Experimental Methods in Kinetics (2014)
|
|
|
- Gyles Bryan
- 6 years ago
- Views:
Transcription
1 IIIa- 35 Experimental Methods in Kinetics (2014) Initial examples assume start R = A 0 + B 0 etc., P = 0 follow reaction forward initial rate, etc. measure [A] vs. t Chemical methods: Could mix and take aliquots, to gain time quench reaction (T, ph, some change.. ) then use chemical analysis Problem slow, requires a lot of material Bio applications, chemical methods not work so well, -- often the change studied not chemically very different (folding, enzymology, ligand or membrane binding,.. ) -- Amounts often very limited (problem for chemical anal.) Physical Methods: Monitor Absorbance, fluorescence, ph, pressure, electrochemistry, scattering, whatever is proportional to concentration e.g. Beer-Lambert law: A = bc, A ~ c ph, pressure, conduct. are ~ conc., low conc. I Fluor ~ c Mixing limitations: Mix reagents in a beaker and stir OK for slow reaction Slow reaction pour together and stir takes time k 2 small (seconds) Speed up by reducing size, depend on diffusion - separation
2 IIIa- 36 Fast reaction need - mix fast Faster mix very small volume rapid mixing small mixer, fast flow & turbulence Stopped flow (Atkins Fig ) fill cell quickly from mixer then stop (cell full of fresh mixed reagents) t = 0 Monitor absorbance, fluorescence or other properties Something proportional concentration Stopped-Flow methods idea is rapid mixing, fill/monitor small volume Drive two components (or more) into mixing chamber and out to a cell for monitoring a physical property typically absorbance or fluorescence A B Mixing can occur in about 1 ms, key is turbulence
3 IIIa- 37 Example: Protein folding rapid mix Lipid vesicle + -lactoglobulin, sheet - helix change (Ning Ge, Kinetic Xiuqi traces Zhang, TAK, Biochemistry, for BLG (0.2mg/ml) with DOPG 2010) Ellipticity(mdeg) -10 N 0.15mM DOPG mM DOPG 0.50mM DOPG mM DOPG 2.00mM DOPG Time/s Relative Intensity Circular Dichroism fit to single exponential [A] = [A0]e-kt Kinetics traces of BLG(0.2mg/ml) with DOPG decay in seconds - apparent 1st order lipid~constant mM DOPG 1.00mM DOPG 0.50mM DOPG 0.25mM DOPG 0.15mM DOPG Time/s Fluorescence senses two steps in mechanism, fit to double exponential [A] = [A0]{a e-k t + a e-k t} k1 k2 k3 N I I U fast decay: 100 ms, slower one: seconds like CD Complex process - use this to postulate a mechanism
4 IIIa- 38 Continuous flow measure along flow x t positions in flow successively later in reaction, x t Faster continuous flow, more material Convert distance to time x t possible to get to 100 s or better with micro fluidics Time scales ~1 ms mixing for small volume: ~ 0.1 ml Faster micro mixers micro etch channels flow idea rate limit mix diffuse A,B together D different constant ~ constant shorter distance between A,B (mixing) Same Concept follow conc. [A] vs. t - physical probe determine rate, vary [A] / determine order
5 Example: rapid mix -lactoglobulin+tfe, transition IIIa- 39
6 IIIa- 40 Speed up initiation if photochemical initate A A* then this can be created in very short time Bio-exam: Hb-CO Hb* + CO recombine after photolysis Rhodopsin Rh* --photo absorb cis-trans conform. change Photocycle light pulse trigger, measure rates for spectral changes, e.g. retinal absorbance, can also trap intermediates with cold or other environment Above focus on loss of reagent (or forming product) If reverse reaction or alternate steps important Need to monitor other species (intermediates) Equilibrium - rate depend on forward and reverse step: R & P
7 IIIa- 41 Equilibrium, if elementary A + B k 1 k -1 K e = k 1 /k -1 = C + D r for = r rev [C] e [D] e [A] e [B] e r r r f k k 1 1 [A] [B] [C] [D] = 1 Disturb equilibrium perturbation then relax back to equil. - change T, P, ph, how? discharge capacitor (T),shock wave (P), Laser flash-(ph precursor) or ex. T-jump (laser absorb by solvent) - system relax to new equilibrium - reaction must go forward/reverse to new state (new equil. conc.) Unimolecular case: A k 1 k -1 B A = A e x B = B e + x x = departure from new equilibrium at time = t, new T, P x = x e e -t/ - relaxation time, x 0 at t =, x = x 0 /e at t = r = k 1 A - k -1 B = k 1 (A e -x) - k -1 (B e +x) -da/dt = dx/dt = -(k 1 +k -1 )x + k 1 A e - k -1 B e = -(k 1 +k -1 )x Since at equil. k 1 A e = k -1 B e, can drop 2nd and 3rd terms dx/x = -(k 1 +k -1 )dt ln[x/x 0 ] = -(k 1 +k -1 )t x = x 0 e -t(k 1 +k -1 ) relaxation time: 1/ = k 1 + k -1 Note: relax faster if either k 1 or k -1 fast (same as rate to equil.) (see Derivation 21.4, p.797 in Atkins, Table 7.4 in Tinoco or Engel Ch ) 2 nd order more complex form (hint: x-small, so x 2 very small, neglect quadratic, 2 nd order)
8 IIIa- 42 Since K eq = k 1 /k -1 get both values k 1, k -1 from & K eq Slow T-jump example: Refold Ribonuclease A, FTIR Monitor change 2 frequencies Alternate Example see Engel p.687 RNA conformation Faster Dyer, Callender and co-workers protein and peptide folding with IR or fluorescence See slides Webpage Peptide folding example
9 IIIa- 43 UIC T-jump example: TrpZip2 hairpin with isotope label Labels let us focus on parts of hairpin fold Variation in Temperature Arrhenius behavior
10 IIIa- 44 Helix-Coil mechanism If a polypeptide is helical, then each residue has local ( ) torsions that fit the helical model If it undergoes a transition to a new form, such as a coil, then the local ( ) values change These could occur in a concerted (all together) manner:... hhhhh ccccc... Or more likely in a stepwise pattern:... hhhhh hchhh hcchh... etc.. ccccc... Residues on the termini are more likely to shift from helix to coil due to solvent exposure and lack of H-bond Shows up as activation energy: E a (hhh..-->chh..) < E a (hhh..-->hch..) E a (ccc..-->chc..) < E a (ccc..-->hcc..) So helices are likely to unfold from the termini, or are more likely to fold from center out to ends start unfolded, random coil
11 IIIa- 45 or start fully extended: key is initiate with H-bond formation, center favored: Taken from: Constant Temperature Simulations of Helix Folding SHEN-SHU SUNG,J. Theor.Biol. (1995), 173,
elementary steps have reaction order like stoichiometry Unimolecular: A k 1 P 1 st order -d[a]/dt = k 1 [A] --> ln [A]/[A 0 ] = -k 1 t
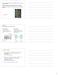
![elementary steps have reaction order like stoichiometry Unimolecular: A k 1 P 1 st order -d[a]/dt = k 1 [A] --> ln [A]/[A 0 ] = -k 1 t elementary steps have reaction order like stoichiometry Unimolecular: A k 1 P 1 st order -d[a]/dt = k 1 [A] --> ln [A]/[A 0 ] = -k 1 t](/thumbs/88/114612848.jpg) B. Mechanism 009 rearrange -- Engel Ch 5.4,0,8 Series of elementary steps (uni-, bimolecular) that when combined give overall reaction and observed rate law elementary steps have reaction order lie stoichiometry
B. Mechanism 009 rearrange -- Engel Ch 5.4,0,8 Series of elementary steps (uni-, bimolecular) that when combined give overall reaction and observed rate law elementary steps have reaction order lie stoichiometry
XV 74. Flouorescence-Polarization-Circular-Dichroism- Jablonski diagram Where does the energy go?
