27 : Distributed Monte Carlo Markov Chain. 1 Recap of MCMC and Naive Parallel Gibbs Sampling
|
|
|
- Sharon Gallagher
- 5 years ago
- Views:
Transcription
1 10-708: Probabilistic Graphical Models , Spring : Distributed Monte Carlo Markov Chain Lecturer: Eric P. Xing Scribes: Pengtao Xie, Khoa Luu In this scribe, we are going to review the Parallel Monte Carlo Markov Chain (MCMC) method. First, we will recap of MCMC methods, particularly the Metropolis-Hasting and Gibbs Sampling algorithms. Then we will show the drawbacks of these classical MCMC methods as well as the Naive Parallel Gibbs Sampling approach. Finally, we will come up with the Sequential Monte Carlo and Parallel Inference for Bayesian nonparametric models, specifically, the Dirichlet Process Mixture model. Numerous kinds of inference techniques are also discussed in this paper. 1 Recap of MCMC and Naive Parallel Gibbs Sampling Monte Carlo methods consist of a wide variety of algorithms that enable us to sample from complicated probability distribution functions without explicitly generating the probabilistic density function (pdf) [1]. These methods are particularly useful when it is computationally intractable to generate the pdf, P (x), which is typical of pdfs defined in high-dimensional spaces. Markov Chain Monte Carlo methods, on the other hand, use adaptive proposals Q(x x) to sample from a given true distribution P (x) and produce new appropriate samples x by using a Markov chain. The distribution P (x) is usually neither simple nor standard one. Therefore, it is unable to be sampled directly in practical applications. Markov Chain Monte Carlo can be applied in these situations where finding the exact solution is intractable. Given the initial probability P (x (0) ) and the transition probabilities T (x x), where x T (x x) = 1 (it is usually called the homogeneous chain). The Markov chain can be computed by simply marginalizing the probability of x as follows: P (x ) = x P (x)t (x x) where x is the current sample. The detailed balance property tells that the probability of picking x from the target density P (x) and transition from x to x is just as likely as picking x and transition T (x x). Thus it implies the invariance of the distribution P (x) and the probability distribution of the chain converges to P (x). MCMC algorithms use Q(x x) for proposal distribution, instead of Q(x). This process therefore generates a Markov chain from samples x(1), x(2)... One of the most popular MCMC method is Metropolis-Hastings, which allows us to specify any proposal Q(x x), although choosing a good Q(x x) requires care. Gibbs sampling is a famous example of Metropolis-Hastings method, which sets the proposal Q(x x) to the conditional distribution P (x x) and the accept rate is always 1. 1
2 2 27 : Distributed Monte Carlo Markov Chain 1.1 Metropolis-Hastings Sampling Algorithm Metropolis-Hastings (MH) sampling algorithm is distinct in the sense that it is a Markov Chain variant of Monte Carlo sampling. It can be seen as a generalization of Metropolis algorithm. Specifically, the proposal is no longer a fixed probability distribution, the functional form changes based on the current state of the sampler. In other words, our proposal is only conditioned on the most recently drawn sample. Therefore, the proposal function can be denoted as the following conditional probability distribution Q(x x (t) ). In order to perform sampling in this manner, we first assume that we can evaluate some unnormalized version of our pdf, P (x), at all possible values of x. Secondly, we define an auxiliary proposal distribution which we denote as Q (x). We define Q (x) such that it is easy to evaluate and to sample from. With these two ingredients, we can sample from P (x) using a variety of Monte Carlo sampling methods assuming certain mathematica properties hold for the proposal distribution. Far apart from Metropolis algorithm where the proposal distribution must be symmetric, e.g. Q(x x (t) ) = Q(x (t) x ), the proposal distribution Q(x x (t) ) in Metropolis-Hastings algorithm doesn t require that property. Given a distribution P (x), it can be sampled using the MH algorithm by performing Alg. 1. Algorithm 1 Metropolis-Hastings Sampling Algorithm Input: Conditional proposal Q (x x), Target P (x) Output: The sampled set of N correlated samples drawn from P (x) Initialize state x (0) Initialize sampled set as Null for t = 1..N do Sample proposal Q ( x x (t)) to obtain candidate x Accept this state with a probability α = P (x )Q(x (t) x ) P (x (t) )Q(x x (t) ) if state is accepted then Set x (t+1) = x else Set x (t+1) = x (t) end if Add x (t+1) to the sampled set. end for return the sampled set Selecting a right proposal distribution Q(x x) for the given distribution P (x) strongly impacts the performance of the algorithm in both convergence rate and the accuracy. When proposal distribution Q(x x) has the form of isotropic Gaussian, smaller variance results in many accepted samples. However, the algorithm will slowly search through the continuous space and possibly ignore some of the models of P (x). On the other hand, for larger variance, the algorithm rejects most of the samples generated from the proposal distribution since p(x) is low. 1.2 Gibbs Sampling Algorithm Gibbs Sampling is also an instance of a Markov Chain Monte Carlo technique. It is a special case of MH algorithm where each random variable x i is sampled, given the rest of variables or the Markov Blanket of x i. The conditional probability distribution P (x x i ) is selected as the proposal distribution Q(x x) to sample = x i for the expected distribution P (x 1:K) = P (x 1, x 2,...; x K ), where x i refers to the ith random variable and x i denotes the set of all random variables except x i. x (t+1) i
3 27 : Distributed Monte Carlo Markov Chain 3 Algorithm 2 Gibbs Sampling Algorithm Input: Conditional proposal Output: The sampled set of N correlated samples drawn from P (x) Initialize state x (0) =< x (0) Initialize sampled set as Null for t = 1..N do for i = 1..K do Sample x (t+1) i 1, x(0) 2,.., x(0) K > P (Z i x (t+1) 1,..., x (t+1) Add x (t+1) to the sampled set. end for end for return the sampled set i 1, x(t) i+1, x(t) K ) For conditional distributions that belong to the family of standard distribution, we can sample directly from the conditionals, otherwise, we can draw samples using MH embedded within the Gibbs sampler. A simple calculation can show that the acceptance probability for each sample in Gibbs sampling is one. A(x x) = P (x )Q(x x ) P (x)q(x x) = P (x x i )P (x i )P (x i x i ) P (x i x i )P (x i )P (x i x i) = 1 where x i = x i. In this case, all the samples generated from a Gibbs sampler are always accepted. 1.3 MCMC in Large Scale Datasets Datasets and models nowadays are usually very large. We may have millions to billions of data points and millions to billions of random variables. Computational time is measured in CPU-years, which means that it is impractical to run on a single thread on a single machine. In addition, we may need memory of GB or TB to store the computing information. An example is the Yahoo web graph which has 1.4 billion nodes and 6.6 billion edges. Imagining that we run a Markov Random Field on the network, it is clearly impossible to run it on a single machine. Without parallelism, large datasets and models cannot be used efficiently. It is notice that the proper use of MCMC actually requires parallelism. To determine convergence, multiple MCMC chains have to be employed. This is the reason why we need to explore the paralleling version in Gibbs sampling algorithm. However, there is a difference between Normal Gibbs sampling approach and the Naive Parallel Gibbs Sampling one when processing multiple MCMC chains. In the Normal Gibbs sampling method, a node is updated based on the current state of all other nodes. Meanwhile, in Naive Parallel Gibbs Sampling approach, a node is updated by using a previous state of all other nodes. Fore example, recall the Alarm Network problem (as shown in Fig. 1). Let s say we initialize all variables at time t to false. The idea is to in parallel Gibbs sample all variables at step t conditioned on t 1. We start to sample B from P (B A, E) and the Markov blanket of B is A and E. Using Bayesian rule to figure out the conditional distribution P (B A, E) P (A B, E)P (B), a sample can be drawn from the posterior. Let s suppose the drawn B is false. Next we sample E from P (E A, B). The values of A, B are still those at t 1. In normal Gibbs sampling, we compute P (E A, B) based on B t=1 and A t=0. In a naive parallel Gibbs sampler, we compute P (E A, B) based on B t=0 and A t=0. This is the key difference. We always condition on the previous iteration, rather than the most recently updated values. We precede to sample A and pick false. Similarly, we precede on and sample J. Finally, we sample M and pick false. That s one iteration of the Naive Gibbs sampling. We just finished sampling variables for t = 1. We run multiple Gibbs sampler on multiple machines with different seeds. They will produce different
4 4 27 : Distributed Monte Carlo Markov Chain Figure 1: Example of Alarm Network Problem Markov chains. Once they are all converged to the same statistics, then we stop and say the chain has converged. If they are not converged, it is no the time to stop. However, taking multiple chains does not solve every issue. If the burn in period is long when you are doing MCMC, chains will take long time to converge. What we really want is to take each sample faster. Using the example above, we can see that the algorithm is only conditioned on variable state at t = 0, which already is known in advance. We therefore can sample every node, e.g. B, E, A, J, and M, on separate processors, without having to send information between processors. In practice, this approach works very well for some graphical models, e.g. collapsed Gibbs Sampling for LDA, just assign different z i s to different processors or machines. However, this Naive Parallel Gibbs Sampling approach may not converge to the stationary distribution since the Naive parallel GS performs poorly on near-discrete distributions. 2 Sequential Importance Sampling When we do Sequential Importance Sampling (SIS), we draw sample x from the proposal, compute the unnormalized weights, resample given the samples and normalized weights to get a set of samples with uniform rate. Then move to the second variable, third variable, so on so forth. To summarize, in parallel Gibbs sampling, there is a naive strategy where we sample all variables at the same time and a correct strategy where we first perform graph coloring and sample same-colored node in parallel before moving to the next color. In sequential Monte Carlo, we use incremental proposal distribution in order to get an algorithm that is truly online, meaning that every new data point comes in, we only do a constant amount of work. That works in MCMC, because we have a proposal that is incremental, we have a way to compute the weights incrementally and a way to precede to the next variable incrementally. It provides a framework for designing online, parallel MCMC algorithms. 3 Parallel Inference for Bayesian Nonparametric Models In this part, we are going to look at the parallel inference for Bayesian nonparametric models, specifically, Dirichlet process mixture model.
