A Machine Learning Approach for Prediction of Gibberellic Acid Metabolic Enzymes in Monocotyledonous Plants
|
|
|
- Roberta Ward
- 5 years ago
- Views:
Transcription
1 A Machine Learning Approach for Prediction of Gibberellic Acid Metabolic Enzymes in Monocotyledonous Plants Sreepriya P. 1, Naganeeswaran S. 1, Hemalatha N. 2, Sreejisha P. 1 and Rajesh M. K. 1,* 1 Bioinformatics Centre, Central Plantation Crops Research Institute, Kasaragod, Kerala, India 2 AIMIT, St. Aloysius College, Mangalore, Karnataka, India sreepriya.pradeep@gmail.com, naganeeswaran@gmail.com, hemasree71@gmail.com, sreeji279@gmail.com, mkraju.cpcri@gmail.com [*Corresponding author: Phone: ; Fax: ] ABSTRACT Gibberellins (GA) are one of the most important phytohormones that control different aspects of plant growth and influence various developments such as seed germination, stem elongation and floral induction. More than 130 GAs have been identified; however, only a small number of them are biologically active. In this study, five enzymes in GA metabolic pathway in monocots have been thoroughly researched namely, ent-copalyl-diphosphate synthase (CPS), ent-kaurene synthase (KS), ent-kaurene oxidase (KO), GA 20-oxidase (GA20ox), and GA 2-oxidase (GA2ox). We have designed and implemented a high performance prediction tool for these enzymes using machine learning algorithms. GAPred is a web-based system to provide a comprehensive collection of enzymes in GA metabolic pathway and a systematic framework for the analysis of these enzymes for monocots. WEKA-based classifiers (Naïve-Bayes) and Support Vector Machine (SVM) based-modules were developed using dipeptide composition and high accuracies were obtained. In addition, BLAST and Hidden Markov Model (HMMER-based model) were also developed for searching sequence databases for homolog s of enzymes of GA metabolic pathway, and for making protein sequence alignments. Keywords: GA, SVM, WEKA, BLAST, HMMER 1 INTRODUCTION Gibberellic acids (GA) are naturally occurring phytohormones that regulate growth and influence various developmental processes, including stem elongation, germination, dormancy, flowering, enzyme induction, and leaf and fruit senescence [1]. They are also involved in the discernment of environmental stimuli, thus are significant not only for a plant s growth and development but also in awareness of its environment. Gibberellins are diterpenoid acids which DOI: /tmlai Publication Date: 28 th August 2014 URL:
2 Transactions on Machine Learning and Artificial Intelligence Volume 2, Issue 4, August 2104 are formed by the terpenoid pathway in plastids and then modifying the endoplasmic reticulum (ER) and cytosol until they are biologically-active [2]. Gibberellins are derived by the entgibberellane skeleton, but are synthesized by ent-kaurene [3]. The GA biosynthetic pathway can be divided into three stages, each stage residing in a different cellular compartment viz. plastid, the endoplasmic reticulum, and the cytosol [4]. A number of experimental studies have explained thoroughly the biosynthetic functions of gibberellic acid. In this study, five enzymes involved in GA metabolic pathway in monocots have been thoroughly researched namely, ent-copalyl-diphosphate synthase (CPS), ent-kaurene synthase (KS), ent-kaurene oxidase (KO), GA 20-oxidase (GA20ox), and GA 2-oxidase (GA2ox) [5]. In this study, we have designed and implemented a high performance prediction tool based on kernel-based Machine Learning Algorithms viz., Support Vector Machine (SVM) and WEKA for prediction of enzymes in gibberellic acid metabolic pathway. In addition, standalone BLAST and Hidden Markov Model (HMMER-based model) were also developed for searching sequence databases for homolog s of enzymes of GA metabolic pathway, and for making protein sequence alignments. GAPred was developed using the evolutionary and sequence features of a protein sequence and the performance of the each model was evaluated using crossvalidation techniques. Based on our study, we have also created and hosted a web server for predicting enzymes involved in GA metabolism. 2.1 Dataset 2 MATERIALS AND METHODS In the present study, two datasets were considered for the development of the prediction tool GAPred. Positive (+ve) dataset comprised of 102 selected GA metabolic enzyme protein sequences from monocots viz., date palm (Phoenix dactylifera), coconut (Cocos nucifera), rice (Oryza sativa), barley (Hordeum vulgare), maize (Zea mays), banana (Musa acuminata), and brachypodium (Brachypodium distachyon), after redundancy elimination by using ClustalW. Similarly negative (-ve) dataset was created by using same numbers of non-ga metabolic enzymes sequences. The sequences were retrieved from NCBI in FASTA format ( Domains of enzymes involved in GA metabolic pathway were identified using Pfam search and PRINTS search and most of the identified enzyme domains are known to be conserved in related species. To avoid the over estimation, we clustered the protein sequences from positive data (+ve) set with a threshold of 30% identity by CD-HIT (Cluster Database at High Identity with Tolerance). Out of 102 GA metabolic enzymes sequence, 62 proteins were randomly selected for the creation of training set. Similarly training set of non-ga metabolic enzymes sequence was created. To test the reliability of the prediction tool, we also prepared a test set of 40 GA metabolic enzymes sequences and non-ga metabolic enzymes sequences which were not the part of training set. Copyright Society for Science and Education United Kingdom 37
3 Sreepriya P., Naganeeswaran S., Hemalatha N., Sreejisha P. and Rajesh M. K. ; A Machine Learning Approach for Prediction of Gibberellic Acid Metabolic Enzymes in Monocotyledonous Plants; Transactions on Machine Learning and Artificial Intelligence, Volume 2 No 4, Aug (2014); pp: Support Vector Machine Support Vector Machines (SVM) are a group of rapid optimization machine learning algorithms with strong theoretical foundation, which have been used for many kinds of pattern recognition [6-8]. SVMs are now extensively used for biological applications and methods such as classifying objects as diverse as protein and DNA sequences, mass spectra and microarray expression profiles [9]. In this work, SVM has been implemented by using SVM multiclass package [10] which possesses two modules: SVM_multiclass_learn and SVM_multiclass_classify. The first module (SVM_multiclass_learn) is concerned for preparing models learned from the training dataset (+ve and -ve) and the final one classifies the data by using the models prepared by SVM_multiclass_learn. Here, we have trained the SVM multiclass by using a set of positive and negative datasets, and produces a model (classifier) that can be used to identify the potential enzymes involved in gibberellic acid metabolic pathway. With the help of this package the user can select various kernel functions (linear, polynomial, radial basis, sigmoid or any other user defined kernel) for preparing models. In SVMs, the kernel function selected must be the most favorable one. Here in the creation of SVM models, we have used three types of kernel functions: linear, polynomial, and radial. The performance of SVM based methods has been optimized by regulating SVM parameters so that maximum accuracy could be obtained. 2.3 WEKA WEKA stands for Waikato Environment for Knowledge Analysis and is a free open source software developed by at the University of Waikato, New Zealand. This popular machine learning software contains a collection of algorithms and visualization tools for data analysis, analytical modeling and also graphical user interfaces for easy access to this functionality. In the given work, we used WEKA classification [11], where different attributes of a protein sequence are analyzed to classify the protein sequence into one of the predefined classes. Both train and test set was used to get the classification of the data set by using better algorithms. The performance of WEKA has been optimized by tuning evaluation parameters and visualization schemes, in order to analyze the accuracy of classifiers. 2.4 Sequence Features Dipeptide composition gives comprehensive information about each protein sequence that possess sequence feature. Generally, the total number of amino acids is 20 and thus the theoretical number of possible dipeptides is 400. A matrix of these 400 dipeptides was generated for each protein and is then given as an input to both SVM and WEKA. Each dipeptide frequency is calculated by the formula: DF ij =N ij /N where, N ij =count of the ijth dipeptide; N=total number of possible dipeptides; i, j = 1-20 amino acid. URL: 38
