overall, among all of the 23 states, the cumulative distributions of daily maximum
First Year Examination Department of Statistics, University of Florida
 First Year Examination Department of Statistics, University of Florida August 19, 010, 8:00 am - 1:00 noon Instructions: 1. You have four hours to answer questions in this examination.. You must show your
First Year Examination Department of Statistics, University of Florida August 19, 010, 8:00 am - 1:00 noon Instructions: 1. You have four hours to answer questions in this examination.. You must show your
HI SUMMER WORK
 HI-201 2018-2019 SUMMER WORK This packet belongs to: Dear Dual Enrollment Student, May 7 th, 2018 Dual Enrollment United States History is a challenging adventure. Though the year holds countless hours
HI-201 2018-2019 SUMMER WORK This packet belongs to: Dear Dual Enrollment Student, May 7 th, 2018 Dual Enrollment United States History is a challenging adventure. Though the year holds countless hours
DIC: Deviance Information Criterion
 (((( Welcome Page Latest News DIC: Deviance Information Criterion Contact us/bugs list WinBUGS New WinBUGS examples FAQs DIC GeoBUGS DIC (Deviance Information Criterion) is a Bayesian method for model
(((( Welcome Page Latest News DIC: Deviance Information Criterion Contact us/bugs list WinBUGS New WinBUGS examples FAQs DIC GeoBUGS DIC (Deviance Information Criterion) is a Bayesian method for model
Bayesian performance
 Bayesian performance Frequentist properties of estimators refer to the performance of an estimator (say the posterior mean) over repeated experiments under the same conditions. The posterior distribution
Bayesian performance Frequentist properties of estimators refer to the performance of an estimator (say the posterior mean) over repeated experiments under the same conditions. The posterior distribution
Metric Predicted Variable on One Group
 Metric Predicted Variable on One Group Tim Frasier Copyright Tim Frasier This work is licensed under the Creative Commons Attribution 4.0 International license. Click here for more information. Prior Homework
Metric Predicted Variable on One Group Tim Frasier Copyright Tim Frasier This work is licensed under the Creative Commons Attribution 4.0 International license. Click here for more information. Prior Homework
A Parameter Expansion Approach to Bayesian SEM Estimation
 A Parameter Expansion Approach to Bayesian SEM Estimation Ed Merkle and Yves Rosseel Utrecht University 24 June 2016 Yves Rosseel A Parameter Expansion Approach to Bayesian SEM Estimation 1 / 51 overview
A Parameter Expansion Approach to Bayesian SEM Estimation Ed Merkle and Yves Rosseel Utrecht University 24 June 2016 Yves Rosseel A Parameter Expansion Approach to Bayesian SEM Estimation 1 / 51 overview
Bayesian Inference for Regression Parameters
 Bayesian Inference for Regression Parameters 1 Bayesian inference for simple linear regression parameters follows the usual pattern for all Bayesian analyses: 1. Form a prior distribution over all unknown
Bayesian Inference for Regression Parameters 1 Bayesian inference for simple linear regression parameters follows the usual pattern for all Bayesian analyses: 1. Form a prior distribution over all unknown
COMPREHENSIVE WRITTEN EXAMINATION, PAPER III FRIDAY AUGUST 26, 2005, 9:00 A.M. 1:00 P.M. STATISTICS 174 QUESTION
 COMPREHENSIVE WRITTEN EXAMINATION, PAPER III FRIDAY AUGUST 26, 2005, 9:00 A.M. 1:00 P.M. STATISTICS 174 QUESTION Answer all parts. Closed book, calculators allowed. It is important to show all working,
COMPREHENSIVE WRITTEN EXAMINATION, PAPER III FRIDAY AUGUST 26, 2005, 9:00 A.M. 1:00 P.M. STATISTICS 174 QUESTION Answer all parts. Closed book, calculators allowed. It is important to show all working,
bivariate correlation bivariate regression multiple regression
 bivariate correlation bivariate regression multiple regression Today Bivariate Correlation Pearson product-moment correlation (r) assesses nature and strength of the linear relationship between two continuous
bivariate correlation bivariate regression multiple regression Today Bivariate Correlation Pearson product-moment correlation (r) assesses nature and strength of the linear relationship between two continuous
Lesson 1 - Pre-Visit Safe at Home: Location, Place, and Baseball
 Lesson 1 - Pre-Visit Safe at Home: Location, Place, and Baseball Objective: Students will be able to: Define location and place, two of the five themes of geography. Give reasons for the use of latitude
Lesson 1 - Pre-Visit Safe at Home: Location, Place, and Baseball Objective: Students will be able to: Define location and place, two of the five themes of geography. Give reasons for the use of latitude
Final Exam Bus 320 Spring 2000 Russell
 Name Final Exam Bus 320 Spring 2000 Russell Do not turn over this page until you are told to do so. You will have 3 hours minutes to complete this exam. The exam has a total of 100 points and is divided
Name Final Exam Bus 320 Spring 2000 Russell Do not turn over this page until you are told to do so. You will have 3 hours minutes to complete this exam. The exam has a total of 100 points and is divided
Plausible Values for Latent Variables Using Mplus
 Plausible Values for Latent Variables Using Mplus Tihomir Asparouhov and Bengt Muthén August 21, 2010 1 1 Introduction Plausible values are imputed values for latent variables. All latent variables can
Plausible Values for Latent Variables Using Mplus Tihomir Asparouhov and Bengt Muthén August 21, 2010 1 1 Introduction Plausible values are imputed values for latent variables. All latent variables can
Why Bayesian approaches? The average height of a rare plant