5 27 : Distributed Monte Carlo Markov Chain Recap of Dirichlet Process First, let s take a recap of Dirichlet process. Before going to the infinite case, let s first see the finite case. Suppose there is a restaurant which has K tables in it. On each table, there is a dish. The dish here is represented by color. People enter the restaurant and pick a table. In this metaphor, table represents cluster and people are items to be clustered. Dish on the table can be thought of parameters, such as the center of each cluster. The things we want to estimate are: 1, what is the dish on each table; 2, the assignments of people to each table. We can use hard k-means to do the estimation. If we change the setting a little bit, that whether people will sit on the table is propositional to the appreciation of the dish on that table, we can use soft k-means to solve this problem. In the generative perspective, we assume each dish has a distribution, from which we can draw items. Here is the whole generative process, from some distribution H, we draw K clusters. For each item, we first draw cluster index and then draw item from the cluster corresponding to the cluster index. Now, in the clustering process, we want to incorporate the richer get richer property, which means a table with more people is more likely to be picked. In this case, the probability that a person chooses a table is not only proportional to his appreciation of the dish on the table, but also how popular the table is. This is what we called finite mixture model. If we look at the generative picture of the finite mixture model, this part is exactly the same as before. The only addition is that now the cluster index are drawn from a multinomial. Figure 2: Restaurant Perspective of Infinite Mixture Model Next, we consider infinite number of tables. People sit on an existing table according to their appreciation of the dish and the popularity of the table. They also pick a new table with certain amount of probability and place a new dish onto that table. This is called Dirichlet process mixture model. Here is an example 4 (Figure 2) where three tables are occupied. The probabilities to pick an occupied table is α 9+α 9+α, 3 9+α, 2 9+α. The important part is people pick a new table with probability of. Note that if we want to pick a particular table, which is unoccupied, that probability is zero, because there is infinitely many of them. This process is called Dirichlet process mixture model is because, if we look at the posterior distribution, it will give we exactly the same thing as this. The next question we are going to answer is what the sample of Dirichlet process looks like. The rectangle denotes the probability of picking a particular table. This distribution is the dish on the table. If we draw a sample from Dirichlet process, then this is what it looks like. The next question is how can we create such a distribution. One way is to create by construction. Note that there will be infinitely many such clusters. But the probability should sum to one. So what we do is we take a stick of unit length, then we break it into two parts. The first part is the probability to pick up a particular cluster. Next, we take the remain of the stick, again break it into two parts. Precede and do this infinitely often, we will get a distribution over infinite number of clusters. To the end, we have a probability to pick a table and a dish over each table. Now it is very easy to give the generative model for Dirichlet process mixture model (Figure 3). This is the stick breaking part, using Beta distribution. This part is to sample dishes from which we can sample items later. The generative process is very similar to finite mixture model. Through the stick breaking process, we will get a multinomial distribution over infinite number of clusters. Just use this multinomial to sample
6 6 27 : Distributed Monte Carlo Markov Chain Figure 3: Graphical Representation of Dirichlet Process Model cluster index and then generate the item from the corresponding cluster. 3.2 Parallel Inference of Dirichlet Process Mixture Model Figure 4: Performance Measure of Different Inference Methods Now we look at different kinds of inference technique. In Gibbs sampling, we sample one random variable given the rest. That is very easy to parallelize on. If we are given V k and ηk, all Z n are independent of each other. What we can do is, we distribute the data into multiple processors. And sample Z n independently. But we have to sample V k and ηk jointly. The problem is it may show very poor mixing. Thus, it is rarely used in practice. The way to solve this problem is to use collapsed Gibbs sampling. We integrate V k and ηk out, which make all Z n inter connected. In this case, it is very difficulty to parallelize inference. There is a large cost in Gibbs sampling. Normally, in Gibbs sampling, if we fix the model size, the time complexity will increase linearly with the data size. If we look at the rate of convergence, it is O(N), where N is the number of data. In Dirichlet process mixture model, since the number of parameters is also increasing, the time taken would be larger. Here is an example (Figure 4). This is synthetic data where we know which cluster each item belongs to. This enables us to plot the accuracy over time. This is the log scale plot. What we see here is that it converges around 2000 minutes. This is for 1 million points, not very large. Another inference technique is variational inference. In variational inference, what we are trying to do is to
7 27 : Distributed Monte Carlo Markov Chain 7 approximate the posterior using some manageable distribution. To make it manageable, we remove some of the dependence structure. This can lead to very simple parallel scheme. But it doesn t perform as well as MCMC techniques. You need a lot of tuning of the parameters. If we parallel the variational inference on eight processors, it converges quite early, before eight minutes or so (Figure 4). But it converges to a point that is not great in accuracy. In sequential Monte Carlo method, we keep a pool of particles. You can update the pool of particles independently, which can be trivially parallelized. These are the results of SMC on 10 particles and 100 particles. Note that to remove the high variance problem, sometime, we have to go back and run MCMC on subset of the data that we have previously seen. But we don t have a good way to distribute that particular step, so usually we just use the naive way. Now we are done with the existing technique which can be trivially distributed, but leads to poor results. Next, we are going to discuss two methods. One is the naive method in which we distribute data on P processor and run the inference algorithm on each processor. Intermediately, we aggregate the sufficient statistics. This will give we a lot of failure. Because the color in one table is independent of the color in another table. But sometimes, for some model, if we keep doing this iteratively, it can converge and work well. There are other problems associated with this. Since this is nonparametric model, each processor can generate new clusters. How to combine the new clusters that have been found is a problem. Figure 5: The Equivalence between Auxiliary Variable Dirichlet Process and Ordinary Dirichlet Process The last parallel MCMC technique for Dirichlet process mixture model is based on the following idea: the Dirichlet Mixture of Dirichlet processes are Dirichlet processes. So what does the Dirichlet mixture of Dirichlet processes mean in Chinese restaurant process mean? Each restaurant has a key and the key can lead us to a particular restaurant. The key is from a multinomial distribution which is Dirichlet distributed. Then choosing the restaurant is another Dirichlet process. How this can help us to parallel the inference of Dirichlet process? Given the keys of all items, each of the processors is independent of other processors. Then we can run the favorite inference algorithm on each of the processor independently and combine the results. In terms of generative process, it is like this: D i the Dirichlet process for individual processor, π i is the key of an item. The new generative model is equivalent to the original Dirichlet process model (Figure 5). The key point of this method is, we introduce an auxiliary variable, which has a conditional independence structure, which help us to distribute the model. If we look at the posterior distribution of the original model and the posterior distribution of the auxiliary variable model, they are equivalent. Once we describe the model, the inference procedure is very simple. If we get the keys, we can run inference algorithm on each processor independently. In the second step, we are going to use MCMC method to infer the keys. We can select a cluster and propose to move the cluster from one processor to another, the move will be accepted based on the likelihood ratio. It seems that this will use a lot of network bandwidth. But in practice, we can do it as less often as we want. What we found in practice is that we can do less than 10 times for this step for 10 million points. As can be seen from Figure 4, the auxiliary variable method leads to great improvement of performance. Now let s look at the measure of perplexity. Perplexity denotes how surprise our model is when seeing new data. The lower, the better. As can be seen from the figure, the AVparallel method is much better than the
8 8 27 : Distributed Monte Carlo Markov Chain naive method. The message is that naive method will not always work. Another point is, if we can utilize the structure of the problem to explore conditional independence, the parallelization can be achieved easily. References [1] J. M. Hammersley et al., Monte Carlo Methods, Vol. 1, Springer, [2] Eric Xing, Lecture Note Approximation Inference: Parallel MCMC, Probabilistic Graphical Models Class, CMU, [3] J. Gonzalez et al., Parallel Gibbs Sampling: From Colored Fields to Thin Junction Trees, AISTATS, [4] Christopher M Bishop et al. Pattern Recognition and Machine Learning, Vol. 1, Springer, New York, 2006.
