Acid-labile ATP and/or ADP/P i Binding to the Tetraprotomeric Form of Na/K-ATPase Accompanying Catalytic Phosphorylation-Dephosphorylation Cycle*
|
|
|
- Martha Wells
- 6 years ago
- Views:
Transcription
1 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 274, No. 45, Issue of November 5, pp , by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Acid-labile ATP and/or ADP/P i Binding to the Tetraprotomeric Form of Na/K-ATPase Accompanying Catalytic Phosphorylation-Dephosphorylation Cycle* (Received for publication, July 30, 1999, and in revised form, August 18, 1999) Takeshi Yokoyama, Shunji Kaya, Kazuhiro Abe, Kazuya Taniguchi, Tsuyoshi Katoh, Michio Yazawa, Yutaro Hayashi, and Sven Mårdh** From Biological Chemistry and Bioorganic Chemistry, Division of Chemistry, Graduate School of Science, Hokkaido University, Sapporo , Japan, the Department of Biochemistry, Kyorin University School of Medicine, Mitaka , Japan, and the **Department of Biomedicine and Surgery, Faculty of Health Sciences, Linköping University, Linköping S , Sweden The Na/K-ATPase has been shown to bind 1 and 0.5 mol of 32 P/mol of -chain in the presence [ - 32 P]ATP and [ - 32 P]ATP, respectively, accompanied by a maximum accumulation of 0.5 mol of ADP-sensitive phosphoenzyme (NaE1P) and potassium-sensitive phosphoenzyme (E2P). The former accumulation was followed by the slow constant liberation of P i, but the latter was accompanied with a rapid 0.25 mol of acid-labile P i burst. The rubidium (potassium congener)-occluded enzyme ( 1.7 mol of rubidium/mol of -chain) completely lost rubidium on the addition of sodium magnesium. Further addition of 100 M [ - 32 P]ATP and [ - 32 P]ATP, both induced 0.5 mol of 32 P-ATP binding to the enzyme and caused accumulation of 1 mol of rubidium/mol of -chain, accompanied by a rapid 0.5 mol of P i burst with no detectable phosphoenzyme under steady state conditions. Electron microscopy of rotary-shadowed soluble and membrane-bound Na/K-ATPases and an antibody-na/k-atpase complex, indicated the presence of tetraprotomeric structures ( ) 4. These and other data suggest that Na/K-ATP hydrolysis occurs via four parallel paths, the sequential appearance of (NaE1P:E ATP) 2, (E2P:E ATP:E2P:E ADP/P i ), and (KE2:E ADP/P i ) 2, each of which has been previously referred to as NaE1P, E2P, and KE2, respectively. It is generally accepted that Na/K-dependent ATP hydrolysis (1) occurs via the sequential appearance of phosphoenzymes and dephosphoenzymes (Scheme I), accompanied by the active transport of 3 sodium and 2 potassium ions across the membranes (2 6). The maximum ligand binding or occlusion and phosphorylation capacity per catalytic subunit ( -chain) reported are generally taken as evidence for the functional subunit being a protomer (7 9). However, the maximum phosphorylation capacity of these enzymes has been unequivocally shown to be 0.5 mol/mol of -chain during sodium-dependent ATP hydrolysis (10) even though the binding stoichiometry is 1 * This work was supported in part by grants-in-aid for Scientific Research and an International Scientific Research Program from the Ministry of Education, Science, Sports and Culture of Japan and for the promotion of science from the Novartis Foundation and The Swedish Medical Research Council (Project 4X-4965). The costs of publication of this article were defrayed in part by the payment of page charges. This article must therefore be hereby marked advertisement in accordance with 18 U.S.C. Section 1734 solely to indicate this fact. To whom correspondence should be addressed: Biological Chemistry, Graduate School of Science, Hokkaido University, Sapporo , Japan. Tel.:/Fax: ; ktan@ccms1.hucc. hokudai.ac.jp. mol of ouabain/mol. Data concerning conformational changes of Na/K-ATPase in real time (11) and a quarter site phosphorylation of the -chain by ATP (10) and P i (12 14) as well as other kinetic data (15, 16) support the hypothesis (11) that four different ATP binding sites to the tetraprotomeric enzyme form, ( ) 4, induce out of phase conformational changes in Na/K-ATPase (14, 17). Therefore, questions arise as to the stoichiometry of ATP binding to the phosphorylated or dephosphorylated (cation-occluded) enzyme states during ATP hydrolysis as well as the gross molecular structure of the enzyme. MATERIALS AND METHODS Methods for the purification of Na/K-ATPase from kidneys of pig (14) and dog (18) and for the estimation of the amount of EP from [ - 32 P]ATP (15) and bound 32 P (19) have been described previously except that the enzyme (2 4 mg of protein/ml) was incubated with 50 l ofaph7.4 solution containing 25 mm imidazole HCl, 0.1 mm EDTA-Tris, 25 mm sucrose, 2 M NaCl, or changing the concentrations of NaCl to maintain the ionic strength at 2 M with choline chloride or 16 mm sodium and 0.43 mm MgCl 2 and various concentrations of [ - or - 32 P]ATP and 50 mm [ 3 H]glucose to estimate water space. Approximately 25 l of the mixture was applied to double membranes, an upper Bio-Rad nitrocellulose membrane with a pore size of 0.45 m with the lower type AA Millipore filter with a pore size of 0.80 m under vacuum. After 5 10 s, the Millipore filters were incubated with 1 ml of a solution containing cold 5mM ATP-Tris and 50 mm glucose. An aliquot of both the sample and the reaction mixture before filtration was taken and mixed with scintillation mixture and counted. To obtain maximum rubidium occlusion, pig enzyme preparations giving 2 nmol of phosphoenzyme/mg of protein were incubated (0.5 mg of protein/ml) with 100 l of a buffer solution at ph 7.4 containing 25 mm imidazole HCl, 0.1 mm EDTA-Tris, 25 mm sucrose with 320 M 86 RbCl without or with 10 mm RbCl at 25 C. After 60 min, the samples were diluted with 20 ml of ice-cold buffer as described above except 320 M 86 Rb was replaced with 30 mm KCl with or without 10 mm RbCl. The rubidium-occluded enzymes were isolated on a Millipore membrane (20) and measured as described previously (19). To estimate rubidium occlusion via E2P, pig kidney enzymes were also incubated with 320 M 86 RbCl in the presence of variable concentrations of ATP in the presence of 16 mm NaCl 1mM CaCl 2 or 160 mm NaCl 0.43 (or 4) mm MgCl 2 for 10 s at 0 C. RESULTS AND DISCUSSION Fig. 1 shows that the amount of bound 32 P in the presence of 0.43 mm magnesium 2 M sodium with increasing concentrations of [ - 32 P]ATP and [ - 32 P]ATP increased to give saturation at 1 and 0.5 mol/mol of -chain (K ), respectively. The maximum amount (10) of NaE1P 1 (14, 17, 21), estimated 1 The abbreviations used are: NaE1P, sodium-occluded ADP-sensitive phosphoenzyme; NaEATP, Na/K-ATPase complexed with sodium and ATP in the presence of magnesium; E2P, potassium-sensitive phosphoenzyme; RbE2, rubidium-occluded enzyme; KE2, potassium-occluded enzyme; E ATP, trichloroacetic acid-labile ATP-Na/K-ATPase This paper is available on line at
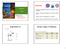
2 Tetraprotomeric Na/K-ATPase FIG. 1. Concentration dependence of [ - 32 P]ATP and [ - 32 P]ATP on the amount of trichloroacetic acid-labile bound 32 P and trichloroacetic acid stable phosphoenzyme (EP). Bound 32 P in the presence of [ - 32 P]ATP (closed circles) or[ - 32 P]ATP (open circles) was estimated from the difference between the counts of the reaction mixture and the counts of the filtrate in the Millipore filter. The amounts of 32 P bound/mg of protein were corrected as mole/mole -chain from the maximum amount of EP to be 0.5 mole/mole -chain (11). The x axis shows the initial concentrations of ATP added. Data shown in this paper are the means S.D. for three or four samples. as trichloroacetic acid-stable E 32 P formed from [ - 32 P]ATP under steady state conditions, was 0.5 mol/mol of -chain (K M) as has been shown previously (10). These data show that 1 mol of 32 P bound in the presence of [ - 32 P]ATP was due to 0.5 mol each of NaE1P and trichloroacetic acid-labile enzymebound ATP or P i. Thus, 0.5 mol of 32 P bound in the presence of [ - 32 P]ATP was due to the trichloroacetic acid-labile enzymebound ATP or ADP. A similar stoichiometry of 32 P binding and phosphorylation was also observed (not shown) in the dog kidney enzyme, giving 3.5 nmol of phosphoenzyme/mg of protein as in the case of the pig kidney enzyme. These data can be explained by the presence of a diprotomer, designated as NaE1P:E ATP or NaE1P:E ADP/P i. The binding of 32 P is enzymatic, as evidenced by the fact that the amount of 32 P bound to pig enzymes at 10 s after the addition of 100 M [ - 32 P]ATP and [ - 32 P]ATP, respectively, decreased to 3 and 20%, after a 15-min incubation at 0 C. To investigate the presence of bound 32 PinE2P, the concentrations of sodium were reduced from 2 M to 16 mm to increase the fraction of E2P (14, 17, 21). Fig. 1, inset, shows that the effect of changing sodium concentrations at a constant ionic strength of 2 M maintained with choline chloride had a negligible effect on the stoichiometry of bound 32 P, which reached nearly 1 mol/mol of -chain in the presence of [ - 32 P]ATP. The phosphoenzyme in the presence of 16 mm sodium 1984 mm choline chloride and 100 M [ - 32 P]ATP with magnesium was shown to be a mixture of 0.25 mol of NaE1P and E2P each, and the amount of 32 P bound in the presence of [ - 32 P]ATP was 0.5 mol/mol of -chain. These data suggest the accumulation of 0.25 mol of either NaE1P:E ATP or NaE1P: E ADP/P i and 0.25 mol of either E2P:E ATP or E2P:E ADP/P i. In other words the nonphosphorylated -chain of E2P also bound ATP or ADP/P i as in the case of NaE1P. To investigate whether bound 32 P is present in a rubidiumoccluded enzyme (RbE2), rubidium occlusion was measured complex; E ADP/P i, trichloroacetic acid-labile ADP/P i -Na/K-ATPase complex; (NaE1P:E ATP) 2, (E2P:E ATP:E2P:E ADP/P i ), and (KE2: E ADP/P i ) 2, teteraprotomer form of each enzyme state, which has been previously referred to as NaE1P, E2P, and KE2, respectively; C 12 E 8, octaethylene glycol dodecyl ether; EDC, 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide; FITC, 5 -isothiocyanate. FIG. 2. Concentration dependence of ATP on the amount of occluded rubidium. Rubidium occlusion of pig kidney enzymes was measured as described under Material and Methods. under various conditions as described under Materials and Methods. A rubidium-occluded enzyme (Fig. 2, closed squares), which is formed directly by incubating the enzyme with 0.32 mm rubidium, lost occluded rubidium nearly completely upon the addition of 160 mm sodium with 0.43 or 4 mm magnesium or 16 mm sodium with 1 mm calcium. With increasing concentrations of ATP from 0 to 5 mm, the amount of rubidium occlusion increased to nearly 1 mol and then appeared to decrease to a constant level of 0.5 mol of rubidium/mol of -chain, especially in the presence of 16 mm sodium 1mM calcium. For both 32 P binding in the presence of 100 M [ - 32 P]ATP and [ - 32 P]ATP, the amount of phosphoenzyme, and that of rubidium occlusion, were measured in the presence of 1 mm calcium, 16 mm sodium, and 0.32 mm rubidium. Both 32 P bindings were shown to be 0.5 mol/mol of -chain with 0.02 mol of E 32 P and 1 mol for the occluded Rb/ -chain. These data mean the stoichiometry of rubidium occlusion/ep dephosphorylation, 2, as shown in the case of another potassium congener, 2 thallium-occluded/ep-dephosphorylated (20). They are consistent with the view that 2 mol of rubidium occlusions occur in 1 mol of the -chain, which was phosphorylated to form E2P just prior the binding of rubidium. These data suggest the presence of stoichiometric amounts of both RbE2 and trichloroacetic acid-labile bound 32 P, such for RbE2:E ATP or RbE2:E ADP/P i in the presence of 100 M ATP with sodium and calcium or magnesium. To measure trichloroacetic acid-labile P i more directly, a time course for the phosphorylation and liberation of P i was followed by quenching the reaction with trichloroacetic acid. The addition of ATP in the presence of 2 M sodium with magnesium (Fig. 3A) induced the accumulation of NaE1P (14, 17, 21), which was followed by a slow steady liberation of P i as also shown at 25 C (21). These data indicate the presence of the trichloroacetic acid-labile binding of ATP, but not of P i,tothe nonphosphorylated -chains of NaE1P such as NaE1P:E ATP. Fig. 3B shows the addition of ATP in the presence of 16 mm sodium with magnesium-induced accumulation of E2P (14, 17, 21), which was accompanied by a rapid clear P i burst estimated from the extrapolation of P i liberation at time 0, mol/mol of -chain. The maximum amount of P i burst, estimated from the burst sizes observed via the additions of 25, 50, 100, and 200 M ATP, appeared to give 0.24 mol/mol of -chain with an apparent K m of 43 M. A similar P i burst has also been shown by the addition of 32 M ATP at 25 C (21). These considerations suggest the presence of both 0.25 mol of trichloroacetic acid-labile ATP and ADP/P i binding to nonphosphorylated -chains of E2P, such as E2P:E ATP:E2P:E ADP/P i.
