Calculated enhancement of open base pair probability downstream
|
|
|
- Stuart Underwood
- 6 years ago
- Views:
Transcription
1 Calculated enhancement of open base pair probability downstream of a (TATA)2 box R. Beger and E. W. Prohofsky Department of Physics, Purdue University, West Lafayette, Indiana ABSTRACT The modified self-consistent phonon approximation is generalized to calculate the open base pair probability of a (TATA)2 insert between two semi-infinite poly(da-dc) poly(dg-dt) helices. An iterative method based entirely on the Green's function method is developed to compute the open interbase hydrogen bond probability of the insert and near bases versus temperature. The open interbase hydrogen bond probability is calculated for two temperatures and compared with perfect homopolymer open base pair probabilities at room temperature. The probability of opening is enhanced by a factor of two for the major groove bond of the AT pair at the transcriptional downstream end of the (TATA)2 insert. The total base pair opening probability of that pair is enhanced by 54%. The probability of the next inline GC base pair to be open is increased by a factor of ten. INTRODUCTION Back in 1910 Lindeman (1) first correlated melting with thermal vibrational motion in his published melting formula. The generalization, the Lindeman criterion has remained a valid phenomenological predictor of melting ever since. The essence of the criterion is that all solids melt when the vibrational amplitude of a bond between atoms reaches = 10% of the bond length. This implication is that thermal motion ultimately leads to atoms escaping the binding associated with rigidly bound structures. We have developed a quantitative theory for bond breaking and melting of the DNA helix. We do find that the interstrand hydrogen bonds are disrupted when the thermal motion along these bonds reaches = 10% of the total hydrogen bond length. This bond separation is not to be confused with helix unwinding which obviously requires additional motion. The melting we refer to is just the breakdown of the rigid bonding associated with the structural models of DNA. This type of DNA melting seems not to be unique in any way and is in line with observations on other materials and the Lindeman criterion. There have been many observations by both Raman scattering and infrared radiation (i.r.) absorption of vibrational motion assigned to motion along the interbase hydrogen bonds (2-5). The principal such band = 85 cm-1 has been shown to be intrahelical as the Raman curves are identical for DNA ranging from those dissolved in dilute solution, fibers with widely ranging hydration levels, nucleosome core particles, to unseparated chromatin (6). The band is seen to soften with increasing temperature and disappear at the melting temperature (2). The band is resonant (not overdamped) and its lifetime has been studied in some detail based on observed line widths (7). In line with these observations of resonant bands in DNA we have developed an effective harmonic theory of vibrational motion for the DNA helix. The theory is based on a modified self-consistent phonon approximation (MSPA) (8). The method predicted the 85-cm-l band (9) and predicts its observed temperature dependence from liquid helium temperatures to melting temperatures (10). Because melting involves displacements up to = 10% of bond lengths simple harmonic theories are inappropriate. The MSPA effective force constants, calculated for the interaction between atoms across the hydrogen bonds, are not constant nor are they the small displacement force constants of traditional harmonic theory. They are thermodynamic parameters that themselves depend on the thermal motion. The parameter can be used however, to give solutions of problems involving H-bond stretch. The functional form for determining the effective force comes from a free energy minimization (8). Such an effective force constant can be compared to an analogous parameter, the compressability of a gas used in calculations of acoustic propagation. That too is a thermodynamic effective parameter that is appropriate to solutions of acoustic problems. The origin of the compressability is quite complex and also depends on thermal motion. The hydrogen bond thermal motion that leads to melting is calculated by this approach. All the earlier applications have dealt with systems with helical symmetry (11, 12). This work applies the bond breaking theory to a system with a specific symmetry breaking base sequence. The results of the earlier work have been in agreement with the /92/ni /92/01/58/05 /58/05 $2.00 $200 Biophys. J. e Biophysical Society Volume 61 January
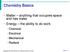
2 observation of fluctuational opening in the premelting region (11) and has shown the proper cooperative behavior in the critical transition region (12). A Green's function technique (13) has been modified to study the vibrational properties of DNA polymers with broken helical symmetry and has been applied to several examples of DNA double helix. These include the junction of AT/GC helices (14), the terminus of a DNA homopolymer (15), a junction of DNA double helix and single strands (16), and defect-mediated melting of DNA homopolymers (8). In these examples, the alterations in interactions between atoms from those of the perfect helix are treated as defects. The presence of such defects breaks the helical symmetry thereby making the description of the normal modes much more difficult. However, for situations of interest, such defects usually exist in a finite region of the polymer and involve a relatively small number of atoms. The problem is then conveniently solved by the Green's function technique. A variety of realistic DNA molecules can be treated using this technique, including those with structural disorder, such as terminal regions, irregular base pair sequences, and various linking arrangements involving DNA double helices or single strands which occur during DNA replication, recombination, RNA transcription, and other biological processes. These defects are believed to play an important role in the melting of the DNA double helix. It has been found that localized modes can exist around defects in DNA polymers, which enhance the vibrational fluctuations in the hydrogen bonds that link the two complimentary strands of the DNA double helix. This can initiate melting of the DNA double helix around the defects (8, 14-16). It is also found experimentally that the stability of a portion of DNA helix can be influenced by the nucleotide sequence of adjacent regions (17, 18). Hence, alterations in the nucleotide sequence in regions adjacent to an actual DNA-protein interaction site could influence the recognition of that site, particularly if fluctuations to a transitory open state of the helix is important for protein binding and initiation of the biological process. It is also believed that A-T rich regions should melt at lower temperatures than G-C rich regions, and this can lead to an initiation of melting from the AT rich sites. Such open regions offer a potential interaction site. In this paper, we apply the Green's function technique to study vibrational properties of a DNA double helix which has a finite region containing base pairs different from the rest of the helix. Breathing modes of the hydrogen bonds are of particular interest as discussed earlier. The Green's function method is used to study the thermal fluctuation in the average hydrogen bond stretch and lead to calculations of the probability of interbase hydrogen bond disruption. The calculation is carried out for a DNA double helix in the B conformation which has (TATA )2 base pair box in between two semi-infinite poly(da-dc) * poly(dg-dt) helices. One side of (TATA)2 has a GC base pair next to the insert and the other, the downstream side, has an AT base pair next to the insert. This gives rise to a TATAA region that corresponds more closely to the consensus inserts (19, 20). Open base pair probability is calculated for the insert base pairs and for five base pairs on both sides of the insert and compared with those of poly(dadc) * poly(dg-dt) and poly(dt-da) * poly(dt-da) in the same conformation. FORMALISM We construct the helix system starting with a perfect double helix (B - poly(da-dc) poly(dg-dt)) and a finite section (2N base pairs) of double helix with the same conformation but different base pairs (TATA )2. Then we replace N consecutive cells of the perfect helix by the finite section. This is achieved by setting the force constants linking the atoms across cell boundary to zero at two junctions N cells apart in the infinite helix and adding corresponding forces to connect the finite section to the two semi-infinite strands as shown in Fig. 1. We choose to cut the 05-C5 bond so that the number of valence forces involved is a minimum. The nonbonded stacking forces are also cut (21). The perfect helix system, consisting of an infinite double helix and a finite section of helix of different base