Computing RMSD and fitting protein structures: how I do it and how others do it
|
|
|
- Carol Cobb
- 5 years ago
- Views:
Transcription
1 Computing RMSD and fitting protein structures: how I do it and how others do it Bertalan Kovács, Pázmány Péter Catholic University 03/08/ Introduction All the following algorithms have been implemented into the java program PDBFitter and are available through its graphical interface. Furthermore, it is briefly reported how different programs (Chimera, MOLMOL) carry out the same calculations and operations in order to clarify possible differences in the final values. 1. Root mean square deviation (RMSD) The root mean square deviation is usually used to compare two or more protein structures and quantify their deviation Model-model RMSD RMSD can be defined for two structures that contain identical number and types of atoms. Let the two sets atomic coordinates be r i and r i (i = 1, 2,, n). In this case, RMSD between the two structures is RMSD = 1 n r i r i 2 n i=1 where r i r i 2 = (r i,x r i,x ) 2 + (r i,y r i,y ) 2 + (r i,z r i,z ) 2 is the square distance between two identical atomic positions. Naturally, r i and r i may refer to a subset of the entire molecule, such as C α atoms, backbone atoms or heavy atoms. Final RMSD value depends on the exact definition of the atom set r i. It is possible to define the weighted RMSD, in which the atomic distances are multiplied by the weights (e.g. atomic masses) w i.
2 n RMSD = 1 n w i r i r i 2 i=1 Structural ensembles of proteins usually contain more than two models of the same molecule. For such ensembles, RMSD can be calculated in three ways. Overall RMSD is usually given as the average and the variance of the single model-model RMSDs. (In the following formulae, assume that the ensemble consist of N models, and the RMSD between models k and l is denoted with RMSD k,l.) 1. RMSD can be calculated in a pairwise manner, and be given as an N N matrix. Such matrix contains as many independent values as the number of elements in the triangular matrix (diagonal not considered, since the RMSD of each model to itself is 0), which is N(N-1)/2. The overall average RMSD is therefore RMSD = 2 N(N 1) RMSD k,l k<l 2. RMSD can be given for each model relative to any chosen model, for example the first one. In this case RMSD = 1 N RMSD k,1 3. RMSD of each model can be calculated relative to the mean structure. It has to be kept in mind, that the mean structure no always represents a chemically realistic structure, especially if the ensemble is composed of more than one significantly different conformation. Let the atomic coordinates of the mean structure be r i and the r i atomic coordinate in model k be r i k. In this case the mean structure is and the overall average RMSD is given by k N r i = 1 N r i k k=1 RMSD = 1 N RMSD k,mean assuming that RMSD k,mean is the RMSD between model k and the mean structure. k 4. Finally, model-model RMSD can be calculated in a similar way as in method 1., but comparison of any two models is preceded by fitting of the same two models on each other. This results in a necessarily lower-or-equal average RMSD than that of method 1., since the pairwise RMSDs are minimized in each step. (Fitting is discussed in detail in Chapter 2.) It is of utmost importance to clearly define how the RMSD was calculated, i.e. based on which subset of the atomic positions (backbone, heavy atoms, etc.) and relative to which structure (pairwise
3 manner, to the mean structure or to a given model). The reason for this is that fitting algorithms (see Chapter 2.) guarantee minimal RMSD only if the same subset and algorithms are used for both fitting and RMSD calculation Local RMSD Internal motion of a specific region within the protein can be quantified with the local RMSD. Local RMSD is often calculated for a single atom or a group of atoms, e.g. a single residue. Similarly, it has to be accurately defined, which atoms of the residue are used for local RMSD calculation. The RMSD of a single atom (e.g. the C α of a given residue) can be calculated in two different ways. 1. In a pairwise manner, i.e. square distances are computed between all the possible pairs of identical atoms in different models. This also results in N(N-1)/2 distances, that have to be averaged. 2 RMSD pairwise (r i ) = N(N 1) r i k r i l 2 k<l where the indexes k and l go over the models of the ensemble. 2. Relative to the mean position of the given atom: It can be shown, that for high N RMSD to mean (r i ) = 1 N r i k r 2 i RMSD pairwise (r i ) = 2 RMSD to mean (r i ) Proof of the above statement includes complicated statistical calculations, which are not discussed here any further. (Also because I wasn t able to come up with a mathematically accurate explanation so far, but I tested it countless times, so I take it for granted.) Local RMSD is usually calculated for more than one atom in each residue. In this case, the local RMSD of a residue (denoted by res i ) is given as RMSD(res i ) = RMSD(r i ) which is the average of the RMSDs of the single atoms within the given residue. 1.3 Maximum distances Instead of the local RMSD, the highest distance can be given between two identical atomic positions between two different models. If more than one atom is considered in a residue, than distances between different types of atoms will not be measured and compared, only between identical atoms in different models. k
4 2. Superposition (fitting) of protein structures For models within structural ensembles, in order to reflect internal motions of proteins, it is necessary to have more or less the same orientation in space. This is most often achieved by fitting the models on each other or on a given model Least square fit The superposition of two sets of identical atomic positions can be formulated in a mathematical way as the problem to minimize the RMSD or the weighted RMSD between them. In this case, for two sets of centered atomic positions r i and r i such a rotation matrix R is needed, that minimizes RMSD: RMSD min = 1 n r i Rr i 2 n i=1 Any operation that results in an appropriate R is generally called as least square fit. In the following, the not weighted RMSD minimization is discussed Kabsch algorithm One of the many and one of the earliest algorithms for achieving a mathematically exact solution for minimum RMSD is described by Wolfgang Kabsch in Two sets of centered atomic positions is given: r i and r i. If the points are not centered, it is important before the operation to translate them in a way that their average coincides with the origin. In the following, the 3 3 rotation matrix R is needed that rotates the points of r i into r i. As the first step, N 3 matrixes P and Q are produced, that contain the coordinates of r i and r i as row vectors, respectively. Then, cross-covariance matrix A is calculated as The rotation matrix is given as x 1 y 1 z 1 x 2 y 2 z 2 P = ( ) x n y n z n A = P T Q R = (A T A) 1/2 A 1 However, A is not guaranteed to have an inverse. To account for all special cases, singular value decomposition (SVD) of matrix A is carried out. This way, the rotation matrix R is given as A = VSW T
5 1 0 0 R = W ( 0 1 0) V T 0 0 d where d = sign(det (WV T )) 2.3. Implementations of least square fit Fitting two or more protein models on each other, needless to say, requires an exact definition of the subset of atoms used for the fitting. In a structural ensemble, the models may be fitted to any of the models (e.g. the first one), or to the mean structure. The difference is significant, since least square fit guarantees minimal RMSD only if it is calculated the same way as the fitting has been carried out. That is, pairwise RMSD, for example, is not guaranteed to be minimal if all the models are fit to the first model. Similarly, if all the models are fit to the first model, the RMSD relative to the mean structure is not guaranteed to be minimal. In some cases, a rotation matrix R for minimizing weighted RMSD is searched for. The above described version of the Kabsch algorithm is not capable of yielding such rotation matrix. Instead, one should implement the algorithm described by McLachlan in Iterative fitting The mean structure of a protein ensemble clearly changes after fitting the ensemble. This also modifies the overall RMSD relative to the mean structure. To guarantee minimal RMSD, the fitting to the mean structure is carried out in an iterative way, and the mean structure is recalculated in each step of the iteration. In some other cases, it is necessary to decide which regions of the proteins should be considered in fitting. Flexible regions for example would spoil the purpose of fitting and impede obtaining a low RMSD value. To avoid such ambiguity, local RMSD may be calculated after fitting, and after deciding, which residues have too high RMSD, the may be left out from the following step of the iterative fitting. Such automatic mechanism is not implemented in PDBFitter, since decision about rather flexible than static region requires thorough consideration. 3. How other programs do it 3.1. Chimera Chimera in general has a fancy graphical user interface, and is capable of performing a large scale of operations, both from command line and on the graphical surface. However, it is often not very customizable, and does not have the advantage of easily modifying the code to obtain a special value Model-model RMSD Chimera easily yields the same fit pairwise model-model table as described in 1.1., algorithm 4. (Tools, Structure Comparison, Ensemble Match.) It also interactively rotates the models onto each other. Furthermore, it can draw an RMSD map, under the option MD movie.
