Genomics Education Partnership Fosmid 16 Harry Quedenfeld. Data and text from this paper is allowed to be included in publications
|
|
|
- Theresa Pope
- 5 years ago
- Views:
Transcription
1 TheCharacterizationofOrthologousCG14561, CG7139 PA,CG7139 PB,CG7133,CG7130andthe IneffectualExclusionofDrosophilamelanogaster ExonicSequenceinRpLP0inDrosophilaerecta GenomicsEducationPartnership Fosmid16 HarryQuedenfeld Dataandtextfromthispaperisallowedtobeincludedinpublications Overview Thearearepresentedbymyfosmidstartsatapproximatelybasepairnumber22,058,504andproceeds44,625 basepairsuntilabout22,103,128indrosophilamelanogaster,thoughfosmid16is40,000bpind.ere.,which showsthateitherthelatterhasdeletionsortheformerinsertions.thefigurebelowshowsthegene containing region,astherestofdna3 ofmsopacontainsnogenes.theintentionofthisprojectwastodeterminegenes withinmyfosmidofdrosophilaerectaandprobablefunctionofthosegenesbasedond.mel.asareference genome.thetranscriptsofthegenescg14561,cg7139,cg7133,cg7130,rplp0,sfp79bandmsopawerefound tohavehomologousregionswithmyfosmidbyuseofblast2,andallconfirmedgeneshadthesameamountof exonsastheirorthologousgenesind.mel(rplp0isanexceptionduetogenechecker sinsufficiencytodetect untranslatedexons).sfp79bandmsopaarethoughttobepseudogenesbecauseoftheirshortpeptidelengthand highdegreeofmutationfromd.mel.evidencefromd.mel.supportsthatsimilargeneswithsimilarfunctional proteinsarefoundind.ere.forallgenesexceptsfp79bandmsopa.rplp0 sfirstexonandaportionofthesecond arepartofthe3 untranslatedregionind.mel.rplp0ind.mel.doesnothaveastartcodoninitsfirstexon,but inthemiddleofitssecond.ind.ere.,however,thereisamutationwhichcausesforaprematurestartand readingofaprematurestopcodon.ifthisnormallyuntranslated1 st exonwastranscribedandtranslated,the proteinwouldbenonfunctionalandapseudogene.ifthe1 st exonisnottranscribed,thentherplp0proteinind. ere.is98%identicaltod.mel. s,whichisexpectedforaribosomalprotein.genescg14561,cg7139andcg7130 ind.ere.havenosignificantdifferencesfromtheird.mel.ortholog.cg7133ind.ere.issimilarlengthbutshows a55%identitywithd.mel. s,soitprobablyhasadifferentfunctionind.ere.thisproteinisnothighlyconserved withincloserelativesofd.ere.andisstillthoughttobeagene,whichevidencesitsprobabilitytobeageneind. ere.aswell.inthespeciesdrosophilasimulans,drosophilasechelliaanddrosophilayakuba,cg7133isnothighly conservedbutisstillaputativegene,thusthisisalsoageneind.ere.thegenesfp79bisapseudogenebecauseit
2 HarryQuedenfeld Genomics Dr.Moss isonly4aminoacidsinlengthduetoamutationthatcodesforanearlystartcodon,whichisfollowedbyastop codon.msopaisalsothoughttobeapseudogenebecauseitisonly64aminoacidslongandhasalargedeletion whencomparedtod.mel. Dere/GG13189 RB Genes CG14561/NM_ CG14561 PAisthenameoftheorthologinmelanogastertothegenefoundinfosmid16ofD.ere.Ithasasingle isoform,cg14561 PA.Thisgenecodesfrom andthestopcodonispresentat infosmid16.It isasingleexongeneandcodesintheplusdirection.itisageneind.mel.andthusisprotein coding,yetits functionandgeneontologyisunknown.theblastpbetweentheproteinforthegeneiannotatedind.ere.and theknowngeneind.mel.yieldsasignificante valueof3e 100andwith89%identity,showingthattheyare ratherhomologous.thefunctionind.mel.islikelythesameasthefunctionind.ere.,sooncethefunctionis found,itissafetoassumeitsfunctionind.ere.isalsofound.thisgenehasorthologsin10closelyrelated Drosophilaspecies,soitoriginatedfromacommonancestorbetweenthese.Oncethereisareferencewithinone ofthese10,itsfunctioncanlikelybeappliedtoallbecausethissequenceismoderatelyconserved.orthologsare named(dspecies/name):dana\gf10942,dgri\gh15124,dmoj\gi13775,dper\gl25424,dpse\ga13080, Dsec\GM22428,Dsim\GD15018,Dvir\GJ13580,Dwil\GK10706andDyak\GE Exon 1 [flybase.org,genomebrowser scaffold_4784: ] Start End Stopcodon CG7139/NM_ ThisgenehastwomRNAproducts,ortwoisoforms,namedCG7139 PAandCG7139 PBinD.mel.Bothisoforms areconservedratherhighlyind.ere.,stillhaving4exonsinisoformaand2exonsinisoformb,bothrunningin theminusdirection.isoformbconsistsofthelasttwoexonsofisoforma,thusismissingthefirsttwoexonsfrom isoforma.thefunctionofthisgene,ineitherisoform,isnotknown.cg7139hasorthologsin10otherdrosophilid species,inwhichthey renamed(dspecies/name):dana\gf23494,dgri\gh16330,dmoj\gi11522,dper\gl25453, Dpse\GA20130,Dsec\GM22101,Dsim\GD12077,Dvir\GJ11777,Dwil\GK12132andDyak\GE22996.The coordinatesinfosmid16forisoformaare , , and ,withastopcodon 2
3 HarryQuedenfeld Genomics Dr.Moss at ThecoordinatesforisoformBare and withastopcodonat The lasttwoexonsinisoformaaretheexactsamepresentinisoformb;sincethefirstcodoninthethirdexonin isoformaisamethionine(astartcodon),isoformbbeginsatthesamelocation.thefirsttwoexonsofisoforma canbeexcludedtoyieldisoformb,thusthisgenecodesfortwomrnaandproteinproductsthatmayplay differingorsimilarrolesinacell.thisistheonlygeneinfosmid16thatdisplaystwomrnasforonegene.the proteinproductsyieldedbymyannotationofd.ere. sgenearesimilartothoseind.mel.blastpshowedanevalueof0.0andanidentityof85%withbothisoforms. CG7139 PA Exon Start End Stopcodon CG7139 PB Exon Start End Stopcodon CG7133/NM_ Thisgeneisasingleexongenetranscribedintheminusdirection.Thecodingregioncoordinatesinfosmid16is andthestopcodonisfrom Ithasasingleisoform,CG7133 PA.Thisgenehasitsgene ontologydefined.itsmolecularfunctionisdescribedasaproteinthatbindstounfoldedprotein.itfunctionsin heatshockprotein(hsp)binding.thusthisproteinisaheatshockproteinthatactsasachaperoneproteintofold unfoldedproteinsthatareeitherrecentlytranslatedorweredenaturedbyheat.thee valuefromablastp betweenthed.mel.cg7133proteinandthed.ere. sorthologousproteinis3e 81,andidentitysitsat55%.Sixtyfivepercentscoreaspositives,meaning10%ofaminoacidsthatdifferaresimilarinboth.Still,thislackof similarityissurprisingforsuchanimportantprotein.whenthed.mel.proteinisblastedagainstotherclosely relateddrosophilidspecies,noneofthemhavehighsimilarity,allhavingidentitiesaroundorlessthan50%.this eithershowsthatthisisanewmutationind.mel.thatcausesforafunctionalhsporitisafunctionalhspinall speciesbutdoesnotneedtoconserve50%ofitspeptidesequencetoremainfunctional.iftheformeristhecase, thenthatsupportsthatthisgeneisapseudogeneind.ere.therearenoothercopiesofasimilargeneind.ere., soidonotbelieveittobeapseudogene;rather,ibelievethatthisgeneisfunctionalinalloftherelatedspecies becauseonlyconservingacertainareaisnecessarytoretainitsfunction.infact,aboutthefirst65aminoacidsin CG7133 sproteinind.mel.showmoderatehomologytomanyotherrelatedspecies,whichsuggeststhisregionis moreselectedforthantheotherresiduesoftheprotein,whichsuggestsitmaybeessentialfortheprotein s interaction.thisgenehas4orthologsincloselyrelatedspecies,meaningitisanewergenethancg14561, CG7139andCG7130. Exon Start End Stopcodon CG7130/NM_
4 HarryQuedenfeld Genomics Dr.Moss Thisgeneisasingleexongenetranscribedintheminusdirection.Thecodingregioncoordinatesinfosmid16is andthestopcodonisfrom Ithasasingleisoform,CG7130 PA.Thisgenehasitsgene ontologydefined.itsmolecularfunctionisdescribedasaproteinthatbindstounfoldedprotein.itfunctionsin HSPbinding.ThusthisproteinisaHSPthatactsasachaperoneproteintofoldunfoldedproteinsthatareeither recentlytranslatedorweredenaturedbyheat.theblastpbetweenthed.mel.cg7130proteinandtheproteini predictedind.ere.yieldsane valueof1e 69anda92%aminoacididentity.SinceCG7130andCG7133arenot highlyparalogous,itisnotlikelythatcg7130cancompensateforalackoffunctionincg7133,whichfurther supportsthatcg7133isnotapseudogene(identity:32%).thispartialidentitycanbeaccountedforbythefact thattheyperformsimilarfunctions,buttheirstructurewouldbedifferentenoughthatonewouldmostlikelynot beabletocompensateforanother.thisgenehas10orthologs,whichmeansthishsphasbeenaroundlonger thancg7133.bothcg7130andcg7133areinthesameproteinfamily,hsp40 ( howevertheirdifferentstructureshavenotyetbeendetermined.atthistimethereisnotenoughinformationto determinewhethertheybindtothesamesubstrates,butihypothesizethattheydonotbecauseoftheirhighlack ofaminoacididentity. CG7130 Exon Start End Stopcodon RpLP0/NM_ ItsgeneontologyisdefinedashavingmolecularfunctionsinrepairingDNA,specificallyasaDNA (apurinicor apyrimidinicsite)lyaseactivity,whichisanenzymethatcutsoutnucleotidestobereplaced.insidethecellitis