4 Transactions on Machine Learning and Artificial Intelligence Volume 2, Issue 4, August Performance Assessment of GAPred With the help of statistical calculations, we generally examine the efficiency of a predictor either using single independent dataset test, cross-validation test or jackknife test. However, jackknife test method takes much longer time to examine a predictor based on SVM and WEKA [12] and therefore, in this work, we have adopted 10-fold cross-validation for WEKA and 5-fold cross-validation for SVM multiclass and independent data set validation techniques were adopted for measuring performance. In 10-fold cross-validation test, the significant dataset was divided randomly into ten equally sized sets. The training and testing methods were carried out ten times with each individual set used for testing and for the nine sets left behind for training. Similarly in 5-fold cross validation, the dataset was partitioned randomly into five equally sized sets. In the independent dataset test, the training dataset used to train the predictor does not contain any data that is to be tested. 2.6 Evaluation Parameters We had made use of five parameters to evaluate the reliability of the prediction tool, they are: Accuracy (Ac), Sensitivity (Sn), Specificity (Sp), Precision (Pr) and Matthew s Correlation Coefficient (MCC). Accuracy defines the proportion of correctly predicted proteins (Eq.1). The sensitivity (Sn) and specificity (Sp) represent the correct prediction ratios of positive (+ve) and negative data (-ve) sets of metabolic enzymes of gibberellic acid sequence respectively (Eq. 2 and 3). Precision is the proportion of the predicted positive cases that were correct (Eq.4). Matthew s correlation coefficient or MCC [13-14] is a statistical parameter which also used to estimate the accuracy of prediction (Eq.5). MCC may range from -1 to +1 and the highest MCC value indicates better prediction [15]. Accuracy = Sensitivity = TP Specificity = Precision = MCC = TP+FN TN TP+TN TP+TN+FP+FN 100 (1) 100 (2) FP+TN TP 100 (3) 100 (4) TP+FP (TP TN) (FP FN) (TP+FP)(TP+FN)(TN+FP)(TN+FN) (5) Where TP=number of true positives; TN=number of true negatives; FP=number of false positives; FN=number of false negatives. In this work, metabolic enzymes of GA sequences are true positives and non- metabolic enzymes of GA sequences are true negatives. Copyright Society for Science and Education United Kingdom 39
5 Sreepriya P., Naganeeswaran S., Hemalatha N., Sreejisha P. and Rajesh M. K. ; A Machine Learning Approach for Prediction of Gibberellic Acid Metabolic Enzymes in Monocotyledonous Plants; Transactions on Machine Learning and Artificial Intelligence, Volume 2 No 4, Aug (2014); pp: Sequence Similarity search using standalone BLAST The standalone BLAST programs are freely provided as open-source software by NCBI. With stand-alone BLAST we can made our own databases to search against. In this study, sequences were searched against the protein non-redundant (nr) database in association with standalone BLASTp and to detect homology of metabolic enzymes of gibberellic acid proteins and result was analysed. 2.7 Sequence Similarity Search using HMMER Profile Hidden Markov models (profile HMMs) techniques are one of the most dominant methods for protein homology detection [16]. HMMER helps to find out protein sequences which are similar in sequence databases and to make protein sequence alignments. HMMER becomes particularly powerful when the query is a multiple sequence alignment of a sequence family rather than for single query sequences. It makes a profile of the query that assigns a position-specific scoring system for substitutions, insertions, and deletions [17]. HMMER profiles are probabilistic models called profile Hidden Markov models (profile HMMs). Because of the strength of its underlying probability models, HMMER aims to be much more accurate and more capable of finding out remote homolog s rather than BLAST, FASTA or any of the other sequence alignment and database search tools based on older scoring methodology [18]. Hence, in this study, we have used HMMER to detect homology of metabolic enzymes of gibberellic acid proteins and a remarkable result was analyzed. 2.8 ROC Curves By making use of ROC curves, a graph created by plotting the fraction of false positives (FPR) against true positives (TPR) at various threshold settings [19], we can explain the performance of multi class classifiers in SVM and WEKA more specifically. TPR is also known as sensitivity, and FPR is 1-specificity or true negative rate. ROC analysis is linked in a direct and natural method to benefit analysis of diagnostic decision making. ROC curves useful for the evaluation of machine learning techniques and data mining research. 2.9 Web-server We have implemented the prediction tool GAPred in a web server. The program is written entirely in HTML, PHP and PERL program in a Linux platform. The tool page serve as the platform for submitting data where users can either paste or upload sequence which should be in standard FASTA format. It also provides a comprehensive collection of enzymes in GA metabolic pathway and introduces a user to gibberellic acid metabolic pathway. URL: 40
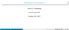
6 Transactions on Machine Learning and Artificial Intelligence Volume 2, Issue 4, August GAPRED 3.1 Evaluation of Performance of GAPred We have carried out 10-fold cross-validation for WEKA and 5-fold cross-validation for SVM and also independent data test validation to evaluate the performance of GAPred (Tables 1-4). Cross validation and independent data test results for SVM from the Tables 1 and 3 shows that cross validation has better result for dipeptide composition methods compared to independent data test. While in the case of WEKA, the independent data set has better result than cross validation method. 3.2 Comparison of GAPred with BLAST and HMMER We have also used standalone BLAST and HMMER to detect homology of metabolic enzymes of gibberellic acid. This was used to compare an input protein sequence with a created database to generate the homology of the given sequence. A comparison of enzyme proteins was conducted with standalone BLAST and HMMER database and an accuracy of 99% and 93% were obtained (Tables 5-6). By making a comparison between SVM, WEKA, BLAST and HMMER from Table 7 and Figure 1, it can be seen that SVM has achieved 100% accuracy and MCC value equal to 1 that an ideal classification method should possess. Hence, SVM was selected to be the best model for GAPred. Table 1 Validation of independent data test results of dipeptide composition of metabolic enzymes of gibberellic acid with SVM multiclass Algorithm Sn Sp Ac Pr MCC Linear Polynomial RBF Table 2 Validation of independent data test results of dipeptide composition of metabolic enzymes of gibberellic acid with WEKA Classifiers Sn Sp Ac Pr MCC Naïve Bayes Bayes Net Decorate Copyright Society for Science and Education United Kingdom 41
7 Sreepriya P., Naganeeswaran S., Hemalatha N., Sreejisha P. and Rajesh M. K. ; A Machine Learning Approach for Prediction of Gibberellic Acid Metabolic Enzymes in Monocotyledonous Plants; Transactions on Machine Learning and Artificial Intelligence, Volume 2 No 4, Aug (2014); pp: Table 3 Comparison of the prediction performance of three kernels of SVMmulticlass with dipeptide composition technique using 5-fold cross validation Algorithm Sn Sp Ac Pr MCC Linear Polynomial RBF Table 4 Comparison of the prediction performance of three classifiers of WEKA with dipeptide composition technique using 10-fold cross validation Classifiers Sn Sp Ac Pr MCC Naïve Bayes Bayes Net Decorate Table 5 Comparison of the prediction performance of standalone BLAST with created database of domains of metabolic enzymes of gibberellic acid Sn Sp Ac Pr MCC BLAST Table 6 Comparison of the prediction performance of HMMER with created database of domains of metabolic enzymes of gibberellic acid Sn Sp Ac Pr MCC HMMER URL: 42
8 Transactions on Machine Learning and Artificial Intelligence Volume 2, Issue 4, August 2104 Table 7 Comparison of the prediction performance of three methods with metabolic enzymes of gibberellic acid sequences Sn Sp Ac Pr MCC SVM WEKA BLAST HMMER Different methods used in GAPred Accuracy HMMER BLAST WEKA SVM Figure 1: Comparison of performance validation of GAPred with different methods 3.3 ROC curve We have plotted the ROC curves for SVM and WEKA based on the independent test performance of the dipeptide compositions. From the ROC curves (Figures 2-3), representing the relationship between sensitivity and (1-specificity) for a class, it is clear that the SVM composition module represents a perfect classifier since the curve obtained is an inverted L, which is a desirable characteristic of an ROC curve. Each point on the ROC curve was plotted based on different threshold scores. Copyright Society for Science and Education United Kingdom 43
9 Sreepriya P., Naganeeswaran S., Hemalatha N., Sreejisha P. and Rajesh M. K. ; A Machine Learning Approach for Prediction of Gibberellic Acid Metabolic Enzymes in Monocotyledonous Plants; Transactions on Machine Learning and Artificial Intelligence, Volume 2 No 4, Aug (2014); pp: ROC curve for SVM multiclass cps Sensitivity ent-kaurene oxidase ent-kaurene synthase GA 2-oxidase 1-specificity GA 20- oxidase Figure 2: ROC curve for dipeptide composition in SVM multiclass using independent test results ROC curve for WEKA 0 Sensitivity 1-specificity Figure 3: ROC curve for dipeptide composition in WEKA using independent test results 3.4 Description of Web Server We have implemented the prediction tool GAPred in a web server. The tool was developed in PERL program and web interface in PHP and HTML to assess the user queries, in Linux platform. The tool page serve as the platform for submitting data where users can either paste or upload sequence which should be in standard FASTA format (Figure 4). It also provides a comprehensive collection of enzymes in GA metabolic pathway and introduces the user about gibberellic acid. The tool is freely available at URL: 44
