Connecting NMR data to biomolecular structure and dynamics
|
|
|
- Miranda Copeland
- 6 years ago
- Views:
Transcription
1 Connecting NMR data to biomolecular structure and dynamics David A. Case Chem 538, Spring, 2014
2 Basics of NMR All nuclei are characterized by a spin quantum number I, which can be 0, 1/2, 1, 3/2, 2... Even atomic mass and number: I=0 ( 12 C, 16 O, 32 S) Even atomic mass, odd atomic number: I = integer ( 14 N, 2 H) Odd atomic mass: I=half integer ( 1 H, 15 N, 13 C, 31 P)
3 Basics of NMR./,0($1*02(0&3',!"# $"# %$&' B o =0 B o >0 $()*(+#,(-./01/(2/,34,567 $" %$8(.594:./(;,:16)+7 %$ z y x
4 Energy gaps are very small./,0($1*02(0&3', 4/(5$3'6'3$5/,$/$7088'*'2)$&+&93/)0+2$:!;<$/27$)5'$7088'*'2('$=')>''2$)5'$)>+$0,$*'3/)'7$ "! #$#% &"'( 4$8+*$ C D$/)$EFF$"D@$:.F$G$HIJ$K;$0,$LIM$N$CF OJ $P(/3QA+3 $$$$! "! #$CIFFFFRE $K5'$,9*&39,$&+&93/)0+2$0,$,A/33$:',&'(0/33-$>5'2$(+A&/*'7$)+$ST$+*$U#;I K5/)$*'27'*,$!"#$/$*/)5'*$02,'2,0)06'$)'(520V9'W
5 ./'$'0'()*+1234')5($,&'()*61 Comparison to other spectroscopies -rays X-rays ultraviolet visible infrared microwave radiofrequency Mossbauer electronic vibrational rotational NMR H 19 F 31 P 13 C /MHz 10 aldehydic 8 aromatic 6 olefinic 4 acetylenic 2 aliphatic 0 /ppm /Hz
6 Chemical shift dispersion!"#$%&'()*+,(+&-./'$0/'12(34$%/25) 1')/-4 &*+)+=, 312>'$&*+)+=, 3*+13)2($*2=? $&*+)+=, 1')/-4'=' &*+)+=, 678$"9:$ ; 9$,&'()*<1$+5$3$,1344$&*+)'2=$ shielding frequency magnetic field
7 Expansion to two (or more) dimensions Protein NMR 2D NOESY
8 Chemical shifts: Environmental effects for protons Nearby groups with magnetic susceptibilities ring currents peptide group contributions paramagnetic metal sites Electric fields bond polarizability models important in hydrogen bonds predicted secondary shift methyl shifts in proteins van der Waals (close contact) deshieldings hydrogen bonds general solvation effects observed secondary shift Ösapay & Case, JACS 113, 9436 (1991) local ( through-bond ) contributions Buckingham, Schaefer, Schneider, JCP 32: 1227, 1960
9 Chemical shielding and magnetic susceptibilities P H 0 y Heff Heff M j x
10 Spin transfer between nearby 2D NOESY spins
11 NMR 101 ( ) 2 E σ ij = µ i B j σ τ cs 2 r 6
12 Fragment B3 of protein G
13 N H correlation functions for GB3 effective decay time (6τ) 1 = e T D e But beware: statistical uncertainty in C(τ) is roughly ( ) 2τ 1/2 [1 C(τ)] T
14 Now consider intenal motions phe 30 ala 48 <P 2 [u(0).u(t)]> leu gly time, ns Hall & Fushman, JBNMR, 27, 261 (2003)
15 Converting distances to structures
16 Metric Matrix Distance Geometry To describe a molecule in terms of the distances between atoms,there are many constraints on the distances, since for N atoms there are N(N 1)/2distances but only 3N coordinates. General considerations for the conditions required to "embed" a set of interatomic distances into a realizable three-dimensional object forms the subject of distance geometry. The basic approach starts from the metric matrix that contains the scalar products of the vectors x i that give the positions of the atoms: g ij x i x j (1) These matrix elements can be expressed in terms of the distances d ij : g ij = 2(d 2 i0 + d 2 j0 d 2 ij ) (2) If the origin ("0") is chosen at the centroid of the atoms, then it can be shown that distances from this point can be computed from the interatomic distances alone. A fundamental theorem of distance geometry states that a set of distances can correspond to a three-dimensional object only if the metric matrix g is rank three, i.e. if it has three positive and N-3 zero eigenvalues. This may be made plausible by thinking of the eigenanalysis as a principal component analysis: all of the distance properties of the molecule should be describable in terms of three "components," which would be the x, y and z coordinates.
17 Metric Matrix Distance Geometry (part 2) If we denote the eigenvector matrix as w and the eigenvalues λ, the metric matrix can be written in two ways: 3 3 g ij = x ik x jk = w ik w jk λ k (3) k=1 k=1 The first equality follows from the definition of the metric tensor, Eq. (1); the upper limit of three in the second summation reflects the fact that a rank three matrix has only three non-zero eigenvalues. Eq. (3) then provides an expression for the coordinates x i in terms of the eigenvalues and eigenvectors of the metric matrix: x ik = λ 1/2 k w ik (4)
18 Using imprecise distances If the input distances are not exact, then in general the metric matrix will have more than three non-zero eigenvalues, but an approximate scheme can be made by using Eq. (4) with the three largest eigenvalues. Since information is lost by discarding the remaining eigenvectors, the resulting distances will not agree with the input distances, but will approximate them in a certain optimal fashion. If one only knows a distance range, then some choice of distance to be used must be made. Considerable attention has been paid recently to improving the performance of distance geometry by examining the ways in which the bounds are "smoothed" and by which distances are selected between the bounds. Triangle bound inequalities can improve consistency among the bounds, and NAB implements the "random pairwise metrization" algorithm developed by Jay Ponder. Methods like these are important especially for underconstrained problems, where a goal is to generate a reasonably random distribution of acceptable structures, and the difference between individual members of the ensemble may be quite large. An alternative procedure, which we call "random embedding", implements the procedure of degroot et al. for satisfying distance constraints. This does not use the embedding idea discussed above, but rather randomly corrects individual distances, ignoring all couplings between distances.
