Part V: Binary response data
|
|
|
- Lambert Moody
- 5 years ago
- Views:
Transcription
1 Part V: Binary response data 275 BIO 233, Spring 2015
2 Western Collaborative Group Study Prospective study of coronary heart disease (CHD) Recruited 3,524 men aged between employed at 10 companies in California baseline survey at intake annual surveys until December 1969 Exclusions: 78 men who were actually outside the pre-specified age range 141 subjects with CHD manifest at intake 106 employees at one firm that excluded itself from follow-up 45 subjects who were lost to follow-up, non-chd death or self-exclusion prior to the first follow-up n = 3,154 study participants at risk for CHD 276 BIO 233, Spring 2015
3 Our primary goal is to investigate the relationship between behavior pattern and risk of CHD Participants were categorized into one of two behavior pattern groups: Type A: characterized by enhanced aggressiveness, ambitiousness, competitive drive, and chronic sense of urgency Type B: characterized by more relaxed and non-competitive Data and documentation are available on the class website > ## > load("wcgs_data.dat") > > dim(wcgs) [1] > names(wcgs) [1] "age" "ht" "wt" "sbp" "dbp" "chol" "ncigs" "behave" [9] "chd" "type" "time" 277 BIO 233, Spring 2015
4 The variables (in column order) are: 1 age age, years 2 ht height, in 3 wt weight, lbs 4 sbp systolic blood pressure, mmhg 5 dbp diastolic blood pressure, mmhg 6 chol cholesterol, mg/dl 7 ncigs number of cigarettes smoked per day 8 behave behavior type 0/1 = B/A 9 chd occurrence of a CHD event during follow-up 10 type type of CHD event 11 time time post-recruitment of the CHD event, days Values for the risk factor covariates are those measured at the intake visit The three CHD-related variables were measured prospectively over an approx. 8.5 years of follow-up 278 BIO 233, Spring 2015
5 Important note: 423 were lost to follow-up 140 men died during the follow-up For our purposes, we are going to ignore these issues and consider the binary outcome of: 1 occurrence of CHD during follow-up Y = 0 otherwise In the dataset, the response variable is chd : > ## > table(wcgs$chd) > round(mean(wcgs$chd) * 100, 1) [1] BIO 233, Spring 2015
6 Primary exposure of interest is behave : > ## > table(wcgs$behave) > round(mean(wcgs$behave) * 100, 1) [1] 50.4 Cross-tabulation and exposure-specific incidence > ## > table(wcgs$behave, wcgs$chd) > round(tapply(wcgs$chd, list(wcgs$behave), FUN=mean) * 100, 1) BIO 233, Spring 2015
7 The probability of the occurrence of CHD during follow-up among type B men is estimated to be expected percentage of type B men who will develop CHD during follow-up is 5.0% The probability of the occurrence of CHD during follow-up among type A men is estimated to be expected percentage of type A men who will develop CHD during follow-up is 11.2% Often use the generic term risk Either way, it s important to remember that these statements are referring to populations of men, rather than the individuals themselves we ve estimated a common or average risk of CHD referred to as the marginal risk marginal in the sense that it does not condition on anything else 281 BIO 233, Spring 2015
8 Contrasts As stated at the start, the primary goal is to investigate the relationship between behavior pattern and risk of CHD We ve characterized risk for each type but the goal requires a comparison of the risks To perform such a comparison we need to choose a contrast Risk difference: RD = = difference in the estimated risk of CHD during follow-up between type A and type B men is (or 6.2%) the way in which the additional risk of CHD of being a type A person manifests through an absolute increase 282 BIO 233, Spring 2015
9 Relative risk: RR = / = 2.24 ratio of the estimated risk of CHD for type A men during follow-up to the estimated risk for type B men the way in which the additional risk of CHD of being a type A person manifests through an relative increase As with the interpretation of the risks themselves, these statements refer to contrasts between populations population of Type A men vs. population of Type B men Contrasts are marginal in the sense that we don t condition on anything else when comparing the two populations i.e. we don t adjust for anything 283 BIO 233, Spring 2015
10 Important to note that the RD and RR are related relationship depends on the value of the response probability for the referent group RD across different combinations of P(Y = 1 X = 0) and RR RR = RR = RR = RR = RR = RR = RR = NA RR = NA NA 284 BIO 233, Spring 2015
11 The RD may be small even if the RR is big for either protective or detrimental effects When the RR is small, the RD is also small unless P(Y = 1 X = 0) is big common outcome However a small RR operating on a large population could correspond to a big public health impact this rationale is often cited in studies of air pollution To move beyond simple contrasts, we need a more general framework for modeling the relationship between the binary response and a vector of covariates 285 BIO 233, Spring 2015
12 GLMs for binary data We ve noted that the Bernoulli distribution is the only possible distribution for binary data Y Bernoulli(µ) f Y (y;µ) = µ y (1 µ) 1 y f Y (y;θ,φ) = exp{yθ log(1+exp{θ})} θ = log ( ) µ 1 µ a(φ) = 1 b(θ) = log(1+exp{θ}) c(y,φ) = BIO 233, Spring 2015
13 The log-likelihood is l(β;y) = = n i=1 n i=1 y i θ i b(θ i ) y i θ i log(1+exp{θ i }) where θ i is a function of β via and g(µ i ) = X T i β µ i = exp{θ i } 1+exp{θ i } 287 BIO 233, Spring 2015
14 The score function for β j is l(β; y) β j = n i=1 µ i η i X j,i µ i (1 µ i ) (y i µ i ) where the expression for µ i / η i is dependent on the choice of the link function g( ) Since the log-likelihood is only a function of β, the expected information matrix is given by the (p+1) (p+1) matrix: I ββ = X T WX where X is the design matrix for the model and W is a diagonal matrix with i th diagonal element W i = ( µi η i ) 2 1 µ i (1 µ i ) 288 BIO 233, Spring 2015
15 Link functions In a GLM, the systematic component is given by g(µ i ) = η i = X T i β We ve noted previously that, for binary data, there are various options for link functions including: linear: g(µ i ) = µ i log: g(µ i ) = log(µ i ) ( ) µi logit: g(µ i ) = log 1 µ i probit: g(µ i ) = probit(µ i ) complementary log-log: g(µ i ) = log{ log(1 µ i )} 289 BIO 233, Spring 2015
16 Q: How do we make a choice from among these options? Balance between interpretability and mathematical properties interpretability of contrasts mathematical properties in terms of fitted values being in the appropriate range 290 BIO 233, Spring 2015
17 Linear (identity) link function µ i = β 0 +β 1 X i Interpret β 0 as the probability of response when X = 0 Interpret β 1 as the change in the probability of response, comparing two populations whose value of X differs by 1 unit The contrast we are modeling the risk difference (RD) As we ve noted, a potential problem is that this specification of the model doesn t respect the fact that the (true) response probability is bounded 291 BIO 233, Spring 2015
18 Log link function log(µ i ) = β 0 +β 1 X i Interpret β 0 as the log of the probability of response when X = 0 exp{β 0 } is the probability of response when X = 0 Interpret β 1 as the change in the log of the probability of response, comparing two populations whose value of X differs by 1 unit exp{β 1 } is the ratio of the probability of response when X = 1 to that when X = 0 The contrast we are modeling the risk ratio (RR) 292 BIO 233, Spring 2015
19 As with the linear link, this choice of link function doesn t necessarily respect the fact that the (true) response probability is bounded Can see this explicitly this by considering the inverse of the link function: µ i = exp{x T i β} which takes values on (0, ) 293 BIO 233, Spring 2015
20 Logit link function logit(µ i ) = log ( µi 1 µ i ) = X T i β The functional µ i = P(Y i = 1 X i ) 1 µ i P(Y i = 0 X i ) is the odds of response Interpret β 0 as the log of the odds of response when X = 0 exp{β 0 } is the odds of response when X = BIO 233, Spring 2015
21 Interpret β 1 as the change in the log of the odds of response, comparing two populations whose value of X differs by 1 unit exp{β 1 } is the ratio of the odds of response when X = 1 to that when X = 0 The contrast we are modeling is the odds ratio (OR) Considering the inverse of the link function yields: µ i = exp{x T i β} 1+exp{X T i β} referred to as the expit function 295 BIO 233, Spring 2015
22 The expit function is the CDF of the standard logistic distribution distribution for a continuous random variable with support on (, ) pdf is given by f X (x) = exp{ x} (1+exp{ x}) 2 The CDF (of any distribution) provides a mapping from the support of the random variable to the (0,1) interval F X ( ) : (, ) (0,1) We could use the inverse CDF of any distribution as a link function F 1 X ( ) : (0,1) (, ) g( ) F 1 ( ) maps µ (0,1) to η (, ) 296 BIO 233, Spring 2015
23 Probit link function probit(µ i ) = Φ 1 (µ i ) = X T i β where Φ( ) is the CDF of the standard normal distribution Interpret β 0 as the probit of probability of response when X = 0 Interpret β 1 as the change in the probit of the probability of response, comparing two populations whose value of X differs by 1 unit Interpretation is tricky contrast is in terms of the inverse CDF of a standard normal distribution no easy way of relating this contrast to more intuitive measures 297 BIO 233, Spring 2015
24 Complementary log-log function log{ log(1 µ i )} = X T i β Inverse CDF of the extreme value (or log-weibull) distribution As with the probit link function, there isn t any intuitive way of interpreting regression parameters based on this link function Has the distinction that it is asymmetric may be useful if the primary purpose is prediction 298 BIO 233, Spring 2015
25 Comparisons Over values of µ (0.1,0.9), models based on the linear, logit and probit link function agree approximately considering their inverse link functions, over the range η i ( 2,2): ( ) η i 4 expit(η 2πηi i) Φ 4 so their fitted values will be approximately equal over this range Also use these relationships to provide approximate relationships between the regression parameters: β linear 1 4 β logit 1 2πβ probit BIO 233, Spring 2015
26 Conditional mean, µ i linear logit probit Linear predictor, η i 300 BIO 233, Spring 2015
27 5 c log log logit probit log g(µ i ) logit(µ i ) 301 BIO 233, Spring 2015
28 From the figures, differences across these link functions manifest primarily in the tails when the probability of response is small or large Also, the logit and probit functions are almost linearly related noted this from the approximations as well For small values of µ i, the complementary log-log, logit and log functions are close to each other equally good for rare events for µ i 0.1 log ( µi 1 µ i ) log(µ i ) log link has the best interpretation OR and RR are close numerically 302 BIO 233, Spring 2015
29 Modeling: WCGS Returning to the WCGS, the dataset has a number of covariates that we might consider including in a model > ## > names(wcgs) [1] "age" "ht" "wt" "sbp" "dbp" "chol" "ncigs" "behave" [9] "chd" "type" "time" Q: How do we approach making decisions about what to include in the model? depends on the purpose of the analysis Towards this, it s useful to classify the analysis into one of two types: association studies prediction studies 303 BIO 233, Spring 2015
30 Association studies The goal is to characterize the relationship between some exposure of interest and the response establish cause-and-effect Understanding the underlying (data generating) mechanisms are crucial need to be attentive to the possibility of alternative explanations control of confounding is crucial Model selection, in terms of the choice of potential confounders, should be based on scientific considerations Despite this ideal, it s not always clear which covariates are confounders and which aren t 304 BIO 233, Spring 2015
