Cluster Distance Geometry of Polypeptide Chains
|
|
|
- Jasmine Ellis
- 6 years ago
- Views:
Transcription
1 Cluster Distance Geometry of Polypeptide Chains GORDON M. CRIPPEN College of Pharmacy, University of Michigan, Ann Arbor, Michigan Received 9 February 004; Accepted 3 March 004 DOI /jcc.0056 Published online in Wiley InterScience ( Abstract: Distance geometry has been a broadly useful tool for dealing with conformational calculations. Customarily each atom is represented as a point, constraints on the distances between some atoms are obtained from experimental or theoretical sources, and then a random sampling of conformations can be calculated that are consistent with the constraints. Although these methods can be applied to small proteins having on the order of 1000 atoms, for some purposes it is advantageous to view the problem at lower resolution. Here distance geometry is generalized to deal with distances between sets of points. In the end, much of the same techniques produce a sampling of different configurations of these sets of points subject to distance constraints, but now the radii of gyration of the different sets play an important role. A simple example is given of how the packing constraints for polypeptide chains combine with loose distance constraints to give good calculated protein conformers at a very low resolution. 004 Wiley Periodicals, Inc. J Comput Chem 5: , 004 Key words: distance geometry; protein conformation; excluded volume; conformational analysis; radius of gyration Introduction Distance geometry refers to a treatment of geometric problems that emphasizes Euclidean distances between points, rather than angles, and so forth. 1 This is sometimes a convenient way to deal with conformational problems, where the points correspond to atoms. 4 Currently the most frequent application is to calculate a sampling of sets of atomic coordinates given constraints on some of the interatomic distances derived from NMR experiments. 5 7 Other applications include protein homology modeling, 8,9 protein structure prediction, 10 1 and more abstract conformation 13 and sequence 14 spaces. A recurring task in all these applications is how to generate a set of coordinates for the points given at least some constraints on some of the interpoint distances. New methods for this continue to be developed, each with its advantages and disadvantages, depending on the type of constraint set. Here we consider a new sort of distance geometry problem where the points may be atoms or even whole amino acid residues, and we want to look at conformations at a very low resolution where the primary objects are clusters or subsets of the points. Much of the standard methodology for single points carries over in an analogous form for clusters, with the advantage of building in the space-filling features of real atoms or residues in a natural way. In order to make this correspondence clear, we first briefly outline the standard distance geometry methods, then show the equivalent procedures for clusters, and finally give a simple demonstration of calculating conformations for a protein given certain constraints. Methods Standard Distance Geometry Suppose we have a set of n distinct points with a matrix of proposed squared Euclidean distances between them, M (d ij ). Obviously M must be symmetric (d ij d ji ), the diagonal must be zero (d ii 0), and the elements must be non-negative (d ij 0). There are in addition some nonobvious requirements that M must fulfill in order to correspond to a realizable arrangement of the points in three-dimensional space that is not confined to some planar subspace. These constraints are expressed in terms of C(M, k), the Cayley-Menger determinant involving the first k points: d 1 d 0 1k C M, k 1 d 1 0 d k (1) 1 d k1 d k Blumenthal s theorem 1 specialized to three dimensions and n 4 requires that there be some ordering of the points such that C(M, Correspondence to: G. M. Crippen; gcrippen@umich.edu Contract/grant sponsor: University of Michigan Bioinformatics Program. Contract/grant sponsor: Howard Hughes Medical Institute. 004 Wiley Periodicals, Inc.
2 1306 Crippen Vol. 5, No. 10 Journal of Computational Chemistry ) 0, C(M, 3) 0, C(M, 4) 0, and for any choice of additional fifth and sixth points that C(M, 5) C(M, 6) 0. The constraints on the lower order determinants can readily be interpreted.,3 Because C(M, ) d 1, the first inequality is trivially satisfied by d 1 0. The second inequality involves factoring the determinant C M, 3 d 3 d 1 d 13 d 1 d 13 d 3 d 1 d 13 d 3 d 1 d 13 d 3 0 () which is satisfied if the triangle inequality, d 13 d 1 d 3, holds as a strict inequality for all three permutations of indices. The triangle inequality becomes an equality when the three points are colinear, so the requirement that C(M, 3) 0 means the first three points are not colinear. The algebra for four points is more complicated, but solving C(M, 4) 0 for d 14 (assuming fixed values of the other distances) gives two real, positive roots, and as long as d 14 lies strictly between these two bounds, we have C(M, 4) 0. This has been called the tetrangle inequality, 4 and satisfying the strict inequality implies the first four points are noncoplanar. Another way to look at the Cayley-Menger determinants is in terms of coordinates of the points in the case that the constraints on the corresponding distances are satisfied and coordinates can consequently be found. Any two points i and j span a linear subspace that we can equip with a coordinate system consisting of an origin and an x-axis. Then the determinant for the ordered pair of points where C M, i, j ij (3) ij 1 1 x i x j x j x i (4) is the orientation of the two ordered points relative to the coordinate system. That is, ij 0 if going from x i to x j is the same direction as going from the origin to the positive x-axis, ij 0if the orientation is opposite, and ij 0 if the two points coincide and hence have no orientation along the x-axis. Similarly, it is straightforward but tedious to verify that for an ordered set of three points, [i, j, k], spanning two dimensions, the corresponding Cayley-Menger determinant is C M, i, j, k 4 ijk where the three-point orientation relative to the x- and y-axes of the coordinate system is ijk x i x j x k y i y j y k (6) and ijk 0 when going from point i to j to k is counterclockwise in the xy-plane. When the three points are colinear, ijk 0. (5) There are other useful quantities that follow directly from the distances without reference to coordinates. Given the matrix of squared distances, M, one can directly calculate d io, the squared distance from point i to the unweighted center of mass of all n points 18. d io n 1 d ij n d jk j j k Because the radius of gyration of the n points is r g (n 1 i d io ) 1/, it follows from eq. (7) that r g n d jk (8) j k An important problem in distance geometry is finding coordinates for the points, if any can exist, that satisfy given bounds on the distances between some of the pairs of points. Thus for every distance we require that l ij d ij u ij, and for some distances the upper and lower bounds may be significant constraints, while for others they may be the trivial l ij 0 and u ij. Bound smoothing 3,4 is a process whereby the information from the tight bounds is spread around to all distances by lowering some upper bounds and raising some lower bounds. At the triangle inequality level, one simply repeatedly examines all triples of points and whenever (7) u ik u ij u jk (9) is violated, u ik is reduced to the right-hand side of the inequality. After no further reductions in upper bounds can be achieved, all triples of points are repeatedly checked for violations of l ik l ij u jk (10) in which case l ik is increased to the right-hand side of the inequality. From the smoothed bounds, one may pick distance values at random in the intervals l ij d ij u ij, independently for each d ij. However, the resulting distances may not necessarily obey the triangle inequality. In the process called metrization, 5 each time a particular d ij is chosen, the corresponding bounds are tightened to l ij d ij u ij, and smoothing of the other bounds by eqs. (9) and (10) may tighten them. When all the distances have been chosen in this way, they do obey the triangle inequality. In order to determine coordinates from the selected distances, the metric matrix G ( g ij ) is determined by g ij 1 d io d jo d ij (11) where the distances to the center of mass come from eq. (7). Then the j x, y, z coordinate vectors of the n points are determined by c j j 1/ w j (1)
3 Distance Geometry of Polypeptide Chains 1307 from the three largest (positive) eigenvalues j and corresponding eigenvectors w j of G. If the other n 3 eigenvalues are relatively large in magnitude, the resulting coordinates may need adjusting in order to obey the original bounds. Cluster Distance Geometry General Relationships Suppose the objects of interest are not individual points, but rather sets of b points. So if n is a multiple of b, we can think of grouping M into b b blocks to form an order n/b matrix M b (D IJ ), where each element D IJ is the sum of the b interpoint squared distances, d ij, within that block. It is still true that M b is a symmetric matrix of non-negative elements, but now the diagonal elements are no longer necessarily zero. Instead, we see from eq. (8) that D II b r g,i (13) where r g,i is the squared radius of gyration of the Ith set of points. The equivalent constraints on the Cayley-Menger determinants apparently hold. The two-block determinant is not as trivial as the two-point determinant: C M b, I, J D IJ D II D JJ 0 (14) It is easy enough to demonstrate that this condition holds when there are coordinates for the points. Consider the case b, involving only four points labeled i, i 1, j, and j 1. Suppose these points have coordinates [ x i, y i, z i ] T, and so forth. Then C M, I, J d ij d i 1, j d i, j 1 d i 1, j 1 d i,i 1 d j, j 1 x i x j x i 1 x j x i x j 1 x i 1 x j 1 x i x i 1 x j x j 1 similar for y and z x i x i 1 x j x j 1 similar for y and z 8d I,J (15) where I and J denote the unweighted centers of mass (or centroids) of the two clusters. In general C M b, I, J b d I,J b I J 0 (16) where I J is the one-dimensional orientation for the two centroids as in eq. (4), and C(M b,[i, J]) 0 only when the two centroids coincide. Clearly for b 1, this reduces to the standard distance geometry case. Similarly, C(M b, 3) 0 is widely observed to be always satisfied for configurations of points in three dimensions, but is not so simply interpreted in terms of something like the triangle inequality. In terms of coordinates in the plane, one can derive in analogy to eq. (15) that C M b, I, J, K 4b I J K 0 (17) and C(M b,[i, J, K]) 0 when the corresponding three centroids are colinear. In terms of distances, like the treatment of the tetrangle inequality above, one can solve C(M b,[i, J, K]) 0 for an upper and lower bound on, say, D IK, given values for all the other matrix elements. This gives D IJ D JK D JJ P 1/ D IK D IJ D JK D JJ P 1/ (18) where P D JK D JJ D KK D IJ D II D JJ C M b, J, K C M b, I, J (19) and the bounds in eq. (18) are always real because both factors in eq. (19) are positive, according to the requirement of eq. (14). In the b 1 limit where D II D JJ D KK 0, this reduces to the usual triangle inequality limits of max d jk d ij, d ij d jk d ik d ij d jk. (0) The equivalent to triangle inequality level bound smoothing is somewhat more complicated than in standard distance geometry. For every pair of blocks there are bounds L IJ D IJ U IJ that may be trivial (L IJ 0 and U IJ ) for some or nontrivial given bounds for others. The diagonal elements may also have nontrivial bounds, rather than d ii 0 for a single point i. Thus the first step is to repeatedly scan all pairs of blocks looking for violations of L IJ L II L JJ / (1) which would require raising L IJ to the right-hand side of the inequality, in accord with eq. (14). Similarly, a less frequently occurring case is violations of U IJ L II U JJ () which requires lowering U JJ to the left-hand side of the inequality. The same holds for I and J exchanged. After no further tightening of bounds can be achieved, the next step is to repeatedly examine all unordered triples of blocks I, J, and K for violations of U IK U IJ U JK L JJ U JK L JJ L KK U IJ L II L JJ 1/ (3) which comes from maximizing the right-hand side of eq. (18). Note that that expression achieves its maximum subject to the constraints that D IJ U IJ, D JK U JK, D II L II, D KK L KK, and D JJ L JJ. The constraints that D JK D JJ D KK 0 and D IJ D II D JJ 0 are not active at the maximum, and the two factors in the square root of eq. (3) are necessarily nonnegative. Here U IK is reduced to the right-hand side, very much like using eq. (9).
