Gene Ontology and overrepresentation analysis
|
|
|
- Joanna Morris
- 5 years ago
- Views:
Transcription
1 Gene Ontology and overrepresentation analysis Kjell Petersen J Express Microarray analysis course Oslo December 2009 Presentation adapted from Endre Anderssen and Vidar Beisvåg NMC Trondheim
2 Overview How can ontologies (and pathway) information help us What is an ontology? The Gene Ontology and how it's structured How to use Interactively Statistically Additional things to think about
3 So here you are Figure of diff exp
4 Gene lists Long list of differentially expressed genes Possibly hundreds of papers describing the functions of the genes Misleading names Different names in dfifferent organisms
5 What s in a name? The same name can be used to describe different concepts What is a cell?
6 Cell
7 Cell
8 Cell Image from
9 Gene Ontology (GO) Ontologies Sequence Ontology (SO) (sequence features) Phenotype and Trait Ontology (PATO) Taxon (NCBI) Anatomy (Penn) Disease (ICD9) Developmental stage (multiple sources)
10 Why Gene Ontology? Gene Ontology (GO) Produce a controlled vocabulary describing aspects of molecular biology, that can be applied to all organisms. Facilitate communication between people and organization. Improve interoperability between systems.
11 Goal of GO Consortium ( Produce a controlled vocabulary describing aspects of molecular biology, that could be applied to all organism. Describe gene products using vocabulary terms (annotation). Develop tools: to query and modify the vocavularies and annotations
12 How does GO work? What information might we want to capture about a gene product? What does the gene product do? Why does it perform these activities? Where does it act?
13 The Gene Ontology (GO) Molecular function: Gene product at biochemical level. Biological process: Cellular events to which the gene product contributes. Cellular component: Location or complex of gene/protein.
14 Molecular Function activities or jobs of a gene product Insulin binding insulin transport activity
15 Molecular Function drug transporter activity
16 Biological Process a commonly recognized series of events cell division
17 Cellular Component where a gene product acts
18 Content of GO Molecular Function 7,309 terms Biological Process 10,041 terms Cellular Component 1,629 terms Total 18, 975 terms Obsolete terms: 992 As of October 2005
19 Ontology Structure Directed acyclic graphs (DAGs) Relationships is a a is a type of b (e.g. truck is a car, or mitochondrion is an organelle) Regulates Positively regulates Negatively regulates part of sub process of (process) physical part of (component) (e.g. engine is part of a car, or mitochondrion membrane is a part of a mitochondrion)
20
21 Term Definitions and Curation The definitions for each GO term are being primarily derived from the Oxford Dictionary of Molecular Biology, or from relevant literature sources (SWISS PROT, PIR, NCBI CGAP, EC...). Curators around the world shifting through genomic and proteomic data then use the definitions and GO terms provided by GO to annotate or curate the genes and proteins in their favorite species. GO is stored as flat files, as XML files and as a relational database implemented in MySQL.
22 GO Annotation Association between gene product and applicable GO terms Provided by member databases. Collaborating databases annotate their gene products (or genes) with GO terms, providing references and indicating what kind of evidence is available to support the annotations. Made by manual or automated methods. GO Annotation Database object: gene or gene product GO term ID Evidence supporting annotation Reference publication or computational method
23 Gene Ontology and Microarrays Hypothesis: Functionally related, differentially expressed genes should accumulate in the corresponding GO group. Problem: Find a method, which scores accumulation of differential gene expression in a node of the Gene Ontology. GO tools can be important in order to answer questions such as: are genes involved in process P overrepresented among the total of differentially expressed genes in an experiment or does treatment A induce more genes involved in process P than treatment B?".
24 Browsing GO in J Express
25 Overrepresentation of GO terms We have a subset of genes List of differentially expressed genes List of genes that cluster together Which biological processes do these genes take part in? Is there an over representation of the number of genes belonging to a particular biological process, compared to what could be expected?
26 Question If we look at the dataset containing all of our genes and see that 10% of these belong to cell cycle. We then do a differentially expressed genes analysis and get a list of genes we believe are significantly changed. How many of the genes in the gene list do you expect belong to cell cycle?
27 Setup We name our subset of interesting genes for test data Reference data And the dataset containing all of our genes, the dataset we extracted the interesting genes Test datafrom and that we want to compare our testdata to, for reference data
28 Gene Ontology Analysis Reference data Test data Statistical comparison between the two GO components
29 Some gotchas P value not corrected for multiple testing Bonferroni correction: multiply by number of terms mapped to your dataset (sum of 3 blue numbers at the 3 top terms) Results depend on Selected cut off for test data (check for consistency over several cut offs) Version of GO obo and association files (keep track of which used) Browse the terms in the neighbourhood of high scoring terms, will often reveal more context.
30 Nice to know Alternative term hierarchies / tools GO slim Panther DB ( DAVID (
Genome Annotation. Bioinformatics and Computational Biology. Genome sequencing Assembly. Gene prediction. Protein targeting.
 Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
Gene Ontology. Shifra Ben-Dor. Weizmann Institute of Science
 Gene Ontology Shifra Ben-Dor Weizmann Institute of Science Outline of Session What is GO (Gene Ontology)? What tools do we use to work with it? Combination of GO with other analyses What is Ontology? 1700s
Gene Ontology Shifra Ben-Dor Weizmann Institute of Science Outline of Session What is GO (Gene Ontology)? What tools do we use to work with it? Combination of GO with other analyses What is Ontology? 1700s
2 GENE FUNCTIONAL SIMILARITY. 2.1 Semantic values of GO terms
 Bioinformatics Advance Access published March 7, 2007 The Author (2007). Published by Oxford University Press. All rights reserved. For Permissions, please email: journals.permissions@oxfordjournals.org
Bioinformatics Advance Access published March 7, 2007 The Author (2007). Published by Oxford University Press. All rights reserved. For Permissions, please email: journals.permissions@oxfordjournals.org
GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón
 GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón What is GO? The Gene Ontology (GO) project is a collaborative effort to address the need for consistent descriptions of gene products in
GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón What is GO? The Gene Ontology (GO) project is a collaborative effort to address the need for consistent descriptions of gene products in
BMD645. Integration of Omics
 BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
Gene Ontology and Functional Enrichment. Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein
 Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Integration of functional genomics data
 Integration of functional genomics data Laboratoire Bordelais de Recherche en Informatique (UMR) Centre de Bioinformatique de Bordeaux (Plateforme) Rennes Oct. 2006 1 Observations and motivations Genomics
Integration of functional genomics data Laboratoire Bordelais de Recherche en Informatique (UMR) Centre de Bioinformatique de Bordeaux (Plateforme) Rennes Oct. 2006 1 Observations and motivations Genomics
Francisco M. Couto Mário J. Silva Pedro Coutinho
 Francisco M. Couto Mário J. Silva Pedro Coutinho DI FCUL TR 03 29 Departamento de Informática Faculdade de Ciências da Universidade de Lisboa Campo Grande, 1749 016 Lisboa Portugal Technical reports are
Francisco M. Couto Mário J. Silva Pedro Coutinho DI FCUL TR 03 29 Departamento de Informática Faculdade de Ciências da Universidade de Lisboa Campo Grande, 1749 016 Lisboa Portugal Technical reports are
ToxiCat: Hybrid Named Entity Recognition services to support curation of the Comparative Toxicogenomic Database
 ToxiCat: Hybrid Named Entity Recognition services to support curation of the Comparative Toxicogenomic Database Dina Vishnyakova 1,2, 4, *, Julien Gobeill 1,3,4, Emilie Pasche 1,2,3,4 and Patrick Ruch
ToxiCat: Hybrid Named Entity Recognition services to support curation of the Comparative Toxicogenomic Database Dina Vishnyakova 1,2, 4, *, Julien Gobeill 1,3,4, Emilie Pasche 1,2,3,4 and Patrick Ruch
Analysis and visualization of protein-protein interactions. Olga Vitek Assistant Professor Statistics and Computer Science
 1 Analysis and visualization of protein-protein interactions Olga Vitek Assistant Professor Statistics and Computer Science 2 Outline 1. Protein-protein interactions 2. Using graph structures to study
1 Analysis and visualization of protein-protein interactions Olga Vitek Assistant Professor Statistics and Computer Science 2 Outline 1. Protein-protein interactions 2. Using graph structures to study
Lecture 2. The Blast2GO annotation framework
 Lecture 2 The Blast2GO annotation framework Annotation steps Modulation of annotation intensity Export/Import Functions Sequence Selection Additional Tools Functional assignment Annotation Transference
Lecture 2 The Blast2GO annotation framework Annotation steps Modulation of annotation intensity Export/Import Functions Sequence Selection Additional Tools Functional assignment Annotation Transference
CSCE555 Bioinformatics. Protein Function Annotation
 CSCE555 Bioinformatics Protein Function Annotation Why we need to do function annotation? Fig from: Network-based prediction of protein function. Molecular Systems Biology 3:88. 2007 What s function? The
CSCE555 Bioinformatics Protein Function Annotation Why we need to do function annotation? Fig from: Network-based prediction of protein function. Molecular Systems Biology 3:88. 2007 What s function? The
Supplementary Information 16
 Supplementary Information 16 Cellular Component % of Genes 50 45 40 35 30 25 20 15 10 5 0 human mouse extracellular other membranes plasma membrane cytosol cytoskeleton mitochondrion ER/Golgi translational
Supplementary Information 16 Cellular Component % of Genes 50 45 40 35 30 25 20 15 10 5 0 human mouse extracellular other membranes plasma membrane cytosol cytoskeleton mitochondrion ER/Golgi translational
SABIO-RK Integration and Curation of Reaction Kinetics Data Ulrike Wittig
 SABIO-RK Integration and Curation of Reaction Kinetics Data http://sabio.villa-bosch.de/sabiork Ulrike Wittig Overview Introduction /Motivation Database content /User interface Data integration Curation
SABIO-RK Integration and Curation of Reaction Kinetics Data http://sabio.villa-bosch.de/sabiork Ulrike Wittig Overview Introduction /Motivation Database content /User interface Data integration Curation
86 Part 4 SUMMARY INTRODUCTION
 86 Part 4 Chapter # AN INTEGRATION OF THE DESCRIPTIONS OF GENE NETWORKS AND THEIR MODELS PRESENTED IN SIGMOID (CELLERATOR) AND GENENET Podkolodny N.L. *1, 2, Podkolodnaya N.N. 1, Miginsky D.S. 1, Poplavsky
86 Part 4 Chapter # AN INTEGRATION OF THE DESCRIPTIONS OF GENE NETWORKS AND THEIR MODELS PRESENTED IN SIGMOID (CELLERATOR) AND GENENET Podkolodny N.L. *1, 2, Podkolodnaya N.N. 1, Miginsky D.S. 1, Poplavsky
Introduction to the EMBL-EBI Ontology Lookup Service
 Introduction to the EMBL-EBI Ontology Lookup Service Simon Jupp jupp@ebi.ac.uk, @simonjupp Samples, Phenotypes and Ontologies Team European Bioinformatics Institute Cambridge, UK. Ontologies in the life
Introduction to the EMBL-EBI Ontology Lookup Service Simon Jupp jupp@ebi.ac.uk, @simonjupp Samples, Phenotypes and Ontologies Team European Bioinformatics Institute Cambridge, UK. Ontologies in the life
Biological Concepts and Information Technology (Systems Biology)
 Biological Concepts and Information Technology (Systems Biology) Janaina de Andréa Dernowsek Postdoctoral at Center for Information Technology Renato Archer Janaina.dernowsek@cti.gov.br Division of 3D