10 Transactions on Machine Learning and Artificial Intelligence Volume 2, Issue 4, August 2104 Figure 4: Web interface of GAPred Figure 5: The architecture of the GAPred server. 4 CONCLUSION In this work, we have described SVM and WEKA-based approaches for the prediction of enzymes in gibberellic acid metabolic pathway based on dipeptide composition. Comparison of standalone BLAST and HMMER- based homology searches with machine learning algorithms revealed that the latter performed better compared to homology-based tools. Based on kernel methods, we have developed and implemented an efficient and easy to use user-friendly prediction server called GAPred for predicting five gibberellic acid metabolic enzymes. The sensitivity and specificity reaches 100% for prediction of gibberellic acid metabolic enzymes. We expect that the tool may be a useful resource for researchers as it is freely available. Copyright Society for Science and Education United Kingdom 45
11 Sreepriya P., Naganeeswaran S., Hemalatha N., Sreejisha P. and Rajesh M. K. ; A Machine Learning Approach for Prediction of Gibberellic Acid Metabolic Enzymes in Monocotyledonous Plants; Transactions on Machine Learning and Artificial Intelligence, Volume 2 No 4, Aug (2014); pp: REFERENCES [1]. Phinney, B.O., The history of gibberellins. In: The Biochemistry and Physiology of Gibberellins, Crozier, A. (Ed.), Praeger Publishers, New York, USA, : p [2]. Lichtenthaler, H.K., Rohmer, M. and Schwender, J., Two independent biochemical pathways for isopentenyl diphosphate biosynthesis in higher plants. Physiologia Plantarum, : p [3]. Graebe, J.E., Gibberellin biosynthesis and control. Annual Review of Plant Physiology, : p [4]. Chappell, J., Biochemistry and molecular biology of the isoprenoid biosynthetic pathway in plants. Annual Review of Plant Physiology and Plant Molecular Biology, : p [5]. MacMillan, J., Biosynthesis of the gibberellin plant hormones. Natural Product Reports, : [6]. Vapnik, V.N., An overview of statistical learning theory, Neural Networks. IEEE Transactions, : p [7]. Cortes, C. and Vapnik, V. Support Vector Networks. Machine Learning, : p [8]. Burges, C.J.C., A Tutorial on Support Vector Machines for Pattern Recognition. Data Mining and Knowledge Discovery, : p [9]. Noble, W.S., Support vector machine applications in computational biology. In: Kernel Methods in Computational Biology. Schoelkopf, B., Tsuda, K. and Vert, J. P. (Eds.), Cambridge, MA: MIT Press, p [10]. Joachims, T., Making large-scale SVM learning practical. In: Advances in Kernel Methods: Support Vector Learning. Schoelkopf, B.,Burges, C. and Smola,A. (Eds.), Cambridge MA: MIT Press, p [11]. Üney, F. and Türkay, M., A mixed-integer programming approach to multi-class data classification problem. European Journal of Operational Research, (3): p [12]. Chou, K.C. and Zhang, C.T., Prediction of protein structural classes. Critical Reviews in Biochemistry and Molecular Biology, : p [13]. Matthews, B.W., Comparison of the predicted and observed secondary structure of T4 phage lysozyme. Biochimica et Biophysica Acta, : p [14]. Baldi,P., Brunak, S., Chauvin, Y., Andersen, C.A.F. and Nielsen, H., Assessing the accuracy of prediction algorithms for classification: an overview. Bioinformatics, : p [15]. Carugo, O., Detailed estimation of bioinformatics prediction reliability through the Fragmented Prediction Performance Plots. BMC Bioinformatics, : p URL: 46
12 Transactions on Machine Learning and Artificial Intelligence Volume 2, Issue 4, August 2104 [16]. Park, J., Karplus, K., Barrett, C., Hughey, R., Haussler, D., Hubbard, T. and Chothia, C., Sequence comparisons using multiple sequences detect three times as many remote homologues as pair-wise methods. Journal of Molecular Biology, : p [17]. Hughey, R. and Krogh, A., Hidden Markov models for sequence analysis: extension and analysis of the basic method. Computer applications in the Biosciences, : p [18]. Mitchison, G.J. and Durbin, R., Tree-based maximal likelihood substitution matrices and hidden Markov models. Journal of Molecular Evolution, : p [19]. Swets, J.A., Measuring the accuracy of diagnostic systems. Science, : p Copyright Society for Science and Education United Kingdom 47
Prediction and Classif ication of Human G-protein Coupled Receptors Based on Support Vector Machines
 Article Prediction and Classif ication of Human G-protein Coupled Receptors Based on Support Vector Machines Yun-Fei Wang, Huan Chen, and Yan-Hong Zhou* Hubei Bioinformatics and Molecular Imaging Key Laboratory,
Article Prediction and Classif ication of Human G-protein Coupled Receptors Based on Support Vector Machines Yun-Fei Wang, Huan Chen, and Yan-Hong Zhou* Hubei Bioinformatics and Molecular Imaging Key Laboratory,
SUB-CELLULAR LOCALIZATION PREDICTION USING MACHINE LEARNING APPROACH
 SUB-CELLULAR LOCALIZATION PREDICTION USING MACHINE LEARNING APPROACH Ashutosh Kumar Singh 1, S S Sahu 2, Ankita Mishra 3 1,2,3 Birla Institute of Technology, Mesra, Ranchi Email: 1 ashutosh.4kumar.4singh@gmail.com,
SUB-CELLULAR LOCALIZATION PREDICTION USING MACHINE LEARNING APPROACH Ashutosh Kumar Singh 1, S S Sahu 2, Ankita Mishra 3 1,2,3 Birla Institute of Technology, Mesra, Ranchi Email: 1 ashutosh.4kumar.4singh@gmail.com,
Model Accuracy Measures
 Model Accuracy Measures Master in Bioinformatics UPF 2017-2018 Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA Barcelona, Spain Variables What we can measure (attributes) Hypotheses
Model Accuracy Measures Master in Bioinformatics UPF 2017-2018 Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA Barcelona, Spain Variables What we can measure (attributes) Hypotheses
Machine Learning Concepts in Chemoinformatics
 Machine Learning Concepts in Chemoinformatics Martin Vogt B-IT Life Science Informatics Rheinische Friedrich-Wilhelms-Universität Bonn BigChem Winter School 2017 25. October Data Mining in Chemoinformatics
Machine Learning Concepts in Chemoinformatics Martin Vogt B-IT Life Science Informatics Rheinische Friedrich-Wilhelms-Universität Bonn BigChem Winter School 2017 25. October Data Mining in Chemoinformatics
CISC 636 Computational Biology & Bioinformatics (Fall 2016)
 CISC 636 Computational Biology & Bioinformatics (Fall 2016) Predicting Protein-Protein Interactions CISC636, F16, Lec22, Liao 1 Background Proteins do not function as isolated entities. Protein-Protein
CISC 636 Computational Biology & Bioinformatics (Fall 2016) Predicting Protein-Protein Interactions CISC636, F16, Lec22, Liao 1 Background Proteins do not function as isolated entities. Protein-Protein
Week 10: Homology Modelling (II) - HHpred
 Week 10: Homology Modelling (II) - HHpred Course: Tools for Structural Biology Fabian Glaser BKU - Technion 1 2 Identify and align related structures by sequence methods is not an easy task All comparative
Week 10: Homology Modelling (II) - HHpred Course: Tools for Structural Biology Fabian Glaser BKU - Technion 1 2 Identify and align related structures by sequence methods is not an easy task All comparative
Machine Learning in Action
 Machine Learning in Action Tatyana Goldberg (goldberg@rostlab.org) August 16, 2016 @ Machine Learning in Biology Beijing Genomics Institute in Shenzhen, China June 2014 GenBank 1 173,353,076 DNA sequences
Machine Learning in Action Tatyana Goldberg (goldberg@rostlab.org) August 16, 2016 @ Machine Learning in Biology Beijing Genomics Institute in Shenzhen, China June 2014 GenBank 1 173,353,076 DNA sequences
Bioinformatics. Dept. of Computational Biology & Bioinformatics
 Bioinformatics Dept. of Computational Biology & Bioinformatics 3 Bioinformatics - play with sequences & structures Dept. of Computational Biology & Bioinformatics 4 ORGANIZATION OF LIFE ROLE OF BIOINFORMATICS
Bioinformatics Dept. of Computational Biology & Bioinformatics 3 Bioinformatics - play with sequences & structures Dept. of Computational Biology & Bioinformatics 4 ORGANIZATION OF LIFE ROLE OF BIOINFORMATICS
Statistical Machine Learning Methods for Bioinformatics II. Hidden Markov Model for Biological Sequences
 Statistical Machine Learning Methods for Bioinformatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD Department of Computer Science University of Missouri 2008 Free for Academic
Statistical Machine Learning Methods for Bioinformatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD Department of Computer Science University of Missouri 2008 Free for Academic
Plan. Lecture: What is Chemoinformatics and Drug Design? Description of Support Vector Machine (SVM) and its used in Chemoinformatics.
