Advanced usage of MOLCAS. Ex. 1. Usage of Symmetry in Molcas
|
|
|
- Cornelius Johnson
- 5 years ago
- Views:
Transcription
1 Advanced usage of MOLCAS Ex 1. Symmetry in Molcas (20 min). Ex 2. Transition state optimization (30 min). Ex 3. Ways to run RASSCF program (30 min). Ex 4. Optional. Using EXPBAS module (20 min) Ex 5. Optional. Using RAS-probing (20 min) Ex 6. Basic CASPT2/RASPT2 calculation: Butadiene (30 min) Ex 7. Basic CASPT2/RASPT2 calculation: Acrolein (20 min) Ex 8. Optional. Visualization of excited state calculations (20 min) Ex 9. If everything above was too trivial... Ex. 1. Usage of Symmetry in Molcas Prepare an input for SCF calculation of water molecule, and run it. Now repeat the calculation but remove(or comment/**/) a line GROUP=C1. The code will determine the symmetry (as a subgroup of D 2h ) and use it. Observe the difference between calculation with and without symmetry. Did you get the same energy, charges, dipole moment? Discuss a possibility when these values can differ! Detect the orbitals which belongs to different irreps. Visualize them! Optionally repeat the excercise for benzene. 1
2 Ex. 2. Transition states and geometry optimization The search for a transition state is more complex in comparison to minimal energy search. Thus, you have to push your system towards to the solution. Prepare the following input: &Gateway coord 3 modified water coordinates O H H basis=ano-s-vdz group=c1 >>> Do while &SCF KSDFT=B3LYP &SLAPAF TS >>> EndDo Note, that coordinates of water are modified! Run the calculation, and use geo.molden file to observe the convergence to the transition state. Optional: Another possibility to search TS is to use constrains in the input: &Gateway coord=water.xyz Basis=ANO-S-VDZ Group=C1 Constraints a = Angle H2 O1 H3 Value a = degree 2
3 End of Constraints >>> Do while &SCF KSDFT=B3LYP &SLAPAF FindTS >>> EndDo Ex. 3. Ways to run RASSCF program Objective: We will learn how to set up CASSCF/RASSCF calculation based on graphically selected active space Create an input for SCF calculation of benzene molecule. coord=benzene.xyz basis=ano-s-vdz group=c1 &SCF ASCII ALL Run it and open the dos.grid file with GV. Detect six π orbitals (What is the difference between σ and π orbitals?), and mark them as active orbitals (use key a for this operation). The header of GV window should say 2:6 (2 is a short name for RAS2, a.k.a. active space). Note that if you did not use keyword ALL in GRID IT, you might be able to find only 5 π orbitals. Save the file by pressing F2. Now you should have a file with extension GvOrb: benzene.dos.gvorb Modify the input file. 3
4 coord=benzene.xyz basis=ano-s-vdz group=c1 fileorb=$currdir/benzene.dos.gvorb Spin=1 Charge=0 ASCII name output Note that $CurrDir is a case sensitive: this is a location of your input file. Do not forget to add to Extra files: Benzene.xyz and benzene.dos.gvorb Run the calculation. Check benzene.output.dos.grid file to verify that orbital space did not change - you should see only 6 π orbitals in the file. If some of the orbitals is a σ character - the calculation converged to a wrong solution. Let s have a quick look at RASSCF output. Identify the number of inactive orbitals and active orbitals. How to compute the total number of electrons? Basis functions Wave function specifications: Number of closed shell electrons 36 Number of electrons in active shells 6 Number of inactive orbitals 18 Number of active orbitals 6 Number of secondary orbitals 42 Identify the number of CSF. Check that the starting point was taken from GvOrb file. Identify convergence, weights of the main configurations, and occupation numbers for active orbitals. printout of CI-coefficients larger than 0.05 for root 1 energy= conf/sym Coeff Weight
5 18 2udud u2d0ud ud2u0d uudd Natural orbitals and occupation numbers for root 1 sym 1: Check MO coefficients for orbitals, and observe that some obrital energy are zero. Optional. Choose another active orbitals, and rerun the calculation. Do you get the same active space at the end? Optional. set up RASSCF calculations. Use 1 and 3 to mark orbitals as belongingtoras1andras3space. AddRASSCFkeyword intorasscf input section, followed by two numbers: maximum amount of holes in RAS1, and maximum amount of electrons in RAS3. Note, that using RAS2 simultaneously with RAS1/3 can become very expensive, and use much more memory. If the code stops, complaining about the allocation of memory add into the input: >> export MOLCASMEM=1000 Example: fileorb=$currdir/benzene.dos.gvorb Spin=1 Charge=0 RASSCF= 1 1 The alternative input to CASSCF could be written in the following form: Inactive=34 Nactel=6 0 0 RAS2=6 Spin=1 5
6 Inactive - the number of doubly occupied orbitals. RAS2 - the number of active orbitals Nactel - number of active electrons, max. number of holes in RAS1, number of electrons in RAS3. Spin - spin multiplicity (1 for singlet) Note! since there is no orbital input, the code will select active space around HOMO-LUMO gap. Ex 4. Optional. Using EXPBAS module Objective: selection of active space is more simple if a small basis set is used. EXPBAS is a module, which convert orbital file, obtained for a small basis set into an orbital file for another basis set. Note, currently this feature works only with ANO type of basis sets. The content of benzene.expbas.input: /* first part of the calculation. Remove this line for the second run */ coord=$molcas/coord/benzene.xyz basis=ano-s-vdz group=c1 &seward &grid_it all; ASCII >>COPY $Project.RunFile $CurrDir/Small.RunFile >>UNIX echo "Please, select 6 active orbitals" >>UNIX molcas gv $Project.grid >>COPY $Project.GvOrb $CurrDir/$Project.GvOrb >>>>>>>>>>> exit <<<<<<<<<<<<<<<<<<< /* Remove this line for the second run */ 6
7 coord=$molcas/coord/benzene.xyz basis=ano-l-vdzp group=c1 >>COPY $CurrDir/Small.RunFile RUNFIL1 >>COPY $CurrDir/$Project.GvOrb INPORB >>COPY $Project.RunFile RUNFIL2 &EXPBAS Spin=1 FileOrb=$Project.ExpOrb The calculation split into two parts. The first one - ANO-S-VDZ calculation of benzene molecule. At the end of this input RunFile is copied into input directory, and GV program is automatically runs. If you run the example on a remote machine, you have to run GV code manually. Using GV code select 6 active orbitals, and save the file (F2). To run the second part of the calculation - comment the first block /* */. (Note, that > EXIT command The second calculation uses larger basis set ANO-L-VDZP. >> COPY command similar to UNIX cp command. EXPBAS needs as an input: old and new RunFiles, and orbital file. It produces.exporb file which is used by RASSCF. Check the log file, and identify starting point for CASSCF calculation. 7
8 Ex 5. Optional. Using RAS-probing Objective: In complicated cases, graphical analysis of orbitals is not conclusive. Small RASSCF calculation can be helpful in identifying orbitals which should be included into active space. We will use a similar to previous example setup: benzene molecule, ANO-S- VDZ basis set. But this time, we will select only RAS1 and RAS3 orbitals (use keys: 1 and 3). We also have to add RASSCF keyword, so we will use only single excitations. And finally, we have to request more memory (MOLCASMEM=1000 means 1Gb of RAM will be used). Please, select not more than 18 orbitals in total. coord=$molcas/coord/benzene.xyz basis=ano-s-vdz group=c1 &seward &grid_it all; ASCII >>UNIX echo "OK. we now select only RAS1 and RAS3 spaces" >>UNIX molcas gv $Project.grid >>COPY $Project.GvOrb $CurrDir/$Project.GvOrb /* note, we need more memory! */ >>export MOLCASMEM=1000 Spin=1 RASSCF=1 1 FileOrb=$Project.GvOrb &grid_it name=ras-probe all; ASCII Observe the output and identify the orbitals which can be included into the active space in following calculations. (Check the occupation numbers first!). 8