Biological Concepts and Information Technology (Systems Biology) Janaina de Andréa Dernowsek Postdoctoral at Center for Information Technology Renato Archer Janaina.dernowsek@cti.gov.br Division of 3D
Annotation Error in Public Databases ALEXANDRA SCHNOES UNIVERSITY OF CALIFORNIA, SAN FRANCISCO OCTOBER 25, 2010
 Annotation Error in Public Databases ALEXANDRA SCHNOES UNIVERSITY OF CALIFORNIA, SAN FRANCISCO OCTOBER 25, 2010 1 New genomes (and metagenomes) sequenced every day... 2 3 3 3 3 3 3 3 3 3 Computational
Annotation Error in Public Databases ALEXANDRA SCHNOES UNIVERSITY OF CALIFORNIA, SAN FRANCISCO OCTOBER 25, 2010 1 New genomes (and metagenomes) sequenced every day... 2 3 3 3 3 3 3 3 3 3 Computational
Ontology Alignment in the Presence of a Domain Ontology
 Ontology Alignment in the Presence of a Domain Ontology Finding Protein Homology by Andrew August Carbonetto B.Sc., McGill University, 2005 A THESIS SUBMITTED IN PARTIAL FULFILMENT OF THE REQUIREMENTS
Ontology Alignment in the Presence of a Domain Ontology Finding Protein Homology by Andrew August Carbonetto B.Sc., McGill University, 2005 A THESIS SUBMITTED IN PARTIAL FULFILMENT OF THE REQUIREMENTS
Introduction to Bioinformatics
 CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
Learning in Bayesian Networks
 Learning in Bayesian Networks Florian Markowetz Max-Planck-Institute for Molecular Genetics Computational Molecular Biology Berlin Berlin: 20.06.2002 1 Overview 1. Bayesian Networks Stochastic Networks
Learning in Bayesian Networks Florian Markowetz Max-Planck-Institute for Molecular Genetics Computational Molecular Biology Berlin Berlin: 20.06.2002 1 Overview 1. Bayesian Networks Stochastic Networks
Cells The Units of Life
 Before You Read Before you read the chapter, respond to these statements. Write an A if you agree with the statement. Write a D if you disagree with the statement. Before You Read Bacteria are the smallest
Before You Read Before you read the chapter, respond to these statements. Write an A if you agree with the statement. Write a D if you disagree with the statement. Before You Read Bacteria are the smallest
Networks & pathways. Hedi Peterson MTAT Bioinformatics
 Networks & pathways Hedi Peterson (peterson@quretec.com) MTAT.03.239 Bioinformatics 03.11.2010 Networks are graphs Nodes Edges Edges Directed, undirected, weighted Nodes Genes Proteins Metabolites Enzymes
Networks & pathways Hedi Peterson (peterson@quretec.com) MTAT.03.239 Bioinformatics 03.11.2010 Networks are graphs Nodes Edges Edges Directed, undirected, weighted Nodes Genes Proteins Metabolites Enzymes
Bioinformatics. Dept. of Computational Biology & Bioinformatics
 Bioinformatics Dept. of Computational Biology & Bioinformatics 3 Bioinformatics - play with sequences & structures Dept. of Computational Biology & Bioinformatics 4 ORGANIZATION OF LIFE ROLE OF BIOINFORMATICS
Bioinformatics Dept. of Computational Biology & Bioinformatics 3 Bioinformatics - play with sequences & structures Dept. of Computational Biology & Bioinformatics 4 ORGANIZATION OF LIFE ROLE OF BIOINFORMATICS
Computational approaches for functional genomics
 Computational approaches for functional genomics Kalin Vetsigian October 31, 2001 The rapidly increasing number of completely sequenced genomes have stimulated the development of new methods for finding
Computational approaches for functional genomics Kalin Vetsigian October 31, 2001 The rapidly increasing number of completely sequenced genomes have stimulated the development of new methods for finding
The EcoCyc Database. January 25, de Nitrógeno, UNAM,Cuernavaca, A.P. 565-A, Morelos, 62100, Mexico;
 The EcoCyc Database Peter D. Karp, Monica Riley, Milton Saier,IanT.Paulsen +, Julio Collado-Vides + Suzanne M. Paley, Alida Pellegrini-Toole,César Bonavides ++, and Socorro Gama-Castro ++ January 25, 2002
The EcoCyc Database Peter D. Karp, Monica Riley, Milton Saier,IanT.Paulsen +, Julio Collado-Vides + Suzanne M. Paley, Alida Pellegrini-Toole,César Bonavides ++, and Socorro Gama-Castro ++ January 25, 2002
Predicting Protein Functions and Domain Interactions from Protein Interactions
 Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Gabriella Rustici EMBL-EBI
 Towards a cellular phenotype ontology Gabriella Rustici EMBL-EBI gabry@ebi.ac.uk Systems Microscopy NoE at a glance The Systems Microscopy NoE (2011 2015) is a FP7 funded project, involving 15 research
Towards a cellular phenotype ontology Gabriella Rustici EMBL-EBI gabry@ebi.ac.uk Systems Microscopy NoE at a glance The Systems Microscopy NoE (2011 2015) is a FP7 funded project, involving 15 research
CS612 - Algorithms in Bioinformatics
 Fall 2017 Databases and Protein Structure Representation October 2, 2017 Molecular Biology as Information Science > 12, 000 genomes sequenced, mostly bacterial (2013) > 5x10 6 unique sequences available
Fall 2017 Databases and Protein Structure Representation October 2, 2017 Molecular Biology as Information Science > 12, 000 genomes sequenced, mostly bacterial (2013) > 5x10 6 unique sequences available
Computational Genomics. Systems biology. Putting it together: Data integration using graphical models
 02-710 Computational Genomics Systems biology Putting it together: Data integration using graphical models High throughput data So far in this class we discussed several different types of high throughput
02-710 Computational Genomics Systems biology Putting it together: Data integration using graphical models High throughput data So far in this class we discussed several different types of high throughput
Science Online Instructional Materials Correlation to the 2010 Biology Standards of Learning and Curriculum Framework
 and Curriculum Framework Provider York County School Divison Course Title Biology Last Updated 2010-11 Course Syllabus URL http://yorkcountyschools.org/virtuallearning/coursecatalog.aspx BIO.1 The student
and Curriculum Framework Provider York County School Divison Course Title Biology Last Updated 2010-11 Course Syllabus URL http://yorkcountyschools.org/virtuallearning/coursecatalog.aspx BIO.1 The student
Hands-On Nine The PAX6 Gene and Protein
 Hands-On Nine The PAX6 Gene and Protein Main Purpose of Hands-On Activity: Using bioinformatics tools to examine the sequences, homology, and disease relevance of the Pax6: a master gene of eye formation.
Hands-On Nine The PAX6 Gene and Protein Main Purpose of Hands-On Activity: Using bioinformatics tools to examine the sequences, homology, and disease relevance of the Pax6: a master gene of eye formation.