pairs, is described by the eigen problem: (F - W2I)q = O0 (1) where F is the force constant matrix, I is a unitary matrix, w and q are eigenvalues and eigenvectors in the mass weighted Cartesian (MWC) coordinates. The infinite helix eigenvalues and eigenvectors are solved by helical lattice methods (22). The finite helix section can be solved exactly if the number of cells, N, is small. To avoid the complexity added due to end effects of the finite section that ultimately go away and to reduce the computing time for large N, we approximate the eigenfrequencies and eigenvectors for the finite segment by those extracted from the eigen values and eigen vectors of the corresponding infinite double helix for only the following 0 values: 2'Trn 0= -vr+ n= 1,2,3,...,N. N' For a finite (TATA )2 section four base pairs long (two unit cells long), the eigenvalues and eigenvectors corresponding to 0 = 0, and Tr from - those of B poly(dt-da) poly(dt-da) are used. These two 0 values for each band of the periodic helix give the correct number of states associated with a two unit cell section. The altered helix system is described by a similar eigen equation (23): (F w'i + C + C)q = 0, (2) where F is the force constant matrix for the unperturbed system. w and q are the new eigenvalues and eigenvectors. The alteration matrix, C, is the change in F necessary to bring about the severing of the infinite helix and the joining of the two semi-infinite helices created, to the finite section. c is the matrix due to the anharmonicity of the hydrogen bonds which soften as the temperature rises or as the thermal motion Beger and Prohofsky Open Base Pair Probability Beger and Prohofsky Open Base Pair Probability 59
3 increases for any reason. Differences in motion are introduced by the C factors. The c will act synergistically to enhance the altered motion arising from C, i.e., the structural defects. The finite alteration C breaks the helical symmetry. The normal mode calculation can no longer use helical lattice methods as these depend on symmetry. However, because C is a matrix of relatively low dimensionality, Eq. 2 can be solved in terms of Green's function of the unperturbed system. The hydrogen bond mean square amplitudes may be directly calculated from normal mode analysis. In the current problem, however, the harmonic approximation fails because any analysis of hydrogen bond disruption necessarily involves large displacement of the hydrogen bonds. In the MSPA theory, an iterative method is introduced to compute the above quantities self-consistently for a Morse-like hydrogen bond potential. In this paper, an initial set of force constants are obtained by fitting to the Lippincott-Schroeder potential (24) for all hydrogen bonds. Then the secular equation is solved for the copolymers. The normal mode frequencies and eigenvectors are used to obtain anharmonic force constants from a Morse potential. For all future calculations, the Morse potential is used to calculate force constants appropriate to the degree of thermal fluctuation at a given temperature. These new force constants are used to calculate a new set of normal mode frequencies and eigenvectors which are used to reevaluate the force constants. This process is repeated until it reaches self-consistency, which is indicated by a convergence in the force constant for each hydrogen bond into a stable value (8). In our approach, the double helix can be described by Eq. 1 and c is the only matrix that depends on temperature. We may define a Green's function for the system at a fixed temperature with only structural defects (i.e., c = 0). calculation related to an equilibrated system with some thermodynamic fluctuations. The force constant corresponding to the hydrogen bond with this mean square stretch amplitude is then calculated from reference 26 as: where I due_'2/2d. d2vi S-h = due-u1/2dj -i du2 V(u) = VO(e2a((R)+u) _2e-a((R)+u)) (9) is the Morse potential. Here V0 and a are parameters that characterize the potential, (R) is the thermal expanded mean hydrogen bond length and is the centroid of the distribution function exp(-u2/2d) used in Eq. 8. It is a function of the mean square stretch amplitude. The determination of (R) is shown in earlier publications (26). The effective force constant + depends on the mean square fluctuation of the hydrogen bond length D and therefore depends on temperature. D in turn depends on the eigenvalues and eigenvectors of the system which are functions of the force constant matrix which contains +. +, (R), and D are all solved iteratively until a self-consistent set of values are obtained. The probability of a single hydrogen bond, being disrupted in the premelting state, is then given by MSPA theory (11, 12) as (8) P- Pi = Ci..m, du exp (-(u -(Ri))212Di), (10) G = (,I- F - C)-, and a Green's function for the complete system, with thermal effects, (3) where Cy'=J,f due-'. (11) It can be shown that (25): where G= (2I-F- C -C)'. G = G + GTG, T =(C'-G)-l The Green's function G is temperature independent except for the Bose Einstein weight factor and needs to be calculated only once from the initial helix system. Then, Eqs. 5 and 6 can be used to compute the Green function C of the complete system at any iterative step for a set of trial hydrogen bond force constants and only this calculation is done iteratively to self-consistency. The inband thermal mean-squared vibrational amplitude for the mth hydrogen bond may be obtained by (14): h - Dm(w) =-Im [G:"'(w2)] ITr~T Ihw\ 2 coth 2-T In calculating the total D we integrate over w. The result is correct for a noncoherent set of contributions from all modes that have some projection on hydrogen bond stretch. It is not just the contribution from one coherent oscillation. The lifetime of the modes involved does not effect this result. The total integrated contribution from a mode is independent of lifetime for the same reason that the total integrated intensity of an absorption band is independent of mode lifetime and width, i.e., what is missed at one frequency is picked up elsewhere. The (4) Here, C, is the normalization factor for the ith hydrogen bond. L j"m' is the maximum value of stretch before the ith hydrogen bond is considered in the open state. Di is the thermal mean fluctuation square (5) stretch of the ith hydrogen bond. (R1) is the equilibrium length of the ith hydrogen bond. hi is determined from the relation given below (11, 12): (6) V((Ri) hi) - = = V(oo) (7) 0. (12) The probability that a given base pair is in the open state is the probability that all the hydrogen bonds of that pair are disrupted simultaneously and in our mean field approximation that is the product of the open probabilities of the individual hydrogen bonds. The Morse parameters and maximum length of all the hydrogen bonds used in this calculation are shown in Table 1 and Table 2 for Adenosine-Thymine and Guanine-Cytosine base pairs, respectively (10, 23). Ro is the position of the minimum of the Morse potential and is equal to (R) at T = 0. The values for the Morse parameters used here give rise to mode calculations that fit the observed spectra (27) and fit TABLE 1 The Morse parameters of AT base pairs a Ro VO Lmax Bond A-l A mdyn A A N(1)-H-N(3) N(6)-H-0(4) niophysical 1992~~~~~~~~~~~~~~~~~~~~~~~~~~~~ Biophysical Journal Volume 61 January 1992
4 TABLE 2 The Morse parameters of GC base pairs TABLE 3 Insert open base pair probabilities a Ro VO Lma Bond A-1 A mdyn A A N(1)-H-N(3) (6)-H-N(4) N(2)-H-0(2) the temperature dependence of the 85-cm-' mode (9) which are derived by Chen, Y. Z., et al. (29). RESULTS AND DISCUSSION We carried out the calculation for an infinite DNA double helix in the B conformation, which has (TATA )2 base pairs in the middle and poly(da-dc) * poly(dgdt) base pairs for the rest of it. We have found that breathing modes exist around 85 wave numbers in both homopolymers (28). We scanned through the entire spectra of both poly(dt-da ) * poly(dt-da) and poly(da-dc) * poly(dg-dt) and found that the average hydrogen bond stretch amplitude is significant only for a few branches of the perfect helix dispersion curves between 5 and 300 cm-'. Therefore, only this frequency range is considered in studying the breathing modes of the altered helix. The open hydrogen bond probability is given in Table 3 for temperatures T = 293 and 310 K. The hydrogen bond numbers refer to the numbers seen at the bottom of Fig. 1. The open hydrogen bond probability (Pi) for the hydrogen bonds upstream of the insert except for hydrogen bond number 4 are smaller than the perfect open hydrogen bond probability (P(per)). Inside the insert the open hydrogen bond probabilities are lower than the perfect AT copolymer open hydrogen bond probability. This is because the C-G base pairs now included on both sides of the insert tend to increase the stability of the insert and lower the open hydrogen bond probability. The open hydrogen bond probabilities are dramatically increased for the hydrogen bonds downstream the insert for number 14 at T = 293 K. The effect