9 Ex 6. Basic CASPT2/RASPT2 calculations: Butadiene Objective: Selection of active space depends on the problem, and can vary even for the same system. Note: the input below requires an intermediate run of GV, and so, it can be executed if you have installation of MOLCAS at your computer Input file butadiene.inp COORD = butadiene.xyz BASIS = ANO-S-VDZP GROUP = C1 &SCF ALL; ASCII >>>>>>>> UNIX molcas gv $Project.grid <<<<<<<< >> COPY $Project.GvOrb $CurrDir/$Project.GvOrb FILEORB=$CurrDir/$Project.GvOrb FILEORB=$Project.RasOrb NAME=RASSCF ALL; ASCII &CASPT2 if Molcas farm is used, the input above should be split: First run: COORD = butadiene.xyz BASIS = ANO-S-VDZP 9
10 GROUP = C1 &SCF ALL; ASCII Grid file is modified by GV. Second run: FILEORB=$Project.GvOrb FILEORB=$Project.RasOrb NAME=RASSCF ALL; ASCII &CASPT2 Note, that if running on farm: GvOrb file should be added as extra file CurrDir and WorkDir are the same directories While selecting the active orbitals we can make different decisions. We will identify π shaped orbitals. The choice among occupied orbitals is straightforward (14, 15), but for the virtual orbitals we can find a pair 16 and 20, but also 22 and 26. Output. What should we look at? before we go... Always look at the top part, it contains echo of your input and details of calculations Check the header of GATEWAY module: it tells you - how much memory has been used, was it serial or parallel run. )()()()()()()()()()()()()()()()()()()()()()()()()()()()()()()()()()()()() MOLCAS executing module GATEWAY with 256 MB of memory at 09:29:35 Thu Aug ()()()()()()()()()()()()()()()()()()()()()()()()()()()()()()()()()()()()() 10
11 Check geometry, and basis set information. in RASSCF output Orbital specification Starting orbitals. It is easy to make a mistake here! Charge (especially if you set up input using Nactel keyword) Convergence (problems with convergence might indicate problems with active space) CI-coefficients Natural orbitals and occupation numbers Total energy :: RASSCF root number 1 Total energy = Properties (e.g. do you have symmetry, and reasonable values in Mulliken charges) in CASPT2 output Orbital specification details of calculation Type of Fock operator to use: STANDARD Type of HZERO operator to use: STANDARD IPEA FINAL CASPT2 RESULT: Reference energy: Total energy: Reference weight: Small reference weight and/or large correction to energy might indicate an intruder state Contributions to the CASPT2 correlation energy Active & Virtual Only: One Inactive Excited: Two Inactive Excited: Report on small energy denominators, large coefficients, and large energy contributions Properties Do we need to include all 6 orbitals, or 4 is enough? Try to do both calcula- 11
12 tions, and compare the results. Ex 7. Basic CASPT2/RASPT2 calculations: Acrolein Objective: Set up of multistate CASSCF/CASPT2 calculations In this exercise we will use Acrolein molecule. Step 1. Preparing the orbitals. coord = $MOLCAS/Coord/Acrolein.xyz basis = ANO-S-VDZ; group= c1 &SCF All; ASCII run GV, and select the minimal active space (5 orbitals: 13, 14, 15, 16, 18) Save the orbital file, and rename it to acrolein.gvorb Now, we can run CASPT2 calculation: coord = $MOLCAS/Coord/Acrolein.xyz basis = ANO-S-VDZ; group= c1 Spin= 1 fileorb=$currdir/acrolein.gvorb All; ASCII &CASPT2 Verify that the orbitals after CASSCF calculation are correct. Check the log 12
13 file. Identify Reference energy, CASPT2 energy, and Reference weight. We now perform multi-state CASPT2 calculation for all 5 roots: Spin= 1 fileorb=$currdir/acrolein.gvorb CiRoot= &CASPT2 Multistate= Compare the results of single state CASPT2 and MS-CASPT2. Ex 8. Optional. Visualization of excited state calculations create a file acrolein.ms-cas-grid.input coord = $MOLCAS/Coord/Acrolein.xyz basis = ANO-S-VDZ; group= c1 Spin= 1 fileorb=$currdir/acrolein.gvorb CiRoot= OutOrbital Natural= 5 FILEORB = $Project.RasOrb.1 NAME = root1; All; ASCII FILEORB = $Project.RasOrb.2 NAME = root2; All; ASCII FILEORB = $Project.RasOrb.3 13
14 NAME = root3; All; ASCII FILEORB = $Project.RasOrb.4 NAME = root4; All; ASCII FILEORB = $Project.RasOrb.5 NAME = root5; All; ASCII RASSCF code produces individual orbital files RasOrb.1 - RasOrb.5, and for each of these files we run GRID IT code, and use keyword NAME to give different names for the grid files. Now, we can visualize the difference between grids, computed for different root. Use the command: gv.exe acrolein.ms-cas-grid.root1.grid -a -1.0 acrolein.ms-cas-grid.root2.grid to get the difference between first grid (the ground state) and the second grid (first excited state). Red color corresponds to an excess of electrons. Hint: use +/- buttons to change the shown iso-value. Ex 9. If everything above was too trivial... Compute Ionization potential for NO molecule. For simplicity, use the same interatomic distance for NO and NO + : Å, use basis set ANO-S-VDZP. Questions: What is the spin state of NO? What is the spin state of NO +? What to do if you don t know the answer? What would be an appropriate orbital spaces? How can you verify that 14
15 statement? The correct answer 9.26 ev (Hubert&Herzberg) Dissociation of trans-butadiene molecule If we would like to study dissociation of butadiene molecule into two vinyl radicals, we also have to break C-C bond. It means, that for our active space we have to addcorresponding σ bond. Identify σ andσ orbitals, andinclude them into active space. Hint. To study a reaction, we need an active space which covers both: reactants and products. Study the dissociation of trans-butadiene molecule. 15
Multiconfigurational methods in MOLCAS
 Multiconfigurational methods in MOLCAS Valera Veryazov Valera.Veryazov@teokem.lu.se Division of Theoretical Chemistry Lund University Sweden CASSCF//CASPT2 p. 1/?? Overview What is correlation? CASPT2
Multiconfigurational methods in MOLCAS Valera Veryazov Valera.Veryazov@teokem.lu.se Division of Theoretical Chemistry Lund University Sweden CASSCF//CASPT2 p. 1/?? Overview What is correlation? CASPT2
Multiconfigurational Quantum Chemistry. Björn O. Roos as told by RL Department of Theoretical Chemistry Chemical Center Lund University Sweden
 Multiconfigurational Quantum Chemistry Björn O. Roos as told by RL Department of Theoretical Chemistry Chemical Center Lund University Sweden April 20, 2009 1 The Slater determinant Using the spin-orbitals,
Multiconfigurational Quantum Chemistry Björn O. Roos as told by RL Department of Theoretical Chemistry Chemical Center Lund University Sweden April 20, 2009 1 The Slater determinant Using the spin-orbitals,
Introduction to Hartree-Fock calculations in Spartan
 EE5 in 2008 Hannes Jónsson Introduction to Hartree-Fock calculations in Spartan In this exercise, you will get to use state of the art software for carrying out calculations of wavefunctions for molecues,
EE5 in 2008 Hannes Jónsson Introduction to Hartree-Fock calculations in Spartan In this exercise, you will get to use state of the art software for carrying out calculations of wavefunctions for molecues,
IFM Chemistry Computational Chemistry 2010, 7.5 hp LAB2. Computer laboratory exercise 1 (LAB2): Quantum chemical calculations
 Computer laboratory exercise 1 (LAB2): Quantum chemical calculations Introduction: The objective of the second computer laboratory exercise is to get acquainted with a program for performing quantum chemical
Computer laboratory exercise 1 (LAB2): Quantum chemical calculations Introduction: The objective of the second computer laboratory exercise is to get acquainted with a program for performing quantum chemical
Molecular Orbitals for Ozone
 Molecular Orbitals for Ozone Purpose: In this exercise you will do semi-empirical molecular orbital calculations on ozone with the goal of understanding the molecular orbital print out provided by Spartan
Molecular Orbitals for Ozone Purpose: In this exercise you will do semi-empirical molecular orbital calculations on ozone with the goal of understanding the molecular orbital print out provided by Spartan
Practical Issues on the Use of the CASPT2/CASSCF Method in Modeling Photochemistry: the Selection and Protection of an Active Space
