As featured in: Nanoscale
|
|
|
- Hugo Francis
- 6 years ago
- Views:
Transcription
1 ISSN PAPER S. Balog, A. Petri-Fink et al. Characterizing nanoparticles in complex biological media and physiological fluids with depolarized dynamic light scattering Volume 7 Number April 2015 Pages Showcasing research from the Department of Mechanical and Aerospace Engineering, The Ohio State University, Columbus, OH, USA. Mechanical design of DNA nanostructures Structural DNA nanotechnology is a rapidly emerging field that enables precise design of nanoscale geometries and incorporation of chemical functionalities. Major advances in the last two decades have included consideration of mechanical behavior such as stiffness, deformation, and structural dynamics. In particular, we have developed a unique approach to make DNA machine components like the ones shown above following macroscopic engineering design concepts resulting in fundamental advances in the ability to control complex motion and mechanical behavior of DNA nanostructures. As featured in: See Carlos E. Castro et al., 2015, 7, Registered charity number:
2 FEATURE ARTICLE View Article Online View Journal View Issue Mechanical design of DNA nanostructures Cite this:, 2015, 7, 5913 Carlos E. Castro,* a,b Hai-Jun Su, a Alexander E. Marras, a Lifeng Zhou a and Joshua Johnson b Received 3rd December 2014, Accepted 24th January 2015 DOI: /c4nr07153k Introduction Biological cells rely on biomolecular machinery to perform essential processes such as physical and chemical communication with their environment. The function of these macromolecules, including proteins, DNA, and RNA, is dependent on specific characteristics, namely geometry, chemical functionality, and mechanical behavior, which are encoded in primary amino acid or nucleotide sequences. Molecular motors, for example, follow tracks with precisely spaced and oriented docking sites, hydrolyze ATP, and generate mechanical forces to transport intracellular cargo or drive cellular motion. Biomolecular nanotechnology has made great strides in developing the capacity to mimic these functional characteristics (geometry, chemical functionality, and mechanical behavior) in designed systems fabricated via molecular selfassembly. Amino acid based design 1,2 affords the long-term possibility of reaching the functional scope of cellular proteins such as molecular motors or enzymes; however, the complexity of interactions that govern amino acid folding and enable chemical functionality make de novo design of precise protein geometries highly challenging. DNA assembly, on the other hand, is governed by well-understood Watson Crick basepairing interactions. 3 While the chemical functionality of DNA is limited relative to proteins, the programmable nature of a Department of Mechanical and Aerospace Engineering, The Ohio State University, Columbus, OH 43210, USA. castro.39@osu.edu b Biophysics Graduate Program, The Ohio State University, Columbus, OH 43210, USA Electronic supplementary information (ESI) available. See DOI: / c4nr07153k Structural DNA nanotechnology is a rapidly emerging field that has demonstrated great potential for applications such as single molecule sensing, drug delivery, and templating molecular components. As the applications of DNA nanotechnology expand, a consideration of their mechanical behavior is becoming essential to understand how these structures will respond to physical interactions. This review considers three major avenues of recent progress in this area: (1) measuring and designing mechanical properties of DNA nanostructures, (2) designing complex nanostructures based on imposed mechanical stresses, and (3) designing and controlling structurally dynamic nanostructures. This work has laid the foundation for mechanically active nanomachines that can generate, transmit, and respond to physical cues in molecular systems. base-pairing assembly has provided a foundation for the rapidly progressing field of structural DNA nanotechnology. Since its inception in the early 1980 s through the foundational work of Nadrian Seeman, 4 structural DNA nanotechnology has evolved as a field of geometric design starting with lattices (geometrically connected immobile DNA junctions 4 6 ), then leading to DNA nanotubes and arrays, 7 11 and more recently developing shapes of almost arbitrary geometric complexity with the development of scaffolded DNA origami 12 and its extension to 3D. 13 The majority of current applications for DNA origami rely primarily on the ability to design complex and mechanically stiff nanoscale geometries. Examples include nanotubes to assist structure alignment for solution NMR studies, 14 nanopores for single molecule sensing, platforms for superresolution fluorescence microscopy, templates for nanotubes, 21 nanoparticles, and vehicles for small molecule drug delivery Other efforts have exploited the ability to modify DNA in order to expand the range of chemical functionality by integrating small molecules, 29,30 DNA aptamers, peptides, 34 proteins, 32,35,36 or other biomolecules. 16,37,38 Recent examples demonstrating the power of these chemical modifications include coupling DNA origami nanostructures with motor proteins for studies of motor cooperativity 39 and integration of DNA origami nanostructures into lipid membranes via carbohydrate modifications 16,38 thus opening possibilities to study cell membrane surfaces and interactions. While the chemical functionality of DNA nanostructures has been expanded through modification, the mechanical functionality beyond designing specific geometries has been less widely explored. DNA origami nanostructures, in particular, This journal is The Royal Society of Chemistry 2015,2015,7,
3 are often designed with a specific number of helices in order to approximate some complex geometry, but with relatively little consideration for the resulting mechanical behavior. Generally, this approach results in sufficient stiffness to minimize thermal fluctuations and maintain a well-defined shape that defines the function of the nanostructure. However, just as incorporating chemical modifications enhances functional scope, specific consideration and design of mechanical behavior can open possibilities to a broad range of new applications that include physical communication with the local environment. Here we highlight recent progress in the area of DNA origami nanotechnology specifically with regards to (1) designing and characterizing mechanical stiffness, (2) exploiting mechanical stress in DNA nanostructure design, and (3) designing and actuating dynamic behavior (i.e. motion and reconfiguration) of DNA nanostructures. Designing and characterizing mechanical stiffness of DNA origami nanostructures The mechanical properties of double-stranded DNA (dsdna) have been widely-studied since they play a critical role in gene regulation and DNA replication and repair. Compared to structural biopolymers (e.g. actin or microtubules), DNA is flexible, which allows it to be tightly packaged into the nucleus and undergo structural dynamics critical for DNA processing. Structural DNA nanotechnology connects several dsdna helices into a compact geometry to create objects that can be several orders of magnitude stiffer than the constituent DNA. Ultimately, the stiffness of designed DNA nanostructures are a result of the physical properties of DNA, the number, locations, and stiffness of connections points, and the geometry of the structure. While the mechanics of DNA are well characterized, the stiffness of DNA junctions has primarily been investigated in terms of physiologically relevant Holliday junctions, 44,45 although recent computational efforts have begun to consider junction stiffness in DNA nanostructures Here we outline recent progress made in quantifying and designing the mechanical behavior of designed DNA nanostructures. We focus our discussion on the persistence length (L P ), defined as L P = K B / k b T (K B is the bending stiffness, k b is Boltzmann s constant, and T is absolute temperature) since mechanical characterization has largely focused on quantifying L P. Direct measurement of the stiffness of designed DNA nanostructures started with the development of DNA nanotubes (circumferentially connected helices) several μm in length. 7,9,10 Rothemund et al. 9 first measured the average persistence length (L P ) of DNA nanotubes 7 20 nm in diameter as, L P = 3.9 μm, approximately two orders of magnitude larger than dsdna (L P,dsDNA =50nm 40,43 ). Yin and co-workers 50 later developed an approach to control the diameter of DNA nanotubes by specifying the number of helices in the cross-section. Fig. 1 Mechanical Properties of DNA origami nanostructures. (a) Bundling dsdna helices into compact cross-sections increases persistence length, L P, similar to bundling of actin filaments. The red data shows persistence length measurements of actin bundles with different concentrations of bundling protein (different shapes) and the corresponding limits of uncoupled (L P scales with N) and rigidly coupled (L P scales with N 2 ) filaments (data reprinted from 54 with permission from Macmillan Publishers Ltd: Nature Materials, copyright 2006 and Dr. Andreas Bausch). The shaded gray region shows similar limits for uncoupled and rigidly coupled DNA helices. DNA nanotubes 50,51 (black circles) closely follow the rigidly coupled limit and can even exceed the N 2 scaling due to swelling of the cross-section. Schiffels et al. 51 (Data is reprinted with permission from (ref. 51), copyright 2013 American Chemical Society and with permission of Dr. Deborah Fygenson.) developed a model to predict L P assuming rigid coupling and accounting for swelling (solid black line). L P of scaffolded DNA origami nanostructures with 4-helix 46 and 6-helix 46,52 cross-sections approximately follow a power law scaling behavior of N 1.94 (dashed black line). Schematics of cross-sections are shown next to the corresponding data. The green and orange cross-sections are potential cross-sections that could have stiffness in the range of actin filaments. (b) We measured shape distributions of DNA origami 6-hb revealing L P = 1.4 μm, in good agreement with other measurements. (c) Stochastic simulations of shape distributions closely match 6- hb experiments, and demonstrate the reduced shape fluctuations for larger cross-sections. Schiffels et al. 51 found L P of these nanotubes ranged from 2.0 µm for a 5-helix tube up to 16.8 µm for a 10-helix tube (Fig. 1a, black circles), closely following the behavior of rigidly coupled helices. Persistence length measurements of scaffolded DNA origami nanostructures have been limited to small cross-sections that form objects >100 nm in length. Liedl and coworkers first measured L P of gel-purified 6-helix bundles (hb) as 1.6 µm, 52 and Kauert et al. 46 used magnetic tweezers to measure L P for 4-hb and 6-hb of 0.74 µm and 1.88 µm, respectively. Here, we present a conformational distribution for similar 6-hb (Fig. 1b), which was used to determine L P = 1.4 μm by measuring the splay width of the conformational distribution as a function of arc length 53 (details in ESI ). Fig. 1a shows a summary of L P measurements for DNA nanotubes and scaffolded DNA origami bundles, and for comparison the L P scaling of actin bundles 54 as a function of the number of polymers (i.e. dsdna helices or actin filaments) in the bundle, N. The gray shaded region shows the limits of loosely coupled (L P N) and rigidly coupled DNA origami nanostructures, which can exceed the typical L P N 2 scaling for rigidly coupled bundles 54 because they are not close packed (i.e. the cross-section contains voids). Direct calcu- 5914,2015,7, This journal is The Royal Society of Chemistry 2015