3 31794 Tetraprotomeric Na/K-ATPase SCHEME I. Sequential model for Na/K-ATPase: Post-Albers mechanism. FIG. 3.Time course of EP formation and P I liberation in the accumulation condition of NaE1P, E2P, and RbE2 via E2P. Pig kidney enzymes (1 mg/ml) were incubated with 50 l of imidazole HCl buffer containing 0.43 mm MgCl 2 with 2 M NaCl, 16 mm NaCl, and 16 mm NaCl 320 M RbCl, respectively, for the ATP-induced accumulation of NaE1P (A), E2P (B), and RbE2 (C). The time course of phosphorylation (solid squares) and P i liberation (open symbols) at 0 C were followed after the addition of 50 l of the buffer containing 200 M [ - 32 P]ATP. The reaction was terminated by trichloroacetic acid, and trichloroacetic acid-labile P i was separated by charcoal (32), and the amount of both P i and EP were measured. The amount of P i burst was estimated from the extrapolation of the time course of P i liberation to time 0. The v/ep was estimated from the slope and the maximum amount of EP, 0.5 mol/mol of -chain. Inset, 5 s after the addition of [ - 32 P]ATP, the reaction mixture was diluted with 2 ml of a solution containing the same concentration of ligands present in the phosphorylation reaction except the enzyme was absent and [ - 32 P]ATP was replaced with 100 M nonradioactive ATP-Tris. The reaction was terminated with trichloroacetic acid. An apparent first order rate constant of EP breakdown was estimated. To investigate the effect of bound ATP and/or ADP/P i on sodium-dependent ATP hydrolysis, the rate constant of EP breakdown and the turnover number, v/ep, was compared. Fig. 3B and the inset show that the rate constant and v/ep are, respectively, around 0.07/s and /s. A value of v/ep, /s, was also obtained in a similar experiment. The rate constant is approximately half the value of v/ep. These data clearly indicate that ATP hydrolysis occurs via both E2P and enzyme-bound ATP and/or ADP/P i. Fig. 3C shows that the addition of ATP to the enzyme in the presence of 16 mm sodium, 0.43 mm magnesium, and 0.32 mm rubidium induced a rapid burst of P i, followed by a slow steady increase of P i production without detectable phosphoenzyme. The magnitude of the burst, 0.42 mol/mol of -chain, was close to the value of the maximum amount of EP in the absence of rubidium, 0.5 mol/mol of -chain. These data show that the trichloroacetic acid-labile binding of ADP/P i occurs to the nonrubidium-occluded -chain of the enzyme in the form of RbE2: E ADP/P i. The data suggest that ATP hydrolysis occurs via KE2:E ADP/P i, which is considered to be a KE2 P i complex, following the breakdown of E2P (Scheme I), as shown from the initial burst of P i production (22). The data described above and other data (10 15) strongly suggest a tetraprotomeric structure for the enzyme. To obtain direct evidence for this, both solubilized and membrane-bound Na/K-ATPases were rotary-shadowed and observed by electron microscopy. Purified membrane-bound dog kidney Na/K- ATPase was solubilized (18) with octaethylene glycol dodecyl ether (C 12 E 8 ) in the presence of 100 mm potassium acetate and subjected to gel filtration at 0 C. The ATP binding capacity 2 appeared as three different peaks with molecular weights of , , and When the first peak fraction was examined, tetrameric, dimeric, and monomeric structures, which may correspond to tetraprotomer, diprotomer, and protomer ( ), respectively, were observed (Fig. 4, A D). The protomer showed a pear-like shape 21 nm in length and 11 nm in the width at the widest portion and appeared to be a unitary structure of the diprotomer and tetraprotomer. The percentages of these structures were 32, 39, and 27%, respectively, and higher oligomeric structures constituted only 2% (n 151). The tetrameric, dimeric, and monomeric molecules were 29, 49, and 21%, respectively, for the second fraction (n 200) and 18, 38, and 43%, respectively, for the third fraction (n 208). When the structure of the molecules contained in the first peak fraction were fixed by crosslinking with a zero-length cross-linker, 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide (EDC), the morphology of these structures was unchanged (Fig. 4, B and E), but their relative amounts were different; the higher oligomeric and tetrameric structures increased to 18 and 39%, respectively, and the monomeric structure decreased to 5% (n 153). Therefore, it is possible that native enzyme could exist in a tetrameric form, which tends to dissociate into dimeric and monomeric forms when solubilized. Electron microscopy of the membrane-bound pig enzyme, which had been treated with EDC prior to solubilization with C 12 E 8, also showed tetrameric, dimeric, and monomeric structures, which were morphologically similar to those observed for the dog enzyme (Fig. 4, A E and F H). To confirm that the molecules observed were, in fact, Na/K-ATPase molecules, the enzyme (10) containing 0.9 mol/mol of -chain of Lys-501 labeled with fluorescein 5 -isothiocyanate (FITC) was com- 2 Y. Hayashi, unpublished data.
4 Tetraprotomeric Na/K-ATPase FIG. 5. Hypothetical model of tetraprotomeric Na/K-ATPase during ATP hydrolysis. Large open circles and triangles, respectively, designate the sodium-bound and potassium-occluded protomer of Na/K-ATPase and the small circles and triangles, respectively, designate 3 sodium and 2 potassium ions, and the corresponding closed symbols designate occluded cations. Shaded symbols designate the difference in the reactivity of NaE1P to acetate (16) or p-nitrophenol (15) and ATP sensitivity of the rubidium occluded form of the enzyme (Fig. 2). Trichloroacetic acid-labile ATP or ADP/P i bound to protomers are also shown. Open circles with ATP- or ADP/P i -bound forms might either contain or occlude sodium, and the KE2 form in the presence of 3 5mM ATP with magnesium and sodium may decrease to a quarter (Fig. 2) with increase in the amount of bound ATP or ADP/P i or phosphoenzyme (37). These remain to be determined. Under nonphosphorylated conditions as shown in the dotted parentheses in the figure, the enzyme bound or occluded a maximum of 3 sodium or 2 potassium ions/mol of -chain, respectively (8, 9). FIG. 4.Rotary-shadowed molecules of Na/K-ATPases. Samples of Na/K-ATPase preparations were diluted to 10 g/ml with 60% glycerol (33), sprayed onto mica, rotary-shadowed with platinum, and observed with a Hitachi H-800 electron microscope operated at 75 kv (34). The solubilized dog kidney enzyme was used directly (A D) or after cross-linking with EDC (E) to fix its structure. The membrane-bound pig enzymes without (F I) or with (J) Anti-FITC antibodies (Molecular Probes, Inc.) were treated with EDC and then solubilized with C 12 E 8. The FITC-labeled enzyme ( 0.9 mol labeled at Lys-501/mol of -chain) was prepared as described previously (10). The EDC treatment of these enzymes was carried out on ice overnight with 5 mm EDC in the presence of 10 mm imidazole (ph 7.0) and was terminated with 10 mm dithiothreitol (34). A, monomeric, dimeric, and tetrameric structures of solubilized enzyme in the first peak fractions on gel filtration are indicated by single, double, and triple arrowheads, respectively; B, E, F, plexed with anti-fitc antibodies (Fig. 4, I and J). The antibodies showed the presence of tetramer with 1 4 molecules of the antibodies attached to the wider end of the pear-like structure, which also indicated that the wider end reflected the cytosolic side (Fig. 4J). No antibody binding was observed for the unlabeled enzyme. The tetraprotomeric nature of Na/K-ATPase was proposed nearly 20 years ago (12, 13, 23). A quarter, half and third to fourth, and full site reactivity have been demonstrated by several independent data, collected by us. 1) The stoichiometries of ATP binding (15, 24), rubidium occlusion (Fig. 2) and phosphorylation by ATP (10), P i (14), acetyl phosphate (16), and p-nitrophenyl phosphate (14) and the ratio of ouabain binding to phosphoenzyme (15, 24); 2) half-site reactivities of NaE1P to acetate (16) and p-nitrophenol (15), namely a quarter site reactivity of the -chain (10); 3) the fact that ATP induced four different conformational changes out of phase (11); 4) 32 P binding and the burst size of P i (Fig. 3, B and C); and 5) electron microscopy (Fig. 4). The experiments described in this paper not only show direct biochemical evidence for the presence of a tetraprotomer structure of Na/K-ATPase during ATP hydrolysis but also indicate quite new aspects of the enzyme forms. All reaction intermediates detectable in the Post-Albers scheme (Scheme I) bind ATP and/or ADP/P i. Further experiments are required to clarify the role of tetrameric structure in Na/K-ATP hydrolysis, although changes in the subunit interaction by physiological ligands were already suggested (4, 5, 10, 11, 18). and I, the enzyme molecules with a tetrameric appearance; C and G, enzyme molecules with a dimeric appearance; D and H, enzyme molecules with a monomeric appearance. E, enzyme fixed by cross-linking with EDC; I, FITC-labeled enzyme; J, FITC-labeled enzyme complexed with anti-fitc antibodies. A structure judged as most probable by eye for its magnified image was drawn under each each electron micrograph in B J. The rotary shadowed antidodies observed as triangula or fat Y -shaped structures, as reported previously (35, 36), were drawn as dotted structures in J. The scale bar for B J is indicated at the bottom.
