Supplementary Figure 1. The workflow of the KinomeXplorer web interface.
|
|
|
- Poppy Allen
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1. The workflow of the KinomeXplorer web interface. a b c d (a) NetworKIN accepts sequences, Gene/Protein names or MaxQuant/ProteomeDiscoverer results as input. In case of names, the Reflect web-service is used to find the best matches in Ensembl v59 and the corresponding sequence will be taken. (b) Sites can be selected in multiple ways: 1) manually by clicking, 2) based on a predictor for phosphorylation probability, 3) known high/low-throughput sites extracted from KinomeXplorer-DB and 4) by uploading a file with a list of sites. Predictions will only be made for the selected sites. (c) In the result page, multiple filtering possibilities are given: 1) filtering by minimum score, 2) filtering by maximum distance from the best scoring kinase for a given site, 3) filtering by tree (user must select which should be shown). For performance reasons, only the best five predictions are shown by default. (d) After having filtered the results, the user can download the result as displayed or select to retrieve the full set of predictions.
2 Supplementary Figure 2. Performance comparison of NetPhorest and NetworKIN (old and new algorithms, highlighting the benefits of adding contextual information NetPhorest NetworKIN (old) NetworKIN (new) 0.8 AUC ATM AuroraA PLK1 CDK1 CDK2 PDHK1 MAPK1 GRK2 MAPKAPK2 InsR MAPK8 Syk Lck Fyn Kinome wide Kinases that have a large enough validation set (at least 20 positive sites) and where STRING could contribute context were taken into consideration. When estimating the NetPhorest performance, we counted a prediction as true if the kinase was part of the kinase family classifier from NetPhorest. The NetworKIN results show that, by adding contextual information, we are able to disambiguate between the potential kinases of one NetPhorest group without losing predictive performance. To evaluate only the effect of the new scoring algorithm, the same NetPhorest engine and benchmark data set were used. It is important to note that while the improvement in AUC may appear modest, the corresponding improvement in practical accuracy is typically several fold.
3 Supplementary Note Dataset NetworKIN integrates two types of data; protein networks from STRING, which combines various types of data based on quality scores in a way that higher quality data has larger weight. The other one consists of kinase-substrate and phospho-binding domain interactions which have been identified in cells and have been reported in public databases (e.g. PhosphoELM, PhosphositePlus and PhosphoGRID) and position specific scoring matrices (PSSMs) which were generated by peptide array experiments in collaboration with the Yaffe 1 and Turk 2 labs. The reported kinasesubstrate relationships have been manually curated from literature mining of peer-reviewed reports of individual kinases and their phosphorylation of specific downstream proteins (part of the Phospho.ELM effort), where the specificity of the kinase-substrate relationships have been carefully tested and validated. In vivo phosphatase interactions were obtained from HuPho 3. The quality of the PSSMs was assessed based on the performance of predicting experimentally observed substrates using a rigorous computational pipeline consisting of homology reduction, data partitioning, and cross-validation as previously described in detail 4, and only high quality data entries were kept and deployed in the current release. All the peptide screening data and in vivo interactions used for training NetPhorest and NetworKIN are listed in Supplementary Table 2. Data organization for training The NetPhorest pipeline organizes datasets based on phylogenetic trees. In the tree-based organization of the training dataset, each node has positive and negatives sets. The data organization process starts from leaf nodes, which are individual kinases. For a kinase, the positive set consists of substrates that are reported to be phosphorylated by this kinase, and the negative set consists of substrates that are reported to be phosphorylated by other kinases. Through the automated selection procedure of the NetPhorest pipeline (as previously described 4 ), positive and negative sets are assembled in a way that they are best distinctive. NetworKIN uses the same data organization to avoid overtraining. Peptide binding data is used only in NetPhorest pipeline. Utilizing STRING context information in KinomeXplorer We previously developed two algorithms to predict kinases for experimentally identified phosphorylation sites, NetPhorest 4 and NetworKIN 5, which are extensively used by researchers worldwide NetPhorest analyses experimentally identified phosphorylation sites, and classifies them according to the linear motif (peptide specificity) of the kinases and phospho-binding domains. Consensus motifs have been shown to be useful in the development of kinase- or kinase-family- specific antibodies, or to detect biases arising from the enrichment procedures commonly used in phospho-proteomics, such as phospho-specific antibodies, IMAC or titanium dioxide (TiO2) 15,16. Complementary to NetPhorest, NetworKIN models kinase-substrate interactions in an integrative manner by combining sequence specificity motifs from NetPhorest with contextual network information (e.g. protein-protein interactions) from STRING 17. This database provides the contextual information we utilize to disambiguate between kinases sharing the same or a similar motif (and thus are grouped into one NetPhorest classifier), in addition to adding more confidence to the NetPhorest predictions. In the KinomeXplorer framework, phosphorylation sites are firstly classified according to the binding motifs of the kinases in the NetPhorest atlas 4, after which these classifications are refined by integrating data about the direct and indirect protein-protein interactions between the predicted kinase and its potential substrates, as this relationship is a good predictor for in vivo association. As data sources of relevant information are sparse, it is preferable to take meta-information as a refinement, thus we decided to select the STRING database as the source for our approach.
4 STRING integrates many different sources of information, including text-mining, pathway databases or co-expression into one probabilistic protein-protein association score, allowing us to accurately integrate the NetPhorest and STRING probabilities. As most kinase-substrate interactions are unknown, we extend the STRING coverage to include also indirect paths, which broadens the association information to e.g. scaffolding processes. When calculating indirect pathways, we penalize for the path lengths as well as for the overall connectivity of intermediate hubs (highly connected nodes). Finally, the score combination and likelihood transformation normalize for the biases introduced by the contextual data (namely study bias and network topology), as the weighting of context and motif information is adjusted specifically for each kinase. Supplementary Fig. 2 demonstrates the additional predictive power gained by the integration of contextual information. Kinome Coverage of KinomeXplorer platform NetPhorest was designed to incorporate new training data as soon as it becomes available 4, and we have thus retrained and benchmarked the algorithm with new experimental data in the form of position-specific scoring matrices (PSSM) and phosphorylation sites for neural network training from the latest Phospho.ELM 18 and PhosphositePlus database releases 19. This has allowed us to expand the coverage from 179 human protein kinases to 222. We also added 22 human phosphatases, which covers around 60% of human tyrosine phosphatases. Additionally, the kinome-wide sequence specificity matrices for all kinases in the budding yeast (Saccharomyces cerevisiae) have been included 20. The yeast version covers 71 of the 122 protein kinases, giving rise to a ~60% kinome-coverage. As only 111 kinases have been isolated as active enzymes in vitro, this resulted in a de-facto coverage of 64%. Because the NetworKIN algorithm builds upon NetPhorest, the improvements described above have also expanded NetworKIN s coverage to >40% of the human kinome. Additionally, due to the high sequence homology, the human versions of NetworKIN and NetPhorest can also be deployed on mouse data 10,21. Naive Bayes method We combine likelihood ratios which are derived from the NetPhorest probability and network proximity scores. The conversion of the NetPhorest probability and network proximity score was done in a kinase specific manner. To convert a score (either NetPhorest probability or network proximity) to a likelihood ratio, positive and negative sets were arranged in decreasing order by the score and a sliding window was applied. The likelihood ratio was calculated as the probability of observing the value in the positive set divided by the probability of observing the value in the negative set: Wn and Ws are windows covering a certain range of NetPhorest probability and network proximity score of String 17. The unified likelihood (Lu) of a certain NetPhorest probability and a certain network proximity score was calculated as the product of Ln and Ls by the joint probability rule: The Lu is presented as a score of a prediction. Predictions with a score higher than the theoretically neutral value of 1 are likely to be true, and the higher the score for a given kinase, the higher the likelihood it is indeed this kinase.