4 lation of the rigidly coupled limit (i.e. infinite stiffness of cross-overs prevents any relative motion of connected helices) for the four cross-sections shown (details in ESI ) reveals an upper limit scaling behavior of L P N 2.42 (note that this scaling behavior would vary with different cross-sectional geometries). A simple power law fit to the measured persistence length values for scaffolded DNA origami nanostructures reveals a scaling of L P N 1.94 suggesting DNA origami crossover connections result in stiff, but not rigid coupling between helices. In other words, some shearing between helices in the bundle may occur, but cross-over connections provide significant resistance to that shearing. Using this scaling, we can estimate the bending stiffness of larger DNA origami honeycomb cross-sections, for example the 18-hb or 54-hb shown as insets in Fig. 1a. This approach provides only rough estimates for L P of larger cross-sections. Previous work has shown that the L P of polymer bundles only follows simple power law scaling in the rigidly coupled or loosely coupled limits, and a more detailed theoretical 54 or computational approach 47,48 would be required to accurately predict the stiffness of larger cross-sections. However, these simple estimates at least indicate that nanostructures as stiff as actin filaments (L P = 18 µm (ref. 55)) and potentially even microtubules (L P = µm (ref. 55, 56)) could be made using scaffolded DNA origami. To illustrate the thermal fluctuations of DNA origami nanostructures, we performed stochastic simulations of DNA nanostructure configurations. Briefly, bundle trajectories were reconstructed as chains of segments where the change in angle between segments, Δθ, was defined according to the tangent angle correlation equation, cos(δθ) = exp( Δs/2L p ). The factor of 2 is present to reproduce 2D conformational distributions. The results for conformational distributions were not sensitive to selection of the segment length provided L segment /L bundle Details of simulations are provided in ESI. Fig. 1c shows simulations of conformational distributions using persistence lengths obtained from the scaling behavior in Fig. 1a for 6-hb, 18-hb, and 54-hb. Here we assumed the total number of base-pairs was limited to a typical DNA origami scaffold (7560 bp), 13,57 hence structures with larger cross-sections are shorter in length. The 6-hb simulation closely matches the experimental data. These simulations illustrate that DNA origami nanostructures containing 18-hb or more helices in the cross-section are not subject to significant thermal fluctuations, and hence maintain a well-defined shape. A few studies have explored the mechanical behavior and design of mechanical properties of DNA origami nanostructures more extensively than by just considering the bending persistence length. For example, Kauert et al. 46 measured the torsional persistence length of 4-helix and 6-helix bundles to be 390 nm and 530 nm, respectively; Plesa and co-workers 58 demonstrated that the bending stiffness of DNA origami nanoplates was critical to their functional ion conductance; and Pfitzner et al. 59 exploited the stiffness of DNA origami beams to improve the resolution of force spectroscopy experiments. In addition, our collaborative team 60 has recently explored the design of deformable (compliant) DNA nanostructures with tunable stiffness. Going forward, consideration of mechanical properties will be a critical factor in the design of devices that must withstand and transmit forces through interactions with other materials and molecules. The development and use of computational tools, such as CANDO 47,57 or recent efforts in molecular dynamics 48 and lattice-free structure prediction, 49 will undoubtedly provide useful tools for predicting and interpreting mechanical behavior of DNA nanostructures. Exploiting mechanical stress in DNA nanostructure design In biological cells, DNA is bent and twisted in order to be packaged into chromatin resulting in mechanical strain energy stored in the deformed polymer. This stored mechanical energy results in local stresses, often referred to as residual stress in macroscopic systems, that play a critical role in the unwrapping required for DNA processing. These residual stresses can be programmed into DNA origami nanostructures to induce additional geometric complexities in design, such as curvature or twist, much like a tensed string results in bending of a bow in a bow and arrow. This is achieved by introducing local tensile, compressive, or torsional stresses that combine to induce deformation of the overall structure. Fig. 2 shows several examples of complex DNA nanostructures where the geometry relies on residual stresses in the constituent DNA helices. While prior efforts had exploited mechanical properties, such as flexibility, in DNA assembly, 61 the use of stored mechanical energy to design deformed structures was first demonstrated by Dietz and co-workers 62 who locally deleted or added base-pairs between DNA origami cross-over connections to introduce stresses that caused bending or twist of the overall structure (Fig. 2a). The degree of bending or twist could be programmed according to the amount of stress (i.e. number of base-pairs added or deleted) and the overall bending or torsional stiffness of the structure. Han and co-workers 63 later established control of curvature in two directions by forcing cross-over connections between helices of different lengths to induce bending in the direction of the shorter helices. By spatially arranging longer and shorter helices in parallel, they created fully closed spherical objects (Fig. 2b). Han et al. 64 also developed a similar design framework by connecting helices in a cross-hatched pattern where perpendicular branched connections could be made between parallel helices of different lengths to induce 1D or 2D curvature (Fig. 2c). The previously described approaches rely on local stresses to create structures with complex curvature or twist that are still mechanically stiff and structurally well-defined (i.e. undergo minimal thermal fluctuations). To access a broader range of mechanical behavior, Liedl et al. 52 exploited the entropic elasticity of single-stranded DNA (ssdna) to develop DNA This journal is The Royal Society of Chemistry 2015,2015,7,
5 Fig. 2 Exploiting mechanical stress in DNA nanostructure design. (a) Curvature or twist can be programmed by adding or deleting base pairs between cross-over connections in DNA origami structures to introduce local stresses (From (ref. 62). Reprinted with permission from AAAS.). Scale bars are 20 nm (top) and 50 nm (bottom). (b) Similarly, forcing connections between mismatched lengths of dsdna with helices arranged in parallel (From (ref. 63). Reprinted with permission from AAAS.) or (c) perpendicularly in a cross-hatched pattern (From (ref. 64). Reprinted with permission from AAAS.) enables programming curvature and making curved shell or spherical objects. (d) The use of ssdna allows application of entropic forces that can induce compression (left) or bending (right) of dsdna components (Reprinted by permission from Macmillan Publishers Ltd: Nature Nanotechnology (ref. 52), copyright 2010). (e) Bending can be directed using anisotropic dsdna components to make components with tunable curvature (Adapted with permission from (ref. 60). Copyright 2014 American Chemical Society.). In addition tuning lengths of ssdna by partially folding it into dsdna, as in (d) on the right, or by shifting ssdna length between to a reservoir loop, as in (e), allows tuning the tension in the ssdna. Combining ssdna and dsdna ultimately allows access to a broader range of mechanical behavior. Scale bars are 50 nm in (b) and 20 nm in (d, e). origami tensegrity structures (Fig. 2d). In their DNA origami tensegrity structures, tension in ssdna components is balanced with compression or bending of dsdna components. The overall stiffness of the structures depends on both dsdna and ssdna components, thereby affording access to a large range of stiffness. Liedl and co-workers further developed a method to tune the length of ssdna components by partially folding them into small DNA origami bundles as depicted in Fig. 2d (right). We have recently expanded on this method to create DNA origami compliant (deformable) nanostructures with tunable geometry and stiffness. 60 Our approach exploits entropic elasticity of ssdna to bend an anisotropic stiff dsdna component, a single layer of 6-helices. Examples are shown in Fig. 2E. Both the stiffness and geometry of the structure can be precisely controlled by modifying the length of the ssdna connections, which we achieved by shifting ssdna between the tension-bearing connections and a neighboring ssdna loop. These two efforts demonstrate the potential to enhance DNA origami design, in particular mechanical functionality, through the incorporation of ssdna. In all of these approaches, the cumulative base-pairing energy outweighs the local mechanical strain energy due to deformation of dsdna so that the structure is still stable, although Dietz et al. 62 noted that structures with higher mechanical strain tended to fold with lower yield. Ultimately, the magnitude of mechanical stress that can be introduced or the localized force that can be applied is limited by the base pairing energy of the structure. However, design modifications (e.g. arranging helices in a shear as opposed to unzipping conformation 65 ), locally integrating more stable components (e.g. peptide nucleic acids 66 ), or even chemical modifications (e.g. local covalent cross-links between strands 67 ) could allow for mechanically enhanced structures that can withstand and transmit larger forces or store more mechanical energy. Furthermore, methods could be developed in the future to release this energy to mimic force generation of molecular motors 68 or biological springs. 69 Designing and actuating dynamic behavior of DNA nanostructures Dynamic behavior including thermal fluctuation, conformational changes, and directed motion, is a key aspect of the function of biomolecular machinery. The ability to design this dynamic behavior in synthetic systems is a central challenge of biomolecular nanotechnology. Although the majority of current applications of DNA nanostructures rely primarily on well-defined geometric design, some significant advances have been made since the early stages of the field in fabricating and controlling dynamic behavior of DNA nanostructures. These dynamic structures have been designed to achieve complex conformational changes (Fig. 3a,b (ref. 70, 71)), detect molecular binding events (Fig. 3c,d (ref. 72, 73)), or encapsulate and then release compounds or molecules (Fig. 3e h (ref. 31, 32, 74, 75)). The first controllable designed DNA nanostructure, introduced by Mao and co-workers, 76 exploited the B-Z transition of DNA, triggered by changing solution conditions, to make switchable devices made from a few strands of DNA. Since their breakthrough, research efforts to fabricate dynamic DNA nanosystems have primarily focused on nanostructures where conformational changes can be triggered by molecular binding events, or by strand invasion (i.e. strand displacement), 77 whereby a DNA strand in the folded structure is exchanged with a DNA input strand from solution usually mediated by initial binding to a ssdna overhang (i.e. toehold). 78 The strand invasion approaches started with Yurke et al. 71 who fabricated DNA tweezers comprising three strands that could be opened or closed with DNA inputs, shown schematically in Fig. 3a. Yan and co-workers extended this approach to demonstrate controlled rotational motion in a DNA nanostructure array. 79 These works laid a foundation for several reconfigurable structures with conformational changes triggered by binding of DNA inputs These systems achieve small ( 1 10 nm) motions that in some cases could be 5916,2015,7, This journal is The Royal Society of Chemistry 2015