 Practical Issues on the Use of the CASPT2/CASSCF Method in Modeling Photochemistry: the Selection and Protection of an Active Space Roland Lindh Dept. of Chemistry Ångström The Theoretical Chemistry Programme
Practical Issues on the Use of the CASPT2/CASSCF Method in Modeling Photochemistry: the Selection and Protection of an Active Space Roland Lindh Dept. of Chemistry Ångström The Theoretical Chemistry Programme
Supporting Information to: Theoretical analysis of the solvent effects on the magnetic exchange coupling in bis-nitroxides
 Supporting Information to: Theoretical analysis of the solvent effects on the magnetic exchange coupling in bis-nitroxides Esther Coulaud, Denis Hagebaum-Reigner, Didier Siri, Paul Tordo and Nicolas Ferré
Supporting Information to: Theoretical analysis of the solvent effects on the magnetic exchange coupling in bis-nitroxides Esther Coulaud, Denis Hagebaum-Reigner, Didier Siri, Paul Tordo and Nicolas Ferré
Hints on Using the Orca Program
 Computational Chemistry Workshops West Ridge Research Building-UAF Campus 9:00am-4:00pm, Room 009 Electronic Structure - July 19-21, 2016 Molecular Dynamics - July 26-28, 2016 Hints on Using the Orca Program
Computational Chemistry Workshops West Ridge Research Building-UAF Campus 9:00am-4:00pm, Room 009 Electronic Structure - July 19-21, 2016 Molecular Dynamics - July 26-28, 2016 Hints on Using the Orca Program
MRCI calculations in MOLPRO
 1 MRCI calculations in MOLPRO Molpro is a software package written in Fortran and maintained by H.J. Werner and P.J. Knowles. It is often used for performing sophisticated electronic structure calculations,
1 MRCI calculations in MOLPRO Molpro is a software package written in Fortran and maintained by H.J. Werner and P.J. Knowles. It is often used for performing sophisticated electronic structure calculations,
LUMO + 1 LUMO. Tómas Arnar Guðmundsson Report 2 Reikniefnafræði G
 Q1: Display all the MOs for N2 in your report and classify each one of them as bonding, antibonding or non-bonding, and say whether the symmetry of the orbital is σ or π. Sketch a molecular orbital diagram
Q1: Display all the MOs for N2 in your report and classify each one of them as bonding, antibonding or non-bonding, and say whether the symmetry of the orbital is σ or π. Sketch a molecular orbital diagram
5.4. Electronic structure of water
 5.4. Electronic structure of water Water belongs to C 2v point group, we have discussed the corresponding character table. Here it is again: C 2v E C 2 σ v (yz) σ v (xz) A 1 1 1 1 1 A 2 1 1-1 -1 B 1 1-1
5.4. Electronic structure of water Water belongs to C 2v point group, we have discussed the corresponding character table. Here it is again: C 2v E C 2 σ v (yz) σ v (xz) A 1 1 1 1 1 A 2 1 1-1 -1 B 1 1-1
Practical Advice for Quantum Chemistry Computations. C. David Sherrill School of Chemistry and Biochemistry Georgia Institute of Technology
 Practical Advice for Quantum Chemistry Computations C. David Sherrill School of Chemistry and Biochemistry Georgia Institute of Technology Choice of Basis Set STO-3G is too small 6-31G* or 6-31G** 6 probably
Practical Advice for Quantum Chemistry Computations C. David Sherrill School of Chemistry and Biochemistry Georgia Institute of Technology Choice of Basis Set STO-3G is too small 6-31G* or 6-31G** 6 probably
计算物理作业二. Excercise 1: Illustration of the convergence of the dissociation energy for H 2 toward HF limit.
 计算物理作业二 Excercise 1: Illustration of the convergence of the dissociation energy for H 2 toward HF limit. In this exercise, basis indicates one of the following basis sets: STO-3G, cc-pvdz, cc-pvtz, cc-pvqz
计算物理作业二 Excercise 1: Illustration of the convergence of the dissociation energy for H 2 toward HF limit. In this exercise, basis indicates one of the following basis sets: STO-3G, cc-pvdz, cc-pvtz, cc-pvqz
Ethene. Introduction. The ethene molecule is planar (i.e. all the six atoms lie in the same plane) and has a high degree of symmetry:
 FY1006 Innføring i kvantefysikk og TFY4215 Kjemisk fysikk og kvantemekanikk Spring 2012 Chemical Physics Exercise 1 To be delivered by Friday 27.04.12 Introduction Ethene. Ethylene, C 2 H 4, or ethene,
FY1006 Innføring i kvantefysikk og TFY4215 Kjemisk fysikk og kvantemekanikk Spring 2012 Chemical Physics Exercise 1 To be delivered by Friday 27.04.12 Introduction Ethene. Ethylene, C 2 H 4, or ethene,
Theoretical UV/VIS Spectroscopy
 Theoretical UV/VIS Spectroscopy Why is a Ruby Red When Chromium Oxide is Green? How Does a Ruby Laser Work? Goals of this Exercise: - Calculation of the energy of electronically excited states - Understanding
Theoretical UV/VIS Spectroscopy Why is a Ruby Red When Chromium Oxide is Green? How Does a Ruby Laser Work? Goals of this Exercise: - Calculation of the energy of electronically excited states - Understanding
Some practical HINTS on how to use MOLCAS in quantum chemistry problems
 Some practical HINTS on how to use MOLCAS in quantum chemistry problems Luis Serrano-Andrés and Valera Veryazov MOLCAS hints p.1/19 Outline Some comments about different programs Planning your research
Some practical HINTS on how to use MOLCAS in quantum chemistry problems Luis Serrano-Andrés and Valera Veryazov MOLCAS hints p.1/19 Outline Some comments about different programs Planning your research
Figure 1: Transition State, Saddle Point, Reaction Pathway
 Computational Chemistry Workshops West Ridge Research Building-UAF Campus 9:00am-4:00pm, Room 009 Electronic Structure - July 19-21, 2016 Molecular Dynamics - July 26-28, 2016 Potential Energy Surfaces
Computational Chemistry Workshops West Ridge Research Building-UAF Campus 9:00am-4:00pm, Room 009 Electronic Structure - July 19-21, 2016 Molecular Dynamics - July 26-28, 2016 Potential Energy Surfaces
The MCSCF Method *, Molecular Orbitals, Reference Spaces and COLUMBUS Input
 The MCSCF Method *, Molecular Orbitals, Reference Spaces and COLUMBUS Input Hans Lischka University of Vienna *Excerpt of a course presented by R. Shepard, Argonne National Laboratory, at the Workshop
The MCSCF Method *, Molecular Orbitals, Reference Spaces and COLUMBUS Input Hans Lischka University of Vienna *Excerpt of a course presented by R. Shepard, Argonne National Laboratory, at the Workshop
ABC of DFT: Hands-on session 1 Introduction into calculations on molecules
 ABC of DFT: Hands-on session 1 Introduction into calculations on molecules Tutor: Alexej Bagrets Wann? 09.11.2012, 11:30-13:00 Wo? KIT Campus Nord, Flachbau Physik, Geb. 30.22, Computerpool, Raum FE-6
ABC of DFT: Hands-on session 1 Introduction into calculations on molecules Tutor: Alexej Bagrets Wann? 09.11.2012, 11:30-13:00 Wo? KIT Campus Nord, Flachbau Physik, Geb. 30.22, Computerpool, Raum FE-6
CASSCF and NEVPT2 calculations: Ground and excited states of multireference systems. A case study of Ni(CO)4 and the magnetic system NArO
 CASSCF and NEVPT2 calculations: Ground and excited states of multireference systems. A case study of Ni(CO)4 and the magnetic system NArO The ground states of many molecules are often well described by
CASSCF and NEVPT2 calculations: Ground and excited states of multireference systems. A case study of Ni(CO)4 and the magnetic system NArO The ground states of many molecules are often well described by
Extended Wavefunction Analysis for Multireference Methods
 Extended Wavefunction Analysis for Multireference Methods Felix Plasser González Research Group Institute for Theoretical Chemistry, University of Vienna, Austria Vienna, 1 st April 2016 Introduction Analysis
Extended Wavefunction Analysis for Multireference Methods Felix Plasser González Research Group Institute for Theoretical Chemistry, University of Vienna, Austria Vienna, 1 st April 2016 Introduction Analysis
Electron Correlation Methods
 Electron Correlation Methods HF method: electron-electron interaction is replaced by an average interaction E HF c = E 0 E HF E 0 exact ground state energy E HF HF energy for a given basis set HF E c