thoughttobeaconstituentofaribosome,anditsbiologicalfunctionsaretranslation,dnarepair,translational elongationandribosomebiogenesis.therewereprobablymistakesmadeinitsgodefinitionbecauseforitto repairdnaandbeapartofaribosomerequiredifferentstructures,forthesameproteintoperformsuch differentfunctionsisunlikely.itisatwo exongenecodingintheplusdirection.thecoordinatesfortheexons withinmyfosmidare: , ,withthestopat ;itcodesintheplusdirection.In D.mel.itisannotatedtohave3exons,howeverthefirstexonandaportionofthesecondarenottranslatedinto aminoacids,andarethusa3 untranslatedregion.the3 untranslatedregion(and1 st exon)isat , withthesecondexonstartingat11410butnotcodinguntil11450.ind.ere.however,thereisamissense mutationthatcausesforaprematurestartcodon,whichresultsinaprematurestopcodonbeingreadanda highlytruncatedanddissimilarprotein,thusthisusuallyuntranslated1 st exonmustnotbeanexonind.ere. becauserplp0ishighlyconservedamongdrosophilids.d.ere. srplp0contains2exons,thefirstisthesameas thecodingregionofthe2 nd exonind.mel.,andthe3 rd hasidenticalintron/exonborders.sincethe1 st exonind. mel.nolongerexistsasanexonind.ere.,theproteinsarethesamelengthandtheiridentityis98%,withanevalueof0.0.thusthisproteinishighlyconservedbetweenspeciesandperformsthesamefunctionind.ere.asin D.mel.RpLP0has10orthologousgenesindifferentDrosophilaspeciesincludingerecta.Theorthologousgenes arein(species/ortholog):dana\gf10946,dere\gg16244,dgri\gh14667,dmoj\gi13777\,dpse\ga20389, Dsec\GM22429,Dsim\GD15019,Dvir\GJ13582,Dwil\GK20443andDyak\RpLP0.Thetwomorecloselyrelated species,[r.1]dsecand[r.2]danaareshownbelow,bothofwhichalsoexcludethe3 untranslatedexoninorder toconservetheproteinsequence.figurer.3showsdmel srplp0nucleotidesequence,withafirstexonnoncodingandpartial2 nd non codingexon,indicatedbyblack,capitalletters.theseuntranslatedregionsmaybeade novomutationthatchangesthestartcodoninthefirstexon.thusdmelprobablyshowsanewinclusionofthis untranslatedexon,insteadofdereshowinganewexclusionbecauseotherspeciesaresimilartodere,notdmel. [Source:DecoratedFASTA,flybase.org] 4
5 [R.1] HarryQuedenfeld Genomics Dr.Moss [R.2] [R.3] Exon Start End Stopcodon Query:D.mel.RpLP0mRNA,Sbjct:fosmid16 Thegreenboxbelowindicatesthemissensemutationandtheprematurestartcodonintheregionhomologousto the1 st exonind.mel.ifanexon,thisnormallyuntranslatedexonwouldbetranslatedind.ere.,thusitmustno 5
6 HarryQuedenfeld Genomics Dr.Moss longerbeanexonind.ere.becauseconservationofthisproteinisessential,with89%aminoacididentityor higheracrossd.sec.,d.ere.,d.ana.andd.yak. [NCBIBlast2P] Ifthisprematurestartcodoniscodedfor,itwillresultinaproteinthatsharesnohomologywithRpLP0,asitis onlycodedforinthefirstexonandtherplp0geneind.mel.andotherspecieshasanuntranslatedfirstexon. Thisproteinwouldonlybe29aminoacidslongandwouldmostlikelybenonfunctional.InorderforD.ere.to survive,ihypothesizethatthisuntranslatedexonisexcludedintranscriptionofmrnaind.ere.thereisnoother proteinthatlookslikerplp0ind.ere.,sothisimportantgeneisnotapseudogenecopy. Sfp79B/ACG69555 NotonlyisSfp79BaveryshortproteininD.mel.ofonly35aminoacids,itistruncatedto4aminoacidsinD.ere., whicheliminatesanypossibilityofthesfp79bgenecodingforafunctionalproteinind.ere.basedonitslength,it mustbeapseudogene.asisseeninthefigurebelow,thereisamissensemutationthatcausesforapremature startcodon,whichafter9aminoacids,isfollowedbyastopcodon(taa).itwouldhavebeenasingleexongene intheplusdirection,iffunctional.sfp79bisaputativeseminalfluidproteinind.mel.becauseithasapredicted structuresimilartootherseminalfluidproteins(thusisnotconfirmed).blastpofthe4aminoacidsequence againstd.mel.yieldednosignificantresults.thisgeneisnotanywhereelseind.ere.,butsincetherearemany seminalfluidproteinsthatcancompensateforitsabsence,itdoesnotaffectfertility.itcouldalsobea pseudogeneind.mel.aswellbecauseitisshort(<100aminoacids)andhasnotbeencurated. 6
7 HarryQuedenfeld Genomics Dr.Moss 7 Query:D.mel. Sbjct:fosmid16(Tool:NCBI sblast2n) Msopa/NM_ Thisfeatureisprobablyapseudogenebecauseitisonly63aminoacidslong.TheD.mel.msopamayalsobea pseudogenebecauseitwasannotatedbylookingfororfsthatmightcodeforproteinsthatlooklikeacertain proteinfamily,andhasnotbeencurated.d.mel. smsopaproteinisonly83aminoacidslong.regardlessofits legitimacyasageneind.mel.,ind.ere.,msopahasa20aminoaciddeletioncomparedtod.mel.,issignificantly shorterandonlyhasa46%identitywithd.mel. s.thesethreefactspointtoitsbeingapseudogene.thisgeneis thoughttobeinvolvedinimmunityordefense,howevernothingisspecifiedbecausethisgenehasnotbeen researched.theproteinisnotknowntoexist,andat83aminoacidsitisunlikelytobeafunctionalgene.foritto existas63aminoacidsisevenmoreunlikely.msopadoesnotoccuranywhereelseinthed.ere.genome,the highestpercentidentityistheregioninmyfosmidwith46%sharedaminoacids.furthermore,msopaisnot conservedbetweenotherspecies(blastp68%identitywithdrosophilasechellia46%indrosophilayakuba,ncbi BlastP).Withsuchalackofconservationandsuchashortlength,itisprobablyapseudogene.Thisevidenceis strongerthanitsinferredexistenceduetopredictedstructuresimilaritybecausetheresearchersdidnotaccount foritslackofconservationinspecies.denovomutationsarehighlyunlikely.thereare4orthologstomsopain theotherdrosophilidspecies:dere\gg16245,dsec\gm22431,dsim\gd15020anddyak\ge22603,howeverin Dsimitis122aminoacidslong,almosttwiceaslongasDere ssupposedmsopaortholog.idonotagreethat GG16245isfunctionalinD.ere.orthatGM22431issimilarinD.sec.becauseitsproteinisonly67aminoacids long,giventhemedianpeptidelengthinalldrosophilidspeciesis373. [ GENSCAN Genscanpredicted6featuresformymaskedfosmid,threeofwhichweresingle exon.genscanpredictedsingleexongenesratherwell.thefirstexampleiscg14561,inwhichgenscanpredictedaregioninmyfosmidthat, whenblastedagainstmelanogaster,turnedouttobethemiddleofcg14561;thestartcodonwaslateandthe stopcodonwasearly,probablyinthewrongreadingframe.itpredictedthethirdexonofcg7139tobeadoubleexongene,whichisunderstandablebecauseitbeginswithastartcodon(asdoesisoformb)andthereisasplice junction(gt)rightbeforethestopcodon(tga),the T ofthestopcodonisthe T ofthe GT splicejunction. Forthethirdfeature,GenscanalsopredictedasingleexongeneinthemiddleofCG7133,butitonlytakesup aboutonethirdoftheactualgene.thisprobablyhappenedbecausegenscanreadinthewrongframe,causinga latestartcodonandaprematurestop.thefourthpredictedfeaturewassingle exonandturnedouttobe homologoustothecenterofcg7130,onlyslightlyshortonbothendsprobablyduetoreadingtheincorrectframe. Thefifthpredictedfeature,whenextractedfromfosmid16andblastedagainstD.mel.,ishighlyhomologousto therplp0gene,onlymissingthefirstexonandpredictingithastwo.genscandidpredictthisaccuratelyby overlookingthefirstexon,whichcontainsamutationforastartcodon.genscanmostlikelyskippedthisbecause theresultingproteinwouldbetooshort,thusitendeduppredictingrplp0correctly.thesixthfeaturepredicted atwo exongenethatspansfrom Whenthesebasesareextractedfromfosmid16,theyare homologouswithregionsaftermsopabuttherearenogenesind.melanogasterinanyofthehomologous regions.thegenscancoordinatesgivenmatchedupwithsplicejunctionsgtandag,butthereisastopcodonin thereadingframeandcutsthefirstpredictedexonshort,withnosplicejunctionnearby.duetothefactthat Genscanpredictedthefirstexonincorrectlyandtheexonsareverydistantfromeachother,thereisnotstrong
8 HarryQuedenfeld Genomics Dr.Moss evidenceforanewgenehereind.ere.thereare4intronspredictedtobebetweenthetwoexons,butthereare 10kbpbetweenthefirstexonandfirstintron,whichisnotlikelytooccurbecauseexonsandintronsareusually continuous.sincenoneofthegenscanpredictedgenesmatchedupwiththerealgenesperfectly,itcannotbe solelyreliedon;however,itprovestobeagoodstartingpoint.blastingtheregionofthepredictedfeatureagainst D.mel.willresultinhomologousregions,andGenomeViewwillshowifitispartofanygene,whichishelpful. CLUSTALanalysis RpLP0analysis AstheimagefromCLUSTALW2illustrates,RpLP0isahighlyconservedproteinbecauseithasanessentialfunction asaconstituentofaribosome.ribosomesarenecessaryineverycellallthetime,iftheymutatethecellwilldie.if anorganismhasamutationinrplp0,itwilllikelydie.thereare5mutationsthroughoutthe4organisms,butthey allresultinasimilaraminoacid.thismeanstheyhavethesamechargeand/orpolarity,sotheprotein sfunction wouldnotbealteredsignificantlybysuchamutation.key:fbpp :d.melanogasterrplp0,fbpp : D.sechelliaGM22429,FBpp :D.ananassaeGF10946,RpLP0 PA_peptide:D.ere. sorthologousproteinof RpLP0fromfosmid16,GG UpstreamRegions Thesealignmentsareperformedusingthe1,000bp5 upstreamfromrplp0ind.mel.anditsorthologsind.ere, D.sec.andD.ana.ThereisaconservedTATAboxinboththeD.ere.andD.mel.5 upstreamregions,shownby theredboxbelow.thetataboxisat inD.ere.and inD.mel.TATAboxusedhereisdefinedas anysequenceupstreamoftheinitiatorwith5of6nucleotidesconformingtotheconsensustataaa.(locations arebasedonthe1000bpextract5 ofrplp0oritsorthologs). 8