5 Parameter optimization To determine the optimal combination of the two parameters used (penalty for path length and node connectivity), we benchmarked against the same collection of known phosphorylation sites used to benchmark NetPhorest. To avoid overtraining we 1) use the partitioning of phosphorylation sites into training, test, and validation sets organized by the NetPhorest training pipeline, 2) use for each site the motif score from the neural network that had the site in the test set, not the training set, 3) limit the information used from the STRING network 17 to only indirect evidence paths, and 4) optimize the parameters on the test set. This ensures that parameter optimization is performed based on sites that were not used to identify the sequence specificity of the kinase in question (i.e. were not used for training the neural networks), and that the STRING network scores are not based on the same evidence that is recorded in Phospho.ELM 18. It also ensures that the validation set has neither been used for the training of NetPhorest or for parameter optimization of NetworKIN; it is hence an independent set on which the predictive performance can be accurately assessed. Web interface The KinomeXplorer web service provides an interactive and intuitive user interface (Supplementary Fig. 1). The user provides either a set of sequences in FASTA format or a list of protein or gene names. This input is then mapped to the protein collection of the STRING database 17, using sequence homology and protein names. In case of ambiguity, the website shows candidate proteins with the functional and structural details that are obtained from the Reflect web service 22, and the user is asked to manually select the correct protein. To remove ambiguity, HGNC symbols are recommended for protein names. If the input proteins only contain phosphorylation-dependent signaling domains (such as kinase, SH2, or phosphatase domains), the user is given the option to skip directly to the results page and view predicted substrates or binding partners of these. Otherwise, the input proteins are assumed to be substrates, and the user is asked to select phosphorylation sites. This can be done by uploading a file with sites, manually selecting individual sites, or using KinomeXplorer-DB, an in-house database which integrates all known phosphorylation sites from the Phospho.ELM 18, PhosphositePlus 19, and PhosphoGRID 23 databases. The web interface also integrates the NetPhos phosphorylation-site predictor 24, which can be used to add sites that are likely to be modified and/or to remove selected sites that are unlikely to be phosphorylated. The latter feature can be used e.g. to filter out likely false positive phosphorylation sites from dated and less accurately curated large-scale data sets. While we recommend the use of the downloadable source code for analyzing large datasets locally, the KinomeXplorer web interface also provides a high-throughput section, which takes the output of phospho-proteomics experiments. Users can either input tab-delimited phosphosite information or the output of peptide search engines such as MaxQuant and ProteomeDiscoverer. When working with this workflow method, it is imperative to select the correct sequence database in the pull-down menu, that was used for assigning the phosphorylation site locations and protein identifiers to guarantee correct protein/phosphosite determination. The result page offers multiple options to filter the NetPhorest or NetworKIN predictions: The primary filtering step is done by an absolute and a relative score threshold. This allows the user to show only predictions that score above a certain threshold and to hide predictions that score considerably worse than the best prediction for the same site. Additionally, one can select for which domain-classes (e.g. kinases or SH2 domains) results should be displayed. The user has the option to download either the filtered or the full set of predictions for further analysis.
6 In case that a FASTA file is uploaded with the respective phosphorylation sites, the protein identifiers can be of any nature, as the protein sequences will be mapped to STRING to guarantee correct protein matching. This is to accommodate the use of any sequence database of choice (e.g. UniProt, Ensembl). The supplied protein identifiers and phosphorylation site locations will also be used in the final output, to allow the user to easily identify their experimentally determined phosphorylation events of interest. This applies to the general prediction section of the KinomeXplorer web interface, and the downloadable package version as described below. KinomeXplorer provides a landing page with a flow-type guide to the user of which sub-module to use for the specific task at hand. For example, if the user is interested in kinase-substrate predictions, it directs the user to the NetworKIN submodule. Furthermore, we have completely redesigned the web interface for both NetPhorest and NetworKIN to make them more intuitive, tightly integrated, and applicable for analyses of large-scale experimental phosphorylation data. The workflow of the new unified interface is highlighted in Supplementary Fig. 1 and Supplementary Note. KinomeXplorer distinguishes itself from other network modeling tools such as ARACNE 25 or SteinerNet 26 by 1) using proteomics data rather than microarray expression data, and 2) by attempting to construct novel kinase-substrate interactions based on experimentally observed phosphorylation sites. To facilitate down-stream interpretation of generated predictions, they can be directly imported into network visualization and interrogation tools such as Cytoscape 27. Finally, to guide the design of follow-up kinase perturbation studies, we have integrated the KinomeSelector resource ( into KinomeXplorer, which provides an interface to construct kinase-panels for inhibition studies. The interface consists of a tree of the human kinome from which the user can select kinases to include in the panel. Additional kinases that fall within a user-customizable threshold are highlighted. These additional kinases are considered likely targets of the same inhibitors and thus need not be screened separately. The final set of selected kinases and kinases similar to them can be downloaded. We also provide a precomputed panel of 100 kinase inhibitors (covering 88% of the human kinome). Local version of NetworKIN and NetPhorest We also provide a local version of NetworKIN and NetPhorest in the Download section of the web interface ( which is our recommended method of choice for large datasets. The package contains a python script of NetworKIN and NetPhorest and the ANSI-C source code of NetPhorest and data files. NetworKIN takes a FASTA and phosphosite file as input, and predicts kinases, phosphatases, and other phospho-binding domains for the supplied phosphosites. The phosphosite file can be either a tab-delimited text file containing protein IDs, positions, and amino acids of phosphorylated residues, or output of MaxQuant and ProteomeDiscoverer. Proteins are mapped to corresponding nodes in the STRING network using sequences in the input FASTA file, not using identifiers. Therefore, in case where standard protein identifiers are ambiguous, use of local package is recommended as the supplied sequences will be mapped to STRING proteins through BLAST. The predicted substrate-kinase list is provided with additional information such as names, protein identifiers, description, and intermediate nodes in the STRING network. Similarly, NetPhorest takes the FASTA and phosphosite file supplied by the user, and predicts kinases, phosphatases and other phospho-binding domains for all the supplied
7 phosphosites. Exact details for installation and running of the package are provided in the README.txt file. References for Supplementary Information 1. Obata, T. et al. J Biol Chem 275, (2000). 2. Hutti, J. E. et al. Nat Methods 1, (2004). 3. Liberti, S. et al. FEBS J 280, (2013). 4. Miller, M. L. et al. Sci Signal 1, ra2 (2008). 5. Linding, R. et al. Cell 129, (2007). 6. Bakal, C. et al. Science 322, (2008). 7. Tan, C. S. et al. Sci Signal 2, ra39 (2009). 8. Van Hoof, D. et al. Cell Stem Cell 5, (2009). 9. Jorgensen, C. et al. Science 326, (2009). 10. Lundby, A. et al. Sci Signal 6, rs11 (2013). 11. Hekmat, O. et al. J Proteome Res 12, (2013). 12. Lundby, A. et al. Sci Signal 6, rs11 (2013). 13. Olsen, J. V. et al. Sci Signal 3, ra3 (2010). 14. Zanivan, S. et al. Cell Rep 3, (2013). 15. Zhou, H. et al. Mol Cell Proteomics 10, M (2011). 16. Rosenqvist, H., Ye, J. & Jensen, O. N. Methods Mol Biol 753, (2011). 17. Franceschini, A. et al. Nucleic Acids Res 41, D (2013). 18. Dinkel, H. et al. Nucleic Acids Res 39, D261-7 (2011). 19. Hornbeck, P. V. et al. Nucleic Acids Res 40, D (2012). 20. Mok, J. et al. Sci Signal 3, ra12 (2010). 21. Zanivan, S. et al. Cell Rep 3, (2013). 22. Pafilis, E. et al. Nat Biotechnol 27(6), (2009). 23. Sadowski, I. et al. Database (Oxford) 2013, bat026 (2013). 24. Blom, N., Gammeltoft, S. & Brunak, S. J Mol Biol 294, (1999). 25. Margolin, A. A. et al. Nat Protoc 1, (2006). 26. Tuncbag, N., McCallum, S., Huang, S. S. & Fraenkel, E. Nucleic Acids Res 40, W505-9 (2012). 27. Smoot, M. E., Ono, K., Ruscheinski, J., Wang, P. L. & Ideker, T. Bioinformatics 27, (2011).