28 : Approximate Inference - Distributed MCMC
 10-708: Probabilistic Graphical Models, Spring 2015 28 : Approximate Inference - Distributed MCMC Lecturer: Avinava Dubey Scribes: Hakim Sidahmed, Aman Gupta 1 Introduction For many interesting problems,
10-708: Probabilistic Graphical Models, Spring 2015 28 : Approximate Inference - Distributed MCMC Lecturer: Avinava Dubey Scribes: Hakim Sidahmed, Aman Gupta 1 Introduction For many interesting problems,
Markov chain Monte Carlo Lecture 9
 Markov chain Monte Carlo Lecture 9 David Sontag New York University Slides adapted from Eric Xing and Qirong Ho (CMU) Limitations of Monte Carlo Direct (unconditional) sampling Hard to get rare events
Markov chain Monte Carlo Lecture 9 David Sontag New York University Slides adapted from Eric Xing and Qirong Ho (CMU) Limitations of Monte Carlo Direct (unconditional) sampling Hard to get rare events
16 : Approximate Inference: Markov Chain Monte Carlo
 10-708: Probabilistic Graphical Models 10-708, Spring 2017 16 : Approximate Inference: Markov Chain Monte Carlo Lecturer: Eric P. Xing Scribes: Yuan Yang, Chao-Ming Yen 1 Introduction As the target distribution
10-708: Probabilistic Graphical Models 10-708, Spring 2017 16 : Approximate Inference: Markov Chain Monte Carlo Lecturer: Eric P. Xing Scribes: Yuan Yang, Chao-Ming Yen 1 Introduction As the target distribution
MCMC and Gibbs Sampling. Kayhan Batmanghelich
 MCMC and Gibbs Sampling Kayhan Batmanghelich 1 Approaches to inference l Exact inference algorithms l l l The elimination algorithm Message-passing algorithm (sum-product, belief propagation) The junction
MCMC and Gibbs Sampling Kayhan Batmanghelich 1 Approaches to inference l Exact inference algorithms l l l The elimination algorithm Message-passing algorithm (sum-product, belief propagation) The junction
18 : Advanced topics in MCMC. 1 Gibbs Sampling (Continued from the last lecture)
 10-708: Probabilistic Graphical Models 10-708, Spring 2014 18 : Advanced topics in MCMC Lecturer: Eric P. Xing Scribes: Jessica Chemali, Seungwhan Moon 1 Gibbs Sampling (Continued from the last lecture)
10-708: Probabilistic Graphical Models 10-708, Spring 2014 18 : Advanced topics in MCMC Lecturer: Eric P. Xing Scribes: Jessica Chemali, Seungwhan Moon 1 Gibbs Sampling (Continued from the last lecture)
Bayesian Methods for Machine Learning
 Bayesian Methods for Machine Learning CS 584: Big Data Analytics Material adapted from Radford Neal s tutorial (http://ftp.cs.utoronto.ca/pub/radford/bayes-tut.pdf), Zoubin Ghahramni (http://hunch.net/~coms-4771/zoubin_ghahramani_bayesian_learning.pdf),
Bayesian Methods for Machine Learning CS 584: Big Data Analytics Material adapted from Radford Neal s tutorial (http://ftp.cs.utoronto.ca/pub/radford/bayes-tut.pdf), Zoubin Ghahramni (http://hunch.net/~coms-4771/zoubin_ghahramani_bayesian_learning.pdf),
CPSC 540: Machine Learning
 CPSC 540: Machine Learning MCMC and Non-Parametric Bayes Mark Schmidt University of British Columbia Winter 2016 Admin I went through project proposals: Some of you got a message on Piazza. No news is
CPSC 540: Machine Learning MCMC and Non-Parametric Bayes Mark Schmidt University of British Columbia Winter 2016 Admin I went through project proposals: Some of you got a message on Piazza. No news is
Machine Learning for Data Science (CS4786) Lecture 24
 Machine Learning for Data Science (CS4786) Lecture 24 Graphical Models: Approximate Inference Course Webpage : http://www.cs.cornell.edu/courses/cs4786/2016sp/ BELIEF PROPAGATION OR MESSAGE PASSING Each
Machine Learning for Data Science (CS4786) Lecture 24 Graphical Models: Approximate Inference Course Webpage : http://www.cs.cornell.edu/courses/cs4786/2016sp/ BELIEF PROPAGATION OR MESSAGE PASSING Each
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistical Sciences! rsalakhu@cs.toronto.edu! h0p://www.cs.utoronto.ca/~rsalakhu/ Lecture 7 Approximate
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistical Sciences! rsalakhu@cs.toronto.edu! h0p://www.cs.utoronto.ca/~rsalakhu/ Lecture 7 Approximate
17 : Markov Chain Monte Carlo
 10-708: Probabilistic Graphical Models, Spring 2015 17 : Markov Chain Monte Carlo Lecturer: Eric P. Xing Scribes: Heran Lin, Bin Deng, Yun Huang 1 Review of Monte Carlo Methods 1.1 Overview Monte Carlo
10-708: Probabilistic Graphical Models, Spring 2015 17 : Markov Chain Monte Carlo Lecturer: Eric P. Xing Scribes: Heran Lin, Bin Deng, Yun Huang 1 Review of Monte Carlo Methods 1.1 Overview Monte Carlo
Sampling Methods (11/30/04)
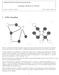 CS281A/Stat241A: Statistical Learning Theory Sampling Methods (11/30/04) Lecturer: Michael I. Jordan Scribe: Jaspal S. Sandhu 1 Gibbs Sampling Figure 1: Undirected and directed graphs, respectively, with
CS281A/Stat241A: Statistical Learning Theory Sampling Methods (11/30/04) Lecturer: Michael I. Jordan Scribe: Jaspal S. Sandhu 1 Gibbs Sampling Figure 1: Undirected and directed graphs, respectively, with
CSC 446 Notes: Lecture 13
 CSC 446 Notes: Lecture 3 The Problem We have already studied how to calculate the probability of a variable or variables using the message passing method. However, there are some times when the structure
CSC 446 Notes: Lecture 3 The Problem We have already studied how to calculate the probability of a variable or variables using the message passing method. However, there are some times when the structure
CSC 2541: Bayesian Methods for Machine Learning
 CSC 2541: Bayesian Methods for Machine Learning Radford M. Neal, University of Toronto, 2011 Lecture 3 More Markov Chain Monte Carlo Methods The Metropolis algorithm isn t the only way to do MCMC. We ll
CSC 2541: Bayesian Methods for Machine Learning Radford M. Neal, University of Toronto, 2011 Lecture 3 More Markov Chain Monte Carlo Methods The Metropolis algorithm isn t the only way to do MCMC. We ll
Non-Parametric Bayes
 Non-Parametric Bayes Mark Schmidt UBC Machine Learning Reading Group January 2016 Current Hot Topics in Machine Learning Bayesian learning includes: Gaussian processes. Approximate inference. Bayesian
Non-Parametric Bayes Mark Schmidt UBC Machine Learning Reading Group January 2016 Current Hot Topics in Machine Learning Bayesian learning includes: Gaussian processes. Approximate inference. Bayesian
Probabilistic Graphical Models Lecture 17: Markov chain Monte Carlo
 Probabilistic Graphical Models Lecture 17: Markov chain Monte Carlo Andrew Gordon Wilson www.cs.cmu.edu/~andrewgw Carnegie Mellon University March 18, 2015 1 / 45 Resources and Attribution Image credits,
Probabilistic Graphical Models Lecture 17: Markov chain Monte Carlo Andrew Gordon Wilson www.cs.cmu.edu/~andrewgw Carnegie Mellon University March 18, 2015 1 / 45 Resources and Attribution Image credits,
Part IV: Monte Carlo and nonparametric Bayes
 Part IV: Monte Carlo and nonparametric Bayes Outline Monte Carlo methods Nonparametric Bayesian models Outline Monte Carlo methods Nonparametric Bayesian models The Monte Carlo principle The expectation
Part IV: Monte Carlo and nonparametric Bayes Outline Monte Carlo methods Nonparametric Bayesian models Outline Monte Carlo methods Nonparametric Bayesian models The Monte Carlo principle The expectation
Lecturer: David Blei Lecture #3 Scribes: Jordan Boyd-Graber and Francisco Pereira October 1, 2007
 COS 597C: Bayesian Nonparametrics Lecturer: David Blei Lecture # Scribes: Jordan Boyd-Graber and Francisco Pereira October, 7 Gibbs Sampling with a DP First, let s recapitulate the model that we re using.