6 Local RMSD Chimera can calculate local RMSD for backbone, Ca and all atoms under Tools, Sequence, and then Structure, Associations, where all the structures have to be associated to the same sequence, and then Headers. The local RMSDs are calculated in a pairwise manner, and yield the same result as in Chapter 1.2, algorithm 1. Note that Chimera includes into the backbone the following atoms: N, C, CA, O Fitting Chimera has a sophisticated fitting tool, which is capable of performing sequence and 3D structure alignment. If identical sequences are superimposed, it is also possible to use an iterative method, in which atom pairs with a distance above a given threshold can be dropped. The fitting tool is available under Tools, Structure Comparison, MatchMaker. The result of the non-iterative fitting can exactly be reproduced through the Kabsch algorithm that was implemented in PDBFitter MOLMOL The MOLMOL 2.6 was released in It has a primitive graphical interface and is capable a number of operations and calculations on proteins. However, usage and algorithms are not well documented. Exact calculations sometimes can only be traced by looking at the source code, which, in the present report, is not discussed Model-model RMSD MOLMOL yields the same model-model tables as described in Chapter 1.1., algorithms 1., 3., and 4., for backbone atoms, heavy atoms and the heavy atoms of the sidechains. The values can be reproduced exactly. It has to be noted that for MOLMOL the backbone consists of C, N and CA only. Also, it gives the variance, where PDBFitter the square of the variance.
7 Local RMSDs MOLMOL computes the local RMSDs as the average of 3 consecutive RMSDs. The exact values cannot be reproduced, for it is unclear what algorithms the program uses. However, the tendencies are the same. Also, pairwise and relative-to-mean RMSDs can be calculated, which show the same relation as described in Chapter 1.2. MOLMOL also computes two other sets of data, which are called global and local displacements. It is unclear what these values represent Fitting The results of least square fit in MOLMOL are the same as in other programs. The fit ensembles can exactly be reproduced with PDBFitter. 4. References Billeter, M., Havel, T. F., & Kuntz, I. D. (1987). A new approach to the problem of docking two molecules: the ellipsoid algorithm. Biopolymers, 26(6), doi: /bip Billeter, M., Kline, A. D., Braun, W., Huber, R., & Wüthrich, K. (1989). Comparison of the highresolution structures of the alpha-amylase inhibitor tendamistat determined by nuclear magnetic resonance in solution and by X-ray diffraction in single crystals. Journal of Molecular Biology, 206(4), Retrieved from Billeter, M., Vendrell, J., Wider, G., Avilés, F. X., Coll, M., Guasch, A., Wüthrich, K. (1992). Comparison of the NMR solution structure with the X-ray crystal structure of the activation domain from procarboxypeptidase B. J. Biomol. NMR, 2(1), doi: /bf Koradi, R., Billeter, M., & Wüthrich, K. (1996). MOLMOL: A program for display and analysis of macromolecular structures. Journal of Molecular Graphics, 14(1), doi: / (96) McLachlan, A. D. (1979). Gene duplications in the structural evolution of chymotrypsin. Journal of Molecular Biology, 128(1), doi: / (79) Kabsch, W. (1976). A solution for the best rotation to relate two sets of vectors. Acta Crystallographica Section A, 32(5), doi: /s Kabsch, W. (1978). A discussion of the solution for the best rotation to relate two sets of vectors. Acta Crystallographica Section A, 34(5), doi: /s
DATA MINING LECTURE 8. Dimensionality Reduction PCA -- SVD
 DATA MINING LECTURE 8 Dimensionality Reduction PCA -- SVD The curse of dimensionality Real data usually have thousands, or millions of dimensions E.g., web documents, where the dimensionality is the vocabulary
DATA MINING LECTURE 8 Dimensionality Reduction PCA -- SVD The curse of dimensionality Real data usually have thousands, or millions of dimensions E.g., web documents, where the dimensionality is the vocabulary
Molecular Modeling lecture 2
 Molecular Modeling 2018 -- lecture 2 Topics 1. Secondary structure 3. Sequence similarity and homology 2. Secondary structure prediction 4. Where do protein structures come from? X-ray crystallography
Molecular Modeling 2018 -- lecture 2 Topics 1. Secondary structure 3. Sequence similarity and homology 2. Secondary structure prediction 4. Where do protein structures come from? X-ray crystallography
Solving distance geometry problems for protein structure determination
 Graduate Theses and Dissertations Iowa State University Capstones, Theses and Dissertations 2010 Solving distance geometry problems for protein structure determination Atilla Sit Iowa State University
Graduate Theses and Dissertations Iowa State University Capstones, Theses and Dissertations 2010 Solving distance geometry problems for protein structure determination Atilla Sit Iowa State University
1. Protein Data Bank (PDB) 1. Protein Data Bank (PDB)
 Protein structure databases; visualization; and classifications 1. Introduction to Protein Data Bank (PDB) 2. Free graphic software for 3D structure visualization 3. Hierarchical classification of protein
Protein structure databases; visualization; and classifications 1. Introduction to Protein Data Bank (PDB) 2. Free graphic software for 3D structure visualization 3. Hierarchical classification of protein
FlexSADRA: Flexible Structural Alignment using a Dimensionality Reduction Approach
 FlexSADRA: Flexible Structural Alignment using a Dimensionality Reduction Approach Shirley Hui and Forbes J. Burkowski University of Waterloo, 200 University Avenue W., Waterloo, Canada ABSTRACT A topic
FlexSADRA: Flexible Structural Alignment using a Dimensionality Reduction Approach Shirley Hui and Forbes J. Burkowski University of Waterloo, 200 University Avenue W., Waterloo, Canada ABSTRACT A topic
I690/B680 Structural Bioinformatics Spring Protein Structure Determination by NMR Spectroscopy
 I690/B680 Structural Bioinformatics Spring 2006 Protein Structure Determination by NMR Spectroscopy Suggested Reading (1) Van Holde, Johnson, Ho. Principles of Physical Biochemistry, 2 nd Ed., Prentice
I690/B680 Structural Bioinformatics Spring 2006 Protein Structure Determination by NMR Spectroscopy Suggested Reading (1) Van Holde, Johnson, Ho. Principles of Physical Biochemistry, 2 nd Ed., Prentice
NMR, X-ray Diffraction, Protein Structure, and RasMol
 NMR, X-ray Diffraction, Protein Structure, and RasMol Introduction So far we have been mostly concerned with the proteins themselves. The techniques (NMR or X-ray diffraction) used to determine a structure
NMR, X-ray Diffraction, Protein Structure, and RasMol Introduction So far we have been mostly concerned with the proteins themselves. The techniques (NMR or X-ray diffraction) used to determine a structure
Basics of protein structure
 Today: 1. Projects a. Requirements: i. Critical review of one paper ii. At least one computational result b. Noon, Dec. 3 rd written report and oral presentation are due; submit via email to bphys101@fas.harvard.edu
Today: 1. Projects a. Requirements: i. Critical review of one paper ii. At least one computational result b. Noon, Dec. 3 rd written report and oral presentation are due; submit via email to bphys101@fas.harvard.edu
Protein structures and comparisons ndrew Torda Bioinformatik, Mai 2008
 Protein structures and comparisons ndrew Torda 67.937 Bioinformatik, Mai 2008 Ultimate aim how to find out the most about a protein what you can get from sequence and structure information On the way..
