The DNA-Based Algorithms of Implementing Arithmetical Operations of Complex Vectors on a Biological Computer
|
|
|
- Cameron Brown
- 5 years ago
- Views:
Transcription
1 IEEE TRANSACTIONS ON NANOBIOSCIENCE VOL 14 NO 8 DECEMBER The DNA-Based Algorithms of Implementing Arithmetical Operations of Complex Vectors on a Biological Computer Weng-Long Chang Athanasios V Vasilakos Michael Shan-HuiHo Abstract Here we show that arithmetical operations of complex vectors can be implemented by means of the proposed DNA-based algorithms Index Terms DNA-based algorithms molecular computing I INTRODUCTION F ROM [1] [2] MANY biological algorithms of solving different problems were introduced From [3] molecular algorithms of implementing biomolecular databases were proposed From [4] quantum algorithms of implementing biomolecular solutions of the vertex cover problem were proposed Our major contributions in this journal paper are as follows We show that addition of complex vectors closure axioms of addition of complex vector can be implemented by means of biological operations DNA strs II THE FORMAL MODEL OF COMPUTATION DNA (deoxyribonucleic acid) in [1] [2] includes polymer chains which are commonly regarded as DNA strs Each str may be made of a sequence of nucleotides or bases attached to a sugar-phosphate backbone The four DNA nucleotides are adenine guanine cytosine thymine commonly abbreviated to A G C T respectively The following biomolecular operations will be applied to develop molecular algorithms of implementing arithmetical operations of complex vectors Their implementation can be found in [1] Definition 2-1: Given set a bit the biomolecular operation Append-Head appends onto the head of every element in set The formal representation is written as Manuscript received May ; revised July ; accepted September Date of publication October ; date of current version January This work was supported by National Science Foundation of Republic of China under Grants E CC3 Asterisk indicates corresponding author *W-L Chang is with the Department of Computer Science Information Engineering National Kaohsiung University of Applied Sciences Kaohsiung 807 Taiwan ( changwl@cckuasedutw) A V Vasilakos is with the Department of Computer Science National Technical University of Athens Greece ( vasilako@athforthnetgr) M Ho is with Computer Center Institute of Electrical Engineering National Taipei University Taiwan ( MHoInCerritos@yahoocom) Color versions of one or more of the figures in this paper are available online at Digital Object Identifier /TNB Definition 2-2: Given set a bit the biomolecular operation Append-Tail appends onto the end of every element in set The formal representation is written as Definition 2-3: Given set the biomolecular operation sets to be an empty set can be represented as Definition 2-4: Given set the biomolecular operation creates a number of identical copies of set then Definition 2-5: Given set a bit if the value of is equal to one then the biomolecular extract operation creates two new sets Otherwise it produces another two new sets Definition 2-6: Given sets the biomolecular merge operation Definition 2-7: Given set the biomolecular operation returns true if Otherwise it returns false Definition 2-8: Given set the biomolecular operation describes any element in Even if contains many different elements the biomolecular operation can give an explicit description of exactly one of them III THE DNA-BASED ALGORITHMS OF IMPLEMENTING ARITHMETICAL OPERATIONS OF COMPLEX VECTORS The following subsections are applied to show how arithmetical operations of complex vectors are implemented by the proposed DNA-based algorithms that are made of biological operations DNA sequences A The DNA-Based Algorithms of Implementing Addition of Complex Vectors Closure Axioms of Addition It is assumed that a nonempty set is equal to It is supposed that the values of in can be subsequently represented as two signed binary number for represent positive sign IEEE Personal use is permitted but republication/redistribution requires IEEE permission See for more information
2 908 IEEE TRANSACTIONS ON NANOBIOSCIENCE VOL 14 NO 8 DECEMBER 2015 represent negative sign It is assumed for that are their integer are their fraction From [1] [2] for every bit twodistinct DNA sequences are designed to encode them It is assumed that denote their value to be 1 defines their value to be 0 defines their value to be 0 or 1 It is supposed for that two binary numbers represent respectively the sumoftherealpart theimaginary part to the th element in that are any element in represent positive sign represent negative sign It is assumed that denotes their value to be 1 defines their value to be 0 The following DNA-based algorithm Algorithm 3-1 is used to implement addition of complex vectors closure axioms of addition In Algorithm 3-1 the initial state of each tube is set to an empty tube Algorithm 3-1: Implement addition of complex vectors closure axioms of addition (2) (3) For down to 1 (3a) (3b) (3c) (3d) (3e) (3f) (3g) (3h) (3i) (3j) (4) For to (5) For to (5a) (5b) (5c) (5d) (5e) (5f) (5g) If (5h) (5i) (5j) ElseIf (5k) (5l) (5m) ElseIf (5n) (5o) (5p) ElseIf (5q) (5r) (6) If (6a) EndAlgorithm Theorem 3-1: Algorithm 3-1 can be used to implement addition of complex vectors closure axioms of addition Proof: After the first execution of Step Step (2) is implemented tube contains DNA sequences encoding all of the elements in tube contains DNA sequences encoding the real part the imaginary part of Next after the th execution of Step (3a) is implemented two opers in are positive real values two opers in are negative real values in the absolute value of the first positive oper is greater than or equal to the absolute value of the second negative oper in the absolute value of the first positive oper is less than the absolute value of the second negative oper in the absolute value of the first negative oper is greater than or equal to the absolute value of the second negative oper in the absolute value of the first negative oper is less than the absolute value of the second positive oper Next after the th execution of Step (3b) through Step (3d) is implemented the required computation for their th elements in the imaginary part of in is completed Next the th execution of Step (3e) pours their contents into Next similar computation to the real part of is completed by means of implementing the th execution of Step (3f) through Step (3j) Next each execution of Step (5a) through Step (5f) yields different tubes with different DNA strs Next each execution of Step (5g) through Step (5r) removes illegal DNA strs reserves legal DNA strs Finally DNA sequences in give the answer of satisfying closure axioms Next the execution of Step (6) Step (6a) completes the process of reading the answer(s) Therefore it is inferred that Algorithm 3-1 can be applied to implement addition of complex vectors closure axioms of addition B Constructing Molecular Solutions to Domain Range of Closure Axioms of Addition isusedto construct molecular solutions of a nonempty set denoted in Section III-A The notations used in are denoted in Section III-A The first parameter is an empty tube For to (1a) (1b) (1c) (2) For down to 1 (2a) (2b) (2c) (2d) (3) For down to 1 (3a) (3b) (3c) (3d)