19 Molecular dynamics-based structure refinement
20 474 Example: cyclophilin bound to cyclosporin A A Spitzfaden, Braun, Wider, Widmer & Wüthrich, J. Biomol. NMR 4, 463
21 NMR 102 ( ) 2 E J ij = µ i µ j R ex = p A p B ( ω) 2 τ ex
22 Three-bond backbone couplings in proteins H Hα H Cβ H C 4 2 C C Case, Scheurer, Brüschweiler, JACS 122:10390, C Cβ C Hα
23 J-couplings across Watson-Crick hydrogen bonds h J(N N) h J(N N) or h J(H N), Hz solid: AT dashed: GC 1h J(H N) H...N distance, Angstroms
24 Getting dynamics from NMR NMR Spectroscopy Protein dynamics
25 Chemical exchange A k k B CH 3 CH 3 N N O k k CH 3 CH 3 N N O
26 Effects of chemical exchange on NMR lineshapes NMR Spectroscopy - Protein dynamics Chemical exchange
27 NMR Spectroscopy - Protein dynamics Hydrogen/deuterium Hydrogen/Deuterium exchange (H/D) exchange kop kch kcl kobs=kop kch/(kop + kcl + kch) EX2: kcl>>kch kobs=kop kch/(kcl) = Kop kch Kop is referred to as the protection factor, P!Gop = RTlnKop
28 Protection factors Hydrogen/Deuterium illustrate slow protein (H/D) exchange dynamics Significant heterogeneity in the magnitude of the protection factors A large number of states define the native-state ensemble
29 NMR Spectroscopy - Protein dynamics There is anotherhydrogen/deuterium kinetic regime (H/D) for H/D exchange exchange kop kch kcl kobs=kop kch/(kop + kcl + kch) EX1: kcl<<kch kobs=kop
30 Hydrogen/Deuterium (H/D) exchange Getting dynamics from NMR N-H (closed) k cl N-H (open) k ch D 2 O k op [k obs =k op ] [k obs =(k op /k cl ) k ch )] k ch (s -1 )
Using distance geomtry to generate structures
 Using distance geomtry to generate structures David A. Case Genomic systems and structures, Spring, 2009 Converting distances to structures Metric Matrix Distance Geometry To describe a molecule in terms
Using distance geomtry to generate structures David A. Case Genomic systems and structures, Spring, 2009 Converting distances to structures Metric Matrix Distance Geometry To describe a molecule in terms
HSQC spectra for three proteins
 HSQC spectra for three proteins SH3 domain from Abp1p Kinase domain from EphB2 apo Calmodulin What do the spectra tell you about the three proteins? HSQC spectra for three proteins Small protein Big protein
HSQC spectra for three proteins SH3 domain from Abp1p Kinase domain from EphB2 apo Calmodulin What do the spectra tell you about the three proteins? HSQC spectra for three proteins Small protein Big protein
Principles of Molecular Spectroscopy: Electromagnetic Radiation and Molecular structure. Nuclear Magnetic Resonance (NMR)
 Principles of Molecular Spectroscopy: Electromagnetic Radiation and Molecular structure Nuclear Magnetic Resonance (NMR) !E = h" Electromagnetic radiation is absorbed when the energy of photon corresponds
Principles of Molecular Spectroscopy: Electromagnetic Radiation and Molecular structure Nuclear Magnetic Resonance (NMR) !E = h" Electromagnetic radiation is absorbed when the energy of photon corresponds
Nuclear Magnetic Resonance H-NMR Part 1 Introduction to NMR, Instrumentation, Sample Prep, Chemical Shift. Dr. Sapna Gupta
 Nuclear Magnetic Resonance H-NMR Part 1 Introduction to NMR, Instrumentation, Sample Prep, Chemical Shift Dr. Sapna Gupta Introduction NMR is the most powerful tool available for organic structure determination.
Nuclear Magnetic Resonance H-NMR Part 1 Introduction to NMR, Instrumentation, Sample Prep, Chemical Shift Dr. Sapna Gupta Introduction NMR is the most powerful tool available for organic structure determination.
CS273: Algorithms for Structure Handout # 13 and Motion in Biology Stanford University Tuesday, 11 May 2003
 CS273: Algorithms for Structure Handout # 13 and Motion in Biology Stanford University Tuesday, 11 May 2003 Lecture #13: 11 May 2004 Topics: Protein Structure Determination Scribe: Minli Zhu We acknowledge
CS273: Algorithms for Structure Handout # 13 and Motion in Biology Stanford University Tuesday, 11 May 2003 Lecture #13: 11 May 2004 Topics: Protein Structure Determination Scribe: Minli Zhu We acknowledge
NMR parameters intensity chemical shift coupling constants 1D 1 H spectra of nucleic acids and proteins
 Lecture #2 M230 NMR parameters intensity chemical shift coupling constants Juli Feigon 1D 1 H spectra of nucleic acids and proteins NMR Parameters A. Intensity (area) 1D NMR spectrum: integrated intensity
Lecture #2 M230 NMR parameters intensity chemical shift coupling constants Juli Feigon 1D 1 H spectra of nucleic acids and proteins NMR Parameters A. Intensity (area) 1D NMR spectrum: integrated intensity
Origin of Scalar Couplings BCMB/CHEM 8190
 Origin of Scalar Couplings BCMB/CHEM 8190 Traditional View of Scalar Coupling Splitting of NMR signals due to through-bond interactions between nuclei is called scalar coupling (or J coupling or through-bond
Origin of Scalar Couplings BCMB/CHEM 8190 Traditional View of Scalar Coupling Splitting of NMR signals due to through-bond interactions between nuclei is called scalar coupling (or J coupling or through-bond
BMB/Bi/Ch 173 Winter 2018
 BMB/Bi/Ch 173 Winter 2018 Homework Set 8.1 (100 Points) Assigned 2-27-18, due 3-6-18 by 10:30 a.m. TA: Rachael Kuintzle. Office hours: SFL 220, Friday 3/2 4:00-5:00pm and SFL 229, Monday 3/5 4:00-5:30pm.
BMB/Bi/Ch 173 Winter 2018 Homework Set 8.1 (100 Points) Assigned 2-27-18, due 3-6-18 by 10:30 a.m. TA: Rachael Kuintzle. Office hours: SFL 220, Friday 3/2 4:00-5:00pm and SFL 229, Monday 3/5 4:00-5:30pm.
1. 3-hour Open book exam. No discussion among yourselves.
 Lecture 13 Review 1. 3-hour Open book exam. No discussion among yourselves. 2. Simple calculations. 3. Terminologies. 4. Decriptive questions. 5. Analyze a pulse program using density matrix approach (omonuclear
Lecture 13 Review 1. 3-hour Open book exam. No discussion among yourselves. 2. Simple calculations. 3. Terminologies. 4. Decriptive questions. 5. Analyze a pulse program using density matrix approach (omonuclear
6 NMR Interactions: Zeeman and CSA
 6 NMR Interactions: Zeeman and CSA 6.1 Zeeman Interaction Up to this point, we have mentioned a number of NMR interactions - Zeeman, quadrupolar, dipolar - but we have not looked at the nature of these
6 NMR Interactions: Zeeman and CSA 6.1 Zeeman Interaction Up to this point, we have mentioned a number of NMR interactions - Zeeman, quadrupolar, dipolar - but we have not looked at the nature of these
Química Orgânica I. Nuclear Magnetic Resonance Spectroscopy (I) Ciências Farmacêuticas Bioquímica Química AFB QO I 2007/08 1 AFB QO I 2007/08 2
 Química Orgânica I Ciências Farmacêuticas Bioquímica Química AFB QO I 2007/08 1 Nuclear Magnetic Resonance Spectroscopy (I) AFB QO I 2007/08 2 1 Adaptado de: Organic Chemistry, 6th Edition; L. G. Wade,
Química Orgânica I Ciências Farmacêuticas Bioquímica Química AFB QO I 2007/08 1 Nuclear Magnetic Resonance Spectroscopy (I) AFB QO I 2007/08 2 1 Adaptado de: Organic Chemistry, 6th Edition; L. G. Wade,
BMB/Bi/Ch 173 Winter 2018
 BMB/Bi/Ch 173 Winter 2018 Homework Set 8.1 (100 Points) Assigned 2-27-18, due 3-6-18 by 10:30 a.m. TA: Rachael Kuintzle. Office hours: SFL 220, Friday 3/2 4-5pm and SFL 229, Monday 3/5 4-5:30pm. 1. NMR
BMB/Bi/Ch 173 Winter 2018 Homework Set 8.1 (100 Points) Assigned 2-27-18, due 3-6-18 by 10:30 a.m. TA: Rachael Kuintzle. Office hours: SFL 220, Friday 3/2 4-5pm and SFL 229, Monday 3/5 4-5:30pm. 1. NMR
Introduction solution NMR
 2 NMR journey Introduction solution NMR Alexandre Bonvin Bijvoet Center for Biomolecular Research with thanks to Dr. Klaartje Houben EMBO Global Exchange course, IHEP, Beijing April 28 - May 5, 20 3 Topics
2 NMR journey Introduction solution NMR Alexandre Bonvin Bijvoet Center for Biomolecular Research with thanks to Dr. Klaartje Houben EMBO Global Exchange course, IHEP, Beijing April 28 - May 5, 20 3 Topics
Nuclear Magnetic Resonance Spectroscopy: Tools for Structure Determination
 Nuclear Magnetic Resonance Spectroscopy: Tools for Structure Determination Chung-Ming Sun Department of Applied Chemistry National Chiao Tung University Hualien 300, Taiwan Introduction NMR (Nuclear Magnetic
Nuclear Magnetic Resonance Spectroscopy: Tools for Structure Determination Chung-Ming Sun Department of Applied Chemistry National Chiao Tung University Hualien 300, Taiwan Introduction NMR (Nuclear Magnetic
NMRis the most valuable spectroscopic technique for organic chemists because it maps the carbon-hydrogen framework of a molecule.