 XV 74 Flouorescence-Polarization-Circular-Dichroism- Jablonski diagram Where does the energy go? 1) Excite system through A Absorbance S 0 S n Excite from ground excited singlet S = 0 could be any of them
XV 74 Flouorescence-Polarization-Circular-Dichroism- Jablonski diagram Where does the energy go? 1) Excite system through A Absorbance S 0 S n Excite from ground excited singlet S = 0 could be any of them
Fluorescence (Notes 16)
 Fluorescence - 2014 (Notes 16) XV 74 Jablonski diagram Where does the energy go? Can be viewed like multistep kinetic pathway 1) Excite system through A Absorbance S 0 S n Excite from ground excited singlet
Fluorescence - 2014 (Notes 16) XV 74 Jablonski diagram Where does the energy go? Can be viewed like multistep kinetic pathway 1) Excite system through A Absorbance S 0 S n Excite from ground excited singlet
Fluorescence 2009 update
 XV 74 Fluorescence 2009 update Jablonski diagram Where does the energy go? Can be viewed like multistep kinetic pathway 1) Excite system through A Absorbance S 0 S n Excite from ground excited singlet
XV 74 Fluorescence 2009 update Jablonski diagram Where does the energy go? Can be viewed like multistep kinetic pathway 1) Excite system through A Absorbance S 0 S n Excite from ground excited singlet
Biophysical Model Building
 Biophysical Model Building Step 1: Come up with a hypothesis about how a system works How many binding sites? Is there cooperativity? Step 2: Translate the qualitative hypotheses into an observable mathematical
Biophysical Model Building Step 1: Come up with a hypothesis about how a system works How many binding sites? Is there cooperativity? Step 2: Translate the qualitative hypotheses into an observable mathematical
Trpzip-based beta hairpin temperature jump IR studies enhanced by sitespecific
 Trpzip-based beta hairpin temperature jump IR studies enhanced by sitespecific isotope labeling Carsten Kretjschi 1, Karin auser 1, Rong uang 2, Tim Keiderling 2 1 Institute of Biophysics, University of
Trpzip-based beta hairpin temperature jump IR studies enhanced by sitespecific isotope labeling Carsten Kretjschi 1, Karin auser 1, Rong uang 2, Tim Keiderling 2 1 Institute of Biophysics, University of
Contents. xiii. Preface v
 Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
or more general example: aa + bb cc + dd r = -1/a da/dt = -1/b db/dt = 1/c dc/dt = 1/d dd/dt
 Chem 344--Physical Chemistry for Biochemists II --F'12 I. Introduction see syllabus II. Experimental Chemical kinetics (Atkins, Ch.6) How fast is reaction? Rate of formation of product or loss of reactant
Chem 344--Physical Chemistry for Biochemists II --F'12 I. Introduction see syllabus II. Experimental Chemical kinetics (Atkins, Ch.6) How fast is reaction? Rate of formation of product or loss of reactant
Principles of Physical Biochemistry
 Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Kinetics Mechanisms (2012) Examples Atkins Ch 7 Tinoco Ch.7 (p ), Engel Ch , Ch
 II 3 Kinetics Mechanisms (01) Examples Atins Ch 7 Tinoco Ch.7 (p.341-354), Engel Ch 5.5-10, Ch 6.1-3 Recall penicillin example basic chemistry, open ring N O R + H O O O We saw observed rate law: 1 st
II 3 Kinetics Mechanisms (01) Examples Atins Ch 7 Tinoco Ch.7 (p.341-354), Engel Ch 5.5-10, Ch 6.1-3 Recall penicillin example basic chemistry, open ring N O R + H O O O We saw observed rate law: 1 st
Chapter 13 Lecture Lecture Presentation. Chapter 13. Chemical Kinetics. Sherril Soman Grand Valley State University Pearson Education, Inc.