31 One strategy is to fit and report the following three models: (1) an unadjusted or minimally adjusted model (2) a model that includes core confounders clear indication from scientific knowledge and/or the literature consensus among investigators (3) a model that includes core confounders plus any potential confounders indication is less certain Report results from model (2) as primary based conclusions on the results of this model interpret models (1) and (3) in terms of sensitivity analyses There are, of course, other philosophies on this! 305 BIO 233, Spring 2015
32 Prediction studies The goal is to estimate the response Y as opposed to the goal of estimating β In contrast to association studies, prediction is typically not hypothesis-driven there is no single exposure or association or parameter that is of interest mechanisms and confounding is less of a concern, if at all Choice of which covariates to include in the model is driven by the extent to which its inclusion improves our ability to predict future outcomes care is needed not to overfit the data These issues typically don t come up in association studies requires different analysis strategies and different statistical tools 306 BIO 233, Spring 2015
33 Confounding The data for the WCGS is observational as a study of Type A vs Type B behavior patterns, the investigators didn t randomize behavior pattern As such, an analysis based on these data may be subject to confounding bias A confounder is defined as a covariate that is (causally) associated with both the exposure of interest and the outcome of interest, while not being on the causal pathway X C? Y 307 BIO 233, Spring 2015
34 Intuitively, from the causal diagram, there is a backdoor association between X and Y, through C If one does not block this pathway then one cannot isolate the (direct) association between X and Y the unadjusted association is spurious in the sense that it is a mixture of the true association and the association characterized by the backdoor pathway confounding bias Note, we haven t introduced any estimators yet we haven t even introduced a contrast yet! As such, confounding is a scientific issue distinct from statistical bias that is an operating characteristic of an estimator 308 BIO 233, Spring 2015
35 The control of confounding bias must, therefore, be approached from a scientific perspective we cannot use statistical techniques to determine whether or not a covariate is a confounder we must use scientific knowledge to make these decisions Given a collection of (potential) confounders, the standard approach to controlling confounding bias is to include them in the linear predictor referred to as regression adjustment e.g., η i = β 0 + β x X i + β c C i interpret β x conditional on C or within strata of C 309 BIO 233, Spring 2015
36 Going back to the causal diagram, conditioning on the confounder blocks the backdoor pathway the effect of including C in the model is to break the association between C and Y X C? Y 310 BIO 233, Spring 2015
37 Exploratory data analysis Whatever the purpose of the study, it is often useful to perform some preliminary exploratory data analysis Q: Why? > ## > apply(wcgs[,1:7], 2, FUN=summary) $age Min. 1st Qu. Median Mean 3rd Qu. Max $ht Min. 1st Qu. Median Mean 3rd Qu. Max $wt Min. 1st Qu. Median Mean 3rd Qu. Max. 311 BIO 233, Spring 2015
38 $sbp Min. 1st Qu. Median Mean 3rd Qu. Max $dbp Min. 1st Qu. Median Mean 3rd Qu. Max $chol Min. 1st Qu. Median Mean 3rd Qu. Max. NA s $ncigs Min. 1st Qu. Median Mean 3rd Qu. Max BIO 233, Spring 2015
39 313 BIO 233, Spring 2015 Frequency Age, years
40 Weight, lbs Height, in 314 BIO 233, Spring 2015
41 Diastolic blood pressure, mmhg 315 BIO 233, Spring 2015 Systolic blood pressure, mmhg
42 Study id Cholesterol, mg/dl 316 BIO 233, Spring 2015 Cholesterol, mg/dl Frequency
43 Frequency Number of cigarettes, per day 317 BIO 233, Spring 2015
44 > ## > table(wcgs$ncigs) Study participants seem to be reporting round numbers likely some misclassification of actual smoking 318 BIO 233, Spring 2015
45 Overall, nothing too worrying pops out Some instances of large values weight of 320lbs diastolic blood pressure of 150 mmhg cholesterol of 645mg/dL smoking 99 cigarettes per day There is also some missingness in the data in a real collaborative setting, we d want to know more about the cholesterol values in particular, why were they missing? only 12 out of 3,154 observations with missing values 319 BIO 233, Spring 2015
46 Based on the EDA, perform the following data manipulations: > ## > wcgs$chol[wcgs$chol > 500] <- NA ## Take out (particularly) strange value > wcgs <- na.omit(wcgs) ## Remove observations with missing chol > > ## Standardize continuous variables to make the intercept interpretable > ## > wcgs$age <- (wcgs$age - 40) / 5 > wcgs$ht <- (wcgs$ht - 70) / 2 > wcgs$wt <- (wcgs$wt - 170) / 10 > wcgs$sbp <- (wcgs$sbp - 125) / 10 > wcgs$dbp <- (wcgs$dbp - 80) / 10 > wcgs$chol <- (wcgs$chol - 200) / 20 > > ## Smoker 0/1 = No/Yes > ## > wcgs$smoker <- as.numeric(wcgs$ncigs > 0) 320 BIO 233, Spring 2015
47 Unadjusted analysis Fit the logistic regression model: logit(µ i ) = β 0 + β 1 behave i > ## > fit0 <- glm(chd ~ behave, family=binomial(), data=wcgs) > summary(fit0) Call: glm(formula = chd ~ behave, family = binomial(), data = wcgs) Deviance Residuals: Min 1Q Median 3Q Max BIO 233, Spring 2015
48 Coefficients: Estimate Std. Error z value Pr(> z ) (Intercept) <2e-16 *** behave e-09 *** --- Signif. codes: 0 *** ** 0.01 * (Dispersion parameter for binomial family taken to be 1) Null deviance: on 3140 degrees of freedom Residual deviance: on 3139 degrees of freedom AIC: Number of Fisher Scoring iterations: 5 > summary(fit0$fitted) Min. 1st Qu. Median Mean 3rd Qu. Max BIO 233, Spring 2015
49 Core adjustment Add core adjustment variables into the linear predictor and fit logit(µ i ) = β 0 + β 1 behave i + β 2 age i + β 3 wt i + β 4 sbp i + β 5 chol i + β 6 smoker i > ## > fit1 <- glm(chd ~ behave + age + wt + sbp + chol + smoker, family=binomial(), data=wcgs) > summary(fit1)... Coefficients: Estimate Std. Error z value Pr(> z ) (Intercept) < 2e-16 *** behave e-06 *** age e-07 *** wt ** sbp e-05 *** 323 BIO 233, Spring 2015
50 chol e-12 *** smoker e-05 *** --- Signif. codes: 0 *** ** 0.01 * (Dispersion parameter for binomial family taken to be 1) Null deviance: on 3140 degrees of freedom Residual deviance: on 3134 degrees of freedom AIC: Number of Fisher Scoring iterations: 6 > summary(fit1$fitted) Min. 1st Qu. Median Mean 3rd Qu. Max BIO 233, Spring 2015
51 Full adjustment Add remaining adjustment variables into the linear predictor and fit logit(µ i ) = β 0 + β 1 behave i + β 2 age i + β 3 wt i + β 4 sbp i + β 5 chol i + β 6 smoker i + β 7 ht i + β 8 dbp i > fit2 <- glm(chd ~ behave + age + wt + sbp + chol + smoker + ht + dbp, family=binomial(), data=wcgs) > summary(fit2)... Coefficients: Estimate Std. Error z value Pr(> z ) (Intercept) < 2e-16 *** behave e-06 *** age e-07 *** wt * sbp ** 325 BIO 233, Spring 2015
52 chol e-12 *** smoker e-05 *** ht dbp Signif. codes: 0 *** ** 0.01 * (Dispersion parameter for binomial family taken to be 1) Null deviance: on 3140 degrees of freedom Residual deviance: on 3132 degrees of freedom AIC: Number of Fisher Scoring iterations: 6 > summary(fit2$fitted) Min. 1st Qu. Median Mean 3rd Qu. Max BIO 233, Spring 2015
53 Interpretation of results Characterizing the effect of behavior type is the primary scientific goal typically report results on the odds ratio scale denote the odds ratio by θ 1 = exp{β 1 } 95% CIs can be obtained in a number of ways (i) compute the 95% CI for ˆβ 1 and exponentiate (ii) compute a 95% CI directly for ˆθ 1 glm() returns the standard error estimates for the ˆβ s use the delta method to get the standard error for ˆθ 1 Approaches are equivalent asymptotically in small samples, first approach results in an asymmetric CI 327 BIO 233, Spring 2015
54 getci() function implements the first approach code is available on the class website > ## > getci(fit0) exp{beta} lower upper (Intercept) behave Interpretation of ˆθ 1 = 2.36: > ## > getci(fit1)[1:2,] exp{beta} lower upper (Intercept) behave Interpretation of ˆθ 1 = 1.99: 328 BIO 233, Spring 2015
55 > ## > getci(fit2)[1:2,] exp{beta} lower upper (Intercept) behave Interpretation of ˆθ 1 = 1.98: 329 BIO 233, Spring 2015
56 Flexible adjustment When we include potential confounders in the model, we are less concerned with their interpretation primary purpose is the control of confounding bias if we don t model the effects of confounders properly, there may be residual confounding Suggest including these covariates into the model in as flexible manner as possible go beyond linearity Two simple strategies for flexibly modeling continuous covariates are (i) including additional polynomial terms (ii) categorization 330 BIO 233, Spring 2015
57 > ## Polynomial > ## > wcgs$age2 <- wcgs$age^2 > wcgs$age3 <- wcgs$age^3... > > ## Categorization > ## > wcgs$cigscat <- 0 > wcgs$cigscat[wcgs$ncigs >= 10] <- 1 > wcgs$cigscat[wcgs$ncigs >= 20] <- 2 > wcgs$cigscat[wcgs$ncigs >= 30] <- 3 > wcgs$cigscat[wcgs$ncigs >= 40] <- 4 > > ## > flex1 <- glm(chd ~ behave + age + age2 + age3 + wt + wt2 + wt3 + sbp + sbp2 + sbp3 + chol + chol2 + chol3 + factor(cigscat), family=binomial(), data=wcgs) 331 BIO 233, Spring 2015
58 > summary(flex1)... Estimate Std. Error z value Pr(> z ) (Intercept) < 2e-16 *** behave e-06 *** age age age wt * wt ** wt ** sbp * sbp sbp chol e-05 *** chol chol factor(cigscat) factor(cigscat) *** factor(cigscat) e-06 *** factor(cigscat) ** > 332 BIO 233, Spring 2015
59 > ## > getci(fit1)[1:2,] exp{beta} lower upper (Intercept) behave > > getci(flex1)[1:2,] exp{beta} lower upper (Intercept) behave > > ## > LRtest(fit1, flex1) Test Statistic = 25.5 on 11 df => p-value = 0.01 [1] 0.01 The likelihood ratio test suggests a better fit but there is virtually no impact on estimation or inference 333 BIO 233, Spring 2015
60 Link functions So far, we ve only considered the logit link function g(µ i ) = log ( µi 1 µ i ) = X T i β By far the most common link function used for GLMs of binary data guaranteed that fitted values are in (0,1) reasonable interpretation of contrasts in terms of odds ratios when the event is rare: OR RR ability to analyze case-control data as if it had been collected prospectively Q: What about other link functions? 334 BIO 233, Spring 2015
61 Potential choices include: linear: g(µ i ) = µ i log: g(µ i ) = log(µ i ) probit: g(µ i ) = probit(µ i ) complementary log-log: g(µ i ) = log{ log(1 µ i )} We ve noted that there is a trade-off between interpretability and mathematical properties For the goal of characterizing the association between behavior type and risk of CHD, interpretability is crucial examine the linear and log link functions If the goals is prediction, then we d be more likely to entertain the probit and complementary log-log link functions 335 BIO 233, Spring 2015
62 In R we use the family argument to change the link other components of the GLM that are functions of the link are appropriately adjusted Let s first consider changing the link function for the unadjusted analysis for the binomial family, the logit link is the default but just to show you how it works > ## > logitf0 <- glm(chd ~ behave, family=binomial(link="logit"), data=wcgs) > summary(logitf0$fitted) Min. 1st Qu. Median Mean 3rd Qu. Max > getci(logitf0) exp{beta} lower upper (Intercept) behave BIO 233, Spring 2015
63 Now let s fit model using the linear link: > ## > linearf0 <- glm(chd ~ behave, family=binomial(link="identity"), data=wcgs) > summary(linearf0$fitted) Min. 1st Qu. Median Mean 3rd Qu. Max > getci(linearf0, expo=false, digits=4) * 100 beta lower upper (Intercept) behave Notice that the fitted values are the same as those obtained using the logit link Q: Why? Interpretation of ˆβ 1 = 6.11: 337 BIO 233, Spring 2015