 Plan Lecture: What is Chemoinformatics and Drug Design? Description of Support Vector Machine (SVM) and its used in Chemoinformatics. Exercise: Example and exercise with herg potassium channel: Use of
Plan Lecture: What is Chemoinformatics and Drug Design? Description of Support Vector Machine (SVM) and its used in Chemoinformatics. Exercise: Example and exercise with herg potassium channel: Use of
Supplementary Materials for mplr-loc Web-server
 Supplementary Materials for mplr-loc Web-server Shibiao Wan and Man-Wai Mak email: shibiao.wan@connect.polyu.hk, enmwmak@polyu.edu.hk June 2014 Back to mplr-loc Server Contents 1 Introduction to mplr-loc
Supplementary Materials for mplr-loc Web-server Shibiao Wan and Man-Wai Mak email: shibiao.wan@connect.polyu.hk, enmwmak@polyu.edu.hk June 2014 Back to mplr-loc Server Contents 1 Introduction to mplr-loc
Statistical Machine Learning Methods for Biomedical Informatics II. Hidden Markov Model for Biological Sequences
 Statistical Machine Learning Methods for Biomedical Informatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD William and Nancy Thompson Missouri Distinguished Professor Department
Statistical Machine Learning Methods for Biomedical Informatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD William and Nancy Thompson Missouri Distinguished Professor Department
PROTEIN FUNCTION PREDICTION WITH AMINO ACID SEQUENCE AND SECONDARY STRUCTURE ALIGNMENT SCORES
 PROTEIN FUNCTION PREDICTION WITH AMINO ACID SEQUENCE AND SECONDARY STRUCTURE ALIGNMENT SCORES Eser Aygün 1, Caner Kömürlü 2, Zafer Aydin 3 and Zehra Çataltepe 1 1 Computer Engineering Department and 2
PROTEIN FUNCTION PREDICTION WITH AMINO ACID SEQUENCE AND SECONDARY STRUCTURE ALIGNMENT SCORES Eser Aygün 1, Caner Kömürlü 2, Zafer Aydin 3 and Zehra Çataltepe 1 1 Computer Engineering Department and 2
HYPERGRAPH BASED SEMI-SUPERVISED LEARNING ALGORITHMS APPLIED TO SPEECH RECOGNITION PROBLEM: A NOVEL APPROACH
 HYPERGRAPH BASED SEMI-SUPERVISED LEARNING ALGORITHMS APPLIED TO SPEECH RECOGNITION PROBLEM: A NOVEL APPROACH Hoang Trang 1, Tran Hoang Loc 1 1 Ho Chi Minh City University of Technology-VNU HCM, Ho Chi
HYPERGRAPH BASED SEMI-SUPERVISED LEARNING ALGORITHMS APPLIED TO SPEECH RECOGNITION PROBLEM: A NOVEL APPROACH Hoang Trang 1, Tran Hoang Loc 1 1 Ho Chi Minh City University of Technology-VNU HCM, Ho Chi
Protein Bioinformatics. Rickard Sandberg Dept. of Cell and Molecular Biology Karolinska Institutet sandberg.cmb.ki.
 Protein Bioinformatics Rickard Sandberg Dept. of Cell and Molecular Biology Karolinska Institutet rickard.sandberg@ki.se sandberg.cmb.ki.se Outline Protein features motifs patterns profiles signals 2 Protein
Protein Bioinformatics Rickard Sandberg Dept. of Cell and Molecular Biology Karolinska Institutet rickard.sandberg@ki.se sandberg.cmb.ki.se Outline Protein features motifs patterns profiles signals 2 Protein
Efficient Remote Homology Detection with Secondary Structure
 Efficient Remote Homology Detection with Secondary Structure 2 Yuna Hou 1, Wynne Hsu 1, Mong Li Lee 1, and Christopher Bystroff 2 1 School of Computing,National University of Singapore,Singapore 117543
Efficient Remote Homology Detection with Secondary Structure 2 Yuna Hou 1, Wynne Hsu 1, Mong Li Lee 1, and Christopher Bystroff 2 1 School of Computing,National University of Singapore,Singapore 117543
Grundlagen der Bioinformatik Summer semester Lecturer: Prof. Daniel Huson
 Grundlagen der Bioinformatik, SS 10, D. Huson, April 12, 2010 1 1 Introduction Grundlagen der Bioinformatik Summer semester 2010 Lecturer: Prof. Daniel Huson Office hours: Thursdays 17-18h (Sand 14, C310a)
Grundlagen der Bioinformatik, SS 10, D. Huson, April 12, 2010 1 1 Introduction Grundlagen der Bioinformatik Summer semester 2010 Lecturer: Prof. Daniel Huson Office hours: Thursdays 17-18h (Sand 14, C310a)
hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference
 CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
Transductive learning with EM algorithm to classify proteins based on phylogenetic profiles
 Int. J. Data Mining and Bioinformatics, Vol. 1, No. 4, 2007 337 Transductive learning with EM algorithm to classify proteins based on phylogenetic profiles Roger A. Craig and Li Liao* Department of Computer
Int. J. Data Mining and Bioinformatics, Vol. 1, No. 4, 2007 337 Transductive learning with EM algorithm to classify proteins based on phylogenetic profiles Roger A. Craig and Li Liao* Department of Computer
Data Mining: Concepts and Techniques. (3 rd ed.) Chapter 8. Chapter 8. Classification: Basic Concepts
 Data Mining: Concepts and Techniques (3 rd ed.) Chapter 8 1 Chapter 8. Classification: Basic Concepts Classification: Basic Concepts Decision Tree Induction Bayes Classification Methods Rule-Based Classification
Data Mining: Concepts and Techniques (3 rd ed.) Chapter 8 1 Chapter 8. Classification: Basic Concepts Classification: Basic Concepts Decision Tree Induction Bayes Classification Methods Rule-Based Classification
Supplementary Materials for R3P-Loc Web-server
 Supplementary Materials for R3P-Loc Web-server Shibiao Wan and Man-Wai Mak email: shibiao.wan@connect.polyu.hk, enmwmak@polyu.edu.hk June 2014 Back to R3P-Loc Server Contents 1 Introduction to R3P-Loc
Supplementary Materials for R3P-Loc Web-server Shibiao Wan and Man-Wai Mak email: shibiao.wan@connect.polyu.hk, enmwmak@polyu.edu.hk June 2014 Back to R3P-Loc Server Contents 1 Introduction to R3P-Loc
Statistical Machine Learning Methods for Bioinformatics IV. Neural Network & Deep Learning Applications in Bioinformatics
 Statistical Machine Learning Methods for Bioinformatics IV. Neural Network & Deep Learning Applications in Bioinformatics Jianlin Cheng, PhD Department of Computer Science University of Missouri, Columbia
Statistical Machine Learning Methods for Bioinformatics IV. Neural Network & Deep Learning Applications in Bioinformatics Jianlin Cheng, PhD Department of Computer Science University of Missouri, Columbia
BANA 7046 Data Mining I Lecture 4. Logistic Regression and Classications 1
 BANA 7046 Data Mining I Lecture 4. Logistic Regression and Classications 1 Shaobo Li University of Cincinnati 1 Partially based on Hastie, et al. (2009) ESL, and James, et al. (2013) ISLR Data Mining I
BANA 7046 Data Mining I Lecture 4. Logistic Regression and Classications 1 Shaobo Li University of Cincinnati 1 Partially based on Hastie, et al. (2009) ESL, and James, et al. (2013) ISLR Data Mining I
Performance Evaluation
 Performance Evaluation David S. Rosenberg Bloomberg ML EDU October 26, 2017 David S. Rosenberg (Bloomberg ML EDU) October 26, 2017 1 / 36 Baseline Models David S. Rosenberg (Bloomberg ML EDU) October 26,
Performance Evaluation David S. Rosenberg Bloomberg ML EDU October 26, 2017 David S. Rosenberg (Bloomberg ML EDU) October 26, 2017 1 / 36 Baseline Models David S. Rosenberg (Bloomberg ML EDU) October 26,
Performance Evaluation
 Performance Evaluation Confusion Matrix: Detected Positive Negative Actual Positive A: True Positive B: False Negative Negative C: False Positive D: True Negative Recall or Sensitivity or True Positive
Performance Evaluation Confusion Matrix: Detected Positive Negative Actual Positive A: True Positive B: False Negative Negative C: False Positive D: True Negative Recall or Sensitivity or True Positive
Visualize Biological Database for Protein in Homosapiens Using Classification Searching Models
 International Journal of Computational Intelligence Research ISSN 0973-1873 Volume 13, Number 2 (2017), pp. 213-224 Research India Publications http://www.ripublication.com Visualize Biological Database
International Journal of Computational Intelligence Research ISSN 0973-1873 Volume 13, Number 2 (2017), pp. 213-224 Research India Publications http://www.ripublication.com Visualize Biological Database
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 07: profile Hidden Markov Model http://bibiserv.techfak.uni-bielefeld.de/sadr2/databasesearch/hmmer/profilehmm.gif Slides adapted from Dr. Shaojie Zhang
EECS730: Introduction to Bioinformatics Lecture 07: profile Hidden Markov Model http://bibiserv.techfak.uni-bielefeld.de/sadr2/databasesearch/hmmer/profilehmm.gif Slides adapted from Dr. Shaojie Zhang
Computational Biology Course Descriptions 12-14
 Computational Biology Course Descriptions 12-14 Course Number and Title INTRODUCTORY COURSES BIO 311C: Introductory Biology I BIO 311D: Introductory Biology II BIO 325: Genetics CH 301: Principles of Chemistry
Computational Biology Course Descriptions 12-14 Course Number and Title INTRODUCTORY COURSES BIO 311C: Introductory Biology I BIO 311D: Introductory Biology II BIO 325: Genetics CH 301: Principles of Chemistry
Shibiao Wan and Man-Wai Mak December 2013 Back to HybridGO-Loc Server
 Shibiao Wan and Man-Wai Mak December 2013 Back to HybridGO-Loc Server Contents 1 Functions of HybridGO-Loc Server 2 1.1 Webserver Interface....................................... 2 1.2 Inputing Protein
Shibiao Wan and Man-Wai Mak December 2013 Back to HybridGO-Loc Server Contents 1 Functions of HybridGO-Loc Server 2 1.1 Webserver Interface....................................... 2 1.2 Inputing Protein