 Chapter 13 Lecture Lecture Presentation Chapter 13 Chemical Kinetics Sherril Soman Grand Valley State University Ectotherms Lizards, and other cold-blooded creatures, are ectotherms animals whose body
Chapter 13 Lecture Lecture Presentation Chapter 13 Chemical Kinetics Sherril Soman Grand Valley State University Ectotherms Lizards, and other cold-blooded creatures, are ectotherms animals whose body
Advanced Physical Chemistry CHAPTER 18 ELEMENTARY CHEMICAL KINETICS
 Experimental Kinetics and Gas Phase Reactions Advanced Physical Chemistry CHAPTER 18 ELEMENTARY CHEMICAL KINETICS Professor Angelo R. Rossi http://homepages.uconn.edu/rossi Department of Chemistry, Room
Experimental Kinetics and Gas Phase Reactions Advanced Physical Chemistry CHAPTER 18 ELEMENTARY CHEMICAL KINETICS Professor Angelo R. Rossi http://homepages.uconn.edu/rossi Department of Chemistry, Room
UV-vis (Electronic) Spectra Ch.13 Atkins, Ch.19 Engel
 XV 74 UV-vis (Electronic) Spectra-2014 -Ch.13 Atkins, Ch.19 Engel Most broadly used analytical tech / especially bio-applic. inexpensive optics / solvent & cell usually not problem intense transitions
XV 74 UV-vis (Electronic) Spectra-2014 -Ch.13 Atkins, Ch.19 Engel Most broadly used analytical tech / especially bio-applic. inexpensive optics / solvent & cell usually not problem intense transitions
Biochemistry 3100 Sample Problems Binding proteins, Kinetics & Catalysis
 (1) Draw an approximate denaturation curve for a typical blood protein (eg myoglobin) as a function of ph. (2) Myoglobin is a simple, single subunit binding protein that has an oxygen storage function
(1) Draw an approximate denaturation curve for a typical blood protein (eg myoglobin) as a function of ph. (2) Myoglobin is a simple, single subunit binding protein that has an oxygen storage function
THE UNIVERSITY OF MANITOBA. PAPER NO: 409 LOCATION: Fr. Kennedy Gold Gym PAGE NO: 1 of 6 DEPARTMENT & COURSE NO: CHEM 4630 TIME: 3 HOURS
 PAPER NO: 409 LOCATION: Fr. Kennedy Gold Gym PAGE NO: 1 of 6 DEPARTMENT & COURSE NO: CHEM 4630 TIME: 3 HOURS EXAMINATION: Biochemistry of Proteins EXAMINER: J. O'Neil Section 1: You must answer all of
PAPER NO: 409 LOCATION: Fr. Kennedy Gold Gym PAGE NO: 1 of 6 DEPARTMENT & COURSE NO: CHEM 4630 TIME: 3 HOURS EXAMINATION: Biochemistry of Proteins EXAMINER: J. O'Neil Section 1: You must answer all of
CHM 5423 Atmospheric Chemistry Notes on kinetics (Chapter 4)
 CHM 5423 Atmospheric Chemistry Notes on kinetics (Chapter 4) Introduction A mechanism is one or a series of elementary reactions that convert reactants into products or otherwise model the chemistry of
CHM 5423 Atmospheric Chemistry Notes on kinetics (Chapter 4) Introduction A mechanism is one or a series of elementary reactions that convert reactants into products or otherwise model the chemistry of
for Single mixing applications
 RAPID KINETICS AND SPECTROSCOPY SFM-20 2 SYRINGE STOPPED-FLOW SYSTEM for Single mixing applications The SFM-20 is a high performance, modular stopped-flow system, designed for single mixing rapid kinetics
RAPID KINETICS AND SPECTROSCOPY SFM-20 2 SYRINGE STOPPED-FLOW SYSTEM for Single mixing applications The SFM-20 is a high performance, modular stopped-flow system, designed for single mixing rapid kinetics
Physical Chemistry. Chemical Kinetics
 Physical Chemistry Chemical Kinetics This chapter introduces the principles of chemical kinetics, the study of reaction rates,by showing how the rates of reactions may be measured and interpreted. The
Physical Chemistry Chemical Kinetics This chapter introduces the principles of chemical kinetics, the study of reaction rates,by showing how the rates of reactions may be measured and interpreted. The
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability
 Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
CD Basis Set of Spectra that is used is that derived from comparing the spectra of globular proteins whose secondary structures are known from X-ray
 CD Basis Set of Spectra that is used is that derived from comparing the spectra of globular proteins whose secondary structures are known from X-ray crystallography An example of the use of CD Modeling
CD Basis Set of Spectra that is used is that derived from comparing the spectra of globular proteins whose secondary structures are known from X-ray crystallography An example of the use of CD Modeling
Circular Dichroism. For students of HI Computational Structural Biology
 T H E U N I V E R S I T Y of T E X A S S C H O O L O F H E A L T H I N F O R M A T I O N S C I E N C E S A T H O U S T O N Circular Dichroism For students of HI 6001-125 Computational Structural Biology
T H E U N I V E R S I T Y of T E X A S S C H O O L O F H E A L T H I N F O R M A T I O N S C I E N C E S A T H O U S T O N Circular Dichroism For students of HI 6001-125 Computational Structural Biology
Effects of Chemical Exchange on NMR Spectra
 Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any process in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any process in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
Visible and IR Absorption Spectroscopy. Andrew Rouff and Kyle Chau
 Visible and IR Absorption Spectroscopy Andrew Rouff and Kyle Chau The Basics wavelength= (λ) original intensity= Ι o sample slab thickness= dl Final intensity= I f ε = molar extinction coefficient -di=
Visible and IR Absorption Spectroscopy Andrew Rouff and Kyle Chau The Basics wavelength= (λ) original intensity= Ι o sample slab thickness= dl Final intensity= I f ε = molar extinction coefficient -di=
To be covered (and why) Spectroscopy of Proteins. UV-Vis Absorption. UV-Vis Absorption. Spectra
 To be covered (and why) Spectroscopy of Proteins General considerations UV-Vis Absorption quantitation Fluorescence hydrophobicity Foldedness FT-Infrared Foldedness ircular Dichroism Foldedness NMR (a
To be covered (and why) Spectroscopy of Proteins General considerations UV-Vis Absorption quantitation Fluorescence hydrophobicity Foldedness FT-Infrared Foldedness ircular Dichroism Foldedness NMR (a
Theoretical Models for Chemical Kinetics
 Theoretical Models for Chemical Kinetics Thus far we have calculated rate laws, rate constants, reaction orders, etc. based on observations of macroscopic properties, but what is happening at the molecular
Theoretical Models for Chemical Kinetics Thus far we have calculated rate laws, rate constants, reaction orders, etc. based on observations of macroscopic properties, but what is happening at the molecular
ion mobility spectrometry IR spectroscopy
 Debasmita Gho 29.10.2016 Introducti on Owing to its accuracy, sensitivity, and speed, mass spectrometry (MS) coupled to fragmentation techniques is the method of choice for determining the primary structure
Debasmita Gho 29.10.2016 Introducti on Owing to its accuracy, sensitivity, and speed, mass spectrometry (MS) coupled to fragmentation techniques is the method of choice for determining the primary structure
Slow symmetric exchange
 Slow symmetric exchange ϕ A k k B t A B There are three things you should notice compared with the Figure on the previous slide: 1) The lines are broader, 2) the intensities are reduced and 3) the peaks