Protein structures and comparisons ndrew Torda 67.937 Bioinformatik, Mai 2008 Ultimate aim how to find out the most about a protein what you can get from sequence and structure information On the way..
Experimental Techniques in Protein Structure Determination
 Experimental Techniques in Protein Structure Determination Homayoun Valafar Department of Computer Science and Engineering, USC Two Main Experimental Methods X-Ray crystallography Nuclear Magnetic Resonance
Experimental Techniques in Protein Structure Determination Homayoun Valafar Department of Computer Science and Engineering, USC Two Main Experimental Methods X-Ray crystallography Nuclear Magnetic Resonance
Automated Assignment of Backbone NMR Data using Artificial Intelligence
 Automated Assignment of Backbone NMR Data using Artificial Intelligence John Emmons στ, Steven Johnson τ, Timothy Urness*, and Adina Kilpatrick* Department of Computer Science and Mathematics Department
Automated Assignment of Backbone NMR Data using Artificial Intelligence John Emmons στ, Steven Johnson τ, Timothy Urness*, and Adina Kilpatrick* Department of Computer Science and Mathematics Department
Visualization of Macromolecular Structures
 Visualization of Macromolecular Structures Present by: Qihang Li orig. author: O Donoghue, et al. Structural biology is rapidly accumulating a wealth of detailed information. Over 60,000 high-resolution
Visualization of Macromolecular Structures Present by: Qihang Li orig. author: O Donoghue, et al. Structural biology is rapidly accumulating a wealth of detailed information. Over 60,000 high-resolution
Copyright Mark Brandt, Ph.D A third method, cryogenic electron microscopy has seen increasing use over the past few years.
 Structure Determination and Sequence Analysis The vast majority of the experimentally determined three-dimensional protein structures have been solved by one of two methods: X-ray diffraction and Nuclear
Structure Determination and Sequence Analysis The vast majority of the experimentally determined three-dimensional protein structures have been solved by one of two methods: X-ray diffraction and Nuclear
Dimension Reduction and Iterative Consensus Clustering
 Dimension Reduction and Iterative Consensus Clustering Southeastern Clustering and Ranking Workshop August 24, 2009 Dimension Reduction and Iterative 1 Document Clustering Geometry of the SVD Centered
Dimension Reduction and Iterative Consensus Clustering Southeastern Clustering and Ranking Workshop August 24, 2009 Dimension Reduction and Iterative 1 Document Clustering Geometry of the SVD Centered
Francisco Melo, Damien Devos, Eric Depiereux and Ernest Feytmans
 From: ISMB-97 Proceedings. Copyright 1997, AAAI (www.aaai.org). All rights reserved. ANOLEA: A www Server to Assess Protein Structures Francisco Melo, Damien Devos, Eric Depiereux and Ernest Feytmans Facultés
From: ISMB-97 Proceedings. Copyright 1997, AAAI (www.aaai.org). All rights reserved. ANOLEA: A www Server to Assess Protein Structures Francisco Melo, Damien Devos, Eric Depiereux and Ernest Feytmans Facultés
Examples of Protein Modeling. Protein Modeling. Primary Structure. Protein Structure Description. Protein Sequence Sources. Importing Sequences to MOE
 Examples of Protein Modeling Protein Modeling Visualization Examination of an experimental structure to gain insight about a research question Dynamics To examine the dynamics of protein structures To
Examples of Protein Modeling Protein Modeling Visualization Examination of an experimental structure to gain insight about a research question Dynamics To examine the dynamics of protein structures To
Fondamenti di Chimica Farmaceutica. Computer Chemistry in Drug Research: Introduction
 Fondamenti di Chimica Farmaceutica Computer Chemistry in Drug Research: Introduction Introduction Introduction Introduction Computer Chemistry in Drug Design Drug Discovery: Target identification Lead
Fondamenti di Chimica Farmaceutica Computer Chemistry in Drug Research: Introduction Introduction Introduction Introduction Computer Chemistry in Drug Design Drug Discovery: Target identification Lead
Dipolar Couplings in Partially Aligned Macromolecules - New Directions in. Structure Determination using Solution State NMR.
 Dipolar Couplings in Partially Aligned Macromolecules - New Directions in Structure Determination using Solution State NMR. Recently developed methods for the partial alignment of macromolecules in dilute
Dipolar Couplings in Partially Aligned Macromolecules - New Directions in Structure Determination using Solution State NMR. Recently developed methods for the partial alignment of macromolecules in dilute
CMPS 6630: Introduction to Computational Biology and Bioinformatics. Structure Comparison
 CMPS 6630: Introduction to Computational Biology and Bioinformatics Structure Comparison Protein Structure Comparison Motivation Understand sequence and structure variability Understand Domain architecture
CMPS 6630: Introduction to Computational Biology and Bioinformatics Structure Comparison Protein Structure Comparison Motivation Understand sequence and structure variability Understand Domain architecture
Selecting protein fuzzy contact maps through information and structure measures
 Selecting protein fuzzy contact maps through information and structure measures Carlos Bousoño-Calzón Signal Processing and Communication Dpt. Univ. Carlos III de Madrid Avda. de la Universidad, 30 28911
Selecting protein fuzzy contact maps through information and structure measures Carlos Bousoño-Calzón Signal Processing and Communication Dpt. Univ. Carlos III de Madrid Avda. de la Universidad, 30 28911
CSE 554 Lecture 7: Alignment
 CSE 554 Lecture 7: Alignment Fall 2012 CSE554 Alignment Slide 1 Review Fairing (smoothing) Relocating vertices to achieve a smoother appearance Method: centroid averaging Simplification Reducing vertex
CSE 554 Lecture 7: Alignment Fall 2012 CSE554 Alignment Slide 1 Review Fairing (smoothing) Relocating vertices to achieve a smoother appearance Method: centroid averaging Simplification Reducing vertex
Singular Value Decomposition and Principal Component Analysis (PCA) I
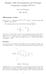 Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Analysis of MD trajectories in GROMACS David van der Spoel
 Analysis of MD trajectories in GROMACS David van der Spoel What does MD produce? Energy terms E(t) Coordinates x(t) Velocities v(t) Forces f(t) Managing your files trjcat - merging trajectories concatenating
Analysis of MD trajectories in GROMACS David van der Spoel What does MD produce? Energy terms E(t) Coordinates x(t) Velocities v(t) Forces f(t) Managing your files trjcat - merging trajectories concatenating
Analysis and Prediction of Protein Structure (I)
 Analysis and Prediction of Protein Structure (I) Jianlin Cheng, PhD School of Electrical Engineering and Computer Science University of Central Florida 2006 Free for academic use. Copyright @ Jianlin Cheng
Analysis and Prediction of Protein Structure (I) Jianlin Cheng, PhD School of Electrical Engineering and Computer Science University of Central Florida 2006 Free for academic use. Copyright @ Jianlin Cheng
Automated Assignment of Simulated and Experimental NOESY Spectra of Proteins by Feedback Filtering and Self-correcting Distance Geometry
 J. Mol. Biol. (1995) 254, 465 480 Automated Assignment of Simulated and Experimental NOESY Spectra of Proteins by Feedback Filtering and Self-correcting Distance Geometry Ch. Mumenthaler 1 and W. Braun
J. Mol. Biol. (1995) 254, 465 480 Automated Assignment of Simulated and Experimental NOESY Spectra of Proteins by Feedback Filtering and Self-correcting Distance Geometry Ch. Mumenthaler 1 and W. Braun
Pairwise & Multiple sequence alignments
 Pairwise & Multiple sequence alignments Urmila Kulkarni-Kale Bioinformatics Centre 411 007 urmila@bioinfo.ernet.in Basis for Sequence comparison Theory of evolution: gene sequences have evolved/derived
Pairwise & Multiple sequence alignments Urmila Kulkarni-Kale Bioinformatics Centre 411 007 urmila@bioinfo.ernet.in Basis for Sequence comparison Theory of evolution: gene sequences have evolved/derived
All measurement has a limit of precision and accuracy, and this must be taken into account when evaluating experimental results.