of the particular sequence studied is to stabilize the base pair bonding to one side of the insert (the upstream side) and to destabilize the base pair bonding on the other side (downstream side). Downstream is the direction that RNA transcription occurs with respect to the (TATA )2 insert in many studied initiation sites. The pair opening probability for AT pairs is farily large. The likely critical element in initiating or elongating an open state, necessary for transcription to occur, is to open the GC pairs. They would form the bottleneck to open state elongation. The first GC pair downstream of the insert is H P(per) HiP(per) P1 HIPi Pi Bondi T= 293 T= 293 T= 293 T= 293 Ratio T= Column 1 is H-bond number. Column 2 is open probability for each H-bond at 293 K, for the perfect system, i.e., the separate homopolymers. Column 3 is the probability of a base pair being open at 293 K for the perfect system. Column 4 is the individual H-bond open probability for the new structure with insert at 293 K. Column S is the open base probability for the new structure. Column 6 is the ratio of open base probability for new structure to perfect system. Column 7 is the H-bond open probability at 310 K. an order of magnitude more likely to fluctuate to an open state than a GC pair in the original perfect copolymer. It is 110 times more likely to open than the first GC pair upstream to the (TA TA )2 insert. The actual numerical values of open bond probability Major Minor Central ACACAC ACAC ACAC...,... TGTGTG TGTG TGTGTG ACACAC TGTGTG TATA ATA \ TA TATA ATAT A C A C A C T GT GT G FIGURE 1 The double helix is constructed by cutting the perfect helix, and joining the finite section (TA TA)2 to the semi-infinite strands. The numbers at the bottom identify the hydrogen bonds of each base pair and are used in Table 3. Beger and Prohofsky Open Base Pair Probability 61 Beger and Prohofsky Open Base Pair Probability 61
5 are strongly dependent on values chosen for L". Our choice of Lmaxs come from observation of the onset of anomalous behavior when exploring the thermal melting of homopolymer DNAs (29). The actual values would be determined by how hydrogen bonds are altered by the presence of solvent atoms and should be fit with greater care. The values chosen do however fit data of bond opening based on analysis of proton exchange for both amino and imino protons in DNA fairly well (11). This work was supported by Office of Naval Research contract N K Received for publication on 29 April 1991 and in final formn 9 August REFERENCES 1. Lindeman, F Thermal melting law. Phys. Z. 11: Urabe, H., and Y. Tominga, Low frequency Raman spectra of DNA. J. Phys. Soc. Jpn. 50: Lamba, 0. P., A. H. J. Wang, and G. J. Thomas, Jr Low frequency dynamics and Raman scattering of crystals, of B-, A-, and Z-DNA and fibers of C-DNA. Biopolymers. 28: Weidlich, T., and S. M. Lindsay Counterion effects on the structure and dynamics of solid DNA. Phys. Rev. Lett. 61: Powell, J. W., G. S. Edwards, L. Genzel, and A. Wittlin Investigation of far-infrared vibrational modes in polynucleotides. Phys. Rev. A. 35: Urabe, H., H. Hayashi, X. Tominga, Y. Nishimura, K. Kubota, and M. Tsuboi Collective vibrational modes in molecular assembly of DNA and its applications to biological systems. Low frequency Raman spectroscopy. J. Chem. Phys. 82: Tominga, Y., M. Shida, K. Kubota, H. Urabe, Y. Nishimura, and M. Tsuboi Coupled dynamics between DNA helix and spectroscopy. J. Chem. Phys. 83: Kim, Y., and E. W. Prohofsky Defect-mediated hydrogenbond instability of poly(dg) * poly(dc). Phys. Rev. B33: Eyster, J. M., and E. W. Prohofsky Lattice vibrational lmodes of Poly(rU) Poly(rA). A coupled single-helical ap- 'proach. Biopolymers. 13: Kim, Y., and E. W. Prohofsky Vibrational modes of a DNA polymer at low temperature. Phys. Rev. B36: Chen, Y. Z., Y. Feng, and E. W. Prohofsky Premelting thermal fluctuational base pair opening probability of poly(da ) * poly(dt) as predicted by the modified self-consistent phonon theory. Biopolymers. 31: Chen, Y. Z., and E. W. Prohofsky A self-consistent mean-field calculation of a homopolymer deoxyribose nucleic acid strand separation: bond breaking in a macromolecule. J. Chem. Phys. 94: Lifshitz, I. M., and S. I. Pekar Tamm connected states of electrons on the surface of a crystal and the surface oscillations of the atoms of a lattice. Usp. Fiziol. Nauk. 56: Feng, Y., R. D. Beger, X. Hua, and E. W. Prohofsky Breathing modes near a junction of DNA double helices. Phys. Rev. 40: Putnam, B. F., L. L. Van Znadt, E. W. Prohofsky, and W. N. Mei Resonant and localized breathing modes in terminal regions of the DNA double helix. Biophys. J. 35: Putnam, B. F., and E. W. Prohofsky Localized vibrational modes at a double-helix single strand junction. Biopolymers. 22: Wells, R. D., R. W. Blakesley, S. C. Hardies, G. T. Horn, J. E. Larson, E. Selsing, J. F. Burd, H. W. Chan, J. B. Dodgson, K. F. Jensen, I. F. Nes, and R. M. Wartell The role of DNA structure in genetic regulation. CRC Critical Reviews in Biochemistry Burd, J. D., R. M. Wartell, J. B. Dodgson, and R. D. Wells Transmission of stability (telestability) in deoxyribonucleic acid. J. Bio. Chem. 250: Hawley, D. K., and W. R. McClure Compilation and analysis of Escherichia coli promoter sequences. Nucleic Acids Res. 11: Cho, K., and C. Yanofsky Development of a trpe promoterstrength measuring system and its use in comparison of the trpedcba, trpr and aroh promoters. J. Mol. BioL 204: Putnam, B. F Calculation of macromolecular force constants and vibrational properties of a semi-infinite strand of the DNA double helix. Ph.D. Thesis, Purdue University, West Lafayette, IN Eyster, J. M Normal modes of helical polymers of biological interest. Ph.D. Thesis, Purdue University, West Lafayette, IN Feng, Y., W. Zhuang, and E. W. Prohofsky Calculation of temperature dependence of interbase breathing motion of a guanine-cytosine DNA double helix with adenine-thymine insert. Phys. Rev. A. 43: Schroeder, R., and E. R. Lippincott Potential function model of hydrogen bonds. II. J. Phys. Chem. 61: Bottger, H Principles of the Theory of Lattice Dynamics. Physik-Verlag, Weinbeim, FRG. 87 pp. 26. Gao, Y., K. V. Prasad, and E. W. Prohofsky A selfconsistent microscopic theory of hydrogen bond melting with application topoly(dg) * poly(dc). J. Chem. Phys. 80: Devi-Prasad, K. V., and E. W. Prohofsky Low-frequency mode predictions in A-DNA compared to experimental observations and significance for A-to-B. Biopolymers. 23: Prohofsky, E. W Biomolecular Stereodynamics IV, Proceedings of the Fourth Conservation in the Discipline Biomolecular Stereodynamics. Albany, NY Chen, Y. Z., Y. Feng, and E. Prohofsky Criteria of thermal denaturation for modified self-consistent-phonon-theory meanfield calculations in DNA polymers. Phys. Rev. B.42: Biophysical Journal Volume 61 January 1992
poly(da) poly(dt) against fluctuational interbase H-bond
 .=) 1992 Oxford University Press Nucleic Acids Research, Vol. 20, No. 3 415-419 The role of a minor groove spine of hydration in stabilizing poly(da) poly(dt) against fluctuational interbase H-bond disruption
.=) 1992 Oxford University Press Nucleic Acids Research, Vol. 20, No. 3 415-419 The role of a minor groove spine of hydration in stabilizing poly(da) poly(dt) against fluctuational interbase H-bond disruption
Principles of Physical Biochemistry
 Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
DISPLACEMENTS OF BACKBONE VIBRATIONAL
 DSPLACEMENTS OF BACKBONE VBRATONAL MODES OF A-DNA AND B-DNA K.-C. Lu, L. L. VAN ZANDT, AND E. W. PROHOFSKY, Department ofphysics, Purdue University, West Lafayette, ndiana 4797 U.S.A. ABSTRACT We display
DSPLACEMENTS OF BACKBONE VBRATONAL MODES OF A-DNA AND B-DNA K.-C. Lu, L. L. VAN ZANDT, AND E. W. PROHOFSKY, Department ofphysics, Purdue University, West Lafayette, ndiana 4797 U.S.A. ABSTRACT We display
Contents. xiii. Preface v
 Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
Synergistic effects in the melting of DNA hydration shell: melting of
 Synergistic effects in the melting of DNA hydration shell: melting of the minor groove hydration spine in poly(da). poly(dt) and its effect on base pair stability Y. Z. Chen and E. W. Prohofsky Department
Synergistic effects in the melting of DNA hydration shell: melting of the minor groove hydration spine in poly(da). poly(dt) and its effect on base pair stability Y. Z. Chen and E. W. Prohofsky Department
Stability of Triple-Helical Poly(dT)-Poly(dA)-Poly(dT) DNA with Counterions
 70 Biophysical Journal Volume 75 July 1998 70 91 Stability of Triple-Helical Poly(dT)-Poly(dA)-Poly(dT) DNA with Counterions Voichita M. Dadarlat and V. K. Saxena Department of Physics, Purdue University,
70 Biophysical Journal Volume 75 July 1998 70 91 Stability of Triple-Helical Poly(dT)-Poly(dA)-Poly(dT) DNA with Counterions Voichita M. Dadarlat and V. K. Saxena Department of Physics, Purdue University,
Chemistry Basics. Matter anything that occupies space and has mass Energy the ability to do work. Chemical Electrical Mechanical Radiant. Slide 2.