Slow symmetric exchange ϕ A k k B t A B There are three things you should notice compared with the Figure on the previous slide: 1) The lines are broader, 2) the intensities are reduced and 3) the peaks
Lecture 27. Transition States and Enzyme Catalysis
 Lecture 27 Transition States and Enzyme Catalysis Reading for Today: Chapter 15 sections B and C Chapter 16 next two lectures 4/8/16 1 Pop Question 9 Binding data for your thesis protein (YTP), binding
Lecture 27 Transition States and Enzyme Catalysis Reading for Today: Chapter 15 sections B and C Chapter 16 next two lectures 4/8/16 1 Pop Question 9 Binding data for your thesis protein (YTP), binding
Biology Chemistry & Physics of Biomolecules. Examination #1. Proteins Module. September 29, Answer Key
 Biology 5357 Chemistry & Physics of Biomolecules Examination #1 Proteins Module September 29, 2017 Answer Key Question 1 (A) (5 points) Structure (b) is more common, as it contains the shorter connection
Biology 5357 Chemistry & Physics of Biomolecules Examination #1 Proteins Module September 29, 2017 Answer Key Question 1 (A) (5 points) Structure (b) is more common, as it contains the shorter connection
Added topics: depend on time/rate - Skipped
 VIa- 1 Added topics: depend on time/rate - Skipped - 2012 a) diffusion rate material moves in medium faster motion more friction affect equilibrium separations like electrophoresis HPLC pharmacological
VIa- 1 Added topics: depend on time/rate - Skipped - 2012 a) diffusion rate material moves in medium faster motion more friction affect equilibrium separations like electrophoresis HPLC pharmacological
Effects of Chemical Exchange on NMR Spectra
 Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any process in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any process in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
Archives of Biochemistry and Biophysics
 Archives of Biochemistry and Biophysics 531 (2013) 24 33 Contents lists available at SciVerse ScienceDirect Archives of Biochemistry and Biophysics journal homepage: www.elsevier.com/locate/yabbi Review
Archives of Biochemistry and Biophysics 531 (2013) 24 33 Contents lists available at SciVerse ScienceDirect Archives of Biochemistry and Biophysics journal homepage: www.elsevier.com/locate/yabbi Review
Protein Folding & Stability. Lecture 11: Margaret A. Daugherty. Fall How do we go from an unfolded polypeptide chain to a
 Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2004 How do we go from an unfolded polypeptide chain to a compact folded protein? (Folding of thioredoxin, F. Richards) Structure - Function
Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2004 How do we go from an unfolded polypeptide chain to a compact folded protein? (Folding of thioredoxin, F. Richards) Structure - Function
Kinetics Mechanisms (2008-rev) Review and Examples
 II 25 Kinetics Mechanisms (2008-rev) Review and Examples Mechanism: series of elementary steps (uni-, bimolecular) that combine to give observed rate law elementary step - reaction order lie stoichiometry
II 25 Kinetics Mechanisms (2008-rev) Review and Examples Mechanism: series of elementary steps (uni-, bimolecular) that combine to give observed rate law elementary step - reaction order lie stoichiometry
Molecular Modelling. part of Bioinformatik von RNA- und Proteinstrukturen. Sonja Prohaska. Leipzig, SS Computational EvoDevo University Leipzig
 part of Bioinformatik von RNA- und Proteinstrukturen Computational EvoDevo University Leipzig Leipzig, SS 2011 Protein Structure levels or organization Primary structure: sequence of amino acids (from
part of Bioinformatik von RNA- und Proteinstrukturen Computational EvoDevo University Leipzig Leipzig, SS 2011 Protein Structure levels or organization Primary structure: sequence of amino acids (from
Singlet. Fluorescence Spectroscopy * LUMO
 Fluorescence Spectroscopy Light can be absorbed and re-emitted by matter luminescence (photo-luminescence). There are two types of luminescence, in this discussion: fluorescence and phosphorescence. A
Fluorescence Spectroscopy Light can be absorbed and re-emitted by matter luminescence (photo-luminescence). There are two types of luminescence, in this discussion: fluorescence and phosphorescence. A
Impact of β-turn Sequence on β-hairpin Dynamics Studied with Infrared-Detected Temperature Jump
 Spectroscopy: An International Journal Volume 27 (2012), Issue 5-6, Pages 557 564 doi:10.1155/2012/102423 Impact of β-turn Sequence on β-airpin Dynamics Studied with Infrared-Detected Temperature Jump
Spectroscopy: An International Journal Volume 27 (2012), Issue 5-6, Pages 557 564 doi:10.1155/2012/102423 Impact of β-turn Sequence on β-airpin Dynamics Studied with Infrared-Detected Temperature Jump
Lecture 21 (11/3/17) Protein Stability, Folding, and Dynamics Hydrophobic effect drives protein folding
 Reading: Ch4; 142-151 Problems: Ch4 (text); 14, 16 Ch6 (text); 1, 4 NEXT (after exam) Reading: Ch8; 310-312, 279-285, 285-289 Ch24; 957-961 Problems: Ch8 (text); 1,2,22 Ch8 (study-guide:facts); 1,2,3,4,5,9,10
Reading: Ch4; 142-151 Problems: Ch4 (text); 14, 16 Ch6 (text); 1, 4 NEXT (after exam) Reading: Ch8; 310-312, 279-285, 285-289 Ch24; 957-961 Problems: Ch8 (text); 1,2,22 Ch8 (study-guide:facts); 1,2,3,4,5,9,10
Protein Folding experiments and theory
 Protein Folding experiments and theory 1, 2,and 3 Protein Structure Fig. 3-16 from Lehninger Biochemistry, 4 th ed. The 3D structure is not encoded at the single aa level Hydrogen Bonding Shared H atom
Protein Folding experiments and theory 1, 2,and 3 Protein Structure Fig. 3-16 from Lehninger Biochemistry, 4 th ed. The 3D structure is not encoded at the single aa level Hydrogen Bonding Shared H atom
Lecture 11: Protein Folding & Stability
 Structure - Function Protein Folding: What we know Lecture 11: Protein Folding & Stability 1). Amino acid sequence dictates structure. 2). The native structure represents the lowest energy state for a
Structure - Function Protein Folding: What we know Lecture 11: Protein Folding & Stability 1). Amino acid sequence dictates structure. 2). The native structure represents the lowest energy state for a
Protein Folding & Stability. Lecture 11: Margaret A. Daugherty. Fall Protein Folding: What we know. Protein Folding
 Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2003 Structure - Function Protein Folding: What we know 1). Amino acid sequence dictates structure. 2). The native structure represents
Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2003 Structure - Function Protein Folding: What we know 1). Amino acid sequence dictates structure. 2). The native structure represents
Transient kinetic methods. Biophysics seminars Kinga Futó
 Transient kinetic methods Biophysics seminars Kinga Futó Fast kinetics Reaction kinetics: temporal description of reactions reactionmechanisms reaction dynamics (processes on molecular level) rate of reaction
Transient kinetic methods Biophysics seminars Kinga Futó Fast kinetics Reaction kinetics: temporal description of reactions reactionmechanisms reaction dynamics (processes on molecular level) rate of reaction
Supplemental Materials and Methods