5 31796 Tetraprotomeric Na/K-ATPase To explain the data described above, we propose a tetramer mechanism for ATP hydrolysis (Fig. 5). ATP binding is followed by four parallel paths, which occur at each of the two half-sites for phosphorylation-dephosphorylation and for ATP hydrolysis without acid-stable phosphoenzyme via (NaE1P:E ATP) 2, (E2P:E ATP:E2P:E ADP/P i ), and (KE2:E ADP/P i ) 2, respectively. The sequential formation of E2P from NaE1P and KE2 from E2P (Scheme I) is accompanied by, respectively, hydrolysis of half of the trichloroacetic acid-labile bound ATP to ADP/P i and of another half of the bound ATP to ADP/P i (Fig. 5). The finding related to the stoichiometric amount of trichloroacetic acid-labile ATP and ADP/P i binding and the P i burst via nonphosphorylated protomer(s) explain the long standing discrepancies between the turnover number of the enzyme, v/ep, and the rate constant for the breakdown of EP (5, 20) as well as the origin of the P i burst (21, 22, 25). Dynamic changes in stoichiometry for the binding of ligands and/or the occlusion of Na/K-ATPase (Fig. 2 and see Refs. 5, 6, and 26) suggest oligomeric interactions between protomers (4 11, 18, 26 31), which might induce different phosphorylation site stoichiometry by synthetic substrates such as chromium tetraaquaadenosine 5 -triphosphate (9) and p-nitrophenyl phosphate (15) and by chymotrypsin cleavage (31). If the simultaneous presence of the sodium- and rubidiumbound form of the enzyme becomes clear as an intermediate of ATP hydrolysis such as the RbE2:NaE1 ADP/P i complex, the determination of one of the two transport mechanisms, i.e. simultaneous or consecutive would be crucial. Further experiments, such as the binding stoichiometry of magnesium, sodium, and potassium to these subunits as shown as E ATP and E ADP/P i (Fig. 5) and the ATP-induced subunit interactions will be required for better understanding the molecular mechanism of ATP hydrolysis. REFERENCES 1. Skou, J. C. (1957) Biochim. Biophys. Acta 23, Post, R. L., Hegyvary, C., and Kume, S. (1972) J. Biol. Chem. 247, Albers, R. W. (1976) in The Enzymes of Biological Membranes (Martonossi, A., ed) Vol. 3, pp , Plenum Publishing Corp., New York 4. Askari, A. (1987) J. Bioenerg. Biomembr. 19, Glynn, I. M. (1985) in The Enzymes of Biological Membranes (Martonossi, A., ed) Vol. 3, pp , Plenum Publishing Corp., New York 6. Glynn, I. M. (1993) J. Physiol. (Lond.) 462, Peters, W. H. M., Swarts, H. G. P., De Pont, J. J. H. H. M., Schuurmans Stekhoven, F. M. A. H., and Bonting, S. L. (1981) Nature 290, Matsui, H., Hayashi. Y., Homareda, H., and Taguchi, M. (1983) Curr. Top. Membr. Transp. 19, Vilsen, B., Andersen, J. C., Petersen, J., and Jorgensen, P. L. (1987) J. Biol. Chem. 262, Tsuda, T., Kaya, S., Yokoyama, T., Hayashi, Y., and Taniguchi, K. (1998) J. Biol. Chem. 273, Tsuda, T., Kaya, S., Yokoyama, T., Hayashi, Y., and Taniguchi, K. (1998) J. Biol. Chem. 273, Askari, A., and Huang, W.-H. (1980) Biochem. Biophys. Res. Commun. 93, Askari, A., and Huang, W.-H. (1981) FEBS Lett. 126, Taniguchi, K., Suzuki, K., and Iida, S. (1982) J. Biol. Chem. 257, Yamazaki, A., Kaya, S., Tsuda, T., Araki, Y., Hayashi, Y., and Taniguchi, K. (1994) J. Biochem. (Tokyo) 116, Taniguchi, K., Tosa, H., Suzuki, K., and Kamo, Y. (1988) J. Biol. Chem. 263, Taniguchi, K., Suzuki, K., Kai, D., Matsuoka, I., Tomita, K., and Iida, S. (1984) J. Biol. Chem. 259, Hayashi, Y., Mimura, K., Matsui, H., and Takagi, T. (1989) Biochim. Biophys. Acta 983, Yamaguchi, M., and Tonomura, Y. (1979) J. Biochem. (Tokyo) 86, Rossi, R. C., and Norby, J. G. (1993) J. Biol. Chem. 268, Taniguchi, K., Suzuki, K., Sasaki, T., Shimokobe, H., and Iida, S. (1986) J. Biochem. (Tokyo) 100, Froehlich, J. P., Albers, R. W., Koval, G. J. Goebel, R., and Berman, M. (1975) J. Biol. Chem. 251, Haase, W., and Koespell, H. (1977) Pflügers Arch. 381, Tsuda, T., Kaya, S., Funatsu, H., Hayashi, Y., and Taniguchi, K (1998) J. Biochem. (Tokyo) 123, Froehlich, J. P., Taniguchi, K., Fendler, K., Mahaney, J. S., Thomas, D., D., and Albers, R. W. (1997) Ann. N. Y. Acad. Sci. 834, Esmann, M., and Skou, J. C. (1985) in the Sodium Pump (Glynn, I., and Ellory, C., eds) pp , The Company of Biologists Ltd., 27. Schoner, W., Thönges, D., Hamer, E., Antolovic, R., Buxbaum, E., Willeke, M., Serpersu, E. H., and Scheiner-Bobis, G. (1994) in The Sodium Pump, Structure, Mechanism, Hormonal Control and Its Role in Disease (Bamberg, E., and Schoner, W., eds) pp , Dietrich Steinkopff Verlag GmbH & Co. KG, Darmstadt, Germany 28. Antolovic, R., Hamer, E., Serpersu, E. H., Kost, H., Linnertz, H., Kovarik, Z., and Schoner. W. (1999) Eur. J. Biochem. 260, Hansen, O., Jensen, J., Norby, J. S., and Ottolenghi, P. (1979) Nature 280, Skou, J. C., and Esmann, M. (1992) J. Bioenerg. Biomembr. 24, Liu, G., Xie, Z., Modyanov, N., and Askari, A (1996) FEBS Lett. 390, Blostein, R. (1988) Methods Enzymol. 156, Tyler, J. M., and Branton, D. (1980) J. Ultrastruct. Res. 71, Katoh, T., Konishi, K., and Yazawa, M. (1998) J. Biol. Chem. 273, Waller, G. S., and Lowey, S. (1985) J. Biol. Chem. 260, Kato, T., and Lowey, S. (1989) J. Cell Biol. 109, Suzuki, K., Taniguchi, K., and Iida, S. (1987) J. Biol. Chem. 262,
Structural and functional aspects of gastric proton pump, H +,K + -ATPase
 Kazuhiro ABE, Ph. D. E-mail:kabe@cespi.nagoya-u.ac.jp Structural and functional aspects of gastric proton pump, H +,K + -ATPase In response to food intake, ph of our stomach reaches around 1. This highly
Kazuhiro ABE, Ph. D. E-mail:kabe@cespi.nagoya-u.ac.jp Structural and functional aspects of gastric proton pump, H +,K + -ATPase In response to food intake, ph of our stomach reaches around 1. This highly
Enzymatic Assay of PROTEIN KINASE C
 PRINCIPLE: Histone +? 32 P-ATP Protein Kinase > [ 32 P]-Phosphorylated Histone + ADP Abbreviations used:? 32 P-ATP = Adenosine 5'-Triphosphate? 32 P-labelled ADP = Adenosine 5'-Diphosphate CONDITIONS:
PRINCIPLE: Histone +? 32 P-ATP Protein Kinase > [ 32 P]-Phosphorylated Histone + ADP Abbreviations used:? 32 P-ATP = Adenosine 5'-Triphosphate? 32 P-labelled ADP = Adenosine 5'-Diphosphate CONDITIONS:
Rate Limitation of the Na,K -ATPase Pump Cycle
 Biophysical Journal Volume 81 October 2001 2069 2081 2069 Rate Limitation of the Na,K -ATPase Pump Cycle Christian Lüpfert,* Ernst Grell, Verena Pintschovius, Hans-Jürgen Apell, Flemming Cornelius, and
Biophysical Journal Volume 81 October 2001 2069 2081 2069 Rate Limitation of the Na,K -ATPase Pump Cycle Christian Lüpfert,* Ernst Grell, Verena Pintschovius, Hans-Jürgen Apell, Flemming Cornelius, and
Kinetics of Na -Dependent Conformational Changes of Rabbit Kidney Na,K -ATPase
 1340 Biophysical Journal Volume 75 September 1998 1340 1353 Kinetics of Na -Dependent Conformational Changes of Rabbit Kidney Na,K -ATPase Ronald J. Clarke,* David J. Kane,* Hans-Jürgen Apell, # Milena
1340 Biophysical Journal Volume 75 September 1998 1340 1353 Kinetics of Na -Dependent Conformational Changes of Rabbit Kidney Na,K -ATPase Ronald J. Clarke,* David J. Kane,* Hans-Jürgen Apell, # Milena
Methods in Enzymology. Volume 156. Biomembranes. Part P ATP-Driven Pumps and Related Transport : The Na,K-Pump. Sidney Fleischer Becca Fleischer
 Methods in Enzymology Volume 156 Biomembranes Part P ATP-Driven Pumps and Related Transport : The Na,K-Pump Sidney Fleischer Becca Fleischer CONTRIBUTORS TO VOLUME 156 PREFACE VOLUMES IN SERIES ix Xii
Methods in Enzymology Volume 156 Biomembranes Part P ATP-Driven Pumps and Related Transport : The Na,K-Pump Sidney Fleischer Becca Fleischer CONTRIBUTORS TO VOLUME 156 PREFACE VOLUMES IN SERIES ix Xii
(Na++ K +)-ATPase in artificial lipid vesicles: influence of the concentration of mono- and divalent cations on the pumping rate
 254 Biochimica et Biophysica Acta 862 (1986) 254-264 Elsevier BBA 72961 (Na++ K +)-ATPase in artificial lipid vesicles: influence of the concentration of mono- and divalent cations on the pumping rate
254 Biochimica et Biophysica Acta 862 (1986) 254-264 Elsevier BBA 72961 (Na++ K +)-ATPase in artificial lipid vesicles: influence of the concentration of mono- and divalent cations on the pumping rate