3 CHANG et al: THE DNA-BASED ALGORITHMS OF IMPLEMENTING ARITHMETICAL OPERATIONS OF COMPLEX VECTORS 909 (4) For to (4a) End Lemma 3-1: The DNA-based algorithm can be applied to construct molecular solutions to domain range of closure axioms of addition Proof: Please refer to the proof of Theorem 3-1 C Constructing Molecular Solutions to Any Two Elements With p-tuples of Complex Numbers It is supposed that where are real numbers for It is supposed that the value of in in can be subsequently represented as two signed binary numbers for represent positive sign represent negative sign It is assumed for that are their integer are their fraction It is also supposed that the value of in in can be also represented as two signed binary numbers for represent positive sign represent negative sign It is assumed for that are their integer are their fraction To it is assumed that denote their values to be one zero The isusedto following DNA-based algorithm construct molecular solutions of The first parameter is an empty tube For to 1 (2) For to (2a) (3) For to (3a) (4) For to 1 (5) For to (5a) (6) For to (6a) End Lemma 3-2: The DNA-based algorithm can be applied to construct molecular solutions of Proof: Please refer to the proof of Theorem 3-1 D Constructing Parallel Comparators of Complex Numbers The following first DNA-based algorithm is proposed to complete the function of a parallel comparator of bits for the imaginary part of complex numbers the following second DNA-based algorithm isalsopresented to complete the function of a parallel comparator of bits for the real part of complex numbers When the two DNA-based algorithms are called by Algorithm 3-1 molecular solutions of are in the first parameter thesecond parameter through the seventh parameter are all empty tubes the values of the eighth parameter the ninth parameter are the value of the index variable of the first single loop in Algorithm 3-1 Notations used in them are denoted in Section III-C End Lemma 3-3: The DNA-based algorithm can be applied to complete the function of a parallel comparator of the imaginary part of complex numbers Proof: Please refer to the proof of Theorem 3-1 End bits for Lemma 3-4: The DNA-based algorithm can be employed to complete the function of a parallel comparator of bits for the real part of complex numbers Proof: Please refer to the proof of Theorem 3-1 E Constructing a Parallel Comparator of Bits for the Real Part the Imaginary Part of Complex Numbers is proposed to complete the function of a parallel comparator of bits for the real part the imaginary part of complex numbers The content from the first parameter to the ninth parameter is the same as that of nine arguments in two callers If it is called for dealing with the imaginary part then the tenth eleventh parameters are replaced by Otherwisetheyare replaced by (2) (3) (4) For to 1
4 910 IEEE TRANSACTIONS ON NANOBIOSCIENCE VOL 14 NO 8 DECEMBER 2015 (4a) (4b) If ( " " ) then (4c) Terminate the execution of the loop (5) (6) End Lemma 3-5: The DNA-based algorithm can be applied to complete the function of a parallel comparator of bits for the real part the imaginary part of molecular solutions of Proof: Please refer to the proof of Theorem 3-1 F Constructing a Parallel Comparator of One Bit for the Real Part the Imaginary Part of Complex Numbers ispresented to finish the function of a parallel comparator of one bit The first parameter includes DNA sequences that have the second parameter contains DNA sequences that have thethird parameter through the sixth parameter are all empty tubes the value of the seventh parameter is the value of the index variable of the first single loop in Algorithm 3-1 the value of the eighth parameter is the value of the index variable of the only single loop in the caller If it is called for dealing with the imaginary part then the ninth tenth parameters are replaced by Otherwise they are replaced by (2) (3) (4) (5) (6) (7) (8) (9) (10) (11) (12) End Lemma 3-6: The DNA-based algorithm can be employed to complete the function of a parallel comparator of one bit Proof: Please refer to the proof of Theorem 3-1 G Constructing Parallel Adders of Complex Numbers For the imaginary part real part of complex numbers are offered to complete the function of a parallel adder of bits DNA sequences in encode two positive opers DNA sequences in encode two negative opers the value of is the value of the index variable of the first single loop in Algorithm 3-1 The notations used in the following two DNA-based algorithms are denoted in the previous subsections End Lemma 3-7: The DNA-based algorithm can be used to complete the function of a parallel adder of bits for the imaginary part of complex numbers Proof: Please refer to the proof of Theorem 3-1 End Lemma 3-8: The DNA-based algorithm can be applied to complete the function of a parallel adder of bits for the real part of complex numbers Proof: Please refer to the proof of Theorem 3-1 H Constructing a Parallel Adder of Bits for the Real Part the Imaginary Part of Complex Numbers is offered to complete the function of a parallel adder of bits for the real part the imaginary part of complex numbers The front three parameters are the same as the two callers If it is called by then the fourth fifth sixth parameters are subsequently replaced by Otherwisethey are subsequently replaced by An adder of onebitcanbeappliedtofigureoutthesumthecarryoftwo input bits a previous carry An adder for two opers with bits can be completed by means of the adder of one bit (0) If (2) (3) For to (3a) (4) (5) (5a) If (6) (7) (8) For to (8a) (9)
5 CHANG et al: THE DNA-BASED ALGORITHMS OF IMPLEMENTING ARITHMETICAL OPERATIONS OF COMPLEX VECTORS 911 (10) End Lemma 3-9: The DNA-based algorithm can be applied to complete the function of a parallel adder of bits for the real part the imaginary part of complex numbers Proof: Please refer to the proof of Theorem 3-1 I Constructing a Parallel Adder of One Bit for the Real Part the Imaginary Part of Complex Numbers isproposedto complete the function of a parallel adder of one bits for the real part the imaginary part of complex numbers If it is called by Step (3a) in then the first parameter contains DNA sequences encoding positive opers If it is called by Step (8a) in then the first parameter contains DNA sequences encoding negative opers The value of the second parameter isthe value of the index variable of the first single loop in Algorithm 3-1 the value of the third parameter is the value of the index variable of the single loop in Step (3) or Step (8) in the caller The last three parameters are the same as the last three arguments in the caller In an adder of one bit it is assumed for that represents the first input represents the first output represents the second input represents the third input represents the second output It is assumed that for are used to represent their values to be one are used to represent their values to be zero (2) (3) (4) (5) (6) (7) (8) (9) (10) (11) (12) (13) (14) (15) (16) End Lemma 3-10: The DNA-based algorithm can be applied to complete the function of a parallel adder of one bit Proof: Please refer to the proof of Theorem 3-1 J Constructing Parallel Subtractors for the Absolute Value of the First Oper Greater Than or Equal to the Absolute Value of the Second Oper in Complex Numbers For the absolute value of the first oper greater than or equal to the absolute value of the second oper in the imaginary real parts of complex numbers are proposed to complete the function of a parallel subtractor of bits DNA sequences in encode the first positive oper the second negative oper in which the absolute value of the first positive oper is greater than or equal to the absolute value of the second negative oper DNA sequences in encode the first negative oper the second positive oper in which the absolute value of the first negative oper is greater than or equal to the absolute value of the second positive oper The value of the third parameter is the value of the index variable of the first single loop in Algorithm 3-1 The notations used in the following two DNA-based algorithms are denoted in the previous subsections End Lemma 3-11: The DNA-based algorithm can be employed to complete the function of a parallel subtractor of bits for the absolute value of the first oper greater than or equal to the absolute value of the second oper in the imaginary part of complex numbers Proof: Please refer to the proof of Theorem 3-1 End Lemma 3-12: The DNA-based algorithm can be used to complete the function of a parallel subtractor of bits for the absolute value of the first oper greater than or equal to the absolute value of the second oper in the real part of complex numbers Proof: Please refer to the proof of Theorem 3-1 K Constructing Parallel Subtractors of Bits for the Absolute Value of the First Oper Greater Than or Equal to the Absolute Value of the Second Oper in the Real Part the Imaginary Part of Complex Numbers is