 Why Bayesian approaches? The average height of a rare plant Estimation and comparison of averages is an important step in many ecological analyses and demographic models. In this demonstration you will
Why Bayesian approaches? The average height of a rare plant Estimation and comparison of averages is an important step in many ecological analyses and demographic models. In this demonstration you will
Correlation (pp. 1 of 6)
 Correlation (pp. 1 of 6) Car dealers want to know how mileage affects price on used Corvettes. Biologists are studying the effects of temperature on cricket chirps. Farmers are trying to determine if there
Correlation (pp. 1 of 6) Car dealers want to know how mileage affects price on used Corvettes. Biologists are studying the effects of temperature on cricket chirps. Farmers are trying to determine if there
Consistent high-dimensional Bayesian variable selection via penalized credible regions
 Consistent high-dimensional Bayesian variable selection via penalized credible regions Howard Bondell bondell@stat.ncsu.edu Joint work with Brian Reich Howard Bondell p. 1 Outline High-Dimensional Variable
Consistent high-dimensional Bayesian variable selection via penalized credible regions Howard Bondell bondell@stat.ncsu.edu Joint work with Brian Reich Howard Bondell p. 1 Outline High-Dimensional Variable
Lecture: Sampling and Standard Error LECTURE 8 1
 Lecture: Sampling and Standard Error 6.0002 LECTURE 8 1 Announcements Relevant reading: Chapter 17 No lecture Wednesday of next week! 6.0002 LECTURE 8 2 Recall Inferential Statistics Inferential statistics:
Lecture: Sampling and Standard Error 6.0002 LECTURE 8 1 Announcements Relevant reading: Chapter 17 No lecture Wednesday of next week! 6.0002 LECTURE 8 2 Recall Inferential Statistics Inferential statistics:
Bayesian variable selection via. Penalized credible regions. Brian Reich, NCSU. Joint work with. Howard Bondell and Ander Wilson
 Bayesian variable selection via penalized credible regions Brian Reich, NC State Joint work with Howard Bondell and Ander Wilson Brian Reich, NCSU Penalized credible regions 1 Motivation big p, small n
Bayesian variable selection via penalized credible regions Brian Reich, NC State Joint work with Howard Bondell and Ander Wilson Brian Reich, NCSU Penalized credible regions 1 Motivation big p, small n
Machine Learning Linear Classification. Prof. Matteo Matteucci
 Machine Learning Linear Classification Prof. Matteo Matteucci Recall from the first lecture 2 X R p Regression Y R Continuous Output X R p Y {Ω 0, Ω 1,, Ω K } Classification Discrete Output X R p Y (X)
Machine Learning Linear Classification Prof. Matteo Matteucci Recall from the first lecture 2 X R p Regression Y R Continuous Output X R p Y {Ω 0, Ω 1,, Ω K } Classification Discrete Output X R p Y (X)
ECN221 Exam 1 VERSION B Fall 2017 (Modules 1-4), ASU-COX VERSION B
 ECN221 Exam 1 VERSION B Fall 2017 (Modules 1-4), ASU-COX VERSION B Choose the best answer. Do not write letters in the margin or communicate with other students in any way; if you do you will receive a
ECN221 Exam 1 VERSION B Fall 2017 (Modules 1-4), ASU-COX VERSION B Choose the best answer. Do not write letters in the margin or communicate with other students in any way; if you do you will receive a
Regression of Inflation on Percent M3 Change
 ECON 497 Final Exam Page of ECON 497: Economic Research and Forecasting Name: Spring 2006 Bellas Final Exam Return this exam to me by midnight on Thursday, April 27. It may be e-mailed to me. It may be
ECON 497 Final Exam Page of ECON 497: Economic Research and Forecasting Name: Spring 2006 Bellas Final Exam Return this exam to me by midnight on Thursday, April 27. It may be e-mailed to me. It may be
High-dimensional regression modeling
 High-dimensional regression modeling David Causeur Department of Statistics and Computer Science Agrocampus Ouest IRMAR CNRS UMR 6625 http://www.agrocampus-ouest.fr/math/causeur/ Course objectives Making
High-dimensional regression modeling David Causeur Department of Statistics and Computer Science Agrocampus Ouest IRMAR CNRS UMR 6625 http://www.agrocampus-ouest.fr/math/causeur/ Course objectives Making
Kathryn Robinson. Grades 3-5. From the Just Turn & Share Centers Series VOLUME 12
 1 2 From the Just Turn & Share Centers Series VOLUME 12 Temperature TM From the Just Turn & Share Centers Series Kathryn Robinson 3 4 M Enterprises WriteMath Enterprises 2303 Marseille Ct. Suite 104 Valrico,
1 2 From the Just Turn & Share Centers Series VOLUME 12 Temperature TM From the Just Turn & Share Centers Series Kathryn Robinson 3 4 M Enterprises WriteMath Enterprises 2303 Marseille Ct. Suite 104 Valrico,
Assumptions in Regression Modeling
 Fall Semester, 2001 Statistics 621 Lecture 2 Robert Stine 1 Assumptions in Regression Modeling Preliminaries Preparing for class Read the casebook prior to class Pace in class is too fast to absorb without
Fall Semester, 2001 Statistics 621 Lecture 2 Robert Stine 1 Assumptions in Regression Modeling Preliminaries Preparing for class Read the casebook prior to class Pace in class is too fast to absorb without
Heriot-Watt University
 Heriot-Watt University Heriot-Watt University Research Gateway Prediction of settlement delay in critical illness insurance claims by using the generalized beta of the second kind distribution Dodd, Erengul;
Heriot-Watt University Heriot-Watt University Research Gateway Prediction of settlement delay in critical illness insurance claims by using the generalized beta of the second kind distribution Dodd, Erengul;
Scaling in Biology. How do properties of living systems change as their size is varied?