64 Finally, let s fit model using the log link: > ## > logf0 <- glm(chd ~ behave, family=binomial(link="log"), data=wcgs) > summary(logf0$fitted) Min. 1st Qu. Median Mean 3rd Qu. Max > getci(logf0) exp{beta} lower upper (Intercept) behave Again, notice that the fitted values are the same Interpretation of ˆθ 1 = 2.21: 338 BIO 233, Spring 2015
65 Q: How does changing the link function impact the adjusted analysis? > ## > logitf1 <- glm(chd ~ behave + age + wt + sbp + chol + smoker, family=binomial(), data=wcgs) > getci(logitf1)[1:2,] exp{beta} lower upper (Intercept) behave > > ## > linearf1 <- glm(chd ~ behave + age + wt + sbp + chol + smoker, family=binomial(link="identity"), data=wcgs) Error: no valid set of coefficients has been found: please supply starting values The IWLS algorithm is having trouble finding valid starting values 339 BIO 233, Spring 2015
66 Taking a closer look at the glm() function > > args(glm) function (formula, family = gaussian, data, weights, subset, na.action, start = NULL, etastart, mustart, offset, control = list(...), model = TRUE, method = "glm.fit", x = FALSE, y = TRUE, contrasts = NULL,...) NULL we can provide our own starting values via start for the regression coefficients, β etastart for the linear predictors, {η 1..., η n } mustart for the fitted value, {µ 1,..., µ n } Use values from some other fit that was successful a fit using some other link function a fit based on a different mean model 340 BIO 233, Spring 2015
67 Using a linear link with binary data we also have to be careful about the mean-variance relationship specified by the binomial() family > > names(binomial()) [1] "family" "link" "linkfun" "linkinv" "variance" [6] "dev.resids" "aic" "mu.eta" "initialize" "validmu" [11] "valideta" "simulate" > > binomial()$variance function (mu) mu * (1 - mu) If, at any point during the IWLS algorithm, one of the fitted values is outside (0,1) then the variance will be negative unlikely that the algorithm will converge 341 BIO 233, Spring 2015
68 An alternative is to use OLS and use an appropriate variance estimator to account for the heteroskedasticity induced by the mean-variance relationship Huber-White variance estimator sandwich estimator robust estimator bootstrap variance estimator In R use the lm() function function robustci(), available on the class website, computes robustand bootstrap-based 95% confidence intervals > ## > linearf1 <- lm(chd ~ behave + age + wt + sbp + chol + smoker, data=wcgs) > robustci(linearf1, digits=4, B=1000) * BIO 233, Spring 2015
69 betahat Naive Lo Naive Up Robust Lo Robust Up Boot Lo Boot Up (Intercept) behave age wt sbp chol smoker Interpretation of ˆβ 1 = 4.59: Q: What about the negative fitted values? > ## > summary(linearf1$fitted) Min. 1st Qu. Median Mean 3rd Qu. Max BIO 233, Spring 2015
70 Fitted values using a logit link 344 BIO 233, Spring 2015 Fitted values using a linear link
71 Clearly, some of the fitted values are < 0 > ## > range(logitf1$fitted[linearf1$fitted <= 0]) [1] Fitted values that are < 0 are all small Fitted values that are > 0 are in a much tighter range of values maximum value of 0.326, as opposed to for the logistic model Turning to the log link: > ## > logf1 <- glm(chd ~ behave + age + wt + sbp + chol + smoker, family=binomial(link="log"), data=wcgs) Error: no valid set of coefficients has been found: please supply starting values 345 BIO 233, Spring 2015
72 This time we can t use the lm() function but we can provide starting values from the (successful) fit of the logistic regression: > ## > logf1 <- glm(chd ~ behave + age + wt + sbp + chol + smoker, family=binomial(link="log"), mustart=fitted(logitf1), data=wcgs) > getci(logf1)[1:2,] exp{beta} lower upper (Intercept) behave > summary(logf1$fitted) Min. 1st Qu. Median Mean 3rd Qu. Max All of the fitted values are in (0,1) Interpretation of ˆθ 1 = 1.78: 346 BIO 233, Spring 2015
73 Fitted values using a logit link 347 BIO 233, Spring 2015 Fitted values using a log link
74 Summary of results: Link Contrast Unadjusted Adjusted function model model logit OR 2.36 (1.79, 3.10) 1.99 (1.50, 2.64) linear RD 6.11 (4.21, 8.01) 4.59 (2.75, 6.43) log RR 2.21 (1.71, 2.85) 1.78 (1.38, 2.29) 95% CI based on the Huber-White robust standard error estimate Convincing evidence of a statistically significant difference between Type A and Type B behavior types in CHD risk however you define the contrast Q: Do you think we can claim clinical significance? 348 BIO 233, Spring 2015
75 The Bayesian Solution GLMs for binary data are specified by: Y i X i Bernoulli(µ i ) g(µ i ) = X T i β The unknown parameters are the regression coefficients: β p + 1 parameters In the absence of prior knowledge, it is typical to adopt a flat prior π(β) BIO 233, Spring 2015
76 Computation Generate samples from the posterior π(β y) L(β; y)π(β) via the Metropolis-Hastings algorithm Use the asymptotic sampling distribution of the MLE as a proposal distribution q(β;y) Normal( β MLE, I 1 ββ ) from the (usual) frequentist fit of the GLM Also use this distribution for starting values 350 BIO 233, Spring 2015
77 ## fit1 <- glm(chd ~ behave + age + wt + sbp + chol + smoker, family=binomial(), data=wcgs) ## betahat <- fit1$coef betavar <- summary(fit1)$cov.unscaled X <- model.matrix(fit1) Y <- model.frame(fit1)[,1] ## 3 chains, each for 1,000 scans ## M <- 3 R < startvals <- rmvnorm(m, betahat, betavar) posterior <- array(na, dim=c(r, length(betahat), M)) accept <- array(0, dim=c(r, M)) for(m in 1:M) { ## beta <- startvals[m,] mu <- as.vector(expit(x %*% beta)) 351 BIO 233, Spring 2015
78 } ## for(r in 1:R) { ## betastar <- as.vector(rmvnorm(1, betahat, betavar)) mustar <- as.vector(expit(x %*% betastar)) ## logpiratio <- sum(dbinom(y, 1, mustar, log=true)) - sum(dbinom(y, 1, mu, log=true)) logqratio <- log(dmvnorm(beta, betahat, betavar)) - log(dmvnorm(betastar, betahat, betavar)) ar <- exp(logpiratio + logqratio) if(runif(1) < ar) { beta <- betastar mu <- mustar accept[r,m] <- 1 } posterior[r,,m] <- beta } 352 BIO 233, Spring 2015
79 Examine trace plots for evidence of convergence (or lack thereof) Intercept, β Scan behave logor, β Scan 353 BIO 233, Spring 2015
80 Acceptance rate for the Metropolis-Hastings algorithm: > ## > accrate <- round(apply(accept, 2, mean) * 100, 1) > accrate [1] Proposal and posterior distribution for the log-or of behave, β 1 proposal posterior BIO 233, Spring 2015
81 Summaries of the posterior distribution potential scale reduction (PSR) results based on the Bayesian analysis pool samples from the 3 chains, each with 10% burn in MLE and 95% confidence interval PSR Median 2.5% 97.5% exp{beta} lower upper (Intercept) behave age wt sbp chol smoker Numerical results based on the Bayesian and frequentist analyses are virtually identical differ in their interpretation 355 BIO 233, Spring 2015
82 Posterior distribution for the OR of behave, θ 1 = exp{β 1 } posterior median/mean and (central) 95% credible interval BIO 233, Spring 2015
83 Log link Suppose we want to model the RR, rather than the OR log link, rather the logit link In terms of the model specification, the only thing that changes is the dependence of the mean on the linear predictor: Y i X i Bernoulli(µ i ) log(µ i ) = X T i β form of the likelihood is the same Retain the flat prior for β even though the parameters are different 357 BIO 233, Spring 2015
84 Operationally we need to modify the Metropolis-Hasthings algorithm: (1) change how the µ i s are calculated to evaluate the likelihood/posterior µ i = expit(x T i β) µ i = exp(x T i β) (2) check that the proposed value of β yields a valid set of µ i s if the proposal yields any µ i / (0,1) then we automatically reject the proposal will have zero posterior probability 358 BIO 233, Spring 2015
85 At the r th scan for the m th chain, the algorithm proceeds as: ## betastar <- as.vector(rmvnorm(1, betahat, betavar)) mustar <- as.vector(exp(x %*% betastar)) ## change to the link ## if(sum(mustar <= 0 mustar >= 1) == 0) { logpiratio <- sum(dbinom(y, 1, mustar, log=true)) - sum(dbinom(y, 1, mu, log=true)) logqratio <- log(dmvnorm(beta, betahat, betavar)) - log(dmvnorm(betastar, betahat, betavar)) ar <- exp(logpiratio + logqratio) if(runif(1) < ar) { beta <- betastar mu <- mustar accept[r,m] <- 1 } posterior[r,,m] <- beta } 359 BIO 233, Spring 2015
86 Examine trace plots for evidence of convergence (or lack thereof) Intercept, β Scan behave logor, β Scan 360 BIO 233, Spring 2015
87 Acceptance rate for the Metropolis-Hastings algorithm: > ## > accrate <- round(apply(accept, 2, mean) * 100, 1) > accrate [1] Results: PSR Median 2.5% 97.5% exp{beta} lower upper (Intercept) behave age wt sbp chol smoker Again, the numerical results are virtually identical although the interpretation differs 361 BIO 233, Spring 2015
88 Confounding and Collapsibility Linear regression For a continuous response variable, consider two models: E[Y X,Z] = β 0 + β 1 X + β 2 Z (1) E[Y X] = α 0 + α 1 X (2) In model (1), β 1 is a conditional parameter contrast conditions on the value of Z In model (2), α 1 is a marginal parameter contrast does not condition on anything Q: How are these parameters related? 362 BIO 233, Spring 2015
89 It s straightforward to show that E[Y X] = E[E[Y X,Z]] = E[Y X,Z]f Z X (Z = z X) z z = β 0 + β 1 X + β 2 E[Z X] So the marginal contrast equals α 1 = E[Y X = (x+1)] E[Y X = x] = β 1 + β 2 {E[Z X = (x+1)] E[Z X = x]} The expression within the brackets is the slope from a linear regression of Z X 363 BIO 233, Spring 2015
90 Using this fact, we can write α 1 = β 1 + β 2 COV[X,Z] V[X] the marginal contrast is the conditional contrast plus a bias term Bias requires both β 2 0 and COV[X,Z] 0 Z is related to Y Z is related to X i.e. Z is a confounder The direction of the bias depends on the interplay between β 2 and COV[X, Z] confounding bias may be positive or negative confounding may result in an estimate that is too big or too small 364 BIO 233, Spring 2015
91 If either β 2 = 0 or COV[X,Z] = 0 then β 1 = α 1 Therefore, if Z is a precision variable then β 1 and α 1 have different interpretations the same numerical value However, as the name suggests, the standard error of β 1 will be smaller than the standard error for α 1 Suggests that adjusting for a precisions variable is a good thing, even if one is interested in the marginal association 365 BIO 233, Spring 2015
92 Logistic regression Q: Does the same hold for logistic regression? how are the marginal and conditional parameters related? For a binary outcome, consider two models: logit E[Y X,Z] = β 0 + β 1 X + β 2 Z (3) logit E[Y X] = α 0 + α 1 X (4) The conditional odds ratio for a binary X is θ c x = exp{β 1 } = E[Y = 1 X = 1,Z] E[Y = 0 X = 1,Z] / E[Y = 1 X = 0,Z] E[Y = 0 X = 0,Z] conditional on the value of Z 366 BIO 233, Spring 2015
93 The marginal odds ratio for X is θ m x = exp{α 1 } = E[Y = 1 X = 1] E[Y = 0 X = 1] / E[Y = 1 X = 0] E[Y = 0 X = 0] where E[Y X] = E[Y X,Z]f Z X (Z = z X) z z The relationship between the conditional contrast θ c x and marginal contrast θ m x is not straightforward no simple, closed-form expression for θ m x as a function of θ c x In particular, unlike in the setting of linear regression, they are not linearly related 367 BIO 233, Spring 2015
94 We can, however, calculate θ m x numerically To do so, from the expression for E[Y X], we need to specify E[Y X,Z] f Z X (Z = z X) The first component is given by the logistic regression model: logit E[Y X,Z] = β 0 + β 1 X + β 2 Z For binary X and Z, it s convenient to represent f Z X (Z = z X) via the logistic regression logit E[Z X] = γ 0 + γ 1 X notationally, let φ XZ = exp{γ 1 } denote the X/Z odds ratio 368 BIO 233, Spring 2015
95 The following slides consider the percent difference: θ m 1 θ c 1 θ c under various scenarios for the conditional odds ratio for X, θ c x the conditional odds ratio for Z, θ c z the X/Z odds ratio, φ XZ Throughout, the following are held fixed P(X = 1) = 0.2 P(Z = 1 X = 0) = 0.2 P(Y = 1) = 0.1 R code is available on the course website 369 BIO 233, Spring 2015