 Chapter 11: Measurement and data processing and analysis 11.1 Uncertainty and error in measurement and results All measurement has a limit of precision and accuracy, and this must be taken into account
Chapter 11: Measurement and data processing and analysis 11.1 Uncertainty and error in measurement and results All measurement has a limit of precision and accuracy, and this must be taken into account
Anisotropy in macromolecular crystal structures. Andrea Thorn July 19 th, 2012
 Anisotropy in macromolecular crystal structures Andrea Thorn July 19 th, 2012 Motivation Courtesy of M. Sawaya Motivation Crystal structures are inherently anisotropic. X-ray diffraction reflects this
Anisotropy in macromolecular crystal structures Andrea Thorn July 19 th, 2012 Motivation Courtesy of M. Sawaya Motivation Crystal structures are inherently anisotropic. X-ray diffraction reflects this
Overview & Applications. T. Lezon Hands-on Workshop in Computational Biophysics Pittsburgh Supercomputing Center 04 June, 2015
 Overview & Applications T. Lezon Hands-on Workshop in Computational Biophysics Pittsburgh Supercomputing Center 4 June, 215 Simulations still take time Bakan et al. Bioinformatics 211. Coarse-grained Elastic
Overview & Applications T. Lezon Hands-on Workshop in Computational Biophysics Pittsburgh Supercomputing Center 4 June, 215 Simulations still take time Bakan et al. Bioinformatics 211. Coarse-grained Elastic
Statistical Machine Learning Methods for Bioinformatics IV. Neural Network & Deep Learning Applications in Bioinformatics
 Statistical Machine Learning Methods for Bioinformatics IV. Neural Network & Deep Learning Applications in Bioinformatics Jianlin Cheng, PhD Department of Computer Science University of Missouri, Columbia
Statistical Machine Learning Methods for Bioinformatics IV. Neural Network & Deep Learning Applications in Bioinformatics Jianlin Cheng, PhD Department of Computer Science University of Missouri, Columbia
Determining Protein Structure BIBC 100
 Determining Protein Structure BIBC 100 Determining Protein Structure X-Ray Diffraction Interactions of x-rays with electrons in molecules in a crystal NMR- Nuclear Magnetic Resonance Interactions of magnetic
Determining Protein Structure BIBC 100 Determining Protein Structure X-Ray Diffraction Interactions of x-rays with electrons in molecules in a crystal NMR- Nuclear Magnetic Resonance Interactions of magnetic
Computational Modeling of Protein Kinase A and Comparison with Nuclear Magnetic Resonance Data
 Computational Modeling of Protein Kinase A and Comparison with Nuclear Magnetic Resonance Data ABSTRACT Keyword Lei Shi 1 Advisor: Gianluigi Veglia 1,2 Department of Chemistry 1, & Biochemistry, Molecular
Computational Modeling of Protein Kinase A and Comparison with Nuclear Magnetic Resonance Data ABSTRACT Keyword Lei Shi 1 Advisor: Gianluigi Veglia 1,2 Department of Chemistry 1, & Biochemistry, Molecular
Calculation and Application of MOPITT Averaging Kernels
 Calculation and Application of MOPITT Averaging Kernels Merritt N. Deeter Atmospheric Chemistry Division National Center for Atmospheric Research Boulder, Colorado 80307 July, 2002 I. Introduction Retrieval
Calculation and Application of MOPITT Averaging Kernels Merritt N. Deeter Atmospheric Chemistry Division National Center for Atmospheric Research Boulder, Colorado 80307 July, 2002 I. Introduction Retrieval
CSE 554 Lecture 6: Deformation I
 CSE 554 Lecture 6: Deformation I Fall 20 CSE554 Deformation I Slide Review Alignment Registering source to target by rotation and translation Methods Rigid-body transformations Aligning principle directions
CSE 554 Lecture 6: Deformation I Fall 20 CSE554 Deformation I Slide Review Alignment Registering source to target by rotation and translation Methods Rigid-body transformations Aligning principle directions
Analysis of protein dynamics using local-dme calculations
 Biochemistry, Biophysics and Molecular Biology Publications Biochemistry, Biophysics and Molecular Biology 2011 Analysis of protein dynamics using local-dme calculations Di Wu Western Kentucky University
Biochemistry, Biophysics and Molecular Biology Publications Biochemistry, Biophysics and Molecular Biology 2011 Analysis of protein dynamics using local-dme calculations Di Wu Western Kentucky University
Computer simulation methods (2) Dr. Vania Calandrini
 Computer simulation methods (2) Dr. Vania Calandrini in the previous lecture: time average versus ensemble average MC versus MD simulations equipartition theorem (=> computing T) virial theorem (=> computing
Computer simulation methods (2) Dr. Vania Calandrini in the previous lecture: time average versus ensemble average MC versus MD simulations equipartition theorem (=> computing T) virial theorem (=> computing
DOCKING TUTORIAL. A. The docking Workflow
 2 nd Strasbourg Summer School on Chemoinformatics VVF Obernai, France, 20-24 June 2010 E. Kellenberger DOCKING TUTORIAL A. The docking Workflow 1. Ligand preparation It consists in the standardization
2 nd Strasbourg Summer School on Chemoinformatics VVF Obernai, France, 20-24 June 2010 E. Kellenberger DOCKING TUTORIAL A. The docking Workflow 1. Ligand preparation It consists in the standardization
Algorithms in Bioinformatics
 Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri RNA Structure Prediction Secondary
Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri RNA Structure Prediction Secondary
Everyday NMR. Innovation with Integrity. Why infer when you can be sure? NMR
 Everyday NMR Why infer when you can be sure? Innovation with Integrity NMR Only NMR gives you definitive answers, on your terms. Over the past half-century, scientists have used nuclear magnetic resonance
Everyday NMR Why infer when you can be sure? Innovation with Integrity NMR Only NMR gives you definitive answers, on your terms. Over the past half-century, scientists have used nuclear magnetic resonance
tconcoord-gui: Visually Supported Conformational Sampling of Bioactive Molecules
 Software News and Updates tconcoord-gui: Visually Supported Conformational Sampling of Bioactive Molecules DANIEL SEELIGER, BERT L. DE GROOT Computational Biomolecular Dynamics Group, Max-Planck-Institute
Software News and Updates tconcoord-gui: Visually Supported Conformational Sampling of Bioactive Molecules DANIEL SEELIGER, BERT L. DE GROOT Computational Biomolecular Dynamics Group, Max-Planck-Institute
Accelerating Biomolecular Nuclear Magnetic Resonance Assignment with A*
 Accelerating Biomolecular Nuclear Magnetic Resonance Assignment with A* Joel Venzke, Paxten Johnson, Rachel Davis, John Emmons, Katherine Roth, David Mascharka, Leah Robison, Timothy Urness and Adina Kilpatrick
Accelerating Biomolecular Nuclear Magnetic Resonance Assignment with A* Joel Venzke, Paxten Johnson, Rachel Davis, John Emmons, Katherine Roth, David Mascharka, Leah Robison, Timothy Urness and Adina Kilpatrick
Homologous proteins have similar structures and structural superposition means to rotate and translate the structures so that corresponding atoms are
 1 Homologous proteins have similar structures and structural superposition means to rotate and translate the structures so that corresponding atoms are as close to each other as possible. Structural similarity
1 Homologous proteins have similar structures and structural superposition means to rotate and translate the structures so that corresponding atoms are as close to each other as possible. Structural similarity
PROTEIN-PROTEIN DOCKING REFINEMENT USING RESTRAINT MOLECULAR DYNAMICS SIMULATIONS
 TASKQUARTERLYvol.20,No4,2016,pp.353 360 PROTEIN-PROTEIN DOCKING REFINEMENT USING RESTRAINT MOLECULAR DYNAMICS SIMULATIONS MARTIN ZACHARIAS Physics Department T38, Technical University of Munich James-Franck-Str.