FEBS Letters 532 (2002) 198^202. of the Na,K-ATPase. Received 25 September 2002; revised 26 October 2002; accepted 28 October 2002
 FEBS Letters 532 (2002) 198^202 FEBS 26781 Do H þ ions obscure electrogenic Na þ and K þ binding in the E 1 state of the Na,K-ATPase? Hans-Ju«rgen Apell, Anna Diller University of Konstanz, Biology, Universita«tsstrasse
FEBS Letters 532 (2002) 198^202 FEBS 26781 Do H þ ions obscure electrogenic Na þ and K þ binding in the E 1 state of the Na,K-ATPase? Hans-Ju«rgen Apell, Anna Diller University of Konstanz, Biology, Universita«tsstrasse
In Situ Gelation-Induced Death of Cancer Cells Based on Proteinosomes
 Supporting information for In Situ Gelation-Induced Death of Cancer Cells Based on Proteinosomes Yuting Zhou, Jianmin Song, Lei Wang*, Xuting Xue, Xiaoman Liu, Hui Xie*, and Xin Huang* MIIT Key Laboratory
Supporting information for In Situ Gelation-Induced Death of Cancer Cells Based on Proteinosomes Yuting Zhou, Jianmin Song, Lei Wang*, Xuting Xue, Xiaoman Liu, Hui Xie*, and Xin Huang* MIIT Key Laboratory
Yoshihiro FUKUSHIMA and Yuji TONOMURA Department of Biology, Faculty of Science, Osaka University, Toyonaka, Osaka
 Two Kinds of High Energy Phosphorylated Intermediate, with and without Bound ADP, in the Reaction of Na+-K+-dependent ATPase* Yoshihiro FUKUSHIMA and Yuji TONOMURA Department of Biology, Faculty of Science,
Two Kinds of High Energy Phosphorylated Intermediate, with and without Bound ADP, in the Reaction of Na+-K+-dependent ATPase* Yoshihiro FUKUSHIMA and Yuji TONOMURA Department of Biology, Faculty of Science,
The Effect of Ionic Strength and Specific Anions on Substrate Binding and Hydrolytic Activities of Na,K-ATPase
 Published Online: 1 May, 1997 Supp Info: http://doi.org/10.1085/jgp.109.5.555 Downloaded from jgp.rupress.org on June 19, 2018 The Effect of Ionic Strength and Specific Anions on Substrate Binding and
Published Online: 1 May, 1997 Supp Info: http://doi.org/10.1085/jgp.109.5.555 Downloaded from jgp.rupress.org on June 19, 2018 The Effect of Ionic Strength and Specific Anions on Substrate Binding and
Enzymatic Assay of ACTOMYOSIN ATPase Assay
 Enzymatic Assay of ACTOMYOSIN PRINCIPLE: ATP + H 2 O ATPase > ADP + P i Abbreviations used: ATP = Adenosine 5'-Triphosphate ATPase = Adenosine 5'-Triphosphatase ADP = Adenosine 5'-Diphosphate P i = Inorganic
Enzymatic Assay of ACTOMYOSIN PRINCIPLE: ATP + H 2 O ATPase > ADP + P i Abbreviations used: ATP = Adenosine 5'-Triphosphate ATPase = Adenosine 5'-Triphosphatase ADP = Adenosine 5'-Diphosphate P i = Inorganic
HoOo-ModificQtion of Na,K-ATPase
 HoOo-ModificQtion of,k-atpase Alterations in External and K Stimulation of K Influx Margaret H. Garner, William H. Garner, and Abraham Specror Studies, at steady state, of the,k-atpase dependent influx
HoOo-ModificQtion of,k-atpase Alterations in External and K Stimulation of K Influx Margaret H. Garner, William H. Garner, and Abraham Specror Studies, at steady state, of the,k-atpase dependent influx
A. 50 mm Sodium Lauryl Sulfate Solution (SDS) (Prepare 10 ml in deionized water using Lauryl Sulfate, Sodium Salt, Sigma Prod. No. L-5750.
 SIGMA QUALITY CONTROL TEST PROCEDURE Enzymatic Assay of PHOSPHOLIPASE D 1 (EC 3.1.4.4) PRINCIPLE: L-α-Phosphatidylcholine + 2H 2 O Phospholipase D > Choline + Phosphatidic Acid 2 Choline + O 2 Choline
SIGMA QUALITY CONTROL TEST PROCEDURE Enzymatic Assay of PHOSPHOLIPASE D 1 (EC 3.1.4.4) PRINCIPLE: L-α-Phosphatidylcholine + 2H 2 O Phospholipase D > Choline + Phosphatidic Acid 2 Choline + O 2 Choline
Molecular Mechanism and Regulation in Cation Transport ATPases and Related Genetic Diseases
 Molecular Mechanism and Regulation in Cation Transport ATPases and Related Genetic Diseases Program Thursday, June 15 at RIHGA ROYAL HOTEL KYOTO 17:00-19:00 Registration for SYMPOSIUM (in front of the
Molecular Mechanism and Regulation in Cation Transport ATPases and Related Genetic Diseases Program Thursday, June 15 at RIHGA ROYAL HOTEL KYOTO 17:00-19:00 Registration for SYMPOSIUM (in front of the
Proton-activated Rubidium Transport Catalyzed by the Sodium Pump*
 THE JOURNAL OF BIOLOGICAL CHEMISTRY 81985 by The American Society of Biological Chemists, Inc. Vol. 260, No. 1, Issue of January 25, pp. 829833,1985 Printed in U.S.A. Protonactivated Rubidium Transport
THE JOURNAL OF BIOLOGICAL CHEMISTRY 81985 by The American Society of Biological Chemists, Inc. Vol. 260, No. 1, Issue of January 25, pp. 829833,1985 Printed in U.S.A. Protonactivated Rubidium Transport
The Pre-steady State of the Myosin-Adenosine Triphosphate System
 /. Biochem., 71, 115-124 (1972) The Pre-steady State of the Myosin-Adenosine Triphosphate System XL Formation and Decomposition of the Reactive Myosin-Phosphate- Complex* Akio INOUE, Kazuko SHIBATA-SEKIYA
/. Biochem., 71, 115-124 (1972) The Pre-steady State of the Myosin-Adenosine Triphosphate System XL Formation and Decomposition of the Reactive Myosin-Phosphate- Complex* Akio INOUE, Kazuko SHIBATA-SEKIYA
Secondary Structure of Heart Sarcolemmal Proteins During Interaction with Metallic Cofactors of (Na + + K + )-ATPase
 Gen. Physiol. Biophys. (1984), 3, 31 7-325 317 Secondary Structure of Heart Sarcolemmal Proteins During Interaction with Metallic Cofactors of (Na K )-ATPase N. VRBJAR 1, J. Soós 2 and A. ZIEGELHÔFFER
Gen. Physiol. Biophys. (1984), 3, 31 7-325 317 Secondary Structure of Heart Sarcolemmal Proteins During Interaction with Metallic Cofactors of (Na K )-ATPase N. VRBJAR 1, J. Soós 2 and A. ZIEGELHÔFFER
Biological Sciences 11 Spring Experiment 4. Protein crosslinking
 Biological Sciences 11 Spring 2000 Experiment 4. Protein crosslinking = C - CH 2 - CH 2 - CH 2 - C = H H GA Cl - H 2 N N H 2 Cl - C - CH 2 - CH 2 - CH 2 - CH 2 - CH 2 - CH 2 - C DMS CH 3 CH 3 N - - C -
Biological Sciences 11 Spring 2000 Experiment 4. Protein crosslinking = C - CH 2 - CH 2 - CH 2 - C = H H GA Cl - H 2 N N H 2 Cl - C - CH 2 - CH 2 - CH 2 - CH 2 - CH 2 - CH 2 - C DMS CH 3 CH 3 N - - C -
CELLS NOT YOUR CELL PHONE HOMEOSTASIS: LESSON 5 OVERVIEW TEKS
 Lesson 5: Active Transport Protein Pumps Objectives: In this lesson the student will: CELLS NOT YOUR CELL PHONE HOMEOSTASIS: LESSON 5 OVERVIEW 1. Identify how the unique structure of the cell membrane
Lesson 5: Active Transport Protein Pumps Objectives: In this lesson the student will: CELLS NOT YOUR CELL PHONE HOMEOSTASIS: LESSON 5 OVERVIEW 1. Identify how the unique structure of the cell membrane
Certificate of Analysis
 Certificate of Analysis 10 Old Barn Road Lake Placid, NY 12946 Technical Support: T: 800 548-7853 F: 518 523-4513 email: techserv@upstate.com Sales Department: T: 800 233-3991 F: 781 890-7738 Licensing
Certificate of Analysis 10 Old Barn Road Lake Placid, NY 12946 Technical Support: T: 800 548-7853 F: 518 523-4513 email: techserv@upstate.com Sales Department: T: 800 233-3991 F: 781 890-7738 Licensing
15,UM for ATP, 65,uM for adenylyl-imidodiphosphate (AMP-PNP) and 180 /ZM for
 J. Physiol. (198), 299, pp. 367-383 367 With 6 text-ftgures Printed in Great Britain THE EQUILIBRIUM BETWEEN DIFFERENT CONFORMATIONS OF THE UNPHOSPHORYLATED SODIUM PUMP: EFFECTS OF ATP AND OF POTASSIUM
J. Physiol. (198), 299, pp. 367-383 367 With 6 text-ftgures Printed in Great Britain THE EQUILIBRIUM BETWEEN DIFFERENT CONFORMATIONS OF THE UNPHOSPHORYLATED SODIUM PUMP: EFFECTS OF ATP AND OF POTASSIUM
Supporting Information. Visualization of Phagosomal Hydrogen Peroxide Production by A Novel Fluorescent Probe That Is Localized via SNAP-tag Labeling
 Supporting Information Visualization of Phagosomal Hydrogen Peroxide Production by A Novel Fluorescent Probe That Is Localized via SNAP-tag Labeling Masahiro Abo, Reiko Minakami, Kei Miyano, Mako Kamiya,
Supporting Information Visualization of Phagosomal Hydrogen Peroxide Production by A Novel Fluorescent Probe That Is Localized via SNAP-tag Labeling Masahiro Abo, Reiko Minakami, Kei Miyano, Mako Kamiya,
SUPPLEMENTARY INFORMATION