96 Strong confounder/exposure association: φ XZ = 0.33 Percentage difference between θ X m and θx c θ X c = 0.20 θ X c = 0.50 θ X c = 0.67 θ X c = 1.00 θ X c = 1.50 θ X c = 2.00 θ X c = Conditional odds ratio for Z, θ Z c 370 BIO 233, Spring 2015
97 Strong confounder/exposure association: φ XZ = 3.00 Percentage difference between θ X m and θx c θ X c = 0.20 θ X c = 0.50 θ X c = 0.67 θ X c = 1.00 θ X c = 1.50 θ X c = 2.00 θ X c = Conditional odds ratio for Z, θ Z c 371 BIO 233, Spring 2015
98 Moderate confounder/exposure association: φ XZ = 0.50 Percentage difference between θ X m and θx c θ X c = 0.20 θ X c = 0.50 θ X c = 0.67 θ X c = 1.00 θ X c = 1.50 θ X c = 2.00 θ X c = Conditional odds ratio for Z, θ Z c 372 BIO 233, Spring 2015
99 Moderate confounder/exposure association: φ XZ = 2.00 Percentage difference between θ X m and θx c θ X c = 0.20 θ X c = 0.50 θ X c = 0.67 θ X c = 1.00 θ X c = 1.50 θ X c = 2.00 θ X c = Conditional odds ratio for Z, θ Z c 373 BIO 233, Spring 2015
100 Weak confounder/exposure association: φ XZ = 0.80 Percentage difference between θ X m and θx c θ X c = 0.20 θ X c = 0.50 θ X c = 0.67 θ X c = 1.00 θ X c = 1.50 θ X c = 2.00 θ X c = Conditional odds ratio for Z, θ Z c 374 BIO 233, Spring 2015
101 Weak confounder/exposure association: φ XZ = 1.20 Percentage difference between θ X m and θx c θ X c = 0.20 θ X c = 0.50 θ X c = 0.67 θ X c = 1.00 θ X c = 1.50 θ X c = 2.00 θ X c = Conditional odds ratio for Z, θ Z c 375 BIO 233, Spring 2015
102 No confounder/exposure association: φ XZ = 1.00 Percentage difference between θ X m and θx c θ X c = 0.20 θ X c = 0.50 θ X c = 0.67 θ X c = 1.00 θ X c = 1.50 θ X c = 2.00 θ X c = Conditional odds ratio for Z, θ Z c 376 BIO 233, Spring 2015
103 As with linear regression, confounding bias may lead to marginal contrasts that are either bigger or smaller than the conditional contrast the true association may be of the opposite sign to the estimated association depends on whether or not the sign of θ c z and φ XZ are the same or opposite The magnitude of confounding bias depends on an interplay between θ c x, θ c z and φ XZ If φ XZ = 0, then θx m may not equal θx c i.e. Z is precision variable this difference is not confounding bias it is due to the non-collapsibility of the odds ratio 377 BIO 233, Spring 2015
104 In contrast to linear regression, if Z is a precision variable then θ m x and θ c x have different interpretations different numerical values Q: How does one choose between the target parameters? 378 BIO 233, Spring 2015
105 Stratified designs So far, we ve considered estimation and inference based on an independent sample of size n, {(X i,y i ); i = 1,...,n} and the likelihood: L = n P(Y i X i ) i=1 parameterize P(Y X) in terms of a regression model, µ = E[Y X;β] learn about the regression coefficients, β Prospective sampling: choose individuals on the basis of their covariates and observe their outcomes Y is random, conditional on X 379 BIO 233, Spring 2015
106 Cross-sectional sampling: choose individuals completely at random and observe their outcomes/ covariates (Y,X) are jointly random, so that the likelihood is L = = n P(Y i,x i ) i=1 n P(Y i X i )P(X i ) i=1 assume that the marginal covariate distribution does not provide information about the prospective association(s) base estimation/inference on L = n P(Y i X i ) i=1 380 BIO 233, Spring 2015
107 In many settings, these sampling schemes are perfectly reasonable However there are settings where we may need a surprisingly large sample size to have reasonable power King County birth weight data: examine power to detect an association between lbw and welfare based on the logistic model: lbw ~ welfare + married + college + age + smoker + wpre use simulation to estimate power under a range of scenarios odds ratio: 1.5, 2.0, and 3.0 sample size: 3,000 8,000 Homework #6 381 BIO 233, Spring 2015
108 Power for the welfare effect Sample size, n with a sample size of n=8,000, we would have an estimated 67% power to detect an odds ratio of BIO 233, Spring 2015
109 That the outcome is rare is a key reason why power is so low incidence of 5.1% in the observed sample controlled in the simulation by manipulating the value β 0 As we draw random samples, we get very few LBW events see the direct impact on the standard error for the odds ratio association between a binary X and binary outcome Y se[ θ] = θ 1 n n n n 11. Q: What happens if we increase the incidence? 383 BIO 233, Spring 2015
110 Repeat simulations for the association between welfare and lbw manipulate β 0 such that the incidence increases from 0.05 to 0.20 fix the sample size at n=4,000 estimated power based on a Wald test: Odds ratio Incidence as incidence increases power increases rate of increase is not dramatic because the exposure of interest (welfare) is also rare 384 BIO 233, Spring 2015
111 In practice, of course, we cannot manipulate incidence But we can manipulate the (relative) number of cases and non-cases that we observe in the data i.e., artificially inflate the observed incidence for example, via a case-control design The problem is that the sample is no longer representative of the target population the sample is non-random But this non-randomness is by design under the control of the researcher such designs referred to as biased sampling schemes use statistical techniques to account for the non-random sampling 385 BIO 233, Spring 2015
112 Case-control studies In a case-control study, we initially stratify the population by outcome status know Y =0/1 for everyone for any given individual, we can (easily) determine Y Proceed by sampling, at random, n 1 cases, i.e. for whom Y = 1 n 0 non-cases or controls, i.e. for whom Y = 0 For all n=n 0 +n 1 sampled individuals, observe the value of their covariates crucial: X is random and not Y 386 BIO 233, Spring 2015
113 The appropriate likelihood is L R = = n P(X i Y i ) i=1 n 0 i=1 P(X i Y i = 0) n 0 +n 1 i=n 0 +1 P(X i Y i = 1) n independent, outcome-specific contributions retrospective likelihood However, the scientific goal is (most often) to learn about prospective associations i.e., P(Y X) Q: How do we learn about prospective associations from the retrospective likelihood? 387 BIO 233, Spring 2015
114 Consider the logistic regression model: logit P(Y = 1 X) = X T β model corresponds the target population of interest As we ve noted, case-control sampling is non-random with respect to the target population Formalize this by introducing a random variable S that indicates selection by the sampling scheme S = 1 selected 0 not selected binary random variable with some probability, P(S = 1) 388 BIO 233, Spring 2015
115 Cross-sectional sampling selection is independent of (Y, X) P(S = 1) is constant Prospective sampling selection depends on the covariate values, X write P(S = 1 X) Case-control sampling selection depends on outcome status, Y write P(S = 1 Y = y) 389 BIO 233, Spring 2015
116 Now consider the distribution of the outcome, conditional on being selected: P(Y = 1 X,S = 1) 390 BIO 233, Spring 2015
117 Using Bayes Theorem and noting that selection depends solely on Y: P(Y = 1 X,S = 1) = P(S = 1 X,Y = 1) P(Y = 1 X) P(S = 1 X) = P(S = 1 X,Y = 1) P(Y = 1 X) 1 P(S = 1 X,Y = y) P(Y = y X) y=0 = = P(S = 1 Y = 1) P(Y = 1 X) 1 P(S = 1 Y = y) P(Y = y X) y=0 π 1 P(Y = 1 X) 1 π y P(Y = y X) y=0 391 BIO 233, Spring 2015
118 dividing the numerator and denominator by: π 0 P(Y = 0 X) = π 1 π 0 exp{x T β} 1+ π 1 π 0 exp{x T β} = exp{β 0 + β 1 X β k X K } 1+exp{β 0 + β 1X β k X K } where β 0 = β 0 + log ( π1 π 0 ) 392 BIO 233, Spring 2015
119 We see that P(Y = 1 X,S = 1) has the same functional form as the desired logistic regression model if P(Y = 1 X) is of logistic form then so is P(Y = 1 X,S = 1) The odds ratio relationships between X and Y are preserved despite the selection process in Homework #5, we saw that bias (for odds ratios) only arises when selection depends on both Y and X The intercept for the two logistic models are different, however 393 BIO 233, Spring 2015
120 All this suggests that, if the primary goal is to learn about odds ratio parameters, estimation/inference could proceed by forming a likelihood using these probabilities: L P = n P(Y i X i,s i = 1) i=1 ignores the fact that the sample was obtained via a case-control scheme i.e., pretend that the sample was obtained prospectively Use L P to learn about {β 0,β 1,...,β K } In principle, we can also learn about the intercept, β 0, if we have information on the probabilities of selection π 0 and π 1 β 0 = β 0 log ( π1 π 0 ) 394 BIO 233, Spring 2015
121 While this seems reasonable, showing that P(Y = 1 X,S = 1) and P(Y = 1 X) have the same functional form is not sufficient Recall the retrospective likelihood: L R = n P(X i Y i ) = i=1 = n P(X i Y i,s i = 1) i=1 n i=1 P(Y i X i,s i = 1) P(X i S i = 1) P(Y i S i = 1) the components of L P correspond to the first component of L R but ignores the other terms Crucially, the P(Y i X i,s i = 1) contributions are not independent of each other as is assumed by L P 395 BIO 233, Spring 2015
122 The true joint distribution of the outcomes {Y 1,...,Y n } is constrained by the sampling scheme the case-control sampling scheme dictates that there will be n 0 controls and n 1 cases so the {Y 1,...,Y n } cannot freely vary To see this more formally, note that L R = n 0 i=1 P(X i Y i = 0) n 0 +n 1 i=n 0 +1 P(X i Y i = 1) = n 0 i=1 P(Y i = 0 X i,s i = 1) P(X i S i = 1) P(Y i = 0 S i = 1) n 0 +n 1 i=n 0 +1 P(Y i = 1 X i,s i = 1) P(X i S i = 1) P(Y i = 1 S i = 1) 396 BIO 233, Spring 2015
Linear Regression Models P8111
 Linear Regression Models P8111 Lecture 25 Jeff Goldsmith April 26, 2016 1 of 37 Today s Lecture Logistic regression / GLMs Model framework Interpretation Estimation 2 of 37 Linear regression Course started
Linear Regression Models P8111 Lecture 25 Jeff Goldsmith April 26, 2016 1 of 37 Today s Lecture Logistic regression / GLMs Model framework Interpretation Estimation 2 of 37 Linear regression Course started
Generalized Linear Models. Last time: Background & motivation for moving beyond linear
 Generalized Linear Models Last time: Background & motivation for moving beyond linear regression - non-normal/non-linear cases, binary, categorical data Today s class: 1. Examples of count and ordered
Generalized Linear Models Last time: Background & motivation for moving beyond linear regression - non-normal/non-linear cases, binary, categorical data Today s class: 1. Examples of count and ordered
Lecture 12: Effect modification, and confounding in logistic regression
 Lecture 12: Effect modification, and confounding in logistic regression Ani Manichaikul amanicha@jhsph.edu 4 May 2007 Today Categorical predictor create dummy variables just like for linear regression
Lecture 12: Effect modification, and confounding in logistic regression Ani Manichaikul amanicha@jhsph.edu 4 May 2007 Today Categorical predictor create dummy variables just like for linear regression
Model Selection in GLMs. (should be able to implement frequentist GLM analyses!) Today: standard frequentist methods for model selection
 Model Selection in GLMs Last class: estimability/identifiability, analysis of deviance, standard errors & confidence intervals (should be able to implement frequentist GLM analyses!) Today: standard frequentist
Model Selection in GLMs Last class: estimability/identifiability, analysis of deviance, standard errors & confidence intervals (should be able to implement frequentist GLM analyses!) Today: standard frequentist
Generalized Linear Models
 Generalized Linear Models Advanced Methods for Data Analysis (36-402/36-608 Spring 2014 1 Generalized linear models 1.1 Introduction: two regressions So far we ve seen two canonical settings for regression.