Lecture Notes for Fall Network Modeling. Ernest Fraenkel
 Lecture Notes for 20.320 Fall 2012 Network Modeling Ernest Fraenkel In this lecture we will explore ways in which network models can help us to understand better biological data. We will explore how networks
Lecture Notes for 20.320 Fall 2012 Network Modeling Ernest Fraenkel In this lecture we will explore ways in which network models can help us to understand better biological data. We will explore how networks
Supplementary Materials for R3P-Loc Web-server
 Supplementary Materials for R3P-Loc Web-server Shibiao Wan and Man-Wai Mak email: shibiao.wan@connect.polyu.hk, enmwmak@polyu.edu.hk June 2014 Back to R3P-Loc Server Contents 1 Introduction to R3P-Loc
Supplementary Materials for R3P-Loc Web-server Shibiao Wan and Man-Wai Mak email: shibiao.wan@connect.polyu.hk, enmwmak@polyu.edu.hk June 2014 Back to R3P-Loc Server Contents 1 Introduction to R3P-Loc
Supplementary Materials for mplr-loc Web-server
 Supplementary Materials for mplr-loc Web-server Shibiao Wan and Man-Wai Mak email: shibiao.wan@connect.polyu.hk, enmwmak@polyu.edu.hk June 2014 Back to mplr-loc Server Contents 1 Introduction to mplr-loc
Supplementary Materials for mplr-loc Web-server Shibiao Wan and Man-Wai Mak email: shibiao.wan@connect.polyu.hk, enmwmak@polyu.edu.hk June 2014 Back to mplr-loc Server Contents 1 Introduction to mplr-loc
OECD QSAR Toolbox v.4.0. Tutorial on how to predict Skin sensitization potential taking into account alert performance
 OECD QSAR Toolbox v.4.0 Tutorial on how to predict Skin sensitization potential taking into account alert performance Outlook Background Objectives Specific Aims Read across and analogue approach The exercise
OECD QSAR Toolbox v.4.0 Tutorial on how to predict Skin sensitization potential taking into account alert performance Outlook Background Objectives Specific Aims Read across and analogue approach The exercise
OECD QSAR Toolbox v.4.1. Tutorial on how to predict Skin sensitization potential taking into account alert performance
 OECD QSAR Toolbox v.4.1 Tutorial on how to predict Skin sensitization potential taking into account alert performance Outlook Background Objectives Specific Aims Read across and analogue approach The exercise
OECD QSAR Toolbox v.4.1 Tutorial on how to predict Skin sensitization potential taking into account alert performance Outlook Background Objectives Specific Aims Read across and analogue approach The exercise
HOWTO, example workflow and data files. (Version )
 HOWTO, example workflow and data files. (Version 20 09 2017) 1 Introduction: SugarQb is a collection of software tools (Nodes) which enable the automated identification of intact glycopeptides from HCD
HOWTO, example workflow and data files. (Version 20 09 2017) 1 Introduction: SugarQb is a collection of software tools (Nodes) which enable the automated identification of intact glycopeptides from HCD
hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference
 CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
SUPPLEMENTARY INFORMATION
 Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Bioinformatics tools for phylogeny and visualization. Yanbin Yin
 Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
Tutorial. Getting started. Sample to Insight. March 31, 2016
 Getting started March 31, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com Getting started
Getting started March 31, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com Getting started
EasySDM: A Spatial Data Mining Platform
 EasySDM: A Spatial Data Mining Platform (User Manual) Authors: Amine Abdaoui and Mohamed Ala Al Chikha, Students at the National Computing Engineering School. Algiers. June 2013. 1. Overview EasySDM is
EasySDM: A Spatial Data Mining Platform (User Manual) Authors: Amine Abdaoui and Mohamed Ala Al Chikha, Students at the National Computing Engineering School. Algiers. June 2013. 1. Overview EasySDM is
Neural Networks for Protein Structure Prediction Brown, JMB CS 466 Saurabh Sinha
 Neural Networks for Protein Structure Prediction Brown, JMB 1999 CS 466 Saurabh Sinha Outline Goal is to predict secondary structure of a protein from its sequence Artificial Neural Network used for this
Neural Networks for Protein Structure Prediction Brown, JMB 1999 CS 466 Saurabh Sinha Outline Goal is to predict secondary structure of a protein from its sequence Artificial Neural Network used for this
Overview - MS Proteomics in One Slide. MS masses of peptides. MS/MS fragments of a peptide. Results! Match to sequence database
 Overview - MS Proteomics in One Slide Obtain protein Digest into peptides Acquire spectra in mass spectrometer MS masses of peptides MS/MS fragments of a peptide Results! Match to sequence database 2 But
Overview - MS Proteomics in One Slide Obtain protein Digest into peptides Acquire spectra in mass spectrometer MS masses of peptides MS/MS fragments of a peptide Results! Match to sequence database 2 But
TMSEG Michael Bernhofer, Jonas Reeb pp1_tmseg
 title: short title: TMSEG Michael Bernhofer, Jonas Reeb pp1_tmseg lecture: Protein Prediction 1 (for Computational Biology) Protein structure TUM summer semester 09.06.2016 1 Last time 2 3 Yet another
title: short title: TMSEG Michael Bernhofer, Jonas Reeb pp1_tmseg lecture: Protein Prediction 1 (for Computational Biology) Protein structure TUM summer semester 09.06.2016 1 Last time 2 3 Yet another
ATLAS of Biochemistry
 ATLAS of Biochemistry USER GUIDE http://lcsb-databases.epfl.ch/atlas/ CONTENT 1 2 3 GET STARTED Create your user account NAVIGATE Curated KEGG reactions ATLAS reactions Pathways Maps USE IT! Fill a gap
ATLAS of Biochemistry USER GUIDE http://lcsb-databases.epfl.ch/atlas/ CONTENT 1 2 3 GET STARTED Create your user account NAVIGATE Curated KEGG reactions ATLAS reactions Pathways Maps USE IT! Fill a gap
Networks & pathways. Hedi Peterson MTAT Bioinformatics
 Networks & pathways Hedi Peterson (peterson@quretec.com) MTAT.03.239 Bioinformatics 03.11.2010 Networks are graphs Nodes Edges Edges Directed, undirected, weighted Nodes Genes Proteins Metabolites Enzymes
Networks & pathways Hedi Peterson (peterson@quretec.com) MTAT.03.239 Bioinformatics 03.11.2010 Networks are graphs Nodes Edges Edges Directed, undirected, weighted Nodes Genes Proteins Metabolites Enzymes
Comparing whole genomes
 BioNumerics Tutorial: Comparing whole genomes 1 Aim The Chromosome Comparison window in BioNumerics has been designed for large-scale comparison of sequences of unlimited length. In this tutorial you will
BioNumerics Tutorial: Comparing whole genomes 1 Aim The Chromosome Comparison window in BioNumerics has been designed for large-scale comparison of sequences of unlimited length. In this tutorial you will
Synteny Portal Documentation
 Synteny Portal Documentation Synteny Portal is a web application portal for visualizing, browsing, searching and building synteny blocks. Synteny Portal provides four main web applications: SynCircos,
Synteny Portal Documentation Synteny Portal is a web application portal for visualizing, browsing, searching and building synteny blocks. Synteny Portal provides four main web applications: SynCircos,
THEORY. Based on sequence Length According to the length of sequence being compared it is of following two types
 Exp 11- THEORY Sequence Alignment is a process of aligning two sequences to achieve maximum levels of identity between them. This help to derive functional, structural and evolutionary relationships between
Exp 11- THEORY Sequence Alignment is a process of aligning two sequences to achieve maximum levels of identity between them. This help to derive functional, structural and evolutionary relationships between
Computational methods for predicting protein-protein interactions
 Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Ligand Scout Tutorials
 Ligand Scout Tutorials Step : Creating a pharmacophore from a protein-ligand complex. Type ke6 in the upper right area of the screen and press the button Download *+. The protein will be downloaded and
Ligand Scout Tutorials Step : Creating a pharmacophore from a protein-ligand complex. Type ke6 in the upper right area of the screen and press the button Download *+. The protein will be downloaded and
2MHR. Protein structure classification is important because it organizes the protein structure universe that is independent of sequence similarity.