4 1308 Crippen Vol. 5, No. 10 Journal of Computational Chemistry Figure 1. A semilog plot of the range of diagonal elements D II as a function of block size b. Plotted values are those observed in a survey of 3 small protein crystal structures for log L II (diamonds) and log U II (crosses) compared to fitted upper bounds from eq. (30) (thick line) and fitted lower bounds from eq. (31) (thin line). Eq. (3) is used to reduce off-diagonal upper bounds, just as in standard distance geometry, but it can also lead to reductions in U II, U JJ, and U KK. For example, if substituting U II for L II in eq. (3) leads to a calculated U IK L IK, then U II must be reduced to U II U IJ L JJ L IK U IJ U JK L JJ U JK L JJ L KK (4) which comes from solving C(M b,[i, J, K]) 0 for D II when D IK, D JJ, D KK are at their lower bounds, and D IJ and D JK are at their upper bounds. Swapping indices I and K in eq. (4) gives the condition for reducing U KK. For reducing U JJ in the equivalent situation, the limit is U JJ U IJ U JK L IK U IJ L KK L IK U JK L II L IK L II L KK L KK L II L IK (5) D IK 1 C M b, I, J C M b, J, K D II D KK (6) which suggests three cases depending on the allowed ranges of C(M b,[i, J]) and C(M b,[j, K]). The first case is when max C(M b,[i, J]) min C(M b,[j, K]) as detected by U IJ L II L JK U KK and L JK U KK L JJ 0. Then the minimal value of D IK is achieved at the combination of upper and lower bounds: L IK U IJ L JK L JJ U IJ L II L JJ L JK L JJ L KK 1/ (7) which involves L KK, not U KK. The second case is when the allowed interval for C(M b,[i, J]) is strictly above that for C(M b, [ J, K]). Simply exchanging indices I and K, one detects that U JK L KK L IJ U II and L IJ U II L JJ 0, resulting in L IK L IJ U JK L JJ The last step is to repeatedly examine all unordered triples of blocks in order to raise L IK by the equivalent of eq. (10). Referring to eq. (18), the question is whether the current value of L IK is less than the minimal value of D IJ D JK D JJ (C(M b, [I, J])C(M b,[j, K])) 1/ subject to C(M b,[i, J]) 0, C(M b,[j, K]) 0, and the upper and lower bounds on all five variables, D II, D JJ, D KK, D IJ, and D JK. It is easy to verify that the left-hand side of eq. (18) can be expressed as L IJ L II L JJ U JK L JJ L KK 1/. (8) The third case is when the ranges overlap. The right-hand side of eq. (6) is minimized when they are equal, and it reduces to the two-block constraint that L IK (L II L KK )/. In the special case of b 1, the bounds on the diagonal elements are all zero, U IJ becomes u ij, and so forth, so that eq. (8) reduces to eq. (10), and eq. (7) reduces to the equivalent relation, l ik l jk u ij.
5 Distance Geometry of Polypeptide Chains 1309 Figure. A semilog plot of the range of diagonal elements D II as a function of block size b, when the corresponding protein residues are all part of a -strand. Values observed in a survey of protein crystal structures for log L II (diamonds) and log U II (crosses) are compared to fitted upper (thick line) and fitted lower bounds (thin line) from eq. (34). Protein Specific Relationships Consider a low resolution representation of protein structure where the n points are the C atoms of the n amino acid residues in a single polypeptide chain, one point per residue. Because of the chemical structure of the chain and its space-filling atoms, there are lower and upper bounds on the cluster distances that are much tighter than the pure geometric constraints. For instance, sequentially adjacent C atoms linked with a trans peptide bond would have a fixed separation of 3.8 Å, assuming standard bond lengths and angles. On the other hand, the mathematical series b i 1 b i j b b 1 6 j 1 so the upper bound on the diagonal blocks U II D II is U II 4.01 Å b b 1 6 (9) (30) for a fully extended polypeptide chain. The scaling factor of 4.01 Å rather than 3.8 Å comes from a survey over crystal structures of small proteins (PDB codes 1A1X, 1ACF, 1BK, 1BM8, 1BYW.A, 1C44.A, 1COA.I, 1CQY.A, 1DHN, 1DT4.A, 1ENH, 1EW4.A, 1G9O.A, 1HEY, 1IT.A, 1JWO.A, 1MIL, 1MJC, 1OPS, 1PGB, 1PHT, 1PTF, 1QAU.A, 1TEN, 1TMY, 1TUL, 1UBI, 1VCC, 1WHI, IGD, 3IL8, and 9MSI.A) and takes into account some experimental variations from standard peptide geometry. Over this dataset, the bounds are tight for b 10, but for larger blocks, finding such fully extended chain segments becomes unlikely, as shown in Figure 1. For the lower bounds on the diagonal blocks, L II D II,we know from eq. (13) that this corresponds to the minimal squared radius of gyration for a contiguous segment of b residues, and we know that the minimal r g,i varies linearly with b 1/3 from an earlier survey. 19 Once again surveying over the same 3 protein crystal structures as for the upper bound, a tight lower bound for 1 b 19 is L II b 3.17 Å b 1/3 1 (31) as shown in Figure 1. If all we know about a protein s structure is that it is a single, connected polypeptide chain, then the diagonal lower bounds are just L II for all I from eq. (31), and the off-diagonal lower bounds L IJ are that same value for all I and J by eq. (1). The default diagonal upper bounds, U(b), come from eq. (30), but these can be used to build up the off-diagonal upper bounds. Note that U I,I 1 U II U I 1,I 1 U(b) and U II U I 1,I 1
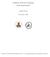
6 1310 Crippen Vol. 5, No. 10 Journal of Computational Chemistry Figure 3. A semilog plot of the range of diagonal elements D II as a function of block size b, when the corresponding protein residues are all part of an -helix. Values observed in a survey of protein crystal structures for log L II (diamonds) and log U II (crosses) are compared to fitted upper (thick line) and fitted lower bounds (thin line) from eq. (35). U(b), so the first off-diagonal upper bounds are all determined. In general U I,I k U I k,i 1 U 1 k b l,m U l,m (3) for k 1,..., (n/b) 1, where the sum runs over the square submatrix I l, m I k except for the two corners, (I, I k) and (I k, I). However, if any additional constraints have tightened some bounds more than those corresponding to a single polypeptide chain, the maximal packing density considerations can raise some off-diagonal lower bounds. Using the same notation as eq. (3), violations of L I,I k L I k,i 1 L 1 k b l,m U l,m (33) require that L I,I k and L I k,i be raised to the value of the righthand side. A knowledge of secondary structure can give tighter bounds on the diagonal elements. If all the residues in a block of b residues are known to be part of a -strand, then a survey over the same set of small proteins finds rather tight bounds: 3.09 Å b b 1 L 6 II U II 3.9 Å b b 1 6 (34) analogous to eq. (30), as shown in Figure. For an -helical block, the dependence on b is of lower order, but not so low as in eq. (31). Cubic dependence fits well as shown in Figure Å b 3 L II U II.86 Å b 3 (35) Embedding Finding coordinates for the centroids of the blocks from distance constraints is somewhat more elaborate than in standard distance geometry, but basically follows the same procedure. Initial bounds are either the trivial L IJ 0 and U IJ, or in the case of a polypeptide chain use eqs. (30), (31), and (3). Additional information further tightens some bounds, such as knowledge of protein secondary structure via eqs. (34) and (35). Subsequent bound smoothing involves exhaustive application of two-block relations [eqs. (1) and (), and for proteins eq. (33)] and three-block relations [eqs. (3), (4), (5), (7), and (8)]. Metrization works as before to choose random diagonal and off-diagonal D IJ consistent with all applicable bound smoothing relations. Conceptually, this amounts to proposed radii of gyration for all the blocks, as well as distances between them. By eq. (16), the (n/b) (n/b) matrix of proposed inter-centroid distances is calculated, which is then converted to coordinates as usual by eqs. (11) and (1). For very low resolution models, such as (n/b) 7, the resulting coordinates often completely satisfy the original bounds without any further adjustment.