Text Mining. Dr. Yanjun Li. Associate Professor. Department of Computer and Information Sciences Fordham University
 Text Mining Dr. Yanjun Li Associate Professor Department of Computer and Information Sciences Fordham University Outline Introduction: Data Mining Part One: Text Mining Part Two: Preprocessing Text Data
Text Mining Dr. Yanjun Li Associate Professor Department of Computer and Information Sciences Fordham University Outline Introduction: Data Mining Part One: Text Mining Part Two: Preprocessing Text Data
Online Estimation of Discrete Densities using Classifier Chains
 Online Estimation of Discrete Densities using Classifier Chains Michael Geilke 1 and Eibe Frank 2 and Stefan Kramer 1 1 Johannes Gutenberg-Universtität Mainz, Germany {geilke,kramer}@informatik.uni-mainz.de
Online Estimation of Discrete Densities using Classifier Chains Michael Geilke 1 and Eibe Frank 2 and Stefan Kramer 1 1 Johannes Gutenberg-Universtität Mainz, Germany {geilke,kramer}@informatik.uni-mainz.de
Combining pairwise sequence similarity and support vector machines for remote protein homology detection
 Combining pairwise sequence similarity and support vector machines for remote protein homology detection Li Liao Central Research & Development E. I. du Pont de Nemours Company li.liao@usa.dupont.com William
Combining pairwise sequence similarity and support vector machines for remote protein homology detection Li Liao Central Research & Development E. I. du Pont de Nemours Company li.liao@usa.dupont.com William
Evaluation & Credibility Issues
 Evaluation & Credibility Issues What measure should we use? accuracy might not be enough. How reliable are the predicted results? How much should we believe in what was learned? Error on the training data
Evaluation & Credibility Issues What measure should we use? accuracy might not be enough. How reliable are the predicted results? How much should we believe in what was learned? Error on the training data
Sequence Alignment Techniques and Their Uses
 Sequence Alignment Techniques and Their Uses Sarah Fiorentino Since rapid sequencing technology and whole genomes sequencing, the amount of sequence information has grown exponentially. With all of this
Sequence Alignment Techniques and Their Uses Sarah Fiorentino Since rapid sequencing technology and whole genomes sequencing, the amount of sequence information has grown exponentially. With all of this
SINGLE-SEQUENCE PROTEIN SECONDARY STRUCTURE PREDICTION BY NEAREST-NEIGHBOR CLASSIFICATION OF PROTEIN WORDS
 SINGLE-SEQUENCE PROTEIN SECONDARY STRUCTURE PREDICTION BY NEAREST-NEIGHBOR CLASSIFICATION OF PROTEIN WORDS Item type Authors Publisher Rights text; Electronic Thesis PORFIRIO, DAVID JONATHAN The University
SINGLE-SEQUENCE PROTEIN SECONDARY STRUCTURE PREDICTION BY NEAREST-NEIGHBOR CLASSIFICATION OF PROTEIN WORDS Item type Authors Publisher Rights text; Electronic Thesis PORFIRIO, DAVID JONATHAN The University
Computational Molecular Biology (
 Computational Molecular Biology (http://cmgm cmgm.stanford.edu/biochem218/) Biochemistry 218/Medical Information Sciences 231 Douglas L. Brutlag, Lee Kozar Jimmy Huang, Josh Silverman Lecture Syllabus
Computational Molecular Biology (http://cmgm cmgm.stanford.edu/biochem218/) Biochemistry 218/Medical Information Sciences 231 Douglas L. Brutlag, Lee Kozar Jimmy Huang, Josh Silverman Lecture Syllabus
HMM applications. Applications of HMMs. Gene finding with HMMs. Using the gene finder
 HMM applications Applications of HMMs Gene finding Pairwise alignment (pair HMMs) Characterizing protein families (profile HMMs) Predicting membrane proteins, and membrane protein topology Gene finding
HMM applications Applications of HMMs Gene finding Pairwise alignment (pair HMMs) Characterizing protein families (profile HMMs) Predicting membrane proteins, and membrane protein topology Gene finding
Combining pairwise sequence similarity and support vector machines for remote protein homology detection
 Combining pairwise sequence similarity and support vector machines for remote protein homology detection Li Liao Central Research & Development E. I. du Pont de Nemours Company li.liao@usa.dupont.com William
Combining pairwise sequence similarity and support vector machines for remote protein homology detection Li Liao Central Research & Development E. I. du Pont de Nemours Company li.liao@usa.dupont.com William
An Artificial Neural Network Classifier for the Prediction of Protein Structural Classes
 International Journal of Current Engineering and Technology E-ISSN 2277 4106, P-ISSN 2347 5161 2017 INPRESSCO, All Rights Reserved Available at http://inpressco.com/category/ijcet Research Article An Artificial
International Journal of Current Engineering and Technology E-ISSN 2277 4106, P-ISSN 2347 5161 2017 INPRESSCO, All Rights Reserved Available at http://inpressco.com/category/ijcet Research Article An Artificial
Protein Structure Prediction Using Neural Networks
 Protein Structure Prediction Using Neural Networks Martha Mercaldi Kasia Wilamowska Literature Review December 16, 2003 The Protein Folding Problem Evolution of Neural Networks Neural networks originally
Protein Structure Prediction Using Neural Networks Martha Mercaldi Kasia Wilamowska Literature Review December 16, 2003 The Protein Folding Problem Evolution of Neural Networks Neural networks originally
Sequence Alignment: A General Overview. COMP Fall 2010 Luay Nakhleh, Rice University
 Sequence Alignment: A General Overview COMP 571 - Fall 2010 Luay Nakhleh, Rice University Life through Evolution All living organisms are related to each other through evolution This means: any pair of
Sequence Alignment: A General Overview COMP 571 - Fall 2010 Luay Nakhleh, Rice University Life through Evolution All living organisms are related to each other through evolution This means: any pair of
Support Vector Machines (SVM) in bioinformatics. Day 1: Introduction to SVM
 1 Support Vector Machines (SVM) in bioinformatics Day 1: Introduction to SVM Jean-Philippe Vert Bioinformatics Center, Kyoto University, Japan Jean-Philippe.Vert@mines.org Human Genome Center, University
1 Support Vector Machines (SVM) in bioinformatics Day 1: Introduction to SVM Jean-Philippe Vert Bioinformatics Center, Kyoto University, Japan Jean-Philippe.Vert@mines.org Human Genome Center, University
Motifs, Profiles and Domains. Michael Tress Protein Design Group Centro Nacional de Biotecnología, CSIC
 Motifs, Profiles and Domains Michael Tress Protein Design Group Centro Nacional de Biotecnología, CSIC Comparing Two Proteins Sequence Alignment Determining the pattern of evolution and identifying conserved
Motifs, Profiles and Domains Michael Tress Protein Design Group Centro Nacional de Biotecnología, CSIC Comparing Two Proteins Sequence Alignment Determining the pattern of evolution and identifying conserved
Protein-Protein Interaction Classification Using Jordan Recurrent Neural Network
 Protein-Protein Interaction Classification Using Jordan Recurrent Neural Network Dilpreet Kaur Department of Computer Science and Engineering PEC University of Technology Chandigarh, India dilpreet.kaur88@gmail.com
Protein-Protein Interaction Classification Using Jordan Recurrent Neural Network Dilpreet Kaur Department of Computer Science and Engineering PEC University of Technology Chandigarh, India dilpreet.kaur88@gmail.com
SVM Kernel Optimization: An Example in Yeast Protein Subcellular Localization Prediction
 SVM Kernel Optimization: An Example in Yeast Protein Subcellular Localization Prediction Ṭaráz E. Buck Computational Biology Program tebuck@andrew.cmu.edu Bin Zhang School of Public Policy and Management
SVM Kernel Optimization: An Example in Yeast Protein Subcellular Localization Prediction Ṭaráz E. Buck Computational Biology Program tebuck@andrew.cmu.edu Bin Zhang School of Public Policy and Management
Using Ensembles of Hidden Markov Models for Grand Challenges in Bioinformatics
 Using Ensembles of Hidden Markov Models for Grand Challenges in Bioinformatics Tandy Warnow Founder Professor of Engineering The University of Illinois at Urbana-Champaign http://tandy.cs.illinois.edu
Using Ensembles of Hidden Markov Models for Grand Challenges in Bioinformatics Tandy Warnow Founder Professor of Engineering The University of Illinois at Urbana-Champaign http://tandy.cs.illinois.edu
Supplementary Information
 Supplementary Information Performance measures A binary classifier, such as SVM, assigns to predicted binding sequences the positive class label (+1) and to sequences predicted as non-binding the negative
Supplementary Information Performance measures A binary classifier, such as SVM, assigns to predicted binding sequences the positive class label (+1) and to sequences predicted as non-binding the negative
Hidden Markov Models (HMMs) and Profiles
 Hidden Markov Models (HMMs) and Profiles Swiss Institute of Bioinformatics (SIB) 26-30 November 2001 Markov Chain Models A Markov Chain Model is a succession of states S i (i = 0, 1,...) connected by transitions.