6 Fig. 3 Mechanically dynamic nanostructures. Examples of dynamic DNA nanostructures include: (1) (a, b) structures designed to demonstrate complex conformational changes, (a) is reprinted by permission of Macmillan Publishers Ltd: Nature (ref. 71), copyright 2000, and (b) is reprinted by permission of Macmillan Publishers Ltd: Nature Nanotechnology (ref. 70), copyright (2) (c, d) devices that detect molecular binding events, (c) is reprinted with permission from (ref. 72), copyright 2010 American Chemical Society, and (d) is reprinted by permission from Macmillan Publishers Ltd: Nature Communications (ref. 73), copyright or (3) (e h) containers intended to encapsulate and subsequently release cargo, (e) is reprinted by permission of Macmillan Publishers Ltd: Nature (ref. 74), copyright (f) is reprinted with permission from (ref. 75), copyright 2012 American Chemical Society. (g) is from (ref. 32) and is reprinted with permission from AAAS. (h) is from (ref. 31) copyright 2013 by John Wiley Sons, Inc and reprinted with permission of John Wiley Sons, Inc. accumulated on a track to add up to 100 nm 84 and transport 85 or even assemble 86 nanoscale components. Similar strand displacement approaches form the basis of DNA-based molecular computing. 87,88 The establishment of DNA origami techniques provided a platform to direct local strand reconfiguration, for example to guide DNA walkers, 84 harness logic steps, 89 control molecular binding, 73 or locally reconfigure strands on a DNA platform. 90 Building on these approaches, strand displacement has been used to control larger-scale structural transitions in DNA origami systems. For example, Han et al. 70 created a DNA origami Möbius strip (one-sided ribbon structure) that could be cut open via strand displacement reactions to approximately double in size, depicted schematically in Fig. 3b. Other recent efforts have expanded this work to achieve reversible actuation of DNA nanostructures. Kuzuya et al. 73 demonstrated reversible actuation in a DNA origami tweezer-like structure that could be actuated into a closed state upon binding of biomolecules (e.g. streptavidin or oligonucleotides) and then reopened by DNA strand displacement (Fig. 3d). Recently, Kuzyk and co-workers 91 used a similar scissor-like device to actuate changes in chirality of gold nanorods enabling reversible control of plasmonic interactions. Feng and co-workers 92 demonstrated that reversible conformational changes could be executed in a lattice system where repeat units in the lattice were extended and subsequently shortened, both via DNA strand invasion. Both Feng and Kuzuya implemented distributed actuation methods where DNA inputs bind to or displace multiple strands on the structure to drive conformational changes. The strand invasion concept has also been implemented to construct DNA containers for drug release 31,32,74,75 (Fig. 3e h). These approaches exploit the energy of DNA basepairing that is achieved during the initial self-assembly process to hold the structure in a closed configuration while molecules are trapped inside. When local connections are released by strand displacement, the system relaxes to an open or unconstrained state where molecules diffuse out of the container or at least are exposed to solution. Douglas et al. 32 (Fig. 3g) and Banerjee et al. 31 (Fig. 3h) expanded the scope of the strand invasion process using DNA aptamers to lock containers closed that could then open in the presence of a non-nucleotide target molecules (i.e. proteins or small molecules). This use of DNA aptamers provides access to a diverse range of interaction targets in vitro or in vivo. Our research team has recently expanded on dynamic DNAbased design by adopting an engineering machine design approach. 93 Like in macroscopic machine design, our approach starts with fabricating joints with constrained motion in specified rotational or translational degrees of freedom. Our general approach for designing constrained motion is to combine precisely designed stiff dsdna components with selectively placed ssdna strands so that the geometry of the stiff components and the placement of the flexible connections allow motion only in specific degrees of freedom. We call these nanostructures DNA origami mechanisms (DOM). Fig. 4a and 4b show schematics and TEM images of DNA origami joints with a single dominant angular (Fig. 4a) or linear (Fig. 4b) degree of freedom. While angular motion has been previously demonstrated in several of the aforementioned studies, our work 93 was the first to demonstrate controlled linear motion. In our framework, more complex motion can be designed by incorporating multiple joints into a single structure. These joints can move independently as in the case of the universal joint presented here (Fig. 4c), which has two angular degrees of freedom (gel analysis and additional TEM images are pro- This journal is The Royal Society of Chemistry 2015,2015,7,
7 Fig. 4 DNA origami machine components. A framework for designing DNA origami mechanisms and machines 93 starts with designing basic joints with motion in specified degrees of freedom. Schematics and TEM images depict joints with (a) a single dominant rotational or (b) linear degree of freedom. (c) DNA origami joints can also exhibit multiple independent degrees of freedom as in the universal joint, or (d) these joints can be integrated into a mechanism which couples rotational to linear motion, as in the slider-crank. These mechanisms can be assembled into larger structures that resemble macroscopic machines as demonstrated by the scissor mechanism in (e). The DNA origami scissor mechanism was constructed by first assembling the structural unit (left) and then polymerizing a 1D array (right) that couples rotational motion of the unit into linear motion of the mechanism. The scale bar in (a) is 20 nm and all other scale bars are 50 nm. The structures in (a), (b) and (d) are reprinted from our previous work. 93 The structures in (c) and (e) are presented here for the first time. The scale bars were already defined. vided in the ESI ); or they can be integrated into a mechanism, such as the crank-slider 93 (Fig. 4d), which comprises components with rotational and linear motion that are coupled resulting in a mechanism with 2D motion with a single degree of freedom. This machine-like design approach opens the door for designing complex devices with well-defined motion and/ or force transmission. For example, the crank-slider could translate a binding event that drives rotational motion ( previously demonstrated by others 72,73,94 ) into linear motion to generate a linear force or facilitate a downstream interaction. Alternatively, it could generate a torque in response to a linear displacement. A major direction in the field of biomolecular nanotechnology is to reproduce the complex function of natural molecular machinery. In contrast, a major cornerstone of our efforts in DNA origami nanotechnology relies on drawing parallels to and implementing ideas from macroscopic engineering structure and machine design. The ability to design stiff components of essentially arbitrary geometric complexity and integrate constrained motion in specified degrees of freedom enables a modular approach to piece components together in machines that resemble macroscopic systems. To illustrate this concept, we have fabricated a DNA origami scissor mechanism (Fig. 4e) that resembles and functions similarly to a macroscopic scissor lift. The DNA origami scissor mechanism is constructed by first decomposing the mechanism into a unit cell that has three rotational degrees of freedom (Fig. 4e left). This unit was polymerized by sticky end interactions to form a 1D array where the rotational degrees of freedom couple to achieve linear motion. The TEM images on the right of Fig. 4e depict an open and an extended state of a 3-mer polymerized scissor mechanism. In this case, this motion was achieved simply by thermal fluctuation along the constrained motion path. Our future work will pursue actuation of this scissor mechanism and other extendable DNA origami mechanisms to couple distributed molecular actuation to force generation and μm scale motion. Full design details, folding protocols, gel analysis, and additional TEM images for the scissor mechanism are available in the ESI. Concluding remarks Here we have detailed major strides in designing and exploiting the mechanical behavior of DNA origami nanostructures, specifically in characterizing and designing mechanical stiffness, exploiting mechanical stress in geometric design, and fabricating and actuating dynamic DNA nanostructures. In particular, the recent progress in DNA origami mechanism (DOM) design paves the way to mimic the macroscopic machines for designing dynamic nanostructures with controllable complex motion. In the future, expanding the range of accessible mechanical behavior of DNA origami nanostructures will greatly expand the scope of potential applications. Already some current applications exploit the 5918,2015,7, This journal is The Royal Society of Chemistry 2015
8 mechanical behavior of DNA nanostructures, for example to obtain improved resolution in force spectroscopy, 59 detect DNA protein binding interactions, 72 or exploit flexibility of ssdna to facilitate enzyme activity. 95 Further advances in understanding and modeling the mechanical behavior of designed DNA nanostructures will enable improved design of mechanical properties, dynamic behavior, and force generation mechanisms similar to natural molecular machines. A promising future direction for mechanical design of DNA nanostructures is the direct design of complex DNA origami energy landscapes that include multiple stable states and energy barriers to regulate conformational dynamics and physical interactions with the local environment. In the future, designed mechanical behavior can be integrated with precise geometry and chemical functionality to create DNA-based nanomachines that can transmit and generate forces, and more generally perform chemical and mechanical work, to achieve complex functions such as the ability to sense and respond to their environment; manipulate, assemble, and transport molecular components; actuate and control chemical reactions; and even store and convert energy. A particularly exciting possibility would be interfacing these nanomachines with DNA-based molecular computing to achieve dynamic mechanical systems that can process and generate information. With the ability to design and control complex mechanical behavior, combined with chemically functionalizing DNA nanostructures and recent advances towards fabricating larger DNA origami systems 96 it seems the functional scope of DNA nanotechnology will continue to expand for years to come. Acknowledgements This work was supported primarily by the National Science Foundation under grant no: CMMI and grant no: CMMI Any opinions, finding, conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of the funding agency. References 1 D. Baker, Philos. Trans. R. Soc., B, 2006, 361, T. Aida, E. W. Meijer and S. I. Stupp, Science, 2012, 335, J. D. Watson and F. H. Crick, Nature, 1953, 171, N. C. Seeman, J. Theor. Biol., 1982, 99, N. C. Seeman and N. R. Kallenbach, Biophys. J., 1983, 44, J. H. Chen and N. C. Seeman, Nature, 1991, 350, D. Liu, S. H. Park, J. H. Reif and T. H. LaBean, Proc. Natl. Acad. Sci. U. S. A., 2004, 101, J. C. Mitchell, J. R. Harris, J. Malo, J. Bath and A. J. Turberfield, J. Am. Chem. Soc., 2004, 126, P. W. Rothemund, A. Ekani-Nkodo, N. Papadakis, A. Kumar, D. K. Fygenson and E. Winfree, J. Am. Chem. Soc., 2004, 126, H. Yan, S. H. Park, G. Finkelstein, J. H. Reif and T. H. LaBean, Science, 2003, 301, D. Liu, M. Wang, Z. Deng, R. Walulu and C. Mao, J. Am. Chem. Soc., 2004, 126, P. W. K. Rothemund, Nature, 2006, 440, S. M. Douglas, H. Dietz, T. Liedl, B. Hogberg, F. Graf and W. M. Shih, Nature, 2009, 459, S. M. Douglas, J. J. Chou and W. M. Shih, Proc. Natl. Acad. Sci. U. S. A., 2007, 104, N. A. Bell, C. R. Engst, M. Ablay, G. Divitini, C. Ducati, T. Liedl and U. F. Keyser, Nano Lett., 2012, 12, M. Langecker, V. Arnaut, T. G. Martin, J. List, S. Renner, M. Mayer, H. Dietz and F. C. Simmel, Science, 2012, 338, R. Wei, T. G. Martin, U. Rant and H. Dietz, Angew. Chem., Int. Ed., 2012, 51, R. Jungmann, M. S. Avendano, J. B. Woehrstein, M. Dai, W. M. Shih and P. Yin, Nat. Methods, 2014, 11, R. Jungmann, C. Steinhauer, M. Scheible, A. Kuzyk, P. Tinnefeld and F. C. Simmel, Nano Lett., 2010, 10, J. J. Schmied, C. Forthmann, E. Pibiri, B. Lalkens, P. Nickels, T. Liedl and P. Tinnefeld, Nano Lett., 2013, 13, H. T. Maune, S. P. Han, R. D. Barish, M. Bockrath, W. A. Iii, P. W. Rothemund and E. Winfree, Nat. Nanotechnol., 2010, 5, H. Bui, C. Onodera, C. Kidwell, Y. Tan, E. Graugnard, W. Kuang, J. Lee, W. B. Knowlton, B. Yurke and W. L. Hughes, Nano Lett., 2010, 10, A. Kuzyk, R. Schreiber, Z. Y. Fan, G. Pardatscher, E. M. Roller, A. Hogele, F. C. Simmel, A. O. Govorov and T. Liedl, Nature, 2012, 483, R. Schreiber, J. Do, E. M. Roller, T. Zhang, V. J. Schuller, P. C. Nickels, J. Feldmann and T. Liedl, Nat. Nanotechnol., 2014, 9, R. Schreiber, S. Kempter, S. Holler, V. Schuller, D. Schiffels, S. S. Simmel, P. C. Nickels and T. Liedl, Small, 2011, 7, Q. Jiang, C. Song, J. Nangreave, X. Liu, L. Lin, D. Qiu, Z. G. Wang, G. Zou, X. Liang, H. Yan and B. Ding, J. Am. Chem. Soc., 2012, 134, Q. Zhang, Q. Jiang, N. Li, L. Dai, Q. Liu, L. Song, J. Wang, Y. Li, J. Tian, B. Ding and Y. Du, ACS Nano, 2014, 8, Y. X. Zhao, A. Shaw, X. Zeng, E. Benson, A. M. Nystrom and B. Hogberg, ACS Nano, 2012, 6, H. Lee, A. K. Lytton-Jean, Y. Chen, K. T. Love, A. I. Park, E. D. Karagiannis, A. Sehgal, W. Querbes, C. S. Zurenko, M. Jayaraman, C. G. Peng, K. Charisse, A. Borodovsky, M. Manoharan, J. S. Donahoe, J. Truelove, M. Nahrendorf, R. Langer and D. G. Anderson, Nat. Nanotechnol., 2012, 7, This journal is The Royal Society of Chemistry 2015,2015,7,