TASKQUARTERLYvol.20,No4,2016,pp.353 360 PROTEIN-PROTEIN DOCKING REFINEMENT USING RESTRAINT MOLECULAR DYNAMICS SIMULATIONS MARTIN ZACHARIAS Physics Department T38, Technical University of Munich James-Franck-Str.
Distance Geometry-Matrix Completion
 Christos Konaxis Algs in Struct BioInfo 2010 Outline Outline Tertiary structure Incomplete data Tertiary structure Measure diffrence of matched sets Def. Root Mean Square Deviation RMSD = 1 n x i y i 2,
Christos Konaxis Algs in Struct BioInfo 2010 Outline Outline Tertiary structure Incomplete data Tertiary structure Measure diffrence of matched sets Def. Root Mean Square Deviation RMSD = 1 n x i y i 2,
Macromolecular X-ray Crystallography
 Protein Structural Models for CHEM 641 Fall 07 Brian Bahnson Department of Chemistry & Biochemistry University of Delaware Macromolecular X-ray Crystallography Purified Protein X-ray Diffraction Data collection
Protein Structural Models for CHEM 641 Fall 07 Brian Bahnson Department of Chemistry & Biochemistry University of Delaware Macromolecular X-ray Crystallography Purified Protein X-ray Diffraction Data collection
Analysis of the simulation
 Analysis of the simulation Marcus Elstner and Tomáš Kubař January 7, 2014 Thermodynamic properties time averages of thermodynamic quantites correspond to ensemble averages (ergodic theorem) some quantities
Analysis of the simulation Marcus Elstner and Tomáš Kubař January 7, 2014 Thermodynamic properties time averages of thermodynamic quantites correspond to ensemble averages (ergodic theorem) some quantities
Conceptual Questions for Review
 Conceptual Questions for Review Chapter 1 1.1 Which vectors are linear combinations of v = (3, 1) and w = (4, 3)? 1.2 Compare the dot product of v = (3, 1) and w = (4, 3) to the product of their lengths.
Conceptual Questions for Review Chapter 1 1.1 Which vectors are linear combinations of v = (3, 1) and w = (4, 3)? 1.2 Compare the dot product of v = (3, 1) and w = (4, 3) to the product of their lengths.
CS47300: Web Information Search and Management
 CS47300: Web Information Search and Management Prof. Chris Clifton 6 September 2017 Material adapted from course created by Dr. Luo Si, now leading Alibaba research group 1 Vector Space Model Disadvantages:
CS47300: Web Information Search and Management Prof. Chris Clifton 6 September 2017 Material adapted from course created by Dr. Luo Si, now leading Alibaba research group 1 Vector Space Model Disadvantages:
Relative binding enthalpies from molecular dynamics simulations. using a direct method
 Relative binding enthalpies from molecular dynamics simulations using a direct method Amitava Roy, 1 Duy P. Hua, 1 Joshua M. Ward, 1 and Carol Beth Post* Department of Medicinal Chemistry, Markey Center
Relative binding enthalpies from molecular dynamics simulations using a direct method Amitava Roy, 1 Duy P. Hua, 1 Joshua M. Ward, 1 and Carol Beth Post* Department of Medicinal Chemistry, Markey Center
User Guide for LeDock
 User Guide for LeDock Hongtao Zhao, PhD Email: htzhao@lephar.com Website: www.lephar.com Copyright 2017 Hongtao Zhao. All rights reserved. Introduction LeDock is flexible small-molecule docking software,
User Guide for LeDock Hongtao Zhao, PhD Email: htzhao@lephar.com Website: www.lephar.com Copyright 2017 Hongtao Zhao. All rights reserved. Introduction LeDock is flexible small-molecule docking software,
HTCondor and macromolecular structure validation
 HTCondor and macromolecular structure validation Vincent Chen John Markley/Eldon Ulrich, NMRFAM/BMRB, UW@Madison David & Jane Richardson, Duke University Macromolecules David S. Goodsell 1999 Two questions
HTCondor and macromolecular structure validation Vincent Chen John Markley/Eldon Ulrich, NMRFAM/BMRB, UW@Madison David & Jane Richardson, Duke University Macromolecules David S. Goodsell 1999 Two questions
StructuralBiology. October 18, 2018
 StructuralBiology October 18, 2018 1 Lecture 18: Computational Structural Biology CBIO (CSCI) 4835/6835: Introduction to Computational Biology 1.1 Overview and Objectives Even though we re officially moving
StructuralBiology October 18, 2018 1 Lecture 18: Computational Structural Biology CBIO (CSCI) 4835/6835: Introduction to Computational Biology 1.1 Overview and Objectives Even though we re officially moving
Docking. GBCB 5874: Problem Solving in GBCB
 Docking Benzamidine Docking to Trypsin Relationship to Drug Design Ligand-based design QSAR Pharmacophore modeling Can be done without 3-D structure of protein Receptor/Structure-based design Molecular
Docking Benzamidine Docking to Trypsin Relationship to Drug Design Ligand-based design QSAR Pharmacophore modeling Can be done without 3-D structure of protein Receptor/Structure-based design Molecular
DISCRETE TUTORIAL. Agustí Emperador. Institute for Research in Biomedicine, Barcelona APPLICATION OF DISCRETE TO FLEXIBLE PROTEIN-PROTEIN DOCKING:
 DISCRETE TUTORIAL Agustí Emperador Institute for Research in Biomedicine, Barcelona APPLICATION OF DISCRETE TO FLEXIBLE PROTEIN-PROTEIN DOCKING: STRUCTURAL REFINEMENT OF DOCKING CONFORMATIONS Emperador
DISCRETE TUTORIAL Agustí Emperador Institute for Research in Biomedicine, Barcelona APPLICATION OF DISCRETE TO FLEXIBLE PROTEIN-PROTEIN DOCKING: STRUCTURAL REFINEMENT OF DOCKING CONFORMATIONS Emperador
CMPS 6630: Introduction to Computational Biology and Bioinformatics. Tertiary Structure Prediction
 CMPS 6630: Introduction to Computational Biology and Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the
CMPS 6630: Introduction to Computational Biology and Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the
NMR Spectroscopy of Polymers
 UNESCO/IUPAC Course 2005/2006 Jiri Brus NMR Spectroscopy of Polymers Brus J 1. part At the very beginning the phenomenon of nuclear spin resonance was studied predominantly by physicists and the application
UNESCO/IUPAC Course 2005/2006 Jiri Brus NMR Spectroscopy of Polymers Brus J 1. part At the very beginning the phenomenon of nuclear spin resonance was studied predominantly by physicists and the application
Karhunen-Loève Transform KLT. JanKees van der Poel D.Sc. Student, Mechanical Engineering