 Scaling in Biology How do properties of living systems change as their size is varied? Example: How does basal metabolic rate (heat radiation) vary as a function of an animal s body mass? Mouse Hamster
Scaling in Biology How do properties of living systems change as their size is varied? Example: How does basal metabolic rate (heat radiation) vary as a function of an animal s body mass? Mouse Hamster
Downloaded from:
 Camacho, A; Kucharski, AJ; Funk, S; Breman, J; Piot, P; Edmunds, WJ (2014) Potential for large outbreaks of Ebola virus disease. Epidemics, 9. pp. 70-8. ISSN 1755-4365 DOI: https://doi.org/10.1016/j.epidem.2014.09.003
Camacho, A; Kucharski, AJ; Funk, S; Breman, J; Piot, P; Edmunds, WJ (2014) Potential for large outbreaks of Ebola virus disease. Epidemics, 9. pp. 70-8. ISSN 1755-4365 DOI: https://doi.org/10.1016/j.epidem.2014.09.003
Linear Regression. Data Model. β, σ 2. Process Model. ,V β. ,s 2. s 1. Parameter Model
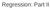 Regression: Part II Linear Regression y~n X, 2 X Y Data Model β, σ 2 Process Model Β 0,V β s 1,s 2 Parameter Model Assumptions of Linear Model Homoskedasticity No error in X variables Error in Y variables
Regression: Part II Linear Regression y~n X, 2 X Y Data Model β, σ 2 Process Model Β 0,V β s 1,s 2 Parameter Model Assumptions of Linear Model Homoskedasticity No error in X variables Error in Y variables
Supplementary Note on Bayesian analysis
 Supplementary Note on Bayesian analysis Structured variability of muscle activations supports the minimal intervention principle of motor control Francisco J. Valero-Cuevas 1,2,3, Madhusudhan Venkadesan
Supplementary Note on Bayesian analysis Structured variability of muscle activations supports the minimal intervention principle of motor control Francisco J. Valero-Cuevas 1,2,3, Madhusudhan Venkadesan
Penalized Loss functions for Bayesian Model Choice
 Penalized Loss functions for Bayesian Model Choice Martyn International Agency for Research on Cancer Lyon, France 13 November 2009 The pure approach For a Bayesian purist, all uncertainty is represented
Penalized Loss functions for Bayesian Model Choice Martyn International Agency for Research on Cancer Lyon, France 13 November 2009 The pure approach For a Bayesian purist, all uncertainty is represented
Name. The data below are airfares to various cities from Baltimore, MD (including the descriptive statistics).
 Name The data below are airfares to various cities from Baltimore, MD (including the descriptive statistics). 178 138 94 278 158 258 198 188 98 179 138 98 N Mean Std. Dev. Min Q 1 Median Q 3 Max 12 166.92
Name The data below are airfares to various cities from Baltimore, MD (including the descriptive statistics). 178 138 94 278 158 258 198 188 98 179 138 98 N Mean Std. Dev. Min Q 1 Median Q 3 Max 12 166.92
Package horseshoe. November 8, 2016
 Title Implementation of the Horseshoe Prior Version 0.1.0 Package horseshoe November 8, 2016 Description Contains functions for applying the horseshoe prior to highdimensional linear regression, yielding
Title Implementation of the Horseshoe Prior Version 0.1.0 Package horseshoe November 8, 2016 Description Contains functions for applying the horseshoe prior to highdimensional linear regression, yielding
Regression Models - Introduction
 Regression Models - Introduction In regression models there are two types of variables that are studied: A dependent variable, Y, also called response variable. It is modeled as random. An independent
Regression Models - Introduction In regression models there are two types of variables that are studied: A dependent variable, Y, also called response variable. It is modeled as random. An independent
Estimation and Confidence Intervals
 Estimation and Confidence Intervals Sections 7.1-7.3 Cathy Poliak, Ph.D. cathy@math.uh.edu Office in Fleming 11c Department of Mathematics University of Houston Lecture 17-3339 Cathy Poliak, Ph.D. cathy@math.uh.edu
Estimation and Confidence Intervals Sections 7.1-7.3 Cathy Poliak, Ph.D. cathy@math.uh.edu Office in Fleming 11c Department of Mathematics University of Houston Lecture 17-3339 Cathy Poliak, Ph.D. cathy@math.uh.edu
36-463/663: Multilevel & Hierarchical Models HW09 Solution