A Protein Ontology from Large-scale Textmining?
 A Protein Ontology from Large-scale Textmining? Protege-Workshop Manchester, 07-07-2003 Kai Kumpf, Juliane Fluck and Martin Hofmann Instructive mistakes: a narrative Aim: Protein ontology that supports
A Protein Ontology from Large-scale Textmining? Protege-Workshop Manchester, 07-07-2003 Kai Kumpf, Juliane Fluck and Martin Hofmann Instructive mistakes: a narrative Aim: Protein ontology that supports
Science Textbook and Instructional Materials Correlation to the 2010 Biology Standards of Learning and Curriculum Framework. Publisher Information
 Publisher Information Copyright date 2013 Contact Carol Kornfeind Phone# 847-486-2065 E-mail carol.kornfeind@pearson.com Biology 1 of 12 Virginia Department of Education Text Miller Levine Biology, Virginia
Publisher Information Copyright date 2013 Contact Carol Kornfeind Phone# 847-486-2065 E-mail carol.kornfeind@pearson.com Biology 1 of 12 Virginia Department of Education Text Miller Levine Biology, Virginia
A Study of Correlations between the Definition and Application of the Gene Ontology
 University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Theses, Dissertations, & Student Research in Computer Electronics & Engineering Electrical & Computer Engineering, Department
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Theses, Dissertations, & Student Research in Computer Electronics & Engineering Electrical & Computer Engineering, Department
What is Systems Biology
 What is Systems Biology 2 CBS, Department of Systems Biology 3 CBS, Department of Systems Biology Data integration In the Big Data era Combine different types of data, describing different things or the
What is Systems Biology 2 CBS, Department of Systems Biology 3 CBS, Department of Systems Biology Data integration In the Big Data era Combine different types of data, describing different things or the
Research Article HomoKinase: A Curated Database of Human Protein Kinases
 ISRN Computational Biology Volume 2013, Article ID 417634, 5 pages http://dx.doi.org/10.1155/2013/417634 Research Article HomoKinase: A Curated Database of Human Protein Kinases Suresh Subramani, Saranya
ISRN Computational Biology Volume 2013, Article ID 417634, 5 pages http://dx.doi.org/10.1155/2013/417634 Research Article HomoKinase: A Curated Database of Human Protein Kinases Suresh Subramani, Saranya
Lab 2 Worksheet. Problems. Problem 1: Geometry and Linear Equations
 Lab 2 Worksheet Problems Problem : Geometry and Linear Equations Linear algebra is, first and foremost, the study of systems of linear equations. You are going to encounter linear systems frequently in
Lab 2 Worksheet Problems Problem : Geometry and Linear Equations Linear algebra is, first and foremost, the study of systems of linear equations. You are going to encounter linear systems frequently in
Lecture 3: A basic statistical concept
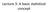 Lecture 3: A basic statistical concept P value In statistical hypothesis testing, the p value is the probability of obtaining a result at least as extreme as the one that was actually observed, assuming
Lecture 3: A basic statistical concept P value In statistical hypothesis testing, the p value is the probability of obtaining a result at least as extreme as the one that was actually observed, assuming
A Database of human biological pathways
 A Database of human biological pathways Steve Jupe - sjupe@ebi.ac.uk 1 Rationale Journal information Nature 407(6805):770-6.The Biochemistry of Apoptosis. Caspase-8 is the key initiator caspase in the
A Database of human biological pathways Steve Jupe - sjupe@ebi.ac.uk 1 Rationale Journal information Nature 407(6805):770-6.The Biochemistry of Apoptosis. Caspase-8 is the key initiator caspase in the
EBI web resources II: Ensembl and InterPro. Yanbin Yin Spring 2013
 EBI web resources II: Ensembl and InterPro Yanbin Yin Spring 2013 1 Outline Intro to genome annotation Protein family/domain databases InterPro, Pfam, Superfamily etc. Genome browser Ensembl Hands on Practice
EBI web resources II: Ensembl and InterPro Yanbin Yin Spring 2013 1 Outline Intro to genome annotation Protein family/domain databases InterPro, Pfam, Superfamily etc. Genome browser Ensembl Hands on Practice
Proteomics. Yeast two hybrid. Proteomics - PAGE techniques. Data obtained. What is it?
 Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
Warm Up. What are some examples of living things? Describe the characteristics of living things
 Warm Up What are some examples of living things? Describe the characteristics of living things Objectives Identify the levels of biological organization and explain their relationships Describe cell structure
Warm Up What are some examples of living things? Describe the characteristics of living things Objectives Identify the levels of biological organization and explain their relationships Describe cell structure
Goal 1: Develop knowledge and understanding of core content in biology
 Indiana State University» College of Arts & Sciences» Biology BA/BS in Biology Standing Requirements s Library Goal 1: Develop knowledge and understanding of core content in biology 1: Illustrate and examine
Indiana State University» College of Arts & Sciences» Biology BA/BS in Biology Standing Requirements s Library Goal 1: Develop knowledge and understanding of core content in biology 1: Illustrate and examine
Outline. Terminologies and Ontologies. Communication and Computation. Communication. Outline. Terminologies and Vocabularies.