9 30 N. V. Voigt, T. Torring, A. Rotaru, M. F. Jacobsen, J. B. Ravnsbaek, R. Subramani, W. Mamdouh, J. Kjems, A. Mokhir, F. Besenbacher and K. V. Gothelf, Nat. Nanotechnol., 2010, 5, A. Banerjee, D. Bhatia, A. Saminathan, S. Chakraborty, S. Kar and Y. Krishnan, Angew. Chem., Int. Ed., 2013, 52, S. M. Douglas, I. Bachelet and G. M. Church, Science, 2012, 335, M. Tintore, I. Gallego, B. Manning, R. Eritja and C. Fabrega, Angew. Chem., Int. Ed., 2013, 52, A. Udomprasert, M. N. Bongiovanni, R. Sha, W. B. Sherman, T. Wang, P. S. Arora, J. W. Canary, S. L. Gras and N. C. Seeman, Nat. Nanotechnol., 2014, 9, B. Sacca, R. Meyer, M. Erkelenz, K. Kiko, A. Arndt, H. Schroeder, K. S. Rabe and C. M. Niemeyer, Angew. Chem., Int. Ed., 2010, 49, A. Shaw, V. Lundin, E. Petrova, F. Fordos, E. Benson, A. Al-Amin, A. Herland, A. Blokzijl, B. Hogberg and A. I. Teixeira, Nat. Methods, 2014, 11, P. Wang, S. H. Ko, C. Tian, C. Hao and C. Mao, Chem. Commun., 2013, 49, A. Johnson-Buck, S. Jiang, H. Yan and N. G. Walter, ACS Nano, 2014, 8, N. D. Derr, B. S. Goodman, R. Jungmann, A. E. Leschziner, W. M. Shih and S. L. Reck-Peterson, Science, 2012, 338, C. Bustamante, S. B. Smith, J. Liphardt and D. Smith, Curr. Opin. Struct. Biol., 2000, 10, J. Lipfert, J. W. Kerssemakers, T. Jager and N. H. Dekker, Nat. Methods, 2010, 7, J. F. Marko and E. D. Siggia, Macromolecules, 1995, 28, M. D. Wang, H. Yin, R. Landick, J. Gelles and S. M. Block, Biophys. J., 1997, 72, S. A. McKinney, A. C. Declais, D. M. Lilley and T. Ha, Nat. Struct. Biol., 2003, 10, M. A. Karymov, M. Chinnaraj, A. Bogdanov, A. R. Srinivasan, G. Zheng, W. K. Olson and Y. L. Lyubchenko, Biophys. J., 2008, 95, D. J. Kauert, T. Kurth, T. Liedl and R. Seidel, Nano Lett., 2011, 11, D. N. Kim, F. Kilchherr, H. Dietz and M. Bathe, Nucleic Acids Res., 2012, 40, J. Yoo and A. Aksimentiev, Proc. Natl. Acad. Sci. U. S. A., 2013, 110, K. Pan, D. N. Kim, F. Zhang, M. R. Adendorff, H. Yan and M. Bathe, Nat. Commun., 2014, 5, P. Yin, R. F. Hariadi, S. Sahu, H. M. Choi, S. H. Park, T. H. Labean and J. H. Reif, Science, 2008, 321, D. Schiffels, T. Liedl and D. K. Fygenson, ACS Nano, 2013, 7, T. Liedl, B. Hogberg, J. Tytell, D. E. Ingber and W. M. Shih, Nat. Nanotechnol., 2010, 5, H. Isambert, P. Venier, A. C. Maggs, A. Fattoum, R. Kassab, D. Pantaloni and M. F. Carlier, J. Biol. Chem., 1995, 270, M. M. Claessens, M. Bathe, E. Frey and A. R. Bausch, Nat. Mater., 2006, 5, F. Gittes, B. Mickey, J. Nettleton and J. Howard, J. Cell Biol., 1993, 120, F. Pampaloni, G. Lattanzi, A. Jonas, T. Surrey, E. Frey and E. L. Florin, Proc. Natl. Acad. Sci. U. S. A., 2006, 103, C. E. Castro, F. Kilchherr, D. N. Kim, E. L. Shiao, T. Wauer, P. Wortmann, M. Bathe and H. Dietz, Nat. Methods, 2011, 8, C. Plesa, A. N. Ananth, V. Linko, C. Gulcher, A. J. Katan, H. Dietz and C. Dekker, ACS Nano, 2014, 8, E. Pfitzner, C. Wachauf, F. Kilchherr, B. Pelz, W. M. Shih, M. Rief and H. Dietz, Angew. Chem., Int. Ed., 2013, 52, L. Zhou, A. E. Marras, H. J. Su and C. E. Castro, ACS Nano, 2014, 8, C. Zhang, M. Su, Y. He, X. Zhao, P. A. Fang, A. E. Ribbe, W. Jiang and C. Mao, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, H. Dietz, S. M. Douglas and W. M. Shih, Science, 2009, 325, D. Han, S. Pal, J. Nangreave, Z. Deng, Y. Liu and H. Yan, Science, 2011, 332, D. Han, S. Pal, Y. Yang, S. Jiang, J. Nangreave, Y. Liu and H. Yan, Science, 2013, 339, M. J. Lang, P. M. Fordyce, A. M. Engh, K. C. Neuman and S. M. Block, Nat. Methods, 2004, 1, M. Eriksson and P. E. Nielsen, Q. R. Biophys., 1996, 29, A. Rajendran, M. Endo, Y. Katsuda, K. Hidaka and H. Sugiyama, J. Am. Chem. Soc., 2011, 133, K. Svoboda and S. M. Block, Cell, 1994, 77, J. H. Shin, B. K. Tam, R. R. Brau, M. J. Lang, L. Mahadevan and P. Matsudaira, Biophys. J., 2007, 92, D. R. Han, S. Pal, Y. Liu and H. Yan, Nat. Nanotechnol., 2010, 5, B. Yurke, A. J. Turberfield, A. P. Mills Jr., F. C. Simmel and J. L. Neumann, Nature, 2000, 406, H. Gu, W. Yang and N. C. Seeman, J. Am. Chem. Soc., 2010, 132, A. Kuzuya, Y. Sakai, T. Yamazaki, Y. Xu and M. Komiyama, Nat. Commun., 2011, 2, E. S. Andersen, M. Dong, M. M. Nielsen, K. Jahn, R. Subramani, W. Mamdouh, M. M. Golas, B. Sander, H. Stark, C. L. Oliveira, J. S. Pedersen, V. Birkedal, F. Besenbacher, K. V. Gothelf and J. Kjems, Nature, 2009, 459, R. M. Zadegan, M. D. Jepsen, K. E. Thomsen, A. H. Okholm, D. H. Schaffert, E. S. Andersen, V. Birkedal and J. Kjems, ACS Nano, 2012, 6, C. Mao, W. Sun, Z. Shen and N. C. Seeman, Nature, 1999, 397, ,2015,7, This journal is The Royal Society of Chemistry 2015
10 77 D. Y. Zhang and G. Seelig, Nat. Chem., 2011, 3, N. Srinivas, T. E. Ouldridge, P. Sulc, J. M. Schaeffer, B. Yurke, A. A. Louis, J. P. Doye and E. Winfree, Nucleic Acids Res., 2013, 41, H. Yan, X. Zhang, Z. Shen and N. C. Seeman, Nature, 2002, 415, D. Y. Zhang, A. J. Turberfield, B. Yurke and E. Winfree, Science, 2007, 318, T. Omabegho, R. Sha and N. C. Seeman, Science, 2009, 324, P. Yin, H. Yan, X. G. Daniell, A. J. Turberfield and J. H. Reif, Angew. Chem., Int. Ed., 2004, 43, B. Ding and N. C. Seeman, Science, 2006, 314, K. Lund, A. J. Manzo, N. Dabby, N. Michelotti, A. Johnson- Buck, J. Nangreave, S. Taylor, R. Pei, M. N. Stojanovic, N. G. Walter, E. Winfree and H. Yan, Nature, 2010, 465, T. G. Cha, J. Pan, H. Chen, J. Salgado, X. Li, C. Mao and J. H. Choi, Nat. Nanotechnol., 2014, 9, H. Gu, J. Chao, S. J. Xiao and N. C. Seeman, Nature, 2010, 465, L. Qian and E. Winfree, Science, 2011, 332, L. Qian, E. Winfree and J. Bruck, Nature, 2011, 475, D. Y. Zhang, R. F. Hariadi, H. M. Choi and E. Winfree, Nat. Commun., 2013, 4, F. Zhang, J. Nangreave, Y. Liu and H. Yan, Nano Lett., 2012, 12, A. Kuzyk, R. Schreiber, H. Zhang, A. O. Govorov, T. Liedl and N. Liu, Nat. Mater., 2014, 13, L. Feng, S. H. Park, J. H. Reif and H. Yan, Angew. Chem., Int. Ed., 2003, 42, A. E. Marras, L. Zhou, H. J. Su and C. E. Castro, Proc. Natl. Acad. Sci. U. S. A., 2015, 112, G. J. Lavella, A. D. Jadhav and M. M. Maharbiz, Nano Lett., 2012, 12, J. Fu, Y. R. Yang, A. Johnson-Buck, M. Liu, Y. Liu, N. G. Walter, N. W. Woodbury and H. Yan, Nat. Nanotechnol., 2014, 9, A. N. Marchi, I. Saaem, B. N. Vogen, S. Brown and T. H. LaBean, Nano Lett., 2014, 14, This journal is The Royal Society of Chemistry 2015,2015,7,
Dedicated softwares for DNA origami designs
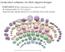 Dedicated softwares for DNA origami designs DAEDALUS (http://daedalus-dna-origami.org/) à independent to M13 ssdna à generates dedicated DNA scaffolds (450-3,400 bases) NANOANDES 2017, November 22-29,
Dedicated softwares for DNA origami designs DAEDALUS (http://daedalus-dna-origami.org/) à independent to M13 ssdna à generates dedicated DNA scaffolds (450-3,400 bases) NANOANDES 2017, November 22-29,
Nucleic acid self-assembly has proven to be a powerful tool
 pubs.acs.org/nanolett Isothermal Self-Assembly of Complex DNA Structures under Diverse and Biocompatible Conditions Cameron Myhrvold,, Mingjie Dai,, Pamela A. Silver,, and Peng Yin*,, Department of Systems
pubs.acs.org/nanolett Isothermal Self-Assembly of Complex DNA Structures under Diverse and Biocompatible Conditions Cameron Myhrvold,, Mingjie Dai,, Pamela A. Silver,, and Peng Yin*,, Department of Systems
Intrinsic DNA Curvature of Double-Crossover Tiles
 Intrinsic DNA Curvature of Double-Crossover Tiles Seungjae Kim, 1 Junghoon Kim, 1 Pengfei Qian, 2 Jihoon Shin, 3 Rashid Amin, 3 Sang Jung Ahn, 4 Thomas H. LaBean, 5 Moon Ki Kim, 2,3 * and Sung Ha Park,
Intrinsic DNA Curvature of Double-Crossover Tiles Seungjae Kim, 1 Junghoon Kim, 1 Pengfei Qian, 2 Jihoon Shin, 3 Rashid Amin, 3 Sang Jung Ahn, 4 Thomas H. LaBean, 5 Moon Ki Kim, 2,3 * and Sung Ha Park,
Interdisciplinary Nanoscience Center University of Aarhus, Denmark. Design and Imaging. Assistant Professor.
 Interdisciplinary Nanoscience Center University of Aarhus, Denmark Design and Imaging DNA Nanostructures Assistant Professor Wael Mamdouh wael@inano.dk Molecular Self-assembly Synthesis, SPM microscopy,
Interdisciplinary Nanoscience Center University of Aarhus, Denmark Design and Imaging DNA Nanostructures Assistant Professor Wael Mamdouh wael@inano.dk Molecular Self-assembly Synthesis, SPM microscopy,
Rational Design of DNA Motors: Fuel Optimization through Single- Molecule Fluorescence
 pubs.acs.org/jacs Rational Design of DNA Motors: Fuel Optimization through Single- Molecule Fluorescence Toma E. Tomov, Roman Tsukanov, Miran Liber, Rula Masoud, Noa Plavner, and Eyal Nir* Department of
pubs.acs.org/jacs Rational Design of DNA Motors: Fuel Optimization through Single- Molecule Fluorescence Toma E. Tomov, Roman Tsukanov, Miran Liber, Rula Masoud, Noa Plavner, and Eyal Nir* Department of
Supramolecular DNA nanotechnology. Faisal A. Aldaye
 Supramolecular DA nanotechnology Faisal A. Aldaye Department of Chemistry, McGill University Current address: Department of Systems Biology, Harvard University 200 Longwood Avenue, Boston, MA 02115, USA
Supramolecular DA nanotechnology Faisal A. Aldaye Department of Chemistry, McGill University Current address: Department of Systems Biology, Harvard University 200 Longwood Avenue, Boston, MA 02115, USA
A Study of DNA Tube Formation Mechanisms Using 4-, 8-, and 12-Helix DNA Nanostructures
 Published on Web 03/02/2006 A Study of DNA Tube Formation Mechanisms Using 4-, 8-, and 12-Helix DNA Nanostructures Yonggang Ke, Yan Liu, Junping Zhang, and Hao Yan* Contribution from the Department of
Published on Web 03/02/2006 A Study of DNA Tube Formation Mechanisms Using 4-, 8-, and 12-Helix DNA Nanostructures Yonggang Ke, Yan Liu, Junping Zhang, and Hao Yan* Contribution from the Department of
Multifactorial Modulation of Binding and Dissociation Kinetics on Two-Dimensional DNA Nanostructures
 pubs.acs.org/nanolett Multifactorial Modulation of Binding and Dissociation Kinetics on Two-Dimensional DNA Nanostructures Alexander Johnson-Buck, Jeanette Nangreave,, Shuoxing Jiang,, Hao Yan,, and Nils
pubs.acs.org/nanolett Multifactorial Modulation of Binding and Dissociation Kinetics on Two-Dimensional DNA Nanostructures Alexander Johnson-Buck, Jeanette Nangreave,, Shuoxing Jiang,, Hao Yan,, and Nils
Overview of New Structures for DNA-Based Nanofabrication and Computation
 Overview of New Structures for DNA-Based Nanofabrication and Computation Thomas H. LaBean, Hao Yan, Sung Ha Park, Liping Feng, Peng Yin, Hanying Li, Sang Jung Ahn, Dage Liu, Xiaoju Guan, and John H. Reif
Overview of New Structures for DNA-Based Nanofabrication and Computation Thomas H. LaBean, Hao Yan, Sung Ha Park, Liping Feng, Peng Yin, Hanying Li, Sang Jung Ahn, Dage Liu, Xiaoju Guan, and John H. Reif
Programmable DNA Self-Assemblies for Nanoscale Organization of Ligands and Proteins
 Programmable DNA Self-Assemblies for Nanoscale Organization of Ligands and Proteins NANO LETTERS 2005 Vol. 5, No. 4 729-733 Sung Ha Park, Peng Yin, Yan Liu, John H. Reif, Thomas H. LaBean,*, and Hao Yan*,
Programmable DNA Self-Assemblies for Nanoscale Organization of Ligands and Proteins NANO LETTERS 2005 Vol. 5, No. 4 729-733 Sung Ha Park, Peng Yin, Yan Liu, John H. Reif, Thomas H. LaBean,*, and Hao Yan*,
Single molecule force spectroscopy reveals a highly compliant helical
 Supplementary Information Single molecule force spectroscopy reveals a highly compliant helical folding for the 30 nm chromatin fiber Maarten Kruithof, Fan-Tso Chien, Andrew Routh, Colin Logie, Daniela
Supplementary Information Single molecule force spectroscopy reveals a highly compliant helical folding for the 30 nm chromatin fiber Maarten Kruithof, Fan-Tso Chien, Andrew Routh, Colin Logie, Daniela
Toward Modular Molecular Composite Nanosystems
 Toward Modular Molecular Composite Nanosystems K. Eric Drexler, PhD U.C. Berkeley 26 April 2009 Intended take-away messages: Paths are now open toward complex, self-assembled, heterogenous nanosystems
Toward Modular Molecular Composite Nanosystems K. Eric Drexler, PhD U.C. Berkeley 26 April 2009 Intended take-away messages: Paths are now open toward complex, self-assembled, heterogenous nanosystems
DNA-Scaffolded Self-Assembling Nano-Circuitry
 DNA-Scaffolded Self-Assembling Nano-Circuitry An Ongoing Research Project with Dr. Soha Hassoun Presentation by Brandon Lucia and Laura Smith DNA-Scaffolded... DNA is special type of molecule Made of a
DNA-Scaffolded Self-Assembling Nano-Circuitry An Ongoing Research Project with Dr. Soha Hassoun Presentation by Brandon Lucia and Laura Smith DNA-Scaffolded... DNA is special type of molecule Made of a
Autonomous Programmable Nanorobotic Devices Using DNAzymes
 Autonomous Programmable Nanorobotic Devices Using DNAzymes John H. Reif and Sudheer Sahu Department of Computer Science, Duke University Box 929, Durham, NC 2778-29, USA {reif,sudheer}@cs.duke.edu Abstract.