Hidden Markov Models (HMMs) and Profiles Swiss Institute of Bioinformatics (SIB) 26-30 November 2001 Markov Chain Models A Markov Chain Model is a succession of states S i (i = 0, 1,...) connected by transitions.
SUPERVISED LEARNING: INTRODUCTION TO CLASSIFICATION
 SUPERVISED LEARNING: INTRODUCTION TO CLASSIFICATION 1 Outline Basic terminology Features Training and validation Model selection Error and loss measures Statistical comparison Evaluation measures 2 Terminology
SUPERVISED LEARNING: INTRODUCTION TO CLASSIFICATION 1 Outline Basic terminology Features Training and validation Model selection Error and loss measures Statistical comparison Evaluation measures 2 Terminology
ROUGH SET PROTEIN CLASSIFIER
 ROUGH SET PROTEIN CLASSIFIER RAMADEVI YELLASIRI 1, C.R.RAO 2 1 Dept. of CSE, Chaitanya Bharathi Institute of Technology, Hyderabad, INDIA. 2 DCIS, School of MCIS, University of Hyderabad, Hyderabad, INDIA.
ROUGH SET PROTEIN CLASSIFIER RAMADEVI YELLASIRI 1, C.R.RAO 2 1 Dept. of CSE, Chaitanya Bharathi Institute of Technology, Hyderabad, INDIA. 2 DCIS, School of MCIS, University of Hyderabad, Hyderabad, INDIA.
Hands-On Nine The PAX6 Gene and Protein
 Hands-On Nine The PAX6 Gene and Protein Main Purpose of Hands-On Activity: Using bioinformatics tools to examine the sequences, homology, and disease relevance of the Pax6: a master gene of eye formation.
Hands-On Nine The PAX6 Gene and Protein Main Purpose of Hands-On Activity: Using bioinformatics tools to examine the sequences, homology, and disease relevance of the Pax6: a master gene of eye formation.
STRUCTURAL BIOINFORMATICS I. Fall 2015
 STRUCTURAL BIOINFORMATICS I Fall 2015 Info Course Number - Classification: Biology 5411 Class Schedule: Monday 5:30-7:50 PM, SERC Room 456 (4 th floor) Instructors: Vincenzo Carnevale - SERC, Room 704C;
STRUCTURAL BIOINFORMATICS I Fall 2015 Info Course Number - Classification: Biology 5411 Class Schedule: Monday 5:30-7:50 PM, SERC Room 456 (4 th floor) Instructors: Vincenzo Carnevale - SERC, Room 704C;
METABOLIC PATHWAY PREDICTION/ALIGNMENT
 COMPUTATIONAL SYSTEMIC BIOLOGY METABOLIC PATHWAY PREDICTION/ALIGNMENT Hofestaedt R*, Chen M Bioinformatics / Medical Informatics, Technische Fakultaet, Universitaet Bielefeld Postfach 10 01 31, D-33501
COMPUTATIONAL SYSTEMIC BIOLOGY METABOLIC PATHWAY PREDICTION/ALIGNMENT Hofestaedt R*, Chen M Bioinformatics / Medical Informatics, Technische Fakultaet, Universitaet Bielefeld Postfach 10 01 31, D-33501
Class 4: Classification. Quaid Morris February 11 th, 2011 ML4Bio
 Class 4: Classification Quaid Morris February 11 th, 211 ML4Bio Overview Basic concepts in classification: overfitting, cross-validation, evaluation. Linear Discriminant Analysis and Quadratic Discriminant
Class 4: Classification Quaid Morris February 11 th, 211 ML4Bio Overview Basic concepts in classification: overfitting, cross-validation, evaluation. Linear Discriminant Analysis and Quadratic Discriminant
Bayesian Decision Theory
 Introduction to Pattern Recognition [ Part 4 ] Mahdi Vasighi Remarks It is quite common to assume that the data in each class are adequately described by a Gaussian distribution. Bayesian classifier is
Introduction to Pattern Recognition [ Part 4 ] Mahdi Vasighi Remarks It is quite common to assume that the data in each class are adequately described by a Gaussian distribution. Bayesian classifier is
A Tutorial on Support Vector Machine
 A Tutorial on School of Computing National University of Singapore Contents Theory on Using with Other s Contents Transforming Theory on Using with Other s What is a classifier? A function that maps instances
A Tutorial on School of Computing National University of Singapore Contents Theory on Using with Other s Contents Transforming Theory on Using with Other s What is a classifier? A function that maps instances
BIOINFORMATICS: An Introduction
 BIOINFORMATICS: An Introduction What is Bioinformatics? The term was first coined in 1988 by Dr. Hwa Lim The original definition was : a collective term for data compilation, organisation, analysis and
BIOINFORMATICS: An Introduction What is Bioinformatics? The term was first coined in 1988 by Dr. Hwa Lim The original definition was : a collective term for data compilation, organisation, analysis and
Truncated Profile Hidden Markov Models
 Boise State University ScholarWorks Electrical and Computer Engineering Faculty Publications and Presentations Department of Electrical and Computer Engineering 11-1-2005 Truncated Profile Hidden Markov
Boise State University ScholarWorks Electrical and Computer Engineering Faculty Publications and Presentations Department of Electrical and Computer Engineering 11-1-2005 Truncated Profile Hidden Markov
Carson Andorf 1,3, Adrian Silvescu 1,3, Drena Dobbs 2,3,4, Vasant Honavar 1,3,4. University, Ames, Iowa, 50010, USA. Ames, Iowa, 50010, USA
 Learning Classifiers for Assigning Protein Sequences to Gene Ontology Functional Families: Combining of Function Annotation Using Sequence Homology With that Based on Amino Acid k-gram Composition Yields
Learning Classifiers for Assigning Protein Sequences to Gene Ontology Functional Families: Combining of Function Annotation Using Sequence Homology With that Based on Amino Acid k-gram Composition Yields
Prediction of the subcellular location of apoptosis proteins based on approximate entropy
 Prediction of the subcellular location of apoptosis proteins based on approximate entropy Chaohong Song Feng Shi* Corresponding author Xuan Ma College of Science, Huazhong Agricultural University, Wuhan,
Prediction of the subcellular location of apoptosis proteins based on approximate entropy Chaohong Song Feng Shi* Corresponding author Xuan Ma College of Science, Huazhong Agricultural University, Wuhan,
Hidden Markov Models in computational biology. Ron Elber Computer Science Cornell
 Hidden Markov Models in computational biology Ron Elber Computer Science Cornell 1 Or: how to fish homolog sequences from a database Many sequences in database RPOBESEQ Partitioned data base 2 An accessible
Hidden Markov Models in computational biology Ron Elber Computer Science Cornell 1 Or: how to fish homolog sequences from a database Many sequences in database RPOBESEQ Partitioned data base 2 An accessible
Computational methods for predicting protein-protein interactions
 Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
IMPORTANCE OF SECONDARY STRUCTURE ELEMENTS FOR PREDICTION OF GO ANNOTATIONS
 IMPORTANCE OF SECONDARY STRUCTURE ELEMENTS FOR PREDICTION OF GO ANNOTATIONS Aslı Filiz 1, Eser Aygün 2, Özlem Keskin 3 and Zehra Cataltepe 2 1 Informatics Institute and 2 Computer Engineering Department,
IMPORTANCE OF SECONDARY STRUCTURE ELEMENTS FOR PREDICTION OF GO ANNOTATIONS Aslı Filiz 1, Eser Aygün 2, Özlem Keskin 3 and Zehra Cataltepe 2 1 Informatics Institute and 2 Computer Engineering Department,
Computational methods for the analysis of bacterial gene regulation Brouwer, Rutger Wubbe Willem
 University of Groningen Computational methods for the analysis of bacterial gene regulation Brouwer, Rutger Wubbe Willem IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's
University of Groningen Computational methods for the analysis of bacterial gene regulation Brouwer, Rutger Wubbe Willem IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's
DETECTING HUMAN ACTIVITIES IN THE ARCTIC OCEAN BY CONSTRUCTING AND ANALYZING SUPER-RESOLUTION IMAGES FROM MODIS DATA INTRODUCTION
 DETECTING HUMAN ACTIVITIES IN THE ARCTIC OCEAN BY CONSTRUCTING AND ANALYZING SUPER-RESOLUTION IMAGES FROM MODIS DATA Shizhi Chen and YingLi Tian Department of Electrical Engineering The City College of