 Chemistry Basics Matter anything that occupies space and has mass Energy the ability to do work Chemical Electrical Mechanical Radiant Slide 2.1 Composition of Matter Elements Fundamental units of matter
Chemistry Basics Matter anything that occupies space and has mass Energy the ability to do work Chemical Electrical Mechanical Radiant Slide 2.1 Composition of Matter Elements Fundamental units of matter
NUMERICAL SIMULATION OF THE IR SPECTRA OF DNA BASES
 BIOPHYSICS NUMERICAL SIMULATION OF THE IR SPECTRA OF DNA BASES C. I. MORARI, CRISTINA MUNTEAN National Institute of Research and Development for Isotopic and Molecular Technologies P.O. Box 700, R-400293
BIOPHYSICS NUMERICAL SIMULATION OF THE IR SPECTRA OF DNA BASES C. I. MORARI, CRISTINA MUNTEAN National Institute of Research and Development for Isotopic and Molecular Technologies P.O. Box 700, R-400293
BIOCHEMISTRY GUIDED NOTES - AP BIOLOGY-
 BIOCHEMISTRY GUIDED NOTES - AP BIOLOGY- ELEMENTS AND COMPOUNDS - anything that has mass and takes up space. - cannot be broken down to other substances. - substance containing two or more different elements
BIOCHEMISTRY GUIDED NOTES - AP BIOLOGY- ELEMENTS AND COMPOUNDS - anything that has mass and takes up space. - cannot be broken down to other substances. - substance containing two or more different elements
Chapter 1. Topic: Overview of basic principles
 Chapter 1 Topic: Overview of basic principles Four major themes of biochemistry I. What are living organism made from? II. How do organism acquire and use energy? III. How does an organism maintain its
Chapter 1 Topic: Overview of basic principles Four major themes of biochemistry I. What are living organism made from? II. How do organism acquire and use energy? III. How does an organism maintain its
arxiv:cond-mat/ v1 [cond-mat.stat-mech] 21 Dec 2004
![arxiv:cond-mat/ v1 [cond-mat.stat-mech] 21 Dec 2004 arxiv:cond-mat/ v1 [cond-mat.stat-mech] 21 Dec 2004](/thumbs/82/86926711.jpg) Theory of Bubble Nucleation and Cooperativity in DNA Melting arxiv:cond-mat/4259v [cond-mat.stat-mech] 2 Dec 24 Saúl Ares,, 2 N. K. Voulgarakis, 3 K. Ø. Rasmussen, and A. R. Bishop Theoretical Division
Theory of Bubble Nucleation and Cooperativity in DNA Melting arxiv:cond-mat/4259v [cond-mat.stat-mech] 2 Dec 24 Saúl Ares,, 2 N. K. Voulgarakis, 3 K. Ø. Rasmussen, and A. R. Bishop Theoretical Division
Introduction to Polymer Physics
 Introduction to Polymer Physics Enrico Carlon, KU Leuven, Belgium February-May, 2016 Enrico Carlon, KU Leuven, Belgium Introduction to Polymer Physics February-May, 2016 1 / 28 Polymers in Chemistry and
Introduction to Polymer Physics Enrico Carlon, KU Leuven, Belgium February-May, 2016 Enrico Carlon, KU Leuven, Belgium Introduction to Polymer Physics February-May, 2016 1 / 28 Polymers in Chemistry and
Human Biology. The Chemistry of Living Things. Concepts and Current Issues. All Matter Consists of Elements Made of Atoms
 2 The Chemistry of Living Things PowerPoint Lecture Slide Presentation Robert J. Sullivan, Marist College Michael D. Johnson Human Biology Concepts and Current Issues THIRD EDITION Copyright 2006 Pearson
2 The Chemistry of Living Things PowerPoint Lecture Slide Presentation Robert J. Sullivan, Marist College Michael D. Johnson Human Biology Concepts and Current Issues THIRD EDITION Copyright 2006 Pearson
Exploring the Sequence Dependent Structure and Dynamics of DNA with Molecular Dynamics Simulation
 Exploring the Sequence Dependent Structure and Dynamics of DNA with Molecular Dynamics Simulation Sarah Harris School of Physics and Astronomy University of Leeds Introduction Calculations of the charge
Exploring the Sequence Dependent Structure and Dynamics of DNA with Molecular Dynamics Simulation Sarah Harris School of Physics and Astronomy University of Leeds Introduction Calculations of the charge
PTYS 214 Spring Announcements. Midterm #1 on Tuesday! Be on time! No one enters after the first person leaves! Do your homework!
 PTYS 214 Spring 2018 Announcements Midterm #1 on Tuesday! Be on time! No one enters after the first person leaves! Do your homework! 1 Last time - Properties of Life Organization, energy utilization, homeostasis,
PTYS 214 Spring 2018 Announcements Midterm #1 on Tuesday! Be on time! No one enters after the first person leaves! Do your homework! 1 Last time - Properties of Life Organization, energy utilization, homeostasis,
DNA Structure. Voet & Voet: Chapter 29 Pages Slide 1
 DNA Structure Voet & Voet: Chapter 29 Pages 1107-1122 Slide 1 Review The four DNA bases and their atom names The four common -D-ribose conformations All B-DNA ribose adopt the C2' endo conformation All
DNA Structure Voet & Voet: Chapter 29 Pages 1107-1122 Slide 1 Review The four DNA bases and their atom names The four common -D-ribose conformations All B-DNA ribose adopt the C2' endo conformation All
Unit 2: Basic Chemistry
 Unit 2: Basic Chemistry I. Matter and Energy A. Matter anything that occupies space and has mass (weight) B. Energy the ability to do work 1. Chemical 2. Electrical 3. Mechanical 4. Radiant C. Composition
Unit 2: Basic Chemistry I. Matter and Energy A. Matter anything that occupies space and has mass (weight) B. Energy the ability to do work 1. Chemical 2. Electrical 3. Mechanical 4. Radiant C. Composition
Stacking and Hydrogen Bonding. DNA Cooperativity at Melting.
 Stacking and Hydrogen Bonding. DNA Cooperativity at Melting. Vladimir F. Morozov *, Artem V. Badasyan, Arsen V. Grigoryan, Mihran A. Sahakyan, Evgeni Sh. Mamasakhlisov. 1 Department of Molecular Physics,
Stacking and Hydrogen Bonding. DNA Cooperativity at Melting. Vladimir F. Morozov *, Artem V. Badasyan, Arsen V. Grigoryan, Mihran A. Sahakyan, Evgeni Sh. Mamasakhlisov. 1 Department of Molecular Physics,
Chapter 2: Chemical Basis of Life
 Chapter 2: Chemical Basis of Life Chemistry is the scientific study of the composition of matter and how composition changes. In order to understand human physiological processes, it is important to understand
Chapter 2: Chemical Basis of Life Chemistry is the scientific study of the composition of matter and how composition changes. In order to understand human physiological processes, it is important to understand
Chapter 2: Chemistry. What does chemistry have to do with biology? Vocabulary BIO 105
 Chapter 2: Chemistry What does chemistry have to do with biology? BIO 105 Vocabulary 1. Matter anything that takes up space and has mass Atoms are the smallest units of matter that can participate in chemical
Chapter 2: Chemistry What does chemistry have to do with biology? BIO 105 Vocabulary 1. Matter anything that takes up space and has mass Atoms are the smallest units of matter that can participate in chemical
 There are two types of polysaccharides in cell: glycogen and starch Starch and glycogen are polysaccharides that function to store energy Glycogen Glucose obtained from primary sources either remains soluble
There are two types of polysaccharides in cell: glycogen and starch Starch and glycogen are polysaccharides that function to store energy Glycogen Glucose obtained from primary sources either remains soluble
2: CHEMICAL COMPOSITION OF THE BODY
 1 2: CHEMICAL COMPOSITION OF THE BODY Although most students of human physiology have had at least some chemistry, this chapter serves very well as a review and as a glossary of chemical terms. In particular,
1 2: CHEMICAL COMPOSITION OF THE BODY Although most students of human physiology have had at least some chemistry, this chapter serves very well as a review and as a glossary of chemical terms. In particular,
The biomolecules of terrestrial life
 Functional groups in biomolecules Groups of atoms that are responsible for the chemical properties of biomolecules The biomolecules of terrestrial life Planets and Astrobiology (2017-2018) G. Vladilo 1
Functional groups in biomolecules Groups of atoms that are responsible for the chemical properties of biomolecules The biomolecules of terrestrial life Planets and Astrobiology (2017-2018) G. Vladilo 1
Base pairing in DNA.
 TFY4215 Kjemisk fysikk og kvantemekanikk Våren 2007 Chemical physics Exercise 3 To be delivered by: Tuesday 08.05. Base pairing in DNA. Introduction DNA, deoxyribonucleic acid are the molecules that contain
TFY4215 Kjemisk fysikk og kvantemekanikk Våren 2007 Chemical physics Exercise 3 To be delivered by: Tuesday 08.05. Base pairing in DNA. Introduction DNA, deoxyribonucleic acid are the molecules that contain
Foundations in Microbiology Seventh Edition
 Lecture PowerPoint to accompany Foundations in Microbiology Seventh Edition Talaro Chapter 2 The Chemistry of Biology Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
Lecture PowerPoint to accompany Foundations in Microbiology Seventh Edition Talaro Chapter 2 The Chemistry of Biology Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
Olle Inganäs: Polymers structure and dynamics. Polymer physics
 Polymer physics Polymers are macromolecules formed by many identical monomers, connected through covalent bonds, to make a linear chain of mers a polymer. The length of the chain specifies the weight of
Polymer physics Polymers are macromolecules formed by many identical monomers, connected through covalent bonds, to make a linear chain of mers a polymer. The length of the chain specifies the weight of
Ch 3: Chemistry of Life. Chemistry Water Macromolecules Enzymes
 Ch 3: Chemistry of Life Chemistry Water Macromolecules Enzymes Chemistry Atom = smallest unit of matter that cannot be broken down by chemical means Element = substances that have similar properties and
Ch 3: Chemistry of Life Chemistry Water Macromolecules Enzymes Chemistry Atom = smallest unit of matter that cannot be broken down by chemical means Element = substances that have similar properties and
CORE MOLIT ACTIVITIES at a glance
 CORE MOLIT ACTIVITIES at a glance 1. Amplification of Biochemical Signals: The ELISA Test http://molit.concord.org/database/activities/248.html The shape of molecules affects the way they function. A test
CORE MOLIT ACTIVITIES at a glance 1. Amplification of Biochemical Signals: The ELISA Test http://molit.concord.org/database/activities/248.html The shape of molecules affects the way they function. A test
1. (5) Draw a diagram of an isomeric molecule to demonstrate a structural, geometric, and an enantiomer organization.