7 Distance Geometry of Polypeptide Chains 1311 In standard distance geometry, one could model this protein as 54 C points linked by virtual bonds and otherwise restricted by lower bounds between all points representing the self-avoiding character of the chain. If the only other constraints were upper bounds among points 1, 9, 18, 7, 36, 45, and 54 taken to be slightly greater than the distances in the crystal structure, then a wide variety of conformations would be possible, including great rearrangements of the general fold. For the cluster distance geometry treatment, we included the a priori constraints for polypeptide chains plus off-diagonal U IJ that were 10% greater than those corresponding to the crystal structure. These explicit constraints plus the maximal packing density restrictions from eqs. (31) and (33) so greatly restrict the possible conformations that metrization sometimes has trouble finding a permitted set of D IJ values. In terms of the root mean squared deviation (RMSD) after optimal superposition of the cluster centers of mass from the crystal structure versus those from embedding, RMSD values ranged from 3 to 6 Å. Figure 6 shows a calculated cluster structure superimposed on the crystal structure clusters where the RMSD in cluster centers of mass was 3.4 Å. Obviously the calculated structure is constrained to be no bigger than the native, but note how some large distances are indirectly enforced by the packing considerations. Clearly there is still some room for conformational variation, as shown in Figure 7 where the traces of several calculated Figure 4. Backbone trace of BPTI (PDB entry 1G6X.A) less the first four residues (curved ribbon). The sequence of cylinders shows the positions of the centers of mass of the six sequential nine-residue segments. Embedding Results As a simple example of cluster distance geometry embedding applied to a protein, consider bovine pancreatic trypsin inhibitor (BPTI, PDB entry 1G6X.A). This consists of a single polypeptide chain of 58 residues. Deleting the N-terminal first four residues permits us to view the chain as a sequence of six clusters each consisting of nine C atoms or points. Figure 4 shows the C trace of the crystal structure and the greatly simplified trace of the six clusters, which nonetheless roughly outline the overall backbone path in space. For the sake of clarity, the radii of gyration of the six clusters are shown separately in Figure 5 as spheres centered at the cluster centers of mass and having the corresponding radii. Note how the C-terminal helix and the tight bend at the top of the illustration in Figure 4 correspond to small sphere in Figure 5, whereas the N-terminal extended strand on the right side has a large sphere. Figure 5. The radii of gyration of the six nine-residue segments of BPTI in the same view as the previous figure. Each segment is represented as a solid sphere centered at its center of mass having radius equal to the radius of gyration.
8 131 Crippen Vol. 5, No. 10 Journal of Computational Chemistry structures are shown superimposed on the crystal structure (not shown) in the same view as the previous figures. Overall, the RMSD of the corresponding cluster radii of gyration ranged from only 1.5 to.5 Å, which varies so little from Figure 5 that it would hardly be noticeable by eye. Conclusion Cluster distance geometry can be applied to a wide variety of geometric problems where a great reduction in resolution is desirable or necessary. Special problem-specific information can be built in, such as chain connectivity, secondary structure, and steric packing limitations, as shown for proteins. In particular, the packing constraints are very naturally incorporated, and their effect on other features is easily propagated. This is an effect that is difficult to include in standard distance geometry, which may make cluster distance geometry a useful approach even in instances that do not require a low resolution treatment. Acknowledgments All calculations were done in MOE using the SVL computer language. 0 Source code is available from the author on request. Figure 7. Superposition of the centers of mass for five calculated BPTI structures in the same view. References Figure 6. Superposition of the centers of mass of the BPTI segments for the native (string of darker cylinders extending slightly lower on the page) and calculated by embedding (other string of cylinders). 1. Blumenthal, L. M. Theory and Applications of Distance Geometry, nd edn.; Chelsea: New York, 1970, Crippen, G. M. J Comput Phys 1977, 4, Crippen, G. M. Distance Geometry and Conformational Calculations; Wiley: New York, Crippen, G. M.; Havel, T. F. Distance Geometry and Molecular Conformation; Wiley: New York, Havel, T. F.; Wüthrich, K. Bull Math Biol 1984, 46, Oezguen, N.; Adamian, L.; Xu, Y.; Rajarathnam, K.; Braun, W. J Biomol NMR 00,, Wider, G.; Wüthrich, K. Curr Opin Struct Biol 1999, 9, Havel, T. F.; Snow, M. E. J Mol Biol 1991, 17, Combet, C.; Jambon, M.; Deleage, G.; Geourjon, C. Bioinformatics 00, 18, Huang, E. S.; Samudrala, R.; Ponder, J. W. J Mol Biol 1999, 90, Xia, Y.; Huang, E. S.; Levitt, M.; Samudrala, R. J Mol Biol 000, 300, Petersen, K.; Taylor, W. R. J Mol Biol 003, 35, Laboulais, C.; Ouali, M.; Le Bret, M.; Gabarro-Arpa, J. Proteins 00, 47, Forster, M.; Heath, A.; Afzal, M. Bioinformatics 1999, 15, Glunt, W.; Hayden, T. Comput Chem 001, 5, Williams, G. A.; Dugan, J. M.; Altman, R. B. J Comput Biol 001, 8, Xu, H.; Izrailev, S.; Agrafiotis, D. K. J Chem Inf Comput Sci 003, 43, Crippen, G. M.; Havel, T. F. Acta Cryst A 1978, 34, Maiorov, V. N.; Crippen, G. M. J Mol Biol 199, 7, Molecular Operating Environment (MOE), Chemical Computing Group, Inc.
Distance Geometry-Matrix Completion
 Christos Konaxis Algs in Struct BioInfo 2010 Outline Outline Tertiary structure Incomplete data Tertiary structure Measure diffrence of matched sets Def. Root Mean Square Deviation RMSD = 1 n x i y i 2,
Christos Konaxis Algs in Struct BioInfo 2010 Outline Outline Tertiary structure Incomplete data Tertiary structure Measure diffrence of matched sets Def. Root Mean Square Deviation RMSD = 1 n x i y i 2,
Alpha-helical Topology and Tertiary Structure Prediction of Globular Proteins Scott R. McAllister Christodoulos A. Floudas Princeton University
 Alpha-helical Topology and Tertiary Structure Prediction of Globular Proteins Scott R. McAllister Christodoulos A. Floudas Princeton University Department of Chemical Engineering Program of Applied and
Alpha-helical Topology and Tertiary Structure Prediction of Globular Proteins Scott R. McAllister Christodoulos A. Floudas Princeton University Department of Chemical Engineering Program of Applied and
Introduction to Comparative Protein Modeling. Chapter 4 Part I
 Introduction to Comparative Protein Modeling Chapter 4 Part I 1 Information on Proteins Each modeling study depends on the quality of the known experimental data. Basis of the model Search in the literature
Introduction to Comparative Protein Modeling Chapter 4 Part I 1 Information on Proteins Each modeling study depends on the quality of the known experimental data. Basis of the model Search in the literature
Universal Similarity Measure for Comparing Protein Structures
 Marcos R. Betancourt Jeffrey Skolnick Laboratory of Computational Genomics, The Donald Danforth Plant Science Center, 893. Warson Rd., Creve Coeur, MO 63141 Universal Similarity Measure for Comparing Protein
Marcos R. Betancourt Jeffrey Skolnick Laboratory of Computational Genomics, The Donald Danforth Plant Science Center, 893. Warson Rd., Creve Coeur, MO 63141 Universal Similarity Measure for Comparing Protein
Secondary Structure. Bioch/BIMS 503 Lecture 2. Structure and Function of Proteins. Further Reading. Φ, Ψ angles alone determine protein structure
 Bioch/BIMS 503 Lecture 2 Structure and Function of Proteins August 28, 2008 Robert Nakamoto rkn3c@virginia.edu 2-0279 Secondary Structure Φ Ψ angles determine protein structure Φ Ψ angles are restricted
Bioch/BIMS 503 Lecture 2 Structure and Function of Proteins August 28, 2008 Robert Nakamoto rkn3c@virginia.edu 2-0279 Secondary Structure Φ Ψ angles determine protein structure Φ Ψ angles are restricted
Protein Structure Determination Using NMR Restraints BCMB/CHEM 8190
 Protein Structure Determination Using NMR Restraints BCMB/CHEM 8190 Programs for NMR Based Structure Determination CNS - Brünger, A. T.; Adams, P. D.; Clore, G. M.; DeLano, W. L.; Gros, P.; Grosse-Kunstleve,
Protein Structure Determination Using NMR Restraints BCMB/CHEM 8190 Programs for NMR Based Structure Determination CNS - Brünger, A. T.; Adams, P. D.; Clore, G. M.; DeLano, W. L.; Gros, P.; Grosse-Kunstleve,
Figure 1. Molecules geometries of 5021 and Each neutral group in CHARMM topology was grouped in dash circle.