6 912 IEEE TRANSACTIONS ON NANOBIOSCIENCE VOL 14 NO 8 DECEMBER 2015 proposed to complete the function of a parallel subtractor of bits for that the absolute value of the first oper is greater than or is equal to the absolute value of the second oper in the real part the imaginary part of complex numbers The front three parameters are the same as the front three arguments of the two callers If it is called by then the fourth fifth sixth parameters are subsequently replaced by Otherwise they are subsequently replaced by A subtractor of one bit can be used to compute the difference bit the borrow bit for two input bits a previous borrow A subtractor for two opers with bits can be completed by means of the subtractor of one bit If (2) (3) (4) For to (4a) (5) (6) (7) If (8) (9) (10) For to (10a) (11) (12) End Lemma 3-13: The DNA-based algorithm can be employed to complete the function of parallel subtractors of bits for that the absolute value of the first oper is greater than or equal to the absolute value of the second oper in the real part the imaginary part of complex numbers Proof: Please refer to the proof of Theorem 3-1 L Constructing Parallel Subtractors of One Bits for the Absolute Value of the First Oper Greater Than or Equal to the Absolute Value of the Second Oper in the Real Part the Imaginary Part of Complex Numbers For the real part the imaginary part of complex numbers the following DNA-based algorithm is offered to complete the function of a parallel subtractor of one bit If it is called by Step (4a) in thenthefirst parameter consists of DNA sequences encoding the first positive oper the second negative oper in which the absolute value of the first positive oper is greater than or equal to the absolute value of the second negative oper If it is called by Step (10a) in then the first parameter contains DNA sequences encoding the first negative oper the second positive oper in which the absolute value of the first negative oper is greater than or equal to the absolute value of the second positive oper The value of the second parameter is the value of the index variable of the first single loop in Algorithm 3-1 the value of the third parameter isthe value of the index variable of the single loop in Step (4) or Step (10) in the caller The last three parameters are the same as the last three arguments of the caller In a subtractor of one bit it is supposed that for represents the first input represents the first output represents the second input represents the third input represents the second output Distinct DNA sequences are designed to encode the value 0 or 1 for for Itisassumedthatfor are applied to represent their values to be one are used to represent their values to be zero (2) (3) (4) (5) (6) (7) (8) (9) (10) (11) (12) (13) (14) (15) (16) End Lemma 3-14: The DNA-based algorithm canbe applied to complete the function of a parallel subtractor with one bit for that the absolute value of the first oper is greater than or equal to the absolute value of the second oper Proof: Please refer to the proof of Theorem 3-1
7 CHANG et al: THE DNA-BASED ALGORITHMS OF IMPLEMENTING ARITHMETICAL OPERATIONS OF COMPLEX VECTORS 913 M Constructing Parallel Subtractors for the Absolute Value of the First Oper Less Than the Absolute Value of the Second Oper in Complex Numbers In the imaginary real parts of complex numbers are proposed to complete the function of a parallel subtractor of bits for that the absolute value of the first oper is less than the absolute value of the second oper When the two DNA-based algorithms are called by Algorithm 3-1 DNA sequences in encode the first positive oper the second negative oper in which the absolute value of the first positive oper is less than the absolute value of the second negative oper DNA sequences in encode the first negative oper the second positive oper in which the absolute value of the first negative oper is less than the absolute value of the second positive oper The value of the third parameter is the value of the index variable of the first single loop in Algorithm 3-1 The notations used in the following two DNA-based algorithms are denoted in the previous subsections End Lemma 3-15: The DNA-based algorithm can be used to complete the function of a parallel subtractor of bits for that the absolute value of the first oper is less than the absolute value of the second oper in the imaginary part of complex numbers Proof: Please refer to the proof of Theorem 3-1 End Lemma 3-16: The DNA-based algorithm can be applied to complete the function of a parallel subtractor of bits for that the absolute value of the first oper is less than the absolute value of the second oper in the real part of complex numbers Proof: Please refer to the proof of Theorem 3-1 N Constructing Parallel Subtractors of Bits for the Absolute Value of the First Oper Less Than the Absolute Value of the Second Oper in the Real Part the Imaginary Part of Complex Numbers is presented to complete the function of a parallel subtractor of bits for that the absolute value of the first oper is less than the absolute value of the second oper in the real part the imaginary part of complex numbers DNA sequences in encode the first positive oper the second negative oper in which the absolute value of the first positive oper is less than the absolute value of the second negative oper DNA sequences in encode the first negative oper the second positive oper in which the absolute value of the first negative oper is less than the absolute value of the second positive oper The value of the third parameter is the value of the index variable of the first single loop in Algorithm 3-1 Thefourth fifth sixth parameters are subsequently replaced by if it is called by Otherwise they are subsequently replaced by A subtractor of one bit can be employed to figure out the difference bit the borrow bit for two input bits a previous borrow A subtractor for two opers with bits can be completed by means of the subtractor of one bit If '' then (2) (3) (4) For to (4a) (5) (6) (7) If '' then (8) (9) (10) For to (10a) (11) (12) End Lemma 3-17: The DNA-based algorithm can be applied to complete the function of parallel subtractors of bits for that the absolute value of the first oper is less than the absolute value of the second oper Proof: Please refer to the proof of Theorem 3-1 O Constructing Parallel Subtractors of One Bits for the Absolute Value of the First Oper Less Than the Absolute Value of the Second Oper in the Real Part the Imaginary Part of Complex Numbers is presented to complete the function of a parallel subtractor of one bit for that the absolute value of the first oper is less than the absolute value of the second oper in the real part the imaginary part of complex numbers If it is called by Step (4a) in then the first parameter includes DNA sequences encoding the first positive oper the second negative oper in which the absolute value of the first positive oper is less than the absolute value of the second negative oper If it is called by Step (10a) in then the first parameter consists of DNA sequences