 Supplemental Materials and Methods Time-resolved FRET (trfret) to probe for changes in the Box A/A stem upon complex assembly U3 MINI was folded and the decay of Fl fluorescence was measured at 20 ºC (see
Supplemental Materials and Methods Time-resolved FRET (trfret) to probe for changes in the Box A/A stem upon complex assembly U3 MINI was folded and the decay of Fl fluorescence was measured at 20 ºC (see
Isothermal experiments characterize time-dependent aggregation and unfolding
 1 Energy Isothermal experiments characterize time-dependent aggregation and unfolding Technical ote Introduction Kinetic measurements have, for decades, given protein scientists insight into the mechanisms
1 Energy Isothermal experiments characterize time-dependent aggregation and unfolding Technical ote Introduction Kinetic measurements have, for decades, given protein scientists insight into the mechanisms
LS1a Fall 2014 Problem Set #2 Due Monday 10/6 at 6 pm in the drop boxes on the Science Center 2 nd Floor
 LS1a Fall 2014 Problem Set #2 Due Monday 10/6 at 6 pm in the drop boxes on the Science Center 2 nd Floor Note: Adequate space is given for each answer. Questions that require a brief explanation should
LS1a Fall 2014 Problem Set #2 Due Monday 10/6 at 6 pm in the drop boxes on the Science Center 2 nd Floor Note: Adequate space is given for each answer. Questions that require a brief explanation should
Chemical kinetics in the gas phase
 Chemical kinetics in the gas phase Chemical kinetics is the study of the rates of transformation of chemical compounds from reactant species into products. The rate of a reaction is defined to be the rate
Chemical kinetics in the gas phase Chemical kinetics is the study of the rates of transformation of chemical compounds from reactant species into products. The rate of a reaction is defined to be the rate
Characterization of the Unfolding of Ribonuclease A by a Pulsed Hydrogen Exchange Study: Evidence for Competing Pathways for Unfolding
 Characterization of the Unfolding of Ribonuclease A by a Pulsed Hydrogen Exchange Study: Evidence for Competing Pathways for Unfolding Juhi Juneja and Jayant B. Udgaonkar* National Centre for Biological
Characterization of the Unfolding of Ribonuclease A by a Pulsed Hydrogen Exchange Study: Evidence for Competing Pathways for Unfolding Juhi Juneja and Jayant B. Udgaonkar* National Centre for Biological
Computational Studies of the Photoreceptor Rhodopsin. Scott E. Feller Wabash College
 Computational Studies of the Photoreceptor Rhodopsin Scott E. Feller Wabash College Rhodopsin Photocycle Dark-adapted Rhodopsin hn Isomerize retinal Photorhodopsin ~200 fs Bathorhodopsin Meta-II ms timescale
Computational Studies of the Photoreceptor Rhodopsin Scott E. Feller Wabash College Rhodopsin Photocycle Dark-adapted Rhodopsin hn Isomerize retinal Photorhodopsin ~200 fs Bathorhodopsin Meta-II ms timescale
A Thesis Presented to The Academic Faculty. Miranda McDaniel
 Proton Coupled Electron Transfer as Explored by the Tryptophan Cation Radical Formation in Biomimetic Peptides A Thesis Presented to The Academic Faculty by Miranda McDaniel In Partial Fulfillment of the
Proton Coupled Electron Transfer as Explored by the Tryptophan Cation Radical Formation in Biomimetic Peptides A Thesis Presented to The Academic Faculty by Miranda McDaniel In Partial Fulfillment of the
Chemistry 524--Final Exam--Keiderling Dec. 12, pm SES
 Chemistry 524--Final Exam--Keiderling Dec. 12, 2002 --4-8 pm -- 238 SES Please answer all questions in the answer book provided. Calculators, rulers, pens and pencils are permitted plus one 8.5 x 11 sheet
Chemistry 524--Final Exam--Keiderling Dec. 12, 2002 --4-8 pm -- 238 SES Please answer all questions in the answer book provided. Calculators, rulers, pens and pencils are permitted plus one 8.5 x 11 sheet
Outline. The ensemble folding kinetics of protein G from an all-atom Monte Carlo simulation. Unfolded Folded. What is protein folding?
 The ensemble folding kinetics of protein G from an all-atom Monte Carlo simulation By Jun Shimada and Eugine Shaknovich Bill Hawse Dr. Bahar Elisa Sandvik and Mehrdad Safavian Outline Background on protein
The ensemble folding kinetics of protein G from an all-atom Monte Carlo simulation By Jun Shimada and Eugine Shaknovich Bill Hawse Dr. Bahar Elisa Sandvik and Mehrdad Safavian Outline Background on protein
Ch 13 Rates of Reaction (Chemical Kinetics)
 Ch 13 Rates of Reaction (Chemical Kinetics) Reaction Rates and Kinetics - The reaction rate is how fast reactants are converted to products. - Chemical kinetics is the study of reaction rates. Kinetics
Ch 13 Rates of Reaction (Chemical Kinetics) Reaction Rates and Kinetics - The reaction rate is how fast reactants are converted to products. - Chemical kinetics is the study of reaction rates. Kinetics
NMR studies of protein folding
 NMR studies of protein folding Juhi Juneja and Jayant B. Udgaonkar* National Centre for Biological Sciences, Tata Institute of Fundamental Research, GKVK Campus, Bangalore 560 065, India NMR spectroscopy
NMR studies of protein folding Juhi Juneja and Jayant B. Udgaonkar* National Centre for Biological Sciences, Tata Institute of Fundamental Research, GKVK Campus, Bangalore 560 065, India NMR spectroscopy
Chapter 14, Chemical Kinetics
 Last wee we covered the following material: Review Vapor Pressure with two volatile components Chapter 14, Chemical Kinetics (continued) Quizzes next wee will be on Chap 14 through section 14.5. 13.6 Colloids
Last wee we covered the following material: Review Vapor Pressure with two volatile components Chapter 14, Chemical Kinetics (continued) Quizzes next wee will be on Chap 14 through section 14.5. 13.6 Colloids
Aleksandr V. Mikhonin, Sanford A. Asher,* Sergei V. Bykov, and Adrian Murza
 3280 J. Phys. Chem. B 2007, 111, 3280-3292 UV Raman Spatially Resolved Melting Dynamics of Isotopically Labeled Polyalanyl Peptide: Slow r-helix Melting Follows 3 10 -Helices and π-bulges Premelting Aleksandr
3280 J. Phys. Chem. B 2007, 111, 3280-3292 UV Raman Spatially Resolved Melting Dynamics of Isotopically Labeled Polyalanyl Peptide: Slow r-helix Melting Follows 3 10 -Helices and π-bulges Premelting Aleksandr
Factors That Affect Rates. Factors That Affect Rates. Factors That Affect Rates. Factors That Affect Rates
 KINETICS Kinetics Study of the speed or rate of a reaction under various conditions Thermodynamically favorable reactions DO NOT mean fast reactions Some reactions take fraction of a second (explosion)
KINETICS Kinetics Study of the speed or rate of a reaction under various conditions Thermodynamically favorable reactions DO NOT mean fast reactions Some reactions take fraction of a second (explosion)
BMB Class 17, November 30, Single Molecule Biophysics (II)
 BMB 178 2018 Class 17, November 30, 2018 15. Single Molecule Biophysics (II) New Advances in Single Molecule Techniques Atomic Force Microscopy Single Molecule Manipulation - optical traps and tweezers
BMB 178 2018 Class 17, November 30, 2018 15. Single Molecule Biophysics (II) New Advances in Single Molecule Techniques Atomic Force Microscopy Single Molecule Manipulation - optical traps and tweezers
The first assumption we will put into our theory of kinetics is that two molecules must collide for a reaction to occur between them.