 Project I Chemistry 8021, Spring 2005/2/23 This document was turned in by a student as a homework paper. 1. Methods First, the cartesian coordinates of 5021 and 8021 molecules (Fig. 1) are generated, in
Project I Chemistry 8021, Spring 2005/2/23 This document was turned in by a student as a homework paper. 1. Methods First, the cartesian coordinates of 5021 and 8021 molecules (Fig. 1) are generated, in
Quivers of Period 2. Mariya Sardarli Max Wimberley Heyi Zhu. November 26, 2014
 Quivers of Period 2 Mariya Sardarli Max Wimberley Heyi Zhu ovember 26, 2014 Abstract A quiver with vertices labeled from 1,..., n is said to have period 2 if the quiver obtained by mutating at 1 and then
Quivers of Period 2 Mariya Sardarli Max Wimberley Heyi Zhu ovember 26, 2014 Abstract A quiver with vertices labeled from 1,..., n is said to have period 2 if the quiver obtained by mutating at 1 and then
Connecting NMR data to biomolecular structure and dynamics
 Connecting NMR data to biomolecular structure and dynamics David A. Case Chem 538, Spring, 2014 Basics of NMR All nuclei are characterized by a spin quantum number I, which can be 0, 1/2, 1, 3/2, 2...
Connecting NMR data to biomolecular structure and dynamics David A. Case Chem 538, Spring, 2014 Basics of NMR All nuclei are characterized by a spin quantum number I, which can be 0, 1/2, 1, 3/2, 2...
CMPS 6630: Introduction to Computational Biology and Bioinformatics. Structure Comparison
 CMPS 6630: Introduction to Computational Biology and Bioinformatics Structure Comparison Protein Structure Comparison Motivation Understand sequence and structure variability Understand Domain architecture
CMPS 6630: Introduction to Computational Biology and Bioinformatics Structure Comparison Protein Structure Comparison Motivation Understand sequence and structure variability Understand Domain architecture
A NOTE ON TENSOR CATEGORIES OF LIE TYPE E 9
 A NOTE ON TENSOR CATEGORIES OF LIE TYPE E 9 ERIC C. ROWELL Abstract. We consider the problem of decomposing tensor powers of the fundamental level 1 highest weight representation V of the affine Kac-Moody
A NOTE ON TENSOR CATEGORIES OF LIE TYPE E 9 ERIC C. ROWELL Abstract. We consider the problem of decomposing tensor powers of the fundamental level 1 highest weight representation V of the affine Kac-Moody
Orientational degeneracy in the presence of one alignment tensor.
 Orientational degeneracy in the presence of one alignment tensor. Rotation about the x, y and z axes can be performed in the aligned mode of the program to examine the four degenerate orientations of two
Orientational degeneracy in the presence of one alignment tensor. Rotation about the x, y and z axes can be performed in the aligned mode of the program to examine the four degenerate orientations of two
Secondary and sidechain structures
 Lecture 2 Secondary and sidechain structures James Chou BCMP201 Spring 2008 Images from Petsko & Ringe, Protein Structure and Function. Branden & Tooze, Introduction to Protein Structure. Richardson, J.
Lecture 2 Secondary and sidechain structures James Chou BCMP201 Spring 2008 Images from Petsko & Ringe, Protein Structure and Function. Branden & Tooze, Introduction to Protein Structure. Richardson, J.
Outline. Levels of Protein Structure. Primary (1 ) Structure. Lecture 6:Protein Architecture II: Secondary Structure or From peptides to proteins
 Lecture 6:Protein Architecture II: Secondary Structure or From peptides to proteins Margaret Daugherty Fall 2004 Outline Four levels of structure are used to describe proteins; Alpha helices and beta sheets
Lecture 6:Protein Architecture II: Secondary Structure or From peptides to proteins Margaret Daugherty Fall 2004 Outline Four levels of structure are used to describe proteins; Alpha helices and beta sheets
Packing of Secondary Structures
 7.88 Lecture Notes - 4 7.24/7.88J/5.48J The Protein Folding and Human Disease Professor Gossard Retrieving, Viewing Protein Structures from the Protein Data Base Helix helix packing Packing of Secondary
7.88 Lecture Notes - 4 7.24/7.88J/5.48J The Protein Folding and Human Disease Professor Gossard Retrieving, Viewing Protein Structures from the Protein Data Base Helix helix packing Packing of Secondary
PROTEIN'STRUCTURE'DETERMINATION'
 PROTEIN'STRUCTURE'DETERMINATION' USING'NMR'RESTRAINTS' BCMB/CHEM'8190' Programs for NMR Based Structure Determination CNS - Brünger, A. T.; Adams, P. D.; Clore, G. M.; DeLano, W. L.; Gros, P.; Grosse-Kunstleve,
PROTEIN'STRUCTURE'DETERMINATION' USING'NMR'RESTRAINTS' BCMB/CHEM'8190' Programs for NMR Based Structure Determination CNS - Brünger, A. T.; Adams, P. D.; Clore, G. M.; DeLano, W. L.; Gros, P.; Grosse-Kunstleve,
Protein Structure. W. M. Grogan, Ph.D. OBJECTIVES
 Protein Structure W. M. Grogan, Ph.D. OBJECTIVES 1. Describe the structure and characteristic properties of typical proteins. 2. List and describe the four levels of structure found in proteins. 3. Relate
Protein Structure W. M. Grogan, Ph.D. OBJECTIVES 1. Describe the structure and characteristic properties of typical proteins. 2. List and describe the four levels of structure found in proteins. 3. Relate
Articles HIGHER BLOCK IFS 2: RELATIONS BETWEEN IFS WITH DIFFERENT LEVELS OF MEMORY
 Fractals, Vol. 8, No. 4 (200) 399 408 c World Scientific Publishing Company DOI: 0.42/S028348X0005044 Articles Fractals 200.8:399-408. Downloaded from www.worldscientific.com HIGHER BLOCK IFS 2: RELATIONS
Fractals, Vol. 8, No. 4 (200) 399 408 c World Scientific Publishing Company DOI: 0.42/S028348X0005044 Articles Fractals 200.8:399-408. Downloaded from www.worldscientific.com HIGHER BLOCK IFS 2: RELATIONS
On Systems of Diagonal Forms II
 On Systems of Diagonal Forms II Michael P Knapp 1 Introduction In a recent paper [8], we considered the system F of homogeneous additive forms F 1 (x) = a 11 x k 1 1 + + a 1s x k 1 s F R (x) = a R1 x k
On Systems of Diagonal Forms II Michael P Knapp 1 Introduction In a recent paper [8], we considered the system F of homogeneous additive forms F 1 (x) = a 11 x k 1 1 + + a 1s x k 1 s F R (x) = a R1 x k
EE731 Lecture Notes: Matrix Computations for Signal Processing
 EE731 Lecture Notes: Matrix Computations for Signal Processing James P. Reilly c Department of Electrical and Computer Engineering McMaster University September 22, 2005 0 Preface This collection of ten
EE731 Lecture Notes: Matrix Computations for Signal Processing James P. Reilly c Department of Electrical and Computer Engineering McMaster University September 22, 2005 0 Preface This collection of ten
Useful background reading
 Overview of lecture * General comment on peptide bond * Discussion of backbone dihedral angles * Discussion of Ramachandran plots * Description of helix types. * Description of structures * NMR patterns
Overview of lecture * General comment on peptide bond * Discussion of backbone dihedral angles * Discussion of Ramachandran plots * Description of helix types. * Description of structures * NMR patterns
1 Matrices and Systems of Linear Equations
 March 3, 203 6-6. Systems of Linear Equations Matrices and Systems of Linear Equations An m n matrix is an array A = a ij of the form a a n a 2 a 2n... a m a mn where each a ij is a real or complex number.
March 3, 203 6-6. Systems of Linear Equations Matrices and Systems of Linear Equations An m n matrix is an array A = a ij of the form a a n a 2 a 2n... a m a mn where each a ij is a real or complex number.