Autonomous Programmable Nanorobotic Devices Using DNAzymes John H. Reif and Sudheer Sahu Department of Computer Science, Duke University Box 929, Durham, NC 2778-29, USA {reif,sudheer}@cs.duke.edu Abstract.
Untangling the Mechanics of Entangled Biopolymers
 Untangling the Mechanics of Entangled Biopolymers Rae M. Robertson-Anderson Physics Department University of San Diego students/postdocs: Cole Chapman, PhD Tobias Falzone, PhD Stephanie Gorczyca, USD 16
Untangling the Mechanics of Entangled Biopolymers Rae M. Robertson-Anderson Physics Department University of San Diego students/postdocs: Cole Chapman, PhD Tobias Falzone, PhD Stephanie Gorczyca, USD 16
Active Tile Self Assembly:
 Active Tile Self Assembly: Simulating Cellular Automata at Temperature 1 Daria Karpenko Department of Mathematics and Statistics, University of South Florida Outline Introduction Overview of DNA self-assembly
Active Tile Self Assembly: Simulating Cellular Automata at Temperature 1 Daria Karpenko Department of Mathematics and Statistics, University of South Florida Outline Introduction Overview of DNA self-assembly
The Design and Fabrication of a Fully Addressable 8-tile DNA Lattice
 The Design and Fabrication of a Fully Addressable 8-tile DNA Lattice Chris Dwyer 1,3, Sung Ha Park 2, Thomas LaBean 3, Alvin Lebeck 3 1 Dept. of Electrical and Computer Engineering, Duke University, Durham,
The Design and Fabrication of a Fully Addressable 8-tile DNA Lattice Chris Dwyer 1,3, Sung Ha Park 2, Thomas LaBean 3, Alvin Lebeck 3 1 Dept. of Electrical and Computer Engineering, Duke University, Durham,
Multimedia : Fibronectin and Titin unfolding simulation movies.
 I LECTURE 21: SINGLE CHAIN ELASTICITY OF BIOMACROMOLECULES: THE GIANT PROTEIN TITIN AND DNA Outline : REVIEW LECTURE #2 : EXTENSIBLE FJC AND WLC... 2 STRUCTURE OF MUSCLE AND TITIN... 3 SINGLE MOLECULE
I LECTURE 21: SINGLE CHAIN ELASTICITY OF BIOMACROMOLECULES: THE GIANT PROTEIN TITIN AND DNA Outline : REVIEW LECTURE #2 : EXTENSIBLE FJC AND WLC... 2 STRUCTURE OF MUSCLE AND TITIN... 3 SINGLE MOLECULE
BOOSTER AWARD FINAL REPORT
 PI: Sinan Keten ISEN BOOSTER AWARD FINAL REPORT April 22, 2013 Project: Simulation-based design of organic subnanoporous thin film separation membranes 1. Introduction Polymer thin films employing nanotubes
PI: Sinan Keten ISEN BOOSTER AWARD FINAL REPORT April 22, 2013 Project: Simulation-based design of organic subnanoporous thin film separation membranes 1. Introduction Polymer thin films employing nanotubes
DNA Structure. Voet & Voet: Chapter 29 Pages Slide 1
 DNA Structure Voet & Voet: Chapter 29 Pages 1107-1122 Slide 1 Review The four DNA bases and their atom names The four common -D-ribose conformations All B-DNA ribose adopt the C2' endo conformation All
DNA Structure Voet & Voet: Chapter 29 Pages 1107-1122 Slide 1 Review The four DNA bases and their atom names The four common -D-ribose conformations All B-DNA ribose adopt the C2' endo conformation All
Autonomous DNA Walking Devices
 1 utonomous DN Walking Devices Peng Yin*, ndrew J. Turberfield, Hao Yan*, John H. Reif* * Department of Computer Science, Duke University Department of Physics, Clarendon Laboratory, University of Oxford
1 utonomous DN Walking Devices Peng Yin*, ndrew J. Turberfield, Hao Yan*, John H. Reif* * Department of Computer Science, Duke University Department of Physics, Clarendon Laboratory, University of Oxford
Adaptive Response of Actin Bundles under Mechanical Stress
 Biophysical Journal, Volume 113 Supplemental Information Adaptive Response of Actin Bundles under Mechanical Stress Florian Rückerl, Martin Lenz, Timo Betz, John Manzi, Jean-Louis Martiel, Mahassine Safouane,
Biophysical Journal, Volume 113 Supplemental Information Adaptive Response of Actin Bundles under Mechanical Stress Florian Rückerl, Martin Lenz, Timo Betz, John Manzi, Jean-Louis Martiel, Mahassine Safouane,
Model Worksheet Teacher Key
 Introduction Despite the complexity of life on Earth, the most important large molecules found in all living things (biomolecules) can be classified into only four main categories: carbohydrates, lipids,
Introduction Despite the complexity of life on Earth, the most important large molecules found in all living things (biomolecules) can be classified into only four main categories: carbohydrates, lipids,
Many proteins spontaneously refold into native form in vitro with high fidelity and high speed.
 Macromolecular Processes 20. Protein Folding Composed of 50 500 amino acids linked in 1D sequence by the polypeptide backbone The amino acid physical and chemical properties of the 20 amino acids dictate
Macromolecular Processes 20. Protein Folding Composed of 50 500 amino acids linked in 1D sequence by the polypeptide backbone The amino acid physical and chemical properties of the 20 amino acids dictate
Sec. 2.1 Filaments in the cell 21 PART I - RODS AND ROPES
 Sec. 2.1 Filaments in the cell 21 PART I - RODS AND ROPES Sec. 2.1 Filaments in the cell 22 CHAPTER 2 - POLYMERS The structural elements of the cell can be broadly classified as filaments or sheets, where
Sec. 2.1 Filaments in the cell 21 PART I - RODS AND ROPES Sec. 2.1 Filaments in the cell 22 CHAPTER 2 - POLYMERS The structural elements of the cell can be broadly classified as filaments or sheets, where
Transfer of Chirality from Molecule to Phase in Self-assembled Chiral Block Copolymers
 Transfer of Chirality from Molecule to Phase in Self-assembled Chiral Block Copolymers Rong-Ming Ho,* Ming-Chia Li, Shih-Chieh Lin, Hsiao-Fang Wang, Yu-Der Lee, Hirokazu Hasegawa, and Edwin L. Thomas Supporting
Transfer of Chirality from Molecule to Phase in Self-assembled Chiral Block Copolymers Rong-Ming Ho,* Ming-Chia Li, Shih-Chieh Lin, Hsiao-Fang Wang, Yu-Der Lee, Hirokazu Hasegawa, and Edwin L. Thomas Supporting
DNA nanotechnology-enabled chiral plasmonics: from static to dynamic
 DNA nanotechnology-enabled chiral plasmonics: from static to dynamic Chao Zhou, Xiaoyang Duan,, and Na Liu, * Max Planck Institute for Intelligent Systems, Heisenbergstrasse 3, D-70569 Stuttgart, Germany
DNA nanotechnology-enabled chiral plasmonics: from static to dynamic Chao Zhou, Xiaoyang Duan,, and Na Liu, * Max Planck Institute for Intelligent Systems, Heisenbergstrasse 3, D-70569 Stuttgart, Germany
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Journal of Materials Chemistry A. This journal is The Royal Society of Chemistry 2016 Electronic Supplementary Information Mesoporous C-coated SnO x nanosheets
Electronic Supplementary Material (ESI) for Journal of Materials Chemistry A. This journal is The Royal Society of Chemistry 2016 Electronic Supplementary Information Mesoporous C-coated SnO x nanosheets
M.J. Buehler, Strength in numbers, Nature Nanotechnology, Vol. 5, pp ,
 1 M.J. Buehler, Strength in numbers, Nature Nanotechnology, Vol. 5, pp. 172-174, 2010 NANOMATERIALS Strength in numbers Self-assembly of proteins commonly associated with neurodegenerative diseases can
1 M.J. Buehler, Strength in numbers, Nature Nanotechnology, Vol. 5, pp. 172-174, 2010 NANOMATERIALS Strength in numbers Self-assembly of proteins commonly associated with neurodegenerative diseases can
Science and Technology, Dalian University of Technology, Dalian , P. R. China b
 Electronic Supplementary Information for Fabrication of Superior-Performance SnO 2 @C Composites for Lithium-Ion Anodes Using Tubular Mesoporous Carbons with Thin Carbon Wall and High Pore Volume Fei Han,
Electronic Supplementary Information for Fabrication of Superior-Performance SnO 2 @C Composites for Lithium-Ion Anodes Using Tubular Mesoporous Carbons with Thin Carbon Wall and High Pore Volume Fei Han,
COMPLEX EFFECTS OF MOLECULAR TOPOLOGY, LENGTH AND CONCENTRATION ON MOLECULAR DYNAMICS IN ENTANGLED DNA BLENDS
 COMPLEX EFFECTS OF MOLECULAR TOPOLOGY, LENGTH AND CONCENTRATION ON MOLECULAR DYNAMICS IN ENTANGLED DNA BLENDS Students Cole E. Chapman Kent Lee Dean Henze Collaborators Doug Smith (UCSD) Sachin Shanbhag
COMPLEX EFFECTS OF MOLECULAR TOPOLOGY, LENGTH AND CONCENTRATION ON MOLECULAR DYNAMICS IN ENTANGLED DNA BLENDS Students Cole E. Chapman Kent Lee Dean Henze Collaborators Doug Smith (UCSD) Sachin Shanbhag
In-Situ Fabrication of CoS and NiS Nanomaterials Anchored on. Reduced Graphene Oxide for Reversible Lithium Storage
 Supporting Information In-Situ Fabrication of CoS and NiS Nanomaterials Anchored on Reduced Graphene Oxide for Reversible Lithium Storage Yingbin Tan, [a] Ming Liang, [b, c] Peili Lou, [a] Zhonghui Cui,
Supporting Information In-Situ Fabrication of CoS and NiS Nanomaterials Anchored on Reduced Graphene Oxide for Reversible Lithium Storage Yingbin Tan, [a] Ming Liang, [b, c] Peili Lou, [a] Zhonghui Cui,
Magnetic Torque Tweezers: measuring torsional stiffness in DNA and RecA DNA filaments
 Magnetic Torque Tweezers: measuring torsional stiffness in DNA and RecA DNA filaments Lipfert, J., Kerssemakers, J. W., Jager, T. & Dekker, N. H. Magnetic torque tweezers: measuring torsional stiffness
Magnetic Torque Tweezers: measuring torsional stiffness in DNA and RecA DNA filaments Lipfert, J., Kerssemakers, J. W., Jager, T. & Dekker, N. H. Magnetic torque tweezers: measuring torsional stiffness
Supplementary Information
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2014 Supplementary Information Large-scale lithography-free metasurface with spectrally tunable super