9 HarryQuedenfeld Genomics Dr.Moss ThefiguretotheleftillustratesthefrequencyofbaseswithinaTATAbox.[Source: bin/jaspar_db.pl] D.ana.alsohasaTATAbox[shownbelow]within20bpofD.ere. sand60ofd.mel. s,howeverd.sec.doesnot possessaqualitytatabox(ithasmorethanonedifferentbasefromtataaa)withinitsequivalent5 regionof DNA,thusitlikelydoesnothaveone.TheTATAboxlocationisat D.ana.Isclearlytheleastsimilaramongthefour;however,D.melandD.ere.showahighersimilarityinthe 1,000bpbeforeRpLP0thantheothersdotothem.ThisisnotinagreementwithwhatisexpectedbecauseD.sec. ismorecloselyrelatedtod.mel.thand.ere.is,howeverthesealignmentsattestthatthis5 upstreamregion havehigherhomologybetweend.mel.andd.ere.accordingtotataboxlocations.thefirst200basepairsofd. sec.andd.ana.notalignwithd.mel.andd.ere.becausethisregionwasnotconservedenoughtoalignbetween 4speciesperfectly,TheremighthavebeenaninsertioninD.secandD.ana.thatdoesnotalignwiththefirst250 basepairsoftheothertwospecies.d.ana.isclearlythemostdistantrelativeoftheother3species,asisvisible withthe showcolors optioninclustal. TheinitiatorCCATTGwasfoundinallfoursequencesinthealignment,indicatedbytheredboxbelow.D.ana.has amissensemutationthatleadsthectobereplacedwithag,butthisisstillaninitiationsequence. 9
10 HarryQuedenfeld Genomics Dr.Moss Adownstreampromoterelement(DPE)isany6nucleotidesequenceatexactly+28to+33with5of6nucleotides conformingtothedpefunctionalrangeseta/g/t C/G A/T C/T A/C/G C/T.Asseeninthefigurebelow, therearenodpeinanyofthespeciesbecausethereisacrichregionhere,whichcausesthethirdnucleotideto alwaysbeacandeliminatesthechanceforadperegion.thed.ana.sequencegcttcawouldworkifthelasta wasat,sod.ana.alsoprobablydoesnothaveadpe oftheinitiator.[blackboxbelowindicates28 35bp aftertheinitiator,wheredpesarepossible]. [Sourceforknownsequences: biology.ucsd.edu/labs/kadonaga/dcpd.html] Repeats [FromRepeatMasker] Mostoffosmid16ishighcomplexity,despitethelast20kbpbeingvacantofanygenesinD.mel.andD.ere.,only 6.31%isrepetitive,withnearly5%ofthatbeinginterspersedrepeats,ofwhichweremostlyDNAtransposons. GiventhattransposonsaccountforahigherpercentageoftheentireDrosophilagenomeonaverage,thelow percentageofdnatransposonsevidencesthatitisapossiblegene richarea.thereisonly0.15%ofmyfosmid composedofretroelements,whicharealsotypicallymuchmorecommon,whichshowsthisareahasbeen conserved.itisexpectedthat,anyorganismsthathadmanytransposonsorretroelementwouldhavemutations inthisareaandincreasetheirchanceofmutatingfatally.theorganismsthatliveddonothavemutationsinthese genescausedbytransposonelements,soweexpectlessrepeatsinagene richregion.however,thefirst20,000 bpoffosmid16actuallycontainnearlytwiceasmanyrepeats[fig2.2].thus,itisnotreliabletojudgewhethera regionisgenerichbytheproportionofrepeatsbecauseinthiscase,thegene richregionwithinalargerarea actuallyhasalmostalloftherepeats;2425of2526repeatswerefoundinthegene richregion.thisshowsthat transposonsandotherrepeatableelementsarepresentbutdonotcauseanyfatalmutations,andshedslightthat thecodingregionsmaynotbeexceptionallystable.[areasinyellowrefertospecificsmentioned] [2.1] Summary: ================================================== file name: RM2_Fosmid16.txt_ sequences: 1 10
11 total length: bp (40000 bp excl N/X-runs) GC level: % bases masked: 2526 bp ( 6.31 %) ================================================== number of length percentage elements* occupied of sequence Retroelements 1 60 bp 0.15 % SINEs: 0 0 bp 0.00 % Penelope 0 0 bp 0.00 % LINEs: 1 60 bp 0.15 % CRE/SLACS 0 0 bp 0.00 % L2/CR1/Rex 0 0 bp 0.00 % R1/LOA/Jockey 0 0 bp 0.00 % R2/R4/NeSL 0 0 bp 0.00 % RTE/Bov-B 0 0 bp 0.00 % L1/CIN4 0 0 bp 0.00 % LTR elements: 0 0 bp 0.00 % BEL/Pao 0 0 bp 0.00 % Ty1/Copia 0 0 bp 0.00 % Gypsy/DIRS1 0 0 bp 0.00 % Retroviral 0 0 bp 0.00 % HarryQuedenfeld Genomics Dr.Moss DNA transposons bp 4.75 % hobo-activator 0 0 bp 0.00 % Tc1-IS630-Pogo 0 0 bp 0.00 % En-Spm 0 0 bp 0.00 % MuDR-IS bp 0.00 % PiggyBac 0 0 bp 0.00 % Tourist/Harbinger 0 0 bp 0.00 % Other (Mirage, bp 1.73 % P-element, Transib) Rolling-circles 0 0 bp 0.00 % Unclassified: 0 0 bp 0.00 % Total interspersed repeats: 1959 bp 4.90 % Small RNA: 0 0 bp 0.00 % Satellites: 0 0 bp 0.00 % Simple repeats: bp 0.61 % Low complexity: bp 0.80 % [2.2] Summary: ================================================== file name: RM2sequpload_ sequences: 1 total length: bp (20000 bp excl N/X-runs) GC level: % bases masked: 2425 bp ( %) ================================================== 11
12 number of length percentage elements* occupied of sequence Retroelements 1 60 bp 0.30 % SINEs: 0 0 bp 0.00 % Penelope 0 0 bp 0.00 % LINEs: 1 60 bp 0.30 % CRE/SLACS 0 0 bp 0.00 % L2/CR1/Rex 0 0 bp 0.00 % R1/LOA/Jockey 0 0 bp 0.00 % R2/R4/NeSL 0 0 bp 0.00 % RTE/Bov-B 0 0 bp 0.00 % L1/CIN4 0 0 bp 0.00 % LTR elements: 0 0 bp 0.00 % BEL/Pao 0 0 bp 0.00 % Ty1/Copia 0 0 bp 0.00 % Gypsy/DIRS1 0 0 bp 0.00 % Retroviral 0 0 bp 0.00 % HarryQuedenfeld Genomics Dr.Moss DNA transposons bp 9.21 % hobo-activator 0 0 bp 0.00 % Tc1-IS630-Pogo 0 0 bp 0.00 % En-Spm 0 0 bp 0.00 % MuDR-IS bp 0.00 % PiggyBac 0 0 bp 0.00 % Tourist/Harbinger 0 0 bp 0.00 % Other (Mirage, bp 3.46 % P-element, Transib) Rolling-circles 0 0 bp 0.00 % Unclassified: 0 0 bp 0.00 % Total interspersed repeats: 1903 bp 9.52 % Small RNA: 0 0 bp 0.00 % Satellites: 0 0 bp 0.00 % Simple repeats: bp 1.04 % Low complexity: bp 1.56 % ================================================== Synteny Key:[Leftmost]verticalredlineatapproximately21735kbpinD.ere.isthestartoffosmid16.Fromthe3 isthe entirecodingregion,all3 aftermsopaisnon codingdnainbothd.mel.andd.ere. [1]OrthologybetweenDereScaffold_4784(top)andDmelChr3L(bottom) 12
13 HarryQuedenfeld Genomics Dr.Moss [flybase.orggbrowser] ThisshowshighsyntenybetweenDereandDmelbecauseallofthegenesinDereareinthesameorderasDmel s, codeinthesamedirectionandaresimilardistancesapart. [2]Fosmid16regioninD.ere. [UCSCGenomeBrowser: [3]Fosmid16regioninD.mel. 13
14 HarryQuedenfeld Genomics Dr.Moss [UCSCGenomeBrowser: Theaboveimagesshowhomology[1]andrepeatelementsinDereandDmel[2,3].Fosmid16isonchromosome 3LinD.melanogasterandScaffold_4784inDere.Syntenyhasbeenpreservedbecausethegenesonfosmid16are allfromthesameregionofthed.melanogastergenome;thatis,theyareinthesameorder,spacedsimilarlyand onthesamechromosomeinbothd.ere.andd.mel.thusthereisnoevidenceofanychromosomalmutations suchasinversionsortranspositionsthathaveoccurredsincethesetwospeciessplit.therearerepeatspresentin Dere[2]betweenthefirstandsecondexon(minusdirection)thatarenotpresentinDmel(bluebox[2]),which suggestsatransposableelementinserteditselfbetweenthoseexons(thefirsttwoincg7139 PA).Thereisalso anotherinsertionofrepeatsinderebetweenrplp0andmsopthatisnotpresentindmel(redboxin[2]).the orangeboxin[3]showsrepeatsthatintervenecg1739andcg1733indmel,butcomeaftercg7133indere, whichevidencesminordnarearrangement;however,thisdoesnotalterthesyntenyofgenesinthisregion. 14
Classification of repetitive elements based on the analysis of protein domains. Pavel Neumann May 2018
 Classification of repetitive elements based on the analysis of protein domains Pavel Neumann May 2018 A unified classification system for eukaryotic transposable elements (Wicker et al. 2007) Repbase classification
Classification of repetitive elements based on the analysis of protein domains Pavel Neumann May 2018 A unified classification system for eukaryotic transposable elements (Wicker et al. 2007) Repbase classification
Annotation of Drosophila grimashawi Contig12