Electron Correlation Methods HF method: electron-electron interaction is replaced by an average interaction E HF c = E 0 E HF E 0 exact ground state energy E HF HF energy for a given basis set HF E c
This is called a singlet or spin singlet, because the so called multiplicity, or number of possible orientations of the total spin, which is
 9. Open shell systems The derivation of Hartree-Fock equations (Chapter 7) was done for a special case of a closed shell systems. Closed shell means that each MO is occupied by two electrons with the opposite
9. Open shell systems The derivation of Hartree-Fock equations (Chapter 7) was done for a special case of a closed shell systems. Closed shell means that each MO is occupied by two electrons with the opposite
Chemistry 543--Final Exam--Keiderling May 5, pm SES
 Chemistry 543--Final Exam--Keiderling May 5,1992 -- 1-5pm -- 174 SES Please answer all questions in the answer book provided. Make sure your name is clearly indicated and that the answers are clearly numbered,
Chemistry 543--Final Exam--Keiderling May 5,1992 -- 1-5pm -- 174 SES Please answer all questions in the answer book provided. Make sure your name is clearly indicated and that the answers are clearly numbered,
Introduction to multiconfigurational quantum chemistry. Emmanuel Fromager
 Institut de Chimie, Strasbourg, France Page 1 Emmanuel Fromager Institut de Chimie de Strasbourg - Laboratoire de Chimie Quantique - Université de Strasbourg /CNRS M2 lecture, Strasbourg, France. Notations
Institut de Chimie, Strasbourg, France Page 1 Emmanuel Fromager Institut de Chimie de Strasbourg - Laboratoire de Chimie Quantique - Université de Strasbourg /CNRS M2 lecture, Strasbourg, France. Notations
Instructions for Using Spartan 14
 Instructions for Using Spartan 14 Log in to the computer with your Colby ID and password. Click on the Spartan 14 icon in the dock at the bottom of your screen. I. Building Molecules Spartan has one main
Instructions for Using Spartan 14 Log in to the computer with your Colby ID and password. Click on the Spartan 14 icon in the dock at the bottom of your screen. I. Building Molecules Spartan has one main
Practical 1: Structure and electronic properties of organic molecules. B/ Structure, electronic and vibrational properties of the water molecule
 D1CH9116 - MMCO Molecular Modelling applied to organic chemistry Practical 1: tructure and electronic properties of organic molecules B/ tructure, electronic and vibrational properties of the water molecule
D1CH9116 - MMCO Molecular Modelling applied to organic chemistry Practical 1: tructure and electronic properties of organic molecules B/ tructure, electronic and vibrational properties of the water molecule
Supplementary Information for: Hydrogen abstraction by photoexcited benzophenone: consequences for DNA photosensitization
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2016 Supplementary Information for: Hydrogen abstraction by photoexcited benzophenone:
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2016 Supplementary Information for: Hydrogen abstraction by photoexcited benzophenone:
Using Web-Based Computations in Organic Chemistry
 10/30/2017 1 Using Web-Based Computations in Organic Chemistry John Keller UAF Department of Chemistry & Biochemistry The UAF WebMO site Practical aspects of computational chemistry theory and nomenclature
10/30/2017 1 Using Web-Based Computations in Organic Chemistry John Keller UAF Department of Chemistry & Biochemistry The UAF WebMO site Practical aspects of computational chemistry theory and nomenclature
The Hückel Approximation Consider a conjugated molecule i.e. a molecule with alternating double and single bonds, as shown in Figure 1.
 The Hückel Approximation In this exercise you will use a program called Hückel to look at the p molecular orbitals in conjugated molecules. The program calculates the energies and shapes of p (pi) molecular
The Hückel Approximation In this exercise you will use a program called Hückel to look at the p molecular orbitals in conjugated molecules. The program calculates the energies and shapes of p (pi) molecular
ABC of DFT: Hands-on session 2 Molecules: structure optimization, visualization of orbitals, charge & spin densities
 ABC of DFT: Hands-on session 2 Molecules: structure optimization, visualization of orbitals, charge & spin densities Tutor: Alexej Bagrets Wann? 16.11.2012, 11:30-13:00 Wo? KIT Campus Nord, Flachbau Physik,
ABC of DFT: Hands-on session 2 Molecules: structure optimization, visualization of orbitals, charge & spin densities Tutor: Alexej Bagrets Wann? 16.11.2012, 11:30-13:00 Wo? KIT Campus Nord, Flachbau Physik,
Department of Chemistry. The Intersection of Computational Chemistry and Experiment
 Department of Chemistry The Intersection of Computational Chemistry and Experiment Structure, Vibrational and Electronic Spectra of Organic Molecules Angelo R. Rossi Department of Chemistry The University
Department of Chemistry The Intersection of Computational Chemistry and Experiment Structure, Vibrational and Electronic Spectra of Organic Molecules Angelo R. Rossi Department of Chemistry The University
NWChem: Coupled Cluster Method (Tensor Contraction Engine)
 NWChem: Coupled Cluster Method (Tensor Contraction Engine) Why CC is important?! Correlation effects are important!! CC is size-extensive theory: can be used to describe dissociation processes.! Higher-order
NWChem: Coupled Cluster Method (Tensor Contraction Engine) Why CC is important?! Correlation effects are important!! CC is size-extensive theory: can be used to describe dissociation processes.! Higher-order
CHEM J-5 June 2014
 CHEM1101 2014-J-5 June 2014 The molecular orbital energy level diagrams for H 2, H 2 +, H 2 and O 2 are shown below. Fill in the valence electrons for each species in its ground state and label the types
CHEM1101 2014-J-5 June 2014 The molecular orbital energy level diagrams for H 2, H 2 +, H 2 and O 2 are shown below. Fill in the valence electrons for each species in its ground state and label the types
Lecture 4: methods and terminology, part II
 So theory guys have got it made in rooms free of pollution. Instead of problems with the reflux, they have only solutions... In other words, experimentalists will likely die of cancer From working hard,
So theory guys have got it made in rooms free of pollution. Instead of problems with the reflux, they have only solutions... In other words, experimentalists will likely die of cancer From working hard,
Table of Contents. Table of Contents Spin-orbit splitting of semiconductor band structures
 Table of Contents Table of Contents Spin-orbit splitting of semiconductor band structures Relavistic effects in Kohn-Sham DFT Silicon band splitting with ATK-DFT LSDA initial guess for the ground state
Table of Contents Table of Contents Spin-orbit splitting of semiconductor band structures Relavistic effects in Kohn-Sham DFT Silicon band splitting with ATK-DFT LSDA initial guess for the ground state
The Hückel Approximation
 The ückel Approximation 1 In this exercise you will use a program called ückel to look at the π molecular orbitals in conjugated molecules. The program calculates the energies and shapes of π (pi) molecular
The ückel Approximation 1 In this exercise you will use a program called ückel to look at the π molecular orbitals in conjugated molecules. The program calculates the energies and shapes of π (pi) molecular
4 Post-Hartree Fock Methods: MPn and Configuration Interaction
 4 Post-Hartree Fock Methods: MPn and Configuration Interaction In the limit of a complete basis, the Hartree-Fock (HF) energy in the complete basis set limit (ECBS HF ) yields an upper boundary to the
4 Post-Hartree Fock Methods: MPn and Configuration Interaction In the limit of a complete basis, the Hartree-Fock (HF) energy in the complete basis set limit (ECBS HF ) yields an upper boundary to the
A Computer Study of Molecular Electronic Structure