8 914 IEEE TRANSACTIONS ON NANOBIOSCIENCE VOL 14 NO 8 DECEMBER 2015 encoding the first negative oper the second positive oper in which the absolute value of the first negative oper is less than the absolute value of the second positive oper The value of the second parameter is the value of the index variable of the first single loop in Algorithm 3-1 the value of the third parameter is the value of the index variable of the single loop in Step (4) or Step (10) in The last three parameters are the same as the last three arguments in the caller Because the absolute value of the first oper is lessthanthe absolute value of the second oper in a subtractor of one bit it is supposed for that represents the first input represents the second input represents the third input represents the first output represents the second output (2) (3) (4) (5) (6) (7) (8) (9) (10) (11) (12) (13) (14) (15) (16) End Lemma 3-18: The DNA-based algorithm can be applied to complete the function of a parallel subtractor of one bit for that the absolute value of the first oper is less than the absolute value of the second oper Proof: Please refer to the proof of Theorem 3-1 IV COMPLEXITY ASSESSMENT Theorem 4-1: For implementing addition closure axioms of addition to complex vectors with -tuples of complex numbers with that the values of each imaginary real parts are encoded as signed binary numbers of bits each bit is encoded by basepairs itstimecomplexityis biological operations its volume complexity is DNA strs its tube complexity is O tubes its longest DNA str is base pairs Proof: The DNA-based algorithm instepof Algorithm 3-1 is implemented by means of biological operations Next the DNA-based algorithm in Step (2) of Algorithm 3-1 is implemented by means of biological operations The four DNA-based algorithms from Step (3a) through Step (3d) of Algorithm 3-1 are implemented by means of biological operations Next Step (3e) of Algorithm 3-1 is implemented by means of O biological operations Similarly the four DNA-based algorithms from Step (3f) through Step (3i) of Algorithm 3-1 are implemented by means of biological operations Next Step (3j) of Algorithm 3-1 is implemented by means of O biological operations Each Step from Step (3a) through Step (3j) in Algorithm 3-1 is implemented times Therefore the total number of biological operations for implementing them is Next the total number of biological operations for implementing Step (5a) through Step (5r) in Algorithm 3-1 is FinallyStep(6)Step(6a) of Algorithm 3-1 are implemented by means of O biological operations Similarly proof can be used to show complexity of volume tube the longest DNA str Therefore it is inferred that its time complexity is biological operations its volume complexity is DNA strs its tube complexity is O tubes its longest DNA str is base pairs V CONCLUSIONS With current biotechnology the time for each operation is at least one second Realistically steps like gel electrophoresis take much longer but for the sake of argument say each biological operation takes one second From Theorem 4-1 if the values of are equal to then we need to take at least which are about REFERENCES [1] M Amos Theoretical Experimental DNA Computation New York: Springer 2005 [2] W-L Chang A V Vasilakos Molecular Computing: Towards a Novel Computing Architecture for Complex Problem Solving New York: Springer 2014 [3] W-L Chang A V Vasilakos Molecular algorithms of implementing bio-molecular databases on a biological computer IEEE Trans NanoBiosci vol 14 no 1 pp Jan 2015 [4] W-L Chang T-T Ren M Feng Quantum algorithms mathematical formulations of bio-molecular solutions of the vertex cover problem in the finite-dimensional Hilbert space IEEE Trans NanoBiosci vol 14 no 1 pp Jan 2015
GENERALIZED ARYABHATA REMAINDER THEOREM
 International Journal of Innovative Computing, Information and Control ICIC International c 2010 ISSN 1349-4198 Volume 6, Number 4, April 2010 pp. 1865 1871 GENERALIZED ARYABHATA REMAINDER THEOREM Chin-Chen
International Journal of Innovative Computing, Information and Control ICIC International c 2010 ISSN 1349-4198 Volume 6, Number 4, April 2010 pp. 1865 1871 GENERALIZED ARYABHATA REMAINDER THEOREM Chin-Chen
Chapter 2 Basic Arithmetic Circuits
 Chapter 2 Basic Arithmetic Circuits This chapter is devoted to the description of simple circuits for the implementation of some of the arithmetic operations presented in Chap. 1. Specifically, the design
Chapter 2 Basic Arithmetic Circuits This chapter is devoted to the description of simple circuits for the implementation of some of the arithmetic operations presented in Chap. 1. Specifically, the design
About the relationship between formal logic and complexity classes
 About the relationship between formal logic and complexity classes Working paper Comments welcome; my email: armandobcm@yahoo.com Armando B. Matos October 20, 2013 1 Introduction We analyze a particular
About the relationship between formal logic and complexity classes Working paper Comments welcome; my email: armandobcm@yahoo.com Armando B. Matos October 20, 2013 1 Introduction We analyze a particular
JANUARY MONDAY TUESDAY WEDNESDAY THURSDAY FRIDAY SATURDAY SUNDAY
 Vocabulary (01) The Calendar (012) In context: Look at the calendar. Then, answer the questions. JANUARY MONDAY TUESDAY WEDNESDAY THURSDAY FRIDAY SATURDAY SUNDAY 1 New 2 3 4 5 6 Year s Day 7 8 9 10 11
Vocabulary (01) The Calendar (012) In context: Look at the calendar. Then, answer the questions. JANUARY MONDAY TUESDAY WEDNESDAY THURSDAY FRIDAY SATURDAY SUNDAY 1 New 2 3 4 5 6 Year s Day 7 8 9 10 11
Structures of the Molecular Components in DNA and RNA with Bond Lengths Interpreted as Sums of Atomic Covalent Radii
 The Open Structural Biology Journal, 2008, 2, 1-7 1 Structures of the Molecular Components in DNA and RNA with Bond Lengths Interpreted as Sums of Atomic Covalent Radii Raji Heyrovska * Institute of Biophysics
The Open Structural Biology Journal, 2008, 2, 1-7 1 Structures of the Molecular Components in DNA and RNA with Bond Lengths Interpreted as Sums of Atomic Covalent Radii Raji Heyrovska * Institute of Biophysics
KEYWORDS: Multiple Valued Logic (MVL), Residue Number System (RNS), Quinary Logic (Q uin), Quinary Full Adder, QFA, Quinary Half Adder, QHA.
 GLOBAL JOURNAL OF ADVANCED ENGINEERING TECHNOLOGIES AND SCIENCES DESIGN OF A QUINARY TO RESIDUE NUMBER SYSTEM CONVERTER USING MULTI-LEVELS OF CONVERSION Hassan Amin Osseily Electrical and Electronics Department,
GLOBAL JOURNAL OF ADVANCED ENGINEERING TECHNOLOGIES AND SCIENCES DESIGN OF A QUINARY TO RESIDUE NUMBER SYSTEM CONVERTER USING MULTI-LEVELS OF CONVERSION Hassan Amin Osseily Electrical and Electronics Department,
Combinational Logic. By : Ali Mustafa
 Combinational Logic By : Ali Mustafa Contents Adder Subtractor Multiplier Comparator Decoder Encoder Multiplexer How to Analyze any combinational circuit like this? Analysis Procedure To obtain the output
Combinational Logic By : Ali Mustafa Contents Adder Subtractor Multiplier Comparator Decoder Encoder Multiplexer How to Analyze any combinational circuit like this? Analysis Procedure To obtain the output
BIOCHEMISTRY GUIDED NOTES - AP BIOLOGY-