 Ultrafast proton transport in sub-1-nm diameter carbon nanotube porins Ramya H. Tunuguntla, 1 Frances I. Allen, 2 Kyunghoon Kim, 1, Allison Belliveau, 1 Aleksandr Noy 1,3, * * Correspondence to AN: noy1@llnl.gov
Ultrafast proton transport in sub-1-nm diameter carbon nanotube porins Ramya H. Tunuguntla, 1 Frances I. Allen, 2 Kyunghoon Kim, 1, Allison Belliveau, 1 Aleksandr Noy 1,3, * * Correspondence to AN: noy1@llnl.gov
Enzymatic Assay of PHOSPHOLIPASE D (EC )
 PRINCIPLE: L-α-Phosphatidylcholine + H 2 O Phospholipase D > Choline + Phosphatidic Acid Choline + O 2 + H 2 O Choline Oxidase > Betaine Aldehyde + H 2 O 2 2H 2 O 2 + 4-AAP + Phenol Peroxidase > 4H 2 O
PRINCIPLE: L-α-Phosphatidylcholine + H 2 O Phospholipase D > Choline + Phosphatidic Acid Choline + O 2 + H 2 O Choline Oxidase > Betaine Aldehyde + H 2 O 2 2H 2 O 2 + 4-AAP + Phenol Peroxidase > 4H 2 O
Action of the Protease from Streptomyces cellulosae on L-Leu-Gly
 /. Biochem. 99, 1625-1630 (1986) Action of the Protease from Streptomyces cellulosae on L-Leu-Gly Tetsuo MURO, Yoshio TOMINAGA, and Shigetaka OKADA Osaka Municipal Technical Research Institute, Joto-ku,
/. Biochem. 99, 1625-1630 (1986) Action of the Protease from Streptomyces cellulosae on L-Leu-Gly Tetsuo MURO, Yoshio TOMINAGA, and Shigetaka OKADA Osaka Municipal Technical Research Institute, Joto-ku,
Orthophosphate-promoted Ouabain Binding to Na/K Pumps of Resealed Red Cell Ghosts
 THE JOURNAL OF BIOLOGICAL CHEMISTRY 0 1992 by The American Society for Biochemistry and Molecular Biology, Inc. Vol. 267, No. 3, Issue of January 25, pp. 1596-1602,1992 Printed in U.S.A. Orthophosphate-promoted
THE JOURNAL OF BIOLOGICAL CHEMISTRY 0 1992 by The American Society for Biochemistry and Molecular Biology, Inc. Vol. 267, No. 3, Issue of January 25, pp. 1596-1602,1992 Printed in U.S.A. Orthophosphate-promoted
Analysis of nucleotide binding to p97 reveals the properties of a tandem AAA hexameric ATPase
 SUPPLEMENTARY INFORMATION Analysis of nucleotide binding to p97 reveals the properties of a tandem AAA hexameric ATPase Louise C Briggs, Geoff S Baldwin, Non Miyata, Hisao Kondo, Xiaodong Zhang, Paul S
SUPPLEMENTARY INFORMATION Analysis of nucleotide binding to p97 reveals the properties of a tandem AAA hexameric ATPase Louise C Briggs, Geoff S Baldwin, Non Miyata, Hisao Kondo, Xiaodong Zhang, Paul S
b) What is the gradient at room temperature? Du = J/molK * 298 K * ln (1/1000) = kj/mol
 Chem350 Practice Problems Membranes 1. a) What is the chemical potential generated by the movement of glucose by passive diffusion established by a 1000 fold concentration gradient at physiological temperature?
Chem350 Practice Problems Membranes 1. a) What is the chemical potential generated by the movement of glucose by passive diffusion established by a 1000 fold concentration gradient at physiological temperature?
CHAPTER 7: Solutions & Colloids 7.2 SOLUBILITY. Degrees of Solution. Page PHYSICAL STATES of SOLUTIONS SOLUTION
 CHAPTER 7: Solutions & Colloids Predict the relative solubility of materials on the basis of polarity Describe solution formation in terms of solutesolvent interactions Calculate solution concentrations
CHAPTER 7: Solutions & Colloids Predict the relative solubility of materials on the basis of polarity Describe solution formation in terms of solutesolvent interactions Calculate solution concentrations
Keywords: dimer dissociation, Streptomyces subtilisin inhibitor, rate constant
 J. Biol. Macromol. 8(2), 38-47 (2008) Article Kinetic study on the dissociation of a dimeric protein, Streptomyces Subtilisin Inhibitor Keiko Momma 1,2, Ben ichiro Tonomura, and Keitaro Hiromi Department
J. Biol. Macromol. 8(2), 38-47 (2008) Article Kinetic study on the dissociation of a dimeric protein, Streptomyces Subtilisin Inhibitor Keiko Momma 1,2, Ben ichiro Tonomura, and Keitaro Hiromi Department
Enzymatic Assay of CASEIN KINASE II
 PRINCIPLE: Casein +?- 32 P-ATP Casein Kinase II > [ 32 P]-Phosphorylated Casein + ADP CONDITIONS: T = 37 C, ph 7.5 METHOD: Radiolabelled Stop Reaction REAGENTS: A. 100 mm HEPES Buffer, ph 7.5 at 37 C (Enzyme
PRINCIPLE: Casein +?- 32 P-ATP Casein Kinase II > [ 32 P]-Phosphorylated Casein + ADP CONDITIONS: T = 37 C, ph 7.5 METHOD: Radiolabelled Stop Reaction REAGENTS: A. 100 mm HEPES Buffer, ph 7.5 at 37 C (Enzyme
A hypothetical model of the influence of inorganic phosphate on the kinetics of pyruvate kinase
 BioSystems 54 (1999) 71 76 www.elsevier.com/locate/biosystems A hypothetical model of the influence of inorganic phosphate on the kinetics of pyruvate kinase Marian Kuczek * Institute of Biology and En
BioSystems 54 (1999) 71 76 www.elsevier.com/locate/biosystems A hypothetical model of the influence of inorganic phosphate on the kinetics of pyruvate kinase Marian Kuczek * Institute of Biology and En
[180]ATP species formation (180-oxygen exchange/cooperativity/photophosphorylation/31p NMR)
![[180]ATP species formation (180-oxygen exchange/cooperativity/photophosphorylation/31p NMR) [180]ATP species formation (180-oxygen exchange/cooperativity/photophosphorylation/31p NMR)](/thumbs/73/68745809.jpg) Proc. Natl. Acad. Sci. USA Vol. 76, No. 8, pp. 3646-365, August 1979 Biochemistry Subunit interaction during catalysis: Alternating site cooperativity in photophosphorylation shown by substrate modulation
Proc. Natl. Acad. Sci. USA Vol. 76, No. 8, pp. 3646-365, August 1979 Biochemistry Subunit interaction during catalysis: Alternating site cooperativity in photophosphorylation shown by substrate modulation
THALLIUM AND CESIUM IN MUSCLE CELLS COMPETE FOR THE ADSORPTION SITES NORMALLY OCCUPlED BY K+
 THALLIUM AND CESIUM IN MUSCLE CELLS COMPETE FOR THE ADSORPTION SITES NORMALLY OCCUPlED BY K+ GILBERT N. LING Department of Molecular Biology. Pennsylvania Hospital. Philadelphia, Pennsylvania 19107 Reprit~red
THALLIUM AND CESIUM IN MUSCLE CELLS COMPETE FOR THE ADSORPTION SITES NORMALLY OCCUPlED BY K+ GILBERT N. LING Department of Molecular Biology. Pennsylvania Hospital. Philadelphia, Pennsylvania 19107 Reprit~red
NUTRITION AND METABOLISM OF MARINE BACTERIA'
 NUTRITION AND METABOLISM OF MARINE BACTERIA' XII. ION ACTIVATION OF ADENOSINE TRIPHOSPHATASE IN MEMBRANES OF MARINE BACTERIAL CELLS GABRIEL R. DRAPEAU AND ROBERT A. MAcLEOD Department of Bacteriology,
NUTRITION AND METABOLISM OF MARINE BACTERIA' XII. ION ACTIVATION OF ADENOSINE TRIPHOSPHATASE IN MEMBRANES OF MARINE BACTERIAL CELLS GABRIEL R. DRAPEAU AND ROBERT A. MAcLEOD Department of Bacteriology,
Mukogawa Women s University Nishinomiya, Hyogo , Japan 2 Hyogo Nutrition Vocational College, Nishinomiya, Hyogo , Japan
 J. Biol. Macromol., 4(1) 13-22 (2004) Article Stimulating Effect of High Concentration of Calcium Ion on the Polymerization of the Tubulin-Colchicine Complex. Relationship between Magnesium and Calcium
J. Biol. Macromol., 4(1) 13-22 (2004) Article Stimulating Effect of High Concentration of Calcium Ion on the Polymerization of the Tubulin-Colchicine Complex. Relationship between Magnesium and Calcium
Extracellular Allosteric Na D Binding to the Na D,K D -ATPase in Cardiac Myocytes
 Biophysical Journal Volume 105 December 2013 2695 2705 2695 Extracellular Allosteric Na D Binding to the Na D,K D -ATPase in Cardiac Myocytes Alvaro Garcia, { Natasha A. S. Fry, { Keyvan Karimi, { Chia-chi
Biophysical Journal Volume 105 December 2013 2695 2705 2695 Extracellular Allosteric Na D Binding to the Na D,K D -ATPase in Cardiac Myocytes Alvaro Garcia, { Natasha A. S. Fry, { Keyvan Karimi, { Chia-chi
Carbamoyl Phosphate Synthetase from Escherichia coli Does Not Catalyze the Dehydration of Bicarbonate to Carbon Dioxide 1
 BIOORGANIC CHEMISTRY 26, 255 268 (1998) ARTICLE NO. BH981103 Carbamoyl Phosphate Synthetase from Escherichia coli Does Not Catalyze the Dehydration of Bicarbonate to Carbon Dioxide 1 Grant E. Gibson, Leisha
BIOORGANIC CHEMISTRY 26, 255 268 (1998) ARTICLE NO. BH981103 Carbamoyl Phosphate Synthetase from Escherichia coli Does Not Catalyze the Dehydration of Bicarbonate to Carbon Dioxide 1 Grant E. Gibson, Leisha
Solutions Solubility. Chapter 14