 36-463/663: Multilevel & Hierarchical Models HW09 Solution November 15, 2016 Quesion 1 Following the derivation given in class, when { n( x µ) 2 L(µ) exp, f(p) exp 2σ 2 0 ( the posterior is also normally
36-463/663: Multilevel & Hierarchical Models HW09 Solution November 15, 2016 Quesion 1 Following the derivation given in class, when { n( x µ) 2 L(µ) exp, f(p) exp 2σ 2 0 ( the posterior is also normally
Review: Second Half of Course Stat 704: Data Analysis I, Fall 2014
 Review: Second Half of Course Stat 704: Data Analysis I, Fall 2014 Tim Hanson, Ph.D. University of South Carolina T. Hanson (USC) Stat 704: Data Analysis I, Fall 2014 1 / 13 Chapter 8: Polynomials & Interactions
Review: Second Half of Course Stat 704: Data Analysis I, Fall 2014 Tim Hanson, Ph.D. University of South Carolina T. Hanson (USC) Stat 704: Data Analysis I, Fall 2014 1 / 13 Chapter 8: Polynomials & Interactions
For final project discussion every afternoon Mark and I will be available
 Worshop report 1. Daniels report is on website 2. Don t expect to write it based on listening to one project (we had 6 only 2 was sufficient quality) 3. I suggest writing it on one presentation. 4. Include
Worshop report 1. Daniels report is on website 2. Don t expect to write it based on listening to one project (we had 6 only 2 was sufficient quality) 3. I suggest writing it on one presentation. 4. Include
Markov Chain Monte Carlo
 Markov Chain Monte Carlo Recall: To compute the expectation E ( h(y ) ) we use the approximation E(h(Y )) 1 n n h(y ) t=1 with Y (1),..., Y (n) h(y). Thus our aim is to sample Y (1),..., Y (n) from f(y).
Markov Chain Monte Carlo Recall: To compute the expectation E ( h(y ) ) we use the approximation E(h(Y )) 1 n n h(y ) t=1 with Y (1),..., Y (n) h(y). Thus our aim is to sample Y (1),..., Y (n) from f(y).
Bayesian Model Diagnostics and Checking
 Earvin Balderama Quantitative Ecology Lab Department of Forestry and Environmental Resources North Carolina State University April 12, 2013 1 / 34 Introduction MCMCMC 2 / 34 Introduction MCMCMC Steps in
Earvin Balderama Quantitative Ecology Lab Department of Forestry and Environmental Resources North Carolina State University April 12, 2013 1 / 34 Introduction MCMCMC 2 / 34 Introduction MCMCMC Steps in
Biostatistics-Lecture 16 Model Selection. Ruibin Xi Peking University School of Mathematical Sciences
 Biostatistics-Lecture 16 Model Selection Ruibin Xi Peking University School of Mathematical Sciences Motivating example1 Interested in factors related to the life expectancy (50 US states,1969-71 ) Per
Biostatistics-Lecture 16 Model Selection Ruibin Xi Peking University School of Mathematical Sciences Motivating example1 Interested in factors related to the life expectancy (50 US states,1969-71 ) Per
GENOMIC SELECTION WORKSHOP: Hands on Practical Sessions (BL)
 GENOMIC SELECTION WORKSHOP: Hands on Practical Sessions (BL) Paulino Pérez 1 José Crossa 2 1 ColPos-México 2 CIMMyT-México September, 2014. SLU,Sweden GENOMIC SELECTION WORKSHOP:Hands on Practical Sessions
GENOMIC SELECTION WORKSHOP: Hands on Practical Sessions (BL) Paulino Pérez 1 José Crossa 2 1 ColPos-México 2 CIMMyT-México September, 2014. SLU,Sweden GENOMIC SELECTION WORKSHOP:Hands on Practical Sessions
Trends in Metropolitan Network Circuity
 Trends in Metropolitan Network Circuity David J. Giacomin Luke S. James David M. Levinson Abstract Because people seek to minimize their time and travel distance (or cost) when commuting, the circuity
Trends in Metropolitan Network Circuity David J. Giacomin Luke S. James David M. Levinson Abstract Because people seek to minimize their time and travel distance (or cost) when commuting, the circuity
Metric Predicted Variable With One Nominal Predictor Variable
 Metric Predicted Variable With One Nominal Predictor Variable Tim Frasier Copyright Tim Frasier This work is licensed under the Creative Commons Attribution 4.0 International license. Click here for more
Metric Predicted Variable With One Nominal Predictor Variable Tim Frasier Copyright Tim Frasier This work is licensed under the Creative Commons Attribution 4.0 International license. Click here for more
Metric Predicted Variable on Two Groups