 BIOCHEMISTRY GUIDED NOTES - AP BIOLOGY- ELEMENTS AND COMPOUNDS - anything that has mass and takes up space. - cannot be broken down to other substances. - substance containing two or more different elements
BIOCHEMISTRY GUIDED NOTES - AP BIOLOGY- ELEMENTS AND COMPOUNDS - anything that has mass and takes up space. - cannot be broken down to other substances. - substance containing two or more different elements
Linear Codes and Syndrome Decoding
 Linear Codes and Syndrome Decoding These notes are intended to be used as supplementary reading to Sections 6.7 9 of Grimaldi s Discrete and Combinatorial Mathematics. The proofs of the theorems are left
Linear Codes and Syndrome Decoding These notes are intended to be used as supplementary reading to Sections 6.7 9 of Grimaldi s Discrete and Combinatorial Mathematics. The proofs of the theorems are left
WAITING-TIME DISTRIBUTION FOR THE r th OCCURRENCE OF A COMPOUND PATTERN IN HIGHER-ORDER MARKOVIAN SEQUENCES
 WAITING-TIME DISTRIBUTION FOR THE r th OCCURRENCE OF A COMPOUND PATTERN IN HIGHER-ORDER MARKOVIAN SEQUENCES Donald E. K. Martin 1 and John A. D. Aston 2 1 Mathematics Department, Howard University, Washington,
WAITING-TIME DISTRIBUTION FOR THE r th OCCURRENCE OF A COMPOUND PATTERN IN HIGHER-ORDER MARKOVIAN SEQUENCES Donald E. K. Martin 1 and John A. D. Aston 2 1 Mathematics Department, Howard University, Washington,
Name: Date: Period: Biology Notes: Biochemistry Directions: Fill this out as we cover the following topics in class
 Name: Date: Period: Biology Notes: Biochemistry Directions: Fill this out as we cover the following topics in class Part I. Water Water Basics Polar: part of a molecule is slightly, while another part
Name: Date: Period: Biology Notes: Biochemistry Directions: Fill this out as we cover the following topics in class Part I. Water Water Basics Polar: part of a molecule is slightly, while another part
Part I: Definitions and Properties
 Turing Machines Part I: Definitions and Properties Finite State Automata Deterministic Automata (DFSA) M = {Q, Σ, δ, q 0, F} -- Σ = Symbols -- Q = States -- q 0 = Initial State -- F = Accepting States
Turing Machines Part I: Definitions and Properties Finite State Automata Deterministic Automata (DFSA) M = {Q, Σ, δ, q 0, F} -- Σ = Symbols -- Q = States -- q 0 = Initial State -- F = Accepting States
arxiv: v2 [cs.ds] 17 Sep 2017
![arxiv: v2 [cs.ds] 17 Sep 2017 arxiv: v2 [cs.ds] 17 Sep 2017](/thumbs/90/103295691.jpg) Two-Dimensional Indirect Binary Search for the Positive One-In-Three Satisfiability Problem arxiv:1708.08377v [cs.ds] 17 Sep 017 Shunichi Matsubara Aoyama Gakuin University, 5-10-1, Fuchinobe, Chuo-ku,
Two-Dimensional Indirect Binary Search for the Positive One-In-Three Satisfiability Problem arxiv:1708.08377v [cs.ds] 17 Sep 017 Shunichi Matsubara Aoyama Gakuin University, 5-10-1, Fuchinobe, Chuo-ku,
CHAPTER 12 Boolean Algebra
 318 Chapter 12 Boolean Algebra CHAPTER 12 Boolean Algebra SECTION 12.1 Boolean Functions 2. a) Since x 1 = x, the only solution is x = 0. b) Since 0 + 0 = 0 and 1 + 1 = 1, the only solution is x = 0. c)
318 Chapter 12 Boolean Algebra CHAPTER 12 Boolean Algebra SECTION 12.1 Boolean Functions 2. a) Since x 1 = x, the only solution is x = 0. b) Since 0 + 0 = 0 and 1 + 1 = 1, the only solution is x = 0. c)
UNIVERSITY OF VICTORIA DECEMBER EXAMINATIONS MATH 122: Logic and Foundations
 UNIVERSITY OF VICTORIA DECEMBER EXAMINATIONS 2013 MATH 122: Logic and Foundations Instructor and section (check one): K. Mynhardt [A01] CRN 12132 G. MacGillivray [A02] CRN 12133 NAME: V00#: Duration: 3
UNIVERSITY OF VICTORIA DECEMBER EXAMINATIONS 2013 MATH 122: Logic and Foundations Instructor and section (check one): K. Mynhardt [A01] CRN 12132 G. MacGillivray [A02] CRN 12133 NAME: V00#: Duration: 3
This is a listing of common symbols found within all branches of mathematics 1. x = y means x and y represent the same thing or value.
 This is a listing of common symbols found within all branches of mathematics 1. Symbol Read as Explanation Examples = is equal to; equals < > + is not equal to does not equal is less than, is greater than
This is a listing of common symbols found within all branches of mathematics 1. Symbol Read as Explanation Examples = is equal to; equals < > + is not equal to does not equal is less than, is greater than
Chapters 12&13 Notes: DNA, RNA & Protein Synthesis
 Chapters 12&13 Notes: DNA, RNA & Protein Synthesis Name Period Words to Know: nucleotides, DNA, complementary base pairing, replication, genes, proteins, mrna, rrna, trna, transcription, translation, codon,
Chapters 12&13 Notes: DNA, RNA & Protein Synthesis Name Period Words to Know: nucleotides, DNA, complementary base pairing, replication, genes, proteins, mrna, rrna, trna, transcription, translation, codon,
Writing and Comparing Numbers Through Hundred Thousands Ordinal Numbers
 LESSON 7 Writing and Comparing Numbers Through Hundred Thousands Ordinal Numbers Power Up facts Power Up A count aloud Count up and down by 20s between 0 and 200. Count up and down by 200s between 0 and
LESSON 7 Writing and Comparing Numbers Through Hundred Thousands Ordinal Numbers Power Up facts Power Up A count aloud Count up and down by 20s between 0 and 200. Count up and down by 200s between 0 and
This article appeared in a journal published by Elsevier. The attached copy is furnished to the author for internal non-commercial research and
 This article appeared in a journal published by Elsevier. The attached copy is furnished to the author for internal non-commercial research and education use, including for instruction at the authors institution
This article appeared in a journal published by Elsevier. The attached copy is furnished to the author for internal non-commercial research and education use, including for instruction at the authors institution
10-2 Arithmetic Sequences and Series
 Determine the common difference, and find the next four terms of each arithmetic sequence. 1. 20, 17, 14, 17 20 = 3 14 17 = 3 The common difference is 3. Add 3 to the third term to find the fourth term,
Determine the common difference, and find the next four terms of each arithmetic sequence. 1. 20, 17, 14, 17 20 = 3 14 17 = 3 The common difference is 3. Add 3 to the third term to find the fourth term,
Introduction to Languages and Computation
 Introduction to Languages and Computation George Voutsadakis 1 1 Mathematics and Computer Science Lake Superior State University LSSU Math 400 George Voutsadakis (LSSU) Languages and Computation July 2014
Introduction to Languages and Computation George Voutsadakis 1 1 Mathematics and Computer Science Lake Superior State University LSSU Math 400 George Voutsadakis (LSSU) Languages and Computation July 2014
IV. Turing Machine. Yuxi Fu. BASICS, Shanghai Jiao Tong University
 IV. Turing Machine Yuxi Fu BASICS, Shanghai Jiao Tong University Alan Turing Alan Turing (23Jun.1912-7Jun.1954), an English student of Church, introduced a machine model for effective calculation in On
IV. Turing Machine Yuxi Fu BASICS, Shanghai Jiao Tong University Alan Turing Alan Turing (23Jun.1912-7Jun.1954), an English student of Church, introduced a machine model for effective calculation in On
Comments and Corrections
 1386 IEEE TRANSACTIONS ON AUTOMATIC CONTROL, VOL 59, NO 5, MAY 2014 Comments and Corrections Corrections to Stochastic Barbalat s Lemma and Its Applications Xin Yu and Zhaojing Wu Abstract The proof of
1386 IEEE TRANSACTIONS ON AUTOMATIC CONTROL, VOL 59, NO 5, MAY 2014 Comments and Corrections Corrections to Stochastic Barbalat s Lemma and Its Applications Xin Yu and Zhaojing Wu Abstract The proof of
The body has three primary lines of defense against changes in hydrogen ion concentration in the body fluids.