Generalized Linear Models Advanced Methods for Data Analysis (36-402/36-608 Spring 2014 1 Generalized linear models 1.1 Introduction: two regressions So far we ve seen two canonical settings for regression.
Logistic Regression. James H. Steiger. Department of Psychology and Human Development Vanderbilt University
 Logistic Regression James H. Steiger Department of Psychology and Human Development Vanderbilt University James H. Steiger (Vanderbilt University) Logistic Regression 1 / 38 Logistic Regression 1 Introduction
Logistic Regression James H. Steiger Department of Psychology and Human Development Vanderbilt University James H. Steiger (Vanderbilt University) Logistic Regression 1 / 38 Logistic Regression 1 Introduction
SCHOOL OF MATHEMATICS AND STATISTICS. Linear and Generalised Linear Models
 SCHOOL OF MATHEMATICS AND STATISTICS Linear and Generalised Linear Models Autumn Semester 2017 18 2 hours Attempt all the questions. The allocation of marks is shown in brackets. RESTRICTED OPEN BOOK EXAMINATION
SCHOOL OF MATHEMATICS AND STATISTICS Linear and Generalised Linear Models Autumn Semester 2017 18 2 hours Attempt all the questions. The allocation of marks is shown in brackets. RESTRICTED OPEN BOOK EXAMINATION
EPSY 905: Fundamentals of Multivariate Modeling Online Lecture #7
 Introduction to Generalized Univariate Models: Models for Binary Outcomes EPSY 905: Fundamentals of Multivariate Modeling Online Lecture #7 EPSY 905: Intro to Generalized In This Lecture A short review
Introduction to Generalized Univariate Models: Models for Binary Outcomes EPSY 905: Fundamentals of Multivariate Modeling Online Lecture #7 EPSY 905: Intro to Generalized In This Lecture A short review
Lecture 5: LDA and Logistic Regression
 Lecture 5: and Logistic Regression Hao Helen Zhang Hao Helen Zhang Lecture 5: and Logistic Regression 1 / 39 Outline Linear Classification Methods Two Popular Linear Models for Classification Linear Discriminant
Lecture 5: and Logistic Regression Hao Helen Zhang Hao Helen Zhang Lecture 5: and Logistic Regression 1 / 39 Outline Linear Classification Methods Two Popular Linear Models for Classification Linear Discriminant
Truck prices - linear model? Truck prices - log transform of the response variable. Interpreting models with log transformation
 Background Regression so far... Lecture 23 - Sta 111 Colin Rundel June 17, 2014 At this point we have covered: Simple linear regression Relationship between numerical response and a numerical or categorical
Background Regression so far... Lecture 23 - Sta 111 Colin Rundel June 17, 2014 At this point we have covered: Simple linear regression Relationship between numerical response and a numerical or categorical
STA216: Generalized Linear Models. Lecture 1. Review and Introduction
 STA216: Generalized Linear Models Lecture 1. Review and Introduction Let y 1,..., y n denote n independent observations on a response Treat y i as a realization of a random variable Y i In the general
STA216: Generalized Linear Models Lecture 1. Review and Introduction Let y 1,..., y n denote n independent observations on a response Treat y i as a realization of a random variable Y i In the general
Regression so far... Lecture 21 - Logistic Regression. Odds. Recap of what you should know how to do... At this point we have covered: Sta102 / BME102
 Background Regression so far... Lecture 21 - Sta102 / BME102 Colin Rundel November 18, 2014 At this point we have covered: Simple linear regression Relationship between numerical response and a numerical
Background Regression so far... Lecture 21 - Sta102 / BME102 Colin Rundel November 18, 2014 At this point we have covered: Simple linear regression Relationship between numerical response and a numerical
Figure 36: Respiratory infection versus time for the first 49 children.
 y BINARY DATA MODELS We devote an entire chapter to binary data since such data are challenging, both in terms of modeling the dependence, and parameter interpretation. We again consider mixed effects
y BINARY DATA MODELS We devote an entire chapter to binary data since such data are challenging, both in terms of modeling the dependence, and parameter interpretation. We again consider mixed effects
Bayesian Inference in GLMs. Frequentists typically base inferences on MLEs, asymptotic confidence
 Bayesian Inference in GLMs Frequentists typically base inferences on MLEs, asymptotic confidence limits, and log-likelihood ratio tests Bayesians base inferences on the posterior distribution of the unknowns
Bayesian Inference in GLMs Frequentists typically base inferences on MLEs, asymptotic confidence limits, and log-likelihood ratio tests Bayesians base inferences on the posterior distribution of the unknowns
Generalized linear models
 Generalized linear models Douglas Bates November 01, 2010 Contents 1 Definition 1 2 Links 2 3 Estimating parameters 5 4 Example 6 5 Model building 8 6 Conclusions 8 7 Summary 9 1 Generalized Linear Models
Generalized linear models Douglas Bates November 01, 2010 Contents 1 Definition 1 2 Links 2 3 Estimating parameters 5 4 Example 6 5 Model building 8 6 Conclusions 8 7 Summary 9 1 Generalized Linear Models
Lecture 14: Introduction to Poisson Regression
 Lecture 14: Introduction to Poisson Regression Ani Manichaikul amanicha@jhsph.edu 8 May 2007 1 / 52 Overview Modelling counts Contingency tables Poisson regression models 2 / 52 Modelling counts I Why
Lecture 14: Introduction to Poisson Regression Ani Manichaikul amanicha@jhsph.edu 8 May 2007 1 / 52 Overview Modelling counts Contingency tables Poisson regression models 2 / 52 Modelling counts I Why
Modelling counts. Lecture 14: Introduction to Poisson Regression. Overview
 Modelling counts I Lecture 14: Introduction to Poisson Regression Ani Manichaikul amanicha@jhsph.edu Why count data? Number of traffic accidents per day Mortality counts in a given neighborhood, per week
Modelling counts I Lecture 14: Introduction to Poisson Regression Ani Manichaikul amanicha@jhsph.edu Why count data? Number of traffic accidents per day Mortality counts in a given neighborhood, per week
MS&E 226: Small Data
 MS&E 226: Small Data Lecture 12: Logistic regression (v1) Ramesh Johari ramesh.johari@stanford.edu Fall 2015 1 / 30 Regression methods for binary outcomes 2 / 30 Binary outcomes For the duration of this
MS&E 226: Small Data Lecture 12: Logistic regression (v1) Ramesh Johari ramesh.johari@stanford.edu Fall 2015 1 / 30 Regression methods for binary outcomes 2 / 30 Binary outcomes For the duration of this
Clinical Trials. Olli Saarela. September 18, Dalla Lana School of Public Health University of Toronto.
 Introduction to Dalla Lana School of Public Health University of Toronto olli.saarela@utoronto.ca September 18, 2014 38-1 : a review 38-2 Evidence Ideal: to advance the knowledge-base of clinical medicine,
Introduction to Dalla Lana School of Public Health University of Toronto olli.saarela@utoronto.ca September 18, 2014 38-1 : a review 38-2 Evidence Ideal: to advance the knowledge-base of clinical medicine,
MS&E 226: Small Data
 MS&E 226: Small Data Lecture 9: Logistic regression (v2) Ramesh Johari ramesh.johari@stanford.edu 1 / 28 Regression methods for binary outcomes 2 / 28 Binary outcomes For the duration of this lecture suppose
MS&E 226: Small Data Lecture 9: Logistic regression (v2) Ramesh Johari ramesh.johari@stanford.edu 1 / 28 Regression methods for binary outcomes 2 / 28 Binary outcomes For the duration of this lecture suppose
STA6938-Logistic Regression Model
 Dr. Ying Zhang STA6938-Logistic Regression Model Topic 2-Multiple Logistic Regression Model Outlines:. Model Fitting 2. Statistical Inference for Multiple Logistic Regression Model 3. Interpretation of
Dr. Ying Zhang STA6938-Logistic Regression Model Topic 2-Multiple Logistic Regression Model Outlines:. Model Fitting 2. Statistical Inference for Multiple Logistic Regression Model 3. Interpretation of
Generalized linear models for binary data. A better graphical exploratory data analysis. The simple linear logistic regression model
 Stat 3302 (Spring 2017) Peter F. Craigmile Simple linear logistic regression (part 1) [Dobson and Barnett, 2008, Sections 7.1 7.3] Generalized linear models for binary data Beetles dose-response example
Stat 3302 (Spring 2017) Peter F. Craigmile Simple linear logistic regression (part 1) [Dobson and Barnett, 2008, Sections 7.1 7.3] Generalized linear models for binary data Beetles dose-response example
Classification. Chapter Introduction. 6.2 The Bayes classifier
 Chapter 6 Classification 6.1 Introduction Often encountered in applications is the situation where the response variable Y takes values in a finite set of labels. For example, the response Y could encode
Chapter 6 Classification 6.1 Introduction Often encountered in applications is the situation where the response variable Y takes values in a finite set of labels. For example, the response Y could encode
UNIVERSITY OF TORONTO Faculty of Arts and Science
 UNIVERSITY OF TORONTO Faculty of Arts and Science December 2013 Final Examination STA442H1F/2101HF Methods of Applied Statistics Jerry Brunner Duration - 3 hours Aids: Calculator Model(s): Any calculator
UNIVERSITY OF TORONTO Faculty of Arts and Science December 2013 Final Examination STA442H1F/2101HF Methods of Applied Statistics Jerry Brunner Duration - 3 hours Aids: Calculator Model(s): Any calculator
STAT 526 Spring Midterm 1. Wednesday February 2, 2011
 STAT 526 Spring 2011 Midterm 1 Wednesday February 2, 2011 Time: 2 hours Name (please print): Show all your work and calculations. Partial credit will be given for work that is partially correct. Points
STAT 526 Spring 2011 Midterm 1 Wednesday February 2, 2011 Time: 2 hours Name (please print): Show all your work and calculations. Partial credit will be given for work that is partially correct. Points
Generalized Linear Models. Kurt Hornik
 Generalized Linear Models Kurt Hornik Motivation Assuming normality, the linear model y = Xβ + e has y = β + ε, ε N(0, σ 2 ) such that y N(μ, σ 2 ), E(y ) = μ = β. Various generalizations, including general
Generalized Linear Models Kurt Hornik Motivation Assuming normality, the linear model y = Xβ + e has y = β + ε, ε N(0, σ 2 ) such that y N(μ, σ 2 ), E(y ) = μ = β. Various generalizations, including general
Various Issues in Fitting Contingency Tables
 Various Issues in Fitting Contingency Tables Statistics 149 Spring 2006 Copyright 2006 by Mark E. Irwin Complete Tables with Zero Entries In contingency tables, it is possible to have zero entries in a
Various Issues in Fitting Contingency Tables Statistics 149 Spring 2006 Copyright 2006 by Mark E. Irwin Complete Tables with Zero Entries In contingency tables, it is possible to have zero entries in a
Logistic Regressions. Stat 430
 Logistic Regressions Stat 430 Final Project Final Project is, again, team based You will decide on a project - only constraint is: you are supposed to use techniques for a solution that are related to
Logistic Regressions Stat 430 Final Project Final Project is, again, team based You will decide on a project - only constraint is: you are supposed to use techniques for a solution that are related to
Introduction to the Analysis of Tabular Data
 Introduction to the Analysis of Tabular Data Anthropological Sciences 192/292 Data Analysis in the Anthropological Sciences James Holland Jones & Ian G. Robertson March 15, 2006 1 Tabular Data Is there
Introduction to the Analysis of Tabular Data Anthropological Sciences 192/292 Data Analysis in the Anthropological Sciences James Holland Jones & Ian G. Robertson March 15, 2006 1 Tabular Data Is there
Exam Applied Statistical Regression. Good Luck!