 Chapter 18 Chemical Kinetics: Mechanisms In the last chapter we went through the mechanics of how you extract rate constants and order parameters from experimental data. In this chapter we will ge tthe
Chapter 18 Chemical Kinetics: Mechanisms In the last chapter we went through the mechanics of how you extract rate constants and order parameters from experimental data. In this chapter we will ge tthe
Short Announcements. 1 st Quiz today: 15 minutes. Homework 3: Due next Wednesday.
 Short Announcements 1 st Quiz today: 15 minutes Homework 3: Due next Wednesday. Next Lecture, on Visualizing Molecular Dynamics (VMD) by Klaus Schulten Today s Lecture: Protein Folding, Misfolding, Aggregation
Short Announcements 1 st Quiz today: 15 minutes Homework 3: Due next Wednesday. Next Lecture, on Visualizing Molecular Dynamics (VMD) by Klaus Schulten Today s Lecture: Protein Folding, Misfolding, Aggregation
Determining Protein Structure BIBC 100
 Determining Protein Structure BIBC 100 Determining Protein Structure X-Ray Diffraction Interactions of x-rays with electrons in molecules in a crystal NMR- Nuclear Magnetic Resonance Interactions of magnetic
Determining Protein Structure BIBC 100 Determining Protein Structure X-Ray Diffraction Interactions of x-rays with electrons in molecules in a crystal NMR- Nuclear Magnetic Resonance Interactions of magnetic
The reaction whose rate constant we are to find is the forward reaction in the following equilibrium. NH + 4 (aq) + OH (aq) K b.
 THE RATES OF CHEMICAL REACTIONS 425 E22.3a The reaction for which pk a is 9.25 is NH + 4 aq + H 2Ol NH 3 aq + H 3 O + aq. The reaction whose rate constant we are to find is the forward reaction in the
THE RATES OF CHEMICAL REACTIONS 425 E22.3a The reaction for which pk a is 9.25 is NH + 4 aq + H 2Ol NH 3 aq + H 3 O + aq. The reaction whose rate constant we are to find is the forward reaction in the
Part One: Reaction Rates. 1. Rates of chemical reactions. (how fast products are formed and/or reactants are used up)
 A. Chemical Kinetics deals with: CHAPTER 13: RATES OF REACTION Part One: Reaction Rates 1. Rates of chemical reactions. (how fast products are formed and/or reactants are used up) 2. Mechanisms of chemical
A. Chemical Kinetics deals with: CHAPTER 13: RATES OF REACTION Part One: Reaction Rates 1. Rates of chemical reactions. (how fast products are formed and/or reactants are used up) 2. Mechanisms of chemical
Macromolecule Stability Curves
 Chem728 page 1 Spring 2012 Macromolecule Stability Curves Macromolecule Transitions - We have discussed in class the factors that determine the spontaneity of processes using conformational transitions
Chem728 page 1 Spring 2012 Macromolecule Stability Curves Macromolecule Transitions - We have discussed in class the factors that determine the spontaneity of processes using conformational transitions
A) at equilibrium B) endergonic C) endothermic D) exergonic E) exothermic.
 CHEM 2770: Elements of Biochemistry Mid Term EXAMINATION VERSION A Date: October 29, 2014 Instructor: H. Perreault Location: 172 Schultz Time: 4 or 6 pm. Duration: 1 hour Instructions Please mark the Answer
CHEM 2770: Elements of Biochemistry Mid Term EXAMINATION VERSION A Date: October 29, 2014 Instructor: H. Perreault Location: 172 Schultz Time: 4 or 6 pm. Duration: 1 hour Instructions Please mark the Answer
Name: BCMB/CHEM 8190, BIOMOLECULAR NMR FINAL EXAM-5/5/10
 Name: BCMB/CHEM 8190, BIOMOLECULAR NMR FINAL EXAM-5/5/10 Instructions: This is an open book, limited time, exam. You may use notes you have from class and any text book you find useful. You may also use
Name: BCMB/CHEM 8190, BIOMOLECULAR NMR FINAL EXAM-5/5/10 Instructions: This is an open book, limited time, exam. You may use notes you have from class and any text book you find useful. You may also use
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015,
 Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Methods for the study of the conformation of folding intermediates
 7.88 Lecture Notes - 9 7.24/7.88J/5.48J The Protein Folding and Human Disease Fluorescence spectroscopy Denaturation and Denaturing agents Denatured State as a random coil (First Approx.) Renaturation/Refolding
7.88 Lecture Notes - 9 7.24/7.88J/5.48J The Protein Folding and Human Disease Fluorescence spectroscopy Denaturation and Denaturing agents Denatured State as a random coil (First Approx.) Renaturation/Refolding
T(K) k(cm 3 /molecule s) 7.37 x x x x x 10-12
 CHM 5423 Atmospheric Chemistry Problem Set 3 Due date: Tuesday, February 19 th. The first hour exam is on Thursday, February 21 st. It will cover material from the first four handouts for the class. Do
CHM 5423 Atmospheric Chemistry Problem Set 3 Due date: Tuesday, February 19 th. The first hour exam is on Thursday, February 21 st. It will cover material from the first four handouts for the class. Do
ph-jump-induced Folding and Unfolding Studies of Barstar: Evidence for Multiple Folding and Unfolding Pathways
 Biochemistry 2001, 40, 15267-15279 15267 ph-jump-induced Folding and Unfolding Studies of Barstar: Evidence for Multiple Folding and Unfolding Pathways Bhadresh R. Rami and Jayant B. Udgaonkar* National
Biochemistry 2001, 40, 15267-15279 15267 ph-jump-induced Folding and Unfolding Studies of Barstar: Evidence for Multiple Folding and Unfolding Pathways Bhadresh R. Rami and Jayant B. Udgaonkar* National
Mechanisms of Inorganic Reactions HS -26
 Mechanisms of Inorganic Reactions HS -26 A. Ligand Substitution Reactions Octahedral Co(III), Cr(III) Dissociative mechanism Square Planar Pt(II) Associative mechanism trans effect in Pt(II) complexes
Mechanisms of Inorganic Reactions HS -26 A. Ligand Substitution Reactions Octahedral Co(III), Cr(III) Dissociative mechanism Square Planar Pt(II) Associative mechanism trans effect in Pt(II) complexes
Electrochemistry & Redox. Voltaic Cells. Electrochemical Cells
 Electrochemistry & Redox An oxidation-reduction (redox) reaction involves the transfer of electrons from the reducing agent to the oxidising agent. OXIDATION - is the LOSS of electrons REDUCTION - is the
Electrochemistry & Redox An oxidation-reduction (redox) reaction involves the transfer of electrons from the reducing agent to the oxidising agent. OXIDATION - is the LOSS of electrons REDUCTION - is the
Biomolecules: lecture 10
 Biomolecules: lecture 10 - understanding in detail how protein 3D structures form - realize that protein molecules are not static wire models but instead dynamic, where in principle every atom moves (yet