Computer simulations of protein folding with a small number of distance restraints
 Vol. 49 No. 3/2002 683 692 QUARTERLY Computer simulations of protein folding with a small number of distance restraints Andrzej Sikorski 1, Andrzej Kolinski 1,2 and Jeffrey Skolnick 2 1 Department of Chemistry,
Vol. 49 No. 3/2002 683 692 QUARTERLY Computer simulations of protein folding with a small number of distance restraints Andrzej Sikorski 1, Andrzej Kolinski 1,2 and Jeffrey Skolnick 2 1 Department of Chemistry,
CS 246 Review of Linear Algebra 01/17/19
 1 Linear algebra In this section we will discuss vectors and matrices. We denote the (i, j)th entry of a matrix A as A ij, and the ith entry of a vector as v i. 1.1 Vectors and vector operations A vector
1 Linear algebra In this section we will discuss vectors and matrices. We denote the (i, j)th entry of a matrix A as A ij, and the ith entry of a vector as v i. 1.1 Vectors and vector operations A vector
DS-GA 1002 Lecture notes 0 Fall Linear Algebra. These notes provide a review of basic concepts in linear algebra.
 DS-GA 1002 Lecture notes 0 Fall 2016 Linear Algebra These notes provide a review of basic concepts in linear algebra. 1 Vector spaces You are no doubt familiar with vectors in R 2 or R 3, i.e. [ ] 1.1
DS-GA 1002 Lecture notes 0 Fall 2016 Linear Algebra These notes provide a review of basic concepts in linear algebra. 1 Vector spaces You are no doubt familiar with vectors in R 2 or R 3, i.e. [ ] 1.1
Foundations of Matrix Analysis
 1 Foundations of Matrix Analysis In this chapter we recall the basic elements of linear algebra which will be employed in the remainder of the text For most of the proofs as well as for the details, the
1 Foundations of Matrix Analysis In this chapter we recall the basic elements of linear algebra which will be employed in the remainder of the text For most of the proofs as well as for the details, the
Math Camp Lecture 4: Linear Algebra. Xiao Yu Wang. Aug 2010 MIT. Xiao Yu Wang (MIT) Math Camp /10 1 / 88
 Math Camp 2010 Lecture 4: Linear Algebra Xiao Yu Wang MIT Aug 2010 Xiao Yu Wang (MIT) Math Camp 2010 08/10 1 / 88 Linear Algebra Game Plan Vector Spaces Linear Transformations and Matrices Determinant
Math Camp 2010 Lecture 4: Linear Algebra Xiao Yu Wang MIT Aug 2010 Xiao Yu Wang (MIT) Math Camp 2010 08/10 1 / 88 Linear Algebra Game Plan Vector Spaces Linear Transformations and Matrices Determinant
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/309/5742/1868/dc1 Supporting Online Material for Toward High-Resolution de Novo Structure Prediction for Small Proteins Philip Bradley, Kira M. S. Misura, David Baker*
www.sciencemag.org/cgi/content/full/309/5742/1868/dc1 Supporting Online Material for Toward High-Resolution de Novo Structure Prediction for Small Proteins Philip Bradley, Kira M. S. Misura, David Baker*
Protein Structure Determination Using NMR Restraints BCMB/CHEM 8190
 Protein Structure Determination Using NMR Restraints BCMB/CHEM 8190 Programs for NMR Based Structure Determination CNS - Brunger, A. T.; Adams, P. D.; Clore, G. M.; DeLano, W. L.; Gros, P.; Grosse-Kunstleve,
Protein Structure Determination Using NMR Restraints BCMB/CHEM 8190 Programs for NMR Based Structure Determination CNS - Brunger, A. T.; Adams, P. D.; Clore, G. M.; DeLano, W. L.; Gros, P.; Grosse-Kunstleve,
Structural and mechanistic insight into the substrate. binding from the conformational dynamics in apo. and substrate-bound DapE enzyme
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 215 Structural and mechanistic insight into the substrate binding from the conformational
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 215 Structural and mechanistic insight into the substrate binding from the conformational
Conformational Geometry of Peptides and Proteins:
 Conformational Geometry of Peptides and Proteins: Before discussing secondary structure, it is important to appreciate the conformational plasticity of proteins. Each residue in a polypeptide has three
Conformational Geometry of Peptides and Proteins: Before discussing secondary structure, it is important to appreciate the conformational plasticity of proteins. Each residue in a polypeptide has three
Lagrange Multipliers
 Optimization with Constraints As long as algebra and geometry have been separated, their progress have been slow and their uses limited; but when these two sciences have been united, they have lent each
Optimization with Constraints As long as algebra and geometry have been separated, their progress have been slow and their uses limited; but when these two sciences have been united, they have lent each
Solving distance geometry problems for protein structure determination
 Graduate Theses and Dissertations Iowa State University Capstones, Theses and Dissertations 2010 Solving distance geometry problems for protein structure determination Atilla Sit Iowa State University
Graduate Theses and Dissertations Iowa State University Capstones, Theses and Dissertations 2010 Solving distance geometry problems for protein structure determination Atilla Sit Iowa State University
What is A + B? What is A B? What is AB? What is BA? What is A 2? and B = QUESTION 2. What is the reduced row echelon matrix of A =
 STUDENT S COMPANIONS IN BASIC MATH: THE ELEVENTH Matrix Reloaded by Block Buster Presumably you know the first part of matrix story, including its basic operations (addition and multiplication) and row
STUDENT S COMPANIONS IN BASIC MATH: THE ELEVENTH Matrix Reloaded by Block Buster Presumably you know the first part of matrix story, including its basic operations (addition and multiplication) and row
Protein Structure Prediction
 Page 1 Protein Structure Prediction Russ B. Altman BMI 214 CS 274 Protein Folding is different from structure prediction --Folding is concerned with the process of taking the 3D shape, usually based on
Page 1 Protein Structure Prediction Russ B. Altman BMI 214 CS 274 Protein Folding is different from structure prediction --Folding is concerned with the process of taking the 3D shape, usually based on
Analysis-3 lecture schemes
 Analysis-3 lecture schemes (with Homeworks) 1 Csörgő István November, 2015 1 A jegyzet az ELTE Informatikai Kar 2015. évi Jegyzetpályázatának támogatásával készült Contents 1. Lesson 1 4 1.1. The Space
Analysis-3 lecture schemes (with Homeworks) 1 Csörgő István November, 2015 1 A jegyzet az ELTE Informatikai Kar 2015. évi Jegyzetpályázatának támogatásával készült Contents 1. Lesson 1 4 1.1. The Space
CAP 5510 Lecture 3 Protein Structures
 CAP 5510 Lecture 3 Protein Structures Su-Shing Chen Bioinformatics CISE 8/19/2005 Su-Shing Chen, CISE 1 Protein Conformation 8/19/2005 Su-Shing Chen, CISE 2 Protein Conformational Structures Hydrophobicity
CAP 5510 Lecture 3 Protein Structures Su-Shing Chen Bioinformatics CISE 8/19/2005 Su-Shing Chen, CISE 1 Protein Conformation 8/19/2005 Su-Shing Chen, CISE 2 Protein Conformational Structures Hydrophobicity
Presentation Outline. Prediction of Protein Secondary Structure using Neural Networks at Better than 70% Accuracy
 Prediction of Protein Secondary Structure using Neural Networks at Better than 70% Accuracy Burkhard Rost and Chris Sander By Kalyan C. Gopavarapu 1 Presentation Outline Major Terminology Problem Method
Prediction of Protein Secondary Structure using Neural Networks at Better than 70% Accuracy Burkhard Rost and Chris Sander By Kalyan C. Gopavarapu 1 Presentation Outline Major Terminology Problem Method
Math 302 Outcome Statements Winter 2013
 Math 302 Outcome Statements Winter 2013 1 Rectangular Space Coordinates; Vectors in the Three-Dimensional Space (a) Cartesian coordinates of a point (b) sphere (c) symmetry about a point, a line, and a
Math 302 Outcome Statements Winter 2013 1 Rectangular Space Coordinates; Vectors in the Three-Dimensional Space (a) Cartesian coordinates of a point (b) sphere (c) symmetry about a point, a line, and a
arxiv:cond-mat/ v1 2 Feb 94
 cond-mat/9402010 Properties and Origins of Protein Secondary Structure Nicholas D. Socci (1), William S. Bialek (2), and José Nelson Onuchic (1) (1) Department of Physics, University of California at San
cond-mat/9402010 Properties and Origins of Protein Secondary Structure Nicholas D. Socci (1), William S. Bialek (2), and José Nelson Onuchic (1) (1) Department of Physics, University of California at San
Better Bond Angles in the Protein Data Bank
 Better Bond Angles in the Protein Data Bank C.J. Robinson and D.B. Skillicorn School of Computing Queen s University {robinson,skill}@cs.queensu.ca Abstract The Protein Data Bank (PDB) contains, at least