 Copyright 2004 by Houghton Mifflin Company. Solutions Chapter 14 All rights reserved. 1 Solutions Solutions are homogeneous mixtures Solvent substance present in the largest amount Solute is the dissolved
Copyright 2004 by Houghton Mifflin Company. Solutions Chapter 14 All rights reserved. 1 Solutions Solutions are homogeneous mixtures Solvent substance present in the largest amount Solute is the dissolved
Kinetic Studies of the Gastric H,K-ATPase
 THE JOURNAL OF BIOLOGICAL CHEMISTRY 1988 by The American Society for Biochemistry and Moleculal Biology, Inc Vol. 263, No. 36, Issue of December 25, pp. 19618-19625,1988 Printed in U.S.A. Kinetic Studies
THE JOURNAL OF BIOLOGICAL CHEMISTRY 1988 by The American Society for Biochemistry and Moleculal Biology, Inc Vol. 263, No. 36, Issue of December 25, pp. 19618-19625,1988 Printed in U.S.A. Kinetic Studies
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09450 Supplementary Table 1 Summary of kinetic parameters. Kinetic parameters were V = V / 1 K / ATP and obtained using the relationships max ( + m [ ]) V d s /( 1/ k [ ATP] + 1 k ) =,
doi:10.1038/nature09450 Supplementary Table 1 Summary of kinetic parameters. Kinetic parameters were V = V / 1 K / ATP and obtained using the relationships max ( + m [ ]) V d s /( 1/ k [ ATP] + 1 k ) =,
Thermal denaturation and autolysis profiles of myofibrillar proteins of mantle muscle of jumbo squid Docidicus gigas
 Blackwell Science, LtdOxford, UK FISFisheries Science0919-92682003 Blackwell Science Asia Pty Ltd 691February 2003 fistest.doc Denaturation and autolysis of jumbo squid K Konno et al. 10.1046/j.0919-9268.2002.fistest.doc.x
Blackwell Science, LtdOxford, UK FISFisheries Science0919-92682003 Blackwell Science Asia Pty Ltd 691February 2003 fistest.doc Denaturation and autolysis of jumbo squid K Konno et al. 10.1046/j.0919-9268.2002.fistest.doc.x
ACTIVE TRANSPORT AND GLUCOSE TRANSPORT. (Chapter 14 and 15, pp and pp )
 ACTIVE TRANSPORT AND GLUCOSE TRANSPORT (Chapter 14 and 15, pp 140-143 and pp 146-151) Overview Active transport is the movement of molecules across a cell membrane in the direction against their concentration
ACTIVE TRANSPORT AND GLUCOSE TRANSPORT (Chapter 14 and 15, pp 140-143 and pp 146-151) Overview Active transport is the movement of molecules across a cell membrane in the direction against their concentration
Effect of Clotrimazole on the Pump Cycle of the Na,K-ATPase
 Biophysical Journal Volume 95 August 2008 1813 1825 1813 Effect of Clotrimazole on the Pump Cycle of the Na,K-ATPase Gianluca Bartolommei,* Nadège Devaux, y Francesco Tadini-Buoninsegni,* MariaRosa Moncelli,*
Biophysical Journal Volume 95 August 2008 1813 1825 1813 Effect of Clotrimazole on the Pump Cycle of the Na,K-ATPase Gianluca Bartolommei,* Nadège Devaux, y Francesco Tadini-Buoninsegni,* MariaRosa Moncelli,*
Biology Unit 2 Chemistry of Life (Ch. 6) Guided Notes
 Name Biology Unit 2 Chemistry of Life (Ch. 6) Guided Notes Atoms, Elements, and Chemical Bonding I can draw atom models and identify the # protons, # neutrons, and # electrons in an atom. I can identify
Name Biology Unit 2 Chemistry of Life (Ch. 6) Guided Notes Atoms, Elements, and Chemical Bonding I can draw atom models and identify the # protons, # neutrons, and # electrons in an atom. I can identify
Enzymatic Assay of CALCINEURIN
 PRINCIPLE: The assay for Calcineurin is based on its ability to bind to Calmodulin. Calmodulin which is bound to Calcineurin is no longer available to activate Phosphodiesterase 3':5'-Cyclic Nucleotide.
PRINCIPLE: The assay for Calcineurin is based on its ability to bind to Calmodulin. Calmodulin which is bound to Calcineurin is no longer available to activate Phosphodiesterase 3':5'-Cyclic Nucleotide.
SUPPLEMENTARY INFORMATION
 Supplementary Table 1: Amplitudes of three current levels. Level 0 (pa) Level 1 (pa) Level 2 (pa) TrkA- TrkH WT 200 K 0.01 ± 0.01 9.5 ± 0.01 18.7 ± 0.03 200 Na * 0.001 ± 0.01 3.9 ± 0.01 12.5 ± 0.03 200
Supplementary Table 1: Amplitudes of three current levels. Level 0 (pa) Level 1 (pa) Level 2 (pa) TrkA- TrkH WT 200 K 0.01 ± 0.01 9.5 ± 0.01 18.7 ± 0.03 200 Na * 0.001 ± 0.01 3.9 ± 0.01 12.5 ± 0.03 200
Supporting Information
 Supporting Information Wiley-VCH 2012 69451 Weinheim, Germany Bound Cations Significantly Stabilize the Structure of Multiprotein Complexes in the Gas Phase** Linjie Han, Suk-Joon Hyung, and Brandon T.
Supporting Information Wiley-VCH 2012 69451 Weinheim, Germany Bound Cations Significantly Stabilize the Structure of Multiprotein Complexes in the Gas Phase** Linjie Han, Suk-Joon Hyung, and Brandon T.
Biochemistry. Biochemistry 9/20/ Bio-Energetics. 4.2) Transport of ions and small molecules across cell membranes
 9/20/15 Biochemistry Biochemistry 4. Bio-Energetics 4.2) Transport of ions and small molecules across cell membranes Aquaporin, the water channel, consists of four identical transmembrane polypeptides
9/20/15 Biochemistry Biochemistry 4. Bio-Energetics 4.2) Transport of ions and small molecules across cell membranes Aquaporin, the water channel, consists of four identical transmembrane polypeptides
Nuclear Medicine Department, Academic Medical Center, Amsterdam, The Netherlands;
 [ 3 H]-Spiperone Competition Binding to Dopamine D2, D3 and D4 Receptors Jan-Peter van Wieringen 1 and Martin C. Michel 2* 1 Nuclear Medicine Department, Academic Medical Center, Amsterdam, The Netherlands;
[ 3 H]-Spiperone Competition Binding to Dopamine D2, D3 and D4 Receptors Jan-Peter van Wieringen 1 and Martin C. Michel 2* 1 Nuclear Medicine Department, Academic Medical Center, Amsterdam, The Netherlands;
Chapter 4: Types of Chemical Reactions and Solution Stoichiometry
 Chapter 4: Types of Chemical Reactions and Solution Stoichiometry 4.1 Water, the Common Solvent 4.2 The Nature of Aqueous Solutions: Strong and Weak Electrolytes 4.3 The Composition of Solutions (MOLARITY!)
Chapter 4: Types of Chemical Reactions and Solution Stoichiometry 4.1 Water, the Common Solvent 4.2 The Nature of Aqueous Solutions: Strong and Weak Electrolytes 4.3 The Composition of Solutions (MOLARITY!)
Structure of (Na',K')-ATPase as Revealed by Electron Microscopy and Image Processing
 Published Online: 1 May, 1984 Supp Info: http://doi.org/10.1083/jcb.98.5.1836 Downloaded from jcb.rupress.org on November 3, 2018 Structure of (Na',K')-ATPase as Revealed by Electron Microscopy and Image
Published Online: 1 May, 1984 Supp Info: http://doi.org/10.1083/jcb.98.5.1836 Downloaded from jcb.rupress.org on November 3, 2018 Structure of (Na',K')-ATPase as Revealed by Electron Microscopy and Image
Enzymatic Assay of FK-BINDING PROTEIN Peptidyl-Prolyl Isomerase Activity
 Enzymatic Assay of FK-BINDING PROTEIN PRINCIPLE: Suc-Ala-Leu-cis-Pro-Phe-pNA FKBP > Suc-Ala-Leu-trans-Pro-PhepNA Suc-Ala-Leu-trans-Pro-Phe-pNA + H2O Chymotrypsin > Suc-Ala-Leu-trans-Pro-Phe + p-nitroaniline
Enzymatic Assay of FK-BINDING PROTEIN PRINCIPLE: Suc-Ala-Leu-cis-Pro-Phe-pNA FKBP > Suc-Ala-Leu-trans-Pro-PhepNA Suc-Ala-Leu-trans-Pro-Phe-pNA + H2O Chymotrypsin > Suc-Ala-Leu-trans-Pro-Phe + p-nitroaniline
Spherotech, Inc Irma Lee Circle, Unit 101, Lake Forest, Illinois PARTICLE COATING PROCEDURES
 SPHERO TM Technical Note STN-1 Rev C. 041106 Introduction Currently, there are several methods of attaching biological ligands to polystyrene particles. These methods include adsorption to plain polystyrene
SPHERO TM Technical Note STN-1 Rev C. 041106 Introduction Currently, there are several methods of attaching biological ligands to polystyrene particles. These methods include adsorption to plain polystyrene
Energy Transformation, Cellular Energy & Enzymes (Outline)
 Energy Transformation, Cellular Energy & Enzymes (Outline) Energy conversions and recycling of matter in the ecosystem. Forms of energy: potential and kinetic energy The two laws of thermodynamic and definitions
Energy Transformation, Cellular Energy & Enzymes (Outline) Energy conversions and recycling of matter in the ecosystem. Forms of energy: potential and kinetic energy The two laws of thermodynamic and definitions
Solutions. Solution: A solution is homogeneous liquid mixture of two or more substances.