 Chapter 13: Nuclear magnetic resonance spectroscopy NMRis the most valuable spectroscopic technique for organic chemists because it maps the carbon-hydrogen framework of a molecule. 13.2 The nature of
Chapter 13: Nuclear magnetic resonance spectroscopy NMRis the most valuable spectroscopic technique for organic chemists because it maps the carbon-hydrogen framework of a molecule. 13.2 The nature of
NUCLEAR MAGNETIC RESONANCE AND INTRODUCTION TO MASS SPECTROMETRY
 NUCLEAR MAGNETIC RESONANCE AND INTRODUCTION TO MASS SPECTROMETRY A STUDENT SHOULD BE ABLE TO: 1. Identify and explain the processes involved in proton ( 1 H) and carbon-13 ( 13 C) nuclear magnetic resonance
NUCLEAR MAGNETIC RESONANCE AND INTRODUCTION TO MASS SPECTROMETRY A STUDENT SHOULD BE ABLE TO: 1. Identify and explain the processes involved in proton ( 1 H) and carbon-13 ( 13 C) nuclear magnetic resonance
NMR spectra of some simple molecules. Effect of spinning: averaging field inhomogeneity (nmr1.pdf pg 2)
 NMR spectra of some simple molecules Effect of spinning: averaging field inhomogeneity (nmr1.pdf pg 2) N S H 0 H o Because the protons have a magnetic field associated with them, the field changes as across
NMR spectra of some simple molecules Effect of spinning: averaging field inhomogeneity (nmr1.pdf pg 2) N S H 0 H o Because the protons have a magnetic field associated with them, the field changes as across
Chapter 13 Spectroscopy
 hapter 13 Spectroscopy Infrared spectroscopy Ultraviolet-Visible spectroscopy Nuclear magnetic resonance spectroscopy Mass Spectrometry 13.1 Principles of Molecular Spectroscopy: Electromagnetic Radiation
hapter 13 Spectroscopy Infrared spectroscopy Ultraviolet-Visible spectroscopy Nuclear magnetic resonance spectroscopy Mass Spectrometry 13.1 Principles of Molecular Spectroscopy: Electromagnetic Radiation
Lecture 2 nmr Spectroscopy
 Lecture 2 nmr Spectroscopy Pages 427 430 and Chapter 13 Molecular Spectroscopy Molecular spectroscopy: the study of the frequencies of electromagnetic radiation that are absorbed or emitted by substances
Lecture 2 nmr Spectroscopy Pages 427 430 and Chapter 13 Molecular Spectroscopy Molecular spectroscopy: the study of the frequencies of electromagnetic radiation that are absorbed or emitted by substances
Molecular Mechanics. I. Quantum mechanical treatment of molecular systems
 Molecular Mechanics I. Quantum mechanical treatment of molecular systems The first principle approach for describing the properties of molecules, including proteins, involves quantum mechanics. For example,
Molecular Mechanics I. Quantum mechanical treatment of molecular systems The first principle approach for describing the properties of molecules, including proteins, involves quantum mechanics. For example,
Experiment 11: NUCLEAR MAGNETIC RESONANCE SPECTROSCOPY
 Experiment 11: NUCLEAR MAGNETIC RESONANCE SPECTROSCOPY Purpose: This is an exercise to introduce the use of nuclear magnetic resonance spectroscopy, in conjunction with infrared spectroscopy, to determine
Experiment 11: NUCLEAR MAGNETIC RESONANCE SPECTROSCOPY Purpose: This is an exercise to introduce the use of nuclear magnetic resonance spectroscopy, in conjunction with infrared spectroscopy, to determine
Chapter 13 Nuclear Magnetic Resonance Spectroscopy
 Organic Chemistry, 6 th Edition L. G. Wade, Jr. Chapter 13 Nuclear Magnetic Resonance Spectroscopy Jo Blackburn Richland College, Dallas, TX Dallas County Community College District 2006, Prentice Hall
Organic Chemistry, 6 th Edition L. G. Wade, Jr. Chapter 13 Nuclear Magnetic Resonance Spectroscopy Jo Blackburn Richland College, Dallas, TX Dallas County Community College District 2006, Prentice Hall
Inorganic Spectroscopic and Structural Methods
 Inorganic Spectroscopic and Structural Methods Electromagnetic spectrum has enormous range of energies. Wide variety of techniques based on absorption of energy e.g. ESR and NMR: radiowaves (MHz) IR vibrations
Inorganic Spectroscopic and Structural Methods Electromagnetic spectrum has enormous range of energies. Wide variety of techniques based on absorption of energy e.g. ESR and NMR: radiowaves (MHz) IR vibrations
Biochemistry 530 NMR Theory and Practice
 Biochemistry 530 NMR Theory and Practice Gabriele Varani Department of Biochemistry and Department of Chemistry University of Washington 1D spectra contain structural information.. but is hard to extract:
Biochemistry 530 NMR Theory and Practice Gabriele Varani Department of Biochemistry and Department of Chemistry University of Washington 1D spectra contain structural information.. but is hard to extract:
NMR journey. Introduction to solution NMR. Alexandre Bonvin. Topics. Why use NMR...? Bijvoet Center for Biomolecular Research
 2 NMR journey Introduction to solution NMR Alexandre Bonvin Bijvoet Center for Biomolecular Research with thanks to Dr. Klaartje Houben EMBO Global Exchange course, CCMB, Hyderabad, India November 29th
2 NMR journey Introduction to solution NMR Alexandre Bonvin Bijvoet Center for Biomolecular Research with thanks to Dr. Klaartje Houben EMBO Global Exchange course, CCMB, Hyderabad, India November 29th
William H. Brown & Christopher S. Foote
 Requests for permission to make copies of any part of the work should be mailed to:permissions Department, Harcourt Brace & Company, 6277 Sea Harbor Drive, Orlando, Florida 32887-6777 William H. Brown
Requests for permission to make copies of any part of the work should be mailed to:permissions Department, Harcourt Brace & Company, 6277 Sea Harbor Drive, Orlando, Florida 32887-6777 William H. Brown
Chapter 9. Nuclear Magnetic Resonance. Ch. 9-1
 Chapter 9 Nuclear Magnetic Resonance Ch. 9-1 1. Introduction Classic methods for organic structure determination Boiling point Refractive index Solubility tests Functional group tests Derivative preparation
Chapter 9 Nuclear Magnetic Resonance Ch. 9-1 1. Introduction Classic methods for organic structure determination Boiling point Refractive index Solubility tests Functional group tests Derivative preparation
7a. Structure Elucidation: IR and 13 C-NMR Spectroscopies (text , , 12.10)
 2009, Department of Chemistry, The University of Western Ontario 7a.1 7a. Structure Elucidation: IR and 13 C-NMR Spectroscopies (text 11.1 11.5, 12.1 12.5, 12.10) A. Electromagnetic Radiation Energy is
2009, Department of Chemistry, The University of Western Ontario 7a.1 7a. Structure Elucidation: IR and 13 C-NMR Spectroscopies (text 11.1 11.5, 12.1 12.5, 12.10) A. Electromagnetic Radiation Energy is
1. neopentyl benzene. 4 of 6
 I. 1 H NMR spectroscopy A. Theory 1. The protons and neutrons in atomic nuclei spin, as does the nucleus itself 2. The circulation of nuclear charge can generate a nuclear magnetic moment, u, along the