Better Bond Angles in the Protein Data Bank C.J. Robinson and D.B. Skillicorn School of Computing Queen s University {robinson,skill}@cs.queensu.ca Abstract The Protein Data Bank (PDB) contains, at least
Motif Prediction in Amino Acid Interaction Networks
 Motif Prediction in Amino Acid Interaction Networks Omar GACI and Stefan BALEV Abstract In this paper we represent a protein as a graph where the vertices are amino acids and the edges are interactions
Motif Prediction in Amino Acid Interaction Networks Omar GACI and Stefan BALEV Abstract In this paper we represent a protein as a graph where the vertices are amino acids and the edges are interactions
Supersecondary Structures (structural motifs)
 Supersecondary Structures (structural motifs) Various Sources Slide 1 Supersecondary Structures (Motifs) Supersecondary Structures (Motifs): : Combinations of secondary structures in specific geometric
Supersecondary Structures (structural motifs) Various Sources Slide 1 Supersecondary Structures (Motifs) Supersecondary Structures (Motifs): : Combinations of secondary structures in specific geometric
Lecture 7. Econ August 18
 Lecture 7 Econ 2001 2015 August 18 Lecture 7 Outline First, the theorem of the maximum, an amazing result about continuity in optimization problems. Then, we start linear algebra, mostly looking at familiar
Lecture 7 Econ 2001 2015 August 18 Lecture 7 Outline First, the theorem of the maximum, an amazing result about continuity in optimization problems. Then, we start linear algebra, mostly looking at familiar
Solving Geometric Constraints with Distance-Based Global Coordinate System
 International Workshop on Geometric Constraint Solving, October 23, 2003, Beijing 1 Solving Geometric Constraints with Distance-Based Global Coordinate System Lu Yang 1) Extended Abstract Distance geometry
International Workshop on Geometric Constraint Solving, October 23, 2003, Beijing 1 Solving Geometric Constraints with Distance-Based Global Coordinate System Lu Yang 1) Extended Abstract Distance geometry
Bio nformatics. Lecture 23. Saad Mneimneh
 Bio nformatics Lecture 23 Protein folding The goal is to determine the three-dimensional structure of a protein based on its amino acid sequence Assumption: amino acid sequence completely and uniquely
Bio nformatics Lecture 23 Protein folding The goal is to determine the three-dimensional structure of a protein based on its amino acid sequence Assumption: amino acid sequence completely and uniquely
Practical Linear Algebra: A Geometry Toolbox
 Practical Linear Algebra: A Geometry Toolbox Third edition Chapter 12: Gauss for Linear Systems Gerald Farin & Dianne Hansford CRC Press, Taylor & Francis Group, An A K Peters Book www.farinhansford.com/books/pla
Practical Linear Algebra: A Geometry Toolbox Third edition Chapter 12: Gauss for Linear Systems Gerald Farin & Dianne Hansford CRC Press, Taylor & Francis Group, An A K Peters Book www.farinhansford.com/books/pla
Linear Programming Redux
 Linear Programming Redux Jim Bremer May 12, 2008 The purpose of these notes is to review the basics of linear programming and the simplex method in a clear, concise, and comprehensive way. The book contains
Linear Programming Redux Jim Bremer May 12, 2008 The purpose of these notes is to review the basics of linear programming and the simplex method in a clear, concise, and comprehensive way. The book contains
Sequential resonance assignments in (small) proteins: homonuclear method 2º structure determination
 Lecture 9 M230 Feigon Sequential resonance assignments in (small) proteins: homonuclear method 2º structure determination Reading resources v Roberts NMR of Macromolecules, Chap 4 by Christina Redfield
Lecture 9 M230 Feigon Sequential resonance assignments in (small) proteins: homonuclear method 2º structure determination Reading resources v Roberts NMR of Macromolecules, Chap 4 by Christina Redfield
Powerful Simulated-Annealing Algorithm Locates Global Minimum of Protein-Folding Potentials from Multiple Starting Conformations
 Powerful Simulated-Annealing Algorithm Locates Global Minimum of Protein-Folding Potentials from Multiple Starting Conformations Mark E. Snow University of Michigan, Scientific Computation Croup, 535 W.
Powerful Simulated-Annealing Algorithm Locates Global Minimum of Protein-Folding Potentials from Multiple Starting Conformations Mark E. Snow University of Michigan, Scientific Computation Croup, 535 W.
Linear Algebra March 16, 2019
 Linear Algebra March 16, 2019 2 Contents 0.1 Notation................................ 4 1 Systems of linear equations, and matrices 5 1.1 Systems of linear equations..................... 5 1.2 Augmented
Linear Algebra March 16, 2019 2 Contents 0.1 Notation................................ 4 1 Systems of linear equations, and matrices 5 1.1 Systems of linear equations..................... 5 1.2 Augmented
1 Matrices and Systems of Linear Equations. a 1n a 2n
 March 31, 2013 16-1 16. Systems of Linear Equations 1 Matrices and Systems of Linear Equations An m n matrix is an array A = (a ij ) of the form a 11 a 21 a m1 a 1n a 2n... a mn where each a ij is a real
March 31, 2013 16-1 16. Systems of Linear Equations 1 Matrices and Systems of Linear Equations An m n matrix is an array A = (a ij ) of the form a 11 a 21 a m1 a 1n a 2n... a mn where each a ij is a real
Chapter 2 Spectra of Finite Graphs
 Chapter 2 Spectra of Finite Graphs 2.1 Characteristic Polynomials Let G = (V, E) be a finite graph on n = V vertices. Numbering the vertices, we write down its adjacency matrix in an explicit form of n
Chapter 2 Spectra of Finite Graphs 2.1 Characteristic Polynomials Let G = (V, E) be a finite graph on n = V vertices. Numbering the vertices, we write down its adjacency matrix in an explicit form of n
Protein folding. α-helix. Lecture 21. An α-helix is a simple helix having on average 10 residues (3 turns of the helix)
 Computat onal Biology Lecture 21 Protein folding The goal is to determine the three-dimensional structure of a protein based on its amino acid sequence Assumption: amino acid sequence completely and uniquely
Computat onal Biology Lecture 21 Protein folding The goal is to determine the three-dimensional structure of a protein based on its amino acid sequence Assumption: amino acid sequence completely and uniquely
Lecture 7: Semidefinite programming
 CS 766/QIC 820 Theory of Quantum Information (Fall 2011) Lecture 7: Semidefinite programming This lecture is on semidefinite programming, which is a powerful technique from both an analytic and computational
CS 766/QIC 820 Theory of Quantum Information (Fall 2011) Lecture 7: Semidefinite programming This lecture is on semidefinite programming, which is a powerful technique from both an analytic and computational
Vectors. January 13, 2013
 Vectors January 13, 2013 The simplest tensors are scalars, which are the measurable quantities of a theory, left invariant by symmetry transformations. By far the most common non-scalars are the vectors,
Vectors January 13, 2013 The simplest tensors are scalars, which are the measurable quantities of a theory, left invariant by symmetry transformations. By far the most common non-scalars are the vectors,
1 Multiply Eq. E i by λ 0: (λe i ) (E i ) 2 Multiply Eq. E j by λ and add to Eq. E i : (E i + λe j ) (E i )
 Direct Methods for Linear Systems Chapter Direct Methods for Solving Linear Systems Per-Olof Persson persson@berkeleyedu Department of Mathematics University of California, Berkeley Math 18A Numerical
Direct Methods for Linear Systems Chapter Direct Methods for Solving Linear Systems Per-Olof Persson persson@berkeleyedu Department of Mathematics University of California, Berkeley Math 18A Numerical
PROJECT C: ELECTRONIC BAND STRUCTURE IN A MODEL SEMICONDUCTOR
 PROJECT C: ELECTRONIC BAND STRUCTURE IN A MODEL SEMICONDUCTOR The aim of this project is to present the student with a perspective on the notion of electronic energy band structures and energy band gaps
PROJECT C: ELECTRONIC BAND STRUCTURE IN A MODEL SEMICONDUCTOR The aim of this project is to present the student with a perspective on the notion of electronic energy band structures and energy band gaps
Discrete Applied Mathematics
 Discrete Applied Mathematics 194 (015) 37 59 Contents lists available at ScienceDirect Discrete Applied Mathematics journal homepage: wwwelseviercom/locate/dam Loopy, Hankel, and combinatorially skew-hankel
Discrete Applied Mathematics 194 (015) 37 59 Contents lists available at ScienceDirect Discrete Applied Mathematics journal homepage: wwwelseviercom/locate/dam Loopy, Hankel, and combinatorially skew-hankel