 Solutions Objectives: 1. Learn the various methods of expressing concentrations of solutions. 2. Learn to make percent and molar solutions from solids, liquids, and stock solutions. 3. Learn the various
Solutions Objectives: 1. Learn the various methods of expressing concentrations of solutions. 2. Learn to make percent and molar solutions from solids, liquids, and stock solutions. 3. Learn the various
The Na,K-ATPase is an essential ion transporter in almost
 This is an open access article published under an ACS AuthorChoice License, which permits copying and redistribution of the article or any adaptations for non-commercial purposes. pubs.acs.org/biochemistry
This is an open access article published under an ACS AuthorChoice License, which permits copying and redistribution of the article or any adaptations for non-commercial purposes. pubs.acs.org/biochemistry
Chapter 6- An Introduction to Metabolism*
 Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Center for Cell Imaging Department of Cell Biology
 Center for Cell Imaging Department of Cell Biology Contents Preparation of Colloidal Gold Conjugates Coupling the Protein A to the Gold Particles Purification of the protein A-gold. Storage Influence of
Center for Cell Imaging Department of Cell Biology Contents Preparation of Colloidal Gold Conjugates Coupling the Protein A to the Gold Particles Purification of the protein A-gold. Storage Influence of
Effect of chaotropic anions on the sodium transport by the Na,K-ATPase
 Eur Biophys J (2006) 35: 247 254 DOI 10.1007/s00249-005-0031-9 ARTICLE Artem G. Ayuyan Æ Valerij S. Sokolov Alexander A. Lenz Æ Hans-Ju rgen Apell Effect of chaotropic anions on the sodium transport by
Eur Biophys J (2006) 35: 247 254 DOI 10.1007/s00249-005-0031-9 ARTICLE Artem G. Ayuyan Æ Valerij S. Sokolov Alexander A. Lenz Æ Hans-Ju rgen Apell Effect of chaotropic anions on the sodium transport by
Biochemistry. Biochemistry 7/11/ Bio-Energetics. 4.2) Transport of ions and small molecules across cell membranes
 Biochemistry Biochemistry 4. Bio-Energetics 4.2) Transport of ions and small molecules across cell membranes Aquaporin, the water channel, consists of four identical transmembrane polypeptides Key Energy
Biochemistry Biochemistry 4. Bio-Energetics 4.2) Transport of ions and small molecules across cell membranes Aquaporin, the water channel, consists of four identical transmembrane polypeptides Key Energy
Highly Sensitive and Selective Colorimetric Visualization of Streptomycin in Raw Milk Using Au Nanoparticles Supramolecular Assembly
 SUPPORTING INFORMATION Highly Sensitive and Selective Colorimetric Visualization of Streptomycin in Raw Milk Using Au Nanoparticles Supramolecular Assembly Jiayu Sun, Jiechao Ge, Weimin Liu, Zhiyuan Fan,
SUPPORTING INFORMATION Highly Sensitive and Selective Colorimetric Visualization of Streptomycin in Raw Milk Using Au Nanoparticles Supramolecular Assembly Jiayu Sun, Jiechao Ge, Weimin Liu, Zhiyuan Fan,
Basic Chemistry. Chapter 2 BIOL1000 Dr. Mohamad H. Termos
 Basic Chemistry Chapter 2 BIOL1000 Dr. Mohamad H. Termos Chapter 2 Objectives Following this chapter, you should be able to describe: - Atoms, molecules, and ions - Composition and properties - Types of
Basic Chemistry Chapter 2 BIOL1000 Dr. Mohamad H. Termos Chapter 2 Objectives Following this chapter, you should be able to describe: - Atoms, molecules, and ions - Composition and properties - Types of
respiration."' 12 Contraction is closely linked to electron flow. It cannot X), yielding a burst of respiration, H+ release and Ca++ transport,
 EVIDENCE FOR ACTIVE PHOSPHATE TRANSPORT IN MIAIZE AHITOCHONDRIA* BY J. B. HANSON AND RAYMOND J. MILLER DEPARTMENT OF AGRONOMY, UNIVERSITY OF ILLINOIS, URBANA Communicated by Sterling B. Hendrick, May 26,
EVIDENCE FOR ACTIVE PHOSPHATE TRANSPORT IN MIAIZE AHITOCHONDRIA* BY J. B. HANSON AND RAYMOND J. MILLER DEPARTMENT OF AGRONOMY, UNIVERSITY OF ILLINOIS, URBANA Communicated by Sterling B. Hendrick, May 26,
Advanced Higher Biology. Unit 1- Cells and Proteins 2c) Membrane Proteins
 Advanced Higher Biology Unit 1- Cells and Proteins 2c) Membrane Proteins Membrane Structure Phospholipid bilayer Transmembrane protein Integral protein Movement of Molecules Across Membranes Phospholipid
Advanced Higher Biology Unit 1- Cells and Proteins 2c) Membrane Proteins Membrane Structure Phospholipid bilayer Transmembrane protein Integral protein Movement of Molecules Across Membranes Phospholipid
LAB. FACTORS INFLUENCING ENZYME ACTIVITY
 AP Biology Date LAB. FACTORS INFLUENCING ENZYME ACTIVITY Background Enzymes are biological catalysts capable of speeding up chemical reactions by lowering activation energy. One benefit of enzyme catalysts
AP Biology Date LAB. FACTORS INFLUENCING ENZYME ACTIVITY Background Enzymes are biological catalysts capable of speeding up chemical reactions by lowering activation energy. One benefit of enzyme catalysts
The photoluminescent graphene oxide serves as an acceptor rather. than a donor in the fluorescence resonance energy transfer pair of
 Supplementary Material (ESI) for Chemical Communications This journal is (c) The Royal Society of Chemistry 20XX The photoluminescent graphene oxide serves as an acceptor rather than a donor in the fluorescence
Supplementary Material (ESI) for Chemical Communications This journal is (c) The Royal Society of Chemistry 20XX The photoluminescent graphene oxide serves as an acceptor rather than a donor in the fluorescence
Membrane Protein Channels
 Membrane Protein Channels Potassium ions queuing up in the potassium channel Pumps: 1000 s -1 Channels: 1000000 s -1 Pumps & Channels The lipid bilayer of biological membranes is intrinsically impermeable
Membrane Protein Channels Potassium ions queuing up in the potassium channel Pumps: 1000 s -1 Channels: 1000000 s -1 Pumps & Channels The lipid bilayer of biological membranes is intrinsically impermeable
WHAT REGULATES RESPIRATION IN MITOCHONDRIA?
 Vol. 39, No. 2, May 1996 BIOCHEMISTRY and MOLECULAR BIOLOGY INTERNATIONAL Pages 415-4 ] 9 WHAT REGULATES RESPIRATION IN MITOCHONDRIA? Bernard Korzeniewski Institute of Molecular Biology, Jagiellonian University,
Vol. 39, No. 2, May 1996 BIOCHEMISTRY and MOLECULAR BIOLOGY INTERNATIONAL Pages 415-4 ] 9 WHAT REGULATES RESPIRATION IN MITOCHONDRIA? Bernard Korzeniewski Institute of Molecular Biology, Jagiellonian University,
April Barry Isralewitz
 1 April 2002 Barry Isralewitz All enzymes are beautiful, but ATP synthase is one of the most: - Beautiful because of its 3D structure, - Unusual because of its structural complexity and reaction mechanism,
1 April 2002 Barry Isralewitz All enzymes are beautiful, but ATP synthase is one of the most: - Beautiful because of its 3D structure, - Unusual because of its structural complexity and reaction mechanism,
JOURNAL RANKING 2014 FOR BIOPHYSICS
 The data source used is Journal Citation Reports (JCR) published by Thomson Reuters. Data generated from JCR Science Edition 2014. Biophysics covers resources that focus on the transfer and effects of
The data source used is Journal Citation Reports (JCR) published by Thomson Reuters. Data generated from JCR Science Edition 2014. Biophysics covers resources that focus on the transfer and effects of
Isolation & Purification of Proteoglycans (PGs) and Glycosaminoglycans (GAGs) PEG Trainee Lecture July 23, 2012
 Isolation & Purification of Proteoglycans (PGs) and Glycosaminoglycans (GAGs) PEG Trainee Lecture July 23, 2012 Most Common Extraction Procedure for PGs 4 M Guanidine-HCl Detergents such as 2% CHAPS or
Isolation & Purification of Proteoglycans (PGs) and Glycosaminoglycans (GAGs) PEG Trainee Lecture July 23, 2012 Most Common Extraction Procedure for PGs 4 M Guanidine-HCl Detergents such as 2% CHAPS or
Energy and Cells. Appendix 1. The two primary energy transformations in plants are photosynthesis and respiration.
 Energy and Cells Appendix 1 Energy transformations play a key role in all physical and chemical processes that occur in plants. Energy by itself is insufficient to drive plant growth and development. Enzymes
Energy and Cells Appendix 1 Energy transformations play a key role in all physical and chemical processes that occur in plants. Energy by itself is insufficient to drive plant growth and development. Enzymes
Energy Transformation and Metabolism (Outline)
 Energy Transformation and Metabolism (Outline) - Definitions & Laws of Thermodynamics - Overview of energy flow ecosystem - Biochemical processes: Anabolic/endergonic & Catabolic/exergonic - Chemical reactions
Energy Transformation and Metabolism (Outline) - Definitions & Laws of Thermodynamics - Overview of energy flow ecosystem - Biochemical processes: Anabolic/endergonic & Catabolic/exergonic - Chemical reactions
Online Supplementary Material. Messenger RNA Interactions in the Decoding Center Control the Rate of Translocation
 Online Supplementary Material Messenger RNA Interactions in the Decoding Center Control the Rate of Translocation Prashant K. Khade and Simpson Joseph Supplementary Figure 1 Dissociation of the f[ 35 S]Met-Phe-tRNA
Online Supplementary Material Messenger RNA Interactions in the Decoding Center Control the Rate of Translocation Prashant K. Khade and Simpson Joseph Supplementary Figure 1 Dissociation of the f[ 35 S]Met-Phe-tRNA
Application Note: A TD-700 Laboratory Fluorometer Method for Alkaline Phosphatase Fluorescence
 1. INTRODUCTION Because of their critical functions in eukaryotic cells, methods for measuring protein phosphatases were established at least as early as 1953 1. In 1965 Fernley and Walker 2 decribed the
1. INTRODUCTION Because of their critical functions in eukaryotic cells, methods for measuring protein phosphatases were established at least as early as 1953 1. In 1965 Fernley and Walker 2 decribed the
Essays. Essay Number 2. Equations of State for Fractional Saturation of Human Hemoglobin with Oxygen
 Essays Essay Number 2 Equations of State for Fractional Saturation of Human Hemoglobin with Oxygen Formulation of the Adair Equation with Equivalent O 2 Binding Sites Francis Knowles Equations of State
Essays Essay Number 2 Equations of State for Fractional Saturation of Human Hemoglobin with Oxygen Formulation of the Adair Equation with Equivalent O 2 Binding Sites Francis Knowles Equations of State
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing
 Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Inhibition of the Na,K-ATPase by Fluoride
 THE JOURNAL OF BIOLOGICAL CHEMISTRY (c) 1992 by The American Society for Biochemistry and Molecular Biology, Inc. Vol. 267, No. 24, Issue of August 25, pp. 16995-17000,1932 Printed in U. S. A. Inhibition
THE JOURNAL OF BIOLOGICAL CHEMISTRY (c) 1992 by The American Society for Biochemistry and Molecular Biology, Inc. Vol. 267, No. 24, Issue of August 25, pp. 16995-17000,1932 Printed in U. S. A. Inhibition
Lecture 12: Burst Substrates and the V vs [S] Experiment
![Lecture 12: Burst Substrates and the V vs [S] Experiment Lecture 12: Burst Substrates and the V vs [S] Experiment](/thumbs/95/124907449.jpg) Biological Chemistry Laboratory Biology 3515/Chemistry 3515 Spring 2019 Lecture 12: Burst Substrates and the V vs [S] Experiment 14 February 2019 c David P. Goldenberg University of Utah goldenberg@biology.utah.edu
Biological Chemistry Laboratory Biology 3515/Chemistry 3515 Spring 2019 Lecture 12: Burst Substrates and the V vs [S] Experiment 14 February 2019 c David P. Goldenberg University of Utah goldenberg@biology.utah.edu
Conformational change of L7/L12 stalk in the different functional states of 50S ribosomes
 J. Biosci., Vol. 11, Numbers 1 4, March 1987, pp. 561-569. Printed in India. Conformational change of L7/L12 stalk in the different functional states of 50S ribosomes DEBABRATA DASH, SUBODH MAHANTI and
J. Biosci., Vol. 11, Numbers 1 4, March 1987, pp. 561-569. Printed in India. Conformational change of L7/L12 stalk in the different functional states of 50S ribosomes DEBABRATA DASH, SUBODH MAHANTI and
Figure ) Letter E represents a nucleic acid building block known as a. Answer: nucleotide Diff: 3 Page Ref: 54
 Essentials of Human Anatomy and Physiology, 10e (Marieb) Chapter 2 Basic Chemistry 2.1 Short Answer Figure 2.1 Using Figure 2.1, identify the following: 1) Which letter represents a carbohydrate polymer?