 Annotation of Drosophila grimashawi Contig12 Marshall Strother April 27, 2009 Contents 1 Overview 3 2 Genes 3 2.1 Genscan Feature 12.4............................................. 3 2.1.1 Genome Browser:
Annotation of Drosophila grimashawi Contig12 Marshall Strother April 27, 2009 Contents 1 Overview 3 2 Genes 3 2.1 Genscan Feature 12.4............................................. 3 2.1.1 Genome Browser:
Supplementary Information for: The genome of the extremophile crucifer Thellungiella parvula
 Supplementary Information for: The genome of the extremophile crucifer Thellungiella parvula Maheshi Dassanayake 1,9, Dong-Ha Oh 1,9, Jeffrey S. Haas 1,2, Alvaro Hernandez 3, Hyewon Hong 1,4, Shahjahan
Supplementary Information for: The genome of the extremophile crucifer Thellungiella parvula Maheshi Dassanayake 1,9, Dong-Ha Oh 1,9, Jeffrey S. Haas 1,2, Alvaro Hernandez 3, Hyewon Hong 1,4, Shahjahan
M.B. Zhou, X.M. Liu, and D.Q. Tang. Corresponding author: D.Q. Tang
 Transposable elements in Phyllostachys pubescens (Poaceae) genome survey sequences and the full-length cdna sequences, and their association with simple-sequence repeats M.B. Zhou, X.M. Liu, and D.Q. Tang
Transposable elements in Phyllostachys pubescens (Poaceae) genome survey sequences and the full-length cdna sequences, and their association with simple-sequence repeats M.B. Zhou, X.M. Liu, and D.Q. Tang
Tandem repeat 16,225 20,284. 0kb 5kb 10kb 15kb 20kb 25kb 30kb 35kb
 Overview Fosmid XAAA112 consists of 34,783 nucleotides. Blat results indicate that this fosmid has significant identity to the 2R chromosome of D.melanogaster. Evidence suggests that fosmid XAAA112 contains
Overview Fosmid XAAA112 consists of 34,783 nucleotides. Blat results indicate that this fosmid has significant identity to the 2R chromosome of D.melanogaster. Evidence suggests that fosmid XAAA112 contains
GEP Annotation Report
 GEP Annotation Report Note: For each gene described in this annotation report, you should also prepare the corresponding GFF, transcript and peptide sequence files as part of your submission. Student name:
GEP Annotation Report Note: For each gene described in this annotation report, you should also prepare the corresponding GFF, transcript and peptide sequence files as part of your submission. Student name:
Genome sequence of Plasmopara viticola and insight into the pathogenic mechanism
 Genome sequence of Plasmopara viticola and insight into the pathogenic mechanism Ling Yin 1,3,, Yunhe An 1,2,, Junjie Qu 3,, Xinlong Li 1, Yali Zhang 1, Ian Dry 5, Huijun Wu 2*, Jiang Lu 1,4** 1 College
Genome sequence of Plasmopara viticola and insight into the pathogenic mechanism Ling Yin 1,3,, Yunhe An 1,2,, Junjie Qu 3,, Xinlong Li 1, Yali Zhang 1, Ian Dry 5, Huijun Wu 2*, Jiang Lu 1,4** 1 College
Carri-Lyn Mead Thursday, January 13, 2005 Terry Fox Laboratory, Dr. Dixie Mager
 Investigating Trends in Transposable Element Insertion within Regulatory Regions Carri-Lyn Mead cmead@bcgsc.ca Thursday, January 13, 2005 Terry Fox Laboratory, Dr. Dixie Mager Outline Transposable Element
Investigating Trends in Transposable Element Insertion within Regulatory Regions Carri-Lyn Mead cmead@bcgsc.ca Thursday, January 13, 2005 Terry Fox Laboratory, Dr. Dixie Mager Outline Transposable Element
Outline. Genome Evolution. Genome. Genome Architecture. Constraints on Genome Evolution. New Evolutionary Synthesis 11/8/16
 Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Biology 644: Bioinformatics
 A stochastic (probabilistic) model that assumes the Markov property Markov property is satisfied when the conditional probability distribution of future states of the process (conditional on both past
A stochastic (probabilistic) model that assumes the Markov property Markov property is satisfied when the conditional probability distribution of future states of the process (conditional on both past
Outline. Genome Evolution. Genome. Genome Architecture. Constraints on Genome Evolution. New Evolutionary Synthesis 11/1/18
 Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Genome Evolution Outline 1. What: Patterns of Genome Evolution Carol Eunmi Lee Evolution 410 University of Wisconsin 2. Why? Evolution of Genome Complexity and the interaction between Natural Selection
Markov Models & DNA Sequence Evolution
 7.91 / 7.36 / BE.490 Lecture #5 Mar. 9, 2004 Markov Models & DNA Sequence Evolution Chris Burge Review of Markov & HMM Models for DNA Markov Models for splice sites Hidden Markov Models - looking under
7.91 / 7.36 / BE.490 Lecture #5 Mar. 9, 2004 Markov Models & DNA Sequence Evolution Chris Burge Review of Markov & HMM Models for DNA Markov Models for splice sites Hidden Markov Models - looking under
Academy of Agricultural Sciences, Anyang , Henan, China. 2 BGI-Shenzhen,
 The draft genome of a diploid cotton Gossypium raimondii Kunbo Wang 1,6, Zhiwen Wang 2,6, Fuguang Li 1,6, Wuwei Ye 1,6, Junyi Wang 2,6, Guoli Song 1,6, Zhen Yue 2, Lin Cong 2, Haihong Shang 1, Shilin Zhu
The draft genome of a diploid cotton Gossypium raimondii Kunbo Wang 1,6, Zhiwen Wang 2,6, Fuguang Li 1,6, Wuwei Ye 1,6, Junyi Wang 2,6, Guoli Song 1,6, Zhen Yue 2, Lin Cong 2, Haihong Shang 1, Shilin Zhu
PLNT2530 (2018) Unit 5 Genomes: Organization and Comparisons
 PLNT2530 (2018) Unit 5 Genomes: Organization and Comparisons Unless otherwise cited or referenced, all content of this presenataion is licensed under the Creative Commons License Attribution Share-Alike
PLNT2530 (2018) Unit 5 Genomes: Organization and Comparisons Unless otherwise cited or referenced, all content of this presenataion is licensed under the Creative Commons License Attribution Share-Alike
Frequently Asked Questions (FAQs)
 Frequently Asked Questions (FAQs) Q1. What is meant by Satellite and Repetitive DNA? Ans: Satellite and repetitive DNA generally refers to DNA whose base sequence is repeated many times throughout the
Frequently Asked Questions (FAQs) Q1. What is meant by Satellite and Repetitive DNA? Ans: Satellite and repetitive DNA generally refers to DNA whose base sequence is repeated many times throughout the
Chapter 18 Active Reading Guide Genomes and Their Evolution
 Name: AP Biology Mr. Croft Chapter 18 Active Reading Guide Genomes and Their Evolution Most AP Biology teachers think this chapter involves an advanced topic. The questions posed here will help you understand
Name: AP Biology Mr. Croft Chapter 18 Active Reading Guide Genomes and Their Evolution Most AP Biology teachers think this chapter involves an advanced topic. The questions posed here will help you understand
Python genome Supplementary Information 1. SUPPLEMENTARY INFORMATION Supporting Information Corrected July 17, 1. SUPPLEMENTARY METHODS
 Python genome Supplementary Information 1 SUPPLEMENTARY INFORMATION Supporting Information Corrected July 17, 2014 1. SUPPLEMENTARY METHODS 1.1 Python Genome Sequencing A single Python molurus bivittatus
Python genome Supplementary Information 1 SUPPLEMENTARY INFORMATION Supporting Information Corrected July 17, 2014 1. SUPPLEMENTARY METHODS 1.1 Python Genome Sequencing A single Python molurus bivittatus
Stochastic processes and
 Stochastic processes and Markov chains (part II) Wessel van Wieringen w.n.van.wieringen@vu.nl wieringen@vu nl Department of Epidemiology and Biostatistics, VUmc & Department of Mathematics, VU University
Stochastic processes and Markov chains (part II) Wessel van Wieringen w.n.van.wieringen@vu.nl wieringen@vu nl Department of Epidemiology and Biostatistics, VUmc & Department of Mathematics, VU University
TE content correlates positively with genome size
 TE content correlates positively with genome size Mb 3000 Genomic DNA 2500 2000 1500 1000 TE DNA Protein-coding DNA 500 0 Feschotte & Pritham 2006 Transposable elements. Variation in gene numbers cannot
TE content correlates positively with genome size Mb 3000 Genomic DNA 2500 2000 1500 1000 TE DNA Protein-coding DNA 500 0 Feschotte & Pritham 2006 Transposable elements. Variation in gene numbers cannot
Prof. Christian MICHEL
 CIRCULAR CODES IN GENES AND GENOMES - 2013 - Prof. Christian MICHEL Theoretical Bioinformatics ICube University of Strasbourg, CNRS France c.michel@unistra.fr http://dpt-info.u-strasbg.fr/~c.michel/ Prof.