 A Computer Study of Molecular Electronic Structure The following exercises are designed to give you a brief introduction to some of the types of information that are now readily accessible from electronic
A Computer Study of Molecular Electronic Structure The following exercises are designed to give you a brief introduction to some of the types of information that are now readily accessible from electronic
Name. Chem Organic Chemistry II Laboratory Exercise Molecular Modeling Part 2
 Name Chem 322 - Organic Chemistry II Laboratory Exercise Molecular Modeling Part 2 Click on Titan in the Start menu. When it boots, click on the right corner to make the window full-screen. icon in the
Name Chem 322 - Organic Chemistry II Laboratory Exercise Molecular Modeling Part 2 Click on Titan in the Start menu. When it boots, click on the right corner to make the window full-screen. icon in the
Assignment A02: Geometry Definition: File Formats, Redundant Coordinates, PES Scans
 Assignment A02: Geometry Definition: File Formats, Redundant Coordinates, PES Scans In Assignments A00 and A01, you familiarized yourself with GaussView and G09W, you learned the basics about input (GJF)
Assignment A02: Geometry Definition: File Formats, Redundant Coordinates, PES Scans In Assignments A00 and A01, you familiarized yourself with GaussView and G09W, you learned the basics about input (GJF)
Example: H 2 O (the car file)
 Example: H 2 O (the car file) As a practical example of DFT methods we calculate the energy and electronic properties of the water molecule. In order to carry out the DFT calculation you will need a set
Example: H 2 O (the car file) As a practical example of DFT methods we calculate the energy and electronic properties of the water molecule. In order to carry out the DFT calculation you will need a set
CHAPTER 9 THEORY OF RESONANCE BY, G.DEEPA
 CHAPTER 9 THEORY OF RESONANCE BY, G.DEEPA Conjugation in Alkadienes and Allylic Systems conjugation a series of overlapping p orbitals The Allyl Group allylic position is the next to a double bond 1 allyl
CHAPTER 9 THEORY OF RESONANCE BY, G.DEEPA Conjugation in Alkadienes and Allylic Systems conjugation a series of overlapping p orbitals The Allyl Group allylic position is the next to a double bond 1 allyl
Electronic Structure Models
 Electronic Structure Models Hückel Model (1933) Basic Assumptions: (a) One orbital per atom contributes to the basis set; all orbitals "equal" (b) The relevant integrals involving the Hamiltonian are α
Electronic Structure Models Hückel Model (1933) Basic Assumptions: (a) One orbital per atom contributes to the basis set; all orbitals "equal" (b) The relevant integrals involving the Hamiltonian are α
indicating the configuration they correspond to and predict their relative energy.
 Problem 1 (1 point) Three center four electron (3c/4e) bonds were introduced in class. John F. Berry (Dalton Trans. 2012, 41, 700-713) discusses the effect of the larger density of states for the 3c/4e
Problem 1 (1 point) Three center four electron (3c/4e) bonds were introduced in class. John F. Berry (Dalton Trans. 2012, 41, 700-713) discusses the effect of the larger density of states for the 3c/4e
Conjugated Systems. With conjugated double bonds resonance structures can be drawn
 Conjugated Systems Double bonds in conjugation behave differently than isolated double bonds With conjugated double bonds resonance structures can be drawn With isolated double bonds cannot draw resonance
Conjugated Systems Double bonds in conjugation behave differently than isolated double bonds With conjugated double bonds resonance structures can be drawn With isolated double bonds cannot draw resonance
Chemistry 5021/8021 Computational Chemistry 3/4 Credits Spring Semester 2004 ( Due 4 / 5 / 04 )
 Chemistry 5021/8021 Computational Chemistry 3/4 Credits Spring Semester 2004 ( Due 4 / 5 / 04 ) This problem set will take longer than the last one in the sense that you may need to submit some jobs, leave,
Chemistry 5021/8021 Computational Chemistry 3/4 Credits Spring Semester 2004 ( Due 4 / 5 / 04 ) This problem set will take longer than the last one in the sense that you may need to submit some jobs, leave,
Benzene Dimer: dispersion forces and electronic correlation
 Benzene Dimer: dispersion forces and electronic correlation Introduction The Benzene dimer is an ideal example of a system bound by π-π interaction, which is in several cases present in many biologically
Benzene Dimer: dispersion forces and electronic correlation Introduction The Benzene dimer is an ideal example of a system bound by π-π interaction, which is in several cases present in many biologically
QUANTUM CHEMISTRY WITH GAUSSIAN : A VERY BRIEF INTRODUCTION (PART 2)
 QUANTUM CHEMISTRY WITH GAUSSIAN : A VERY BRIEF INTRODUCTION (PART 2) TARAS V. POGORELOV AND MIKE HALLOCK SCHOOL OF CHEMICAL SCIENCES, UIUC This tutorial continues introduction to Gaussian [2]. Here we
QUANTUM CHEMISTRY WITH GAUSSIAN : A VERY BRIEF INTRODUCTION (PART 2) TARAS V. POGORELOV AND MIKE HALLOCK SCHOOL OF CHEMICAL SCIENCES, UIUC This tutorial continues introduction to Gaussian [2]. Here we
Literature values: ΔH f, gas = % error Source: ΔH f, solid = % error. For comparison, your experimental value was ΔH f = phase:
 1 Molecular Calculations Lab: Some guideline given at the bottom of page 3. 1. Use the semi-empirical AM1 method to calculate ΔH f for the compound you used in the heat of combustion experiment. Be sure
1 Molecular Calculations Lab: Some guideline given at the bottom of page 3. 1. Use the semi-empirical AM1 method to calculate ΔH f for the compound you used in the heat of combustion experiment. Be sure
Modeling the UV-Vis Absorption of a Series of Dyes CH342L: Spectroscopy February 15, 2016
 Modeling the UV-Vis Absorption of a Series of Dyes CH342L: Spectroscopy February 15, 2016 We ll correlate the absorbance maximum of a series of dyes with structural changes between them 1. Chemicals absorb
Modeling the UV-Vis Absorption of a Series of Dyes CH342L: Spectroscopy February 15, 2016 We ll correlate the absorbance maximum of a series of dyes with structural changes between them 1. Chemicals absorb
MENA 9520 FME Modelling Tutorial 5 ( )
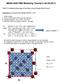 MENA 9520 FME Modelling Tutorial 5 (25.02.2011) Task 5: Understanding type of bonding using charge-density plots Exercise 5.1: Visualizing charge density in Si: 1. mkdir charge 2. Copy a converged CTRL
MENA 9520 FME Modelling Tutorial 5 (25.02.2011) Task 5: Understanding type of bonding using charge-density plots Exercise 5.1: Visualizing charge density in Si: 1. mkdir charge 2. Copy a converged CTRL
Chemistry 14CL. Worksheet for the Molecular Modeling Workshop. (Revised FULL Version 2012 J.W. Pang) (Modified A. A. Russell)
 Chemistry 14CL Worksheet for the Molecular Modeling Workshop (Revised FULL Version 2012 J.W. Pang) (Modified A. A. Russell) Structure of the Molecular Modeling Assignment The molecular modeling assignment
Chemistry 14CL Worksheet for the Molecular Modeling Workshop (Revised FULL Version 2012 J.W. Pang) (Modified A. A. Russell) Structure of the Molecular Modeling Assignment The molecular modeling assignment
Computational Chemistry Using the University of Alaska WebMO Site
 2/7/2017 1 Computational Chemistry Using the University of Alaska WebMO Site John Keller Department of Chemistry & Biochemistry University of Alaska Fairbanks Intro and operation of WebMO and MOPAC Basic
2/7/2017 1 Computational Chemistry Using the University of Alaska WebMO Site John Keller Department of Chemistry & Biochemistry University of Alaska Fairbanks Intro and operation of WebMO and MOPAC Basic
Molecular Modeling Lecture 7. Homology modeling insertions/deletions manual realignment
 Molecular Modeling 2018-- Lecture 7 Homology modeling insertions/deletions manual realignment Homology modeling also called comparative modeling Sequences that have similar sequence have similar structure.