 Dr. M. Dettling Summer 2011 Exam Applied Statistical Regression Approved: Tables: Note: Any written material, calculator (without communication facility). Attached. All tests have to be done at the 5%-level.
Dr. M. Dettling Summer 2011 Exam Applied Statistical Regression Approved: Tables: Note: Any written material, calculator (without communication facility). Attached. All tests have to be done at the 5%-level.
Semiparametric Generalized Linear Models
 Semiparametric Generalized Linear Models North American Stata Users Group Meeting Chicago, Illinois Paul Rathouz Department of Health Studies University of Chicago prathouz@uchicago.edu Liping Gao MS Student
Semiparametric Generalized Linear Models North American Stata Users Group Meeting Chicago, Illinois Paul Rathouz Department of Health Studies University of Chicago prathouz@uchicago.edu Liping Gao MS Student
Stat 5101 Lecture Notes
 Stat 5101 Lecture Notes Charles J. Geyer Copyright 1998, 1999, 2000, 2001 by Charles J. Geyer May 7, 2001 ii Stat 5101 (Geyer) Course Notes Contents 1 Random Variables and Change of Variables 1 1.1 Random
Stat 5101 Lecture Notes Charles J. Geyer Copyright 1998, 1999, 2000, 2001 by Charles J. Geyer May 7, 2001 ii Stat 5101 (Geyer) Course Notes Contents 1 Random Variables and Change of Variables 1 1.1 Random
Administration. Homework 1 on web page, due Feb 11 NSERC summer undergraduate award applications due Feb 5 Some helpful books
 STA 44/04 Jan 6, 00 / 5 Administration Homework on web page, due Feb NSERC summer undergraduate award applications due Feb 5 Some helpful books STA 44/04 Jan 6, 00... administration / 5 STA 44/04 Jan 6,
STA 44/04 Jan 6, 00 / 5 Administration Homework on web page, due Feb NSERC summer undergraduate award applications due Feb 5 Some helpful books STA 44/04 Jan 6, 00... administration / 5 STA 44/04 Jan 6,
22s:152 Applied Linear Regression. Example: Study on lead levels in children. Ch. 14 (sec. 1) and Ch. 15 (sec. 1 & 4): Logistic Regression
 22s:52 Applied Linear Regression Ch. 4 (sec. and Ch. 5 (sec. & 4: Logistic Regression Logistic Regression When the response variable is a binary variable, such as 0 or live or die fail or succeed then
22s:52 Applied Linear Regression Ch. 4 (sec. and Ch. 5 (sec. & 4: Logistic Regression Logistic Regression When the response variable is a binary variable, such as 0 or live or die fail or succeed then
Logistic Regression - problem 6.14
 Logistic Regression - problem 6.14 Let x 1, x 2,, x m be given values of an input variable x and let Y 1,, Y m be independent binomial random variables whose distributions depend on the corresponding values
Logistic Regression - problem 6.14 Let x 1, x 2,, x m be given values of an input variable x and let Y 1,, Y m be independent binomial random variables whose distributions depend on the corresponding values
Linear Regression. Data Model. β, σ 2. Process Model. ,V β. ,s 2. s 1. Parameter Model
 Regression: Part II Linear Regression y~n X, 2 X Y Data Model β, σ 2 Process Model Β 0,V β s 1,s 2 Parameter Model Assumptions of Linear Model Homoskedasticity No error in X variables Error in Y variables
Regression: Part II Linear Regression y~n X, 2 X Y Data Model β, σ 2 Process Model Β 0,V β s 1,s 2 Parameter Model Assumptions of Linear Model Homoskedasticity No error in X variables Error in Y variables
1. Hypothesis testing through analysis of deviance. 3. Model & variable selection - stepwise aproaches
 Sta 216, Lecture 4 Last Time: Logistic regression example, existence/uniqueness of MLEs Today s Class: 1. Hypothesis testing through analysis of deviance 2. Standard errors & confidence intervals 3. Model
Sta 216, Lecture 4 Last Time: Logistic regression example, existence/uniqueness of MLEs Today s Class: 1. Hypothesis testing through analysis of deviance 2. Standard errors & confidence intervals 3. Model
STA 4504/5503 Sample Exam 1 Spring 2011 Categorical Data Analysis. 1. Indicate whether each of the following is true (T) or false (F).
 STA 4504/5503 Sample Exam 1 Spring 2011 Categorical Data Analysis 1. Indicate whether each of the following is true (T) or false (F). (a) T In 2 2 tables, statistical independence is equivalent to a population
STA 4504/5503 Sample Exam 1 Spring 2011 Categorical Data Analysis 1. Indicate whether each of the following is true (T) or false (F). (a) T In 2 2 tables, statistical independence is equivalent to a population
STA 216: GENERALIZED LINEAR MODELS. Lecture 1. Review and Introduction. Much of statistics is based on the assumption that random
 STA 216: GENERALIZED LINEAR MODELS Lecture 1. Review and Introduction Much of statistics is based on the assumption that random variables are continuous & normally distributed. Normal linear regression
STA 216: GENERALIZED LINEAR MODELS Lecture 1. Review and Introduction Much of statistics is based on the assumption that random variables are continuous & normally distributed. Normal linear regression
Introduction to General and Generalized Linear Models
 Introduction to General and Generalized Linear Models Generalized Linear Models - part III Henrik Madsen Poul Thyregod Informatics and Mathematical Modelling Technical University of Denmark DK-2800 Kgs.
Introduction to General and Generalized Linear Models Generalized Linear Models - part III Henrik Madsen Poul Thyregod Informatics and Mathematical Modelling Technical University of Denmark DK-2800 Kgs.
Statistical Methods III Statistics 212. Problem Set 2 - Answer Key
 Statistical Methods III Statistics 212 Problem Set 2 - Answer Key 1. (Analysis to be turned in and discussed on Tuesday, April 24th) The data for this problem are taken from long-term followup of 1423
Statistical Methods III Statistics 212 Problem Set 2 - Answer Key 1. (Analysis to be turned in and discussed on Tuesday, April 24th) The data for this problem are taken from long-term followup of 1423
ECLT 5810 Linear Regression and Logistic Regression for Classification. Prof. Wai Lam
 ECLT 5810 Linear Regression and Logistic Regression for Classification Prof. Wai Lam Linear Regression Models Least Squares Input vectors is an attribute / feature / predictor (independent variable) The
ECLT 5810 Linear Regression and Logistic Regression for Classification Prof. Wai Lam Linear Regression Models Least Squares Input vectors is an attribute / feature / predictor (independent variable) The
Logistic Regression and Generalized Linear Models
 Logistic Regression and Generalized Linear Models Sridhar Mahadevan mahadeva@cs.umass.edu University of Massachusetts Sridhar Mahadevan: CMPSCI 689 p. 1/2 Topics Generative vs. Discriminative models In
Logistic Regression and Generalized Linear Models Sridhar Mahadevan mahadeva@cs.umass.edu University of Massachusetts Sridhar Mahadevan: CMPSCI 689 p. 1/2 Topics Generative vs. Discriminative models In
Sparse Linear Models (10/7/13)
 STA56: Probabilistic machine learning Sparse Linear Models (0/7/) Lecturer: Barbara Engelhardt Scribes: Jiaji Huang, Xin Jiang, Albert Oh Sparsity Sparsity has been a hot topic in statistics and machine
STA56: Probabilistic machine learning Sparse Linear Models (0/7/) Lecturer: Barbara Engelhardt Scribes: Jiaji Huang, Xin Jiang, Albert Oh Sparsity Sparsity has been a hot topic in statistics and machine
Chapter 11. Regression with a Binary Dependent Variable
 Chapter 11 Regression with a Binary Dependent Variable 2 Regression with a Binary Dependent Variable (SW Chapter 11) So far the dependent variable (Y) has been continuous: district-wide average test score
Chapter 11 Regression with a Binary Dependent Variable 2 Regression with a Binary Dependent Variable (SW Chapter 11) So far the dependent variable (Y) has been continuous: district-wide average test score
Generalized Linear Models 1
 Generalized Linear Models 1 STA 2101/442: Fall 2012 1 See last slide for copyright information. 1 / 24 Suggested Reading: Davison s Statistical models Exponential families of distributions Sec. 5.2 Chapter
Generalized Linear Models 1 STA 2101/442: Fall 2012 1 See last slide for copyright information. 1 / 24 Suggested Reading: Davison s Statistical models Exponential families of distributions Sec. 5.2 Chapter
Hierarchical Generalized Linear Models. ERSH 8990 REMS Seminar on HLM Last Lecture!
 Hierarchical Generalized Linear Models ERSH 8990 REMS Seminar on HLM Last Lecture! Hierarchical Generalized Linear Models Introduction to generalized models Models for binary outcomes Interpreting parameter
Hierarchical Generalized Linear Models ERSH 8990 REMS Seminar on HLM Last Lecture! Hierarchical Generalized Linear Models Introduction to generalized models Models for binary outcomes Interpreting parameter
Introduction to logistic regression
 Introduction to logistic regression Tuan V. Nguyen Professor and NHMRC Senior Research Fellow Garvan Institute of Medical Research University of New South Wales Sydney, Australia What we are going to learn
Introduction to logistic regression Tuan V. Nguyen Professor and NHMRC Senior Research Fellow Garvan Institute of Medical Research University of New South Wales Sydney, Australia What we are going to learn
Machine Learning Linear Classification. Prof. Matteo Matteucci
 Machine Learning Linear Classification Prof. Matteo Matteucci Recall from the first lecture 2 X R p Regression Y R Continuous Output X R p Y {Ω 0, Ω 1,, Ω K } Classification Discrete Output X R p Y (X)
Machine Learning Linear Classification Prof. Matteo Matteucci Recall from the first lecture 2 X R p Regression Y R Continuous Output X R p Y {Ω 0, Ω 1,, Ω K } Classification Discrete Output X R p Y (X)
Previous lecture. P-value based combination. Fixed vs random effects models. Meta vs. pooled- analysis. New random effects testing.