Biomolecules: lecture 10 - understanding in detail how protein 3D structures form - realize that protein molecules are not static wire models but instead dynamic, where in principle every atom moves (yet
How fast reactants turn into products. Usually measured in Molarity per second units. Kinetics
 How fast reactants turn into products. Usually measured in Molarity per second units. Kinetics Reaction rated are fractions of a second for fireworks to explode. Reaction Rates takes years for a metal
How fast reactants turn into products. Usually measured in Molarity per second units. Kinetics Reaction rated are fractions of a second for fireworks to explode. Reaction Rates takes years for a metal
Membrane Proteins: 1. Integral proteins: 2. Peripheral proteins: 3. Amphitropic proteins:
 Membrane Proteins: 1. Integral proteins: proteins that insert into/span the membrane bilayer; or covalently linked to membrane lipids. (Interact with the hydrophobic part of the membrane) 2. Peripheral
Membrane Proteins: 1. Integral proteins: proteins that insert into/span the membrane bilayer; or covalently linked to membrane lipids. (Interact with the hydrophobic part of the membrane) 2. Peripheral
Proteorhodopsin phototrophy in the ocean
 Nature 411, 786-789 (14 June 2001) doi:10.1038/35081051; Received 3 January 2001; Accepted 26 March 2001 Proteorhodopsin phototrophy in the ocean Oded Béjà, Elena N. Spudich, John L. Spudich, Marion Leclerc
Nature 411, 786-789 (14 June 2001) doi:10.1038/35081051; Received 3 January 2001; Accepted 26 March 2001 Proteorhodopsin phototrophy in the ocean Oded Béjà, Elena N. Spudich, John L. Spudich, Marion Leclerc
Lecture Presentation. Chapter 14. James F. Kirby Quinnipiac University Hamden, CT. Chemical Kinetics Pearson Education, Inc.
 Lecture Presentation Chapter 14 James F. Kirby Quinnipiac University Hamden, CT In chemical kinetics we study the rate (or speed) at which a chemical process occurs. Besides information about the speed
Lecture Presentation Chapter 14 James F. Kirby Quinnipiac University Hamden, CT In chemical kinetics we study the rate (or speed) at which a chemical process occurs. Besides information about the speed
BIOPHYSICAL CHARACTERIZATION OF PROTEIN FOLDING AND MISFOLDING
 BIOPHYSICAL CHARACTERIZATION OF PROTEIN FOLDING AND MISFOLDING A Dissertation by JASON PETER SCHMITTSCHMITT Submitted to the Office of Graduate Studies of Texas A&M University in partial fulfillment of
BIOPHYSICAL CHARACTERIZATION OF PROTEIN FOLDING AND MISFOLDING A Dissertation by JASON PETER SCHMITTSCHMITT Submitted to the Office of Graduate Studies of Texas A&M University in partial fulfillment of
Modeling Background; Donald J. Jacobs, University of North Carolina at Charlotte Page 1 of 8
 Modeling Background; Donald J. Jacobs, University of North Carolina at Charlotte Page 1 of 8 Depending on thermodynamic and solvent conditions, the interrelationships between thermodynamic stability of
Modeling Background; Donald J. Jacobs, University of North Carolina at Charlotte Page 1 of 8 Depending on thermodynamic and solvent conditions, the interrelationships between thermodynamic stability of
Chem 524 Lecture Notes CD (Section 18) update 2011
 Chem 5 Lecture Notes CD (Section 8) update For HTML of 5 notes, click here XV. Circular Dichroism A. Differential absorption of left and right circular polarized light by molecular transition. Measure
Chem 5 Lecture Notes CD (Section 8) update For HTML of 5 notes, click here XV. Circular Dichroism A. Differential absorption of left and right circular polarized light by molecular transition. Measure
Biochemistry - I SPRING Mondays and Wednesdays 9:30-10:45 AM (MR-1307) Lectures 3-4. Based on Profs. Kevin Gardner & Reza Khayat
 Biochemistry - I Mondays and Wednesdays 9:30-10:45 AM (MR-1307) SPRING 2017 Lectures 3-4 Based on Profs. Kevin Gardner & Reza Khayat 1 Outline Overview of protein structure Peptide bonds Secondary structure
Biochemistry - I Mondays and Wednesdays 9:30-10:45 AM (MR-1307) SPRING 2017 Lectures 3-4 Based on Profs. Kevin Gardner & Reza Khayat 1 Outline Overview of protein structure Peptide bonds Secondary structure
Chemical Exchange and Ligand Binding
 Chemical Exchange and Ligand Binding NMR time scale Fast exchange for binding constants Slow exchange for tight binding Single vs. multiple binding mode Calcium binding process of calcium binding proteins
Chemical Exchange and Ligand Binding NMR time scale Fast exchange for binding constants Slow exchange for tight binding Single vs. multiple binding mode Calcium binding process of calcium binding proteins
Energetics and Thermodynamics
 DNA/Protein structure function analysis and prediction Protein Folding and energetics: Introduction to folding Folding and flexibility (Ch. 6) Energetics and Thermodynamics 1 Active protein conformation
DNA/Protein structure function analysis and prediction Protein Folding and energetics: Introduction to folding Folding and flexibility (Ch. 6) Energetics and Thermodynamics 1 Active protein conformation
type GroEL-GroES complex. Crystals were grown in buffer D (100 mm HEPES, ph 7.5,
 Supplementary Material Supplementary Materials and Methods Structure Determination of SR1-GroES-ADP AlF x SR1-GroES-ADP AlF x was purified as described in Materials and Methods for the wild type GroEL-GroES
Supplementary Material Supplementary Materials and Methods Structure Determination of SR1-GroES-ADP AlF x SR1-GroES-ADP AlF x was purified as described in Materials and Methods for the wild type GroEL-GroES
Chemistry Notes for class 12 Chapter 4 Chemical Kinetics
 1 P a g e Chemistry Notes for class 12 Chapter 4 Chemical Kinetics The branch of chemistry, which deals with the rate of chemical reactions. the factors affecting the rate of reactions and the mechanism
1 P a g e Chemistry Notes for class 12 Chapter 4 Chemical Kinetics The branch of chemistry, which deals with the rate of chemical reactions. the factors affecting the rate of reactions and the mechanism
Presenter: She Zhang
 Presenter: She Zhang Introduction Dr. David Baker Introduction Why design proteins de novo? It is not clear how non-covalent interactions favor one specific native structure over many other non-native
Presenter: She Zhang Introduction Dr. David Baker Introduction Why design proteins de novo? It is not clear how non-covalent interactions favor one specific native structure over many other non-native
e - Galvanic Cell 1. Voltage Sources 1.1 Polymer Electrolyte Membrane (PEM) Fuel Cell
 Galvanic cells convert different forms of energy (chemical fuel, sunlight, mechanical pressure, etc.) into electrical energy and heat. In this lecture, we are interested in some examples of galvanic cells.