 Metric Predicted Variable on Two Groups Tim Frasier Copyright Tim Frasier This work is licensed under the Creative Commons Attribution 4.0 International license. Click here for more information. Goals
Metric Predicted Variable on Two Groups Tim Frasier Copyright Tim Frasier This work is licensed under the Creative Commons Attribution 4.0 International license. Click here for more information. Goals
C Further Concepts in Statistics
 Appendix C.1 Representing Data and Linear Modeling C1 C Further Concepts in Statistics C.1 Representing Data and Linear Modeling Use stem-and-leaf plots to organize and compare sets of data. Use histograms
Appendix C.1 Representing Data and Linear Modeling C1 C Further Concepts in Statistics C.1 Representing Data and Linear Modeling Use stem-and-leaf plots to organize and compare sets of data. Use histograms
Information. Hierarchical Models - Statistical Methods. References. Outline
 Information Hierarchical Models - Statistical Methods Sarah Filippi 1 University of Oxford Hilary Term 2015 Webpage: http://www.stats.ox.ac.uk/~filippi/msc_ hierarchicalmodels_2015.html Lectures: Week
Information Hierarchical Models - Statistical Methods Sarah Filippi 1 University of Oxford Hilary Term 2015 Webpage: http://www.stats.ox.ac.uk/~filippi/msc_ hierarchicalmodels_2015.html Lectures: Week
Simple Linear Regression: One Quantitative IV
 Simple Linear Regression: One Quantitative IV Linear regression is frequently used to explain variation observed in a dependent variable (DV) with theoretically linked independent variables (IV). For example,
Simple Linear Regression: One Quantitative IV Linear regression is frequently used to explain variation observed in a dependent variable (DV) with theoretically linked independent variables (IV). For example,
Bayesian Graphical Models
 Graphical Models and Inference, Lecture 16, Michaelmas Term 2009 December 4, 2009 Parameter θ, data X = x, likelihood L(θ x) p(x θ). Express knowledge about θ through prior distribution π on θ. Inference
Graphical Models and Inference, Lecture 16, Michaelmas Term 2009 December 4, 2009 Parameter θ, data X = x, likelihood L(θ x) p(x θ). Express knowledge about θ through prior distribution π on θ. Inference
Bayesian Networks in Educational Assessment
 Bayesian Networks in Educational Assessment Estimating Parameters with MCMC Bayesian Inference: Expanding Our Context Roy Levy Arizona State University Roy.Levy@asu.edu 2017 Roy Levy MCMC 1 MCMC 2 Posterior
Bayesian Networks in Educational Assessment Estimating Parameters with MCMC Bayesian Inference: Expanding Our Context Roy Levy Arizona State University Roy.Levy@asu.edu 2017 Roy Levy MCMC 1 MCMC 2 Posterior
American Tour: Climate Objective To introduce contour maps as data displays.
 American Tour: Climate Objective To introduce contour maps as data displays. www.everydaymathonline.com epresentations etoolkit Algorithms Practice EM Facts Workshop Game Family Letters Assessment Management
American Tour: Climate Objective To introduce contour maps as data displays. www.everydaymathonline.com epresentations etoolkit Algorithms Practice EM Facts Workshop Game Family Letters Assessment Management
Stat 544 Final Exam. May 2, I have neither given nor received unauthorized assistance on this examination.
 Stat 544 Final Exam May, 006 I have neither given nor received unauthorized assistance on this examination. signature date 1 1. Below is a directed acyclic graph that represents a joint distribution for
Stat 544 Final Exam May, 006 I have neither given nor received unauthorized assistance on this examination. signature date 1 1. Below is a directed acyclic graph that represents a joint distribution for
Approximate Bayesian binary and ordinal regression for prediction with structured uncertainty in the inputs
 Approximate Bayesian binary and ordinal regression for prediction with structured uncertainty in the inputs Aleksandar Dimitriev Erik Štrumbelj Abstract We present a novel approach to binary and ordinal
Approximate Bayesian binary and ordinal regression for prediction with structured uncertainty in the inputs Aleksandar Dimitriev Erik Štrumbelj Abstract We present a novel approach to binary and ordinal
North American Geography. Lesson 5: Barnstorm Like a Tennis Player!