 Page 1 Outline 1. Why do we need terminologies and ontologies? Terminologies and Ontologies Iwei Yeh yeh@smi.stanford.edu 04/16/2002 2. Controlled Terminologies Enzyme Classification Gene Ontology 3. Ontologies
Page 1 Outline 1. Why do we need terminologies and ontologies? Terminologies and Ontologies Iwei Yeh yeh@smi.stanford.edu 04/16/2002 2. Controlled Terminologies Enzyme Classification Gene Ontology 3. Ontologies
ATLAS of Biochemistry
 ATLAS of Biochemistry USER GUIDE http://lcsb-databases.epfl.ch/atlas/ CONTENT 1 2 3 GET STARTED Create your user account NAVIGATE Curated KEGG reactions ATLAS reactions Pathways Maps USE IT! Fill a gap
ATLAS of Biochemistry USER GUIDE http://lcsb-databases.epfl.ch/atlas/ CONTENT 1 2 3 GET STARTED Create your user account NAVIGATE Curated KEGG reactions ATLAS reactions Pathways Maps USE IT! Fill a gap
ADVANCED PLACEMENT BIOLOGY
 ADVANCED PLACEMENT BIOLOGY Description Advanced Placement Biology is designed to be the equivalent of a two-semester college introductory course for Biology majors. The course meets seven periods per week
ADVANCED PLACEMENT BIOLOGY Description Advanced Placement Biology is designed to be the equivalent of a two-semester college introductory course for Biology majors. The course meets seven periods per week
Updated: 10/11/2018 Page 1 of 5
 A. Academic Division: Health Sciences B. Discipline: Biology C. Course Number and Title: BIOL1230 Biology I MASTER SYLLABUS 2018-2019 D. Course Coordinator: Justin Tickhill Assistant Dean: Melinda Roepke,
A. Academic Division: Health Sciences B. Discipline: Biology C. Course Number and Title: BIOL1230 Biology I MASTER SYLLABUS 2018-2019 D. Course Coordinator: Justin Tickhill Assistant Dean: Melinda Roepke,
Grundlagen der Bioinformatik Summer semester Lecturer: Prof. Daniel Huson
 Grundlagen der Bioinformatik, SS 10, D. Huson, April 12, 2010 1 1 Introduction Grundlagen der Bioinformatik Summer semester 2010 Lecturer: Prof. Daniel Huson Office hours: Thursdays 17-18h (Sand 14, C310a)
Grundlagen der Bioinformatik, SS 10, D. Huson, April 12, 2010 1 1 Introduction Grundlagen der Bioinformatik Summer semester 2010 Lecturer: Prof. Daniel Huson Office hours: Thursdays 17-18h (Sand 14, C310a)
Comparative Network Analysis
 Comparative Network Analysis BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by
Comparative Network Analysis BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by
The Role of Network Science in Biology and Medicine. Tiffany J. Callahan Computational Bioscience Program Hunter/Kahn Labs
 The Role of Network Science in Biology and Medicine Tiffany J. Callahan Computational Bioscience Program Hunter/Kahn Labs Network Analysis Working Group 09.28.2017 Network-Enabled Wisdom (NEW) empirically
The Role of Network Science in Biology and Medicine Tiffany J. Callahan Computational Bioscience Program Hunter/Kahn Labs Network Analysis Working Group 09.28.2017 Network-Enabled Wisdom (NEW) empirically
FCModeler: Dynamic Graph Display and Fuzzy Modeling of Regulatory and Metabolic Maps
 FCModeler: Dynamic Graph Display and Fuzzy Modeling of Regulatory and Metabolic Maps Julie Dickerson 1, Zach Cox 1 and Andy Fulmer 2 1 Iowa State University and 2 Proctor & Gamble. FCModeler Goals Capture
FCModeler: Dynamic Graph Display and Fuzzy Modeling of Regulatory and Metabolic Maps Julie Dickerson 1, Zach Cox 1 and Andy Fulmer 2 1 Iowa State University and 2 Proctor & Gamble. FCModeler Goals Capture
EBI web resources II: Ensembl and InterPro
 EBI web resources II: Ensembl and InterPro Yanbin Yin http://www.ebi.ac.uk/training/online/course/ 1 Homework 3 Go to http://www.ebi.ac.uk/interpro/training.htmland finish the second online training course
EBI web resources II: Ensembl and InterPro Yanbin Yin http://www.ebi.ac.uk/training/online/course/ 1 Homework 3 Go to http://www.ebi.ac.uk/interpro/training.htmland finish the second online training course
The geneticist s questions
 The geneticist s questions a) What is consequence of reduced gene function? 1) gene knockout (deletion, RNAi) b) What is the consequence of increased gene function? 2) gene overexpression c) What does
The geneticist s questions a) What is consequence of reduced gene function? 1) gene knockout (deletion, RNAi) b) What is the consequence of increased gene function? 2) gene overexpression c) What does
Student Handout Fruit Fly Ethomics & Genomics
 Student Handout Fruit Fly Ethomics & Genomics Summary of Laboratory Exercise In this laboratory unit, students will connect behavioral phenotypes to their underlying genes and molecules in the model genetic
Student Handout Fruit Fly Ethomics & Genomics Summary of Laboratory Exercise In this laboratory unit, students will connect behavioral phenotypes to their underlying genes and molecules in the model genetic
Cross Discipline Analysis made possible with Data Pipelining. J.R. Tozer SciTegic
 Cross Discipline Analysis made possible with Data Pipelining J.R. Tozer SciTegic System Genesis Pipelining tool created to automate data processing in cheminformatics Modular system built with generic
Cross Discipline Analysis made possible with Data Pipelining J.R. Tozer SciTegic System Genesis Pipelining tool created to automate data processing in cheminformatics Modular system built with generic
Carri-Lyn Mead Thursday, January 13, 2005 Terry Fox Laboratory, Dr. Dixie Mager
 Investigating Trends in Transposable Element Insertion within Regulatory Regions Carri-Lyn Mead cmead@bcgsc.ca Thursday, January 13, 2005 Terry Fox Laboratory, Dr. Dixie Mager Outline Transposable Element
Investigating Trends in Transposable Element Insertion within Regulatory Regions Carri-Lyn Mead cmead@bcgsc.ca Thursday, January 13, 2005 Terry Fox Laboratory, Dr. Dixie Mager Outline Transposable Element
Total
 Student Performance by Question Biology (Multiple-Choice ONLY) Teacher: Core 1 / S-14 Scientific Investigation Life at the Molecular and Cellular Level Analysis of Performance by Question of each student
Student Performance by Question Biology (Multiple-Choice ONLY) Teacher: Core 1 / S-14 Scientific Investigation Life at the Molecular and Cellular Level Analysis of Performance by Question of each student
The MANTiS Manual. Contents. MANTiS Version 1.1