 Organic Chemistry Assignment Score. Name Sec.. Date. Working by yourself or in a group, answer the following questions about the Organic Chemistry material. This assignment is worth 35 points with the
Organic Chemistry Assignment Score. Name Sec.. Date. Working by yourself or in a group, answer the following questions about the Organic Chemistry material. This assignment is worth 35 points with the
Lesson Overview The Structure of DNA
 12.2 THINK ABOUT IT The DNA molecule must somehow specify how to assemble proteins, which are needed to regulate the various functions of each cell. What kind of structure could serve this purpose without
12.2 THINK ABOUT IT The DNA molecule must somehow specify how to assemble proteins, which are needed to regulate the various functions of each cell. What kind of structure could serve this purpose without
Hie-Joon Kim. Professor Emeritus Seoul National University. Experience. Representative Publications
 Hie-Joon Kim Professor Emeritus Seoul National University B.S. Chemistry, Seoul National University, Korea, 1970 Ph.D. Chemistry, University of Chicago, USA, 1977 Experience Professor, Department of Chemistry
Hie-Joon Kim Professor Emeritus Seoul National University B.S. Chemistry, Seoul National University, Korea, 1970 Ph.D. Chemistry, University of Chicago, USA, 1977 Experience Professor, Department of Chemistry
INTRODUCTION TO MODERN VIBRATIONAL SPECTROSCOPY
 INTRODUCTION TO MODERN VIBRATIONAL SPECTROSCOPY MAX DIEM Department of Chemistry City University of New York Hunter College A Wiley-Interscience Publication JOHN WILEY & SONS New York Chichester Brisbane
INTRODUCTION TO MODERN VIBRATIONAL SPECTROSCOPY MAX DIEM Department of Chemistry City University of New York Hunter College A Wiley-Interscience Publication JOHN WILEY & SONS New York Chichester Brisbane
Chapters 12&13 Notes: DNA, RNA & Protein Synthesis
 Chapters 12&13 Notes: DNA, RNA & Protein Synthesis Name Period Words to Know: nucleotides, DNA, complementary base pairing, replication, genes, proteins, mrna, rrna, trna, transcription, translation, codon,
Chapters 12&13 Notes: DNA, RNA & Protein Synthesis Name Period Words to Know: nucleotides, DNA, complementary base pairing, replication, genes, proteins, mrna, rrna, trna, transcription, translation, codon,
A Thermodynamic Investigation into the Stabilization of Poly(dA) [poly(dt)] 2 Triple Helical DNA by Various Divalent Metal Ions
![A Thermodynamic Investigation into the Stabilization of Poly(dA) [poly(dt)] 2 Triple Helical DNA by Various Divalent Metal Ions A Thermodynamic Investigation into the Stabilization of Poly(dA) [poly(dt)] 2 Triple Helical DNA by Various Divalent Metal Ions](/thumbs/90/103085351.jpg) Thermodynamics on the Poly(dA) [poly(dt)] Triplex Formation Bull. Korean Chem. Soc. 009, Vol. 30, No. 11 691 A Thermodynamic Investigation into the Stabilization of Poly(dA) [poly(dt)] Triple Helical DNA
Thermodynamics on the Poly(dA) [poly(dt)] Triplex Formation Bull. Korean Chem. Soc. 009, Vol. 30, No. 11 691 A Thermodynamic Investigation into the Stabilization of Poly(dA) [poly(dt)] Triple Helical DNA
ARTICLES. Normal-mode analysis without the Hessian: A driven molecular-dynamics approach
 JOURNAL OF CHEMICAL PHYSICS VOLUME 119, NUMBER 2 8 JULY 2003 ARTICLES Normal-mode analysis without the Hessian: A driven molecular-dynamics approach Joel M. Bowman, a) Xiubin Zhang, and Alex Brown Cherry
JOURNAL OF CHEMICAL PHYSICS VOLUME 119, NUMBER 2 8 JULY 2003 ARTICLES Normal-mode analysis without the Hessian: A driven molecular-dynamics approach Joel M. Bowman, a) Xiubin Zhang, and Alex Brown Cherry
Statistical Thermodynamics of DNA Denaturation. processes
 Statistical Thermodynamics of DNA Denaturation processes Course: Statistical Thermodynamics Advisor: Prof. Raissa D Souza Spring quarter 2012 By: Sebastian Frey 1 Nomenclature F Free Energy [J] x Coordinate
Statistical Thermodynamics of DNA Denaturation processes Course: Statistical Thermodynamics Advisor: Prof. Raissa D Souza Spring quarter 2012 By: Sebastian Frey 1 Nomenclature F Free Energy [J] x Coordinate
Dynamics of Solitary Waves Induced by Shock Impulses in a Linear Atomic Chain*
 Dynamics of Solitary Waves Induced by Shock Impulses in a Linear Atomic Chain* PHUOC X. TRAN, DONALD W. BRENNER, and C. T. WHITE Naval Research Laboratory, Washington, DC 20375-5000 Abstract The propagation
Dynamics of Solitary Waves Induced by Shock Impulses in a Linear Atomic Chain* PHUOC X. TRAN, DONALD W. BRENNER, and C. T. WHITE Naval Research Laboratory, Washington, DC 20375-5000 Abstract The propagation
A low-frequency Raman study of six nucleosides
 The University of Toledo The University of Toledo Digital Repository Theses and Dissertations 2011 A low-frequency Raman study of six nucleosides Craig Koontz The University of Toledo Follow this and additional
The University of Toledo The University of Toledo Digital Repository Theses and Dissertations 2011 A low-frequency Raman study of six nucleosides Craig Koontz The University of Toledo Follow this and additional
Circular dichroism spectroscopic studies on structures formed by telomeric DNA sequences in vitro
 Circular dichroism spectroscopic studies on structures formed by telomeric DNA sequences in vitro ZHANG Xiaoyan 1, CAO Enhua 1, SUN Xueguang 1 & BAI Chunli 2 1. Institute of Biophysics, Chinese Academy
Circular dichroism spectroscopic studies on structures formed by telomeric DNA sequences in vitro ZHANG Xiaoyan 1, CAO Enhua 1, SUN Xueguang 1 & BAI Chunli 2 1. Institute of Biophysics, Chinese Academy
Chapter 6- An Introduction to Metabolism*
 Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
LAFAYETTE IND DEPT OF PHYSICS V V PRRGHU ET AL. UNCLR~~lll' 16I MAR *6 N K-0252 / 62 IN
 A-MI93 11? HELICAL LATTICE VIBRATIONAL MODES IN DNA(U) PURDUE UNIV l' LAFAYETTE IND DEPT OF PHYSICS V V PRRGHU ET AL. UNCLR~~lll' 16I MAR *6 N99914-86-K-0252 / 62 IN U~LS IFIDFG 2 N p 1111l IIl 11111 25
A-MI93 11? HELICAL LATTICE VIBRATIONAL MODES IN DNA(U) PURDUE UNIV l' LAFAYETTE IND DEPT OF PHYSICS V V PRRGHU ET AL. UNCLR~~lll' 16I MAR *6 N99914-86-K-0252 / 62 IN U~LS IFIDFG 2 N p 1111l IIl 11111 25
2.4 DNA structure. S(l) bl + c log l + d, with c 1.8k B. (2.72)
 2.4 DNA structure DNA molecules come in a wide range of length scales, from roughly 50,000 monomers in a λ-phage, 6 0 9 for human, to 9 0 0 nucleotides in the lily. The latter would be around thirty meters
2.4 DNA structure DNA molecules come in a wide range of length scales, from roughly 50,000 monomers in a λ-phage, 6 0 9 for human, to 9 0 0 nucleotides in the lily. The latter would be around thirty meters
the spatial arrangement of atoms in a molecule and the chemical bonds that hold the atoms together Chemical structure Covalent bond Ionic bond
 Chemical structure the spatial arrangement of atoms in a molecule and the chemical bonds that hold the atoms together Covalent bond bond formed by the sharing of valence electrons between atoms Ionic bond
Chemical structure the spatial arrangement of atoms in a molecule and the chemical bonds that hold the atoms together Covalent bond bond formed by the sharing of valence electrons between atoms Ionic bond
Chaotic Modeling and Simulation (CMSIM) 2: , 2017
 Chaotic Modeling and Simulation (CMSIM) 2: 205-212, 2017 Bifurcation based parameter selection of PBD model in DNA Sohrab Behnia 1, Robabeh Panahinia 1 1 Faculty of physics, Urmia University of Technology,
Chaotic Modeling and Simulation (CMSIM) 2: 205-212, 2017 Bifurcation based parameter selection of PBD model in DNA Sohrab Behnia 1, Robabeh Panahinia 1 1 Faculty of physics, Urmia University of Technology,
Chemistry in Biology. Section 1. Atoms, Elements, and Compounds
 Section 1 Atoms, Elements, and Compounds Atoms! Chemistry is the study of matter.! Atoms are the building blocks of matter.! Neutrons and protons are located at the center of the atom.! Protons are positively
Section 1 Atoms, Elements, and Compounds Atoms! Chemistry is the study of matter.! Atoms are the building blocks of matter.! Neutrons and protons are located at the center of the atom.! Protons are positively
Ch. 2 BASIC CHEMISTRY. Copyright 2010 Pearson Education, Inc.