CIRCULAR CODES IN GENES AND GENOMES - 2013 - Prof. Christian MICHEL Theoretical Bioinformatics ICube University of Strasbourg, CNRS France c.michel@unistra.fr http://dpt-info.u-strasbg.fr/~c.michel/ Prof.
O 3 O 4 O 5. q 3. q 4. Transition
 Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Interpolated Markov Models for Gene Finding. BMI/CS 776 Spring 2015 Colin Dewey
 Interpolated Markov Models for Gene Finding BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2015 Colin Dewey cdewey@biostat.wisc.edu Goals for Lecture the key concepts to understand are the following the
Interpolated Markov Models for Gene Finding BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2015 Colin Dewey cdewey@biostat.wisc.edu Goals for Lecture the key concepts to understand are the following the
Lecture 20 DNA Repair and Genetic Recombination (Chapter 16 and Chapter 15 Genes X)
 Lecture 20 DNA Repair and Genetic Recombination (Chapter 16 and Chapter 15 Genes X) Retrotransposons of the viral superfamily are transposons that mobilize via an RNA that does not form an infectious particle.
Lecture 20 DNA Repair and Genetic Recombination (Chapter 16 and Chapter 15 Genes X) Retrotransposons of the viral superfamily are transposons that mobilize via an RNA that does not form an infectious particle.
Lecture 3: Markov chains.
 1 BIOINFORMATIK II PROBABILITY & STATISTICS Summer semester 2008 The University of Zürich and ETH Zürich Lecture 3: Markov chains. Prof. Andrew Barbour Dr. Nicolas Pétrélis Adapted from a course by Dr.
1 BIOINFORMATIK II PROBABILITY & STATISTICS Summer semester 2008 The University of Zürich and ETH Zürich Lecture 3: Markov chains. Prof. Andrew Barbour Dr. Nicolas Pétrélis Adapted from a course by Dr.
Scalable and reproducible genome analysis in the age of next-generation genome sequencing
 Graduate Theses and Dissertations Graduate College 2016 Scalable and reproducible genome analysis in the age of next-generation genome sequencing Daniel Scott Standage Iowa State University Follow this
Graduate Theses and Dissertations Graduate College 2016 Scalable and reproducible genome analysis in the age of next-generation genome sequencing Daniel Scott Standage Iowa State University Follow this
The nature of genomes. Viral genomes. Prokaryotic genome. Nonliving particle. DNA or RNA. Compact genomes with little spacer DNA
 The nature of genomes Genomics: study of structure and function of genomes Genome size variable, by orders of magnitude number of genes roughly proportional to genome size Plasmids symbiotic DNA molecules,
The nature of genomes Genomics: study of structure and function of genomes Genome size variable, by orders of magnitude number of genes roughly proportional to genome size Plasmids symbiotic DNA molecules,
Genomes Comparision via de Bruijn graphs
 Genomes Comparision via de Bruijn graphs Student: Ilya Minkin Advisor: Son Pham St. Petersburg Academic University June 4, 2012 1 / 19 Synteny Blocks: Algorithmic challenge Suppose that we are given two
Genomes Comparision via de Bruijn graphs Student: Ilya Minkin Advisor: Son Pham St. Petersburg Academic University June 4, 2012 1 / 19 Synteny Blocks: Algorithmic challenge Suppose that we are given two
Department of Forensic Psychiatry, School of Medicine & Forensics, Xi'an Jiaotong University, Xi'an, China;
 Title: Evaluation of genetic susceptibility of common variants in CACNA1D with schizophrenia in Han Chinese Author names and affiliations: Fanglin Guan a,e, Lu Li b, Chuchu Qiao b, Gang Chen b, Tinglin
Title: Evaluation of genetic susceptibility of common variants in CACNA1D with schizophrenia in Han Chinese Author names and affiliations: Fanglin Guan a,e, Lu Li b, Chuchu Qiao b, Gang Chen b, Tinglin
HIGH PERFORMANCE CLUSTER AND GRID COMPUTING SOLUTIONS FOR SCIENCE UMESHKUMAR KESWANI. Presented to the Faculty of the Graduate School of
 HIGH PERFORMANCE CLUSTER AND GRID COMPUTING SOLUTIONS FOR SCIENCE By UMESHKUMAR KESWANI Presented to the Faculty of the Graduate School of The University of Texas at Arlington in Partial Fulfillment of
HIGH PERFORMANCE CLUSTER AND GRID COMPUTING SOLUTIONS FOR SCIENCE By UMESHKUMAR KESWANI Presented to the Faculty of the Graduate School of The University of Texas at Arlington in Partial Fulfillment of
The Gene The gene; Genes Genes Allele;
 Gene, genetic code and regulation of the gene expression, Regulating the Metabolism, The Lac- Operon system,catabolic repression, The Trp Operon system: regulating the biosynthesis of the tryptophan. Mitesh
Gene, genetic code and regulation of the gene expression, Regulating the Metabolism, The Lac- Operon system,catabolic repression, The Trp Operon system: regulating the biosynthesis of the tryptophan. Mitesh
Introduction to Hidden Markov Models for Gene Prediction ECE-S690
 Introduction to Hidden Markov Models for Gene Prediction ECE-S690 Outline Markov Models The Hidden Part How can we use this for gene prediction? Learning Models Want to recognize patterns (e.g. sequence
Introduction to Hidden Markov Models for Gene Prediction ECE-S690 Outline Markov Models The Hidden Part How can we use this for gene prediction? Learning Models Want to recognize patterns (e.g. sequence
Special Topics on Genetics
 ARISTOTLE UNIVERSITY OF THESSALONIKI OPEN COURSES Section 9: Transposable elements Drosopoulou E License The offered educational material is subject to Creative Commons licensing. For educational material,
ARISTOTLE UNIVERSITY OF THESSALONIKI OPEN COURSES Section 9: Transposable elements Drosopoulou E License The offered educational material is subject to Creative Commons licensing. For educational material,
Proteomics. 2 nd semester, Department of Biotechnology and Bioinformatics Laboratory of Nano-Biotechnology and Artificial Bioengineering
 Proteomics 2 nd semester, 2013 1 Text book Principles of Proteomics by R. M. Twyman, BIOS Scientific Publications Other Reference books 1) Proteomics by C. David O Connor and B. David Hames, Scion Publishing
Proteomics 2 nd semester, 2013 1 Text book Principles of Proteomics by R. M. Twyman, BIOS Scientific Publications Other Reference books 1) Proteomics by C. David O Connor and B. David Hames, Scion Publishing
Introduction to Hidden Markov Models (HMMs)
 Introduction to Hidden Markov Models (HMMs) But first, some probability and statistics background Important Topics 1.! Random Variables and Probability 2.! Probability Distributions 3.! Parameter Estimation
Introduction to Hidden Markov Models (HMMs) But first, some probability and statistics background Important Topics 1.! Random Variables and Probability 2.! Probability Distributions 3.! Parameter Estimation
Supplemental Tables for Genomic Legacy of the African Cheetah, Acinonyx jubatus
 Supplemental Tables for Genomic Legacy of the African Cheetah, Acinonyx jubatus 1 List of Tables Table S1: Sequenced cheetah reads for de novo genome assembly 3 Table S2: Re-sequenced cheetah reads for
Supplemental Tables for Genomic Legacy of the African Cheetah, Acinonyx jubatus 1 List of Tables Table S1: Sequenced cheetah reads for de novo genome assembly 3 Table S2: Re-sequenced cheetah reads for
23/01/2018. PiRATE: a Pipeline to Retrieve and Annotate TEs of non-model organisms. Transposable elements (TEs) Impact of TEs on genomes
 Transposable elements () PiRATE: a Pipeline to Retrieve and Annotate of non-model organisms are DNAsequences able to move (= transposition) into the host genome of eucaryotic and procaryotic organisms
Transposable elements () PiRATE: a Pipeline to Retrieve and Annotate of non-model organisms are DNAsequences able to move (= transposition) into the host genome of eucaryotic and procaryotic organisms
Genome Assembly. Sequencing Output. High Throughput Sequencing
 Genome High Throughput Sequencing Sequencing Output Example applications: Sequencing a genome (DNA) Sequencing a transcriptome and gene expression studies (RNA) ChIP (chromatin immunoprecipitation) Example
Genome High Throughput Sequencing Sequencing Output Example applications: Sequencing a genome (DNA) Sequencing a transcriptome and gene expression studies (RNA) ChIP (chromatin immunoprecipitation) Example
Lecture 15: Programming Example: TASEP
 Carl Kingsford, 0-0, Fall 0 Lecture : Programming Example: TASEP The goal for this lecture is to implement a reasonably large program from scratch. The task we will program is to simulate ribosomes moving
Carl Kingsford, 0-0, Fall 0 Lecture : Programming Example: TASEP The goal for this lecture is to implement a reasonably large program from scratch. The task we will program is to simulate ribosomes moving
A unified classification system for eukaryotic transposable elements
 Nature Reviews Genetics AOP, published online 6 November 2007; doi:10.1038/nrg2165 Perspectives g u i d e l i n e s A unified classification system for eukaryotic transposable elements Thomas Wicker, François
Nature Reviews Genetics AOP, published online 6 November 2007; doi:10.1038/nrg2165 Perspectives g u i d e l i n e s A unified classification system for eukaryotic transposable elements Thomas Wicker, François
RNA- seq read mapping
 RNA- seq read mapping Pär Engström SciLifeLab RNA- seq workshop October 216 IniDal steps in RNA- seq data processing 1. Quality checks on reads 2. Trim 3' adapters (opdonal (for species with a reference
RNA- seq read mapping Pär Engström SciLifeLab RNA- seq workshop October 216 IniDal steps in RNA- seq data processing 1. Quality checks on reads 2. Trim 3' adapters (opdonal (for species with a reference
November 13, 2009 Bioe 109 Fall 2009 Lecture 20 Evolutionary Genomics
 November 13, 2009 Bioe 109 Fall 2009 Lecture 20 Evolutionary Genomics - we have now entered the genomics age - the number of complete genomes continues to rise rapidly each year, now numbering about 200.