 North American Geography Lesson 5: Barnstorm Like a Tennis Player! Unit Overview: As students work through the activities in this unit they will be introduced to the United States in general, different
North American Geography Lesson 5: Barnstorm Like a Tennis Player! Unit Overview: As students work through the activities in this unit they will be introduced to the United States in general, different
Bayesian Analysis of Multivariate Normal Models when Dimensions are Absent
 Bayesian Analysis of Multivariate Normal Models when Dimensions are Absent Robert Zeithammer University of Chicago Peter Lenk University of Michigan http://webuser.bus.umich.edu/plenk/downloads.htm SBIES
Bayesian Analysis of Multivariate Normal Models when Dimensions are Absent Robert Zeithammer University of Chicago Peter Lenk University of Michigan http://webuser.bus.umich.edu/plenk/downloads.htm SBIES
Introduction to Machine Learning Midterm, Tues April 8
 Introduction to Machine Learning 10-701 Midterm, Tues April 8 [1 point] Name: Andrew ID: Instructions: You are allowed a (two-sided) sheet of notes. Exam ends at 2:45pm Take a deep breath and don t spend
Introduction to Machine Learning 10-701 Midterm, Tues April 8 [1 point] Name: Andrew ID: Instructions: You are allowed a (two-sided) sheet of notes. Exam ends at 2:45pm Take a deep breath and don t spend
Inference in Normal Regression Model. Dr. Frank Wood
 Inference in Normal Regression Model Dr. Frank Wood Remember We know that the point estimator of b 1 is b 1 = (Xi X )(Y i Ȳ ) (Xi X ) 2 Last class we derived the sampling distribution of b 1, it being
Inference in Normal Regression Model Dr. Frank Wood Remember We know that the point estimator of b 1 is b 1 = (Xi X )(Y i Ȳ ) (Xi X ) 2 Last class we derived the sampling distribution of b 1, it being
Package spatial.gev.bma
 Type Package Package spatial.gev.bma February 20, 2015 Title Hierarchical spatial generalized extreme value (GEV) modeling with Bayesian Model Averaging (BMA) Version 1.0 Date 2014-03-11 Author Alex Lenkoski
Type Package Package spatial.gev.bma February 20, 2015 Title Hierarchical spatial generalized extreme value (GEV) modeling with Bayesian Model Averaging (BMA) Version 1.0 Date 2014-03-11 Author Alex Lenkoski
STAT 425: Introduction to Bayesian Analysis
 STAT 425: Introduction to Bayesian Analysis Marina Vannucci Rice University, USA Fall 2017 Marina Vannucci (Rice University, USA) Bayesian Analysis (Part 2) Fall 2017 1 / 19 Part 2: Markov chain Monte
STAT 425: Introduction to Bayesian Analysis Marina Vannucci Rice University, USA Fall 2017 Marina Vannucci (Rice University, USA) Bayesian Analysis (Part 2) Fall 2017 1 / 19 Part 2: Markov chain Monte
Probabilistic machine learning group, Aalto University Bayesian theory and methods, approximative integration, model
 Aki Vehtari, Aalto University, Finland Probabilistic machine learning group, Aalto University http://research.cs.aalto.fi/pml/ Bayesian theory and methods, approximative integration, model assessment and
Aki Vehtari, Aalto University, Finland Probabilistic machine learning group, Aalto University http://research.cs.aalto.fi/pml/ Bayesian theory and methods, approximative integration, model assessment and
Markov Chain Monte Carlo
 Markov Chain Monte Carlo Jamie Monogan University of Georgia Spring 2013 For more information, including R programs, properties of Markov chains, and Metropolis-Hastings, please see: http://monogan.myweb.uga.edu/teaching/statcomp/mcmc.pdf
Markov Chain Monte Carlo Jamie Monogan University of Georgia Spring 2013 For more information, including R programs, properties of Markov chains, and Metropolis-Hastings, please see: http://monogan.myweb.uga.edu/teaching/statcomp/mcmc.pdf
NAWIC. National Association of Women in Construction. Membership Report. August 2009
 NAWIC National Association of Women in Construction Membership Report August 2009 Core Purpose: To enhance the success of women in the construction industry Region 1 67 Gr Washington, DC 9 16 2 3 1 0 0
NAWIC National Association of Women in Construction Membership Report August 2009 Core Purpose: To enhance the success of women in the construction industry Region 1 67 Gr Washington, DC 9 16 2 3 1 0 0
ASSESSING ACCURACY: HOTEL HORIZONS FORECASTS
 ASSESSING ACCURACY: HOTEL HORIZONS FORECASTS April 13, 2016 EXECUTIVE SUMMARY The US hotel industry had another strong year in 2015 with RevPAR up 6.3 percent over the prior year. In this report, we examine
ASSESSING ACCURACY: HOTEL HORIZONS FORECASTS April 13, 2016 EXECUTIVE SUMMARY The US hotel industry had another strong year in 2015 with RevPAR up 6.3 percent over the prior year. In this report, we examine
ESTIMATING THE MEAN LEVEL OF FINE PARTICULATE MATTER: AN APPLICATION OF SPATIAL STATISTICS
 ESTIMATING THE MEAN LEVEL OF FINE PARTICULATE MATTER: AN APPLICATION OF SPATIAL STATISTICS Richard L. Smith Department of Statistics and Operations Research University of North Carolina Chapel Hill, N.C.,
ESTIMATING THE MEAN LEVEL OF FINE PARTICULATE MATTER: AN APPLICATION OF SPATIAL STATISTICS Richard L. Smith Department of Statistics and Operations Research University of North Carolina Chapel Hill, N.C.,
Determining the Spread of a Distribution