I. 1 H NMR spectroscopy A. Theory 1. The protons and neutrons in atomic nuclei spin, as does the nucleus itself 2. The circulation of nuclear charge can generate a nuclear magnetic moment, u, along the
Magnetic Nuclei other than 1 H
 Magnetic Nuclei other than 1 H 2 H (Deuterium): I = 1 H,D-Exchange might be used to simplify 1 H-NMR spectra since H-D couplings are generally small; - - - -O- - - -D 2 -O- triplet of triplets slightly
Magnetic Nuclei other than 1 H 2 H (Deuterium): I = 1 H,D-Exchange might be used to simplify 1 H-NMR spectra since H-D couplings are generally small; - - - -O- - - -D 2 -O- triplet of triplets slightly
Chapter 13: Nuclear Magnetic Resonance (NMR) Spectroscopy direct observation of the H s and C s of a molecules
 hapter 13: Nuclear Magnetic Resonance (NMR) Spectroscopy direct observation of the s and s of a molecules Nuclei are positively charged and spin on an axis; they create a tiny magnetic field + + Not all
hapter 13: Nuclear Magnetic Resonance (NMR) Spectroscopy direct observation of the s and s of a molecules Nuclei are positively charged and spin on an axis; they create a tiny magnetic field + + Not all
Principles of NMR Protein Spectroscopy. 2) Assignment of chemical shifts in a protein ( 1 H, 13 C, 15 N) 3) Three dimensional structure determination
 1) Protein preparation (>50 aa) 2) Assignment of chemical shifts in a protein ( 1 H, 13 C, 15 N) 3) Three dimensional structure determination Protein Expression overexpression in E. coli - BL21(DE3) 1
1) Protein preparation (>50 aa) 2) Assignment of chemical shifts in a protein ( 1 H, 13 C, 15 N) 3) Three dimensional structure determination Protein Expression overexpression in E. coli - BL21(DE3) 1
(Refer Slide Time: 1:03)
 Principles and Applications of NMR spectroscopy Professor Hanudatta S. Atreya NMR Research Centre Indian Institute of Science Bangalore Module 1 Lecture No 05 Welcome back! In the last class we looked
Principles and Applications of NMR spectroscopy Professor Hanudatta S. Atreya NMR Research Centre Indian Institute of Science Bangalore Module 1 Lecture No 05 Welcome back! In the last class we looked
Introduction to Relaxation Theory James Keeler
 EUROMAR Zürich, 24 Introduction to Relaxation Theory James Keeler University of Cambridge Department of Chemistry What is relaxation? Why might it be interesting? relaxation is the process which drives
EUROMAR Zürich, 24 Introduction to Relaxation Theory James Keeler University of Cambridge Department of Chemistry What is relaxation? Why might it be interesting? relaxation is the process which drives
4) protons experience a net magnetic field strength that is smaller than the applied magnetic field.
 1) Which of the following CANNOT be probed by an NMR spectrometer? See sect 15.1 Chapter 15: 1 A) nucleus with odd number of protons & odd number of neutrons B) nucleus with odd number of protons &even
1) Which of the following CANNOT be probed by an NMR spectrometer? See sect 15.1 Chapter 15: 1 A) nucleus with odd number of protons & odd number of neutrons B) nucleus with odd number of protons &even
Nuclear Magnetic Resonance
 Nuclear Magnetic Resonance PRINCIPLES OF NMR SPECTROSCOPY Contents Principles of nuclear magnetic resonance The nmr spectrometer Basic principles in nmr application NMR tools used to obtain information
Nuclear Magnetic Resonance PRINCIPLES OF NMR SPECTROSCOPY Contents Principles of nuclear magnetic resonance The nmr spectrometer Basic principles in nmr application NMR tools used to obtain information
Infrared spectroscopy Basic theory
 Infrared spectroscopy Basic theory Dr. Davide Ferri Paul Scherrer Institut 056 310 27 81 davide.ferri@psi.ch Importance of IR spectroscopy in catalysis IR Raman NMR XAFS UV-Vis EPR 0 200 400 600 800 1000
Infrared spectroscopy Basic theory Dr. Davide Ferri Paul Scherrer Institut 056 310 27 81 davide.ferri@psi.ch Importance of IR spectroscopy in catalysis IR Raman NMR XAFS UV-Vis EPR 0 200 400 600 800 1000
Module 13: Chemical Shift and Its Measurement
 Subject Chemistry Paper No and Title Module No and Title Module Tag Paper 12: Organic Spectroscopy CHE_P12_M13_e-Text TABLE OF CONTENTS 1. Learning Outcomes 2. Introduction 3. Shielding and deshielding
Subject Chemistry Paper No and Title Module No and Title Module Tag Paper 12: Organic Spectroscopy CHE_P12_M13_e-Text TABLE OF CONTENTS 1. Learning Outcomes 2. Introduction 3. Shielding and deshielding
Nuclear Magnetic Resonance Spectroscopy
 Nuclear Magnetic Resonance Spectroscopy Features: Used to identify products of reactions Also gives information about chemical environment, connectivity and bonding of nuclei Requirements: Pure or mostly
Nuclear Magnetic Resonance Spectroscopy Features: Used to identify products of reactions Also gives information about chemical environment, connectivity and bonding of nuclei Requirements: Pure or mostly
( ) x10 8 m. The energy in a mole of 400 nm photons is calculated by: ' & sec( ) ( & % ) 6.022x10 23 photons' E = h! = hc & 6.
 Introduction to Spectroscopy Spectroscopic techniques are widely used to detect molecules, to measure the concentration of a species in solution, and to determine molecular structure. For proteins, most
Introduction to Spectroscopy Spectroscopic techniques are widely used to detect molecules, to measure the concentration of a species in solution, and to determine molecular structure. For proteins, most
Objective 4. Determine (characterize) the structure of a compound using IR, NMR, MS.
 Objective 4. Determine (characterize) the structure of a compound using IR, NMR, MS. Skills: Draw structure IR: match bond type to IR peak NMR: ID number of non-equivalent H s, relate peak splitting to
Objective 4. Determine (characterize) the structure of a compound using IR, NMR, MS. Skills: Draw structure IR: match bond type to IR peak NMR: ID number of non-equivalent H s, relate peak splitting to
Basic principles of multidimensional NMR in solution
 Basic principles of multidimensional NMR in solution 19.03.2008 The program 2/93 General aspects Basic principles Parameters in NMR spectroscopy Multidimensional NMR-spectroscopy Protein structures NMR-spectra
Basic principles of multidimensional NMR in solution 19.03.2008 The program 2/93 General aspects Basic principles Parameters in NMR spectroscopy Multidimensional NMR-spectroscopy Protein structures NMR-spectra
Biophysical Chemistry: NMR Spectroscopy
 Relaxation & Multidimensional Spectrocopy Vrije Universiteit Brussel 9th December 2011 Outline 1 Relaxation 2 Principles 3 Outline 1 Relaxation 2 Principles 3 Establishment of Thermal Equilibrium As previously
Relaxation & Multidimensional Spectrocopy Vrije Universiteit Brussel 9th December 2011 Outline 1 Relaxation 2 Principles 3 Outline 1 Relaxation 2 Principles 3 Establishment of Thermal Equilibrium As previously
4) protons experience a net magnetic field strength that is smaller than the applied magnetic field.