COS 597C: Bayesian Nonparametrics Lecturer: David Blei Lecture # Scribes: Jordan Boyd-Graber and Francisco Pereira October, 7 Gibbs Sampling with a DP First, let s recapitulate the model that we re using.
Computer Vision Group Prof. Daniel Cremers. 10a. Markov Chain Monte Carlo
 Group Prof. Daniel Cremers 10a. Markov Chain Monte Carlo Markov Chain Monte Carlo In high-dimensional spaces, rejection sampling and importance sampling are very inefficient An alternative is Markov Chain
Group Prof. Daniel Cremers 10a. Markov Chain Monte Carlo Markov Chain Monte Carlo In high-dimensional spaces, rejection sampling and importance sampling are very inefficient An alternative is Markov Chain
Reminder of some Markov Chain properties:
 Reminder of some Markov Chain properties: 1. a transition from one state to another occurs probabilistically 2. only state that matters is where you currently are (i.e. given present, future is independent
Reminder of some Markov Chain properties: 1. a transition from one state to another occurs probabilistically 2. only state that matters is where you currently are (i.e. given present, future is independent
Markov Chains and MCMC
 Markov Chains and MCMC CompSci 590.02 Instructor: AshwinMachanavajjhala Lecture 4 : 590.02 Spring 13 1 Recap: Monte Carlo Method If U is a universe of items, and G is a subset satisfying some property,
Markov Chains and MCMC CompSci 590.02 Instructor: AshwinMachanavajjhala Lecture 4 : 590.02 Spring 13 1 Recap: Monte Carlo Method If U is a universe of items, and G is a subset satisfying some property,
Graphical Models and Kernel Methods
 Graphical Models and Kernel Methods Jerry Zhu Department of Computer Sciences University of Wisconsin Madison, USA MLSS June 17, 2014 1 / 123 Outline Graphical Models Probabilistic Inference Directed vs.
Graphical Models and Kernel Methods Jerry Zhu Department of Computer Sciences University of Wisconsin Madison, USA MLSS June 17, 2014 1 / 123 Outline Graphical Models Probabilistic Inference Directed vs.
Computational statistics
 Computational statistics Markov Chain Monte Carlo methods Thierry Denœux March 2017 Thierry Denœux Computational statistics March 2017 1 / 71 Contents of this chapter When a target density f can be evaluated
Computational statistics Markov Chain Monte Carlo methods Thierry Denœux March 2017 Thierry Denœux Computational statistics March 2017 1 / 71 Contents of this chapter When a target density f can be evaluated
Spatial Normalized Gamma Process
 Spatial Normalized Gamma Process Vinayak Rao Yee Whye Teh Presented at NIPS 2009 Discussion and Slides by Eric Wang June 23, 2010 Outline Introduction Motivation The Gamma Process Spatial Normalized Gamma
Spatial Normalized Gamma Process Vinayak Rao Yee Whye Teh Presented at NIPS 2009 Discussion and Slides by Eric Wang June 23, 2010 Outline Introduction Motivation The Gamma Process Spatial Normalized Gamma
13: Variational inference II
 10-708: Probabilistic Graphical Models, Spring 2015 13: Variational inference II Lecturer: Eric P. Xing Scribes: Ronghuo Zheng, Zhiting Hu, Yuntian Deng 1 Introduction We started to talk about variational
10-708: Probabilistic Graphical Models, Spring 2015 13: Variational inference II Lecturer: Eric P. Xing Scribes: Ronghuo Zheng, Zhiting Hu, Yuntian Deng 1 Introduction We started to talk about variational
Introduction to Machine Learning CMU-10701
 Introduction to Machine Learning CMU-10701 Markov Chain Monte Carlo Methods Barnabás Póczos & Aarti Singh Contents Markov Chain Monte Carlo Methods Goal & Motivation Sampling Rejection Importance Markov
Introduction to Machine Learning CMU-10701 Markov Chain Monte Carlo Methods Barnabás Póczos & Aarti Singh Contents Markov Chain Monte Carlo Methods Goal & Motivation Sampling Rejection Importance Markov
Bagging During Markov Chain Monte Carlo for Smoother Predictions
 Bagging During Markov Chain Monte Carlo for Smoother Predictions Herbert K. H. Lee University of California, Santa Cruz Abstract: Making good predictions from noisy data is a challenging problem. Methods
Bagging During Markov Chain Monte Carlo for Smoother Predictions Herbert K. H. Lee University of California, Santa Cruz Abstract: Making good predictions from noisy data is a challenging problem. Methods
CSC 2541: Bayesian Methods for Machine Learning
 CSC 2541: Bayesian Methods for Machine Learning Radford M. Neal, University of Toronto, 2011 Lecture 4 Problem: Density Estimation We have observed data, y 1,..., y n, drawn independently from some unknown
CSC 2541: Bayesian Methods for Machine Learning Radford M. Neal, University of Toronto, 2011 Lecture 4 Problem: Density Estimation We have observed data, y 1,..., y n, drawn independently from some unknown
16 : Markov Chain Monte Carlo (MCMC)
 10-708: Probabilistic Graphical Models 10-708, Spring 2014 16 : Markov Chain Monte Carlo MCMC Lecturer: Matthew Gormley Scribes: Yining Wang, Renato Negrinho 1 Sampling from low-dimensional distributions
10-708: Probabilistic Graphical Models 10-708, Spring 2014 16 : Markov Chain Monte Carlo MCMC Lecturer: Matthew Gormley Scribes: Yining Wang, Renato Negrinho 1 Sampling from low-dimensional distributions
CS 343: Artificial Intelligence
 CS 343: Artificial Intelligence Bayes Nets: Sampling Prof. Scott Niekum The University of Texas at Austin [These slides based on those of Dan Klein and Pieter Abbeel for CS188 Intro to AI at UC Berkeley.
CS 343: Artificial Intelligence Bayes Nets: Sampling Prof. Scott Niekum The University of Texas at Austin [These slides based on those of Dan Klein and Pieter Abbeel for CS188 Intro to AI at UC Berkeley.