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2014 Supplementary Information Large-scale lithography-free metasurface with spectrally tunable super
Integer and Vector Multiplication by Using DNA
 Int. J. Open Problems Compt. Math., Vol. 1, No. 2, September 2008 Integer and Vector Multiplication by Using DNA Essam Al-Daoud 1, Belal Zaqaibeh 2, and Feras Al-Hanandeh 3 1 Computer Science Department,
Int. J. Open Problems Compt. Math., Vol. 1, No. 2, September 2008 Integer and Vector Multiplication by Using DNA Essam Al-Daoud 1, Belal Zaqaibeh 2, and Feras Al-Hanandeh 3 1 Computer Science Department,
Engineering of Hollow Core-Shell Interlinked Carbon Spheres for Highly Stable Lithium-Sulfur Batteries
 SUPPLEMENTARY INFORMATION Engineering of Hollow Core-Shell Interlinked Carbon Spheres for Highly Stable Lithium-Sulfur Batteries Qiang Sun, Bin He, Xiang-Qian Zhang, and An-Hui Lu* State Key Laboratory
SUPPLEMENTARY INFORMATION Engineering of Hollow Core-Shell Interlinked Carbon Spheres for Highly Stable Lithium-Sulfur Batteries Qiang Sun, Bin He, Xiang-Qian Zhang, and An-Hui Lu* State Key Laboratory
Investigation of dsdna Stretching Meso-Mechanics Using LS-DYNA
 8 th International LS-DYNA Users Conference Simulation echnology (3) Investigation of dsdna Stretching Meso-Mechanics Using LS-DYNA C. A. Yuan and K. N. Chiang Department of Power Mechanical Engineering,
8 th International LS-DYNA Users Conference Simulation echnology (3) Investigation of dsdna Stretching Meso-Mechanics Using LS-DYNA C. A. Yuan and K. N. Chiang Department of Power Mechanical Engineering,
Magnetic tweezers and its application to DNA mechanics
 Biotechnological Center Research group DNA motors (Seidel group) Handout for Practical Course Magnetic tweezers and its application to DNA mechanics When: 9.00 am Where: Biotec, 3 rd Level, Room 317 Tutors:
Biotechnological Center Research group DNA motors (Seidel group) Handout for Practical Course Magnetic tweezers and its application to DNA mechanics When: 9.00 am Where: Biotec, 3 rd Level, Room 317 Tutors:
Ionic Permeability and Mechanical Properties of DNA Origami Nanoplates on Solid-State Nanopores
 Ionic Permeability and Mechanical Properties of DNA Origami Nanoplates on Solid-State Nanopores Calin Plesa, Adithya N. Ananth, Veikko Linko, Coen Gülcher, Allard J. Katan, Hendrik Dietz, and Cees Dekker,
Ionic Permeability and Mechanical Properties of DNA Origami Nanoplates on Solid-State Nanopores Calin Plesa, Adithya N. Ananth, Veikko Linko, Coen Gülcher, Allard J. Katan, Hendrik Dietz, and Cees Dekker,
Elasticity of biological gels
 Seminar II Elasticity of biological gels Author: Gašper Gregorič Mentor: assoc. prof. Primož Ziherl Ljubljana, February 2014 Abstract In the seminar we discuss the elastic behavior of biological gels,
Seminar II Elasticity of biological gels Author: Gašper Gregorič Mentor: assoc. prof. Primož Ziherl Ljubljana, February 2014 Abstract In the seminar we discuss the elastic behavior of biological gels,
SECOND PUBLIC EXAMINATION. Honour School of Physics Part C: 4 Year Course. Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS
 2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2013 Monday, 17 June, 2.30 pm 5.45 pm 15
2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2013 Monday, 17 June, 2.30 pm 5.45 pm 15
3 Biopolymers Uncorrelated chains - Freely jointed chain model
 3.4 Entropy When we talk about biopolymers, it is important to realize that the free energy of a biopolymer in thermal equilibrium is not constant. Unlike solids, biopolymers are characterized through
3.4 Entropy When we talk about biopolymers, it is important to realize that the free energy of a biopolymer in thermal equilibrium is not constant. Unlike solids, biopolymers are characterized through
small
 FULL PAPER DNA Origami Rhombic-Shaped Nanostructures and Mechanical Properties of 2D DNA Origami Constructed with Different Crossover/Nick Designs Zhipeng Ma,* Yunfei Huang, Seongsu Park, Kentaro Kawai,
FULL PAPER DNA Origami Rhombic-Shaped Nanostructures and Mechanical Properties of 2D DNA Origami Constructed with Different Crossover/Nick Designs Zhipeng Ma,* Yunfei Huang, Seongsu Park, Kentaro Kawai,
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/314/5801/1001/dc1 Supporting Online Material for Direct Measurement of the Full, Sequence-Dependent Folding Landscape of a Nucleic Acid Michael T. Woodside, Peter C.
www.sciencemag.org/cgi/content/full/314/5801/1001/dc1 Supporting Online Material for Direct Measurement of the Full, Sequence-Dependent Folding Landscape of a Nucleic Acid Michael T. Woodside, Peter C.
Advanced Drug Delivery Reviews
 Advanced Drug Delivery Reviews 62 (2010) 617 625 Contents lists available at ScienceDirect Advanced Drug Delivery Reviews journal homepage: www.elsevier.com/locate/addr DNA Self-assembly for Nanomedicine
Advanced Drug Delivery Reviews 62 (2010) 617 625 Contents lists available at ScienceDirect Advanced Drug Delivery Reviews journal homepage: www.elsevier.com/locate/addr DNA Self-assembly for Nanomedicine
Members Subjected to Torsional Loads
 Members Subjected to Torsional Loads Torsion of circular shafts Definition of Torsion: Consider a shaft rigidly clamped at one end and twisted at the other end by a torque T = F.d applied in a plane perpendicular
Members Subjected to Torsional Loads Torsion of circular shafts Definition of Torsion: Consider a shaft rigidly clamped at one end and twisted at the other end by a torque T = F.d applied in a plane perpendicular
Table of contents. Saturday 24 June Sunday 25 June Monday 26 June
 Table of contents Saturday 24 June 2017... 1 Sunday 25 June 2017... 3 Monday 26 June 2017... 5 i Saturday 24 June 2017 Quantitative Biology 2017: Computational and Single-Molecule Biophysics Saturday 24
Table of contents Saturday 24 June 2017... 1 Sunday 25 June 2017... 3 Monday 26 June 2017... 5 i Saturday 24 June 2017 Quantitative Biology 2017: Computational and Single-Molecule Biophysics Saturday 24
Bio-inspired self-assembly. Anant K. Paravastu Department of Chemical and Biomedical Engineering Florida A&M University and Florida State University
 Bio-inspired self-assembly Anant K. Paravastu Department of Chemical and Biomedical Engineering Florida A&M University and Florida State University Self-assembly in biology Multi-scale extracellular matrices
Bio-inspired self-assembly Anant K. Paravastu Department of Chemical and Biomedical Engineering Florida A&M University and Florida State University Self-assembly in biology Multi-scale extracellular matrices
Nanotechnology? Source: National Science Foundation (NSF), USA
 2 2 Nanotechnology? Ability to work at the atomic, molecular and even sub-molecular levels in order to create and use material structures, devices and systems with new properties and functions Source:
2 2 Nanotechnology? Ability to work at the atomic, molecular and even sub-molecular levels in order to create and use material structures, devices and systems with new properties and functions Source:
Supplementary Information. The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes
 Supplementary Information The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes Xuesong Shi, a Lin Huang, b David M. J. Lilley, b Pehr B. Harbury a,c and Daniel Herschlag
Supplementary Information The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes Xuesong Shi, a Lin Huang, b David M. J. Lilley, b Pehr B. Harbury a,c and Daniel Herschlag
Design, Simulation, and Experimental Demonstration of Self-assembled DNA Nanostructures and Motors
 Design, Simulation, and Experimental Demonstration of Self-assembled DNA Nanostructures and Motors John H. Reif, Thomas H. LaBean, Sudheer Sahu, Hao Yan, and Peng Yin Department of Computer Science, Duke
Design, Simulation, and Experimental Demonstration of Self-assembled DNA Nanostructures and Motors John H. Reif, Thomas H. LaBean, Sudheer Sahu, Hao Yan, and Peng Yin Department of Computer Science, Duke
PROCEEDINGS OF SPIE. Nanoparticle sorting in silicon waveguide arrays. H. T. Zhao, Y. Zhang, L. K. Chin, P. H. Yap, K. Wang, et al.
 PROCEEDINGS OF SPIE SPIEDigitalLibrary.org/conference-proceedings-of-spie Nanoparticle sorting in silicon waveguide arrays H. T. Zhao, Y. Zhang, L. K. Chin, P. H. Yap, K. Wang, et al. H. T. Zhao, Y. Zhang,
PROCEEDINGS OF SPIE SPIEDigitalLibrary.org/conference-proceedings-of-spie Nanoparticle sorting in silicon waveguide arrays H. T. Zhao, Y. Zhang, L. K. Chin, P. H. Yap, K. Wang, et al. H. T. Zhao, Y. Zhang,
Chapter 1. DNA is made from the building blocks adenine, guanine, cytosine, and. Answer: d
 Chapter 1 1. Matching Questions DNA is made from the building blocks adenine, guanine, cytosine, and. Answer: d 2. Matching Questions : Unbranched polymer that, when folded into its three-dimensional shape,
Chapter 1 1. Matching Questions DNA is made from the building blocks adenine, guanine, cytosine, and. Answer: d 2. Matching Questions : Unbranched polymer that, when folded into its three-dimensional shape,
Novel Nanoparticles for Ultrasensitive Detection and Spectroscopy
 Final Technical Report (DOE-FG02-98ER14873) Project Officer: Dr. Richard Gordon / Dr. John Miller Novel Nanoparticles for Ultrasensitive Detection and Spectroscopy Shuming Nie Indiana University P. 0.
Final Technical Report (DOE-FG02-98ER14873) Project Officer: Dr. Richard Gordon / Dr. John Miller Novel Nanoparticles for Ultrasensitive Detection and Spectroscopy Shuming Nie Indiana University P. 0.