 Previous lecture P-value based combination. Fixed vs random effects models. Meta vs. pooled- analysis. New random effects testing. Interaction Outline: Definition of interaction Additive versus multiplicative
Previous lecture P-value based combination. Fixed vs random effects models. Meta vs. pooled- analysis. New random effects testing. Interaction Outline: Definition of interaction Additive versus multiplicative
9 Generalized Linear Models
 9 Generalized Linear Models The Generalized Linear Model (GLM) is a model which has been built to include a wide range of different models you already know, e.g. ANOVA and multiple linear regression models
9 Generalized Linear Models The Generalized Linear Model (GLM) is a model which has been built to include a wide range of different models you already know, e.g. ANOVA and multiple linear regression models
Review. Timothy Hanson. Department of Statistics, University of South Carolina. Stat 770: Categorical Data Analysis
 Review Timothy Hanson Department of Statistics, University of South Carolina Stat 770: Categorical Data Analysis 1 / 22 Chapter 1: background Nominal, ordinal, interval data. Distributions: Poisson, binomial,
Review Timothy Hanson Department of Statistics, University of South Carolina Stat 770: Categorical Data Analysis 1 / 22 Chapter 1: background Nominal, ordinal, interval data. Distributions: Poisson, binomial,
Single-level Models for Binary Responses
 Single-level Models for Binary Responses Distribution of Binary Data y i response for individual i (i = 1,..., n), coded 0 or 1 Denote by r the number in the sample with y = 1 Mean and variance E(y) =
Single-level Models for Binary Responses Distribution of Binary Data y i response for individual i (i = 1,..., n), coded 0 or 1 Denote by r the number in the sample with y = 1 Mean and variance E(y) =
Linear Regression. In this lecture we will study a particular type of regression model: the linear regression model
 1 Linear Regression 2 Linear Regression In this lecture we will study a particular type of regression model: the linear regression model We will first consider the case of the model with one predictor
1 Linear Regression 2 Linear Regression In this lecture we will study a particular type of regression model: the linear regression model We will first consider the case of the model with one predictor
Econometrics II. Seppo Pynnönen. Spring Department of Mathematics and Statistics, University of Vaasa, Finland
 Department of Mathematics and Statistics, University of Vaasa, Finland Spring 2018 Part III Limited Dependent Variable Models As of Jan 30, 2017 1 Background 2 Binary Dependent Variable The Linear Probability
Department of Mathematics and Statistics, University of Vaasa, Finland Spring 2018 Part III Limited Dependent Variable Models As of Jan 30, 2017 1 Background 2 Binary Dependent Variable The Linear Probability
Part 6: Multivariate Normal and Linear Models
 Part 6: Multivariate Normal and Linear Models 1 Multiple measurements Up until now all of our statistical models have been univariate models models for a single measurement on each member of a sample of
Part 6: Multivariate Normal and Linear Models 1 Multiple measurements Up until now all of our statistical models have been univariate models models for a single measurement on each member of a sample of
Hypothesis Testing. Part I. James J. Heckman University of Chicago. Econ 312 This draft, April 20, 2006
 Hypothesis Testing Part I James J. Heckman University of Chicago Econ 312 This draft, April 20, 2006 1 1 A Brief Review of Hypothesis Testing and Its Uses values and pure significance tests (R.A. Fisher)
Hypothesis Testing Part I James J. Heckman University of Chicago Econ 312 This draft, April 20, 2006 1 1 A Brief Review of Hypothesis Testing and Its Uses values and pure significance tests (R.A. Fisher)
7/28/15. Review Homework. Overview. Lecture 6: Logistic Regression Analysis
 Lecture 6: Logistic Regression Analysis Christopher S. Hollenbeak, PhD Jane R. Schubart, PhD The Outcomes Research Toolbox Review Homework 2 Overview Logistic regression model conceptually Logistic regression
Lecture 6: Logistic Regression Analysis Christopher S. Hollenbeak, PhD Jane R. Schubart, PhD The Outcomes Research Toolbox Review Homework 2 Overview Logistic regression model conceptually Logistic regression
Regression models. Generalized linear models in R. Normal regression models are not always appropriate. Generalized linear models. Examples.
 Regression models Generalized linear models in R Dr Peter K Dunn http://www.usq.edu.au Department of Mathematics and Computing University of Southern Queensland ASC, July 00 The usual linear regression
Regression models Generalized linear models in R Dr Peter K Dunn http://www.usq.edu.au Department of Mathematics and Computing University of Southern Queensland ASC, July 00 The usual linear regression
Hypothesis testing, part 2. With some material from Howard Seltman, Blase Ur, Bilge Mutlu, Vibha Sazawal
 Hypothesis testing, part 2 With some material from Howard Seltman, Blase Ur, Bilge Mutlu, Vibha Sazawal 1 CATEGORICAL IV, NUMERIC DV 2 Independent samples, one IV # Conditions Normal/Parametric Non-parametric
Hypothesis testing, part 2 With some material from Howard Seltman, Blase Ur, Bilge Mutlu, Vibha Sazawal 1 CATEGORICAL IV, NUMERIC DV 2 Independent samples, one IV # Conditions Normal/Parametric Non-parametric
Generalized Linear Models for Non-Normal Data
 Generalized Linear Models for Non-Normal Data Today s Class: 3 parts of a generalized model Models for binary outcomes Complications for generalized multivariate or multilevel models SPLH 861: Lecture
Generalized Linear Models for Non-Normal Data Today s Class: 3 parts of a generalized model Models for binary outcomes Complications for generalized multivariate or multilevel models SPLH 861: Lecture
ST3241 Categorical Data Analysis I Generalized Linear Models. Introduction and Some Examples
 ST3241 Categorical Data Analysis I Generalized Linear Models Introduction and Some Examples 1 Introduction We have discussed methods for analyzing associations in two-way and three-way tables. Now we will
ST3241 Categorical Data Analysis I Generalized Linear Models Introduction and Some Examples 1 Introduction We have discussed methods for analyzing associations in two-way and three-way tables. Now we will
Part 8: GLMs and Hierarchical LMs and GLMs
 Part 8: GLMs and Hierarchical LMs and GLMs 1 Example: Song sparrow reproductive success Arcese et al., (1992) provide data on a sample from a population of 52 female song sparrows studied over the course
Part 8: GLMs and Hierarchical LMs and GLMs 1 Example: Song sparrow reproductive success Arcese et al., (1992) provide data on a sample from a population of 52 female song sparrows studied over the course
ECLT 5810 Linear Regression and Logistic Regression for Classification. Prof. Wai Lam
 ECLT 5810 Linear Regression and Logistic Regression for Classification Prof. Wai Lam Linear Regression Models Least Squares Input vectors is an attribute / feature / predictor (independent variable) The
ECLT 5810 Linear Regression and Logistic Regression for Classification Prof. Wai Lam Linear Regression Models Least Squares Input vectors is an attribute / feature / predictor (independent variable) The
Latent Variable Models for Binary Data. Suppose that for a given vector of explanatory variables x, the latent
 Latent Variable Models for Binary Data Suppose that for a given vector of explanatory variables x, the latent variable, U, has a continuous cumulative distribution function F (u; x) and that the binary
Latent Variable Models for Binary Data Suppose that for a given vector of explanatory variables x, the latent variable, U, has a continuous cumulative distribution function F (u; x) and that the binary
Fall 2017 STAT 532 Homework Peter Hoff. 1. Let P be a probability measure on a collection of sets A.
 1. Let P be a probability measure on a collection of sets A. (a) For each n N, let H n be a set in A such that H n H n+1. Show that P (H n ) monotonically converges to P ( k=1 H k) as n. (b) For each n
1. Let P be a probability measure on a collection of sets A. (a) For each n N, let H n be a set in A such that H n H n+1. Show that P (H n ) monotonically converges to P ( k=1 H k) as n. (b) For each n
Gov 2000: 9. Regression with Two Independent Variables
 Gov 2000: 9. Regression with Two Independent Variables Matthew Blackwell Harvard University mblackwell@gov.harvard.edu Where are we? Where are we going? Last week: we learned about how to calculate a simple
Gov 2000: 9. Regression with Two Independent Variables Matthew Blackwell Harvard University mblackwell@gov.harvard.edu Where are we? Where are we going? Last week: we learned about how to calculate a simple
Simple logistic regression
 Simple logistic regression Biometry 755 Spring 2009 Simple logistic regression p. 1/47 Model assumptions 1. The observed data are independent realizations of a binary response variable Y that follows a
Simple logistic regression Biometry 755 Spring 2009 Simple logistic regression p. 1/47 Model assumptions 1. The observed data are independent realizations of a binary response variable Y that follows a
Marginal versus conditional effects: does it make a difference? Mireille Schnitzer, PhD Université de Montréal
 Marginal versus conditional effects: does it make a difference? Mireille Schnitzer, PhD Université de Montréal Overview In observational and experimental studies, the goal may be to estimate the effect
Marginal versus conditional effects: does it make a difference? Mireille Schnitzer, PhD Université de Montréal Overview In observational and experimental studies, the goal may be to estimate the effect
STA 450/4000 S: January
 STA 450/4000 S: January 6 005 Notes Friday tutorial on R programming reminder office hours on - F; -4 R The book Modern Applied Statistics with S by Venables and Ripley is very useful. Make sure you have
STA 450/4000 S: January 6 005 Notes Friday tutorial on R programming reminder office hours on - F; -4 R The book Modern Applied Statistics with S by Venables and Ripley is very useful. Make sure you have
STA 4504/5503 Sample Exam 1 Spring 2011 Categorical Data Analysis. 1. Indicate whether each of the following is true (T) or false (F).
 STA 4504/5503 Sample Exam 1 Spring 2011 Categorical Data Analysis 1. Indicate whether each of the following is true (T) or false (F). (a) (b) (c) (d) (e) In 2 2 tables, statistical independence is equivalent
STA 4504/5503 Sample Exam 1 Spring 2011 Categorical Data Analysis 1. Indicate whether each of the following is true (T) or false (F). (a) (b) (c) (d) (e) In 2 2 tables, statistical independence is equivalent
Lecture 4 Multiple linear regression
 Lecture 4 Multiple linear regression BIOST 515 January 15, 2004 Outline 1 Motivation for the multiple regression model Multiple regression in matrix notation Least squares estimation of model parameters
Lecture 4 Multiple linear regression BIOST 515 January 15, 2004 Outline 1 Motivation for the multiple regression model Multiple regression in matrix notation Least squares estimation of model parameters
12 Modelling Binomial Response Data
 c 2005, Anthony C. Brooms Statistical Modelling and Data Analysis 12 Modelling Binomial Response Data 12.1 Examples of Binary Response Data Binary response data arise when an observation on an individual
c 2005, Anthony C. Brooms Statistical Modelling and Data Analysis 12 Modelling Binomial Response Data 12.1 Examples of Binary Response Data Binary response data arise when an observation on an individual
Linear model A linear model assumes Y X N(µ(X),σ 2 I), And IE(Y X) = µ(x) = X β, 2/52
 Statistics for Applications Chapter 10: Generalized Linear Models (GLMs) 1/52 Linear model A linear model assumes Y X N(µ(X),σ 2 I), And IE(Y X) = µ(x) = X β, 2/52 Components of a linear model The two
Statistics for Applications Chapter 10: Generalized Linear Models (GLMs) 1/52 Linear model A linear model assumes Y X N(µ(X),σ 2 I), And IE(Y X) = µ(x) = X β, 2/52 Components of a linear model The two
HST.582J / 6.555J / J Biomedical Signal and Image Processing Spring 2007
 MIT OpenCourseWare http://ocw.mit.edu HST.582J / 6.555J / 16.456J Biomedical Signal and Image Processing Spring 2007 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.582J / 6.555J / 16.456J Biomedical Signal and Image Processing Spring 2007 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Neural networks (not in book)
 (not in book) Another approach to classification is neural networks. were developed in the 1980s as a way to model how learning occurs in the brain. There was therefore wide interest in neural networks
(not in book) Another approach to classification is neural networks. were developed in the 1980s as a way to model how learning occurs in the brain. There was therefore wide interest in neural networks
Biostatistics Advanced Methods in Biostatistics IV
 Biostatistics 140.754 Advanced Methods in Biostatistics IV Jeffrey Leek Assistant Professor Department of Biostatistics jleek@jhsph.edu 1 / 35 Tip + Paper Tip Meet with seminar speakers. When you go on
Biostatistics 140.754 Advanced Methods in Biostatistics IV Jeffrey Leek Assistant Professor Department of Biostatistics jleek@jhsph.edu 1 / 35 Tip + Paper Tip Meet with seminar speakers. When you go on
CPSC 340: Machine Learning and Data Mining. MLE and MAP Fall 2017
 CPSC 340: Machine Learning and Data Mining MLE and MAP Fall 2017 Assignment 3: Admin 1 late day to hand in tonight, 2 late days for Wednesday. Assignment 4: Due Friday of next week. Last Time: Multi-Class
CPSC 340: Machine Learning and Data Mining MLE and MAP Fall 2017 Assignment 3: Admin 1 late day to hand in tonight, 2 late days for Wednesday. Assignment 4: Due Friday of next week. Last Time: Multi-Class
Lecture 10: Alternatives to OLS with limited dependent variables. PEA vs APE Logit/Probit Poisson
 Lecture 10: Alternatives to OLS with limited dependent variables PEA vs APE Logit/Probit Poisson PEA vs APE PEA: partial effect at the average The effect of some x on y for a hypothetical case with sample
Lecture 10: Alternatives to OLS with limited dependent variables PEA vs APE Logit/Probit Poisson PEA vs APE PEA: partial effect at the average The effect of some x on y for a hypothetical case with sample
R Hints for Chapter 10
 R Hints for Chapter 10 The multiple logistic regression model assumes that the success probability p for a binomial random variable depends on independent variables or design variables x 1, x 2,, x k.