Pattern Recognition and Machine Learning
 Christopher M. Bishop Pattern Recognition and Machine Learning ÖSpri inger Contents Preface Mathematical notation Contents vii xi xiii 1 Introduction 1 1.1 Example: Polynomial Curve Fitting 4 1.2 Probability
Christopher M. Bishop Pattern Recognition and Machine Learning ÖSpri inger Contents Preface Mathematical notation Contents vii xi xiii 1 Introduction 1 1.1 Example: Polynomial Curve Fitting 4 1.2 Probability
Lecture 7 and 8: Markov Chain Monte Carlo
 Lecture 7 and 8: Markov Chain Monte Carlo 4F13: Machine Learning Zoubin Ghahramani and Carl Edward Rasmussen Department of Engineering University of Cambridge http://mlg.eng.cam.ac.uk/teaching/4f13/ Ghahramani
Lecture 7 and 8: Markov Chain Monte Carlo 4F13: Machine Learning Zoubin Ghahramani and Carl Edward Rasmussen Department of Engineering University of Cambridge http://mlg.eng.cam.ac.uk/teaching/4f13/ Ghahramani
The Particle Filter. PD Dr. Rudolph Triebel Computer Vision Group. Machine Learning for Computer Vision
 The Particle Filter Non-parametric implementation of Bayes filter Represents the belief (posterior) random state samples. by a set of This representation is approximate. Can represent distributions that
The Particle Filter Non-parametric implementation of Bayes filter Represents the belief (posterior) random state samples. by a set of This representation is approximate. Can represent distributions that
Bayes Nets: Sampling
 Bayes Nets: Sampling [These slides were created by Dan Klein and Pieter Abbeel for CS188 Intro to AI at UC Berkeley. All CS188 materials are available at http://ai.berkeley.edu.] Approximate Inference:
Bayes Nets: Sampling [These slides were created by Dan Klein and Pieter Abbeel for CS188 Intro to AI at UC Berkeley. All CS188 materials are available at http://ai.berkeley.edu.] Approximate Inference:
Lecture 13 : Variational Inference: Mean Field Approximation
 10-708: Probabilistic Graphical Models 10-708, Spring 2017 Lecture 13 : Variational Inference: Mean Field Approximation Lecturer: Willie Neiswanger Scribes: Xupeng Tong, Minxing Liu 1 Problem Setup 1.1
10-708: Probabilistic Graphical Models 10-708, Spring 2017 Lecture 13 : Variational Inference: Mean Field Approximation Lecturer: Willie Neiswanger Scribes: Xupeng Tong, Minxing Liu 1 Problem Setup 1.1
19 : Bayesian Nonparametrics: The Indian Buffet Process. 1 Latent Variable Models and the Indian Buffet Process
 10-708: Probabilistic Graphical Models, Spring 2015 19 : Bayesian Nonparametrics: The Indian Buffet Process Lecturer: Avinava Dubey Scribes: Rishav Das, Adam Brodie, and Hemank Lamba 1 Latent Variable
10-708: Probabilistic Graphical Models, Spring 2015 19 : Bayesian Nonparametrics: The Indian Buffet Process Lecturer: Avinava Dubey Scribes: Rishav Das, Adam Brodie, and Hemank Lamba 1 Latent Variable
Computer Vision Group Prof. Daniel Cremers. 11. Sampling Methods: Markov Chain Monte Carlo
 Group Prof. Daniel Cremers 11. Sampling Methods: Markov Chain Monte Carlo Markov Chain Monte Carlo In high-dimensional spaces, rejection sampling and importance sampling are very inefficient An alternative
Group Prof. Daniel Cremers 11. Sampling Methods: Markov Chain Monte Carlo Markov Chain Monte Carlo In high-dimensional spaces, rejection sampling and importance sampling are very inefficient An alternative
13 : Variational Inference: Loopy Belief Propagation and Mean Field
 10-708: Probabilistic Graphical Models 10-708, Spring 2012 13 : Variational Inference: Loopy Belief Propagation and Mean Field Lecturer: Eric P. Xing Scribes: Peter Schulam and William Wang 1 Introduction
10-708: Probabilistic Graphical Models 10-708, Spring 2012 13 : Variational Inference: Loopy Belief Propagation and Mean Field Lecturer: Eric P. Xing Scribes: Peter Schulam and William Wang 1 Introduction
Introduction to Machine Learning
 Introduction to Machine Learning Brown University CSCI 1950-F, Spring 2012 Prof. Erik Sudderth Lecture 25: Markov Chain Monte Carlo (MCMC) Course Review and Advanced Topics Many figures courtesy Kevin
Introduction to Machine Learning Brown University CSCI 1950-F, Spring 2012 Prof. Erik Sudderth Lecture 25: Markov Chain Monte Carlo (MCMC) Course Review and Advanced Topics Many figures courtesy Kevin
Sampling Algorithms for Probabilistic Graphical models
 Sampling Algorithms for Probabilistic Graphical models Vibhav Gogate University of Washington References: Chapter 12 of Probabilistic Graphical models: Principles and Techniques by Daphne Koller and Nir
Sampling Algorithms for Probabilistic Graphical models Vibhav Gogate University of Washington References: Chapter 12 of Probabilistic Graphical models: Principles and Techniques by Daphne Koller and Nir
CS281B / Stat 241B : Statistical Learning Theory Lecture: #22 on 19 Apr Dirichlet Process I
 X i Ν CS281B / Stat 241B : Statistical Learning Theory Lecture: #22 on 19 Apr 2004 Dirichlet Process I Lecturer: Prof. Michael Jordan Scribe: Daniel Schonberg dschonbe@eecs.berkeley.edu 22.1 Dirichlet
X i Ν CS281B / Stat 241B : Statistical Learning Theory Lecture: #22 on 19 Apr 2004 Dirichlet Process I Lecturer: Prof. Michael Jordan Scribe: Daniel Schonberg dschonbe@eecs.berkeley.edu 22.1 Dirichlet
Phylogenetics: Bayesian Phylogenetic Analysis. COMP Spring 2015 Luay Nakhleh, Rice University
 Phylogenetics: Bayesian Phylogenetic Analysis COMP 571 - Spring 2015 Luay Nakhleh, Rice University Bayes Rule P(X = x Y = y) = P(X = x, Y = y) P(Y = y) = P(X = x)p(y = y X = x) P x P(X = x 0 )P(Y = y X
Phylogenetics: Bayesian Phylogenetic Analysis COMP 571 - Spring 2015 Luay Nakhleh, Rice University Bayes Rule P(X = x Y = y) = P(X = x, Y = y) P(Y = y) = P(X = x)p(y = y X = x) P x P(X = x 0 )P(Y = y X
MCMC: Markov Chain Monte Carlo
 I529: Machine Learning in Bioinformatics (Spring 2013) MCMC: Markov Chain Monte Carlo Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Review of Markov
I529: Machine Learning in Bioinformatics (Spring 2013) MCMC: Markov Chain Monte Carlo Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2013 Contents Review of Markov
CS242: Probabilistic Graphical Models Lecture 7B: Markov Chain Monte Carlo & Gibbs Sampling
 CS242: Probabilistic Graphical Models Lecture 7B: Markov Chain Monte Carlo & Gibbs Sampling Professor Erik Sudderth Brown University Computer Science October 27, 2016 Some figures and materials courtesy
CS242: Probabilistic Graphical Models Lecture 7B: Markov Chain Monte Carlo & Gibbs Sampling Professor Erik Sudderth Brown University Computer Science October 27, 2016 Some figures and materials courtesy
Bayesian Networks BY: MOHAMAD ALSABBAGH
 Bayesian Networks BY: MOHAMAD ALSABBAGH Outlines Introduction Bayes Rule Bayesian Networks (BN) Representation Size of a Bayesian Network Inference via BN BN Learning Dynamic BN Introduction Conditional
Bayesian Networks BY: MOHAMAD ALSABBAGH Outlines Introduction Bayes Rule Bayesian Networks (BN) Representation Size of a Bayesian Network Inference via BN BN Learning Dynamic BN Introduction Conditional
April 20th, Advanced Topics in Machine Learning California Institute of Technology. Markov Chain Monte Carlo for Machine Learning
 for for Advanced Topics in California Institute of Technology April 20th, 2017 1 / 50 Table of Contents for 1 2 3 4 2 / 50 History of methods for Enrico Fermi used to calculate incredibly accurate predictions
for for Advanced Topics in California Institute of Technology April 20th, 2017 1 / 50 Table of Contents for 1 2 3 4 2 / 50 History of methods for Enrico Fermi used to calculate incredibly accurate predictions
Probabilistic Graphical Models
 2016 Robert Nowak Probabilistic Graphical Models 1 Introduction We have focused mainly on linear models for signals, in particular the subspace model x = Uθ, where U is a n k matrix and θ R k is a vector
2016 Robert Nowak Probabilistic Graphical Models 1 Introduction We have focused mainly on linear models for signals, in particular the subspace model x = Uθ, where U is a n k matrix and θ R k is a vector
Markov Chain Monte Carlo
 Markov Chain Monte Carlo (and Bayesian Mixture Models) David M. Blei Columbia University October 14, 2014 We have discussed probabilistic modeling, and have seen how the posterior distribution is the critical
Markov Chain Monte Carlo (and Bayesian Mixture Models) David M. Blei Columbia University October 14, 2014 We have discussed probabilistic modeling, and have seen how the posterior distribution is the critical