 1) Which of the following CANNOT be probed by an spectrometer? See sect 15.1 Chapter 15: 1 A) nucleus with odd number of protons & odd number of neutrons B) nucleus with odd number of protons &even number
1) Which of the following CANNOT be probed by an spectrometer? See sect 15.1 Chapter 15: 1 A) nucleus with odd number of protons & odd number of neutrons B) nucleus with odd number of protons &even number
I690/B680 Structural Bioinformatics Spring Protein Structure Determination by NMR Spectroscopy
 I690/B680 Structural Bioinformatics Spring 2006 Protein Structure Determination by NMR Spectroscopy Suggested Reading (1) Van Holde, Johnson, Ho. Principles of Physical Biochemistry, 2 nd Ed., Prentice
I690/B680 Structural Bioinformatics Spring 2006 Protein Structure Determination by NMR Spectroscopy Suggested Reading (1) Van Holde, Johnson, Ho. Principles of Physical Biochemistry, 2 nd Ed., Prentice
Magnetic Resonance Lectures for Chem 341 James Aramini, PhD. CABM 014A
 Magnetic Resonance Lectures for Chem 341 James Aramini, PhD. CABM 014A jma@cabm.rutgers.edu " J.A. 12/11/13 Dec. 4 Dec. 9 Dec. 11" " Outline" " 1. Introduction / Spectroscopy Overview 2. NMR Spectroscopy
Magnetic Resonance Lectures for Chem 341 James Aramini, PhD. CABM 014A jma@cabm.rutgers.edu " J.A. 12/11/13 Dec. 4 Dec. 9 Dec. 11" " Outline" " 1. Introduction / Spectroscopy Overview 2. NMR Spectroscopy
NMR = Nuclear Magnetic Resonance
 NMR = Nuclear Magnetic Resonance NMR spectroscopy is the most powerful technique available to organic chemists for determining molecular structures. Looks at nuclei with odd mass numbers or odd number
NMR = Nuclear Magnetic Resonance NMR spectroscopy is the most powerful technique available to organic chemists for determining molecular structures. Looks at nuclei with odd mass numbers or odd number
Slow symmetric exchange
 Slow symmetric exchange ϕ A k k B t A B There are three things you should notice compared with the Figure on the previous slide: 1) The lines are broader, 2) the intensities are reduced and 3) the peaks
Slow symmetric exchange ϕ A k k B t A B There are three things you should notice compared with the Figure on the previous slide: 1) The lines are broader, 2) the intensities are reduced and 3) the peaks
CHEM 241 UNIT 5: PART A DETERMINATION OF ORGANIC STRUCTURES BY SPECTROSCOPIC METHODS [MASS SPECTROMETRY]
![CHEM 241 UNIT 5: PART A DETERMINATION OF ORGANIC STRUCTURES BY SPECTROSCOPIC METHODS [MASS SPECTROMETRY] CHEM 241 UNIT 5: PART A DETERMINATION OF ORGANIC STRUCTURES BY SPECTROSCOPIC METHODS [MASS SPECTROMETRY]](/thumbs/83/88348834.jpg) CHEM 241 UNIT 5: PART A DETERMINATION OF ORGANIC STRUCTURES BY SPECTROSCOPIC METHODS [MASS SPECTROMETRY] 1 Introduction Outline Mass spectrometry (MS) 2 INTRODUCTION The analysis of the outcome of a reaction
CHEM 241 UNIT 5: PART A DETERMINATION OF ORGANIC STRUCTURES BY SPECTROSCOPIC METHODS [MASS SPECTROMETRY] 1 Introduction Outline Mass spectrometry (MS) 2 INTRODUCTION The analysis of the outcome of a reaction
Your Name: Question 1. 2D-NMR: C 6 H 10 O 2. (20 points)
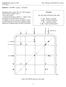 Question 1. 2D-NMR: C 6 H 10 O 2. (20 points) Integrations show signals 3H 1 & 5, 2H for signal 4, and 1H each for signals 2 and 3. - Draw the structure. - Assign the hydrogens to signals 1 5 (that is,
Question 1. 2D-NMR: C 6 H 10 O 2. (20 points) Integrations show signals 3H 1 & 5, 2H for signal 4, and 1H each for signals 2 and 3. - Draw the structure. - Assign the hydrogens to signals 1 5 (that is,
Infrared Spectroscopy
 Infrared Spectroscopy Introduction Spectroscopy is an analytical technique which helps determine structure. It destroys little or no sample. The amount of light absorbed by the sample is measured as wavelength
Infrared Spectroscopy Introduction Spectroscopy is an analytical technique which helps determine structure. It destroys little or no sample. The amount of light absorbed by the sample is measured as wavelength
Chapter 15 Lecture Outline
 Organic Chemistry, First Edition Janice Gorzynski Smith University of Hawaii Chapter 5 Lecture Outline Introduction to NMR Two common types of NMR spectroscopy are used to characterize organic structure:
Organic Chemistry, First Edition Janice Gorzynski Smith University of Hawaii Chapter 5 Lecture Outline Introduction to NMR Two common types of NMR spectroscopy are used to characterize organic structure:
Quantification of Dynamics in the Solid-State
 Bernd Reif Quantification of Dynamics in the Solid-State Technische Universität München Helmholtz-Zentrum München Biomolecular Solid-State NMR Winter School Stowe, VT January 0-5, 206 Motivation. Solid
Bernd Reif Quantification of Dynamics in the Solid-State Technische Universität München Helmholtz-Zentrum München Biomolecular Solid-State NMR Winter School Stowe, VT January 0-5, 206 Motivation. Solid
Chapter 13 Structure t Determination: Nuclear Magnetic Resonance Spectroscopy
 John E. McMurry www.cengage.com/chemistry/mcmurry Chapter 13 Structure t Determination: ti Nuclear Magnetic Resonance Spectroscopy Revisions by Dr. Daniel Holmes MSU Paul D. Adams University of Arkansas
John E. McMurry www.cengage.com/chemistry/mcmurry Chapter 13 Structure t Determination: ti Nuclear Magnetic Resonance Spectroscopy Revisions by Dr. Daniel Holmes MSU Paul D. Adams University of Arkansas
Spectroscopy. Empirical Formula: Chemical Formula: Index of Hydrogen Deficiency (IHD)
 Spectroscopy Empirical Formula: Chemical Formula: Index of Hydrogen Deficiency (IHD) A)From a structure: B)From a molecular formula, C c H h N n O o X x, Formula for saturated hydrocarbons: Subtract the
Spectroscopy Empirical Formula: Chemical Formula: Index of Hydrogen Deficiency (IHD) A)From a structure: B)From a molecular formula, C c H h N n O o X x, Formula for saturated hydrocarbons: Subtract the
NMR-spectroscopy of proteins in solution. Peter Schmieder
 NMR-spectroscopy of proteins in solution Basic aspects of NMR-Spektroskopie Basic aspects of NMR-spectroscopy 3/84 Prerequisite for NMR-spectroscopy is a nuclear spin that can be thought of as a mixture
NMR-spectroscopy of proteins in solution Basic aspects of NMR-Spektroskopie Basic aspects of NMR-spectroscopy 3/84 Prerequisite for NMR-spectroscopy is a nuclear spin that can be thought of as a mixture
OAT Organic Chemistry - Problem Drill 19: NMR Spectroscopy and Mass Spectrometry
 OAT Organic Chemistry - Problem Drill 19: NMR Spectroscopy and Mass Spectrometry Question No. 1 of 10 Question 1. Which statement concerning NMR spectroscopy is incorrect? Question #01 (A) Only nuclei
OAT Organic Chemistry - Problem Drill 19: NMR Spectroscopy and Mass Spectrometry Question No. 1 of 10 Question 1. Which statement concerning NMR spectroscopy is incorrect? Question #01 (A) Only nuclei
Sequential resonance assignments in (small) proteins: homonuclear method 2º structure determination
 Lecture 9 M230 Feigon Sequential resonance assignments in (small) proteins: homonuclear method 2º structure determination Reading resources v Roberts NMR of Macromolecules, Chap 4 by Christina Redfield
Lecture 9 M230 Feigon Sequential resonance assignments in (small) proteins: homonuclear method 2º structure determination Reading resources v Roberts NMR of Macromolecules, Chap 4 by Christina Redfield