Molecular Modeling 2018-- Lecture 7 Homology modeling insertions/deletions manual realignment Homology modeling also called comparative modeling Sequences that have similar sequence have similar structure.
Covalent Bonding 10/29/2013
 Bond Energies or Bond Dissociation Energies Tables 8.4 and 8.5 on page 72 gives a list of the energy required to dissociate or break bonds. This value is used to determine whether covalent bonds will form
Bond Energies or Bond Dissociation Energies Tables 8.4 and 8.5 on page 72 gives a list of the energy required to dissociate or break bonds. This value is used to determine whether covalent bonds will form
Calculating Bond Enthalpies of the Hydrides
 Proposed Exercise for the General Chemistry Section of the Teaching with Cache Workbook: Calculating Bond Enthalpies of the Hydrides Contributed by James Foresman, Rachel Fogle, and Jeremy Beck, York College
Proposed Exercise for the General Chemistry Section of the Teaching with Cache Workbook: Calculating Bond Enthalpies of the Hydrides Contributed by James Foresman, Rachel Fogle, and Jeremy Beck, York College
Due: since the calculation takes longer than before, we ll make it due on 02/05/2016, Friday
 Homework 3 Due: since the calculation takes longer than before, we ll make it due on 02/05/2016, Friday Email to: jqian@caltech.edu Introduction In this assignment, you will be using a commercial periodic
Homework 3 Due: since the calculation takes longer than before, we ll make it due on 02/05/2016, Friday Email to: jqian@caltech.edu Introduction In this assignment, you will be using a commercial periodic
Conformational energy analysis
 Lab 3 Conformational energy analysis Objective This computational project deals with molecular conformations the spatial arrangement of atoms of molecules. Conformations are determined by energy, so the
Lab 3 Conformational energy analysis Objective This computational project deals with molecular conformations the spatial arrangement of atoms of molecules. Conformations are determined by energy, so the
Athena Visual Software, Inc. 1
 Athena Visual Studio Visual Kinetics Tutorial VisualKinetics is an integrated tool within the Athena Visual Studio software environment, which allows scientists and engineers to simulate the dynamic behavior
Athena Visual Studio Visual Kinetics Tutorial VisualKinetics is an integrated tool within the Athena Visual Studio software environment, which allows scientists and engineers to simulate the dynamic behavior
A TUTORIAL FOR COLUMBUS
 A TUTORIAL FOR COLUMBUS Bernhard Sellner, Jaroslaw Szymczak, Felix Plasser, Reed Nieman, Mario Barbatti Institute for Theoretical Chemistry University of Vienna www.univie.ac.at/columbus/tutorial Vienna,
A TUTORIAL FOR COLUMBUS Bernhard Sellner, Jaroslaw Szymczak, Felix Plasser, Reed Nieman, Mario Barbatti Institute for Theoretical Chemistry University of Vienna www.univie.ac.at/columbus/tutorial Vienna,
Introduction to Hartree-Fock calculations using ORCA and Chemcraft
 Introduction to Hartree-Fock calculations using ORCA and Chemcraft In this exercise, you will get to use software for carrying out calculations of wavefunctions for molecules, the ORCA program. While ORCA
Introduction to Hartree-Fock calculations using ORCA and Chemcraft In this exercise, you will get to use software for carrying out calculations of wavefunctions for molecules, the ORCA program. While ORCA
Molecular Modeling and Conformational Analysis with PC Spartan
 Molecular Modeling and Conformational Analysis with PC Spartan Introduction Molecular modeling can be done in a variety of ways, from using simple hand-held models to doing sophisticated calculations on
Molecular Modeling and Conformational Analysis with PC Spartan Introduction Molecular modeling can be done in a variety of ways, from using simple hand-held models to doing sophisticated calculations on
Chemistry 5021/8021 Computational Chemistry 3/4 Credits Spring Semester 2000 ( Due 4 / 3 / 00 )
 Chemistry 5021/8021 Computational Chemistry 3/4 Credits Spring Semester 2000 ( Due 4 / 3 / 00 ) This problem set will take longer than the last one in the sense that you may need to submit some jobs, leave,
Chemistry 5021/8021 Computational Chemistry 3/4 Credits Spring Semester 2000 ( Due 4 / 3 / 00 ) This problem set will take longer than the last one in the sense that you may need to submit some jobs, leave,
CHEM 545 Theory and Practice of Molecular Electronic Structure. Anna I. Krylov. DON T PANIC.
 CHEM 545 Theory and Practice of Molecular Electronic Structure Anna I. Krylov http://iopenshell.usc.edu/chem545/ DON T PANIC USC Fall 2014 Things to do: 1. Install IQmol (by this Thursday). http://iqmol.org/.
CHEM 545 Theory and Practice of Molecular Electronic Structure Anna I. Krylov http://iopenshell.usc.edu/chem545/ DON T PANIC USC Fall 2014 Things to do: 1. Install IQmol (by this Thursday). http://iqmol.org/.
Reikniefnafræði - Verkefni 2 Haustmisseri 2013 Kennari - Hannes Jónsson
 Háskóli Íslands, raunvísindasvið Reikniefnafræði - Verkefni 2 Haustmisseri 2013 Kennari - Hannes Jónsson Guðjón Henning 18. september 2013 1 A. Molecular orbitals of N 2 Q1: Display all the MOs for N 2
Háskóli Íslands, raunvísindasvið Reikniefnafræði - Verkefni 2 Haustmisseri 2013 Kennari - Hannes Jónsson Guðjón Henning 18. september 2013 1 A. Molecular orbitals of N 2 Q1: Display all the MOs for N 2
Performance of Hartree-Fock and Correlated Methods
 Chemistry 460 Fall 2017 Dr. Jean M. Standard December 4, 2017 Performance of Hartree-Fock and Correlated Methods Hartree-Fock Methods Hartree-Fock methods generally yield optimized geomtries and molecular
Chemistry 460 Fall 2017 Dr. Jean M. Standard December 4, 2017 Performance of Hartree-Fock and Correlated Methods Hartree-Fock Methods Hartree-Fock methods generally yield optimized geomtries and molecular
S N 2 reaction. Br- + ClCH 3 BrCH 3 + Cl-
 FY1006 Innføring i kvantefysikk og TFY4215 Kjemisk fysikk og kvantemekanikk Spring 2012 Chemical Physics Exercise 2 To be delivered by Friday 27.04.12 S N 2 reaction. Introduction Many chemical reactions
FY1006 Innføring i kvantefysikk og TFY4215 Kjemisk fysikk og kvantemekanikk Spring 2012 Chemical Physics Exercise 2 To be delivered by Friday 27.04.12 S N 2 reaction. Introduction Many chemical reactions
Designing a Quilt with GIMP 2011
 Planning your quilt and want to see what it will look like in the fabric you just got from your LQS? You don t need to purchase a super expensive program. Try this and the best part it s FREE!!! *** Please
Planning your quilt and want to see what it will look like in the fabric you just got from your LQS? You don t need to purchase a super expensive program. Try this and the best part it s FREE!!! *** Please
HOW TO USE MIKANA. 1. Decompress the zip file MATLAB.zip. This will create the directory MIKANA.