Approximate inference, Sampling & Variational inference Fall Cours 9 November 25
 Approimate inference, Sampling & Variational inference Fall 2015 Cours 9 November 25 Enseignant: Guillaume Obozinski Scribe: Basile Clément, Nathan de Lara 9.1 Approimate inference with MCMC 9.1.1 Gibbs
Approimate inference, Sampling & Variational inference Fall 2015 Cours 9 November 25 Enseignant: Guillaume Obozinski Scribe: Basile Clément, Nathan de Lara 9.1 Approimate inference with MCMC 9.1.1 Gibbs
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 11 Project
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 11 Project
Introduction to Bayesian Statistics and Markov Chain Monte Carlo Estimation. EPSY 905: Multivariate Analysis Spring 2016 Lecture #10: April 6, 2016
 Introduction to Bayesian Statistics and Markov Chain Monte Carlo Estimation EPSY 905: Multivariate Analysis Spring 2016 Lecture #10: April 6, 2016 EPSY 905: Intro to Bayesian and MCMC Today s Class An
Introduction to Bayesian Statistics and Markov Chain Monte Carlo Estimation EPSY 905: Multivariate Analysis Spring 2016 Lecture #10: April 6, 2016 EPSY 905: Intro to Bayesian and MCMC Today s Class An
DAG models and Markov Chain Monte Carlo methods a short overview
 DAG models and Markov Chain Monte Carlo methods a short overview Søren Højsgaard Institute of Genetics and Biotechnology University of Aarhus August 18, 2008 Printed: August 18, 2008 File: DAGMC-Lecture.tex
DAG models and Markov Chain Monte Carlo methods a short overview Søren Højsgaard Institute of Genetics and Biotechnology University of Aarhus August 18, 2008 Printed: August 18, 2008 File: DAGMC-Lecture.tex
Probabilistic Graphical Models
 School of Computer Science Probabilistic Graphical Models Infinite Feature Models: The Indian Buffet Process Eric Xing Lecture 21, April 2, 214 Acknowledgement: slides first drafted by Sinead Williamson
School of Computer Science Probabilistic Graphical Models Infinite Feature Models: The Indian Buffet Process Eric Xing Lecture 21, April 2, 214 Acknowledgement: slides first drafted by Sinead Williamson
Outline. Binomial, Multinomial, Normal, Beta, Dirichlet. Posterior mean, MAP, credible interval, posterior distribution
 Outline A short review on Bayesian analysis. Binomial, Multinomial, Normal, Beta, Dirichlet Posterior mean, MAP, credible interval, posterior distribution Gibbs sampling Revisit the Gaussian mixture model
Outline A short review on Bayesian analysis. Binomial, Multinomial, Normal, Beta, Dirichlet Posterior mean, MAP, credible interval, posterior distribution Gibbs sampling Revisit the Gaussian mixture model
Computer Vision Group Prof. Daniel Cremers. 14. Sampling Methods
 Prof. Daniel Cremers 14. Sampling Methods Sampling Methods Sampling Methods are widely used in Computer Science as an approximation of a deterministic algorithm to represent uncertainty without a parametric
Prof. Daniel Cremers 14. Sampling Methods Sampling Methods Sampling Methods are widely used in Computer Science as an approximation of a deterministic algorithm to represent uncertainty without a parametric
Monte Carlo Methods. Geoff Gordon February 9, 2006
 Monte Carlo Methods Geoff Gordon ggordon@cs.cmu.edu February 9, 2006 Numerical integration problem 5 4 3 f(x,y) 2 1 1 0 0.5 0 X 0.5 1 1 0.8 0.6 0.4 Y 0.2 0 0.2 0.4 0.6 0.8 1 x X f(x)dx Used for: function
Monte Carlo Methods Geoff Gordon ggordon@cs.cmu.edu February 9, 2006 Numerical integration problem 5 4 3 f(x,y) 2 1 1 0 0.5 0 X 0.5 1 1 0.8 0.6 0.4 Y 0.2 0 0.2 0.4 0.6 0.8 1 x X f(x)dx Used for: function
Particle-Based Approximate Inference on Graphical Model
 article-based Approimate Inference on Graphical Model Reference: robabilistic Graphical Model Ch. 2 Koller & Friedman CMU, 0-708, Fall 2009 robabilistic Graphical Models Lectures 8,9 Eric ing attern Recognition
article-based Approimate Inference on Graphical Model Reference: robabilistic Graphical Model Ch. 2 Koller & Friedman CMU, 0-708, Fall 2009 robabilistic Graphical Models Lectures 8,9 Eric ing attern Recognition
arxiv: v1 [stat.co] 23 Apr 2018
![arxiv: v1 [stat.co] 23 Apr 2018 arxiv: v1 [stat.co] 23 Apr 2018](/thumbs/93/112368199.jpg) Bayesian Updating and Uncertainty Quantification using Sequential Tempered MCMC with the Rank-One Modified Metropolis Algorithm Thomas A. Catanach and James L. Beck arxiv:1804.08738v1 [stat.co] 23 Apr
Bayesian Updating and Uncertainty Quantification using Sequential Tempered MCMC with the Rank-One Modified Metropolis Algorithm Thomas A. Catanach and James L. Beck arxiv:1804.08738v1 [stat.co] 23 Apr
Pattern Recognition and Machine Learning. Bishop Chapter 11: Sampling Methods
 Pattern Recognition and Machine Learning Chapter 11: Sampling Methods Elise Arnaud Jakob Verbeek May 22, 2008 Outline of the chapter 11.1 Basic Sampling Algorithms 11.2 Markov Chain Monte Carlo 11.3 Gibbs
Pattern Recognition and Machine Learning Chapter 11: Sampling Methods Elise Arnaud Jakob Verbeek May 22, 2008 Outline of the chapter 11.1 Basic Sampling Algorithms 11.2 Markov Chain Monte Carlo 11.3 Gibbs
Markov Chain Monte Carlo Inference. Siamak Ravanbakhsh Winter 2018
 Graphical Models Markov Chain Monte Carlo Inference Siamak Ravanbakhsh Winter 2018 Learning objectives Markov chains the idea behind Markov Chain Monte Carlo (MCMC) two important examples: Gibbs sampling
Graphical Models Markov Chain Monte Carlo Inference Siamak Ravanbakhsh Winter 2018 Learning objectives Markov chains the idea behind Markov Chain Monte Carlo (MCMC) two important examples: Gibbs sampling
Advanced Machine Learning
 Advanced Machine Learning Nonparametric Bayesian Models --Learning/Reasoning in Open Possible Worlds Eric Xing Lecture 7, August 4, 2009 Reading: Eric Xing Eric Xing @ CMU, 2006-2009 Clustering Eric Xing
Advanced Machine Learning Nonparametric Bayesian Models --Learning/Reasoning in Open Possible Worlds Eric Xing Lecture 7, August 4, 2009 Reading: Eric Xing Eric Xing @ CMU, 2006-2009 Clustering Eric Xing
STA 4273H: Sta-s-cal Machine Learning
 STA 4273H: Sta-s-cal Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistical Sciences! rsalakhu@cs.toronto.edu! h0p://www.cs.utoronto.ca/~rsalakhu/ Lecture 2 In our
STA 4273H: Sta-s-cal Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistical Sciences! rsalakhu@cs.toronto.edu! h0p://www.cs.utoronto.ca/~rsalakhu/ Lecture 2 In our
Lecture 6: Markov Chain Monte Carlo
 Lecture 6: Markov Chain Monte Carlo D. Jason Koskinen koskinen@nbi.ku.dk Photo by Howard Jackman University of Copenhagen Advanced Methods in Applied Statistics Feb - Apr 2016 Niels Bohr Institute 2 Outline
Lecture 6: Markov Chain Monte Carlo D. Jason Koskinen koskinen@nbi.ku.dk Photo by Howard Jackman University of Copenhagen Advanced Methods in Applied Statistics Feb - Apr 2016 Niels Bohr Institute 2 Outline
Bayesian networks: approximate inference
 Bayesian networks: approximate inference Machine Intelligence Thomas D. Nielsen September 2008 Approximative inference September 2008 1 / 25 Motivation Because of the (worst-case) intractability of exact
Bayesian networks: approximate inference Machine Intelligence Thomas D. Nielsen September 2008 Approximative inference September 2008 1 / 25 Motivation Because of the (worst-case) intractability of exact
David B. Dahl. Department of Statistics, and Department of Biostatistics & Medical Informatics University of Wisconsin Madison
 AN IMPROVED MERGE-SPLIT SAMPLER FOR CONJUGATE DIRICHLET PROCESS MIXTURE MODELS David B. Dahl dbdahl@stat.wisc.edu Department of Statistics, and Department of Biostatistics & Medical Informatics University
AN IMPROVED MERGE-SPLIT SAMPLER FOR CONJUGATE DIRICHLET PROCESS MIXTURE MODELS David B. Dahl dbdahl@stat.wisc.edu Department of Statistics, and Department of Biostatistics & Medical Informatics University
19 : Slice Sampling and HMC
 10-708: Probabilistic Graphical Models 10-708, Spring 2018 19 : Slice Sampling and HMC Lecturer: Kayhan Batmanghelich Scribes: Boxiang Lyu 1 MCMC (Auxiliary Variables Methods) In inference, we are often
10-708: Probabilistic Graphical Models 10-708, Spring 2018 19 : Slice Sampling and HMC Lecturer: Kayhan Batmanghelich Scribes: Boxiang Lyu 1 MCMC (Auxiliary Variables Methods) In inference, we are often
Bayesian Nonparametrics: Dirichlet Process