November 13, 2009 Bioe 109 Fall 2009 Lecture 20 Evolutionary Genomics - we have now entered the genomics age - the number of complete genomes continues to rise rapidly each year, now numbering about 200.
Towards More Effective Formulations of the Genome Assembly Problem
 Towards More Effective Formulations of the Genome Assembly Problem Alexandru Tomescu Department of Computer Science University of Helsinki, Finland DACS June 26, 2015 1 / 25 2 / 25 CENTRAL DOGMA OF BIOLOGY
Towards More Effective Formulations of the Genome Assembly Problem Alexandru Tomescu Department of Computer Science University of Helsinki, Finland DACS June 26, 2015 1 / 25 2 / 25 CENTRAL DOGMA OF BIOLOGY
Losing identity: structural diversity of transposable elements belonging to different classes in the genome of Anopheles gambiae
 Fernández-Medina et al. BMC Genomics 2012, 13:272 RESEARCH ARTICLE Losing identity: structural diversity of transposable elements belonging to different classes in the genome of Anopheles gambiae Rita
Fernández-Medina et al. BMC Genomics 2012, 13:272 RESEARCH ARTICLE Losing identity: structural diversity of transposable elements belonging to different classes in the genome of Anopheles gambiae Rita
Stochastic processes and Markov chains (part II)
 Stochastic processes and Markov chains (part II) Wessel van Wieringen w.n.van.wieringen@vu.nl Department of Epidemiology and Biostatistics, VUmc & Department of Mathematics, VU University Amsterdam, The
Stochastic processes and Markov chains (part II) Wessel van Wieringen w.n.van.wieringen@vu.nl Department of Epidemiology and Biostatistics, VUmc & Department of Mathematics, VU University Amsterdam, The
Principles of Genetics
 Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
1/22/13. Example: CpG Island. Question 2: Finding CpG Islands
 I529: Machine Learning in Bioinformatics (Spring 203 Hidden Markov Models Yuzhen Ye School of Informatics and Computing Indiana Univerty, Bloomington Spring 203 Outline Review of Markov chain & CpG island
I529: Machine Learning in Bioinformatics (Spring 203 Hidden Markov Models Yuzhen Ye School of Informatics and Computing Indiana Univerty, Bloomington Spring 203 Outline Review of Markov chain & CpG island
( 1 ) Show that P ( a, b + c ), Q ( b, c + a ) and R ( c, a + b ) are collinear.
 Problems 01 - POINT Page 1 ( 1 ) Show that P ( a, b + c ), Q ( b, c + a ) and R ( c, a + b ) are collinear. ( ) Prove that the two lines joining the mid-points of the pairs of opposite sides and the line
Problems 01 - POINT Page 1 ( 1 ) Show that P ( a, b + c ), Q ( b, c + a ) and R ( c, a + b ) are collinear. ( ) Prove that the two lines joining the mid-points of the pairs of opposite sides and the line
Stochastic processes and
 Stochastic processes and Markov chains (part I) Wessel van Wieringen w.n.van.wieringen@vu.nl wieringen@vu nl Department of Epidemiology and Biostatistics, VUmc & Department of Mathematics, VU University
Stochastic processes and Markov chains (part I) Wessel van Wieringen w.n.van.wieringen@vu.nl wieringen@vu nl Department of Epidemiology and Biostatistics, VUmc & Department of Mathematics, VU University
Pattern Matching (Exact Matching) Overview
 CSI/BINF 5330 Pattern Matching (Exact Matching) Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Pattern Matching Exhaustive Search DFA Algorithm KMP Algorithm
CSI/BINF 5330 Pattern Matching (Exact Matching) Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Pattern Matching Exhaustive Search DFA Algorithm KMP Algorithm
Ion Torrent. The chip is the machine
 Ion Torrent Introduction The Ion Personal Genome Machine [PGM] is simple, more costeffective, and more scalable than any other sequencing technology. Founded in 2007 by Jonathan Rothberg. Part of Life
Ion Torrent Introduction The Ion Personal Genome Machine [PGM] is simple, more costeffective, and more scalable than any other sequencing technology. Founded in 2007 by Jonathan Rothberg. Part of Life
Lecture 7 Mutation and genetic variation
 Lecture 7 Mutation and genetic variation Thymidine dimer Natural selection at a single locus 2. Purifying selection a form of selection acting to eliminate harmful (deleterious) alleles from natural populations.
Lecture 7 Mutation and genetic variation Thymidine dimer Natural selection at a single locus 2. Purifying selection a form of selection acting to eliminate harmful (deleterious) alleles from natural populations.
Phylogenetic Assumptions
 Substitution Models and the Phylogenetic Assumptions Vivek Jayaswal Lars S. Jermiin COMMONWEALTH OF AUSTRALIA Copyright htregulation WARNING This material has been reproduced and communicated to you by
Substitution Models and the Phylogenetic Assumptions Vivek Jayaswal Lars S. Jermiin COMMONWEALTH OF AUSTRALIA Copyright htregulation WARNING This material has been reproduced and communicated to you by
Comparative genomics: Overview & Tools + MUMmer algorithm
 Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Comparative genomics: Overview & Tools + MUMmer algorithm Urmila Kulkarni-Kale Bioinformatics Centre University of Pune, Pune 411 007. urmila@bioinfo.ernet.in Genome sequence: Fact file 1995: The first
Regulatory Sequence Analysis. Sequence models (Bernoulli and Markov models)
 Regulatory Sequence Analysis Sequence models (Bernoulli and Markov models) 1 Why do we need random models? Any pattern discovery relies on an underlying model to estimate the random expectation. This model
Regulatory Sequence Analysis Sequence models (Bernoulli and Markov models) 1 Why do we need random models? Any pattern discovery relies on an underlying model to estimate the random expectation. This model
3/1/17. Content. TWINSCAN model. Example. TWINSCAN algorithm. HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM
 I529: Machine Learning in Bioinformatics (Spring 2017) Content HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM Yuzhen Ye School of Informatics and Computing Indiana University,
I529: Machine Learning in Bioinformatics (Spring 2017) Content HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM Yuzhen Ye School of Informatics and Computing Indiana University,
Chapter 15 Active Reading Guide Regulation of Gene Expression
 Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
15.12 Applications of Suffix Trees
 248 Algorithms in Bioinformatis II, SoSe 07, ZBIT, D. Huson, May 14, 2007 15.12 Appliations of Suffix Trees 1. Searhing for exat patterns 2. Minimal unique substrings 3. Maximum unique mathes 4. Maximum
248 Algorithms in Bioinformatis II, SoSe 07, ZBIT, D. Huson, May 14, 2007 15.12 Appliations of Suffix Trees 1. Searhing for exat patterns 2. Minimal unique substrings 3. Maximum unique mathes 4. Maximum
Hidden Markov Models. music recognition. deal with variations in - pitch - timing - timbre 2
 Hidden Markov Models based on chapters from the book Durbin, Eddy, Krogh and Mitchison Biological Sequence Analysis Shamir s lecture notes and Rabiner s tutorial on HMM 1 music recognition deal with variations
Hidden Markov Models based on chapters from the book Durbin, Eddy, Krogh and Mitchison Biological Sequence Analysis Shamir s lecture notes and Rabiner s tutorial on HMM 1 music recognition deal with variations
Name: SBI 4U. Gene Expression Quiz. Overall Expectation:
 Gene Expression Quiz Overall Expectation: - Demonstrate an understanding of concepts related to molecular genetics, and how genetic modification is applied in industry and agriculture Specific Expectation(s):
Gene Expression Quiz Overall Expectation: - Demonstrate an understanding of concepts related to molecular genetics, and how genetic modification is applied in industry and agriculture Specific Expectation(s):
Graph Algorithms in Bioinformatics
 Graph Algorithms in Bioinformatics Outline 1. Introduction to Graph Theory 2. The Hamiltonian & Eulerian Cycle Problems 3. Basic Biological Applications of Graph Theory 4. DNA Sequencing 5. Shortest Superstring
Graph Algorithms in Bioinformatics Outline 1. Introduction to Graph Theory 2. The Hamiltonian & Eulerian Cycle Problems 3. Basic Biological Applications of Graph Theory 4. DNA Sequencing 5. Shortest Superstring
RGP finder: prediction of Genomic Islands
 Training courses on MicroScope platform RGP finder: prediction of Genomic Islands Dynamics of bacterial genomes Gene gain Horizontal gene transfer Gene loss Deletion of one or several genes Duplication
Training courses on MicroScope platform RGP finder: prediction of Genomic Islands Dynamics of bacterial genomes Gene gain Horizontal gene transfer Gene loss Deletion of one or several genes Duplication
FUNDAMENTALS OF MOLECULAR EVOLUTION
 FUNDAMENTALS OF MOLECULAR EVOLUTION Second Edition Dan Graur TELAVIV UNIVERSITY Wen-Hsiung Li UNIVERSITY OF CHICAGO SINAUER ASSOCIATES, INC., Publishers Sunderland, Massachusetts Contents Preface xiii
FUNDAMENTALS OF MOLECULAR EVOLUTION Second Edition Dan Graur TELAVIV UNIVERSITY Wen-Hsiung Li UNIVERSITY OF CHICAGO SINAUER ASSOCIATES, INC., Publishers Sunderland, Massachusetts Contents Preface xiii
Sum and Product Rules