 The MANTiS Manual MANTiS Version 1.1 Contents Connection to the MANTiS database... 2 Memory settings... 2 Main functionalities... 2 Character Mapping View... 4 Genome content View... 5 Biological processes
The MANTiS Manual MANTiS Version 1.1 Contents Connection to the MANTiS database... 2 Memory settings... 2 Main functionalities... 2 Character Mapping View... 4 Genome content View... 5 Biological processes
Proteomics Systems Biology
 Dr. Sanjeeva Srivastava IIT Bombay Proteomics Systems Biology IIT Bombay 2 1 DNA Genomics RNA Transcriptomics Global Cellular Protein Proteomics Global Cellular Metabolite Metabolomics Global Cellular
Dr. Sanjeeva Srivastava IIT Bombay Proteomics Systems Biology IIT Bombay 2 1 DNA Genomics RNA Transcriptomics Global Cellular Protein Proteomics Global Cellular Metabolite Metabolomics Global Cellular
Bioinformatics 2. Yeast two hybrid. Proteomics. Proteomics
 GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
Computational methods for predicting protein-protein interactions
 Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Molecular and cellular biology is about studying cell structure and function
 Chapter 1 Exploring the World of the Cell In This Chapter Discovering the microscopic world Getting matter and energy Reading the genetic code Molecular and cellular biology is about studying cell structure
Chapter 1 Exploring the World of the Cell In This Chapter Discovering the microscopic world Getting matter and energy Reading the genetic code Molecular and cellular biology is about studying cell structure
Chetek-Weyerhaeuser High School
 Chetek-Weyerhaeuser High School Unit 1 The Science of Biology (5 days) Biology I Units and s Biology I A s 1. I can design a scientific experiment that includes a control group, experimental group, constants,
Chetek-Weyerhaeuser High School Unit 1 The Science of Biology (5 days) Biology I Units and s Biology I A s 1. I can design a scientific experiment that includes a control group, experimental group, constants,
RCPS Curriculum Pacing Guide Subject: Biology. Remembering, Understanding, Applying, Analyzing, Evaluating, Creating
 RCPS Curriculum Pacing Guide 2013 2014 Subject: Biology Week of: SOL # Unit Bloom s Objectives Week 1 and throughout the semester #BIO1 Scientific reasoning, logic and the nature of science Chapter 1 Biology:
RCPS Curriculum Pacing Guide 2013 2014 Subject: Biology Week of: SOL # Unit Bloom s Objectives Week 1 and throughout the semester #BIO1 Scientific reasoning, logic and the nature of science Chapter 1 Biology:
Dynamical Modeling in Biology: a semiotic perspective. Junior Barrera BIOINFO-USP
 Dynamical Modeling in Biology: a semiotic perspective Junior Barrera BIOINFO-USP Layout Introduction Dynamical Systems System Families System Identification Genetic networks design Cell Cycle Modeling
Dynamical Modeling in Biology: a semiotic perspective Junior Barrera BIOINFO-USP Layout Introduction Dynamical Systems System Families System Identification Genetic networks design Cell Cycle Modeling
CISC 636 Computational Biology & Bioinformatics (Fall 2016)
 CISC 636 Computational Biology & Bioinformatics (Fall 2016) Predicting Protein-Protein Interactions CISC636, F16, Lec22, Liao 1 Background Proteins do not function as isolated entities. Protein-Protein
CISC 636 Computational Biology & Bioinformatics (Fall 2016) Predicting Protein-Protein Interactions CISC636, F16, Lec22, Liao 1 Background Proteins do not function as isolated entities. Protein-Protein
Bayesian Hierarchical Classification. Seminar on Predicting Structured Data Jukka Kohonen
 Bayesian Hierarchical Classification Seminar on Predicting Structured Data Jukka Kohonen 17.4.2008 Overview Intro: The task of hierarchical gene annotation Approach I: SVM/Bayes hybrid Barutcuoglu et al:
Bayesian Hierarchical Classification Seminar on Predicting Structured Data Jukka Kohonen 17.4.2008 Overview Intro: The task of hierarchical gene annotation Approach I: SVM/Bayes hybrid Barutcuoglu et al:
Browsing Genomic Information with Ensembl Plants
 Browsing Genomic Information with Ensembl Plants Etienne de Villiers, PhD (Adapted from slides by Bert Overduin EMBL-EBI) Outline of workshop Brief introduction to Ensembl Plants History Content Tutorial
Browsing Genomic Information with Ensembl Plants Etienne de Villiers, PhD (Adapted from slides by Bert Overduin EMBL-EBI) Outline of workshop Brief introduction to Ensembl Plants History Content Tutorial
Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis
 Title Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis Author list Yu Han 1, Huihua Wan 1, Tangren Cheng 1, Jia Wang 1, Weiru Yang 1, Huitang Pan 1* & Qixiang
Title Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis Author list Yu Han 1, Huihua Wan 1, Tangren Cheng 1, Jia Wang 1, Weiru Yang 1, Huitang Pan 1* & Qixiang
Computational Biology Course Descriptions 12-14
 Computational Biology Course Descriptions 12-14 Course Number and Title INTRODUCTORY COURSES BIO 311C: Introductory Biology I BIO 311D: Introductory Biology II BIO 325: Genetics CH 301: Principles of Chemistry
Computational Biology Course Descriptions 12-14 Course Number and Title INTRODUCTORY COURSES BIO 311C: Introductory Biology I BIO 311D: Introductory Biology II BIO 325: Genetics CH 301: Principles of Chemistry
BIOINFORMATICS: An Introduction
 BIOINFORMATICS: An Introduction What is Bioinformatics? The term was first coined in 1988 by Dr. Hwa Lim The original definition was : a collective term for data compilation, organisation, analysis and
BIOINFORMATICS: An Introduction What is Bioinformatics? The term was first coined in 1988 by Dr. Hwa Lim The original definition was : a collective term for data compilation, organisation, analysis and
Differential Modeling for Cancer Microarray Data
 Differential Modeling for Cancer Microarray Data Omar Odibat Department of Computer Science Feb, 01, 2011 1 Outline Introduction Cancer Microarray data Problem Definition Differential analysis Existing
Differential Modeling for Cancer Microarray Data Omar Odibat Department of Computer Science Feb, 01, 2011 1 Outline Introduction Cancer Microarray data Problem Definition Differential analysis Existing
Discovering molecular pathways from protein interaction and ge
 Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
BIOLOGY 111. CHAPTER 1: An Introduction to the Science of Life
 BIOLOGY 111 CHAPTER 1: An Introduction to the Science of Life An Introduction to the Science of Life: Chapter Learning Outcomes 1.1) Describe the properties of life common to all living things. (Module
BIOLOGY 111 CHAPTER 1: An Introduction to the Science of Life An Introduction to the Science of Life: Chapter Learning Outcomes 1.1) Describe the properties of life common to all living things. (Module
Erzsébet Ravasz Advisor: Albert-László Barabási
 Hierarchical Networks Erzsébet Ravasz Advisor: Albert-László Barabási Introduction to networks How to model complex networks? Clustering and hierarchy Hierarchical organization of cellular metabolism The
Hierarchical Networks Erzsébet Ravasz Advisor: Albert-László Barabási Introduction to networks How to model complex networks? Clustering and hierarchy Hierarchical organization of cellular metabolism The
10-810: Advanced Algorithms and Models for Computational Biology. microrna and Whole Genome Comparison
 10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
A taxonomy of visualization tasks for the analysis of biological pathway data
 The Author(s) BMC Bioinformatics 2016, 18(Suppl 2):21 DOI 10.1186/s12859-016-1443-5 RESEARCH A taxonomy of visualization tasks for the analysis of biological pathway data Paul Murray 1*,FintanMcGee 2 and
The Author(s) BMC Bioinformatics 2016, 18(Suppl 2):21 DOI 10.1186/s12859-016-1443-5 RESEARCH A taxonomy of visualization tasks for the analysis of biological pathway data Paul Murray 1*,FintanMcGee 2 and
Intro Secondary structure Transmembrane proteins Function End. Last time. Domains Hidden Markov Models
 Last time Domains Hidden Markov Models Today Secondary structure Transmembrane proteins Structure prediction NAD-specific glutamate dehydrogenase Hard Easy >P24295 DHE2_CLOSY MSKYVDRVIAEVEKKYADEPEFVQTVEEVL
Last time Domains Hidden Markov Models Today Secondary structure Transmembrane proteins Structure prediction NAD-specific glutamate dehydrogenase Hard Easy >P24295 DHE2_CLOSY MSKYVDRVIAEVEKKYADEPEFVQTVEEVL
Today. Last time. Secondary structure Transmembrane proteins. Domains Hidden Markov Models. Structure prediction. Secondary structure
 Last time Today Domains Hidden Markov Models Structure prediction NAD-specific glutamate dehydrogenase Hard Easy >P24295 DHE2_CLOSY MSKYVDRVIAEVEKKYADEPEFVQTVEEVL SSLGPVVDAHPEYEEVALLERMVIPERVIE FRVPWEDDNGKVHVNTGYRVQFNGAIGPYK
Last time Today Domains Hidden Markov Models Structure prediction NAD-specific glutamate dehydrogenase Hard Easy >P24295 DHE2_CLOSY MSKYVDRVIAEVEKKYADEPEFVQTVEEVL SSLGPVVDAHPEYEEVALLERMVIPERVIE FRVPWEDDNGKVHVNTGYRVQFNGAIGPYK
CATEGORY a TERM COUNT b P VALUE GENE FAMILIES: REPRESENTATIVE GENE SYMBOLS c. Annotation Cluster 1 Enrichment Score: 1.
 Table S7 GO term and functional classification enrichment analysis using DAVID for gene families that are expanded in the Drosophila suzukii genome as compared to 14 Drosophila species analyzed in this
Table S7 GO term and functional classification enrichment analysis using DAVID for gene families that are expanded in the Drosophila suzukii genome as compared to 14 Drosophila species analyzed in this
Protein Interaction Mapping: Use of Osprey to map Survival of Motor Neuron Protein interactions
 Protein Interaction Mapping: Use of Osprey to map Survival of Motor Neuron Protein interactions Presented by: Meg Barnhart Computational Biosciences Arizona State University The Spinal Muscular Atrophy
Protein Interaction Mapping: Use of Osprey to map Survival of Motor Neuron Protein interactions Presented by: Meg Barnhart Computational Biosciences Arizona State University The Spinal Muscular Atrophy
Comparison of Human Protein-Protein Interaction Maps
 Comparison of Human Protein-Protein Interaction Maps Matthias E. Futschik 1, Gautam Chaurasia 1,2, Erich Wanker 2 and Hanspeter Herzel 1 1 Institute for Theoretical Biology, Charité, Humboldt-Universität
Comparison of Human Protein-Protein Interaction Maps Matthias E. Futschik 1, Gautam Chaurasia 1,2, Erich Wanker 2 and Hanspeter Herzel 1 1 Institute for Theoretical Biology, Charité, Humboldt-Universität
Genome-wide multilevel spatial interactome model of rice
 Sino-German Workshop on Multiscale Spatial Computational Systems Biology, Beijing, Oct 8-12, 2015 Genome-wide multilevel spatial interactome model of rice Ming CHEN ( 陈铭 ) mchen@zju.edu.cn College of Life
Sino-German Workshop on Multiscale Spatial Computational Systems Biology, Beijing, Oct 8-12, 2015 Genome-wide multilevel spatial interactome model of rice Ming CHEN ( 陈铭 ) mchen@zju.edu.cn College of Life
The Drosophila Interactions Database: Integrating The Interactome And Transcriptome
 Wayne State University Wayne State University Dissertations 1-2-2013 The Drosophila Interactions Database: Integrating The Interactome And Transcriptome Thilakam Murali Wayne State University, Follow this
Wayne State University Wayne State University Dissertations 1-2-2013 The Drosophila Interactions Database: Integrating The Interactome And Transcriptome Thilakam Murali Wayne State University, Follow this
Tandem Mass Spectrometry: Generating function, alignment and assembly
 Tandem Mass Spectrometry: Generating function, alignment and assembly With slides from Sangtae Kim and from Jones & Pevzner 2004 Determining reliability of identifications Can we use Target/Decoy to estimate
Tandem Mass Spectrometry: Generating function, alignment and assembly With slides from Sangtae Kim and from Jones & Pevzner 2004 Determining reliability of identifications Can we use Target/Decoy to estimate
Week 10: Homology Modelling (II) - HHpred
 Week 10: Homology Modelling (II) - HHpred Course: Tools for Structural Biology Fabian Glaser BKU - Technion 1 2 Identify and align related structures by sequence methods is not an easy task All comparative
Week 10: Homology Modelling (II) - HHpred Course: Tools for Structural Biology Fabian Glaser BKU - Technion 1 2 Identify and align related structures by sequence methods is not an easy task All comparative
I. Molecules & Cells. A. Unit One: The Nature of Science. B. Unit Two: The Chemistry of Life. C. Unit Three: The Biology of the Cell.