DETECTING HUMAN ACTIVITIES IN THE ARCTIC OCEAN BY CONSTRUCTING AND ANALYZING SUPER-RESOLUTION IMAGES FROM MODIS DATA Shizhi Chen and YingLi Tian Department of Electrical Engineering The City College of
K-means-based Feature Learning for Protein Sequence Classification
 K-means-based Feature Learning for Protein Sequence Classification Paul Melman and Usman W. Roshan Department of Computer Science, NJIT Newark, NJ, 07102, USA pm462@njit.edu, usman.w.roshan@njit.edu Abstract
K-means-based Feature Learning for Protein Sequence Classification Paul Melman and Usman W. Roshan Department of Computer Science, NJIT Newark, NJ, 07102, USA pm462@njit.edu, usman.w.roshan@njit.edu Abstract
Application of Associative Matrices to Recognize DNA Sequences in Bioinformatics
 Application of Associative Matrices to Recognize DNA Sequences in Bioinformatics 1. Introduction. Jorge L. Ortiz Department of Electrical and Computer Engineering College of Engineering University of Puerto
Application of Associative Matrices to Recognize DNA Sequences in Bioinformatics 1. Introduction. Jorge L. Ortiz Department of Electrical and Computer Engineering College of Engineering University of Puerto
A Novel Prediction Method of Protein Structural Classes Based on Protein Super-Secondary Structure
 Journal of Computer and Communications, 2016, 4, 54-62 http://www.scirp.org/journal/jcc ISSN Online: 2327-5227 ISSN Print: 2327-5219 A Novel Prediction Method of Protein Structural Classes Based on Protein
Journal of Computer and Communications, 2016, 4, 54-62 http://www.scirp.org/journal/jcc ISSN Online: 2327-5227 ISSN Print: 2327-5219 A Novel Prediction Method of Protein Structural Classes Based on Protein
Microarray Data Analysis: Discovery
 Microarray Data Analysis: Discovery Lecture 5 Classification Classification vs. Clustering Classification: Goal: Placing objects (e.g. genes) into meaningful classes Supervised Clustering: Goal: Discover
Microarray Data Analysis: Discovery Lecture 5 Classification Classification vs. Clustering Classification: Goal: Placing objects (e.g. genes) into meaningful classes Supervised Clustering: Goal: Discover
Support Vector Machine & Its Applications
 Support Vector Machine & Its Applications A portion (1/3) of the slides are taken from Prof. Andrew Moore s SVM tutorial at http://www.cs.cmu.edu/~awm/tutorials Mingyue Tan The University of British Columbia
Support Vector Machine & Its Applications A portion (1/3) of the slides are taken from Prof. Andrew Moore s SVM tutorial at http://www.cs.cmu.edu/~awm/tutorials Mingyue Tan The University of British Columbia
ALL LECTURES IN SB Introduction
 1. Introduction 2. Molecular Architecture I 3. Molecular Architecture II 4. Molecular Simulation I 5. Molecular Simulation II 6. Bioinformatics I 7. Bioinformatics II 8. Prediction I 9. Prediction II ALL
1. Introduction 2. Molecular Architecture I 3. Molecular Architecture II 4. Molecular Simulation I 5. Molecular Simulation II 6. Bioinformatics I 7. Bioinformatics II 8. Prediction I 9. Prediction II ALL
A Hybrid Support Vector Machine Method for Protein Remote Homology Detection
 , March 14-16, 18, Hong Kong A Hybrid Support Vector Machine Method for Protein Remote Homology Detection Jiang Xie*, Dongfang Lu, Junhui Shu, Jiao Wang*, Haiya Wang, Chao Meng, Wu Zhang Abstract Remote
, March 14-16, 18, Hong Kong A Hybrid Support Vector Machine Method for Protein Remote Homology Detection Jiang Xie*, Dongfang Lu, Junhui Shu, Jiao Wang*, Haiya Wang, Chao Meng, Wu Zhang Abstract Remote
Building 3D models of proteins
 Building 3D models of proteins Why make a structural model for your protein? The structure can provide clues to the function through structural similarity with other proteins With a structure it is easier
Building 3D models of proteins Why make a structural model for your protein? The structure can provide clues to the function through structural similarity with other proteins With a structure it is easier
Why Bother? Predicting the Cellular Localization Sites of Proteins Using Bayesian Model Averaging. Yetian Chen
 Predicting the Cellular Localization Sites of Proteins Using Bayesian Model Averaging Yetian Chen 04-27-2010 Why Bother? 2008 Nobel Prize in Chemistry Roger Tsien Osamu Shimomura Martin Chalfie Green Fluorescent
Predicting the Cellular Localization Sites of Proteins Using Bayesian Model Averaging Yetian Chen 04-27-2010 Why Bother? 2008 Nobel Prize in Chemistry Roger Tsien Osamu Shimomura Martin Chalfie Green Fluorescent
The Homology Kernel: A Biologically Motivated Sequence Embedding into Euclidean Space
 The Homology Kernel: A Biologically Motivated Sequence Embedding into Euclidean Space Eleazar Eskin Department of Computer Science Engineering University of California, San Diego eeskin@cs.ucsd.edu Sagi
The Homology Kernel: A Biologically Motivated Sequence Embedding into Euclidean Space Eleazar Eskin Department of Computer Science Engineering University of California, San Diego eeskin@cs.ucsd.edu Sagi
Data Mining and Knowledge Discovery: Practice Notes
 Data Mining and Knowledge Discovery: Practice Notes dr. Petra Kralj Novak Petra.Kralj.Novak@ijs.si 7.11.2017 1 Course Prof. Bojan Cestnik Data preparation Prof. Nada Lavrač: Data mining overview Advanced
Data Mining and Knowledge Discovery: Practice Notes dr. Petra Kralj Novak Petra.Kralj.Novak@ijs.si 7.11.2017 1 Course Prof. Bojan Cestnik Data preparation Prof. Nada Lavrač: Data mining overview Advanced
Gene Expression Data Classification with Revised Kernel Partial Least Squares Algorithm
 Gene Expression Data Classification with Revised Kernel Partial Least Squares Algorithm Zhenqiu Liu, Dechang Chen 2 Department of Computer Science Wayne State University, Market Street, Frederick, MD 273,
Gene Expression Data Classification with Revised Kernel Partial Least Squares Algorithm Zhenqiu Liu, Dechang Chen 2 Department of Computer Science Wayne State University, Market Street, Frederick, MD 273,
Neural Networks for Protein Structure Prediction Brown, JMB CS 466 Saurabh Sinha
 Neural Networks for Protein Structure Prediction Brown, JMB 1999 CS 466 Saurabh Sinha Outline Goal is to predict secondary structure of a protein from its sequence Artificial Neural Network used for this
Neural Networks for Protein Structure Prediction Brown, JMB 1999 CS 466 Saurabh Sinha Outline Goal is to predict secondary structure of a protein from its sequence Artificial Neural Network used for this
SUPPORT VECTOR REGRESSION WITH A GENERALIZED QUADRATIC LOSS
 SUPPORT VECTOR REGRESSION WITH A GENERALIZED QUADRATIC LOSS Filippo Portera and Alessandro Sperduti Dipartimento di Matematica Pura ed Applicata Universit a di Padova, Padova, Italy {portera,sperduti}@math.unipd.it
SUPPORT VECTOR REGRESSION WITH A GENERALIZED QUADRATIC LOSS Filippo Portera and Alessandro Sperduti Dipartimento di Matematica Pura ed Applicata Universit a di Padova, Padova, Italy {portera,sperduti}@math.unipd.it
Moving Average Rules to Find. Confusion Matrix. CC283 Intelligent Problem Solving 05/11/2010. Edward Tsang (all rights reserved) 1
 Machine Learning Overview Supervised Learning Training esting Te Unseen data Data Observed x 1 x 2... x n 1.6 7.1... 2.7 1.4 6.8... 3.1 2.1 5.4... 2.8... Machine Learning Patterns y = f(x) Target y Buy
Machine Learning Overview Supervised Learning Training esting Te Unseen data Data Observed x 1 x 2... x n 1.6 7.1... 2.7 1.4 6.8... 3.1 2.1 5.4... 2.8... Machine Learning Patterns y = f(x) Target y Buy
Stephen Scott.