 Protein structure classification is important because it organizes the protein structure universe that is independent of sequence similarity. A global picture of the protein universe will help us to understand
Protein structure classification is important because it organizes the protein structure universe that is independent of sequence similarity. A global picture of the protein universe will help us to understand
The shortest path to chemistry data and literature
 R&D SOLUTIONS Reaxys Fact Sheet The shortest path to chemistry data and literature Designed to support the full range of chemistry research, including pharmaceutical development, environmental health &
R&D SOLUTIONS Reaxys Fact Sheet The shortest path to chemistry data and literature Designed to support the full range of chemistry research, including pharmaceutical development, environmental health &
Research Article HomoKinase: A Curated Database of Human Protein Kinases
 ISRN Computational Biology Volume 2013, Article ID 417634, 5 pages http://dx.doi.org/10.1155/2013/417634 Research Article HomoKinase: A Curated Database of Human Protein Kinases Suresh Subramani, Saranya
ISRN Computational Biology Volume 2013, Article ID 417634, 5 pages http://dx.doi.org/10.1155/2013/417634 Research Article HomoKinase: A Curated Database of Human Protein Kinases Suresh Subramani, Saranya
Models of transcriptional regulation
 Models of transcriptional regulation We have already discussed four simple mechanisms of transcriptional regulation, nuclear exclusion nuclear concentration modification of bound activator redirection
Models of transcriptional regulation We have already discussed four simple mechanisms of transcriptional regulation, nuclear exclusion nuclear concentration modification of bound activator redirection
EBI web resources II: Ensembl and InterPro
 EBI web resources II: Ensembl and InterPro Yanbin Yin http://www.ebi.ac.uk/training/online/course/ 1 Homework 3 Go to http://www.ebi.ac.uk/interpro/training.htmland finish the second online training course
EBI web resources II: Ensembl and InterPro Yanbin Yin http://www.ebi.ac.uk/training/online/course/ 1 Homework 3 Go to http://www.ebi.ac.uk/interpro/training.htmland finish the second online training course
SUPPLEMENTARY INFORMATION
 Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
Evidence for dynamically organized modularity in the yeast protein-protein interaction network
 Evidence for dynamically organized modularity in the yeast protein-protein interaction network Sari Bombino Helsinki 27.3.2007 UNIVERSITY OF HELSINKI Department of Computer Science Seminar on Computational
Evidence for dynamically organized modularity in the yeast protein-protein interaction network Sari Bombino Helsinki 27.3.2007 UNIVERSITY OF HELSINKI Department of Computer Science Seminar on Computational
Last updated: Copyright
 Last updated: 2012-08-20 Copyright 2004-2012 plabel (v2.4) User s Manual by Bioinformatics Group, Institute of Computing Technology, Chinese Academy of Sciences Tel: 86-10-62601016 Email: zhangkun01@ict.ac.cn,
Last updated: 2012-08-20 Copyright 2004-2012 plabel (v2.4) User s Manual by Bioinformatics Group, Institute of Computing Technology, Chinese Academy of Sciences Tel: 86-10-62601016 Email: zhangkun01@ict.ac.cn,
BMD645. Integration of Omics
 BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
OECD QSAR Toolbox v.3.4
 OECD QSAR Toolbox v.3.4 Predicting developmental and reproductive toxicity of Diuron (CAS 330-54-1) based on DART categorization tool and DART SAR model Outlook Background Objectives The exercise Workflow
OECD QSAR Toolbox v.3.4 Predicting developmental and reproductive toxicity of Diuron (CAS 330-54-1) based on DART categorization tool and DART SAR model Outlook Background Objectives The exercise Workflow
Hands-On Nine The PAX6 Gene and Protein
 Hands-On Nine The PAX6 Gene and Protein Main Purpose of Hands-On Activity: Using bioinformatics tools to examine the sequences, homology, and disease relevance of the Pax6: a master gene of eye formation.
Hands-On Nine The PAX6 Gene and Protein Main Purpose of Hands-On Activity: Using bioinformatics tools to examine the sequences, homology, and disease relevance of the Pax6: a master gene of eye formation.