 I. Molecules & Cells A. Unit One: The Nature of Science a. How is the scientific method used to solve problems? b. What is the importance of controls? c. How does Darwin s theory of evolution illustrate
I. Molecules & Cells A. Unit One: The Nature of Science a. How is the scientific method used to solve problems? b. What is the importance of controls? c. How does Darwin s theory of evolution illustrate
TEST SUMMARY AND FRAMEWORK TEST SUMMARY
 Washington Educator Skills Tests Endorsements (WEST E) TEST SUMMARY AND FRAMEWORK TEST SUMMARY BIOLOGY Copyright 2014 by the Washington Professional Educator Standards Board 1 Washington Educator Skills
Washington Educator Skills Tests Endorsements (WEST E) TEST SUMMARY AND FRAMEWORK TEST SUMMARY BIOLOGY Copyright 2014 by the Washington Professional Educator Standards Board 1 Washington Educator Skills
Predicting rice (Oryza sativa) metabolism
 Predicting rice (Oryza sativa) metabolism Sudip Kundu Department of Biophysics, Molecular Biology & Bioinformatics, University of Calcutta, WB, India. skbmbg@caluniv.ac.in Collaborators: Mark G Poolman
Predicting rice (Oryza sativa) metabolism Sudip Kundu Department of Biophysics, Molecular Biology & Bioinformatics, University of Calcutta, WB, India. skbmbg@caluniv.ac.in Collaborators: Mark G Poolman
Written Exam 15 December Course name: Introduction to Systems Biology Course no
 Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Chemical Ontologies. Chemical Ontologies. ChemAxon UGM May 23, 2012
 Chemical Ontologies ChemAxon UGM May 23, 2012 Chemical Ontologies OntoChem GmbH Heinrich-Damerow-Str. 4 06120 Halle (Saale) Germany Tel. +49 345 4780472 Fax: +49 345 4780471 mail: info(at)ontochem.com
Chemical Ontologies ChemAxon UGM May 23, 2012 Chemical Ontologies OntoChem GmbH Heinrich-Damerow-Str. 4 06120 Halle (Saale) Germany Tel. +49 345 4780472 Fax: +49 345 4780471 mail: info(at)ontochem.com
International Journal of Scientific & Engineering Research, Volume 6, Issue 2, February ISSN
 International Journal of Scientific & Engineering Research, Volume 6, Issue 2, February-2015 273 Analogizing And Investigating Some Applications of Metabolic Pathway Analysis Methods Gourav Mukherjee 1,
International Journal of Scientific & Engineering Research, Volume 6, Issue 2, February-2015 273 Analogizing And Investigating Some Applications of Metabolic Pathway Analysis Methods Gourav Mukherjee 1,
From networks of protein interactions to networks of functional dependencies
 Luciani and Bazzoni BMC Systems Biology 2012, 6:44 RESEARCH ARTICLE From networks of protein interactions to networks of functional depencies Davide Luciani 1 and Gianfranco Bazzoni 2* Open Access Abstract
Luciani and Bazzoni BMC Systems Biology 2012, 6:44 RESEARCH ARTICLE From networks of protein interactions to networks of functional depencies Davide Luciani 1 and Gianfranco Bazzoni 2* Open Access Abstract
Context dependent visualization of protein function
 Article III Context dependent visualization of protein function In: Juho Rousu, Samuel Kaski and Esko Ukkonen (eds.). Probabilistic Modeling and Machine Learning in Structural and Systems Biology. 2006,
Article III Context dependent visualization of protein function In: Juho Rousu, Samuel Kaski and Esko Ukkonen (eds.). Probabilistic Modeling and Machine Learning in Structural and Systems Biology. 2006,
Map of AP-Aligned Bio-Rad Kits with Learning Objectives
 Map of AP-Aligned Bio-Rad Kits with Learning Objectives Cover more than one AP Biology Big Idea with these AP-aligned Bio-Rad kits. Big Idea 1 Big Idea 2 Big Idea 3 Big Idea 4 ThINQ! pglo Transformation
Map of AP-Aligned Bio-Rad Kits with Learning Objectives Cover more than one AP Biology Big Idea with these AP-aligned Bio-Rad kits. Big Idea 1 Big Idea 2 Big Idea 3 Big Idea 4 ThINQ! pglo Transformation
Miller & Levine Biology 2014
 A Correlation of Miller & Levine Biology To the Essential Standards for Biology High School Introduction This document demonstrates how meets the North Carolina Essential Standards for Biology, grades
A Correlation of Miller & Levine Biology To the Essential Standards for Biology High School Introduction This document demonstrates how meets the North Carolina Essential Standards for Biology, grades
MTopGO: a tool for module identification in PPI Networks
 MTopGO: a tool for module identification in PPI Networks Danila Vella 1,2, Simone Marini 3,4, Francesca Vitali 5,6,7, Riccardo Bellazzi 1,4 1 Clinical Scientific Institute Maugeri, Pavia, Italy, 2 Department
MTopGO: a tool for module identification in PPI Networks Danila Vella 1,2, Simone Marini 3,4, Francesca Vitali 5,6,7, Riccardo Bellazzi 1,4 1 Clinical Scientific Institute Maugeri, Pavia, Italy, 2 Department
Range of Competencies
 BIOLOGY Content Domain Range of Competencies l. Nature of Science 0001 0003 20% ll. Biochemistry and Cell Biology 0004 0005 13% lll. Genetics and Evolution 0006 0009 27% lv. Biological Unity and Diversity
BIOLOGY Content Domain Range of Competencies l. Nature of Science 0001 0003 20% ll. Biochemistry and Cell Biology 0004 0005 13% lll. Genetics and Evolution 0006 0009 27% lv. Biological Unity and Diversity
Computational Systems Biology
 Computational Systems Biology Vasant Honavar Artificial Intelligence Research Laboratory Bioinformatics and Computational Biology Graduate Program Center for Computational Intelligence, Learning, & Discovery
Computational Systems Biology Vasant Honavar Artificial Intelligence Research Laboratory Bioinformatics and Computational Biology Graduate Program Center for Computational Intelligence, Learning, & Discovery