 Bayesian Nonparametrics: Dirichlet Process Yee Whye Teh Gatsby Computational Neuroscience Unit, UCL http://www.gatsby.ucl.ac.uk/~ywteh/teaching/npbayes2012 Dirichlet Process Cornerstone of modern Bayesian
Bayesian Nonparametrics: Dirichlet Process Yee Whye Teh Gatsby Computational Neuroscience Unit, UCL http://www.gatsby.ucl.ac.uk/~ywteh/teaching/npbayes2012 Dirichlet Process Cornerstone of modern Bayesian
CSCI 5822 Probabilistic Model of Human and Machine Learning. Mike Mozer University of Colorado
 CSCI 5822 Probabilistic Model of Human and Machine Learning Mike Mozer University of Colorado Topics Language modeling Hierarchical processes Pitman-Yor processes Based on work of Teh (2006), A hierarchical
CSCI 5822 Probabilistic Model of Human and Machine Learning Mike Mozer University of Colorado Topics Language modeling Hierarchical processes Pitman-Yor processes Based on work of Teh (2006), A hierarchical
CS 188: Artificial Intelligence. Bayes Nets
 CS 188: Artificial Intelligence Probabilistic Inference: Enumeration, Variable Elimination, Sampling Pieter Abbeel UC Berkeley Many slides over this course adapted from Dan Klein, Stuart Russell, Andrew
CS 188: Artificial Intelligence Probabilistic Inference: Enumeration, Variable Elimination, Sampling Pieter Abbeel UC Berkeley Many slides over this course adapted from Dan Klein, Stuart Russell, Andrew
Bayes Networks. CS540 Bryan R Gibson University of Wisconsin-Madison. Slides adapted from those used by Prof. Jerry Zhu, CS540-1
 Bayes Networks CS540 Bryan R Gibson University of Wisconsin-Madison Slides adapted from those used by Prof. Jerry Zhu, CS540-1 1 / 59 Outline Joint Probability: great for inference, terrible to obtain
Bayes Networks CS540 Bryan R Gibson University of Wisconsin-Madison Slides adapted from those used by Prof. Jerry Zhu, CS540-1 1 / 59 Outline Joint Probability: great for inference, terrible to obtain
Bayesian Estimation with Sparse Grids
 Bayesian Estimation with Sparse Grids Kenneth L. Judd and Thomas M. Mertens Institute on Computational Economics August 7, 27 / 48 Outline Introduction 2 Sparse grids Construction Integration with sparse
Bayesian Estimation with Sparse Grids Kenneth L. Judd and Thomas M. Mertens Institute on Computational Economics August 7, 27 / 48 Outline Introduction 2 Sparse grids Construction Integration with sparse
p L yi z n m x N n xi
 y i z n x n N x i Overview Directed and undirected graphs Conditional independence Exact inference Latent variables and EM Variational inference Books statistical perspective Graphical Models, S. Lauritzen
y i z n x n N x i Overview Directed and undirected graphs Conditional independence Exact inference Latent variables and EM Variational inference Books statistical perspective Graphical Models, S. Lauritzen
Robert Collins CSE586, PSU Intro to Sampling Methods
 Robert Collins Intro to Sampling Methods CSE586 Computer Vision II Penn State Univ Robert Collins A Brief Overview of Sampling Monte Carlo Integration Sampling and Expected Values Inverse Transform Sampling
Robert Collins Intro to Sampling Methods CSE586 Computer Vision II Penn State Univ Robert Collins A Brief Overview of Sampling Monte Carlo Integration Sampling and Expected Values Inverse Transform Sampling
STA 414/2104: Machine Learning
 STA 414/2104: Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistics! rsalakhu@cs.toronto.edu! http://www.cs.toronto.edu/~rsalakhu/ Lecture 9 Sequential Data So far
STA 414/2104: Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistics! rsalakhu@cs.toronto.edu! http://www.cs.toronto.edu/~rsalakhu/ Lecture 9 Sequential Data So far
Bayesian Inference and MCMC
 Bayesian Inference and MCMC Aryan Arbabi Partly based on MCMC slides from CSC412 Fall 2018 1 / 18 Bayesian Inference - Motivation Consider we have a data set D = {x 1,..., x n }. E.g each x i can be the
Bayesian Inference and MCMC Aryan Arbabi Partly based on MCMC slides from CSC412 Fall 2018 1 / 18 Bayesian Inference - Motivation Consider we have a data set D = {x 1,..., x n }. E.g each x i can be the
1 Probabilities. 1.1 Basics 1 PROBABILITIES
 1 PROBABILITIES 1 Probabilities Probability is a tricky word usually meaning the likelyhood of something occuring or how frequent something is. Obviously, if something happens frequently, then its probability
1 PROBABILITIES 1 Probabilities Probability is a tricky word usually meaning the likelyhood of something occuring or how frequent something is. Obviously, if something happens frequently, then its probability
Markov chain Monte Carlo methods in atmospheric remote sensing
 1 / 45 Markov chain Monte Carlo methods in atmospheric remote sensing Johanna Tamminen johanna.tamminen@fmi.fi ESA Summer School on Earth System Monitoring and Modeling July 3 Aug 11, 212, Frascati July,
1 / 45 Markov chain Monte Carlo methods in atmospheric remote sensing Johanna Tamminen johanna.tamminen@fmi.fi ESA Summer School on Earth System Monitoring and Modeling July 3 Aug 11, 212, Frascati July,
26 : Spectral GMs. Lecturer: Eric P. Xing Scribes: Guillermo A Cidre, Abelino Jimenez G.
 10-708: Probabilistic Graphical Models, Spring 2015 26 : Spectral GMs Lecturer: Eric P. Xing Scribes: Guillermo A Cidre, Abelino Jimenez G. 1 Introduction A common task in machine learning is to work with
10-708: Probabilistic Graphical Models, Spring 2015 26 : Spectral GMs Lecturer: Eric P. Xing Scribes: Guillermo A Cidre, Abelino Jimenez G. 1 Introduction A common task in machine learning is to work with
Markov Chain Monte Carlo
 1 Motivation 1.1 Bayesian Learning Markov Chain Monte Carlo Yale Chang In Bayesian learning, given data X, we make assumptions on the generative process of X by introducing hidden variables Z: p(z): prior
1 Motivation 1.1 Bayesian Learning Markov Chain Monte Carlo Yale Chang In Bayesian learning, given data X, we make assumptions on the generative process of X by introducing hidden variables Z: p(z): prior
Probabilistic Machine Learning
 Probabilistic Machine Learning Bayesian Nets, MCMC, and more Marek Petrik 4/18/2017 Based on: P. Murphy, K. (2012). Machine Learning: A Probabilistic Perspective. Chapter 10. Conditional Independence Independent
Probabilistic Machine Learning Bayesian Nets, MCMC, and more Marek Petrik 4/18/2017 Based on: P. Murphy, K. (2012). Machine Learning: A Probabilistic Perspective. Chapter 10. Conditional Independence Independent
References. Markov-Chain Monte Carlo. Recall: Sampling Motivation. Problem. Recall: Sampling Methods. CSE586 Computer Vision II
 References Markov-Chain Monte Carlo CSE586 Computer Vision II Spring 2010, Penn State Univ. Recall: Sampling Motivation If we can generate random samples x i from a given distribution P(x), then we can
References Markov-Chain Monte Carlo CSE586 Computer Vision II Spring 2010, Penn State Univ. Recall: Sampling Motivation If we can generate random samples x i from a given distribution P(x), then we can
Monte Carlo Methods. Leon Gu CSD, CMU
 Monte Carlo Methods Leon Gu CSD, CMU Approximate Inference EM: y-observed variables; x-hidden variables; θ-parameters; E-step: q(x) = p(x y, θ t 1 ) M-step: θ t = arg max E q(x) [log p(y, x θ)] θ Monte
Monte Carlo Methods Leon Gu CSD, CMU Approximate Inference EM: y-observed variables; x-hidden variables; θ-parameters; E-step: q(x) = p(x y, θ t 1 ) M-step: θ t = arg max E q(x) [log p(y, x θ)] θ Monte
Answers and expectations
 Answers and expectations For a function f(x) and distribution P(x), the expectation of f with respect to P is The expectation is the average of f, when x is drawn from the probability distribution P E
Answers and expectations For a function f(x) and distribution P(x), the expectation of f with respect to P is The expectation is the average of f, when x is drawn from the probability distribution P E
SAMPLING ALGORITHMS. In general. Inference in Bayesian models
 SAMPLING ALGORITHMS SAMPLING ALGORITHMS In general A sampling algorithm is an algorithm that outputs samples x 1, x 2,... from a given distribution P or density p. Sampling algorithms can for example be
SAMPLING ALGORITHMS SAMPLING ALGORITHMS In general A sampling algorithm is an algorithm that outputs samples x 1, x 2,... from a given distribution P or density p. Sampling algorithms can for example be
Markov Chain Monte Carlo
 Markov Chain Monte Carlo Recall: To compute the expectation E ( h(y ) ) we use the approximation E(h(Y )) 1 n n h(y ) t=1 with Y (1),..., Y (n) h(y). Thus our aim is to sample Y (1),..., Y (n) from f(y).