 ph and Nucleic acids Hydrogen Ion (H+) concentration is precisely regulated. The H+ concentration in the extracellular fluid is maintained at a very low level, averaging 0.00000004Eq/L. normal variations
ph and Nucleic acids Hydrogen Ion (H+) concentration is precisely regulated. The H+ concentration in the extracellular fluid is maintained at a very low level, averaging 0.00000004Eq/L. normal variations
The L Machines are very high-level, in two senses:
 What is a Computer? State of the machine. CMPSCI 630: Programming Languages An Abstract Machine for Control Spring 2009 (with thanks to Robert Harper) Internal registers, memory, etc. Initial and final
What is a Computer? State of the machine. CMPSCI 630: Programming Languages An Abstract Machine for Control Spring 2009 (with thanks to Robert Harper) Internal registers, memory, etc. Initial and final
ONTARIO REGULATION 156/06. made under the CONSERVATION AUTHORITIES ACT
 ONTARIO REGULATION 156/06 made under the CONSERVATION AUTHORITIES ACT Made: April 28, 2006 Approved: May 4, 2006 Filed: May 4, 2006 Published on e-laws: May 8, 2006 Printed in The Ontario Gazette: May
ONTARIO REGULATION 156/06 made under the CONSERVATION AUTHORITIES ACT Made: April 28, 2006 Approved: May 4, 2006 Filed: May 4, 2006 Published on e-laws: May 8, 2006 Printed in The Ontario Gazette: May
The natural numbers. The natural numbers come with an addition +, a multiplication and an order < p < q, q < p, p = q.
 The natural numbers N = {0, 1,, 3, } The natural numbers come with an addition +, a multiplication and an order < p, q N, p + q N. p, q N, p q N. p, q N, exactly one of the following holds: p < q, q
The natural numbers N = {0, 1,, 3, } The natural numbers come with an addition +, a multiplication and an order < p, q N, p + q N. p, q N, p q N. p, q N, exactly one of the following holds: p < q, q
Chapter 3. Regular grammars
 Chapter 3 Regular grammars 59 3.1 Introduction Other view of the concept of language: not the formalization of the notion of effective procedure, but set of words satisfying a given set of rules Origin
Chapter 3 Regular grammars 59 3.1 Introduction Other view of the concept of language: not the formalization of the notion of effective procedure, but set of words satisfying a given set of rules Origin
Name: Date: Hour: Unit Four: Cell Cycle, Mitosis and Meiosis. Monomer Polymer Example Drawing Function in a cell DNA
 Unit Four: Cell Cycle, Mitosis and Meiosis I. Concept Review A. Why is carbon often called the building block of life? B. List the four major macromolecules. C. Complete the chart below. Monomer Polymer
Unit Four: Cell Cycle, Mitosis and Meiosis I. Concept Review A. Why is carbon often called the building block of life? B. List the four major macromolecules. C. Complete the chart below. Monomer Polymer
COMBINATIONAL LOGIC FUNCTIONS
 COMBINATIONAL LOGIC FUNCTIONS Digital logic circuits can be classified as either combinational or sequential circuits. A combinational circuit is one where the output at any time depends only on the present
COMBINATIONAL LOGIC FUNCTIONS Digital logic circuits can be classified as either combinational or sequential circuits. A combinational circuit is one where the output at any time depends only on the present
The max flow problem. Ford-Fulkerson method. A cut. Lemma Corollary Max Flow Min Cut Theorem. Max Flow Min Cut Theorem
 The max flow problem Ford-Fulkerson method 7 11 Ford-Fulkerson(G) f = 0 while( simple path p from s to t in G f ) 10-2 2 1 f := f + f p output f 4 9 1 2 A cut Lemma 26. + Corollary 26.6 Let f be a flow
The max flow problem Ford-Fulkerson method 7 11 Ford-Fulkerson(G) f = 0 while( simple path p from s to t in G f ) 10-2 2 1 f := f + f p output f 4 9 1 2 A cut Lemma 26. + Corollary 26.6 Let f be a flow
Integer and Vector Multiplication by Using DNA
 Int. J. Open Problems Compt. Math., Vol. 1, No. 2, September 2008 Integer and Vector Multiplication by Using DNA Essam Al-Daoud 1, Belal Zaqaibeh 2, and Feras Al-Hanandeh 3 1 Computer Science Department,
Int. J. Open Problems Compt. Math., Vol. 1, No. 2, September 2008 Integer and Vector Multiplication by Using DNA Essam Al-Daoud 1, Belal Zaqaibeh 2, and Feras Al-Hanandeh 3 1 Computer Science Department,
Classroom Activity to Make Fraction Strips
 Classroom Activity to Make Fraction Strips This resource provides instructions on teaching your students how to make fraction strips out of paper. Fraction strips made from copy paper are an excellent
Classroom Activity to Make Fraction Strips This resource provides instructions on teaching your students how to make fraction strips out of paper. Fraction strips made from copy paper are an excellent
Interpret Astrology The House Combinations. By Michael Erlewine
 Interpret Astrology The House Combinations By Michael Erlewine An e-book from Startypes.com 315 Marion Avenue Big Rapids, Michigan 49307 First published 2006 2006 Michael Erlewine/StarTypes.com ISBN 978-0-9798328-1-9
Interpret Astrology The House Combinations By Michael Erlewine An e-book from Startypes.com 315 Marion Avenue Big Rapids, Michigan 49307 First published 2006 2006 Michael Erlewine/StarTypes.com ISBN 978-0-9798328-1-9
Model Worksheet Teacher Key
 Introduction Despite the complexity of life on Earth, the most important large molecules found in all living things (biomolecules) can be classified into only four main categories: carbohydrates, lipids,
Introduction Despite the complexity of life on Earth, the most important large molecules found in all living things (biomolecules) can be classified into only four main categories: carbohydrates, lipids,
New Minimal Weight Representations for Left-to-Right Window Methods
 New Minimal Weight Representations for Left-to-Right Window Methods James A. Muir 1 and Douglas R. Stinson 2 1 Department of Combinatorics and Optimization 2 School of Computer Science University of Waterloo
New Minimal Weight Representations for Left-to-Right Window Methods James A. Muir 1 and Douglas R. Stinson 2 1 Department of Combinatorics and Optimization 2 School of Computer Science University of Waterloo
Lesson Overview The Structure of DNA
 12.2 THINK ABOUT IT The DNA molecule must somehow specify how to assemble proteins, which are needed to regulate the various functions of each cell. What kind of structure could serve this purpose without
12.2 THINK ABOUT IT The DNA molecule must somehow specify how to assemble proteins, which are needed to regulate the various functions of each cell. What kind of structure could serve this purpose without
Coloring Vertices and Edges of a Path by Nonempty Subsets of a Set
 Coloring Vertices and Edges of a Path by Nonempty Subsets of a Set P.N. Balister E. Győri R.H. Schelp April 28, 28 Abstract A graph G is strongly set colorable if V (G) E(G) can be assigned distinct nonempty
Coloring Vertices and Edges of a Path by Nonempty Subsets of a Set P.N. Balister E. Győri R.H. Schelp April 28, 28 Abstract A graph G is strongly set colorable if V (G) E(G) can be assigned distinct nonempty
Theory of Computing Tamás Herendi
 Theory of Computing Tamás Herendi Theory of Computing Tamás Herendi Publication date 2014 Table of Contents 1 Preface 1 2 Formal languages 2 3 Order of growth rate 9 4 Turing machines 16 1 The definition