An Advanced Anode Material for Sodium Ion. Batteries
 Layered-Structure SbPO 4 /Reduced Graphene Oxide: An Advanced Anode Material for Sodium Ion Batteries Jun Pan, Shulin Chen, # Qiang Fu, Yuanwei Sun, # Yuchen Zhang, Na Lin, Peng Gao,* # Jian Yang,* and
Layered-Structure SbPO 4 /Reduced Graphene Oxide: An Advanced Anode Material for Sodium Ion Batteries Jun Pan, Shulin Chen, # Qiang Fu, Yuanwei Sun, # Yuchen Zhang, Na Lin, Peng Gao,* # Jian Yang,* and
Supplementary Figures
 Fracture Strength (GPa) Supplementary Figures a b 10 R=0.88 mm 1 0.1 Gordon et al Zhu et al Tang et al im et al 5 7 6 4 This work 5 50 500 Si Nanowire Diameter (nm) Supplementary Figure 1: (a) TEM image
Fracture Strength (GPa) Supplementary Figures a b 10 R=0.88 mm 1 0.1 Gordon et al Zhu et al Tang et al im et al 5 7 6 4 This work 5 50 500 Si Nanowire Diameter (nm) Supplementary Figure 1: (a) TEM image
Terahertz Wave Propagation in a Nanotube Conveying Fluid Taking into Account Surface Effect
 Materials 13, 6, 393-399; doi:1.339/ma66393 Article OPEN ACCE materials IN 1996-1944 www.mdpi.com/journal/materials Terahertz Wave Propagation in a Nanotube Conveying Fluid Taking into Account urface Effect
Materials 13, 6, 393-399; doi:1.339/ma66393 Article OPEN ACCE materials IN 1996-1944 www.mdpi.com/journal/materials Terahertz Wave Propagation in a Nanotube Conveying Fluid Taking into Account urface Effect
Self-Assembly of DNA Arrays into Multilayer Stacks
 Article Subscriber access provided by HOUSTON ACADEMY OF MEDICINE Self-Assembly of DNA Arrays into Multilayer Stacks Alexey Y. Koyfman, Sergei N. Magonov, and Norbert O. Reich Langmuir, 2009, 25 (2), 1091-1096
Article Subscriber access provided by HOUSTON ACADEMY OF MEDICINE Self-Assembly of DNA Arrays into Multilayer Stacks Alexey Y. Koyfman, Sergei N. Magonov, and Norbert O. Reich Langmuir, 2009, 25 (2), 1091-1096
bifunctional electrocatalyst for overall water splitting
 Electronic Supplementary Material (ESI) for Journal of Materials Chemistry A. This journal is The Royal Society of Chemistry 2017 Hierarchical Ni/NiTiO 3 derived from NiTi LDHs: a bifunctional electrocatalyst
Electronic Supplementary Material (ESI) for Journal of Materials Chemistry A. This journal is The Royal Society of Chemistry 2017 Hierarchical Ni/NiTiO 3 derived from NiTi LDHs: a bifunctional electrocatalyst
Tuning the Shell Number of Multi-Shelled Metal Oxide. Hollow Fibers for Optimized Lithium Ion Storage
 Supporting Information Tuning the Shell Number of Multi-Shelled Metal Oxide Hollow Fibers for Optimized Lithium Ion Storage Jin Sun, Chunxiao Lv, Fan Lv, ǁ Shuai Chen, Daohao Li, Ziqi Guo, Wei Han, Dongjiang
Supporting Information Tuning the Shell Number of Multi-Shelled Metal Oxide Hollow Fibers for Optimized Lithium Ion Storage Jin Sun, Chunxiao Lv, Fan Lv, ǁ Shuai Chen, Daohao Li, Ziqi Guo, Wei Han, Dongjiang
Solution-Controlled Conformational Switching of an Anchored Wireframe DNA Nanostructure
 FULL PAPER DNA Origami Solution-Controlled Conformational Switching of an Anchored Wireframe DNA Nanostructure Ian T. Hoffecker, Sijie Chen, Andreas Gådin, Alessandro Bosco, Ana I. Teixeira, and Björn
FULL PAPER DNA Origami Solution-Controlled Conformational Switching of an Anchored Wireframe DNA Nanostructure Ian T. Hoffecker, Sijie Chen, Andreas Gådin, Alessandro Bosco, Ana I. Teixeira, and Björn
Kinetically-Enhanced Polysulfide Redox Reactions by Nb2O5. Nanocrystal for High-Rate Lithium Sulfur Battery
 Electronic Supplementary Material (ESI) for Energy & Environmental Science. This journal is The Royal Society of Chemistry 2016 Electronic Supplementary Information (ESI) Kinetically-Enhanced Polysulfide
Electronic Supplementary Material (ESI) for Energy & Environmental Science. This journal is The Royal Society of Chemistry 2016 Electronic Supplementary Information (ESI) Kinetically-Enhanced Polysulfide
Nanopores and Nanofluidics for Single DNA Studies Derek Stein Department of Physics Brown University
 Nanopores and Nanofluidics for Single DNA Studies Derek Stein Department of Physics Brown University Overview Motivation: biological, physical, and technological The program: develop and characterize new
Nanopores and Nanofluidics for Single DNA Studies Derek Stein Department of Physics Brown University Overview Motivation: biological, physical, and technological The program: develop and characterize new
Sophie Dumont BP204 March
 Sophie Dumont BP204 March 9 2015 1 Molecules Mechanics What can we learn? Tools? Framework? Challenges? Looking ahead? structure, chemistry Cells 2 What biological activities are associated with force?
Sophie Dumont BP204 March 9 2015 1 Molecules Mechanics What can we learn? Tools? Framework? Challenges? Looking ahead? structure, chemistry Cells 2 What biological activities are associated with force?
Introduction to MD simulations of DNA systems. Aleksei Aksimentiev
 Introduction to MD simulations of DNA systems Aleksei Aksimentiev Biological Modeling at Different Scales spanning orders of magnitude in space and time The computational microscope Massive parallel computer
Introduction to MD simulations of DNA systems Aleksei Aksimentiev Biological Modeling at Different Scales spanning orders of magnitude in space and time The computational microscope Massive parallel computer
Lecture 4: viscoelasticity and cell mechanics
 Teaser movie: flexible robots! R. Shepherd, Whitesides group, Harvard 1 Lecture 4: viscoelasticity and cell mechanics S-RSI Physics Lectures: Soft Condensed Matter Physics Jacinta C. Conrad University
Teaser movie: flexible robots! R. Shepherd, Whitesides group, Harvard 1 Lecture 4: viscoelasticity and cell mechanics S-RSI Physics Lectures: Soft Condensed Matter Physics Jacinta C. Conrad University
Transport of single molecules along the periodic parallel lattices with coupling
 THE JOURNAL OF CHEMICAL PHYSICS 124 204901 2006 Transport of single molecules along the periodic parallel lattices with coupling Evgeny B. Stukalin The James Franck Institute The University of Chicago
THE JOURNAL OF CHEMICAL PHYSICS 124 204901 2006 Transport of single molecules along the periodic parallel lattices with coupling Evgeny B. Stukalin The James Franck Institute The University of Chicago
Stress in Flip-Chip Solder Bumps due to Package Warpage -- Matt Pharr
 Stress in Flip-Chip Bumps due to Package Warpage -- Matt Pharr Introduction As the size of microelectronic devices continues to decrease, interconnects in the devices are scaling down correspondingly.
Stress in Flip-Chip Bumps due to Package Warpage -- Matt Pharr Introduction As the size of microelectronic devices continues to decrease, interconnects in the devices are scaling down correspondingly.
Self-Assembled DNA Hydrogels with Designable Thermal and Enzymatic Responsiveness
 Self-Assembled DNA Hydrogels with Designable Thermal and Enzymatic Responsiveness Yongzheng Xing, Enjun Cheng, Yang Yang, Ping Chen, Tao Zhang, Yawei Sun, Zhongqiang Yang, and Dongsheng Liu * DNA has received
Self-Assembled DNA Hydrogels with Designable Thermal and Enzymatic Responsiveness Yongzheng Xing, Enjun Cheng, Yang Yang, Ping Chen, Tao Zhang, Yawei Sun, Zhongqiang Yang, and Dongsheng Liu * DNA has received
Supporting Infromation
 Supporting Infromation Transparent and Flexible Self-Charging Power Film and Its Application in Sliding-Unlock System in Touchpad Technology Jianjun Luo 1,#, Wei Tang 1,#, Feng Ru Fan 1, Chaofeng Liu 1,
Supporting Infromation Transparent and Flexible Self-Charging Power Film and Its Application in Sliding-Unlock System in Touchpad Technology Jianjun Luo 1,#, Wei Tang 1,#, Feng Ru Fan 1, Chaofeng Liu 1,
Structural DNA nanotechnology designs nanostructures. Article
 Observing and Controlling the Folding Pathway of DNA Origami at the Nanoscale Jonathan Lee Tin Wah,, Christophe David, Sergii Rudiuk,,,# Damien Baigl,,,# and Andre Estevez-Torres*,, Laboratoire Jean Perrin,
Observing and Controlling the Folding Pathway of DNA Origami at the Nanoscale Jonathan Lee Tin Wah,, Christophe David, Sergii Rudiuk,,,# Damien Baigl,,,# and Andre Estevez-Torres*,, Laboratoire Jean Perrin,
Simulations of Self-Assembly of Polypeptide-Based Copolymers
 East China University of Science and Technology Theory, Algorithms and Applications of Dissipative Particle Dynamics Simulations of Self-Assembly of Polypeptide-Based Copolymers Jiaping LIN ( 林嘉平 ) East
East China University of Science and Technology Theory, Algorithms and Applications of Dissipative Particle Dynamics Simulations of Self-Assembly of Polypeptide-Based Copolymers Jiaping LIN ( 林嘉平 ) East
Supporting Information
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2015 Supporting Information Synthesis and electrochemical properties of spherical and hollow-structured
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2015 Supporting Information Synthesis and electrochemical properties of spherical and hollow-structured
DNA Computing by Self Assembly. Presented by Mohammed Ashraf Ali
 DNA Computing by Self Assembly Presented by Mohammed Ashraf Ali Outline Self- Assembly Examples and behaviour of self-assembly DNA Computing Tiling Theory Nanotechnology DNA Self-assembly Visualization
DNA Computing by Self Assembly Presented by Mohammed Ashraf Ali Outline Self- Assembly Examples and behaviour of self-assembly DNA Computing Tiling Theory Nanotechnology DNA Self-assembly Visualization
F. Piazza Center for Molecular Biophysics and University of Orléans, France. Selected topic in Physical Biology. Lecture 1
 Zhou Pei-Yuan Centre for Applied Mathematics, Tsinghua University November 2013 F. Piazza Center for Molecular Biophysics and University of Orléans, France Selected topic in Physical Biology Lecture 1
Zhou Pei-Yuan Centre for Applied Mathematics, Tsinghua University November 2013 F. Piazza Center for Molecular Biophysics and University of Orléans, France Selected topic in Physical Biology Lecture 1
Prediction of Elastic Constants on 3D Four-directional Braided
 Prediction of Elastic Constants on 3D Four-directional Braided Composites Prediction of Elastic Constants on 3D Four-directional Braided Composites Liang Dao Zhou 1,2,* and Zhuo Zhuang 1 1 School of Aerospace,
Prediction of Elastic Constants on 3D Four-directional Braided Composites Prediction of Elastic Constants on 3D Four-directional Braided Composites Liang Dao Zhou 1,2,* and Zhuo Zhuang 1 1 School of Aerospace,
Quantitative Analysis of Forces in Cells
 Quantitative Analysis of Forces in Cells Anders Carlsson Washington University in St Louis Basic properties of forces in cells Measurement methods and magnitudes of particular types of forces: Polymerization
Quantitative Analysis of Forces in Cells Anders Carlsson Washington University in St Louis Basic properties of forces in cells Measurement methods and magnitudes of particular types of forces: Polymerization
Supporting Information
 Supporting Information MoSe2 embedded CNT-Reduced Graphene Oxide (rgo) Composite Microsphere with Superior Sodium Ion Storage and Electrocatalytic Hydrogen Evolution Performances Gi Dae Park, Jung Hyun
Supporting Information MoSe2 embedded CNT-Reduced Graphene Oxide (rgo) Composite Microsphere with Superior Sodium Ion Storage and Electrocatalytic Hydrogen Evolution Performances Gi Dae Park, Jung Hyun
There have been some notable successes
 B IOCOMPUTATION DNA LATTICES: A METHODFOR MOLECULAR-SCALE PATTERNING AND COMPUTATION DNA lattice research can provide unprecedented capabilities for molecular-scale computation and programmable pattern
B IOCOMPUTATION DNA LATTICES: A METHODFOR MOLECULAR-SCALE PATTERNING AND COMPUTATION DNA lattice research can provide unprecedented capabilities for molecular-scale computation and programmable pattern
Anatoly B. Kolomeisky. Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS?