Chapter 6 Part 1 Structure of the atom
 Chapter 6 Part 1 Structure of the atom What IS the structure of an atom? What are the properties of atoms? REMEMBER: structure affects function! Important questions: Where are the electrons? What is the
Chapter 6 Part 1 Structure of the atom What IS the structure of an atom? What are the properties of atoms? REMEMBER: structure affects function! Important questions: Where are the electrons? What is the
CHEM311 FALL 2005 Practice Exam #3
 CHEM311 FALL 2005 Practice Exam #3 Instructions: This is a multiple choice / short answer practice exam. For the multiple-choice questions, there may be more than one correct answer. If so, then circle
CHEM311 FALL 2005 Practice Exam #3 Instructions: This is a multiple choice / short answer practice exam. For the multiple-choice questions, there may be more than one correct answer. If so, then circle
Timescales of Protein Dynamics
 Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Summary of 1D Experiment time domain data Fourier Transform (FT) frequency domain data or Transverse Relaxation Ensemble of Nuclear
Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Summary of 1D Experiment time domain data Fourier Transform (FT) frequency domain data or Transverse Relaxation Ensemble of Nuclear
Chapter 12 Mass Spectrometry and Infrared Spectroscopy
 Organic Chemistry, 6 th Edition L. G. Wade, Jr. Chapter 12 Mass Spectrometry and Infrared Spectroscopy Jo Blackburn Richland College, Dallas, TX Dallas County Community College District 2006, Prentice
Organic Chemistry, 6 th Edition L. G. Wade, Jr. Chapter 12 Mass Spectrometry and Infrared Spectroscopy Jo Blackburn Richland College, Dallas, TX Dallas County Community College District 2006, Prentice
Biochemistry 530 NMR Theory and Practice
 Biochemistry 530 NMR Theory and Practice David Baker Autumn Quarter 2014 Slides Courtesy of Gabriele Varani Recommended NMR Textbooks Derome, A. E. (1987) Modern NMR Techniques for Chemistry Research,
Biochemistry 530 NMR Theory and Practice David Baker Autumn Quarter 2014 Slides Courtesy of Gabriele Varani Recommended NMR Textbooks Derome, A. E. (1987) Modern NMR Techniques for Chemistry Research,
Unit 11 Instrumentation. Mass, Infrared and NMR Spectroscopy
 Unit 11 Instrumentation Mass, Infrared and NMR Spectroscopy Spectroscopic identification of organic compounds Qualitative analysis: presence but not quantity (i.e. PEDs) Quantitative analysis: quantity
Unit 11 Instrumentation Mass, Infrared and NMR Spectroscopy Spectroscopic identification of organic compounds Qualitative analysis: presence but not quantity (i.e. PEDs) Quantitative analysis: quantity
C NMR Spectroscopy
 13.14 13 C NMR Spectroscopy 1 H and 13 C NMR compared: both give us information about the number of chemically nonequivalent nuclei (nonequivalent hydrogens or nonequivalent carbons) both give us information
13.14 13 C NMR Spectroscopy 1 H and 13 C NMR compared: both give us information about the number of chemically nonequivalent nuclei (nonequivalent hydrogens or nonequivalent carbons) both give us information
Nuclear Magnetic Resonance (NMR)
 Nuclear Magnetic Resonance (NMR) E E increases with increasing magnetic field strength Boltzmann distribution at thermal equilibrium: N (m=-1/2) /N (m=+1/2) = e ( E/kT) with E = γ(h/2π)b o NMR Physical
Nuclear Magnetic Resonance (NMR) E E increases with increasing magnetic field strength Boltzmann distribution at thermal equilibrium: N (m=-1/2) /N (m=+1/2) = e ( E/kT) with E = γ(h/2π)b o NMR Physical
Spectroscopy in Organic Chemistry. Types of Spectroscopy in Organic
 Spectroscopy in Organic Chemistry Spectroscopy Spectrum dealing with light, or more specifically, radiation Scope to see Organic Spectroscopy therefore deals with examining how organic molecules interact
Spectroscopy in Organic Chemistry Spectroscopy Spectrum dealing with light, or more specifically, radiation Scope to see Organic Spectroscopy therefore deals with examining how organic molecules interact
Timescales of Protein Dynamics
 Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Dynamics from NMR Show spies Amide Nitrogen Spies Report On Conformational Dynamics Amide Hydrogen Transverse Relaxation Ensemble
Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Dynamics from NMR Show spies Amide Nitrogen Spies Report On Conformational Dynamics Amide Hydrogen Transverse Relaxation Ensemble
Molecular mechanics. classical description of molecules. Marcus Elstner and Tomáš Kubař. April 29, 2016
 classical description of molecules April 29, 2016 Chemical bond Conceptual and chemical basis quantum effect solution of the SR numerically expensive (only small molecules can be treated) approximations
classical description of molecules April 29, 2016 Chemical bond Conceptual and chemical basis quantum effect solution of the SR numerically expensive (only small molecules can be treated) approximations
9. Nuclear Magnetic Resonance
 9. Nuclear Magnetic Resonance Nuclear Magnetic Resonance (NMR) is a method that can be used to find structures of proteins. NMR spectroscopy is the observation of spins of atoms and electrons in a molecule
9. Nuclear Magnetic Resonance Nuclear Magnetic Resonance (NMR) is a method that can be used to find structures of proteins. NMR spectroscopy is the observation of spins of atoms and electrons in a molecule
MOLECULAR SPECTROSCOPY AND PHOTOCHEMISTRY
 20 CHAPTER MOLECULAR SPECTROSCOPY AND PHOTOCHEMISTRY 20.1 Introduction to Molecular Spectroscopy 20.2 Experimental Methods in Molecular Spectroscopy 20.3 Rotational and Vibrational Spectroscopy 20.4 Nuclear
20 CHAPTER MOLECULAR SPECTROSCOPY AND PHOTOCHEMISTRY 20.1 Introduction to Molecular Spectroscopy 20.2 Experimental Methods in Molecular Spectroscopy 20.3 Rotational and Vibrational Spectroscopy 20.4 Nuclear
Protein Structure Determination Using NMR Restraints BCMB/CHEM 8190
 Protein Structure Determination Using NMR Restraints BCMB/CHEM 8190 Programs for NMR Based Structure Determination CNS - Brünger, A. T.; Adams, P. D.; Clore, G. M.; DeLano, W. L.; Gros, P.; Grosse-Kunstleve,
Protein Structure Determination Using NMR Restraints BCMB/CHEM 8190 Programs for NMR Based Structure Determination CNS - Brünger, A. T.; Adams, P. D.; Clore, G. M.; DeLano, W. L.; Gros, P.; Grosse-Kunstleve,
H B. θ = 90 o. Lecture notes Part 4: Spin-Spin Coupling. θ θ
 Lecture notes Part 4: Spin-Spin Coupling F. olger Försterling October 4, 2011 So far, spins were regarded spins isolated from each other. owever, the magnetic moment of nuclear spins also have effect on
Lecture notes Part 4: Spin-Spin Coupling F. olger Försterling October 4, 2011 So far, spins were regarded spins isolated from each other. owever, the magnetic moment of nuclear spins also have effect on
NMR in Medicine and Biology
 NMR in Medicine and Biology http://en.wikipedia.org/wiki/nmr_spectroscopy MRI- Magnetic Resonance Imaging (water) In-vivo spectroscopy (metabolites) Solid-state t NMR (large structures) t Solution NMR
NMR in Medicine and Biology http://en.wikipedia.org/wiki/nmr_spectroscopy MRI- Magnetic Resonance Imaging (water) In-vivo spectroscopy (metabolites) Solid-state t NMR (large structures) t Solution NMR
4) protons experience a net magnetic field strength that is smaller than the applied magnetic field.