Polypeptide Folding Using Monte Carlo Sampling, Concerted Rotation, and Continuum Solvation
 Polypeptide Folding Using Monte Carlo Sampling, Concerted Rotation, and Continuum Solvation Jakob P. Ulmschneider and William L. Jorgensen J.A.C.S. 2004, 126, 1849-1857 Presented by Laura L. Thomas and
Polypeptide Folding Using Monte Carlo Sampling, Concerted Rotation, and Continuum Solvation Jakob P. Ulmschneider and William L. Jorgensen J.A.C.S. 2004, 126, 1849-1857 Presented by Laura L. Thomas and
Design of a Novel Globular Protein Fold with Atomic-Level Accuracy
 Design of a Novel Globular Protein Fold with Atomic-Level Accuracy Brian Kuhlman, Gautam Dantas, Gregory C. Ireton, Gabriele Varani, Barry L. Stoddard, David Baker Presented by Kate Stafford 4 May 05 Protein
Design of a Novel Globular Protein Fold with Atomic-Level Accuracy Brian Kuhlman, Gautam Dantas, Gregory C. Ireton, Gabriele Varani, Barry L. Stoddard, David Baker Presented by Kate Stafford 4 May 05 Protein
Copositive Plus Matrices
 Copositive Plus Matrices Willemieke van Vliet Master Thesis in Applied Mathematics October 2011 Copositive Plus Matrices Summary In this report we discuss the set of copositive plus matrices and their
Copositive Plus Matrices Willemieke van Vliet Master Thesis in Applied Mathematics October 2011 Copositive Plus Matrices Summary In this report we discuss the set of copositive plus matrices and their
Outline. Levels of Protein Structure. Primary (1 ) Structure. Lecture 6:Protein Architecture II: Secondary Structure or From peptides to proteins
 Lecture 6:Protein Architecture II: Secondary Structure or From peptides to proteins Margaret Daugherty Fall 2003 Outline Four levels of structure are used to describe proteins; Alpha helices and beta sheets
Lecture 6:Protein Architecture II: Secondary Structure or From peptides to proteins Margaret Daugherty Fall 2003 Outline Four levels of structure are used to describe proteins; Alpha helices and beta sheets
GROUP THEORY PRIMER. New terms: so(2n), so(2n+1), symplectic algebra sp(2n)
 GROUP THEORY PRIMER New terms: so(2n), so(2n+1), symplectic algebra sp(2n) 1. Some examples of semi-simple Lie algebras In the previous chapter, we developed the idea of understanding semi-simple Lie algebras
GROUP THEORY PRIMER New terms: so(2n), so(2n+1), symplectic algebra sp(2n) 1. Some examples of semi-simple Lie algebras In the previous chapter, we developed the idea of understanding semi-simple Lie algebras
Triangular Plate Displacement Elements
 Triangular Plate Displacement Elements Chapter : TRIANGULAR PLATE DISPLACEMENT ELEMENTS TABLE OF CONTENTS Page. Introduction...................... Triangular Element Properties................ Triangle
Triangular Plate Displacement Elements Chapter : TRIANGULAR PLATE DISPLACEMENT ELEMENTS TABLE OF CONTENTS Page. Introduction...................... Triangular Element Properties................ Triangle
Self-Organizing Superimposition Algorithm for Conformational Sampling
 Self-Organizing Superimposition Algorithm for Conformational Sampling FANGQIANG ZHU, DIMITRIS K. AGRAFIOTIS Johnson & Johnson Pharmaceutical Research & Development, L.L.C., 665 Stockton Drive, Exton, Pennsylvania
Self-Organizing Superimposition Algorithm for Conformational Sampling FANGQIANG ZHU, DIMITRIS K. AGRAFIOTIS Johnson & Johnson Pharmaceutical Research & Development, L.L.C., 665 Stockton Drive, Exton, Pennsylvania
114 Grundlagen der Bioinformatik, SS 09, D. Huson, July 6, 2009
 114 Grundlagen der Bioinformatik, SS 09, D. Huson, July 6, 2009 9 Protein tertiary structure Sources for this chapter, which are all recommended reading: D.W. Mount. Bioinformatics: Sequences and Genome
114 Grundlagen der Bioinformatik, SS 09, D. Huson, July 6, 2009 9 Protein tertiary structure Sources for this chapter, which are all recommended reading: D.W. Mount. Bioinformatics: Sequences and Genome
n-step mingling inequalities: new facets for the mixed-integer knapsack set
 Math. Program., Ser. A (2012) 132:79 98 DOI 10.1007/s10107-010-0382-6 FULL LENGTH PAPER n-step mingling inequalities: new facets for the mixed-integer knapsack set Alper Atamtürk Kiavash Kianfar Received:
Math. Program., Ser. A (2012) 132:79 98 DOI 10.1007/s10107-010-0382-6 FULL LENGTH PAPER n-step mingling inequalities: new facets for the mixed-integer knapsack set Alper Atamtürk Kiavash Kianfar Received:
DATE A DAtabase of TIM Barrel Enzymes
 DATE A DAtabase of TIM Barrel Enzymes 2 2.1 Introduction.. 2.2 Objective and salient features of the database 2.2.1 Choice of the dataset.. 2.3 Statistical information on the database.. 2.4 Features....
DATE A DAtabase of TIM Barrel Enzymes 2 2.1 Introduction.. 2.2 Objective and salient features of the database 2.2.1 Choice of the dataset.. 2.3 Statistical information on the database.. 2.4 Features....
Boolean Inner-Product Spaces and Boolean Matrices
 Boolean Inner-Product Spaces and Boolean Matrices Stan Gudder Department of Mathematics, University of Denver, Denver CO 80208 Frédéric Latrémolière Department of Mathematics, University of Denver, Denver
Boolean Inner-Product Spaces and Boolean Matrices Stan Gudder Department of Mathematics, University of Denver, Denver CO 80208 Frédéric Latrémolière Department of Mathematics, University of Denver, Denver
Chapter 7. Extremal Problems. 7.1 Extrema and Local Extrema
 Chapter 7 Extremal Problems No matter in theoretical context or in applications many problems can be formulated as problems of finding the maximum or minimum of a function. Whenever this is the case, advanced
Chapter 7 Extremal Problems No matter in theoretical context or in applications many problems can be formulated as problems of finding the maximum or minimum of a function. Whenever this is the case, advanced
Topological Data Analysis - Spring 2018
 Topological Data Analysis - Spring 2018 Simplicial Homology Slightly rearranged, but mostly copy-pasted from Harer s and Edelsbrunner s Computational Topology, Verovsek s class notes. Gunnar Carlsson s
Topological Data Analysis - Spring 2018 Simplicial Homology Slightly rearranged, but mostly copy-pasted from Harer s and Edelsbrunner s Computational Topology, Verovsek s class notes. Gunnar Carlsson s
Molecular Modeling Lecture 7. Homology modeling insertions/deletions manual realignment
 Molecular Modeling 2018-- Lecture 7 Homology modeling insertions/deletions manual realignment Homology modeling also called comparative modeling Sequences that have similar sequence have similar structure.
Molecular Modeling 2018-- Lecture 7 Homology modeling insertions/deletions manual realignment Homology modeling also called comparative modeling Sequences that have similar sequence have similar structure.
Biomolecules: lecture 10
 Biomolecules: lecture 10 - understanding in detail how protein 3D structures form - realize that protein molecules are not static wire models but instead dynamic, where in principle every atom moves (yet
Biomolecules: lecture 10 - understanding in detail how protein 3D structures form - realize that protein molecules are not static wire models but instead dynamic, where in principle every atom moves (yet
This pre-publication material is for review purposes only. Any typographical or technical errors will be corrected prior to publication.
 This pre-publication material is for review purposes only. Any typographical or technical errors will be corrected prior to publication. Copyright Pearson Canada Inc. All rights reserved. Copyright Pearson
This pre-publication material is for review purposes only. Any typographical or technical errors will be corrected prior to publication. Copyright Pearson Canada Inc. All rights reserved. Copyright Pearson
Ramachandran and his Map
 Ramachandran and his Map C Ramakrishnan Introduction C Ramakrishnan, retired professor from Molecular Biophysics Unit, Indian Institute of Science, Bangalore, has been associated with Professor G N Ramachandran
Ramachandran and his Map C Ramakrishnan Introduction C Ramakrishnan, retired professor from Molecular Biophysics Unit, Indian Institute of Science, Bangalore, has been associated with Professor G N Ramachandran
Introduction to" Protein Structure
 Introduction to" Protein Structure Function, evolution & experimental methods Thomas Blicher, Center for Biological Sequence Analysis Learning Objectives Outline the basic levels of protein structure.
Introduction to" Protein Structure Function, evolution & experimental methods Thomas Blicher, Center for Biological Sequence Analysis Learning Objectives Outline the basic levels of protein structure.