Theory of Computing Tamás Herendi Theory of Computing Tamás Herendi Publication date 2014 Table of Contents 1 Preface 1 2 Formal languages 2 3 Order of growth rate 9 4 Turing machines 16 1 The definition
A Simple Left-to-Right Algorithm for Minimal Weight Signed Radix-r Representations
 IEEE TRANSACTIONS ON INFORMATION THEORY, VOL. XX, NO. X, MONTH 2007 1 A Simple Left-to-Right Algorithm for Minimal Weight Signed Radix-r Representations James A. Muir Abstract We present a simple algorithm
IEEE TRANSACTIONS ON INFORMATION THEORY, VOL. XX, NO. X, MONTH 2007 1 A Simple Left-to-Right Algorithm for Minimal Weight Signed Radix-r Representations James A. Muir Abstract We present a simple algorithm
九州工業大学学術機関リポジトリ PROCEDURES FOR LOGIC AND ARITHMETIC OPERATIONS WITH DNA MOLECULES. Author(s) Fujiwara, Akihiro; Matsumoto, Kenic
 九州工業大学学術機関リポジトリ Title PROCEDURES FOR LOGIC AND ARITHMETIC OPERATIONS WITH DNA MOLECULES Author(s) Fujiwara Akihiro; Matsumoto Kenic Issue Date 2004-06 URL http://hdl.handle.net/10228/473 RightsCopyright
九州工業大学学術機関リポジトリ Title PROCEDURES FOR LOGIC AND ARITHMETIC OPERATIONS WITH DNA MOLECULES Author(s) Fujiwara Akihiro; Matsumoto Kenic Issue Date 2004-06 URL http://hdl.handle.net/10228/473 RightsCopyright
Statistical Thermodynamics of DNA Denaturation. processes
 Statistical Thermodynamics of DNA Denaturation processes Course: Statistical Thermodynamics Advisor: Prof. Raissa D Souza Spring quarter 2012 By: Sebastian Frey 1 Nomenclature F Free Energy [J] x Coordinate
Statistical Thermodynamics of DNA Denaturation processes Course: Statistical Thermodynamics Advisor: Prof. Raissa D Souza Spring quarter 2012 By: Sebastian Frey 1 Nomenclature F Free Energy [J] x Coordinate
Distributed Algorithms of Finding the Unique Minimum Distance Dominating Set in Directed Split-Stars
 Distributed Algorithms of Finding the Unique Minimum Distance Dominating Set in Directed Split-Stars Fu Hsing Wang 1 Jou Ming Chang 2 Yue Li Wang 1, 1 Department of Information Management, National Taiwan
Distributed Algorithms of Finding the Unique Minimum Distance Dominating Set in Directed Split-Stars Fu Hsing Wang 1 Jou Ming Chang 2 Yue Li Wang 1, 1 Department of Information Management, National Taiwan
High Speed Time Efficient Reversible ALU Based Logic Gate Structure on Vertex Family
 International Journal of Engineering Research and Development e-issn: 2278-067X, p-issn: 2278-800X, www.ijerd.com Volume 11, Issue 04 (April 2015), PP.72-77 High Speed Time Efficient Reversible ALU Based
International Journal of Engineering Research and Development e-issn: 2278-067X, p-issn: 2278-800X, www.ijerd.com Volume 11, Issue 04 (April 2015), PP.72-77 High Speed Time Efficient Reversible ALU Based
Bio10 Cell and Molecular Lecture Notes SRJC
 Basic Chemistry Atoms Smallest particles that retain properties of an element Made up of subatomic particles: Protons (+) Electrons (-) Neutrons (no charge) Isotopes Atoms of an element with different
Basic Chemistry Atoms Smallest particles that retain properties of an element Made up of subatomic particles: Protons (+) Electrons (-) Neutrons (no charge) Isotopes Atoms of an element with different
Logic and Computer Design Fundamentals. Chapter 5 Arithmetic Functions and Circuits
 Logic and Computer Design Fundamentals Chapter 5 Arithmetic Functions and Circuits Arithmetic functions Operate on binary vectors Use the same subfunction in each bit position Can design functional block
Logic and Computer Design Fundamentals Chapter 5 Arithmetic Functions and Circuits Arithmetic functions Operate on binary vectors Use the same subfunction in each bit position Can design functional block
Lecture 9 : PPAD and the Complexity of Equilibrium Computation. 1 Complexity Class PPAD. 1.1 What does PPAD mean?
 CS 599: Algorithmic Game Theory October 20, 2010 Lecture 9 : PPAD and the Complexity of Equilibrium Computation Prof. Xi Chen Scribes: Cheng Lu and Sasank Vijayan 1 Complexity Class PPAD 1.1 What does
CS 599: Algorithmic Game Theory October 20, 2010 Lecture 9 : PPAD and the Complexity of Equilibrium Computation Prof. Xi Chen Scribes: Cheng Lu and Sasank Vijayan 1 Complexity Class PPAD 1.1 What does
Quantum transport simulations: sensing DNA
 Quantum transport simulations: sensing DNA Hauptseminar SS2016 Frank Schulz Tutor : Ganesh Sivaraman 1 1 Introduction Quantum transport simulations are a very important concept for simulating electronic
Quantum transport simulations: sensing DNA Hauptseminar SS2016 Frank Schulz Tutor : Ganesh Sivaraman 1 1 Introduction Quantum transport simulations are a very important concept for simulating electronic
Cancer Genome Assembly and Alignment Michael Shan-Hui Ho , Kun-Yu Hung , Yu-Shiang Gen , Chaochang Chiu and Tsung-Hua Lu Abstract
 Cancer Genome Assembly and Alignment Michael Shan-Hui Ho 1, Kun-Yu Hung 2, Yu-Shiang Gen 1, Chaochang Chiu 3 and Tsung-Hua Lu 1 1 Department of Electric Engineering, NTPU, New Taipei City, Taiwan, ROC
Cancer Genome Assembly and Alignment Michael Shan-Hui Ho 1, Kun-Yu Hung 2, Yu-Shiang Gen 1, Chaochang Chiu 3 and Tsung-Hua Lu 1 1 Department of Electric Engineering, NTPU, New Taipei City, Taiwan, ROC
A VLSI Algorithm for Modular Multiplication/Division
 A VLSI Algorithm for Modular Multiplication/Division Marcelo E. Kaihara and Naofumi Takagi Department of Information Engineering Nagoya University Nagoya, 464-8603, Japan mkaihara@takagi.nuie.nagoya-u.ac.jp
A VLSI Algorithm for Modular Multiplication/Division Marcelo E. Kaihara and Naofumi Takagi Department of Information Engineering Nagoya University Nagoya, 464-8603, Japan mkaihara@takagi.nuie.nagoya-u.ac.jp
A High-Speed Realization of Chinese Remainder Theorem
 Proceedings of the 2007 WSEAS Int. Conference on Circuits, Systems, Signal and Telecommunications, Gold Coast, Australia, January 17-19, 2007 97 A High-Speed Realization of Chinese Remainder Theorem Shuangching
Proceedings of the 2007 WSEAS Int. Conference on Circuits, Systems, Signal and Telecommunications, Gold Coast, Australia, January 17-19, 2007 97 A High-Speed Realization of Chinese Remainder Theorem Shuangching
A Projection Decoding of a Binary Extremal Self-Dual Code of Length 40
 A Projection Decoding of a Binary Extremal Self-Dual Code of Length 4 arxiv:7.48v [cs.it] 6 Jan 27 Jon-Lark Kim Department of Mathematics Sogang University Seoul, 2-742, South Korea jlkim@sogang.ac.kr
A Projection Decoding of a Binary Extremal Self-Dual Code of Length 4 arxiv:7.48v [cs.it] 6 Jan 27 Jon-Lark Kim Department of Mathematics Sogang University Seoul, 2-742, South Korea jlkim@sogang.ac.kr
Chapter 2: Chemistry. What does chemistry have to do with biology? Vocabulary BIO 105
 Chapter 2: Chemistry What does chemistry have to do with biology? BIO 105 Vocabulary 1. Matter anything that takes up space and has mass Atoms are the smallest units of matter that can participate in chemical
Chapter 2: Chemistry What does chemistry have to do with biology? BIO 105 Vocabulary 1. Matter anything that takes up space and has mass Atoms are the smallest units of matter that can participate in chemical
Looking at a two binary digit sum shows what we need to extend addition to multiple binary digits.