Essentials of Human Anatomy and Physiology, 10e (Marieb) Chapter 2 Basic Chemistry 2.1 Short Answer Figure 2.1 Using Figure 2.1, identify the following: 1) Which letter represents a carbohydrate polymer?
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10458 Active Site Remodeling in the Bifunctional Fructose-1,6- bisphosphate aldolase/phosphatase Juan Du, Rafael F. Say, Wei Lü, Georg Fuchs & Oliver Einsle SUPPLEMENTARY FIGURES Figure
doi:10.1038/nature10458 Active Site Remodeling in the Bifunctional Fructose-1,6- bisphosphate aldolase/phosphatase Juan Du, Rafael F. Say, Wei Lü, Georg Fuchs & Oliver Einsle SUPPLEMENTARY FIGURES Figure
Hole s Human Anatomy and Physiology Tenth Edition. Chapter 2
 PowerPoint Lecture Outlines to accompany Hole s Human Anatomy and Physiology Tenth Edition Shier w Butler w Lewis Chapter 2 Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction
PowerPoint Lecture Outlines to accompany Hole s Human Anatomy and Physiology Tenth Edition Shier w Butler w Lewis Chapter 2 Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction
Title. CitationChemical Communications, 48(53): Issue Date Doc URL. Type. Additional There Information.
 Title A heme degradation enzyme, HutZ, from Vibrio cholera Author(s)Uchida, Takeshi; Sekine, Yukari; Matsui, Toshitaka; CitationChemical Communications, 48(53): 6741-6743 Issue Date 2012-06 Doc URL http://hdl.handle.net/2115/68577
Title A heme degradation enzyme, HutZ, from Vibrio cholera Author(s)Uchida, Takeshi; Sekine, Yukari; Matsui, Toshitaka; CitationChemical Communications, 48(53): 6741-6743 Issue Date 2012-06 Doc URL http://hdl.handle.net/2115/68577
Presentation Microcalorimetry for Life Science Research
 Presentation Microcalorimetry for Life Science Research MicroCalorimetry The Universal Detector Heat is either generated or absorbed in every chemical process Capable of thermal measurements over a wide
Presentation Microcalorimetry for Life Science Research MicroCalorimetry The Universal Detector Heat is either generated or absorbed in every chemical process Capable of thermal measurements over a wide
Chapter 8: An Introduction to Metabolism. 1. Energy & Chemical Reactions 2. ATP 3. Enzymes & Metabolic Pathways
 Chapter 8: An Introduction to Metabolism 1. Energy & Chemical Reactions 2. ATP 3. Enzymes & Metabolic Pathways 1. Energy & Chemical Reactions 2 Basic Forms of Energy Kinetic Energy (KE) energy in motion
Chapter 8: An Introduction to Metabolism 1. Energy & Chemical Reactions 2. ATP 3. Enzymes & Metabolic Pathways 1. Energy & Chemical Reactions 2 Basic Forms of Energy Kinetic Energy (KE) energy in motion
Involvement of Protons in the Ion Transport Cycle of Na+,K+ -ATPase l
 Involvement of Protons in the Ion Transport Cycle of Na+,K+ -ATPase l K. O. Grishanin, V. Yu. Tashkino, A. A. Lenz b, H. J. ApeUC, and V. S. SokoloV" a Frumkin Institute of Physical Chemisliy and Electrochemis/ly,
Involvement of Protons in the Ion Transport Cycle of Na+,K+ -ATPase l K. O. Grishanin, V. Yu. Tashkino, A. A. Lenz b, H. J. ApeUC, and V. S. SokoloV" a Frumkin Institute of Physical Chemisliy and Electrochemis/ly,
BMB Lecture 7. Allostery and Cooperativity
 BMB 178 2017 Lecture 7 October 18, 2017 Allostery and Cooperativity A means for exquisite control Allostery: the basis of enzymatic control From the Greek: allos = other stereos = solid or space Action
BMB 178 2017 Lecture 7 October 18, 2017 Allostery and Cooperativity A means for exquisite control Allostery: the basis of enzymatic control From the Greek: allos = other stereos = solid or space Action
Essentials of Human Anatomy and Physiology, 12e (Marieb) Chapter 2 Basic Chemistry. 2.1 Multiple Choice Part I Questions
 Essentials of Human Anatomy and Physiology 12th Edition Marieb TEST BANK Full download at: https://testbankrealcom/download/essentialshuman-anatomy-physiology-12th-edition-mariebtest-bank/ Essentials of
Essentials of Human Anatomy and Physiology 12th Edition Marieb TEST BANK Full download at: https://testbankrealcom/download/essentialshuman-anatomy-physiology-12th-edition-mariebtest-bank/ Essentials of
Pathway of ATP Hydrolysis by Monomeric Kinesin Eg5
 12334 Biochemistry 2006, 45, 12334-12344 Pathway of ATP Hydrolysis by Monomeric Kinesin Eg5 Jared C. Cochran, Troy C. Krzysiak, and Susan P. Gilbert* Department of Biological Sciences, UniVersity of Pittsburgh,
12334 Biochemistry 2006, 45, 12334-12344 Pathway of ATP Hydrolysis by Monomeric Kinesin Eg5 Jared C. Cochran, Troy C. Krzysiak, and Susan P. Gilbert* Department of Biological Sciences, UniVersity of Pittsburgh,
Malachite Green Phosphate Detection Kit Catalog Number: DY996
 Malachite Green Phosphate Detection Kit Catalog Number: DY996 This Malachite Green Phosphate Detection Kit employs a simple, sensitive, reproducible, and non-radioactive method for measuring inorganic
Malachite Green Phosphate Detection Kit Catalog Number: DY996 This Malachite Green Phosphate Detection Kit employs a simple, sensitive, reproducible, and non-radioactive method for measuring inorganic
Chemistry 5.07SC Biological Chemistry I Fall Semester, 2013
 Chemistry 5.07SC Biological Chemistry I Fall Semester, 2013 Lecture 10. Biochemical Transformations II. Phosphoryl transfer and the kinetics and thermodynamics of energy currency in the cell: ATP and GTP.
Chemistry 5.07SC Biological Chemistry I Fall Semester, 2013 Lecture 10. Biochemical Transformations II. Phosphoryl transfer and the kinetics and thermodynamics of energy currency in the cell: ATP and GTP.
Membrane Physiology. Dr. Hiwa Shafiq Oct-18 1
 Membrane Physiology Dr. Hiwa Shafiq 22-10-2018 29-Oct-18 1 Chemical compositions of extracellular and intracellular fluids. 29-Oct-18 2 Transport through the cell membrane occurs by one of two basic processes:
Membrane Physiology Dr. Hiwa Shafiq 22-10-2018 29-Oct-18 1 Chemical compositions of extracellular and intracellular fluids. 29-Oct-18 2 Transport through the cell membrane occurs by one of two basic processes:
Fructose 1,6-Diphosphate Aldolase from Rabbit Muscle
 Btochem. J. (1975) 151, 167-172 Printed in Great Britain 167 Fructose 1,6-Diphosphate Aldolase from Rabbit Muscle EFFECT OF ph ON THE RATE OF FORMATION AND ON THE EQUILIBRIUM CONCENTRATION OF THE CARBANION
Btochem. J. (1975) 151, 167-172 Printed in Great Britain 167 Fructose 1,6-Diphosphate Aldolase from Rabbit Muscle EFFECT OF ph ON THE RATE OF FORMATION AND ON THE EQUILIBRIUM CONCENTRATION OF THE CARBANION
Chapter 6. The interaction of Src SH2 with the focal adhesion kinase catalytic domain studied by NMR
 The interaction of Src SH2 with the focal adhesion kinase catalytic domain studied by NMR 103 Abstract The interaction of the Src SH2 domain with the catalytic domain of FAK, including the Y397 SH2 domain
The interaction of Src SH2 with the focal adhesion kinase catalytic domain studied by NMR 103 Abstract The interaction of the Src SH2 domain with the catalytic domain of FAK, including the Y397 SH2 domain
the erythrocyte membrane (fluorescent probes)
 Proc. Natl. Acad. Sci. USA Vol. 77, No. 12, pp. 7147-7151, December 1980 Biochemistry Kinetic study of the interaction of oxy- and deoxyhemoglobins with the erythrocyte membrane (fluorescent probes) N.
Proc. Natl. Acad. Sci. USA Vol. 77, No. 12, pp. 7147-7151, December 1980 Biochemistry Kinetic study of the interaction of oxy- and deoxyhemoglobins with the erythrocyte membrane (fluorescent probes) N.
Solutions. Experiment 11. Various Types of Solutions. Solution: A homogenous mixture consisting of ions or molecules
 Solutions Solution: A homogenous mixture consisting of ions or molecules -Assignment: Ch 15 Questions & Problems : 5, (15b,d), (17a, c), 19, 21, 23, 27, (33b,c), 39, (43c,d),45b, 47, (49b,d), (55a,b),
Solutions Solution: A homogenous mixture consisting of ions or molecules -Assignment: Ch 15 Questions & Problems : 5, (15b,d), (17a, c), 19, 21, 23, 27, (33b,c), 39, (43c,d),45b, 47, (49b,d), (55a,b),
Questions Maximum Score Candidate s Score
 Name:...Index Number Date... Candidate s Signature:... 233/1 CHEMISTRY Theory PAPER 1 July 2013 2 Hours STAREHE GIRLS CENTRE FORM 4 MOCK EXAMS, JULY 2013 CHEMISTRY PAPER 1 2 Hours Instructions to candidates:
Name:...Index Number Date... Candidate s Signature:... 233/1 CHEMISTRY Theory PAPER 1 July 2013 2 Hours STAREHE GIRLS CENTRE FORM 4 MOCK EXAMS, JULY 2013 CHEMISTRY PAPER 1 2 Hours Instructions to candidates:
Try this one Calculate the ph of a solution containing M nitrous acid (Ka = 4.5 E -4) and 0.10 M potassium nitrite.
 Chapter 17 Applying equilibrium 17.1 The Common Ion Effect When the salt with the anion of a is added to that acid, it reverses the dissociation of the acid. Lowers the of the acid. The same principle
Chapter 17 Applying equilibrium 17.1 The Common Ion Effect When the salt with the anion of a is added to that acid, it reverses the dissociation of the acid. Lowers the of the acid. The same principle