Galvanic cells convert different forms of energy (chemical fuel, sunlight, mechanical pressure, etc.) into electrical energy and heat. In this lecture, we are interested in some examples of galvanic cells.
INSTRUCTOR RESOURCES
 Kinetic Studies of the Ferroin Complex INSTRUCTOR RESOURCES The CCLI Initiative Learning Objectives The purpose of this experiment is to... to determine the rate of a chemical reaction. to determine the
Kinetic Studies of the Ferroin Complex INSTRUCTOR RESOURCES The CCLI Initiative Learning Objectives The purpose of this experiment is to... to determine the rate of a chemical reaction. to determine the
Transient UV Raman Spectroscopy Finds No Crossing Barrier between the Peptide R-Helix and Fully Random Coil Conformation
 2388 J. Am. Chem. Soc. 2001, 123, 2388-2392 Transient UV Raman Spectroscopy Finds No Crossing Barrier between the Peptide R-Helix and Fully Random Coil Conformation Igor K. Lednev, Anton S. Karnoup, Mark
2388 J. Am. Chem. Soc. 2001, 123, 2388-2392 Transient UV Raman Spectroscopy Finds No Crossing Barrier between the Peptide R-Helix and Fully Random Coil Conformation Igor K. Lednev, Anton S. Karnoup, Mark
Lecture 2-3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability
 Lecture 2-3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions Van der Waals Interactions
Lecture 2-3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions Van der Waals Interactions
Part One: Reaction Rates. 1. Even though a reaction is thermodynamically favorable it may not occur at all if it is kinetically very slow.
 CHAPTER 13: RATES OF REACTION Part One: Reaction Rates A. Chemical Kinetics deals with: 1. 2. B. Importance: 1. Even though a reaction is thermodynamically favorable it may not occur at all if it is kinetically
CHAPTER 13: RATES OF REACTION Part One: Reaction Rates A. Chemical Kinetics deals with: 1. 2. B. Importance: 1. Even though a reaction is thermodynamically favorable it may not occur at all if it is kinetically
Chemistry 524--Final Exam--Keiderling May 4, :30 -?? pm SES
 Chemistry 524--Final Exam--Keiderling May 4, 2011 3:30 -?? pm -- 4286 SES Please answer all questions in the answer book provided. Calculators, rulers, pens and pencils are permitted. No open books or
Chemistry 524--Final Exam--Keiderling May 4, 2011 3:30 -?? pm -- 4286 SES Please answer all questions in the answer book provided. Calculators, rulers, pens and pencils are permitted. No open books or
NMR in Structural Biology
 NMR in Structural Biology Exercise session 2 1. a. List 3 NMR observables that report on structure. b. Also indicate whether the information they give is short/medium or long-range, or perhaps all three?
NMR in Structural Biology Exercise session 2 1. a. List 3 NMR observables that report on structure. b. Also indicate whether the information they give is short/medium or long-range, or perhaps all three?
K ex. Conformational equilibrium. equilibrium K B
 Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any yprocess in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any yprocess in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
CHEM Outline (Part 15) - Luminescence 2013
 CHEM 524 -- Outline (Part 15) - Luminescence 2013 XI. Molecular Luminescence Spectra (Chapter 15) Kinetic process, competing pathways fluorescence, phosphorescence, non-radiative decay Jablonski diagram
CHEM 524 -- Outline (Part 15) - Luminescence 2013 XI. Molecular Luminescence Spectra (Chapter 15) Kinetic process, competing pathways fluorescence, phosphorescence, non-radiative decay Jablonski diagram
Protein folding. Today s Outline
 Protein folding Today s Outline Review of previous sessions Thermodynamics of folding and unfolding Determinants of folding Techniques for measuring folding The folding process The folding problem: Prediction
Protein folding Today s Outline Review of previous sessions Thermodynamics of folding and unfolding Determinants of folding Techniques for measuring folding The folding process The folding problem: Prediction
Chemistry 1B Fall 2016
 Chemistry 1B Fall 2016 Topic 23 [more] Chemical Kinetics 1 goals for topic 23 kinetics and mechanism of chemical reaction energy profile and reaction coordinate activation energy and temperature dependence
Chemistry 1B Fall 2016 Topic 23 [more] Chemical Kinetics 1 goals for topic 23 kinetics and mechanism of chemical reaction energy profile and reaction coordinate activation energy and temperature dependence
GAS PHASE CHEMICAL KINETICS : EXPERIMENTAL ADVANCES AND PROSPECTS
 GAS PHASE CHEMICAL KINETICS : EXPERIMENTAL ADVANCES AND PROSPECTS Sébastien Le Picard France Astrophysique de Laboratoire Institut de Physique de Rennes Université de Rennes 1 Gas phase chemistry in the
GAS PHASE CHEMICAL KINETICS : EXPERIMENTAL ADVANCES AND PROSPECTS Sébastien Le Picard France Astrophysique de Laboratoire Institut de Physique de Rennes Université de Rennes 1 Gas phase chemistry in the
COMBUSTION CHEMISTRY COMBUSTION AND FUELS
 COMBUSTION CHEMISTRY CHEMICAL REACTION AND THE RATE OF REACTION General chemical reaction αa + βb = γc + δd A and B are substracts and C and are products, α, β, γ and δ are stoichiometric coefficients.
COMBUSTION CHEMISTRY CHEMICAL REACTION AND THE RATE OF REACTION General chemical reaction αa + βb = γc + δd A and B are substracts and C and are products, α, β, γ and δ are stoichiometric coefficients.
Introduction to Relaxation Theory James Keeler
 EUROMAR Zürich, 24 Introduction to Relaxation Theory James Keeler University of Cambridge Department of Chemistry What is relaxation? Why might it be interesting? relaxation is the process which drives
EUROMAR Zürich, 24 Introduction to Relaxation Theory James Keeler University of Cambridge Department of Chemistry What is relaxation? Why might it be interesting? relaxation is the process which drives
Chemical Kinetics AP Chemistry Lecture Outline
 Chemical Kinetics AP Chemistry Lecture Outline Name: Factors that govern rates of reactions. Generally... (1)...as the concentration of reactants increases, rate (2)...as temperature increases, rate (3)...with
Chemical Kinetics AP Chemistry Lecture Outline Name: Factors that govern rates of reactions. Generally... (1)...as the concentration of reactants increases, rate (2)...as temperature increases, rate (3)...with