 Sum and Product Rules Exercise. Consider tossing a coin five times. What is the probability of getting the same result on the first two tosses or the last two tosses? Solution. Let E be the event that
Sum and Product Rules Exercise. Consider tossing a coin five times. What is the probability of getting the same result on the first two tosses or the last two tosses? Solution. Let E be the event that
Genomes of coral dinoflagellate symbionts highlight evolutionary adaptations conducive to a symbiotic lifestyle
 1 Genomes of coral dinoflagellate symbionts highlight evolutionary adaptations conducive to a symbiotic lifestyle Short title: Comparative analysis of coral dinoflagellate genomes M. Aranda a, Y. Li a,
1 Genomes of coral dinoflagellate symbionts highlight evolutionary adaptations conducive to a symbiotic lifestyle Short title: Comparative analysis of coral dinoflagellate genomes M. Aranda a, Y. Li a,
6.047 / Computational Biology: Genomes, Networks, Evolution Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Transposon diversity is higher in amphioxus than in vertebrates: functional and evolutionary inferences Cristian Can estro and Ricard Albalat
 BRIEFINGS IN FUNCTIONAL GENOMICS. VOL 11. NO 2. 131^141 doi:10.1093/bfgp/els010 Transposon diversity is higher in amphioxus than in vertebrates: functional and evolutionary inferences Cristian Can estro
BRIEFINGS IN FUNCTIONAL GENOMICS. VOL 11. NO 2. 131^141 doi:10.1093/bfgp/els010 Transposon diversity is higher in amphioxus than in vertebrates: functional and evolutionary inferences Cristian Can estro
arxiv:q-bio/ v1 [q-bio.pe] 23 Jan 2006
![arxiv:q-bio/ v1 [q-bio.pe] 23 Jan 2006 arxiv:q-bio/ v1 [q-bio.pe] 23 Jan 2006](/thumbs/81/83508874.jpg) arxiv:q-bio/0601039v1 [q-bio.pe] 23 Jan 2006 Food-chain competition influences gene s size. Marta Dembska 1, Miros law R. Dudek 1 and Dietrich Stauffer 2 1 Instituteof Physics, Zielona Góra University,
arxiv:q-bio/0601039v1 [q-bio.pe] 23 Jan 2006 Food-chain competition influences gene s size. Marta Dembska 1, Miros law R. Dudek 1 and Dietrich Stauffer 2 1 Instituteof Physics, Zielona Góra University,
G4120: Introduction to Computational Biology
 ICB Fall 2009 G4120: Introduction to Computational Biology Oliver Jovanovic, Ph.D. Columbia University Department of Microbiology & Immunology Copyright 2008 Oliver Jovanovic, All Rights Reserved. Genome
ICB Fall 2009 G4120: Introduction to Computational Biology Oliver Jovanovic, Ph.D. Columbia University Department of Microbiology & Immunology Copyright 2008 Oliver Jovanovic, All Rights Reserved. Genome
Topology. 1 Introduction. 2 Chromosomes Topology & Counts. 3 Genome size. 4 Replichores and gene orientation. 5 Chirochores.
 Topology 1 Introduction 2 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 G+C content 7 Codon usage 27 marc.bailly-bechet@univ-lyon1.fr The big picture Eukaryota Bacteria Many linear chromosomes
Topology 1 Introduction 2 3 Genome size 4 Replichores and gene orientation 5 Chirochores 6 G+C content 7 Codon usage 27 marc.bailly-bechet@univ-lyon1.fr The big picture Eukaryota Bacteria Many linear chromosomes
The Developmental Transcriptome of the Mosquito Aedes aegypti, an invasive species and major arbovirus vector.
 The Developmental Transcriptome of the Mosquito Aedes aegypti, an invasive species and major arbovirus vector. Omar S. Akbari*, Igor Antoshechkin*, Henry Amrhein, Brian Williams, Race Diloreto, Jeremy
The Developmental Transcriptome of the Mosquito Aedes aegypti, an invasive species and major arbovirus vector. Omar S. Akbari*, Igor Antoshechkin*, Henry Amrhein, Brian Williams, Race Diloreto, Jeremy
10-810: Advanced Algorithms and Models for Computational Biology. microrna and Whole Genome Comparison
 10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
Capacity and Expressiveness of Genomic Tandem Duplication
 Capacity and Expressiveness of Genomic Tandem Duplication Siddharth Jain sidjain@caltech.edu Farzad Farnoud (Hassanzadeh) farnoud@caltech.edu Jehoshua Bruck bruck@caltech.edu Abstract The majority of the
Capacity and Expressiveness of Genomic Tandem Duplication Siddharth Jain sidjain@caltech.edu Farzad Farnoud (Hassanzadeh) farnoud@caltech.edu Jehoshua Bruck bruck@caltech.edu Abstract The majority of the
MOBILE ELEMENTS AND EVOLUTION OF MOLECULAR REGULATORY SYSTEMS. Evelina Daskalova*
 PROCEEDINGS OF THE BALKAN SCIENTIFIC CONFERENCE OF BIOLOGY IN PLOVDIV (BULGARIA) FROM 19 TH TILL 21 ST OF MAY 2005 (EDS B. GRUEV, M. NIKOLOVA AND A. DONEV), 2005 (P. 79 89) MOBILE ELEMENTS AND EVOLUTION
PROCEEDINGS OF THE BALKAN SCIENTIFIC CONFERENCE OF BIOLOGY IN PLOVDIV (BULGARIA) FROM 19 TH TILL 21 ST OF MAY 2005 (EDS B. GRUEV, M. NIKOLOVA AND A. DONEV), 2005 (P. 79 89) MOBILE ELEMENTS AND EVOLUTION
Grundlagen der Bioinformatik Summer semester Lecturer: Prof. Daniel Huson
 Grundlagen der Bioinformatik, SS 10, D. Huson, April 12, 2010 1 1 Introduction Grundlagen der Bioinformatik Summer semester 2010 Lecturer: Prof. Daniel Huson Office hours: Thursdays 17-18h (Sand 14, C310a)
Grundlagen der Bioinformatik, SS 10, D. Huson, April 12, 2010 1 1 Introduction Grundlagen der Bioinformatik Summer semester 2010 Lecturer: Prof. Daniel Huson Office hours: Thursdays 17-18h (Sand 14, C310a)
AP Bio Module 16: Bacterial Genetics and Operons, Student Learning Guide
 Name: Period: Date: AP Bio Module 6: Bacterial Genetics and Operons, Student Learning Guide Getting started. Work in pairs (share a computer). Make sure that you log in for the first quiz so that you get
Name: Period: Date: AP Bio Module 6: Bacterial Genetics and Operons, Student Learning Guide Getting started. Work in pairs (share a computer). Make sure that you log in for the first quiz so that you get
The Evolution and Diversity of DNA Transposons in the Genome of the Lizard Anolis carolinensis
 The Evolution and Diversity of DNA Transposons in the Genome of the Lizard Anolis carolinensis Peter A. Novick 1,2, Jeremy D. Smith 3, Mark Floumanhaft 1, David A. Ray 3, and Stéphane Boissinot*,1,2 1
The Evolution and Diversity of DNA Transposons in the Genome of the Lizard Anolis carolinensis Peter A. Novick 1,2, Jeremy D. Smith 3, Mark Floumanhaft 1, David A. Ray 3, and Stéphane Boissinot*,1,2 1
Bacterial Genetics & Operons
 Bacterial Genetics & Operons The Bacterial Genome Because bacteria have simple genomes, they are used most often in molecular genetics studies Most of what we know about bacterial genetics comes from the
Bacterial Genetics & Operons The Bacterial Genome Because bacteria have simple genomes, they are used most often in molecular genetics studies Most of what we know about bacterial genetics comes from the
Assembly improvement: based on Ragout approach. student: Anna Lioznova scientific advisor: Son Pham
 Assembly improvement: based on Ragout approach student: Anna Lioznova scientific advisor: Son Pham Plan Ragout overview Datasets Assembly improvements Quality overlap graph paired-end reads Coverage Plan
Assembly improvement: based on Ragout approach student: Anna Lioznova scientific advisor: Son Pham Plan Ragout overview Datasets Assembly improvements Quality overlap graph paired-end reads Coverage Plan
Conditional Probability and Bayes Theorem (2.4) Independence (2.5)
 Conditional Probability and Bayes Theorem (2.4) Independence (2.5) Prof. Tesler Math 186 Winter 2019 Prof. Tesler Conditional Probability and Bayes Theorem Math 186 / Winter 2019 1 / 38 Scenario: Flip
Conditional Probability and Bayes Theorem (2.4) Independence (2.5) Prof. Tesler Math 186 Winter 2019 Prof. Tesler Conditional Probability and Bayes Theorem Math 186 / Winter 2019 1 / 38 Scenario: Flip
Essentiality in B. subtilis
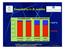 Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Essentiality in B. subtilis 100% 75% Essential genes Non-essential genes Lagging 50% 25% Leading 0% non-highly expressed highly expressed non-highly expressed highly expressed 1 http://www.pasteur.fr/recherche/unites/reg/
Hidden Markov Models (HMMs) November 14, 2017
 Hidden Markov Models (HMMs) November 14, 2017 inferring a hidden truth 1) You hear a static-filled radio transmission. how can you determine what did the sender intended to say? 2) You know that genes
Hidden Markov Models (HMMs) November 14, 2017 inferring a hidden truth 1) You hear a static-filled radio transmission. how can you determine what did the sender intended to say? 2) You know that genes
A DNA Sequence 2017/12/6 1
 A DNA Sequence ccgtacgtacgtagagtgctagtctagtcgtagcgccgtagtcgatcgtgtgg gtagtagctgatatgatgcgaggtaggggataggatagcaacagatgagc ggatgctgagtgcagtggcatgcgatgtcgatgatagcggtaggtagacttc gcgcataaagctgcgcgagatgattgcaaagragttagatgagctgatgcta
A DNA Sequence ccgtacgtacgtagagtgctagtctagtcgtagcgccgtagtcgatcgtgtgg gtagtagctgatatgatgcgaggtaggggataggatagcaacagatgagc ggatgctgagtgcagtggcatgcgatgtcgatgatagcggtaggtagacttc gcgcataaagctgcgcgagatgattgcaaagragttagatgagctgatgcta
Co-ordination occurs in multiple layers Intracellular regulation: self-regulation Intercellular regulation: coordinated cell signalling e.g.