 1) Which of the following CANNOT be probed by an spectrometer? See sect 16.1 Chapter 16: 1 A) nucleus with odd number of protons & odd number of neutrons B) nucleus with odd number of protons &even number
1) Which of the following CANNOT be probed by an spectrometer? See sect 16.1 Chapter 16: 1 A) nucleus with odd number of protons & odd number of neutrons B) nucleus with odd number of protons &even number
Ch 7 Quantum Theory of the Atom (light and atomic structure)
 Ch 7 Quantum Theory of the Atom (light and atomic structure) Electromagnetic Radiation - Electromagnetic radiation consists of oscillations in electric and magnetic fields. The oscillations can be described
Ch 7 Quantum Theory of the Atom (light and atomic structure) Electromagnetic Radiation - Electromagnetic radiation consists of oscillations in electric and magnetic fields. The oscillations can be described
16.1 Introduction to NMR Spectroscopy. Spectroscopy. Spectroscopy. Spectroscopy. Spectroscopy. Spectroscopy 4/11/2013
 What is spectroscopy? NUCLEAR MAGNETIC RESONANCE (NMR) spectroscopy may be the most powerful method of gaining structural information about organic compounds. NMR involves an interaction between electromagnetic
What is spectroscopy? NUCLEAR MAGNETIC RESONANCE (NMR) spectroscopy may be the most powerful method of gaining structural information about organic compounds. NMR involves an interaction between electromagnetic
Longitudinal-relaxation enhanced fast-pulsing techniques: New tools for biomolecular NMR spectroscopy
 Longitudinal-relaxation enhanced fast-pulsing techniques: New tools for biomolecular NMR spectroscopy Bernhard Brutscher Laboratoire de Résonance Magnétique Nucléaire Institut de Biologie Structurale -
Longitudinal-relaxation enhanced fast-pulsing techniques: New tools for biomolecular NMR spectroscopy Bernhard Brutscher Laboratoire de Résonance Magnétique Nucléaire Institut de Biologie Structurale -
Labelling strategies in the NMR structure determination of larger proteins
 Labelling strategies in the NMR structure determination of larger proteins - Difficulties of studying larger proteins - The effect of deuteration on spectral complexity and relaxation rates - NMR expts
Labelling strategies in the NMR structure determination of larger proteins - Difficulties of studying larger proteins - The effect of deuteration on spectral complexity and relaxation rates - NMR expts
Rotations and vibrations of polyatomic molecules
 Rotations and vibrations of polyatomic molecules When the potential energy surface V( R 1, R 2,..., R N ) is known we can compute the energy levels of the molecule. These levels can be an effect of: Rotation
Rotations and vibrations of polyatomic molecules When the potential energy surface V( R 1, R 2,..., R N ) is known we can compute the energy levels of the molecule. These levels can be an effect of: Rotation
Lecture 6: Physical Methods II. UV Vis (electronic spectroscopy) Electron Spin Resonance Mossbauer Spectroscopy
 Lecture 6: Physical Methods II UV Vis (electronic spectroscopy) Electron Spin Resonance Mossbauer Spectroscopy Physical Methods used in bioinorganic chemistry X ray crystallography X ray absorption (XAS)
Lecture 6: Physical Methods II UV Vis (electronic spectroscopy) Electron Spin Resonance Mossbauer Spectroscopy Physical Methods used in bioinorganic chemistry X ray crystallography X ray absorption (XAS)
3D NMR 3D NMR 3D NMR. Visualising 3D NMR spectra. strip plots. preparation mixing mixing t1 t2 t3 I S T I
 3D NMR 3D NMR Chris Waudby c.waudby@ucl.ac.uk 3D NMR Visualising 3D NMR spectra preparation mixing mixing t1 t2 t3 I S T I 2 indirect dimensions, independently incremented evolution times Much longer acquisition
3D NMR 3D NMR Chris Waudby c.waudby@ucl.ac.uk 3D NMR Visualising 3D NMR spectra preparation mixing mixing t1 t2 t3 I S T I 2 indirect dimensions, independently incremented evolution times Much longer acquisition
where, c is the speed of light, ν is the frequency in wave numbers (cm -1 ) and µ is the reduced mass (in amu) of A and B given by the equation: ma
 Vibrational Spectroscopy A rough definition of spectroscopy is the study of the interaction of matter with energy (radiation in the electromagnetic spectrum). A molecular vibration is a periodic distortion
Vibrational Spectroscopy A rough definition of spectroscopy is the study of the interaction of matter with energy (radiation in the electromagnetic spectrum). A molecular vibration is a periodic distortion
ORGANIC - CLUTCH CH ANALYTICAL TECHNIQUES: IR, NMR, MASS SPECT
 !! www.clutchprep.com CONCEPT: PURPOSE OF ANALYTICAL TECHNIQUES Classical Methods (Wet Chemistry): Chemists needed to run dozens of chemical reactions to determine the type of molecules in a compound.
!! www.clutchprep.com CONCEPT: PURPOSE OF ANALYTICAL TECHNIQUES Classical Methods (Wet Chemistry): Chemists needed to run dozens of chemical reactions to determine the type of molecules in a compound.