 HOW TO USE MIKANA MIKANA (Method to Infer Kinetics And Network Architecture) is a novel computational method to infer reaction mechanisms and estimate the kinetic parameters of biochemical pathways from
HOW TO USE MIKANA MIKANA (Method to Infer Kinetics And Network Architecture) is a novel computational method to infer reaction mechanisms and estimate the kinetic parameters of biochemical pathways from
Chapter: 22. Visualization: Making INPUT File and Processing of Output Results
 Chapter: 22 Visualization: Making INPUT File and Processing of Output Results Keywords: visualization, input and output structure, molecular orbital, electron density. In the previous chapters, we have
Chapter: 22 Visualization: Making INPUT File and Processing of Output Results Keywords: visualization, input and output structure, molecular orbital, electron density. In the previous chapters, we have
Molecular Simulation I
 Molecular Simulation I Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences
Molecular Simulation I Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences
An Introduction to Basis Sets:
 An Introduction to Basis Sets: The "Holy Grail" of computational chemistry is the calculation of the molecular orbitals (MOs) for a given molecule. IF we can calculate the MOs for a molecule, THEN we can
An Introduction to Basis Sets: The "Holy Grail" of computational chemistry is the calculation of the molecular orbitals (MOs) for a given molecule. IF we can calculate the MOs for a molecule, THEN we can
Symmetry III: Molecular Orbital Theory. Reading: Shriver and Atkins and , 6.10
 Lecture 9 Symmetry III: Molecular Orbital Theory Reading: Shriver and Atkins 2.7-2.9 and g 6.6-6.7, 6.10 The orbitals of molecules H H The electron energy in each H atom is -13.6 ev below vacuum. What
Lecture 9 Symmetry III: Molecular Orbital Theory Reading: Shriver and Atkins 2.7-2.9 and g 6.6-6.7, 6.10 The orbitals of molecules H H The electron energy in each H atom is -13.6 ev below vacuum. What
Last Name or Student ID
 11/05/18, Chem433 Exam # 2 Last ame or Student ID 1. (2 pts) 2. (9 pts) 3. (2 pts) 4. (2 pts) 5. (2 pts) 6. (2 pts) 7. (2 pts) 8. (4 pts) 9. (14 pts) 10. (10 pts) 11. (26/31 pts) 12. (25/27 pts) Extra
11/05/18, Chem433 Exam # 2 Last ame or Student ID 1. (2 pts) 2. (9 pts) 3. (2 pts) 4. (2 pts) 5. (2 pts) 6. (2 pts) 7. (2 pts) 8. (4 pts) 9. (14 pts) 10. (10 pts) 11. (26/31 pts) 12. (25/27 pts) Extra
Q-Chem Workshop. Doubletree Hotel 2085 S. Harbor Boulevard Anaheim, CA March 26, Schedule
 Q-Chem Workshop Doubletree Hotel 2085 S. Harbor Boulevard Anaheim, CA 92802 March 26, 2011 1 8:30 Schedule Welcome remarks, Prof. Peter Gill, Australian National Univ and President of Q-Chem 8:45-9:15
Q-Chem Workshop Doubletree Hotel 2085 S. Harbor Boulevard Anaheim, CA 92802 March 26, 2011 1 8:30 Schedule Welcome remarks, Prof. Peter Gill, Australian National Univ and President of Q-Chem 8:45-9:15
Tutorial I: IQ MOL and Basic DFT and MP2 Calculations 1 / 30
 Tutorial I: IQ MOL and Basic DFT and MP2 Calculations Q-Chem User Workshop, Denver March 21, 2015 1 / 30 2 / 30 Introduction to IQMOL DFT and MP2 Calculations 3 / 30 IQMOL and Q-CHEM IQMOL is an open-source
Tutorial I: IQ MOL and Basic DFT and MP2 Calculations Q-Chem User Workshop, Denver March 21, 2015 1 / 30 2 / 30 Introduction to IQMOL DFT and MP2 Calculations 3 / 30 IQMOL and Q-CHEM IQMOL is an open-source
ACES2 Labs. Part B: Calculating electronically excited, ionized, and electro-attached states using equation of motion coupled cluster calculations.
 ACES2 Labs. Part B: Calculating electronically excited, ionized, and electro-attached states using equation of motion coupled cluster calculations. I. Electronically excited states. Purpose: - Calculation
ACES2 Labs. Part B: Calculating electronically excited, ionized, and electro-attached states using equation of motion coupled cluster calculations. I. Electronically excited states. Purpose: - Calculation
Electric Fields and Equipotentials
 OBJECTIVE Electric Fields and Equipotentials To study and describe the two-dimensional electric field. To map the location of the equipotential surfaces around charged electrodes. To study the relationship
OBJECTIVE Electric Fields and Equipotentials To study and describe the two-dimensional electric field. To map the location of the equipotential surfaces around charged electrodes. To study the relationship
Modeling of S-N Bond Breaking in an Aromatic Sulfilimine. By Jacob Brunsvold & Katrina Hanson Advisor: Stacey Stoffregen
 Modeling of S-N Bond Breaking in an Aromatic Sulfilimine By Jacob Brunsvold & Katrina Hanson Advisor: Stacey Stoffregen Outline! Background Photochemical Reaction! Introduction to Photochemistry and Quantum
Modeling of S-N Bond Breaking in an Aromatic Sulfilimine By Jacob Brunsvold & Katrina Hanson Advisor: Stacey Stoffregen Outline! Background Photochemical Reaction! Introduction to Photochemistry and Quantum
Quick Tutorial on Natural Bond Order 3 Calculations Within Gaussian 09
 Quick Tutorial on Natural Bond Order 3 Calculations Within Gaussian 09 Benjamin Rudshteyn, and Victor S. Batista* victor.batista@yale.edu Yale University, Department of Chemistry 225 Prospect Street, New
Quick Tutorial on Natural Bond Order 3 Calculations Within Gaussian 09 Benjamin Rudshteyn, and Victor S. Batista* victor.batista@yale.edu Yale University, Department of Chemistry 225 Prospect Street, New
What are we going to learn today?