Markov Chain Monte Carlo Recall: To compute the expectation E ( h(y ) ) we use the approximation E(h(Y )) 1 n n h(y ) t=1 with Y (1),..., Y (n) h(y). Thus our aim is to sample Y (1),..., Y (n) from f(y).
The Recycling Gibbs Sampler for Efficient Learning
 The Recycling Gibbs Sampler for Efficient Learning L. Martino, V. Elvira, G. Camps-Valls Universidade de São Paulo, São Carlos (Brazil). Télécom ParisTech, Université Paris-Saclay. (France), Universidad
The Recycling Gibbs Sampler for Efficient Learning L. Martino, V. Elvira, G. Camps-Valls Universidade de São Paulo, São Carlos (Brazil). Télécom ParisTech, Université Paris-Saclay. (France), Universidad
14 : Approximate Inference Monte Carlo Methods
 10-708: Probabilistic Graphical Models 10-708, Spring 2018 14 : Approximate Inference Monte Carlo Methods Lecturer: Kayhan Batmanghelich Scribes: Biswajit Paria, Prerna Chiersal 1 Introduction We have
10-708: Probabilistic Graphical Models 10-708, Spring 2018 14 : Approximate Inference Monte Carlo Methods Lecturer: Kayhan Batmanghelich Scribes: Biswajit Paria, Prerna Chiersal 1 Introduction We have
Sequential Monte Carlo Methods for Bayesian Computation
 Sequential Monte Carlo Methods for Bayesian Computation A. Doucet Kyoto Sept. 2012 A. Doucet (MLSS Sept. 2012) Sept. 2012 1 / 136 Motivating Example 1: Generic Bayesian Model Let X be a vector parameter
Sequential Monte Carlo Methods for Bayesian Computation A. Doucet Kyoto Sept. 2012 A. Doucet (MLSS Sept. 2012) Sept. 2012 1 / 136 Motivating Example 1: Generic Bayesian Model Let X be a vector parameter
Computer Vision Group Prof. Daniel Cremers. 11. Sampling Methods
 Prof. Daniel Cremers 11. Sampling Methods Sampling Methods Sampling Methods are widely used in Computer Science as an approximation of a deterministic algorithm to represent uncertainty without a parametric
Prof. Daniel Cremers 11. Sampling Methods Sampling Methods Sampling Methods are widely used in Computer Science as an approximation of a deterministic algorithm to represent uncertainty without a parametric
Markov-Chain Monte Carlo
 Markov-Chain Monte Carlo CSE586 Computer Vision II Spring 2010, Penn State Univ. References Recall: Sampling Motivation If we can generate random samples x i from a given distribution P(x), then we can
Markov-Chain Monte Carlo CSE586 Computer Vision II Spring 2010, Penn State Univ. References Recall: Sampling Motivation If we can generate random samples x i from a given distribution P(x), then we can
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 3 Linear
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 3 Linear
Recent Advances in Bayesian Inference Techniques
 Recent Advances in Bayesian Inference Techniques Christopher M. Bishop Microsoft Research, Cambridge, U.K. research.microsoft.com/~cmbishop SIAM Conference on Data Mining, April 2004 Abstract Bayesian
Recent Advances in Bayesian Inference Techniques Christopher M. Bishop Microsoft Research, Cambridge, U.K. research.microsoft.com/~cmbishop SIAM Conference on Data Mining, April 2004 Abstract Bayesian
Machine Learning Techniques for Computer Vision
 Machine Learning Techniques for Computer Vision Part 2: Unsupervised Learning Microsoft Research Cambridge x 3 1 0.5 0.2 0 0.5 0.3 0 0.5 1 ECCV 2004, Prague x 2 x 1 Overview of Part 2 Mixture models EM
Machine Learning Techniques for Computer Vision Part 2: Unsupervised Learning Microsoft Research Cambridge x 3 1 0.5 0.2 0 0.5 0.3 0 0.5 1 ECCV 2004, Prague x 2 x 1 Overview of Part 2 Mixture models EM
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 7 Approximate
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 7 Approximate
13 : Variational Inference: Loopy Belief Propagation
 10-708: Probabilistic Graphical Models 10-708, Spring 2014 13 : Variational Inference: Loopy Belief Propagation Lecturer: Eric P. Xing Scribes: Rajarshi Das, Zhengzhong Liu, Dishan Gupta 1 Introduction
10-708: Probabilistic Graphical Models 10-708, Spring 2014 13 : Variational Inference: Loopy Belief Propagation Lecturer: Eric P. Xing Scribes: Rajarshi Das, Zhengzhong Liu, Dishan Gupta 1 Introduction
MCMC notes by Mark Holder
 MCMC notes by Mark Holder Bayesian inference Ultimately, we want to make probability statements about true values of parameters, given our data. For example P(α 0 < α 1 X). According to Bayes theorem:
MCMC notes by Mark Holder Bayesian inference Ultimately, we want to make probability statements about true values of parameters, given our data. For example P(α 0 < α 1 X). According to Bayes theorem:
Bayesian nonparametrics
 Bayesian nonparametrics 1 Some preliminaries 1.1 de Finetti s theorem We will start our discussion with this foundational theorem. We will assume throughout all variables are defined on the probability
Bayesian nonparametrics 1 Some preliminaries 1.1 de Finetti s theorem We will start our discussion with this foundational theorem. We will assume throughout all variables are defined on the probability
Inference in Bayesian Networks
 Andrea Passerini passerini@disi.unitn.it Machine Learning Inference in graphical models Description Assume we have evidence e on the state of a subset of variables E in the model (i.e. Bayesian Network)
Andrea Passerini passerini@disi.unitn.it Machine Learning Inference in graphical models Description Assume we have evidence e on the state of a subset of variables E in the model (i.e. Bayesian Network)
Stat 451 Lecture Notes Markov Chain Monte Carlo. Ryan Martin UIC
 Stat 451 Lecture Notes 07 12 Markov Chain Monte Carlo Ryan Martin UIC www.math.uic.edu/~rgmartin 1 Based on Chapters 8 9 in Givens & Hoeting, Chapters 25 27 in Lange 2 Updated: April 4, 2016 1 / 42 Outline
Stat 451 Lecture Notes 07 12 Markov Chain Monte Carlo Ryan Martin UIC www.math.uic.edu/~rgmartin 1 Based on Chapters 8 9 in Givens & Hoeting, Chapters 25 27 in Lange 2 Updated: April 4, 2016 1 / 42 Outline
Gaussian Mixture Model
 Case Study : Document Retrieval MAP EM, Latent Dirichlet Allocation, Gibbs Sampling Machine Learning/Statistics for Big Data CSE599C/STAT59, University of Washington Emily Fox 0 Emily Fox February 5 th,
Case Study : Document Retrieval MAP EM, Latent Dirichlet Allocation, Gibbs Sampling Machine Learning/Statistics for Big Data CSE599C/STAT59, University of Washington Emily Fox 0 Emily Fox February 5 th,
Stochastic Simulation
 Stochastic Simulation Idea: probabilities samples Get probabilities from samples: X count x 1 n 1. x k total. n k m X probability x 1. n 1 /m. x k n k /m If we could sample from a variable s (posterior)
Stochastic Simulation Idea: probabilities samples Get probabilities from samples: X count x 1 n 1. x k total. n k m X probability x 1. n 1 /m. x k n k /m If we could sample from a variable s (posterior)