 Determining the Spread of a Distribution 1.3-1.5 Cathy Poliak, Ph.D. cathy@math.uh.edu Department of Mathematics University of Houston Lecture 3-2311 Lecture 3-2311 1 / 58 Outline 1 Describing Quantitative
Determining the Spread of a Distribution 1.3-1.5 Cathy Poliak, Ph.D. cathy@math.uh.edu Department of Mathematics University of Houston Lecture 3-2311 Lecture 3-2311 1 / 58 Outline 1 Describing Quantitative
Milk components rebounding across all western regions
 DV Monitors Milk components rebounding across all western regions By W.K. (Bill) Sanchez, Ph.D., Dipl. ACAN Technical Service Director Dairy Diamond V As published in From DV Monitors data through the
DV Monitors Milk components rebounding across all western regions By W.K. (Bill) Sanchez, Ph.D., Dipl. ACAN Technical Service Director Dairy Diamond V As published in From DV Monitors data through the
Public Library Use and Economic Hard Times: Analysis of Recent Data
 Public Library Use and Economic Hard Times: Analysis of Recent Data A Report Prepared for The American Library Association by The Library Research Center University of Illinois at Urbana Champaign April
Public Library Use and Economic Hard Times: Analysis of Recent Data A Report Prepared for The American Library Association by The Library Research Center University of Illinois at Urbana Champaign April
Determining the Spread of a Distribution
 Determining the Spread of a Distribution 1.3-1.5 Cathy Poliak, Ph.D. cathy@math.uh.edu Department of Mathematics University of Houston Lecture 3-2311 Lecture 3-2311 1 / 58 Outline 1 Describing Quantitative
Determining the Spread of a Distribution 1.3-1.5 Cathy Poliak, Ph.D. cathy@math.uh.edu Department of Mathematics University of Houston Lecture 3-2311 Lecture 3-2311 1 / 58 Outline 1 Describing Quantitative
(5) Multi-parameter models - Gibbs sampling. ST440/540: Applied Bayesian Analysis
 Summarizing a posterior Given the data and prior the posterior is determined Summarizing the posterior gives parameter estimates, intervals, and hypothesis tests Most of these computations are integrals
Summarizing a posterior Given the data and prior the posterior is determined Summarizing the posterior gives parameter estimates, intervals, and hypothesis tests Most of these computations are integrals
Correlation and Linear Regression
 Correlation and Linear Regression Correlation: Relationships between Variables So far, nearly all of our discussion of inferential statistics has focused on testing for differences between group means
Correlation and Linear Regression Correlation: Relationships between Variables So far, nearly all of our discussion of inferential statistics has focused on testing for differences between group means
Online Appendix: Bayesian versus maximum likelihood estimation of treatment effects in bivariate probit instrumental variable models
 Online Appendix: Bayesian versus maximum likelihood estimation of treatment effects in bivariate probit instrumental variable models A. STAN CODE // STAN code for Bayesian bivariate model // based on code
Online Appendix: Bayesian versus maximum likelihood estimation of treatment effects in bivariate probit instrumental variable models A. STAN CODE // STAN code for Bayesian bivariate model // based on code
ADVENTURES IN THE FLIPPED CLASSROOM FOR INTRODUCTORY
 ADVENTURES IN THE FLIPPED CLASSROOM FOR INTRODUCTORY A M Y N U S S B A U M A N D M O N N I E M C G E E STATISTICS S R C O S J U N E 5, 2 0 1 3 Amy Nussbaum and Monnie McGee SRCOS, June 5, 2013 FLIPPED
ADVENTURES IN THE FLIPPED CLASSROOM FOR INTRODUCTORY A M Y N U S S B A U M A N D M O N N I E M C G E E STATISTICS S R C O S J U N E 5, 2 0 1 3 Amy Nussbaum and Monnie McGee SRCOS, June 5, 2013 FLIPPED
Part 8: GLMs and Hierarchical LMs and GLMs
 Part 8: GLMs and Hierarchical LMs and GLMs 1 Example: Song sparrow reproductive success Arcese et al., (1992) provide data on a sample from a population of 52 female song sparrows studied over the course
Part 8: GLMs and Hierarchical LMs and GLMs 1 Example: Song sparrow reproductive success Arcese et al., (1992) provide data on a sample from a population of 52 female song sparrows studied over the course
Bayesian Analysis of Latent Variable Models using Mplus
 Bayesian Analysis of Latent Variable Models using Mplus Tihomir Asparouhov and Bengt Muthén Version 2 June 29, 2010 1 1 Introduction In this paper we describe some of the modeling possibilities that are
Bayesian Analysis of Latent Variable Models using Mplus Tihomir Asparouhov and Bengt Muthén Version 2 June 29, 2010 1 1 Introduction In this paper we describe some of the modeling possibilities that are
JUPITER MILLER BUSINESS CENTER 746,400 SF FOR LEASE
 746,400 SF FOR LEASE Three LEED Certified Cross-Dock Buildings 54,600 Square Feet to 746,400 Square Feet Available Dallas City of Tax Incentives Available 36 Clear Height (Over 25% More Pallet Positions
746,400 SF FOR LEASE Three LEED Certified Cross-Dock Buildings 54,600 Square Feet to 746,400 Square Feet Available Dallas City of Tax Incentives Available 36 Clear Height (Over 25% More Pallet Positions
Supplemental Appendix to Media Bias and Reputation
 Supplemental Appendix to Media Bias and Reputation Matthew Gentzkow Graduate School of Business University of Chicago Jesse M. Shapiro University of Chicago and NBER September 14, 2005 A Extensions and