 Karhunen-Loève Transform KLT JanKees van der Poel D.Sc. Student, Mechanical Engineering Karhunen-Loève Transform Has many names cited in literature: Karhunen-Loève Transform (KLT); Karhunen-Loève Decomposition
Karhunen-Loève Transform KLT JanKees van der Poel D.Sc. Student, Mechanical Engineering Karhunen-Loève Transform Has many names cited in literature: Karhunen-Loève Transform (KLT); Karhunen-Loève Decomposition
Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability
 Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability Dr. Andrew Lee UNC School of Pharmacy (Div. Chemical Biology and Medicinal Chemistry) UNC Med
Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability Dr. Andrew Lee UNC School of Pharmacy (Div. Chemical Biology and Medicinal Chemistry) UNC Med
Introducing Hippy: A visualization tool for understanding the α-helix pair interface
 Introducing Hippy: A visualization tool for understanding the α-helix pair interface Robert Fraser and Janice Glasgow School of Computing, Queen s University, Kingston ON, Canada, K7L3N6 {robert,janice}@cs.queensu.ca
Introducing Hippy: A visualization tool for understanding the α-helix pair interface Robert Fraser and Janice Glasgow School of Computing, Queen s University, Kingston ON, Canada, K7L3N6 {robert,janice}@cs.queensu.ca
Better Bond Angles in the Protein Data Bank
 Better Bond Angles in the Protein Data Bank C.J. Robinson and D.B. Skillicorn School of Computing Queen s University {robinson,skill}@cs.queensu.ca Abstract The Protein Data Bank (PDB) contains, at least
Better Bond Angles in the Protein Data Bank C.J. Robinson and D.B. Skillicorn School of Computing Queen s University {robinson,skill}@cs.queensu.ca Abstract The Protein Data Bank (PDB) contains, at least
Pose and affinity prediction by ICM in D3R GC3. Max Totrov Molsoft
 Pose and affinity prediction by ICM in D3R GC3 Max Totrov Molsoft Pose prediction method: ICM-dock ICM-dock: - pre-sampling of ligand conformers - multiple trajectory Monte-Carlo with gradient minimization
Pose and affinity prediction by ICM in D3R GC3 Max Totrov Molsoft Pose prediction method: ICM-dock ICM-dock: - pre-sampling of ligand conformers - multiple trajectory Monte-Carlo with gradient minimization
T 1, T 2, NOE (reminder)
 T 1, T 2, NOE (reminder) T 1 is the time constant for longitudinal relaxation - the process of re-establishing the Boltzmann distribution of the energy level populations of the system following perturbation
T 1, T 2, NOE (reminder) T 1 is the time constant for longitudinal relaxation - the process of re-establishing the Boltzmann distribution of the energy level populations of the system following perturbation
Multivariate T-Squared Control Chart
 Multivariate T-Squared Control Chart Summary... 1 Data Input... 3 Analysis Summary... 4 Analysis Options... 5 T-Squared Chart... 6 Multivariate Control Chart Report... 7 Generalized Variance Chart... 8
Multivariate T-Squared Control Chart Summary... 1 Data Input... 3 Analysis Summary... 4 Analysis Options... 5 T-Squared Chart... 6 Multivariate Control Chart Report... 7 Generalized Variance Chart... 8
Dimension Reduc-on. Example: height of iden-cal twins. PCA, SVD, MDS, and clustering [ RI ] Twin 2 (inches away from avg)
![Dimension Reduc-on. Example: height of iden-cal twins. PCA, SVD, MDS, and clustering [ RI ] Twin 2 (inches away from avg) Dimension Reduc-on. Example: height of iden-cal twins. PCA, SVD, MDS, and clustering [ RI ] Twin 2 (inches away from avg)](/thumbs/74/70760823.jpg) Dimension Reduc-on PCA, SVD, MDS, and clustering Example: height of iden-cal twins Twin (inches away from avg) 0 5 0 5 0 5 0 5 0 Twin (inches away from avg) Expression between two ethnic groups Frequency
Dimension Reduc-on PCA, SVD, MDS, and clustering Example: height of iden-cal twins Twin (inches away from avg) 0 5 0 5 0 5 0 5 0 Twin (inches away from avg) Expression between two ethnic groups Frequency
Lecture Outline. Biost 518 Applied Biostatistics II. Choice of Model for Analysis. Choice of Model. Choice of Model. Lecture 10: Multiple Regression:
 Biost 518 Applied Biostatistics II Scott S. Emerson, M.D., Ph.D. Professor of Biostatistics University of Washington Lecture utline Choice of Model Alternative Models Effect of data driven selection of
Biost 518 Applied Biostatistics II Scott S. Emerson, M.D., Ph.D. Professor of Biostatistics University of Washington Lecture utline Choice of Model Alternative Models Effect of data driven selection of
Life Science Webinar Series
 Life Science Webinar Series Elegant protein- protein docking in Discovery Studio Francisco Hernandez-Guzman, Ph.D. November 20, 2007 Sr. Solutions Scientist fhernandez@accelrys.com Agenda In silico protein-protein
Life Science Webinar Series Elegant protein- protein docking in Discovery Studio Francisco Hernandez-Guzman, Ph.D. November 20, 2007 Sr. Solutions Scientist fhernandez@accelrys.com Agenda In silico protein-protein
NMR in Medicine and Biology
 NMR in Medicine and Biology http://en.wikipedia.org/wiki/nmr_spectroscopy MRI- Magnetic Resonance Imaging (water) In-vivo spectroscopy (metabolites) Solid-state t NMR (large structures) t Solution NMR
NMR in Medicine and Biology http://en.wikipedia.org/wiki/nmr_spectroscopy MRI- Magnetic Resonance Imaging (water) In-vivo spectroscopy (metabolites) Solid-state t NMR (large structures) t Solution NMR
An engineered scorpion toxin analogue with improved Kv1.3 selectivity displays reduced conformational flexibility
 An engineered scorpion toxin analogue with improved Kv1.3 selectivity displays reduced conformational flexibility Adam Bartok a,#, Krisztina Fehér b,c,#,, Andrea Bodor d, Kinga Rákosi e, Gábor K. Tóth
An engineered scorpion toxin analogue with improved Kv1.3 selectivity displays reduced conformational flexibility Adam Bartok a,#, Krisztina Fehér b,c,#,, Andrea Bodor d, Kinga Rákosi e, Gábor K. Tóth
Pymol Practial Guide
 Pymol Practial Guide Pymol is a powerful visualizor very convenient to work with protein molecules. Its interface may seem complex at first, but you will see that with a little practice is simple and powerful