OECD QSAR Toolbox v.4.1. Tutorial illustrating new options for grouping with metabolism
 OECD QSAR Toolbox v.4.1 Tutorial illustrating new options for grouping with metabolism Outlook Background Objectives Specific Aims The exercise Workflow 2 Background Grouping with metabolism is a procedure
OECD QSAR Toolbox v.4.1 Tutorial illustrating new options for grouping with metabolism Outlook Background Objectives Specific Aims The exercise Workflow 2 Background Grouping with metabolism is a procedure
SUPPLEMENTARY MATERIALS
 SUPPLEMENTARY MATERIALS Enhanced Recognition of Transmembrane Protein Domains with Prediction-based Structural Profiles Baoqiang Cao, Aleksey Porollo, Rafal Adamczak, Mark Jarrell and Jaroslaw Meller Contact:
SUPPLEMENTARY MATERIALS Enhanced Recognition of Transmembrane Protein Domains with Prediction-based Structural Profiles Baoqiang Cao, Aleksey Porollo, Rafal Adamczak, Mark Jarrell and Jaroslaw Meller Contact:
Bioinformatics. Dept. of Computational Biology & Bioinformatics
 Bioinformatics Dept. of Computational Biology & Bioinformatics 3 Bioinformatics - play with sequences & structures Dept. of Computational Biology & Bioinformatics 4 ORGANIZATION OF LIFE ROLE OF BIOINFORMATICS
Bioinformatics Dept. of Computational Biology & Bioinformatics 3 Bioinformatics - play with sequences & structures Dept. of Computational Biology & Bioinformatics 4 ORGANIZATION OF LIFE ROLE OF BIOINFORMATICS
Predicting the Binding Patterns of Hub Proteins: A Study Using Yeast Protein Interaction Networks
 : A Study Using Yeast Protein Interaction Networks Carson M. Andorf 1, Vasant Honavar 1,2, Taner Z. Sen 2,3,4 * 1 Department of Computer Science, Iowa State University, Ames, Iowa, United States of America,
: A Study Using Yeast Protein Interaction Networks Carson M. Andorf 1, Vasant Honavar 1,2, Taner Z. Sen 2,3,4 * 1 Department of Computer Science, Iowa State University, Ames, Iowa, United States of America,
GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón
 GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón What is GO? The Gene Ontology (GO) project is a collaborative effort to address the need for consistent descriptions of gene products in
GENE ONTOLOGY (GO) Wilver Martínez Martínez Giovanny Silva Rincón What is GO? The Gene Ontology (GO) project is a collaborative effort to address the need for consistent descriptions of gene products in
Tiffany Samaroo MB&B 452a December 8, Take Home Final. Topic 1
 Tiffany Samaroo MB&B 452a December 8, 2003 Take Home Final Topic 1 Prior to 1970, protein and DNA sequence alignment was limited to visual comparison. This was a very tedious process; even proteins with
Tiffany Samaroo MB&B 452a December 8, 2003 Take Home Final Topic 1 Prior to 1970, protein and DNA sequence alignment was limited to visual comparison. This was a very tedious process; even proteins with
Using Bioinformatics to Study Evolutionary Relationships Instructions
 3 Using Bioinformatics to Study Evolutionary Relationships Instructions Student Researcher Background: Making and Using Multiple Sequence Alignments One of the primary tasks of genetic researchers is comparing
3 Using Bioinformatics to Study Evolutionary Relationships Instructions Student Researcher Background: Making and Using Multiple Sequence Alignments One of the primary tasks of genetic researchers is comparing
Mathangi Thiagarajan Rice Genome Annotation Workshop May 23rd, 2007
 -2 Transcript Alignment Assembly and Automated Gene Structure Improvements Using PASA-2 Mathangi Thiagarajan mathangi@jcvi.org Rice Genome Annotation Workshop May 23rd, 2007 About PASA PASA is an open
-2 Transcript Alignment Assembly and Automated Gene Structure Improvements Using PASA-2 Mathangi Thiagarajan mathangi@jcvi.org Rice Genome Annotation Workshop May 23rd, 2007 About PASA PASA is an open
Systematic prediction of gene function in Arabidopsis thaliana using a probabilistic functional gene network
 Systematic prediction of gene function in Arabidopsis thaliana using a probabilistic functional gene network Sohyun Hwang 1, Seung Y Rhee 2, Edward M Marcotte 3,4 & Insuk Lee 1 protocol 1 Department of
Systematic prediction of gene function in Arabidopsis thaliana using a probabilistic functional gene network Sohyun Hwang 1, Seung Y Rhee 2, Edward M Marcotte 3,4 & Insuk Lee 1 protocol 1 Department of
CSD. CSD-Enterprise. Access the CSD and ALL CCDC application software
 CSD CSD-Enterprise Access the CSD and ALL CCDC application software CSD-Enterprise brings it all: access to the Cambridge Structural Database (CSD), the world s comprehensive and up-to-date database of
CSD CSD-Enterprise Access the CSD and ALL CCDC application software CSD-Enterprise brings it all: access to the Cambridge Structural Database (CSD), the world s comprehensive and up-to-date database of
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 SUMOylation of proteins changes drastically upon heat shock, MG-132 treatment and PR-619 treatment. (a) Schematic overview of all SUMOylation proteins identified to be differentially
Supplementary Figure 1 SUMOylation of proteins changes drastically upon heat shock, MG-132 treatment and PR-619 treatment. (a) Schematic overview of all SUMOylation proteins identified to be differentially
Multiple Sequence Alignment, Gunnar Klau, December 9, 2005, 17:
 Multiple Sequence Alignment, Gunnar Klau, December 9, 2005, 17:50 5001 5 Multiple Sequence Alignment The first part of this exposition is based on the following sources, which are recommended reading:
Multiple Sequence Alignment, Gunnar Klau, December 9, 2005, 17:50 5001 5 Multiple Sequence Alignment The first part of this exposition is based on the following sources, which are recommended reading:
ncounter PlexSet Data Analysis Guidelines
 ncounter PlexSet Data Analysis Guidelines NanoString Technologies, Inc. 530 airview Ave North Seattle, Washington 98109 USA Telephone: 206.378.6266 888.358.6266 E-mail: info@nanostring.com Molecules That
ncounter PlexSet Data Analysis Guidelines NanoString Technologies, Inc. 530 airview Ave North Seattle, Washington 98109 USA Telephone: 206.378.6266 888.358.6266 E-mail: info@nanostring.com Molecules That
Cross Discipline Analysis made possible with Data Pipelining. J.R. Tozer SciTegic
 Cross Discipline Analysis made possible with Data Pipelining J.R. Tozer SciTegic System Genesis Pipelining tool created to automate data processing in cheminformatics Modular system built with generic
Cross Discipline Analysis made possible with Data Pipelining J.R. Tozer SciTegic System Genesis Pipelining tool created to automate data processing in cheminformatics Modular system built with generic
Bioinformatics. Proteins II. - Pattern, Profile, & Structure Database Searching. Robert Latek, Ph.D. Bioinformatics, Biocomputing
 Bioinformatics Proteins II. - Pattern, Profile, & Structure Database Searching Robert Latek, Ph.D. Bioinformatics, Biocomputing WIBR Bioinformatics Course, Whitehead Institute, 2002 1 Proteins I.-III.
Bioinformatics Proteins II. - Pattern, Profile, & Structure Database Searching Robert Latek, Ph.D. Bioinformatics, Biocomputing WIBR Bioinformatics Course, Whitehead Institute, 2002 1 Proteins I.-III.