Supplemental Appendix to Media Bias and Reputation Matthew Gentzkow Graduate School of Business University of Chicago Jesse M. Shapiro University of Chicago and NBER September 14, 2005 A Extensions and
SMAM 319 Exam1 Name. a B.The equation of a line is 3x + y =6. The slope is a. -3 b.3 c.6 d.1/3 e.-1/3
 SMAM 319 Exam1 Name 1. Pick the best choice. (10 points-2 each) _c A. A data set consisting of fifteen observations has the five number summary 4 11 12 13 15.5. For this data set it is definitely true
SMAM 319 Exam1 Name 1. Pick the best choice. (10 points-2 each) _c A. A data set consisting of fifteen observations has the five number summary 4 11 12 13 15.5. For this data set it is definitely true
Name: Biostatistics 1 st year Comprehensive Examination: Applied in-class exam. June 8 th, 2016: 9am to 1pm
 Name: Biostatistics 1 st year Comprehensive Examination: Applied in-class exam June 8 th, 2016: 9am to 1pm Instructions: 1. This is exam is to be completed independently. Do not discuss your work with
Name: Biostatistics 1 st year Comprehensive Examination: Applied in-class exam June 8 th, 2016: 9am to 1pm Instructions: 1. This is exam is to be completed independently. Do not discuss your work with
Statistical Data Mining and Machine Learning Hilary Term 2016
 Statistical Data Mining and Machine Learning Hilary Term 2016 Dino Sejdinovic Department of Statistics Oxford Slides and other materials available at: http://www.stats.ox.ac.uk/~sejdinov/sdmml Naïve Bayes
Statistical Data Mining and Machine Learning Hilary Term 2016 Dino Sejdinovic Department of Statistics Oxford Slides and other materials available at: http://www.stats.ox.ac.uk/~sejdinov/sdmml Naïve Bayes
Multiple Linear Regression
 Andrew Lonardelli December 20, 2013 Multiple Linear Regression 1 Table Of Contents Introduction: p.3 Multiple Linear Regression Model: p.3 Least Squares Estimation of the Parameters: p.4-5 The matrix approach
Andrew Lonardelli December 20, 2013 Multiple Linear Regression 1 Table Of Contents Introduction: p.3 Multiple Linear Regression Model: p.3 Least Squares Estimation of the Parameters: p.4-5 The matrix approach
Machine Learning, Fall 2009: Midterm
 10-601 Machine Learning, Fall 009: Midterm Monday, November nd hours 1. Personal info: Name: Andrew account: E-mail address:. You are permitted two pages of notes and a calculator. Please turn off all
10-601 Machine Learning, Fall 009: Midterm Monday, November nd hours 1. Personal info: Name: Andrew account: E-mail address:. You are permitted two pages of notes and a calculator. Please turn off all
q3_3 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question.
 q3_3 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. Provide an appropriate response. 1) In 2007, the number of wins had a mean of 81.79 with a standard
q3_3 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. Provide an appropriate response. 1) In 2007, the number of wins had a mean of 81.79 with a standard
Generalized Linear Models
 Generalized Linear Models Assumptions of Linear Model Homoskedasticity Model variance No error in X variables Errors in variables No missing data Missing data model Normally distributed error Error in
Generalized Linear Models Assumptions of Linear Model Homoskedasticity Model variance No error in X variables Errors in variables No missing data Missing data model Normally distributed error Error in
CHAPTER 13. Multiple Regression MODEL ASSUMPTIONS 13.5 TESTING FOR SIGNIFICANCE t Test CONTENTS
 CHAPTER 13 Multiple Regression CONTENTS STATISTICS IN PRACTICE: INTERNATIONAL PAPER 13.1 MULTIPLE REGRESSION MODEL Regression Model and Regression Equation Estimated Multiple Regression Equation 13.2 LEAST
CHAPTER 13 Multiple Regression CONTENTS STATISTICS IN PRACTICE: INTERNATIONAL PAPER 13.1 MULTIPLE REGRESSION MODEL Regression Model and Regression Equation Estimated Multiple Regression Equation 13.2 LEAST
Lecture 19. Spatial GLM + Point Reference Spatial Data. Colin Rundel 04/03/2017
 Lecture 19 Spatial GLM + Point Reference Spatial Data Colin Rundel 04/03/2017 1 Spatial GLM Models 2 Scottish Lip Cancer Data Observed Expected 60 N 59 N 58 N 57 N 56 N value 80 60 40 20 0 55 N 8 W 6 W
Lecture 19 Spatial GLM + Point Reference Spatial Data Colin Rundel 04/03/2017 1 Spatial GLM Models 2 Scottish Lip Cancer Data Observed Expected 60 N 59 N 58 N 57 N 56 N value 80 60 40 20 0 55 N 8 W 6 W
Robust Bayesian Regression
 Readings: Hoff Chapter 9, West JRSSB 1984, Fúquene, Pérez & Pericchi 2015 Duke University November 17, 2016 Body Fat Data: Intervals w/ All Data Response % Body Fat and Predictor Waist Circumference 95%
Readings: Hoff Chapter 9, West JRSSB 1984, Fúquene, Pérez & Pericchi 2015 Duke University November 17, 2016 Body Fat Data: Intervals w/ All Data Response % Body Fat and Predictor Waist Circumference 95%
Statistics 203: Introduction to Regression and Analysis of Variance Penalized models
 Statistics 203: Introduction to Regression and Analysis of Variance Penalized models Jonathan Taylor - p. 1/15 Today s class Bias-Variance tradeoff. Penalized regression. Cross-validation. - p. 2/15 Bias-variance
Statistics 203: Introduction to Regression and Analysis of Variance Penalized models Jonathan Taylor - p. 1/15 Today s class Bias-Variance tradeoff. Penalized regression. Cross-validation. - p. 2/15 Bias-variance