 UNIT3DAY4-LaB Page 1 UNIT3DAY4-LaB Tuesday, October 23, 2012 8:29 AM Vanden Bout/LaBrake CH301 WHY IS EVERYTHING SO DIFFERENT? MORE ON BONDING THEORIES UNIT 3 Day 4 Important Information LM22 DUE Th 9AM
UNIT3DAY4-LaB Page 1 UNIT3DAY4-LaB Tuesday, October 23, 2012 8:29 AM Vanden Bout/LaBrake CH301 WHY IS EVERYTHING SO DIFFERENT? MORE ON BONDING THEORIES UNIT 3 Day 4 Important Information LM22 DUE Th 9AM
Special 5: Wavefunction Analysis and Visualization
 Special 5: Wavefunction Analysis and Visualization Felix Plasser Institute for Theoretical Chemistry, University of Vienna COLUMBUS in China Tianjin, October 10 14, 2016 F. Plasser Wavefunction Analysis
Special 5: Wavefunction Analysis and Visualization Felix Plasser Institute for Theoretical Chemistry, University of Vienna COLUMBUS in China Tianjin, October 10 14, 2016 F. Plasser Wavefunction Analysis
Introduction to Computational Chemistry
 Introduction to Computational Chemistry Vesa Hänninen Laboratory of Physical Chemistry Chemicum 4th floor vesa.hanninen@helsinki.fi September 10, 2013 Lecture 3. Electron correlation methods September
Introduction to Computational Chemistry Vesa Hänninen Laboratory of Physical Chemistry Chemicum 4th floor vesa.hanninen@helsinki.fi September 10, 2013 Lecture 3. Electron correlation methods September
Chm 331 Fall 2015, Exercise Set 4 NMR Review Problems
 Chm 331 Fall 015, Exercise Set 4 NMR Review Problems Mr. Linck Version.0. Compiled December 1, 015 at 11:04:44 4.1 Diagonal Matrix Elements for the nmr H 0 Find the diagonal matrix elements for H 0 (the
Chm 331 Fall 015, Exercise Set 4 NMR Review Problems Mr. Linck Version.0. Compiled December 1, 015 at 11:04:44 4.1 Diagonal Matrix Elements for the nmr H 0 Find the diagonal matrix elements for H 0 (the
Preparing a PDB File
 Figure 1: Schematic view of the ligand-binding domain from the vitamin D receptor (PDB file 1IE9). The crystallographic waters are shown as small spheres and the bound ligand is shown as a CPK model. HO
Figure 1: Schematic view of the ligand-binding domain from the vitamin D receptor (PDB file 1IE9). The crystallographic waters are shown as small spheres and the bound ligand is shown as a CPK model. HO
Ab initio study on the low-lying excited states of retinal
 Ab initio study on the low-lying excited states of retinal Manuela Merchán and Remedios González-Luque Departamento de Química Física, Universitat de València, Dr. Moliner 50, Burjassot, E-46100 Valencia,
Ab initio study on the low-lying excited states of retinal Manuela Merchán and Remedios González-Luque Departamento de Química Física, Universitat de València, Dr. Moliner 50, Burjassot, E-46100 Valencia,
Learning to Use Scigress Wagner, Eugene P. (revised May 15, 2018)
 Learning to Use Scigress Wagner, Eugene P. (revised May 15, 2018) Abstract Students are introduced to basic features of Scigress by building molecules and performing calculations on them using semi-empirical
Learning to Use Scigress Wagner, Eugene P. (revised May 15, 2018) Abstract Students are introduced to basic features of Scigress by building molecules and performing calculations on them using semi-empirical
Ranking of HIV-protease inhibitors using AutoDock
 Ranking of HIV-protease inhibitors using AutoDock 1. Task Calculate possible binding modes and estimate the binding free energies for 1 3 inhibitors of HIV-protease. You will learn: Some of the theory
Ranking of HIV-protease inhibitors using AutoDock 1. Task Calculate possible binding modes and estimate the binding free energies for 1 3 inhibitors of HIV-protease. You will learn: Some of the theory
26 Group Theory Basics
 26 Group Theory Basics 1. Reference: Group Theory and Quantum Mechanics by Michael Tinkham. 2. We said earlier that we will go looking for the set of operators that commute with the molecular Hamiltonian.
26 Group Theory Basics 1. Reference: Group Theory and Quantum Mechanics by Michael Tinkham. 2. We said earlier that we will go looking for the set of operators that commute with the molecular Hamiltonian.
4 Diatomic molecules
 s manual for Burrows et.al. Chemistry 3 Third edition 4 Diatomic molecules Answers to worked examples WE 4.1 The Lewis model (on p. 174 in Chemistry 3 ) Use the Lewis model to describe the bonding in (a)
s manual for Burrows et.al. Chemistry 3 Third edition 4 Diatomic molecules Answers to worked examples WE 4.1 The Lewis model (on p. 174 in Chemistry 3 ) Use the Lewis model to describe the bonding in (a)
5.03 In-Class Exam 2
 5.03 In-Class Exam 2 Christopher C. Cummins March 12, 2010 Instructions Clearly write your name at the top of this front page, but otherwise do not write on this front page as it will be used for scoring.
5.03 In-Class Exam 2 Christopher C. Cummins March 12, 2010 Instructions Clearly write your name at the top of this front page, but otherwise do not write on this front page as it will be used for scoring.
QUANTUM CHEMISTRY FOR TRANSITION METALS
 QUANTUM CHEMISTRY FOR TRANSITION METALS Outline I Introduction II Correlation Static correlation effects MC methods DFT III Relativity Generalities From 4 to 1 components Effective core potential Outline
QUANTUM CHEMISTRY FOR TRANSITION METALS Outline I Introduction II Correlation Static correlation effects MC methods DFT III Relativity Generalities From 4 to 1 components Effective core potential Outline
Lecture B6 Molecular Orbital Theory. Sometimes it's good to be alone.
 Lecture B6 Molecular Orbital Theory Sometimes it's good to be alone. Covalent Bond Theories 1. VSEPR (valence shell electron pair repulsion model). A set of empirical rules for predicting a molecular geometry
Lecture B6 Molecular Orbital Theory Sometimes it's good to be alone. Covalent Bond Theories 1. VSEPR (valence shell electron pair repulsion model). A set of empirical rules for predicting a molecular geometry
Chem 1140; Molecular Modeling
 P. Wipf 1 Chem 1140 $E = -! (q a + q b )" ab S ab + ab!k
P. Wipf 1 Chem 1140 $E = -! (q a + q b )" ab S ab + ab!k
MO Calculation for a Diatomic Molecule. /4 0 ) i=1 j>i (1/r ij )
 MO Calculation for a Diatomic Molecule Introduction The properties of any molecular system can in principle be found by looking at the solutions to the corresponding time independent Schrodinger equation
MO Calculation for a Diatomic Molecule Introduction The properties of any molecular system can in principle be found by looking at the solutions to the corresponding time independent Schrodinger equation
Computational Material Science Part II-1: introduction. Horng-Tay Jeng ( 鄭弘泰 ) Institute of Physics, Academia Sinica
 Computational Material Science Part II-1: introduction Horng-Tay Jeng ( 鄭弘泰 ) Institute of Physics, Academia Sinica Outline Introduction of Computational Material Science (CMS) Density Functional Theory
Computational Material Science Part II-1: introduction Horng-Tay Jeng ( 鄭弘泰 ) Institute of Physics, Academia Sinica Outline Introduction of Computational Material Science (CMS) Density Functional Theory
Multiconfigurational methods for the f-elements
 Multiconfigurational methods for the f-elements Andy Kerridge Winter School in Theoretical f-element Chemistry Helsinki, Finland, December 5 th -8 th, 204 Overview CASPT2: reference weights and intruder
Multiconfigurational methods for the f-elements Andy Kerridge Winter School in Theoretical f-element Chemistry Helsinki, Finland, December 5 th -8 th, 204 Overview CASPT2: reference weights and intruder
Chemistry 4021/8021 Computational Chemistry 3/4 Credits Spring Semester 2013 ( Due 4 / 10 / 12 )
 Chemistry 4021/8021 Computational Chemistry 3/4 Credits Spring Semester 2013 ( Due 4 / 10 / 12 ) This problem set will take longer than the last one in the sense that you will almost certainly need to
Chemistry 4021/8021 Computational Chemistry 3/4 Credits Spring Semester 2013 ( Due 4 / 10 / 12 ) This problem set will take longer than the last one in the sense that you will almost certainly need to
Chemistry 112 Laboratory Experiment 5: Visualizing Molecular Orbitals: A MacSpartan Pro Experience (This experiment will be conducted in OR341)
 Chemistry 112 Laboratory Experiment 5: Visualizing Molecular Orbitals: A MacSpartan Pro Experience (This experiment will be conducted in OR341) Introduction In class we have discussed Lewis structures,
Chemistry 112 Laboratory Experiment 5: Visualizing Molecular Orbitals: A MacSpartan Pro Experience (This experiment will be conducted in OR341) Introduction In class we have discussed Lewis structures,