 Ch. 2 BASIC CHEMISTRY Matter and Composition of Matter Definition: Anything that has mass and occupies space Matter is made up of elements An element cannot be broken down by ordinary chemical means Atoms
Ch. 2 BASIC CHEMISTRY Matter and Composition of Matter Definition: Anything that has mass and occupies space Matter is made up of elements An element cannot be broken down by ordinary chemical means Atoms
CHM Physical Chemistry II Chapter 12 - Supplementary Material. 1. Einstein A and B coefficients
 CHM 3411 - Physical Chemistry II Chapter 12 - Supplementary Material 1. Einstein A and B coefficients Consider two singly degenerate states in an atom, molecule, or ion, with wavefunctions 1 (for the lower
CHM 3411 - Physical Chemistry II Chapter 12 - Supplementary Material 1. Einstein A and B coefficients Consider two singly degenerate states in an atom, molecule, or ion, with wavefunctions 1 (for the lower
2/25/2013. Electronic Configurations
 1 2 3 4 5 Chapter 2 Chemical Principles The Structure of Atoms Chemistry is the study of interactions between atoms and molecules The atom is the smallest unit of matter that enters into chemical reactions
1 2 3 4 5 Chapter 2 Chemical Principles The Structure of Atoms Chemistry is the study of interactions between atoms and molecules The atom is the smallest unit of matter that enters into chemical reactions
Mike Towrie Central Laser Facility Rutherford Appleton Laboratory. Diamond DIAMOND. Tony Parker, Pavel Matousek
 Ultrafast deactivation of the electronic excited states of DNA bases and polynucleotides following 267 nm laser excitation explored using picosecond time-resolved infrared spectroscopy 1 Mike Towrie (m.towrie@rl.ac.uk)
Ultrafast deactivation of the electronic excited states of DNA bases and polynucleotides following 267 nm laser excitation explored using picosecond time-resolved infrared spectroscopy 1 Mike Towrie (m.towrie@rl.ac.uk)
Introduction to Molecular Vibrations and Infrared Spectroscopy
 hemistry 362 Spring 2017 Dr. Jean M. Standard February 15, 2017 Introduction to Molecular Vibrations and Infrared Spectroscopy Vibrational Modes For a molecule with N atoms, the number of vibrational modes
hemistry 362 Spring 2017 Dr. Jean M. Standard February 15, 2017 Introduction to Molecular Vibrations and Infrared Spectroscopy Vibrational Modes For a molecule with N atoms, the number of vibrational modes
Chapter 9 DNA recognition by eukaryotic transcription factors
 Chapter 9 DNA recognition by eukaryotic transcription factors TRANSCRIPTION 101 Eukaryotic RNA polymerases RNA polymerase RNA polymerase I RNA polymerase II RNA polymerase III RNA polymerase IV Function
Chapter 9 DNA recognition by eukaryotic transcription factors TRANSCRIPTION 101 Eukaryotic RNA polymerases RNA polymerase RNA polymerase I RNA polymerase II RNA polymerase III RNA polymerase IV Function
Structure and Dynamics : An Atomic View of Materials
 Structure and Dynamics : An Atomic View of Materials MARTIN T. DOVE Department ofearth Sciences University of Cambridge OXFORD UNIVERSITY PRESS Contents 1 Introduction 1 1.1 Observations 1 1.1.1 Microscopic
Structure and Dynamics : An Atomic View of Materials MARTIN T. DOVE Department ofearth Sciences University of Cambridge OXFORD UNIVERSITY PRESS Contents 1 Introduction 1 1.1 Observations 1 1.1.1 Microscopic
Microbiology with Diseases by Taxonomy, 5e (Bauman) Chapter 2 The Chemistry of Microbiology. 2.1 Multiple Choice Questions
 Microbiology with Diseases by Taxonomy, 5e (Bauman) Chapter 2 The Chemistry of Microbiology 2.1 Multiple Choice Questions 1) Which of the following does not contribute significantly to the mass of an atom?
Microbiology with Diseases by Taxonomy, 5e (Bauman) Chapter 2 The Chemistry of Microbiology 2.1 Multiple Choice Questions 1) Which of the following does not contribute significantly to the mass of an atom?
(DPHY 21) 1) a) Discuss the propagation of light in conducting surface. b) Discuss about the metallic reflection at oblique incidence.
 (DPHY 21) ASSIGNMENT - 1, MAY - 2015. PAPER- V : ELECTROMAGNETIC THEORY AND MODERN OPTICS 1) a) Discuss the propagation of light in conducting surface. b) Discuss about the metallic reflection at oblique
(DPHY 21) ASSIGNMENT - 1, MAY - 2015. PAPER- V : ELECTROMAGNETIC THEORY AND MODERN OPTICS 1) a) Discuss the propagation of light in conducting surface. b) Discuss about the metallic reflection at oblique
Physical principles of IR and Raman. Infrared Spectroscopy
 Physical principles of IR and Raman IR results from the absorption of energy by vibrating chemical bonds. Raman scattering results from the same types of transitions, but the selection rules are different
Physical principles of IR and Raman IR results from the absorption of energy by vibrating chemical bonds. Raman scattering results from the same types of transitions, but the selection rules are different
Model Worksheet Teacher Key
 Introduction Despite the complexity of life on Earth, the most important large molecules found in all living things (biomolecules) can be classified into only four main categories: carbohydrates, lipids,
Introduction Despite the complexity of life on Earth, the most important large molecules found in all living things (biomolecules) can be classified into only four main categories: carbohydrates, lipids,
Structural and mechanistic insight into the substrate. binding from the conformational dynamics in apo. and substrate-bound DapE enzyme
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 215 Structural and mechanistic insight into the substrate binding from the conformational
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 215 Structural and mechanistic insight into the substrate binding from the conformational
Biomolecules. Energetics in biology. Biomolecules inside the cell
 Biomolecules Energetics in biology Biomolecules inside the cell Energetics in biology The production of energy, its storage, and its use are central to the economy of the cell. Energy may be defined as
Biomolecules Energetics in biology Biomolecules inside the cell Energetics in biology The production of energy, its storage, and its use are central to the economy of the cell. Energy may be defined as
Chemical Basis of Life
 Chemical Basis of Life Jan 30 11:42 AM In order to understand digestion and nutrition, we need some basic biochemistry Chemistry studies the composition of matter and its changes as well as the change
Chemical Basis of Life Jan 30 11:42 AM In order to understand digestion and nutrition, we need some basic biochemistry Chemistry studies the composition of matter and its changes as well as the change
Ranjit P. Bahadur Assistant Professor Department of Biotechnology Indian Institute of Technology Kharagpur, India. 1 st November, 2013
 Hydration of protein-rna recognition sites Ranjit P. Bahadur Assistant Professor Department of Biotechnology Indian Institute of Technology Kharagpur, India 1 st November, 2013 Central Dogma of life DNA
Hydration of protein-rna recognition sites Ranjit P. Bahadur Assistant Professor Department of Biotechnology Indian Institute of Technology Kharagpur, India 1 st November, 2013 Central Dogma of life DNA
S(l) bl + c log l + d, with c 1.8k B. (2.71)
 2.4 DNA structure DNA molecules come in a wide range of length scales, from roughly 50,000 monomers in a λ-phage, 6 0 9 for human, to 9 0 0 nucleotides in the lily. The latter would be around thirty meters
2.4 DNA structure DNA molecules come in a wide range of length scales, from roughly 50,000 monomers in a λ-phage, 6 0 9 for human, to 9 0 0 nucleotides in the lily. The latter would be around thirty meters
Molecular dynamics simulations of anti-aggregation effect of ibuprofen. Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov
 Biophysical Journal, Volume 98 Supporting Material Molecular dynamics simulations of anti-aggregation effect of ibuprofen Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov Supplemental
Biophysical Journal, Volume 98 Supporting Material Molecular dynamics simulations of anti-aggregation effect of ibuprofen Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov Supplemental
Hydration of Nucleotides Thomas Wyttenbach, Dengfeng Liu, and Michael T. Bowers
 http://bowers.chem.ucsb.edu/ ydration of ucleotides Thomas Wyttenbach, Dengfeng Liu, and Michael T. Bowers ASMS 2006 Why study hydration? Is a certain property of a molecule (e.g. conformation) inherent
http://bowers.chem.ucsb.edu/ ydration of ucleotides Thomas Wyttenbach, Dengfeng Liu, and Michael T. Bowers ASMS 2006 Why study hydration? Is a certain property of a molecule (e.g. conformation) inherent
Part II Construction and comparison with the structure of DNA
 Part II Construction and comparison with the structure of DNA Desired functional properties 1) We look for a stable structure (stable enough to account for the permanency of biological structures on geological
Part II Construction and comparison with the structure of DNA Desired functional properties 1) We look for a stable structure (stable enough to account for the permanency of biological structures on geological
Videos. Bozeman, transcription and translation: https://youtu.be/h3b9arupxzg Crashcourse: Transcription and Translation - https://youtu.