Pymol Practial Guide Pymol is a powerful visualizor very convenient to work with protein molecules. Its interface may seem complex at first, but you will see that with a little practice is simple and powerful
Principles of Physical Biochemistry
 Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Assessing the potential of atomistic molecular dynamics simulations to probe reversible. protein-protein recognition and binding
 Supporting information for Assessing the potential of atomistic molecular dynamics simulations to probe reversible protein-protein recognition and binding Luciano A. Abriata and Matteo Dal Peraro Laboratory
Supporting information for Assessing the potential of atomistic molecular dynamics simulations to probe reversible protein-protein recognition and binding Luciano A. Abriata and Matteo Dal Peraro Laboratory
Protein Structure Determination
 Protein Structure Determination Given a protein sequence, determine its 3D structure 1 MIKLGIVMDP IANINIKKDS SFAMLLEAQR RGYELHYMEM GDLYLINGEA 51 RAHTRTLNVK QNYEEWFSFV GEQDLPLADL DVILMRKDPP FDTEFIYATY 101
Protein Structure Determination Given a protein sequence, determine its 3D structure 1 MIKLGIVMDP IANINIKKDS SFAMLLEAQR RGYELHYMEM GDLYLINGEA 51 RAHTRTLNVK QNYEEWFSFV GEQDLPLADL DVILMRKDPP FDTEFIYATY 101
Data envelopment analysis
 15 Data envelopment analysis The purpose of data envelopment analysis (DEA) is to compare the operating performance of a set of units such as companies, university departments, hospitals, bank branch offices,
15 Data envelopment analysis The purpose of data envelopment analysis (DEA) is to compare the operating performance of a set of units such as companies, university departments, hospitals, bank branch offices,
Prediction and refinement of NMR structures from sparse experimental data
 Prediction and refinement of NMR structures from sparse experimental data Jeff Skolnick Director Center for the Study of Systems Biology School of Biology Georgia Institute of Technology Overview of talk
Prediction and refinement of NMR structures from sparse experimental data Jeff Skolnick Director Center for the Study of Systems Biology School of Biology Georgia Institute of Technology Overview of talk
Visualizing and Discriminating Atom Intersections Within the Spiegel Visualization Framework
 Visualizing and Discriminating Atom Intersections Within the Spiegel Visualization Framework Ian McIntosh May 19, 2006 Contents Abstract. 3 Overview 1.1 Spiegel. 3 1.2 Atom Intersections... 3 Intersections
Visualizing and Discriminating Atom Intersections Within the Spiegel Visualization Framework Ian McIntosh May 19, 2006 Contents Abstract. 3 Overview 1.1 Spiegel. 3 1.2 Atom Intersections... 3 Intersections
Protein Structure Determination Using NMR Restraints BCMB/CHEM 8190
 Protein Structure Determination Using NMR Restraints BCMB/CHEM 8190 Programs for NMR Based Structure Determination CNS - Brünger, A. T.; Adams, P. D.; Clore, G. M.; DeLano, W. L.; Gros, P.; Grosse-Kunstleve,
Protein Structure Determination Using NMR Restraints BCMB/CHEM 8190 Programs for NMR Based Structure Determination CNS - Brünger, A. T.; Adams, P. D.; Clore, G. M.; DeLano, W. L.; Gros, P.; Grosse-Kunstleve,
Timescales of Protein Dynamics
 Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Summary of 1D Experiment time domain data Fourier Transform (FT) frequency domain data or Transverse Relaxation Ensemble of Nuclear
Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Summary of 1D Experiment time domain data Fourier Transform (FT) frequency domain data or Transverse Relaxation Ensemble of Nuclear
Lecture: Face Recognition and Feature Reduction
 Lecture: Face Recognition and Feature Reduction Juan Carlos Niebles and Ranjay Krishna Stanford Vision and Learning Lab 1 Recap - Curse of dimensionality Assume 5000 points uniformly distributed in the
Lecture: Face Recognition and Feature Reduction Juan Carlos Niebles and Ranjay Krishna Stanford Vision and Learning Lab 1 Recap - Curse of dimensionality Assume 5000 points uniformly distributed in the
PCA, Kernel PCA, ICA
 PCA, Kernel PCA, ICA Learning Representations. Dimensionality Reduction. Maria-Florina Balcan 04/08/2015 Big & High-Dimensional Data High-Dimensions = Lot of Features Document classification Features per
PCA, Kernel PCA, ICA Learning Representations. Dimensionality Reduction. Maria-Florina Balcan 04/08/2015 Big & High-Dimensional Data High-Dimensions = Lot of Features Document classification Features per
CS273: Algorithms for Structure Handout # 2 and Motion in Biology Stanford University Thursday, 1 April 2004
 CS273: Algorithms for Structure Handout # 2 and Motion in Biology Stanford University Thursday, 1 April 2004 Lecture #2: 1 April 2004 Topics: Kinematics : Concepts and Results Kinematics of Ligands and
CS273: Algorithms for Structure Handout # 2 and Motion in Biology Stanford University Thursday, 1 April 2004 Lecture #2: 1 April 2004 Topics: Kinematics : Concepts and Results Kinematics of Ligands and
Autodock tutorial VINA with UCSF Chimera
 Autodock tutorial VINA with UCSF Chimera José R. Valverde CNB/CSIC jrvalverde@cnb.csic.es José R. Valverde, 2014 CC-BY-NC-SA Loading the receptor Open UCSF Chimera and then load the protein: File Open
Autodock tutorial VINA with UCSF Chimera José R. Valverde CNB/CSIC jrvalverde@cnb.csic.es José R. Valverde, 2014 CC-BY-NC-SA Loading the receptor Open UCSF Chimera and then load the protein: File Open
Summary of Experimental Protein Structure Determination. Key Elements
 Programme 8.00-8.20 Summary of last week s lecture and quiz 8.20-9.00 Structure validation 9.00-9.15 Break 9.15-11.00 Exercise: Structure validation tutorial 11.00-11.10 Break 11.10-11.40 Summary & discussion
Programme 8.00-8.20 Summary of last week s lecture and quiz 8.20-9.00 Structure validation 9.00-9.15 Break 9.15-11.00 Exercise: Structure validation tutorial 11.00-11.10 Break 11.10-11.40 Summary & discussion
Focus was on solving matrix inversion problems Now we look at other properties of matrices Useful when A represents a transformations.