 Gene Expression- Overview Differentiating cells Achieved through changes in gene expression All cells contain the same whole genome A typical differentiated cell only expresses ~50% of its total gene Overview
Gene Expression- Overview Differentiating cells Achieved through changes in gene expression All cells contain the same whole genome A typical differentiated cell only expresses ~50% of its total gene Overview
A SINE in the genome of the cephalochordate amphioxus is an Alu element
 Int. J. Biol. Sci. 2006, 2 61 Research paper International Journal of Biological Sciences ISSN 1449-2288 www.biolsci.org 2006 2(2):61-65 2006 Ivyspring International Publisher. All rights reserved A SINE
Int. J. Biol. Sci. 2006, 2 61 Research paper International Journal of Biological Sciences ISSN 1449-2288 www.biolsci.org 2006 2(2):61-65 2006 Ivyspring International Publisher. All rights reserved A SINE
Biology. Biology. Slide 1 of 26. End Show. Copyright Pearson Prentice Hall
 Biology Biology 1 of 26 Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 2 of 26 Gene Regulation: An Example Gene Regulation: An Example
Biology Biology 1 of 26 Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 2 of 26 Gene Regulation: An Example Gene Regulation: An Example
Abstract. comment reviews reports deposited research refereed research interactions information
 http://genomebiology.com/2002/3/12/research/0086.1 Research Assessing the impact of comparative genomic sequence data on the functional annotation of the Drosophila genome Casey M Bergman*, Barret D Pfeiffer*,
http://genomebiology.com/2002/3/12/research/0086.1 Research Assessing the impact of comparative genomic sequence data on the functional annotation of the Drosophila genome Casey M Bergman*, Barret D Pfeiffer*,
Genotyping By Sequencing (GBS) Method Overview
 enotyping By Sequencing (BS) Method Overview RJ Elshire, JC laubitz, Q Sun, JV Harriman ES Buckler, and SE Mitchell http://wwwmaizegeneticsnet/ Topics Presented Background/oals BS lab protocol Illumina
enotyping By Sequencing (BS) Method Overview RJ Elshire, JC laubitz, Q Sun, JV Harriman ES Buckler, and SE Mitchell http://wwwmaizegeneticsnet/ Topics Presented Background/oals BS lab protocol Illumina
HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM
 I529: Machine Learning in Bioinformatics (Spring 2017) HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington
I529: Machine Learning in Bioinformatics (Spring 2017) HMM for modeling aligned multiple sequences: phylo-hmm & multivariate HMM Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington
CHAPTER : Prokaryotic Genetics
 CHAPTER 13.3 13.5: Prokaryotic Genetics 1. Most bacteria are not pathogenic. Identify several important roles they play in the ecosystem and human culture. 2. How do variations arise in bacteria considering
CHAPTER 13.3 13.5: Prokaryotic Genetics 1. Most bacteria are not pathogenic. Identify several important roles they play in the ecosystem and human culture. 2. How do variations arise in bacteria considering
Predicting RNA Secondary Structure
 7.91 / 7.36 / BE.490 Lecture #6 Mar. 11, 2004 Predicting RNA Secondary Structure Chris Burge Review of Markov Models & DNA Evolution CpG Island HMM The Viterbi Algorithm Real World HMMs Markov Models for
7.91 / 7.36 / BE.490 Lecture #6 Mar. 11, 2004 Predicting RNA Secondary Structure Chris Burge Review of Markov Models & DNA Evolution CpG Island HMM The Viterbi Algorithm Real World HMMs Markov Models for
Statistics for Differential Expression in Sequencing Studies. Naomi Altman
 Statistics for Differential Expression in Sequencing Studies Naomi Altman naomi@stat.psu.edu Outline Preliminaries what you need to do before the DE analysis Stat Background what you need to know to understand
Statistics for Differential Expression in Sequencing Studies Naomi Altman naomi@stat.psu.edu Outline Preliminaries what you need to do before the DE analysis Stat Background what you need to know to understand
TRANSPOSABLE elements (TEs) are mobile genetic
 Copyright Ó 2007 by the Genetics Society of America DOI: 10.1534/genetics.107.081109 Note Evolution and Horizontal Transfer of a DD37E DNA Transposon in Mosquitoes James K. Biedler, 1,2 Hongguang Shao
Copyright Ó 2007 by the Genetics Society of America DOI: 10.1534/genetics.107.081109 Note Evolution and Horizontal Transfer of a DD37E DNA Transposon in Mosquitoes James K. Biedler, 1,2 Hongguang Shao
opulation genetics undamentals for SNP datasets
 opulation genetics undamentals for SNP datasets with crocodiles) Sam Banks Charles Darwin University sam.banks@cdu.edu.au I ve got a SNP genotype dataset, now what? Do my data meet the requirements of
opulation genetics undamentals for SNP datasets with crocodiles) Sam Banks Charles Darwin University sam.banks@cdu.edu.au I ve got a SNP genotype dataset, now what? Do my data meet the requirements of
Genotyping By Sequencing (GBS) Method Overview
 enotyping By Sequencing (BS) Method Overview Sharon E Mitchell Institute for enomic Diversity Cornell University http://wwwmaizegeneticsnet/ Topics Presented Background/oals BS lab protocol Illumina sequencing
enotyping By Sequencing (BS) Method Overview Sharon E Mitchell Institute for enomic Diversity Cornell University http://wwwmaizegeneticsnet/ Topics Presented Background/oals BS lab protocol Illumina sequencing
CBSE X Mathematics 2012 Solution (SET 1) Section B
 CBSE X Mathematics 01 Solution (SET 1) Section B Q11. Find the value(s) of k so that the quadratic equation x kx + k = 0 has equal roots. Given equation is x kx k 0 For the given equation to have equal
CBSE X Mathematics 01 Solution (SET 1) Section B Q11. Find the value(s) of k so that the quadratic equation x kx + k = 0 has equal roots. Given equation is x kx k 0 For the given equation to have equal
A rigid-base model for DNA structure prediction. O. Gonzalez
 A rigid-base model for DNA structure prediction O. Gonzalez Introduction Objective. To develop a model to predict the structure and flexibility of standard, B-form DNA from its sequence. Introduction Objective.
A rigid-base model for DNA structure prediction O. Gonzalez Introduction Objective. To develop a model to predict the structure and flexibility of standard, B-form DNA from its sequence. Introduction Objective.
Multiple Choice Review- Eukaryotic Gene Expression
 Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Modelling and Analysis in Bioinformatics. Lecture 1: Genomic k-mer Statistics
 582746 Modelling and Analysis in Bioinformatics Lecture 1: Genomic k-mer Statistics Juha Kärkkäinen 06.09.2016 Outline Course introduction Genomic k-mers 1-Mers 2-Mers 3-Mers k-mers for Larger k Outline
582746 Modelling and Analysis in Bioinformatics Lecture 1: Genomic k-mer Statistics Juha Kärkkäinen 06.09.2016 Outline Course introduction Genomic k-mers 1-Mers 2-Mers 3-Mers k-mers for Larger k Outline
Is KIT locus polymorphism rs related to white belt phenotype in Krškopolje pig?
 Is KIT locus polymorphism rs328592739 related to white belt phenotype in Krškopolje pig? Jernej Ogorevc, Minja Zorc, Martin Škrlep, Riccardo Bozzi, Matthias Petig, Luca Fontanesi, Marjeta Čandek-Potokar,
Is KIT locus polymorphism rs328592739 related to white belt phenotype in Krškopolje pig? Jernej Ogorevc, Minja Zorc, Martin Škrlep, Riccardo Bozzi, Matthias Petig, Luca Fontanesi, Marjeta Čandek-Potokar,
Evolutionary analysis of the well characterized endo16 promoter reveals substantial variation within functional sites
 Evolutionary analysis of the well characterized endo16 promoter reveals substantial variation within functional sites Paper by: James P. Balhoff and Gregory A. Wray Presentation by: Stephanie Lucas Reviewed
Evolutionary analysis of the well characterized endo16 promoter reveals substantial variation within functional sites Paper by: James P. Balhoff and Gregory A. Wray Presentation by: Stephanie Lucas Reviewed