 Anatoly B. Kolomeisky Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS? Motor Proteins Enzymes that convert the chemical energy into mechanical
Anatoly B. Kolomeisky Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS? Motor Proteins Enzymes that convert the chemical energy into mechanical
Supplementary Information
 Supplementary Information Mechanical unzipping and rezipping of a single SNARE complex reveals hysteresis as a force-generating mechanism Duyoung Min, Kipom Kim, Changbong Hyeon, Yong Hoon Cho, Yeon-Kyun
Supplementary Information Mechanical unzipping and rezipping of a single SNARE complex reveals hysteresis as a force-generating mechanism Duyoung Min, Kipom Kim, Changbong Hyeon, Yong Hoon Cho, Yeon-Kyun
New Measurements of DNA Twist Elasticity
 University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 5-1998 New Measurements of DNA Twist Elasticity Philip C. Nelson University of Pennsylvania, nelson@physics.upenn.edu
University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 5-1998 New Measurements of DNA Twist Elasticity Philip C. Nelson University of Pennsylvania, nelson@physics.upenn.edu
Structural investigation of single biomolecules
 Structural investigation of single biomolecules NMR spectroscopy and X-ray crystallography are currently the most common techniques capable of determining the structures of biological macromolecules like
Structural investigation of single biomolecules NMR spectroscopy and X-ray crystallography are currently the most common techniques capable of determining the structures of biological macromolecules like
Supporting Information. Co 4 N Nanosheets Assembled Mesoporous Sphere as a Matrix for Ultrahigh Sulfur Content Lithium Sulfur Batteries
 Supporting Information Co 4 N Nanosheets Assembled Mesoporous Sphere as a Matrix for Ultrahigh Sulfur Content Lithium Sulfur Batteries Ding-Rong Deng, Fei Xue, Yue-Ju Jia, Jian-Chuan Ye, Cheng-Dong Bai,
Supporting Information Co 4 N Nanosheets Assembled Mesoporous Sphere as a Matrix for Ultrahigh Sulfur Content Lithium Sulfur Batteries Ding-Rong Deng, Fei Xue, Yue-Ju Jia, Jian-Chuan Ye, Cheng-Dong Bai,
Large-Area and Uniform Surface-Enhanced Raman. Saturation
 Supporting Information Large-Area and Uniform Surface-Enhanced Raman Spectroscopy Substrate Optimized by Enhancement Saturation Daejong Yang 1, Hyunjun Cho 2, Sukmo Koo 1, Sagar R. Vaidyanathan 2, Kelly
Supporting Information Large-Area and Uniform Surface-Enhanced Raman Spectroscopy Substrate Optimized by Enhancement Saturation Daejong Yang 1, Hyunjun Cho 2, Sukmo Koo 1, Sagar R. Vaidyanathan 2, Kelly
Nanochannel-Assisted Perovskite Nanowires: Growth Mechanisms. to Photodetector Applications
 Supplementary Information: Nanochannel-Assisted Perovskite Nanowires: Growth Mechanisms to Photodetector Applications Qitao Zhou, Jun Gyu Park, Riming Nie, Ashish Kumar Thokchom, Dogyeong Ha, Jing Pan,
Supplementary Information: Nanochannel-Assisted Perovskite Nanowires: Growth Mechanisms to Photodetector Applications Qitao Zhou, Jun Gyu Park, Riming Nie, Ashish Kumar Thokchom, Dogyeong Ha, Jing Pan,
QUESTION BANK SEMESTER: III SUBJECT NAME: MECHANICS OF SOLIDS
 QUESTION BANK SEMESTER: III SUBJECT NAME: MECHANICS OF SOLIDS UNIT 1- STRESS AND STRAIN PART A (2 Marks) 1. Define longitudinal strain and lateral strain. 2. State Hooke s law. 3. Define modular ratio,
QUESTION BANK SEMESTER: III SUBJECT NAME: MECHANICS OF SOLIDS UNIT 1- STRESS AND STRAIN PART A (2 Marks) 1. Define longitudinal strain and lateral strain. 2. State Hooke s law. 3. Define modular ratio,
Supramolecular Chemistry of Nanomaterials
 Supramolecular Chemistry of Nanomaterials Joachim Steinke Ramon Vilar Lecture 6 Towards the Development of Molecular Machines Department of Chemistry Imperial College of Science, Technology and Medicine
Supramolecular Chemistry of Nanomaterials Joachim Steinke Ramon Vilar Lecture 6 Towards the Development of Molecular Machines Department of Chemistry Imperial College of Science, Technology and Medicine
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09450 Supplementary Table 1 Summary of kinetic parameters. Kinetic parameters were V = V / 1 K / ATP and obtained using the relationships max ( + m [ ]) V d s /( 1/ k [ ATP] + 1 k ) =,
doi:10.1038/nature09450 Supplementary Table 1 Summary of kinetic parameters. Kinetic parameters were V = V / 1 K / ATP and obtained using the relationships max ( + m [ ]) V d s /( 1/ k [ ATP] + 1 k ) =,
DNA-based Self-Assembly of Chiral Plasmonic Nanostructures with Tailored Optical Response
 1 DNA-based Self-Assembly of Chiral Plasmonic Nanostructures with Tailored Optical Response Anton Kuzyk 1,*, Robert Schreiber 2,*, Zhiyuan Fan 3, Günther Pardatscher 1, Eva Maria Roller 2, Alexander Högele
1 DNA-based Self-Assembly of Chiral Plasmonic Nanostructures with Tailored Optical Response Anton Kuzyk 1,*, Robert Schreiber 2,*, Zhiyuan Fan 3, Günther Pardatscher 1, Eva Maria Roller 2, Alexander Högele
Villa et al. (2005) Structural dynamics of the lac repressor-dna complex revealed by a multiscale simulation. PNAS 102:
 Villa et al. (2005) Structural dynamics of the lac repressor-dna complex revealed by a multiscale simulation. PNAS 102: 6783-6788. Background: The lac operon is a cluster of genes in the E. coli genome
Villa et al. (2005) Structural dynamics of the lac repressor-dna complex revealed by a multiscale simulation. PNAS 102: 6783-6788. Background: The lac operon is a cluster of genes in the E. coli genome
MOLECULAR, CELLULAR, & TISSUE BIOMECHANICS
 MOLECULAR, CELLULAR, & TISSUE BIOMECHANICS Spring 2015 Problem Set #6 - Cytoskeleton mechanics Distributed: Wednesday, April 15, 2015 Due: Thursday, April 23, 2015 Problem 1: Transmigration A critical
MOLECULAR, CELLULAR, & TISSUE BIOMECHANICS Spring 2015 Problem Set #6 - Cytoskeleton mechanics Distributed: Wednesday, April 15, 2015 Due: Thursday, April 23, 2015 Problem 1: Transmigration A critical
The Chinese University of Hong Kong Department of Chemistry
 Prof. Yue Zhao University of Sherbrooke Canada Control of Stimult-Responsive Polymers by New Methods December 1, 2014 (Monday) 2:30 p.m. Room G35 Lady Shaw Building Prof. Chi Wu Prof. Petr Štěpánek Institute
Prof. Yue Zhao University of Sherbrooke Canada Control of Stimult-Responsive Polymers by New Methods December 1, 2014 (Monday) 2:30 p.m. Room G35 Lady Shaw Building Prof. Chi Wu Prof. Petr Štěpánek Institute
Supporting Information for
 Supporting Information for Multilayer CuO@NiO Hollow Spheres: Microwave-Assisted Metal-Organic-Framework Derivation and Highly Reversible Structure-Matched Stepwise Lithium Storage Wenxiang Guo, Weiwei
Supporting Information for Multilayer CuO@NiO Hollow Spheres: Microwave-Assisted Metal-Organic-Framework Derivation and Highly Reversible Structure-Matched Stepwise Lithium Storage Wenxiang Guo, Weiwei
micromachines ISSN X
 Micromachines 2014, 5, 359-372; doi:10.3390/mi5020359 Article OPEN ACCESS micromachines ISSN 2072-666X www.mdpi.com/journal/micromachines Laser Micro Bending Process of Ti6Al4V Square Bar Gang Chen and
Micromachines 2014, 5, 359-372; doi:10.3390/mi5020359 Article OPEN ACCESS micromachines ISSN 2072-666X www.mdpi.com/journal/micromachines Laser Micro Bending Process of Ti6Al4V Square Bar Gang Chen and
Active Plasmonic Nanostructures in Biosensing and Imaging. Bjoern M. Reinhard Department of Chemistry
 Active Plasmonic Nanostructures in Biosensing and Imaging Bjoern M. Reinhard Department of Chemistry Noble Metal Nanoparticles Light The alternating surface charges effectively form an oscillating dipole,
Active Plasmonic Nanostructures in Biosensing and Imaging Bjoern M. Reinhard Department of Chemistry Noble Metal Nanoparticles Light The alternating surface charges effectively form an oscillating dipole,
NANOMEDICINE. WILEY A John Wiley and Sons, Ltd., Publication DESIGN AND APPLICATIONS OF MAGNETIC NANOMATERIALS, NANOSENSORS AND NANOSYSTEMS
 NANOMEDICINE DESIGN AND APPLICATIONS OF MAGNETIC NANOMATERIALS, NANOSENSORS AND NANOSYSTEMS Vijay K. Varadan Linfeng Chen Jining Xie WILEY A John Wiley and Sons, Ltd., Publication Preface About the Authors
NANOMEDICINE DESIGN AND APPLICATIONS OF MAGNETIC NANOMATERIALS, NANOSENSORS AND NANOSYSTEMS Vijay K. Varadan Linfeng Chen Jining Xie WILEY A John Wiley and Sons, Ltd., Publication Preface About the Authors
Nuclear import of DNA: genetic modification of plants
 Nuclear import of DNA: genetic modification of plants gene delivery by Agrobacterium tumefaciens T. Tzfira & V. Citovsky. 2001. Trends in Cell Biol. 12: 121-129 VirE2 binds ssdna in vitro forms helical
Nuclear import of DNA: genetic modification of plants gene delivery by Agrobacterium tumefaciens T. Tzfira & V. Citovsky. 2001. Trends in Cell Biol. 12: 121-129 VirE2 binds ssdna in vitro forms helical
Nanoparticles, nanorods, nanowires
 Nanoparticles, nanorods, nanowires Nanoparticles, nanocrystals, nanospheres, quantum dots, etc. Drugs, proteins, etc. Nanorods, nanowires. Optical and electronic properties. Organization using biomolecules.
Nanoparticles, nanorods, nanowires Nanoparticles, nanocrystals, nanospheres, quantum dots, etc. Drugs, proteins, etc. Nanorods, nanowires. Optical and electronic properties. Organization using biomolecules.
arxiv: v1 [physics.optics] 21 Nov 2011
![arxiv: v1 [physics.optics] 21 Nov 2011 arxiv: v1 [physics.optics] 21 Nov 2011](/thumbs/85/92542022.jpg) Polarization manipulation holographic lithography by single refracting prism Yi Xu, Man Wu, Xiuming Lan, Xiaoxu Lu,Sheng Lan and Lijun Wu arxiv:1111.4860v1 [physics.optics] 21 Nov 2011 Laboratory of Photonic
Polarization manipulation holographic lithography by single refracting prism Yi Xu, Man Wu, Xiuming Lan, Xiaoxu Lu,Sheng Lan and Lijun Wu arxiv:1111.4860v1 [physics.optics] 21 Nov 2011 Laboratory of Photonic