3.15 Nuclear Magnetic Resonance Spectroscopy, NMR
 3.15 Nuclear Magnetic Resonance Spectroscopy, NMR What is Nuclear Magnetic Resonance - NMR Developed by chemists and physicists together it works by the interaction of magnetic properties of certain nuclei
3.15 Nuclear Magnetic Resonance Spectroscopy, NMR What is Nuclear Magnetic Resonance - NMR Developed by chemists and physicists together it works by the interaction of magnetic properties of certain nuclei
Electron Spin Resonance, Basic principle of NMR, Application of NMR in the study of Biomolecules, NMR imaging and in vivo NMR spectromicroscopy
 Electron Spin Resonance, Basic principle of NMR, Application of NMR in the study of Biomolecules, NMR imaging and in vivo NMR spectromicroscopy Mitesh Shrestha Electron Spin Resonance Electron paramagnetic
Electron Spin Resonance, Basic principle of NMR, Application of NMR in the study of Biomolecules, NMR imaging and in vivo NMR spectromicroscopy Mitesh Shrestha Electron Spin Resonance Electron paramagnetic
Chapter 14 Spectroscopy
 hapter 14 Spectroscopy There are four major analytical techniques used for identifying the structure of organic molecules 1. Nuclear Magnetic Resonance or NMR is the single most important technique for
hapter 14 Spectroscopy There are four major analytical techniques used for identifying the structure of organic molecules 1. Nuclear Magnetic Resonance or NMR is the single most important technique for
The importance of constants
 The importance of constants Sourendu Gupta TIFR Graduate School Quantum Mechanics 1 September 22, 2008 Sourendu Gupta (TIFR Graduate School) The importance of constants QM I 1 / 13 Outline 1 Universal
The importance of constants Sourendu Gupta TIFR Graduate School Quantum Mechanics 1 September 22, 2008 Sourendu Gupta (TIFR Graduate School) The importance of constants QM I 1 / 13 Outline 1 Universal
7. Nuclear Magnetic Resonance
 7. Nuclear Magnetic Resonance Nuclear Magnetic Resonance (NMR) is another method besides crystallography that can be used to find structures of proteins. NMR spectroscopy is the observation of spins of
7. Nuclear Magnetic Resonance Nuclear Magnetic Resonance (NMR) is another method besides crystallography that can be used to find structures of proteins. NMR spectroscopy is the observation of spins of
Chapter 7. Nuclear Magnetic Resonance Spectroscopy
 Chapter 7 Nuclear Magnetic Resonance Spectroscopy I. Introduction 1924, W. Pauli proposed that certain atomic nuclei have spin and magnetic moment and exposure to magnetic field would lead to energy level
Chapter 7 Nuclear Magnetic Resonance Spectroscopy I. Introduction 1924, W. Pauli proposed that certain atomic nuclei have spin and magnetic moment and exposure to magnetic field would lead to energy level
Sensitive NMR Approach for Determining the Binding Mode of Tightly Binding Ligand Molecules to Protein Targets
 Supporting information Sensitive NMR Approach for Determining the Binding Mode of Tightly Binding Ligand Molecules to Protein Targets Wan-Na Chen, Christoph Nitsche, Kala Bharath Pilla, Bim Graham, Thomas
Supporting information Sensitive NMR Approach for Determining the Binding Mode of Tightly Binding Ligand Molecules to Protein Targets Wan-Na Chen, Christoph Nitsche, Kala Bharath Pilla, Bim Graham, Thomas
The resonance frequency of the H b protons is dependent upon the orientation of the H a protons with respect to the external magnetic field:
 Spin-Spin Splitting in Alkanes The signal arising from a proton or set of protons is split into (N+1) lines by the presence of N adjacent nuclei Example 1: Bromoethane The resonance frequency of the H
Spin-Spin Splitting in Alkanes The signal arising from a proton or set of protons is split into (N+1) lines by the presence of N adjacent nuclei Example 1: Bromoethane The resonance frequency of the H
- 1/2. = kb o = hνν + 1/2. B o increasing magnetic field strength. degenerate at B o = 0
 NMR EXPERIMENT When magnetically active nuclei are placed into an external magnetic field, the magnetic fields align themselves with the external field into two orientations. During the experiment, electromagnetic
NMR EXPERIMENT When magnetically active nuclei are placed into an external magnetic field, the magnetic fields align themselves with the external field into two orientations. During the experiment, electromagnetic
Magnetic Resonance Spectroscopy
 INTRODUCTION TO Magnetic Resonance Spectroscopy ESR, NMR, NQR D. N. SATHYANARAYANA Formerly, Chairman Department of Inorganic and Physical Chemistry Indian Institute of Science, Bangalore % I.K. International
INTRODUCTION TO Magnetic Resonance Spectroscopy ESR, NMR, NQR D. N. SATHYANARAYANA Formerly, Chairman Department of Inorganic and Physical Chemistry Indian Institute of Science, Bangalore % I.K. International
NMR-spectroscopy in solution - an introduction. Peter Schmieder
 NMR-spectroscopy in solution - an introduction 2/92 Advanced Bioanalytics NMR-Spectroscopy Introductory session (11:00 12:30) Basic aspects of NMR-spectroscopy NMR parameter Multidimensional NMR-spectroscopy
NMR-spectroscopy in solution - an introduction 2/92 Advanced Bioanalytics NMR-Spectroscopy Introductory session (11:00 12:30) Basic aspects of NMR-spectroscopy NMR parameter Multidimensional NMR-spectroscopy
A GENERAL TRANSFORMATION TO CANONICAL FORM FOR POTENTIALS IN PAIRWISE INTERMOLECULAR INTERACTIONS
 1 A GENERAL TRANSFORMATION TO CANONICAL FORM FOR POTENTIALS IN PAIRWISE INTERMOLECULAR INTERACTIONS Jay R. Walton Department of Mathematics, Texas A & M University College Station, TX, USA Luis A. Rivera-Rivera,
1 A GENERAL TRANSFORMATION TO CANONICAL FORM FOR POTENTIALS IN PAIRWISE INTERMOLECULAR INTERACTIONS Jay R. Walton Department of Mathematics, Texas A & M University College Station, TX, USA Luis A. Rivera-Rivera,
In a solution, there are thousands of atoms generating magnetic fields, all in random directions.
 Nuclear Magnetic Resonance Spectroscopy: Purpose: onnectivity, Map of - framework Process: In nuclear magnetic resonance spectroscopy, we are studying nuclei. onsider this circle to represent a nucleus
Nuclear Magnetic Resonance Spectroscopy: Purpose: onnectivity, Map of - framework Process: In nuclear magnetic resonance spectroscopy, we are studying nuclei. onsider this circle to represent a nucleus
NMR Spectroscopy. for 1 st B.Tech INTRODUCTION Lecture -1 Indian Institute of Technology, Dhanbad
 NMR Spectroscopy for 1 st B.Tech Lecture -1 Indian Institute of Technology, Dhanbad by Dr. R P John & Dr. C. Halder INTRODUCTION Nucleus of any atom has protons and neutrons Both Proton and Neutron has
NMR Spectroscopy for 1 st B.Tech Lecture -1 Indian Institute of Technology, Dhanbad by Dr. R P John & Dr. C. Halder INTRODUCTION Nucleus of any atom has protons and neutrons Both Proton and Neutron has
4) protons experience a net magnetic field strength that is smaller than the applied magnetic field.
 1) Which of the following CANNOT be probed by an spectrometer? See sect 16.1 Chapter 16: 1 A) nucleus with odd number of protons & odd number of neutrons B) nucleus with odd number of protons &even number
1) Which of the following CANNOT be probed by an spectrometer? See sect 16.1 Chapter 16: 1 A) nucleus with odd number of protons & odd number of neutrons B) nucleus with odd number of protons &even number
Model-Free Approach to Internal Motions in Proteins
 Model-Free Approach to Internal Motions in Proteins Lipari & Szabo, JACS 104, 4546 (1982) Palmer AG. Ann. Rev. Biophys. Biomol. Struc., 30, 129-155 (2001) Palmer AG, Kroenke CD, Loria JP, Meth. Enzymol.
Model-Free Approach to Internal Motions in Proteins Lipari & Szabo, JACS 104, 4546 (1982) Palmer AG. Ann. Rev. Biophys. Biomol. Struc., 30, 129-155 (2001) Palmer AG, Kroenke CD, Loria JP, Meth. Enzymol.