R Hints for Chapter 10 The multiple logistic regression model assumes that the success probability p for a binomial random variable depends on independent variables or design variables x 1, x 2,, x k.
LOGISTIC REGRESSION Joseph M. Hilbe
 LOGISTIC REGRESSION Joseph M. Hilbe Arizona State University Logistic regression is the most common method used to model binary response data. When the response is binary, it typically takes the form of
LOGISTIC REGRESSION Joseph M. Hilbe Arizona State University Logistic regression is the most common method used to model binary response data. When the response is binary, it typically takes the form of
Proportional hazards regression
 Proportional hazards regression Patrick Breheny October 8 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/28 Introduction The model Solving for the MLE Inference Today we will begin discussing regression
Proportional hazards regression Patrick Breheny October 8 Patrick Breheny Survival Data Analysis (BIOS 7210) 1/28 Introduction The model Solving for the MLE Inference Today we will begin discussing regression
IP WEIGHTING AND MARGINAL STRUCTURAL MODELS (CHAPTER 12) BIOS IPW and MSM
 IP WEIGHTING AND MARGINAL STRUCTURAL MODELS (CHAPTER 12) BIOS 776 1 12 IPW and MSM IP weighting and marginal structural models ( 12) Outline 12.1 The causal question 12.2 Estimating IP weights via modeling
IP WEIGHTING AND MARGINAL STRUCTURAL MODELS (CHAPTER 12) BIOS 776 1 12 IPW and MSM IP weighting and marginal structural models ( 12) Outline 12.1 The causal question 12.2 Estimating IP weights via modeling
Generalized Linear Models. stat 557 Heike Hofmann
 Generalized Linear Models stat 557 Heike Hofmann Outline Intro to GLM Exponential Family Likelihood Equations GLM for Binomial Response Generalized Linear Models Three components: random, systematic, link
Generalized Linear Models stat 557 Heike Hofmann Outline Intro to GLM Exponential Family Likelihood Equations GLM for Binomial Response Generalized Linear Models Three components: random, systematic, link
STA102 Class Notes Chapter Logistic Regression
 STA0 Class Notes Chapter 0 0. Logistic Regression We continue to study the relationship between a response variable and one or more eplanatory variables. For SLR and MLR (Chapters 8 and 9), our response
STA0 Class Notes Chapter 0 0. Logistic Regression We continue to study the relationship between a response variable and one or more eplanatory variables. For SLR and MLR (Chapters 8 and 9), our response
BIOS 312: Precision of Statistical Inference
 and Power/Sample Size and Standard Errors BIOS 312: of Statistical Inference Chris Slaughter Department of Biostatistics, Vanderbilt University School of Medicine January 3, 2013 Outline Overview and Power/Sample
and Power/Sample Size and Standard Errors BIOS 312: of Statistical Inference Chris Slaughter Department of Biostatistics, Vanderbilt University School of Medicine January 3, 2013 Outline Overview and Power/Sample
General Regression Model
 Scott S. Emerson, M.D., Ph.D. Department of Biostatistics, University of Washington, Seattle, WA 98195, USA January 5, 2015 Abstract Regression analysis can be viewed as an extension of two sample statistical
Scott S. Emerson, M.D., Ph.D. Department of Biostatistics, University of Washington, Seattle, WA 98195, USA January 5, 2015 Abstract Regression analysis can be viewed as an extension of two sample statistical
Generalized Estimating Equations
 Outline Review of Generalized Linear Models (GLM) Generalized Linear Model Exponential Family Components of GLM MLE for GLM, Iterative Weighted Least Squares Measuring Goodness of Fit - Deviance and Pearson
Outline Review of Generalized Linear Models (GLM) Generalized Linear Model Exponential Family Components of GLM MLE for GLM, Iterative Weighted Least Squares Measuring Goodness of Fit - Deviance and Pearson
Poisson regression: Further topics
 Poisson regression: Further topics April 21 Overdispersion One of the defining characteristics of Poisson regression is its lack of a scale parameter: E(Y ) = Var(Y ), and no parameter is available to
Poisson regression: Further topics April 21 Overdispersion One of the defining characteristics of Poisson regression is its lack of a scale parameter: E(Y ) = Var(Y ), and no parameter is available to
Statistical Distribution Assumptions of General Linear Models
 Statistical Distribution Assumptions of General Linear Models Applied Multilevel Models for Cross Sectional Data Lecture 4 ICPSR Summer Workshop University of Colorado Boulder Lecture 4: Statistical Distributions
Statistical Distribution Assumptions of General Linear Models Applied Multilevel Models for Cross Sectional Data Lecture 4 ICPSR Summer Workshop University of Colorado Boulder Lecture 4: Statistical Distributions
POLI 8501 Introduction to Maximum Likelihood Estimation
 POLI 8501 Introduction to Maximum Likelihood Estimation Maximum Likelihood Intuition Consider a model that looks like this: Y i N(µ, σ 2 ) So: E(Y ) = µ V ar(y ) = σ 2 Suppose you have some data on Y,
POLI 8501 Introduction to Maximum Likelihood Estimation Maximum Likelihood Intuition Consider a model that looks like this: Y i N(µ, σ 2 ) So: E(Y ) = µ V ar(y ) = σ 2 Suppose you have some data on Y,
Generalized Linear Models Introduction
 Generalized Linear Models Introduction Statistics 135 Autumn 2005 Copyright c 2005 by Mark E. Irwin Generalized Linear Models For many problems, standard linear regression approaches don t work. Sometimes,
Generalized Linear Models Introduction Statistics 135 Autumn 2005 Copyright c 2005 by Mark E. Irwin Generalized Linear Models For many problems, standard linear regression approaches don t work. Sometimes,
Today. HW 1: due February 4, pm. Aspects of Design CD Chapter 2. Continue with Chapter 2 of ELM. In the News:
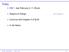 Today HW 1: due February 4, 11.59 pm. Aspects of Design CD Chapter 2 Continue with Chapter 2 of ELM In the News: STA 2201: Applied Statistics II January 14, 2015 1/35 Recap: data on proportions data: y
Today HW 1: due February 4, 11.59 pm. Aspects of Design CD Chapter 2 Continue with Chapter 2 of ELM In the News: STA 2201: Applied Statistics II January 14, 2015 1/35 Recap: data on proportions data: y
Recap. HW due Thursday by 5 pm Next HW coming on Thursday Logistic regression: Pr(G = k X) linear on the logit scale Linear discriminant analysis:
 1 / 23 Recap HW due Thursday by 5 pm Next HW coming on Thursday Logistic regression: Pr(G = k X) linear on the logit scale Linear discriminant analysis: Pr(G = k X) Pr(X G = k)pr(g = k) Theory: LDA more
1 / 23 Recap HW due Thursday by 5 pm Next HW coming on Thursday Logistic regression: Pr(G = k X) linear on the logit scale Linear discriminant analysis: Pr(G = k X) Pr(X G = k)pr(g = k) Theory: LDA more
Ninth ARTNeT Capacity Building Workshop for Trade Research "Trade Flows and Trade Policy Analysis"
 Ninth ARTNeT Capacity Building Workshop for Trade Research "Trade Flows and Trade Policy Analysis" June 2013 Bangkok, Thailand Cosimo Beverelli and Rainer Lanz (World Trade Organization) 1 Selected econometric
Ninth ARTNeT Capacity Building Workshop for Trade Research "Trade Flows and Trade Policy Analysis" June 2013 Bangkok, Thailand Cosimo Beverelli and Rainer Lanz (World Trade Organization) 1 Selected econometric
8 Nominal and Ordinal Logistic Regression
 8 Nominal and Ordinal Logistic Regression 8.1 Introduction If the response variable is categorical, with more then two categories, then there are two options for generalized linear models. One relies on
8 Nominal and Ordinal Logistic Regression 8.1 Introduction If the response variable is categorical, with more then two categories, then there are two options for generalized linear models. One relies on
Multiple Regression Analysis
 Multiple Regression Analysis y = 0 + 1 x 1 + x +... k x k + u 6. Heteroskedasticity What is Heteroskedasticity?! Recall the assumption of homoskedasticity implied that conditional on the explanatory variables,
Multiple Regression Analysis y = 0 + 1 x 1 + x +... k x k + u 6. Heteroskedasticity What is Heteroskedasticity?! Recall the assumption of homoskedasticity implied that conditional on the explanatory variables,
Multiple Regression Analysis
 Multiple Regression Analysis y = β 0 + β 1 x 1 + β 2 x 2 +... β k x k + u 2. Inference 0 Assumptions of the Classical Linear Model (CLM)! So far, we know: 1. The mean and variance of the OLS estimators
Multiple Regression Analysis y = β 0 + β 1 x 1 + β 2 x 2 +... β k x k + u 2. Inference 0 Assumptions of the Classical Linear Model (CLM)! So far, we know: 1. The mean and variance of the OLS estimators
Stat 579: Generalized Linear Models and Extensions
 Stat 579: Generalized Linear Models and Extensions Yan Lu Jan, 2018, week 3 1 / 67 Hypothesis tests Likelihood ratio tests Wald tests Score tests 2 / 67 Generalized Likelihood ratio tests Let Y = (Y 1,
Stat 579: Generalized Linear Models and Extensions Yan Lu Jan, 2018, week 3 1 / 67 Hypothesis tests Likelihood ratio tests Wald tests Score tests 2 / 67 Generalized Likelihood ratio tests Let Y = (Y 1,
Beyond GLM and likelihood
 Stat 6620: Applied Linear Models Department of Statistics Western Michigan University Statistics curriculum Core knowledge (modeling and estimation) Math stat 1 (probability, distributions, convergence
Stat 6620: Applied Linear Models Department of Statistics Western Michigan University Statistics curriculum Core knowledge (modeling and estimation) Math stat 1 (probability, distributions, convergence