The Schrödinger KNIME extensions
 The Schrödinger KNIME extensions Computational Chemistry and Cheminformatics in a workflow environment Jean-Christophe Mozziconacci Volker Eyrich Topics What are the Schrödinger extensions? Workflow application
The Schrödinger KNIME extensions Computational Chemistry and Cheminformatics in a workflow environment Jean-Christophe Mozziconacci Volker Eyrich Topics What are the Schrödinger extensions? Workflow application
CSCE555 Bioinformatics. Protein Function Annotation
 CSCE555 Bioinformatics Protein Function Annotation Why we need to do function annotation? Fig from: Network-based prediction of protein function. Molecular Systems Biology 3:88. 2007 What s function? The
CSCE555 Bioinformatics Protein Function Annotation Why we need to do function annotation? Fig from: Network-based prediction of protein function. Molecular Systems Biology 3:88. 2007 What s function? The
OECD QSAR Toolbox v.3.3. Predicting skin sensitisation potential of a chemical using skin sensitization data extracted from ECHA CHEM database
 OECD QSAR Toolbox v.3.3 Predicting skin sensitisation potential of a chemical using skin sensitization data extracted from ECHA CHEM database Outlook Background The exercise Workflow Save prediction 23.02.2015
OECD QSAR Toolbox v.3.3 Predicting skin sensitisation potential of a chemical using skin sensitization data extracted from ECHA CHEM database Outlook Background The exercise Workflow Save prediction 23.02.2015
OECD QSAR Toolbox v.4.1. Step-by-step example for building QSAR model
 OECD QSAR Toolbox v.4.1 Step-by-step example for building QSAR model Background Objectives The exercise Workflow of the exercise Outlook 2 Background This is a step-by-step presentation designed to take
OECD QSAR Toolbox v.4.1 Step-by-step example for building QSAR model Background Objectives The exercise Workflow of the exercise Outlook 2 Background This is a step-by-step presentation designed to take
08/21/2017 BLAST. Multiple Sequence Alignments: Clustal Omega
 BLAST Multiple Sequence Alignments: Clustal Omega What does basic BLAST do (e.g. what is input sequence and how does BLAST look for matches?) Susan Parrish McDaniel College Multiple Sequence Alignments
BLAST Multiple Sequence Alignments: Clustal Omega What does basic BLAST do (e.g. what is input sequence and how does BLAST look for matches?) Susan Parrish McDaniel College Multiple Sequence Alignments
OECD QSAR Toolbox v.3.3. Step-by-step example of how to build a userdefined
 OECD QSAR Toolbox v.3.3 Step-by-step example of how to build a userdefined QSAR Background Objectives The exercise Workflow of the exercise Outlook 2 Background This is a step-by-step presentation designed
OECD QSAR Toolbox v.3.3 Step-by-step example of how to build a userdefined QSAR Background Objectives The exercise Workflow of the exercise Outlook 2 Background This is a step-by-step presentation designed
Large-Scale Genomic Surveys
 Bioinformatics Subtopics Fold Recognition Secondary Structure Prediction Docking & Drug Design Protein Geometry Protein Flexibility Homology Modeling Sequence Alignment Structure Classification Gene Prediction
Bioinformatics Subtopics Fold Recognition Secondary Structure Prediction Docking & Drug Design Protein Geometry Protein Flexibility Homology Modeling Sequence Alignment Structure Classification Gene Prediction
OECD QSAR Toolbox v.4.1. Step-by-step example for predicting skin sensitization accounting for abiotic activation of chemicals
 OECD QSAR Toolbox v.4.1 Step-by-step example for predicting skin sensitization accounting for abiotic activation of chemicals Background Outlook Objectives The exercise Workflow 2 Background This is a
OECD QSAR Toolbox v.4.1 Step-by-step example for predicting skin sensitization accounting for abiotic activation of chemicals Background Outlook Objectives The exercise Workflow 2 Background This is a
Machine Learning in Action
 Machine Learning in Action Tatyana Goldberg (goldberg@rostlab.org) August 16, 2016 @ Machine Learning in Biology Beijing Genomics Institute in Shenzhen, China June 2014 GenBank 1 173,353,076 DNA sequences
Machine Learning in Action Tatyana Goldberg (goldberg@rostlab.org) August 16, 2016 @ Machine Learning in Biology Beijing Genomics Institute in Shenzhen, China June 2014 GenBank 1 173,353,076 DNA sequences
Bioinformatics 2. Yeast two hybrid. Proteomics. Proteomics
 GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
Problem Set 5 Question 1
 2.32 Problem Set 5 Question As discussed in class, drug discovery often involves screening large libraries of small molecules to identify those that have favorable interactions with a certain druggable
2.32 Problem Set 5 Question As discussed in class, drug discovery often involves screening large libraries of small molecules to identify those that have favorable interactions with a certain druggable
Z -LYTE Assay Setup Guide on the BMG LABTECH CLARIOstar Reader
 08 Jan 15 Page 1 of 18 Z -LYTE Assay Setup Guide on the BMG LABTECH CLARIOstar Reader The BMG LABTECH CLARIOstar Microplate Readers were tested for compatibility with the Life Technologies Z -LYTE Assay
08 Jan 15 Page 1 of 18 Z -LYTE Assay Setup Guide on the BMG LABTECH CLARIOstar Reader The BMG LABTECH CLARIOstar Microplate Readers were tested for compatibility with the Life Technologies Z -LYTE Assay
Protein Structure Prediction II Lecturer: Serafim Batzoglou Scribe: Samy Hamdouche
 Protein Structure Prediction II Lecturer: Serafim Batzoglou Scribe: Samy Hamdouche The molecular structure of a protein can be broken down hierarchically. The primary structure of a protein is simply its
Protein Structure Prediction II Lecturer: Serafim Batzoglou Scribe: Samy Hamdouche The molecular structure of a protein can be broken down hierarchically. The primary structure of a protein is simply its
Evaluating Physical, Chemical, and Biological Impacts from the Savannah Harbor Expansion Project Cooperative Agreement Number W912HZ
 Evaluating Physical, Chemical, and Biological Impacts from the Savannah Harbor Expansion Project Cooperative Agreement Number W912HZ-13-2-0013 Annual Report FY 2018 Submitted by Sergio Bernardes and Marguerite
Evaluating Physical, Chemical, and Biological Impacts from the Savannah Harbor Expansion Project Cooperative Agreement Number W912HZ-13-2-0013 Annual Report FY 2018 Submitted by Sergio Bernardes and Marguerite
Lecture 2. The Blast2GO annotation framework
 Lecture 2 The Blast2GO annotation framework Annotation steps Modulation of annotation intensity Export/Import Functions Sequence Selection Additional Tools Functional assignment Annotation Transference
Lecture 2 The Blast2GO annotation framework Annotation steps Modulation of annotation intensity Export/Import Functions Sequence Selection Additional Tools Functional assignment Annotation Transference
OECD QSAR Toolbox v.3.0
 OECD QSAR Toolbox v.3.0 Step-by-step example of how to categorize an inventory by mechanistic behaviour of the chemicals which it consists Background Objectives Specific Aims Trend analysis The exercise
OECD QSAR Toolbox v.3.0 Step-by-step example of how to categorize an inventory by mechanistic behaviour of the chemicals which it consists Background Objectives Specific Aims Trend analysis The exercise
FuncNet a distributed platform for high-throughput protein function analysis. Andrew Clegg University College London. funcnet.eu
 FuncNet a distributed platform for high-throughput protein function analysis Andrew Clegg University College London Outline of talk Introduction and background Working with FuncNet APIs and extensions
FuncNet a distributed platform for high-throughput protein function analysis Andrew Clegg University College London Outline of talk Introduction and background Working with FuncNet APIs and extensions
OECD QSAR Toolbox v.3.3. Predicting acute aquatic toxicity to fish of Dodecanenitrile (CAS ) taking into account tautomerism
 OECD QSAR Toolbox v.3.3 Predicting acute aquatic toxicity to fish of Dodecanenitrile (CAS 2437-25-4) taking into account tautomerism Outlook Background Objectives The exercise Workflow Save prediction
OECD QSAR Toolbox v.3.3 Predicting acute aquatic toxicity to fish of Dodecanenitrile (CAS 2437-25-4) taking into account tautomerism Outlook Background Objectives The exercise Workflow Save prediction
TUTORIAL EXERCISES WITH ANSWERS
 TUTORIAL EXERCISES WITH ANSWERS Tutorial 1 Settings 1. What is the exact monoisotopic mass difference for peptides carrying a 13 C (and NO additional 15 N) labelled C-terminal lysine residue? a. 6.020129
TUTORIAL EXERCISES WITH ANSWERS Tutorial 1 Settings 1. What is the exact monoisotopic mass difference for peptides carrying a 13 C (and NO additional 15 N) labelled C-terminal lysine residue? a. 6.020129
Genome Annotation. Bioinformatics and Computational Biology. Genome sequencing Assembly. Gene prediction. Protein targeting.
 Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
OECD QSAR Toolbox v.3.3
 OECD QSAR Toolbox v.3.3 Step-by-step example on how to predict the skin sensitisation potential of a chemical by read-across based on an analogue approach Outlook Background Objectives Specific Aims Read
OECD QSAR Toolbox v.3.3 Step-by-step example on how to predict the skin sensitisation potential of a chemical by read-across based on an analogue approach Outlook Background Objectives Specific Aims Read
OECD QSAR Toolbox v.3.3. Step-by-step example of how to categorize an inventory by mechanistic behaviour of the chemicals which it consists
 OECD QSAR Toolbox v.3.3 Step-by-step example of how to categorize an inventory by mechanistic behaviour of the chemicals which it consists Background Objectives Specific Aims Trend analysis The exercise
OECD QSAR Toolbox v.3.3 Step-by-step example of how to categorize an inventory by mechanistic behaviour of the chemicals which it consists Background Objectives Specific Aims Trend analysis The exercise
Multiple sequence alignment
 Multiple sequence alignment Multiple sequence alignment: today s goals to define what a multiple sequence alignment is and how it is generated; to describe profile HMMs to introduce databases of multiple
Multiple sequence alignment Multiple sequence alignment: today s goals to define what a multiple sequence alignment is and how it is generated; to describe profile HMMs to introduce databases of multiple
Reaxys Medicinal Chemistry Fact Sheet
 R&D SOLUTIONS FOR PHARMA & LIFE SCIENCES Reaxys Medicinal Chemistry Fact Sheet Essential data for lead identification and optimization Reaxys Medicinal Chemistry empowers early discovery in drug development
R&D SOLUTIONS FOR PHARMA & LIFE SCIENCES Reaxys Medicinal Chemistry Fact Sheet Essential data for lead identification and optimization Reaxys Medicinal Chemistry empowers early discovery in drug development
Dock Ligands from a 2D Molecule Sketch
 Dock Ligands from a 2D Molecule Sketch March 31, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com
Dock Ligands from a 2D Molecule Sketch March 31, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com
Investigation 3: Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST
 Investigation 3: Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST Introduction Bioinformatics is a powerful tool which can be used to determine evolutionary relationships and
Investigation 3: Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST Introduction Bioinformatics is a powerful tool which can be used to determine evolutionary relationships and
HOW TO USE MIKANA. 1. Decompress the zip file MATLAB.zip. This will create the directory MIKANA.
 HOW TO USE MIKANA MIKANA (Method to Infer Kinetics And Network Architecture) is a novel computational method to infer reaction mechanisms and estimate the kinetic parameters of biochemical pathways from
HOW TO USE MIKANA MIKANA (Method to Infer Kinetics And Network Architecture) is a novel computational method to infer reaction mechanisms and estimate the kinetic parameters of biochemical pathways from
A New Similarity Measure among Protein Sequences
 A New Similarity Measure among Protein Sequences Kuen-Pin Wu, Hsin-Nan Lin, Ting-Yi Sung and Wen-Lian Hsu * Institute of Information Science Academia Sinica, Taipei 115, Taiwan Abstract Protein sequence
A New Similarity Measure among Protein Sequences Kuen-Pin Wu, Hsin-Nan Lin, Ting-Yi Sung and Wen-Lian Hsu * Institute of Information Science Academia Sinica, Taipei 115, Taiwan Abstract Protein sequence
Introduction to Bioinformatics Online Course: IBT
 Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec1 Building a Multiple Sequence Alignment Learning Outcomes 1- Understanding Why multiple
Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec1 Building a Multiple Sequence Alignment Learning Outcomes 1- Understanding Why multiple
Proteomics. Yeast two hybrid. Proteomics - PAGE techniques. Data obtained. What is it?
 Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
Session 5: Phylogenomics
 Session 5: Phylogenomics B.- Phylogeny based orthology assignment REMINDER: Gene tree reconstruction is divided in three steps: homology search, multiple sequence alignment and model selection plus tree
Session 5: Phylogenomics B.- Phylogeny based orthology assignment REMINDER: Gene tree reconstruction is divided in three steps: homology search, multiple sequence alignment and model selection plus tree
OECD QSAR Toolbox v.3.4
 OECD QSAR Toolbox v.3.4 Step-by-step example on how to predict the skin sensitisation potential approach of a chemical by read-across based on an analogue approach Outlook Background Objectives Specific
OECD QSAR Toolbox v.3.4 Step-by-step example on how to predict the skin sensitisation potential approach of a chemical by read-across based on an analogue approach Outlook Background Objectives Specific
An optimized energy potential can predict SH2 domainpeptide
 An optimized energy potential can predict SH2 domainpeptide interactions Running Title Predicting SH2 interactions Authors Zeba Wunderlich 1, Leonid A. Mirny 2 Affiliations 1. Biophysics Program, Harvard
An optimized energy potential can predict SH2 domainpeptide interactions Running Title Predicting SH2 interactions Authors Zeba Wunderlich 1, Leonid A. Mirny 2 Affiliations 1. Biophysics Program, Harvard
Integration of functional genomics data
 Integration of functional genomics data Laboratoire Bordelais de Recherche en Informatique (UMR) Centre de Bioinformatique de Bordeaux (Plateforme) Rennes Oct. 2006 1 Observations and motivations Genomics
Integration of functional genomics data Laboratoire Bordelais de Recherche en Informatique (UMR) Centre de Bioinformatique de Bordeaux (Plateforme) Rennes Oct. 2006 1 Observations and motivations Genomics
1-D Predictions. Prediction of local features: Secondary structure & surface exposure
 1-D Predictions Prediction of local features: Secondary structure & surface exposure 1 Learning Objectives After today s session you should be able to: Explain the meaning and usage of the following local
1-D Predictions Prediction of local features: Secondary structure & surface exposure 1 Learning Objectives After today s session you should be able to: Explain the meaning and usage of the following local
Some Problems from Enzyme Families
 Some Problems from Enzyme Families Greg Butler Department of Computer Science Concordia University, Montreal www.cs.concordia.ca/~faculty/gregb gregb@cs.concordia.ca Abstract I will discuss some problems
Some Problems from Enzyme Families Greg Butler Department of Computer Science Concordia University, Montreal www.cs.concordia.ca/~faculty/gregb gregb@cs.concordia.ca Abstract I will discuss some problems
Chapter 5. Proteomics and the analysis of protein sequence Ⅱ
 Proteomics Chapter 5. Proteomics and the analysis of protein sequence Ⅱ 1 Pairwise similarity searching (1) Figure 5.5: manual alignment One of the amino acids in the top sequence has no equivalent and
Proteomics Chapter 5. Proteomics and the analysis of protein sequence Ⅱ 1 Pairwise similarity searching (1) Figure 5.5: manual alignment One of the amino acids in the top sequence has no equivalent and
Systems biology and biological networks
 Systems Biology Workshop Systems biology and biological networks Center for Biological Sequence Analysis Networks in electronics Radio kindly provided by Lazebnik, Cancer Cell, 2002 Systems Biology Workshop,