 Translation Translation Videos Bozeman, transcription and translation: https://youtu.be/h3b9arupxzg Crashcourse: Transcription and Translation - https://youtu.be/itsb2sqr-r0 Translation Translation The
Translation Translation Videos Bozeman, transcription and translation: https://youtu.be/h3b9arupxzg Crashcourse: Transcription and Translation - https://youtu.be/itsb2sqr-r0 Translation Translation The
Protein Structure. W. M. Grogan, Ph.D. OBJECTIVES
 Protein Structure W. M. Grogan, Ph.D. OBJECTIVES 1. Describe the structure and characteristic properties of typical proteins. 2. List and describe the four levels of structure found in proteins. 3. Relate
Protein Structure W. M. Grogan, Ph.D. OBJECTIVES 1. Describe the structure and characteristic properties of typical proteins. 2. List and describe the four levels of structure found in proteins. 3. Relate
F. Piazza Center for Molecular Biophysics and University of Orléans, France. Selected topic in Physical Biology. Lecture 1
 Zhou Pei-Yuan Centre for Applied Mathematics, Tsinghua University November 2013 F. Piazza Center for Molecular Biophysics and University of Orléans, France Selected topic in Physical Biology Lecture 1
Zhou Pei-Yuan Centre for Applied Mathematics, Tsinghua University November 2013 F. Piazza Center for Molecular Biophysics and University of Orléans, France Selected topic in Physical Biology Lecture 1
Phonons and lattice dynamics
 Chapter Phonons and lattice dynamics. Vibration modes of a cluster Consider a cluster or a molecule formed of an assembly of atoms bound due to a specific potential. First, the structure must be relaxed
Chapter Phonons and lattice dynamics. Vibration modes of a cluster Consider a cluster or a molecule formed of an assembly of atoms bound due to a specific potential. First, the structure must be relaxed
Spectroscopic techniques: why, when, where,and how Dr. Roberto GIANGIACOMO
 Spectroscopic techniques: why, when, where,and how Dr. Roberto GIANGIACOMO BASIC INFORMATION Spectroscopy uses light to analyze substances or products by describing the energy transfer between light and
Spectroscopic techniques: why, when, where,and how Dr. Roberto GIANGIACOMO BASIC INFORMATION Spectroscopy uses light to analyze substances or products by describing the energy transfer between light and
2) Matter composed of a single type of atom is known as a(n) 2) A) element. B) mineral. C) electron. D) compound. E) molecule.
 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Which of the following is a particle found in the nucleus of an atom and that has no electrical
MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Which of the following is a particle found in the nucleus of an atom and that has no electrical
Structures of the Molecular Components in DNA and RNA with Bond Lengths Interpreted as Sums of Atomic Covalent Radii
 The Open Structural Biology Journal, 2008, 2, 1-7 1 Structures of the Molecular Components in DNA and RNA with Bond Lengths Interpreted as Sums of Atomic Covalent Radii Raji Heyrovska * Institute of Biophysics
The Open Structural Biology Journal, 2008, 2, 1-7 1 Structures of the Molecular Components in DNA and RNA with Bond Lengths Interpreted as Sums of Atomic Covalent Radii Raji Heyrovska * Institute of Biophysics
Chapter 2 The Chemistry of Biology. Dr. Ramos BIO 370
 Chapter 2 The Chemistry of Biology Dr. Ramos BIO 370 2 Atoms, Bonds, and Molecules Matter - all materials that occupy space and have mass Matter is composed of atoms. Atom simplest form of matter not divisible
Chapter 2 The Chemistry of Biology Dr. Ramos BIO 370 2 Atoms, Bonds, and Molecules Matter - all materials that occupy space and have mass Matter is composed of atoms. Atom simplest form of matter not divisible
Advanced Cell Biology. Lecture 7
 Advanced Cell Biology. Lecture 7 Alexey Shipunov Minot State University January 25, 2013 Shipunov (MSU) Advanced Cell Biology. Lecture 7 January 25, 2013 1 / 43 Outline Questions and answers Structure
Advanced Cell Biology. Lecture 7 Alexey Shipunov Minot State University January 25, 2013 Shipunov (MSU) Advanced Cell Biology. Lecture 7 January 25, 2013 1 / 43 Outline Questions and answers Structure
CHAPTER 2 THE CHEMICAL BASIS OF LIFE
 CHAPTER 2 THE CHEMICAL BASIS OF LIFE CHAPTER OVERVIEW This chapter introduces very basic concepts of chemistry, emphasizing the structure of atoms and how they combine (form bonds). The types of bonds,
CHAPTER 2 THE CHEMICAL BASIS OF LIFE CHAPTER OVERVIEW This chapter introduces very basic concepts of chemistry, emphasizing the structure of atoms and how they combine (form bonds). The types of bonds,
Name: Date: Hour: Unit Four: Cell Cycle, Mitosis and Meiosis. Monomer Polymer Example Drawing Function in a cell DNA
 Unit Four: Cell Cycle, Mitosis and Meiosis I. Concept Review A. Why is carbon often called the building block of life? B. List the four major macromolecules. C. Complete the chart below. Monomer Polymer
Unit Four: Cell Cycle, Mitosis and Meiosis I. Concept Review A. Why is carbon often called the building block of life? B. List the four major macromolecules. C. Complete the chart below. Monomer Polymer
Spectroscopy in Inorganic Chemistry. Vibration and Rotation Spectroscopy
 Spectroscopy in Inorganic Chemistry Vibrational energy levels in a diatomic molecule f = k r r V = ½kX 2 Force constant r Displacement from equilibrium point 2 X= r=r-r eq V = ½kX 2 Fundamental Vibrational
Spectroscopy in Inorganic Chemistry Vibrational energy levels in a diatomic molecule f = k r r V = ½kX 2 Force constant r Displacement from equilibrium point 2 X= r=r-r eq V = ½kX 2 Fundamental Vibrational
Physical Biochemistry. Kwan Hee Lee, Ph.D. Handong Global University
 Physical Biochemistry Kwan Hee Lee, Ph.D. Handong Global University Week 9 CHAPTER 4 Physical Equilibria Basic Concepts Biological organisms are highly inhomogeneous. Different regions of each cell have
Physical Biochemistry Kwan Hee Lee, Ph.D. Handong Global University Week 9 CHAPTER 4 Physical Equilibria Basic Concepts Biological organisms are highly inhomogeneous. Different regions of each cell have
Lattice Vibrations. Chris J. Pickard. ω (cm -1 ) 200 W L Γ X W K K W
 Lattice Vibrations Chris J. Pickard 500 400 300 ω (cm -1 ) 200 100 L K W X 0 W L Γ X W K The Breakdown of the Static Lattice Model The free electron model was refined by introducing a crystalline external
Lattice Vibrations Chris J. Pickard 500 400 300 ω (cm -1 ) 200 100 L K W X 0 W L Γ X W K The Breakdown of the Static Lattice Model The free electron model was refined by introducing a crystalline external
The vibrational spectroscopy of polymers
 D. I. BOWER Reader in Polymer Spectroscopy Interdisciplinary Research Centre in Polymer Science & Technology Department of Physics, University of Leeds W.F. MADDAMS Senior Visiting Fellow Department of
D. I. BOWER Reader in Polymer Spectroscopy Interdisciplinary Research Centre in Polymer Science & Technology Department of Physics, University of Leeds W.F. MADDAMS Senior Visiting Fellow Department of
Single molecule force spectroscopy reveals a highly compliant helical
 Supplementary Information Single molecule force spectroscopy reveals a highly compliant helical folding for the 30 nm chromatin fiber Maarten Kruithof, Fan-Tso Chien, Andrew Routh, Colin Logie, Daniela
Supplementary Information Single molecule force spectroscopy reveals a highly compliant helical folding for the 30 nm chromatin fiber Maarten Kruithof, Fan-Tso Chien, Andrew Routh, Colin Logie, Daniela
DEPARTMENT OF PHYSICS
 Department of Physics 1 DEPARTMENT OF PHYSICS Office in Engineering Building, Room 124 (970) 491-6206 physics.colostate.edu (http://www.physics.colostate.edu) Professor Jacob Roberts, Chair Undergraduate
Department of Physics 1 DEPARTMENT OF PHYSICS Office in Engineering Building, Room 124 (970) 491-6206 physics.colostate.edu (http://www.physics.colostate.edu) Professor Jacob Roberts, Chair Undergraduate
Chapter 17. From Gene to Protein. Biology Kevin Dees