 1 / 35 (Adapted from Ethem Alpaydin and Tom Mitchell) sscott@cse.unl.edu In Homework 1, you are (supposedly) 1 Choosing a data set 2 Extracting a test set of size > 30 3 Building a tree on the training
1 / 35 (Adapted from Ethem Alpaydin and Tom Mitchell) sscott@cse.unl.edu In Homework 1, you are (supposedly) 1 Choosing a data set 2 Extracting a test set of size > 30 3 Building a tree on the training
Comparing Robustness of Pairwise and Multiclass Neural-Network Systems for Face Recognition
 Comparing Robustness of Pairwise and Multiclass Neural-Network Systems for Face Recognition J. Uglov, V. Schetinin, C. Maple Computing and Information System Department, University of Bedfordshire, Luton,
Comparing Robustness of Pairwise and Multiclass Neural-Network Systems for Face Recognition J. Uglov, V. Schetinin, C. Maple Computing and Information System Department, University of Bedfordshire, Luton,
CS612 - Algorithms in Bioinformatics
 Fall 2017 Databases and Protein Structure Representation October 2, 2017 Molecular Biology as Information Science > 12, 000 genomes sequenced, mostly bacterial (2013) > 5x10 6 unique sequences available
Fall 2017 Databases and Protein Structure Representation October 2, 2017 Molecular Biology as Information Science > 12, 000 genomes sequenced, mostly bacterial (2013) > 5x10 6 unique sequences available
Lecture 4 Discriminant Analysis, k-nearest Neighbors
 Lecture 4 Discriminant Analysis, k-nearest Neighbors Fredrik Lindsten Division of Systems and Control Department of Information Technology Uppsala University. Email: fredrik.lindsten@it.uu.se fredrik.lindsten@it.uu.se
Lecture 4 Discriminant Analysis, k-nearest Neighbors Fredrik Lindsten Division of Systems and Control Department of Information Technology Uppsala University. Email: fredrik.lindsten@it.uu.se fredrik.lindsten@it.uu.se
Data Privacy in Biomedicine. Lecture 11b: Performance Measures for System Evaluation
 Data Privacy in Biomedicine Lecture 11b: Performance Measures for System Evaluation Bradley Malin, PhD (b.malin@vanderbilt.edu) Professor of Biomedical Informatics, Biostatistics, & Computer Science Vanderbilt
Data Privacy in Biomedicine Lecture 11b: Performance Measures for System Evaluation Bradley Malin, PhD (b.malin@vanderbilt.edu) Professor of Biomedical Informatics, Biostatistics, & Computer Science Vanderbilt
O 3 O 4 O 5. q 3. q 4. Transition
 Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
CAP 5510 Lecture 3 Protein Structures
 CAP 5510 Lecture 3 Protein Structures Su-Shing Chen Bioinformatics CISE 8/19/2005 Su-Shing Chen, CISE 1 Protein Conformation 8/19/2005 Su-Shing Chen, CISE 2 Protein Conformational Structures Hydrophobicity
CAP 5510 Lecture 3 Protein Structures Su-Shing Chen Bioinformatics CISE 8/19/2005 Su-Shing Chen, CISE 1 Protein Conformation 8/19/2005 Su-Shing Chen, CISE 2 Protein Conformational Structures Hydrophobicity
Overview of IslandPick pipeline and the generation of GI datasets
 Overview of IslandPick pipeline and the generation of GI datasets Predicting GIs using comparative genomics By using whole genome alignments we can identify regions that are present in one genome but not
Overview of IslandPick pipeline and the generation of GI datasets Predicting GIs using comparative genomics By using whole genome alignments we can identify regions that are present in one genome but not
Detecting Distant Homologs Using Phylogenetic Tree-Based HMMs
 PROTEINS: Structure, Function, and Genetics 52:446 453 (2003) Detecting Distant Homologs Using Phylogenetic Tree-Based HMMs Bin Qian 1 and Richard A. Goldstein 1,2 * 1 Biophysics Research Division, University
PROTEINS: Structure, Function, and Genetics 52:446 453 (2003) Detecting Distant Homologs Using Phylogenetic Tree-Based HMMs Bin Qian 1 and Richard A. Goldstein 1,2 * 1 Biophysics Research Division, University
Syllabus of BIOINF 528 (2017 Fall, Bioinformatics Program)
 Syllabus of BIOINF 528 (2017 Fall, Bioinformatics Program) Course Name: Structural Bioinformatics Course Description: Instructor: This course introduces fundamental concepts and methods for structural
Syllabus of BIOINF 528 (2017 Fall, Bioinformatics Program) Course Name: Structural Bioinformatics Course Description: Instructor: This course introduces fundamental concepts and methods for structural
08/21/2017 BLAST. Multiple Sequence Alignments: Clustal Omega
 BLAST Multiple Sequence Alignments: Clustal Omega What does basic BLAST do (e.g. what is input sequence and how does BLAST look for matches?) Susan Parrish McDaniel College Multiple Sequence Alignments
BLAST Multiple Sequence Alignments: Clustal Omega What does basic BLAST do (e.g. what is input sequence and how does BLAST look for matches?) Susan Parrish McDaniel College Multiple Sequence Alignments
Bioinformatics. Proteins II. - Pattern, Profile, & Structure Database Searching. Robert Latek, Ph.D. Bioinformatics, Biocomputing
 Bioinformatics Proteins II. - Pattern, Profile, & Structure Database Searching Robert Latek, Ph.D. Bioinformatics, Biocomputing WIBR Bioinformatics Course, Whitehead Institute, 2002 1 Proteins I.-III.
Bioinformatics Proteins II. - Pattern, Profile, & Structure Database Searching Robert Latek, Ph.D. Bioinformatics, Biocomputing WIBR Bioinformatics Course, Whitehead Institute, 2002 1 Proteins I.-III.
Chapter 5. Proteomics and the analysis of protein sequence Ⅱ
 Proteomics Chapter 5. Proteomics and the analysis of protein sequence Ⅱ 1 Pairwise similarity searching (1) Figure 5.5: manual alignment One of the amino acids in the top sequence has no equivalent and
Proteomics Chapter 5. Proteomics and the analysis of protein sequence Ⅱ 1 Pairwise similarity searching (1) Figure 5.5: manual alignment One of the amino acids in the top sequence has no equivalent and
Smart Home Health Analytics Information Systems University of Maryland Baltimore County
 Smart Home Health Analytics Information Systems University of Maryland Baltimore County 1 IEEE Expert, October 1996 2 Given sample S from all possible examples D Learner L learns hypothesis h based on
Smart Home Health Analytics Information Systems University of Maryland Baltimore County 1 IEEE Expert, October 1996 2 Given sample S from all possible examples D Learner L learns hypothesis h based on
6.047 / Computational Biology: Genomes, Networks, Evolution Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, etworks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, etworks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Bioinformatics Chapter 1. Introduction
 Bioinformatics Chapter 1. Introduction Outline! Biological Data in Digital Symbol Sequences! Genomes Diversity, Size, and Structure! Proteins and Proteomes! On the Information Content of Biological Sequences!
Bioinformatics Chapter 1. Introduction Outline! Biological Data in Digital Symbol Sequences! Genomes Diversity, Size, and Structure! Proteins and Proteomes! On the Information Content of Biological Sequences!
SUPPLEMENTARY INFORMATION
 Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
Index of Balanced Accuracy: A Performance Measure for Skewed Class Distributions
 Index of Balanced Accuracy: A Performance Measure for Skewed Class Distributions V. García 1,2, R.A. Mollineda 2, and J.S. Sánchez 2 1 Lab. Reconocimiento de Patrones, Instituto Tecnológico de Toluca Av.
Index of Balanced Accuracy: A Performance Measure for Skewed Class Distributions V. García 1,2, R.A. Mollineda 2, and J.S. Sánchez 2 1 Lab. Reconocimiento de Patrones, Instituto Tecnológico de Toluca Av.
Linear Classifiers as Pattern Detectors
 Intelligent Systems: Reasoning and Recognition James L. Crowley ENSIMAG 2 / MoSIG M1 Second Semester 2014/2015 Lesson 16 8 April 2015 Contents Linear Classifiers as Pattern Detectors Notation...2 Linear
Intelligent Systems: Reasoning and Recognition James L. Crowley ENSIMAG 2 / MoSIG M1 Second Semester 2014/2015 Lesson 16 8 April 2015 Contents Linear Classifiers as Pattern Detectors Notation...2 Linear
Content. Learning. Regression vs Classification. Regression a.k.a. function approximation and Classification a.k.a. pattern recognition
 Content Andrew Kusiak Intelligent Systems Laboratory 239 Seamans Center The University of Iowa Iowa City, IA 52242-527 andrew-kusiak@uiowa.edu http://www.icaen.uiowa.edu/~ankusiak Introduction to learning
Content Andrew Kusiak Intelligent Systems Laboratory 239 Seamans Center The University of Iowa Iowa City, IA 52242-527 andrew-kusiak@uiowa.edu http://www.icaen.uiowa.edu/~ankusiak Introduction to learning