Using distance geomtry to generate structures
 Using distance geomtry to generate structures David A. Case Genomic systems and structures, Spring, 2009 Converting distances to structures Metric Matrix Distance Geometry To describe a molecule in terms
Using distance geomtry to generate structures David A. Case Genomic systems and structures, Spring, 2009 Converting distances to structures Metric Matrix Distance Geometry To describe a molecule in terms
CS273: Algorithms for Structure Handout # 2 and Motion in Biology Stanford University Thursday, 1 April 2004
 CS273: Algorithms for Structure Handout # 2 and Motion in Biology Stanford University Thursday, 1 April 2004 Lecture #2: 1 April 2004 Topics: Kinematics : Concepts and Results Kinematics of Ligands and
CS273: Algorithms for Structure Handout # 2 and Motion in Biology Stanford University Thursday, 1 April 2004 Lecture #2: 1 April 2004 Topics: Kinematics : Concepts and Results Kinematics of Ligands and
Algorithm for Rapid Reconstruction of Protein Backbone from Alpha Carbon Coordinates
 Algorithm for Rapid Reconstruction of Protein Backbone from Alpha Carbon Coordinates MARIUSZ MILIK, 1 *, ANDRZEJ KOLINSKI, 1, 2 and JEFFREY SKOLNICK 1 1 The Scripps Research Institute, Department of Molecular
Algorithm for Rapid Reconstruction of Protein Backbone from Alpha Carbon Coordinates MARIUSZ MILIK, 1 *, ANDRZEJ KOLINSKI, 1, 2 and JEFFREY SKOLNICK 1 1 The Scripps Research Institute, Department of Molecular
CMPS 3110: Bioinformatics. Tertiary Structure Prediction
 CMPS 3110: Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the laws of physics! Conformation space is finite
CMPS 3110: Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the laws of physics! Conformation space is finite
Incompatibility Paradoxes
 Chapter 22 Incompatibility Paradoxes 22.1 Simultaneous Values There is never any difficulty in supposing that a classical mechanical system possesses, at a particular instant of time, precise values of
Chapter 22 Incompatibility Paradoxes 22.1 Simultaneous Values There is never any difficulty in supposing that a classical mechanical system possesses, at a particular instant of time, precise values of
CMPS 6630: Introduction to Computational Biology and Bioinformatics. Tertiary Structure Prediction
 CMPS 6630: Introduction to Computational Biology and Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the
CMPS 6630: Introduction to Computational Biology and Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the
Exploring the energy landscape
 Exploring the energy landscape ChE210D Today's lecture: what are general features of the potential energy surface and how can we locate and characterize minima on it Derivatives of the potential energy
Exploring the energy landscape ChE210D Today's lecture: what are general features of the potential energy surface and how can we locate and characterize minima on it Derivatives of the potential energy
RNA and Protein Structure Prediction
 RNA and Protein Structure Prediction Bioinformatics: Issues and Algorithms CSE 308-408 Spring 2007 Lecture 18-1- Outline Multi-Dimensional Nature of Life RNA Secondary Structure Prediction Protein Structure
RNA and Protein Structure Prediction Bioinformatics: Issues and Algorithms CSE 308-408 Spring 2007 Lecture 18-1- Outline Multi-Dimensional Nature of Life RNA Secondary Structure Prediction Protein Structure
The Jordan Canonical Form
 The Jordan Canonical Form The Jordan canonical form describes the structure of an arbitrary linear transformation on a finite-dimensional vector space over an algebraically closed field. Here we develop
The Jordan Canonical Form The Jordan canonical form describes the structure of an arbitrary linear transformation on a finite-dimensional vector space over an algebraically closed field. Here we develop
Sequential Assignment Strategies in Proteins
 Sequential Assignment Strategies in Proteins NMR assignments in order to determine a structure by traditional, NOE-based 1 H- 1 H distance-based methods, the chemical shifts of the individual 1 H nuclei
Sequential Assignment Strategies in Proteins NMR assignments in order to determine a structure by traditional, NOE-based 1 H- 1 H distance-based methods, the chemical shifts of the individual 1 H nuclei
Stereochemistry ofpolypeptide Chain Configurations
 J. Mol. Biol. (1963) 7,95-99 Stereochemistry ofpolypeptide Chain Configurations Various types of polypeptide chain configurations have been proposed in recent years for proteins as well as polypeptides,
J. Mol. Biol. (1963) 7,95-99 Stereochemistry ofpolypeptide Chain Configurations Various types of polypeptide chain configurations have been proposed in recent years for proteins as well as polypeptides,
Analysis and Linear Algebra. Lectures 1-3 on the mathematical tools that will be used in C103
 Analysis and Linear Algebra Lectures 1-3 on the mathematical tools that will be used in C103 Set Notation A, B sets AcB union A1B intersection A\B the set of objects in A that are not in B N. Empty set
Analysis and Linear Algebra Lectures 1-3 on the mathematical tools that will be used in C103 Set Notation A, B sets AcB union A1B intersection A\B the set of objects in A that are not in B N. Empty set
Prediction and refinement of NMR structures from sparse experimental data
 Prediction and refinement of NMR structures from sparse experimental data Jeff Skolnick Director Center for the Study of Systems Biology School of Biology Georgia Institute of Technology Overview of talk
Prediction and refinement of NMR structures from sparse experimental data Jeff Skolnick Director Center for the Study of Systems Biology School of Biology Georgia Institute of Technology Overview of talk
REVIEW FOR EXAM II. The exam covers sections , the part of 3.7 on Markov chains, and
 REVIEW FOR EXAM II The exam covers sections 3.4 3.6, the part of 3.7 on Markov chains, and 4.1 4.3. 1. The LU factorization: An n n matrix A has an LU factorization if A = LU, where L is lower triangular
REVIEW FOR EXAM II The exam covers sections 3.4 3.6, the part of 3.7 on Markov chains, and 4.1 4.3. 1. The LU factorization: An n n matrix A has an LU factorization if A = LU, where L is lower triangular
Nitrogenase MoFe protein from Clostridium pasteurianum at 1.08 Å resolution: comparison with the Azotobacter vinelandii MoFe protein
 Acta Cryst. (2015). D71, 274-282, doi:10.1107/s1399004714025243 Supporting information Volume 71 (2015) Supporting information for article: Nitrogenase MoFe protein from Clostridium pasteurianum at 1.08
Acta Cryst. (2015). D71, 274-282, doi:10.1107/s1399004714025243 Supporting information Volume 71 (2015) Supporting information for article: Nitrogenase MoFe protein from Clostridium pasteurianum at 1.08
A note on the minimal volume of almost cubic parallelepipeds
 A note on the minimal volume of almost cubic parallelepipeds Daniele Micciancio Abstract We prove that the best way to reduce the volume of the n-dimensional unit cube by a linear transformation that maps
A note on the minimal volume of almost cubic parallelepipeds Daniele Micciancio Abstract We prove that the best way to reduce the volume of the n-dimensional unit cube by a linear transformation that maps
Diagonalization. MATH 1502 Calculus II Notes. November 4, 2008
 Diagonalization MATH 1502 Calculus II Notes November 4, 2008 We want to understand all linear transformations L : R n R m. The basic case is the one in which n = m. That is to say, the case in which the
Diagonalization MATH 1502 Calculus II Notes November 4, 2008 We want to understand all linear transformations L : R n R m. The basic case is the one in which n = m. That is to say, the case in which the
Esser et al. Crystal Structures of R. sphaeroides bc 1
 Esser et al. Crystal Structures of R. sphaeroides bc Supplementary Information Trivariate Gaussian Probability Analysis The superposition of six structures results in sextets of 3D coordinates for every
Esser et al. Crystal Structures of R. sphaeroides bc Supplementary Information Trivariate Gaussian Probability Analysis The superposition of six structures results in sextets of 3D coordinates for every
Computing RMSD and fitting protein structures: how I do it and how others do it
 Computing RMSD and fitting protein structures: how I do it and how others do it Bertalan Kovács, Pázmány Péter Catholic University 03/08/2016 0. Introduction All the following algorithms have been implemented
Computing RMSD and fitting protein structures: how I do it and how others do it Bertalan Kovács, Pázmány Péter Catholic University 03/08/2016 0. Introduction All the following algorithms have been implemented
Protein Folding & Stability. Lecture 11: Margaret A. Daugherty. Fall How do we go from an unfolded polypeptide chain to a
 Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2004 How do we go from an unfolded polypeptide chain to a compact folded protein? (Folding of thioredoxin, F. Richards) Structure - Function
Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2004 How do we go from an unfolded polypeptide chain to a compact folded protein? (Folding of thioredoxin, F. Richards) Structure - Function
SUPPLEMENTARY INFORMATION
 www.nature.com/nature 1 Figure S1 Sequence alignment. a Structure based alignment of the plgic of E. chrysanthemi (ELIC), the acetylcholine binding protein from the snail Lymnea stagnalis (AchBP, PDB code
www.nature.com/nature 1 Figure S1 Sequence alignment. a Structure based alignment of the plgic of E. chrysanthemi (ELIC), the acetylcholine binding protein from the snail Lymnea stagnalis (AchBP, PDB code
Molecular dynamics simulations of anti-aggregation effect of ibuprofen. Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov
 Biophysical Journal, Volume 98 Supporting Material Molecular dynamics simulations of anti-aggregation effect of ibuprofen Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov Supplemental
Biophysical Journal, Volume 98 Supporting Material Molecular dynamics simulations of anti-aggregation effect of ibuprofen Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov Supplemental
14 Singular Value Decomposition
 14 Singular Value Decomposition For any high-dimensional data analysis, one s first thought should often be: can I use an SVD? The singular value decomposition is an invaluable analysis tool for dealing
14 Singular Value Decomposition For any high-dimensional data analysis, one s first thought should often be: can I use an SVD? The singular value decomposition is an invaluable analysis tool for dealing