 Chapter 17 From Gene to Protein DNA The information molecule Sequences of bases is a code DNA organized in to chromosomes Chromosomes are organized into genes What do the genes actually say??? Reflecting
Chapter 17 From Gene to Protein DNA The information molecule Sequences of bases is a code DNA organized in to chromosomes Chromosomes are organized into genes What do the genes actually say??? Reflecting
Chem Homework = cm -1, HF; cm -1, H 35 Cl; cm -1, H 81 Br; cm -1, H 127 I
 1. Chem 344 - Homework 10 2. 3. 4. 0 = 4141.3 cm -1, HF; 2988.9 cm -1, H 35 Cl; 2649.7 cm -1, H 81 Br; 2309.5 cm -1, H 127 I 5. 6. 7. Q16.26,27,28,29) Identify the molecular orbitals for F 2 in the images
1. Chem 344 - Homework 10 2. 3. 4. 0 = 4141.3 cm -1, HF; 2988.9 cm -1, H 35 Cl; 2649.7 cm -1, H 81 Br; 2309.5 cm -1, H 127 I 5. 6. 7. Q16.26,27,28,29) Identify the molecular orbitals for F 2 in the images
Ch. 2 Chemistry Comes to Life
 BIOL 164 Human Biology Ch 2 Chemistry Ch. 2 Chemistry Comes to Life Basic Chemistry Helps Us Understand Human Biology Chemistry Science of the composi9on and proper9es of ma:er Carbohydrates, Lipids, Proteins,
BIOL 164 Human Biology Ch 2 Chemistry Ch. 2 Chemistry Comes to Life Basic Chemistry Helps Us Understand Human Biology Chemistry Science of the composi9on and proper9es of ma:er Carbohydrates, Lipids, Proteins,
Low-frequency Vibrations of DNA and Base Pair Opening
 442 Acta Chim. Slov. 2011, 58, 442 447 Scientific paper Low-frequency Vibrations of DNA and Base Pair Opening Franci Merzel 1, * and Mark R. Johnson 2 1 Laboratory for Molecular Modeling, National Institute
442 Acta Chim. Slov. 2011, 58, 442 447 Scientific paper Low-frequency Vibrations of DNA and Base Pair Opening Franci Merzel 1, * and Mark R. Johnson 2 1 Laboratory for Molecular Modeling, National Institute
2.1 Atoms, Ions, and Molecules. 2.1 Atoms, Ions, and Molecules. 2.1 Atoms, Ions, and Molecules. 2.1 Atoms, Ions, and Molecules
 All living things are based on atoms and their interactions. Living things consist of atoms of different elements. An atom is the smallest basic unit of matter. An element is one type of atom. ydrogen
All living things are based on atoms and their interactions. Living things consist of atoms of different elements. An atom is the smallest basic unit of matter. An element is one type of atom. ydrogen
Analysis of the simulation
 Analysis of the simulation Marcus Elstner and Tomáš Kubař January 7, 2014 Thermodynamic properties time averages of thermodynamic quantites correspond to ensemble averages (ergodic theorem) some quantities
Analysis of the simulation Marcus Elstner and Tomáš Kubař January 7, 2014 Thermodynamic properties time averages of thermodynamic quantites correspond to ensemble averages (ergodic theorem) some quantities
Full file at
 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Which of the following is an uncharged particle found in the nucleus of 1) an atom and which has
MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Which of the following is an uncharged particle found in the nucleus of 1) an atom and which has
Cellular Neuroanatomy I The Prototypical Neuron: Soma. Reading: BCP Chapter 2
 Cellular Neuroanatomy I The Prototypical Neuron: Soma Reading: BCP Chapter 2 Functional Unit of the Nervous System The functional unit of the nervous system is the neuron. Neurons are cells specialized
Cellular Neuroanatomy I The Prototypical Neuron: Soma Reading: BCP Chapter 2 Functional Unit of the Nervous System The functional unit of the nervous system is the neuron. Neurons are cells specialized
Spectroscopy: Tinoco Chapter 10 (but vibration, Ch.9)
 Spectroscopy: Tinoco Chapter 10 (but vibration, Ch.9) XIV 67 Vibrational Spectroscopy (Typical for IR and Raman) Born-Oppenheimer separate electron-nuclear motion ψ (rr) = χ υ (R) φ el (r,r) -- product
Spectroscopy: Tinoco Chapter 10 (but vibration, Ch.9) XIV 67 Vibrational Spectroscopy (Typical for IR and Raman) Born-Oppenheimer separate electron-nuclear motion ψ (rr) = χ υ (R) φ el (r,r) -- product
Anatoly B. Kolomeisky. Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS?
 Anatoly B. Kolomeisky Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS? Motor Proteins Enzymes that convert the chemical energy into mechanical
Anatoly B. Kolomeisky Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS? Motor Proteins Enzymes that convert the chemical energy into mechanical
Reading Assignments. A. Genes and the Synthesis of Polypeptides. Lecture Series 7 From DNA to Protein: Genotype to Phenotype
 Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Chiral selection in wrapping, crossover, and braiding of DNA mediated by asymmetric bend-writhe elasticity
 http://www.aimspress.com/ AIMS Biophysics, 2(4): 666-694. DOI: 10.3934/biophy.2015.4.666 Received date 28 August 2015, Accepted date 29 October 2015, Published date 06 November 2015 Research article Chiral
http://www.aimspress.com/ AIMS Biophysics, 2(4): 666-694. DOI: 10.3934/biophy.2015.4.666 Received date 28 August 2015, Accepted date 29 October 2015, Published date 06 November 2015 Research article Chiral
Computational Biology: Basics & Interesting Problems
 Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
Chemical Principles and Biomolecules (Chapter 2) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Eastern Campus
 Chemical Principles and Biomolecules (Chapter 2) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Eastern Campus Primary Source for figures and content: Tortora, G.J. Microbiology
Chemical Principles and Biomolecules (Chapter 2) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Eastern Campus Primary Source for figures and content: Tortora, G.J. Microbiology
Matter and Substances Section 3-1
 Matter and Substances Section 3-1 Key Idea: All matter is made up of atoms. An atom has a positively charges core surrounded by a negatively charged region. An atom is the smallest unit of matter that
Matter and Substances Section 3-1 Key Idea: All matter is made up of atoms. An atom has a positively charges core surrounded by a negatively charged region. An atom is the smallest unit of matter that
MOLECULAR SPECTROSCOPY
 MOLECULAR SPECTROSCOPY First Edition Jeanne L. McHale University of Idaho PRENTICE HALL, Upper Saddle River, New Jersey 07458 CONTENTS PREFACE xiii 1 INTRODUCTION AND REVIEW 1 1.1 Historical Perspective
MOLECULAR SPECTROSCOPY First Edition Jeanne L. McHale University of Idaho PRENTICE HALL, Upper Saddle River, New Jersey 07458 CONTENTS PREFACE xiii 1 INTRODUCTION AND REVIEW 1 1.1 Historical Perspective
2.1 Atoms, Ions, and Molecules
 2.1 Atoms, Ions, and Molecules Living things consist of atoms of different elements. An atom is the smallest basic unit of matter. An element is one type of atom. 6 elements make up 99% of all living things
2.1 Atoms, Ions, and Molecules Living things consist of atoms of different elements. An atom is the smallest basic unit of matter. An element is one type of atom. 6 elements make up 99% of all living things
MBLG lecture 5. The EGG! Visualising Molecules. Dr. Dale Hancock Lab 715
 MBLG lecture 5 Dr. Dale Hancock D.Hancock@mmb.usyd.edu.au Lab 715 The EGG! Visualising Molecules In molecular biology and biochemistry it is better to view molecules as killer pythons rather than smarties.
MBLG lecture 5 Dr. Dale Hancock D.Hancock@mmb.usyd.edu.au Lab 715 The EGG! Visualising Molecules In molecular biology and biochemistry it is better to view molecules as killer pythons rather than smarties.
Name: Date: Period: Biology Notes: Biochemistry Directions: Fill this out as we cover the following topics in class
 Name: Date: Period: Biology Notes: Biochemistry Directions: Fill this out as we cover the following topics in class Part I. Water Water Basics Polar: part of a molecule is slightly, while another part
Name: Date: Period: Biology Notes: Biochemistry Directions: Fill this out as we cover the following topics in class Part I. Water Water Basics Polar: part of a molecule is slightly, while another part