Systems Biology Workshop Systems biology and biological networks Center for Biological Sequence Analysis Networks in electronics Radio kindly provided by Lazebnik, Cancer Cell, 2002 Systems Biology Workshop,
PTM Bioinformatics Yu Xue
 PTM Bioinformatics Yu Xue Huazhong University of Science and Technology Wuhan, China Huazhong University of Science and Technology http://www.uniprot.org/docs/ptmlist Post-translational modification POST:
PTM Bioinformatics Yu Xue Huazhong University of Science and Technology Wuhan, China Huazhong University of Science and Technology http://www.uniprot.org/docs/ptmlist Post-translational modification POST:
The Solution to Assignment 6
 The Solution to Assignment 6 Problem 1: Use the 2-fold cross-validation to evaluate the Decision Tree Model for trees up to 2 levels deep (that is, the maximum path length from the root to the leaves is
The Solution to Assignment 6 Problem 1: Use the 2-fold cross-validation to evaluate the Decision Tree Model for trees up to 2 levels deep (that is, the maximum path length from the root to the leaves is
Ensembl focuses on metazoan (animal) genomes. The genomes currently available at the Ensembl site are:
 Comparative genomics and proteomics Species available Ensembl focuses on metazoan (animal) genomes. The genomes currently available at the Ensembl site are: Vertebrates: human, chimpanzee, mouse, rat,
Comparative genomics and proteomics Species available Ensembl focuses on metazoan (animal) genomes. The genomes currently available at the Ensembl site are: Vertebrates: human, chimpanzee, mouse, rat,
Towards Detecting Protein Complexes from Protein Interaction Data
 Towards Detecting Protein Complexes from Protein Interaction Data Pengjun Pei 1 and Aidong Zhang 1 Department of Computer Science and Engineering State University of New York at Buffalo Buffalo NY 14260,
Towards Detecting Protein Complexes from Protein Interaction Data Pengjun Pei 1 and Aidong Zhang 1 Department of Computer Science and Engineering State University of New York at Buffalo Buffalo NY 14260,
CHEM 463: Advanced Inorganic Chemistry Modeling Metalloproteins for Structural Analysis
 CHEM 463: Advanced Inorganic Chemistry Modeling Metalloproteins for Structural Analysis Purpose: The purpose of this laboratory is to introduce some of the basic visualization and modeling tools for viewing
CHEM 463: Advanced Inorganic Chemistry Modeling Metalloproteins for Structural Analysis Purpose: The purpose of this laboratory is to introduce some of the basic visualization and modeling tools for viewing
Jeremy Chang Identifying protein protein interactions with statistical coupling analysis
 Jeremy Chang Identifying protein protein interactions with statistical coupling analysis Abstract: We used an algorithm known as statistical coupling analysis (SCA) 1 to create a set of features for building
Jeremy Chang Identifying protein protein interactions with statistical coupling analysis Abstract: We used an algorithm known as statistical coupling analysis (SCA) 1 to create a set of features for building
CMPS 6630: Introduction to Computational Biology and Bioinformatics. Structure Comparison
 CMPS 6630: Introduction to Computational Biology and Bioinformatics Structure Comparison Protein Structure Comparison Motivation Understand sequence and structure variability Understand Domain architecture
CMPS 6630: Introduction to Computational Biology and Bioinformatics Structure Comparison Protein Structure Comparison Motivation Understand sequence and structure variability Understand Domain architecture
MassHunter TOF/QTOF Users Meeting
 MassHunter TOF/QTOF Users Meeting 1 Qualitative Analysis Workflows Workflows in Qualitative Analysis allow the user to only see and work with the areas and dialog boxes they need for their specific tasks
MassHunter TOF/QTOF Users Meeting 1 Qualitative Analysis Workflows Workflows in Qualitative Analysis allow the user to only see and work with the areas and dialog boxes they need for their specific tasks
OPSIAL Manual. v Xiaofeng Tan. All Rights Reserved
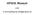 OPSIAL Manual v1.0 2016 Xiaofeng Tan. All Rights Reserved 1. Introduction... 3 1.1 Spectral Calculator & Fitter (SCF)... 3 1.2 Automated Analyzer (AA)... 3 2. Working Principles and Workflows of OPSIAL...
OPSIAL Manual v1.0 2016 Xiaofeng Tan. All Rights Reserved 1. Introduction... 3 1.1 Spectral Calculator & Fitter (SCF)... 3 1.2 Automated Analyzer (AA)... 3 2. Working Principles and Workflows of OPSIAL...
ALGORITHMS FOR BIOLOGICAL NETWORK ALIGNMENT
 ALGORITHMS FOR BIOLOGICAL NETWORK ALIGNMENT A DISSERTATION SUBMITTED TO THE DEPARTMENT OF COMPUTER SCIENCE AND THE COMMITTEE ON GRADUATE STUDIES OF STANFORD UNIVERSITY IN PARTIAL FULFILLMENT OF THE REQUIREMENTS
ALGORITHMS FOR BIOLOGICAL NETWORK ALIGNMENT A DISSERTATION SUBMITTED TO THE DEPARTMENT OF COMPUTER SCIENCE AND THE COMMITTEE ON GRADUATE STUDIES OF STANFORD UNIVERSITY IN PARTIAL FULFILLMENT OF THE REQUIREMENTS
Analysis of N-terminal Acetylation data with Kernel-Based Clustering
 Analysis of N-terminal Acetylation data with Kernel-Based Clustering Ying Liu Department of Computational Biology, School of Medicine University of Pittsburgh yil43@pitt.edu 1 Introduction N-terminal acetylation
Analysis of N-terminal Acetylation data with Kernel-Based Clustering Ying Liu Department of Computational Biology, School of Medicine University of Pittsburgh yil43@pitt.edu 1 Introduction N-terminal acetylation
Keywords: intrinsic disorder, IDP, protein, protein structure, PTM
 PostDoc Journal Vol.2, No.1, January 2014 Journal of Postdoctoral Research www.postdocjournal.com Protein Structure and Function: Methods for Prediction and Analysis Ravi Ramesh Pathak Morsani College
PostDoc Journal Vol.2, No.1, January 2014 Journal of Postdoctoral Research www.postdocjournal.com Protein Structure and Function: Methods for Prediction and Analysis Ravi Ramesh Pathak Morsani College
Agilent MassHunter Profinder: Solving the Challenge of Isotopologue Extraction for Qualitative Flux Analysis
 Agilent MassHunter Profinder: Solving the Challenge of Isotopologue Extraction for Qualitative Flux Analysis Technical Overview Introduction Metabolomics studies measure the relative abundance of metabolites
Agilent MassHunter Profinder: Solving the Challenge of Isotopologue Extraction for Qualitative Flux Analysis Technical Overview Introduction Metabolomics studies measure the relative abundance of metabolites
Tutorial: Structural Analysis of a Protein-Protein Complex
 Molecular Modeling Section (MMS) Department of Pharmaceutical and Pharmacological Sciences University of Padova Via Marzolo 5-35131 Padova (IT) @contact: stefano.moro@unipd.it Tutorial: Structural Analysis
Molecular Modeling Section (MMS) Department of Pharmaceutical and Pharmacological Sciences University of Padova Via Marzolo 5-35131 Padova (IT) @contact: stefano.moro@unipd.it Tutorial: Structural Analysis
Protein Quantitation II: Multiple Reaction Monitoring. Kelly Ruggles New York University
 Protein Quantitation II: Multiple Reaction Monitoring Kelly Ruggles kelly@fenyolab.org New York University Traditional Affinity-based proteomics Use antibodies to quantify proteins Western Blot Immunohistochemistry
Protein Quantitation II: Multiple Reaction Monitoring Kelly Ruggles kelly@fenyolab.org New York University Traditional Affinity-based proteomics Use antibodies to quantify proteins Western Blot Immunohistochemistry