 Previously Focus was on solving matrix inversion problems Now we look at other properties of matrices Useful when A represents a transformations y = Ax Or A simply represents data Notion of eigenvectors,
Previously Focus was on solving matrix inversion problems Now we look at other properties of matrices Useful when A represents a transformations y = Ax Or A simply represents data Notion of eigenvectors,
Unit 2, Section 3: Linear Combinations, Spanning, and Linear Independence Linear Combinations, Spanning, and Linear Independence
 Linear Combinations Spanning and Linear Independence We have seen that there are two operations defined on a given vector space V :. vector addition of two vectors and. scalar multiplication of a vector
Linear Combinations Spanning and Linear Independence We have seen that there are two operations defined on a given vector space V :. vector addition of two vectors and. scalar multiplication of a vector
Using a Hopfield Network: A Nuts and Bolts Approach
 Using a Hopfield Network: A Nuts and Bolts Approach November 4, 2013 Gershon Wolfe, Ph.D. Hopfield Model as Applied to Classification Hopfield network Training the network Updating nodes Sequencing of
Using a Hopfield Network: A Nuts and Bolts Approach November 4, 2013 Gershon Wolfe, Ph.D. Hopfield Model as Applied to Classification Hopfield network Training the network Updating nodes Sequencing of
Integer and Constraint Programming for Batch Annealing Process Planning
 Integer and Constraint Programming for Batch Annealing Process Planning Willem-Jan van Hoeve and Sridhar Tayur Tepper School of Business, Carnegie Mellon University 5000 Forbes Avenue, Pittsburgh, PA 15213,
Integer and Constraint Programming for Batch Annealing Process Planning Willem-Jan van Hoeve and Sridhar Tayur Tepper School of Business, Carnegie Mellon University 5000 Forbes Avenue, Pittsburgh, PA 15213,
CMPS 3110: Bioinformatics. Tertiary Structure Prediction
 CMPS 3110: Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the laws of physics! Conformation space is finite
CMPS 3110: Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the laws of physics! Conformation space is finite
NMR Resonance Assignment Assisted by Mass Spectrometry
 NMR Resonance Assignment Assisted by Mass Spectrometry This lecture talked about a NMR resonance assignment assisted by mass spectrometry [1, 2]. 1 Motivation Nuclear magnetic resonance (NMR) provides
NMR Resonance Assignment Assisted by Mass Spectrometry This lecture talked about a NMR resonance assignment assisted by mass spectrometry [1, 2]. 1 Motivation Nuclear magnetic resonance (NMR) provides
IgE binds asymmetrically to its B cell receptor CD23
 Supplementary Information IgE binds asymmetrically to its B cell receptor CD23 Balvinder Dhaliwal 1*, Marie O. Y. Pang 2, Anthony H. Keeble 2,3, Louisa K. James 2,4, Hannah J. Gould 2, James M. McDonnell
Supplementary Information IgE binds asymmetrically to its B cell receptor CD23 Balvinder Dhaliwal 1*, Marie O. Y. Pang 2, Anthony H. Keeble 2,3, Louisa K. James 2,4, Hannah J. Gould 2, James M. McDonnell
Statistical discovery of site inter-dependencies in sub-molecular hierarchical protein structuring
 Durston et al. EURASIP Journal on Bioinformatics and Systems Biology 2012, 2012:8 RESEARCH Open Access Statistical discovery of site inter-dependencies in sub-molecular hierarchical protein structuring
Durston et al. EURASIP Journal on Bioinformatics and Systems Biology 2012, 2012:8 RESEARCH Open Access Statistical discovery of site inter-dependencies in sub-molecular hierarchical protein structuring
PROTEIN'STRUCTURE'DETERMINATION'
 PROTEIN'STRUCTURE'DETERMINATION' USING'NMR'RESTRAINTS' BCMB/CHEM'8190' Programs for NMR Based Structure Determination CNS - Brünger, A. T.; Adams, P. D.; Clore, G. M.; DeLano, W. L.; Gros, P.; Grosse-Kunstleve,
PROTEIN'STRUCTURE'DETERMINATION' USING'NMR'RESTRAINTS' BCMB/CHEM'8190' Programs for NMR Based Structure Determination CNS - Brünger, A. T.; Adams, P. D.; Clore, G. M.; DeLano, W. L.; Gros, P.; Grosse-Kunstleve,
Protein Structure Determination Using NMR Restraints BCMB/CHEM 8190
 Protein Structure Determination Using NMR Restraints BCMB/CHEM 8190 Programs for NMR Based Structure Determination CNS - Brunger, A. T.; Adams, P. D.; Clore, G. M.; DeLano, W. L.; Gros, P.; Grosse-Kunstleve,
Protein Structure Determination Using NMR Restraints BCMB/CHEM 8190 Programs for NMR Based Structure Determination CNS - Brunger, A. T.; Adams, P. D.; Clore, G. M.; DeLano, W. L.; Gros, P.; Grosse-Kunstleve,
INFO 4300 / CS4300 Information Retrieval. slides adapted from Hinrich Schütze s, linked from
 INFO 4300 / CS4300 Information Retrieval slides adapted from Hinrich Schütze s, linked from http://informationretrieval.org/ IR 8: Evaluation & SVD Paul Ginsparg Cornell University, Ithaca, NY 20 Sep 2011
INFO 4300 / CS4300 Information Retrieval slides adapted from Hinrich Schütze s, linked from http://informationretrieval.org/ IR 8: Evaluation & SVD Paul Ginsparg Cornell University, Ithaca, NY 20 Sep 2011
Chapter 4 Analysis of a cantilever
 Chapter 4 Analysis of a cantilever Before a complex structure is studied performing a seismic analysis, the behaviour of simpler ones should be fully understood. To achieve this knowledge we will start
Chapter 4 Analysis of a cantilever Before a complex structure is studied performing a seismic analysis, the behaviour of simpler ones should be fully understood. To achieve this knowledge we will start
CLASS NOTES Computational Methods for Engineering Applications I Spring 2015
 CLASS NOTES Computational Methods for Engineering Applications I Spring 2015 Petros Koumoutsakos Gerardo Tauriello (Last update: July 27, 2015) IMPORTANT DISCLAIMERS 1. REFERENCES: Much of the material
CLASS NOTES Computational Methods for Engineering Applications I Spring 2015 Petros Koumoutsakos Gerardo Tauriello (Last update: July 27, 2015) IMPORTANT DISCLAIMERS 1. REFERENCES: Much of the material
T H E J O U R N A L O F G E N E R A L P H Y S I O L O G Y. jgp
 S u p p l e m e n ta l m at e r i a l jgp Lee et al., http://www.jgp.org/cgi/content/full/jgp.201411219/dc1 T H E J O U R N A L O F G E N E R A L P H Y S I O L O G Y S u p p l e m e n ta l D I S C U S
S u p p l e m e n ta l m at e r i a l jgp Lee et al., http://www.jgp.org/cgi/content/full/jgp.201411219/dc1 T H E J O U R N A L O F G E N E R A L P H Y S I O L O G Y S u p p l e m e n ta l D I S C U S
Deep Learning Book Notes Chapter 2: Linear Algebra
 Deep Learning Book Notes Chapter 2: Linear Algebra Compiled By: Abhinaba Bala, Dakshit Agrawal, Mohit Jain Section 2.1: Scalars, Vectors, Matrices and Tensors Scalar Single Number Lowercase names in italic
Deep Learning Book Notes Chapter 2: Linear Algebra Compiled By: Abhinaba Bala, Dakshit Agrawal, Mohit Jain Section 2.1: Scalars, Vectors, Matrices and Tensors Scalar Single Number Lowercase names in italic
Protein structure sampling based on molecular dynamics and improvement of docking prediction
 1 1 1, 2 1 NMR X Protein structure sampling based on molecular dynamics and improvement of docking prediction Yusuke Matsuzaki, 1 Yuri Matsuzaki, 1 Masakazu Sekijima 1, 2 and Yutaka Akiyama 1 When a protein
1 1 1, 2 1 NMR X Protein structure sampling based on molecular dynamics and improvement of docking prediction Yusuke Matsuzaki, 1 Yuri Matsuzaki, 1 Masakazu Sekijima 1, 2 and Yutaka Akiyama 1 When a protein
Advanced ensemble modelling of flexible macromolecules using X- ray solution scattering
 Supporting information IUCrJ Volume 2 (2015) Supporting information for article: Advanced ensemble modelling of flexible macromolecules using X- ray solution scattering Giancarlo Tria, Haydyn D. T. Mertens,
Supporting information IUCrJ Volume 2 (2015) Supporting information for article: Advanced ensemble modelling of flexible macromolecules using X- ray solution scattering Giancarlo Tria, Haydyn D. T. Mertens,
Methods of Protein Structure Comparison. Irina Kufareva and Ruben Abagyan
 Chapter 1 Methods of Protein Structure Comparison Irina Kufareva and Ruben Abagyan Abstract Despite its apparent simplicity, the problem of quantifying the differences between two structures of the same
Chapter 1 Methods of Protein Structure Comparison Irina Kufareva and Ruben Abagyan Abstract Despite its apparent simplicity, the problem of quantifying the differences between two structures of the same