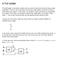 A Full Adder The half-adder is extremely useful until you want to add more that one binary digit quantities. The slow way to develop a two binary digit adders would be to make a truth table and reduce
A Full Adder The half-adder is extremely useful until you want to add more that one binary digit quantities. The slow way to develop a two binary digit adders would be to make a truth table and reduce
Genetic Algorithms. Donald Richards Penn State University
 Genetic Algorithms Donald Richards Penn State University Easy problem: Find the point which maximizes f(x, y) = [16 x(1 x)y(1 y)] 2, x, y [0,1] z (16*x*y*(1-x)*(1-y))**2 0.829 0.663 0.497 0.331 0.166 1
Genetic Algorithms Donald Richards Penn State University Easy problem: Find the point which maximizes f(x, y) = [16 x(1 x)y(1 y)] 2, x, y [0,1] z (16*x*y*(1-x)*(1-y))**2 0.829 0.663 0.497 0.331 0.166 1
Sequences and Summations. CS/APMA 202 Rosen section 3.2 Aaron Bloomfield
 Sequences and Summations CS/APMA 202 Rosen section 3.2 Aaron Bloomfield 1 Definitions Sequence: an ordered list of elements Like a set, but: Elements can be duplicated Elements are ordered 2 Sequences
Sequences and Summations CS/APMA 202 Rosen section 3.2 Aaron Bloomfield 1 Definitions Sequence: an ordered list of elements Like a set, but: Elements can be duplicated Elements are ordered 2 Sequences
( c ) E p s t e i n, C a r t e r a n d B o l l i n g e r C h a p t e r 1 7 : I n f o r m a t i o n S c i e n c e P a g e 1
 ( c ) E p s t e i n, C a r t e r a n d B o l l i n g e r 2 0 1 6 C h a p t e r 1 7 : I n f o r m a t i o n S c i e n c e P a g e 1 CHAPTER 17: Information Science In this chapter, we learn how data can
( c ) E p s t e i n, C a r t e r a n d B o l l i n g e r 2 0 1 6 C h a p t e r 1 7 : I n f o r m a t i o n S c i e n c e P a g e 1 CHAPTER 17: Information Science In this chapter, we learn how data can
K 4 -free graphs with no odd holes
 K 4 -free graphs with no odd holes Maria Chudnovsky 1 Columbia University, New York NY 10027 Neil Robertson 2 Ohio State University, Columbus, Ohio 43210 Paul Seymour 3 Princeton University, Princeton
K 4 -free graphs with no odd holes Maria Chudnovsky 1 Columbia University, New York NY 10027 Neil Robertson 2 Ohio State University, Columbus, Ohio 43210 Paul Seymour 3 Princeton University, Princeton
Analysis of Absorbing Sets and Fully Absorbing Sets of Array-Based LDPC Codes
 Analysis of Absorbing Sets and Fully Absorbing Sets of Array-Based LDPC Codes Lara Dolecek, Zhengya Zhang, Venkat Anantharam, Martin J. Wainwright, and Borivoje Nikolić dolecek@mit.edu; {zyzhang,ananth,wainwrig,bora}@eecs.berkeley.edu
Analysis of Absorbing Sets and Fully Absorbing Sets of Array-Based LDPC Codes Lara Dolecek, Zhengya Zhang, Venkat Anantharam, Martin J. Wainwright, and Borivoje Nikolić dolecek@mit.edu; {zyzhang,ananth,wainwrig,bora}@eecs.berkeley.edu
Standard 2: Algebra Benchmark 1: Patterns
 Organizer Kindergarten First Grade Second Grade Third Grade Fourth Grade Fifth Grade Sixth Grade Seventh Grade Eighth Grade Ninth and Tenth Grade classifies and/or sorts... K.5...concrete objects by similar
Organizer Kindergarten First Grade Second Grade Third Grade Fourth Grade Fifth Grade Sixth Grade Seventh Grade Eighth Grade Ninth and Tenth Grade classifies and/or sorts... K.5...concrete objects by similar
Statistics 251/551 spring 2013 Homework # 4 Due: Wednesday 20 February
 1 Statistics 251/551 spring 2013 Homework # 4 Due: Wednesday 20 February If you are not able to solve a part of a problem, you can still get credit for later parts: Just assume the truth of what you were
1 Statistics 251/551 spring 2013 Homework # 4 Due: Wednesday 20 February If you are not able to solve a part of a problem, you can still get credit for later parts: Just assume the truth of what you were
Turing Machines. Lecture 8
 Turing Machines Lecture 8 1 Course Trajectory We will see algorithms, what can be done. But what cannot be done? 2 Computation Problem: To compute a function F that maps each input (a string) to an output
Turing Machines Lecture 8 1 Course Trajectory We will see algorithms, what can be done. But what cannot be done? 2 Computation Problem: To compute a function F that maps each input (a string) to an output
Language Recognition Power of Watson-Crick Automata
 01 Third International Conference on Netorking and Computing Language Recognition Poer of Watson-Crick Automata - Multiheads and Sensing - Kunio Aizaa and Masayuki Aoyama Dept. of Mathematics and Computer
01 Third International Conference on Netorking and Computing Language Recognition Poer of Watson-Crick Automata - Multiheads and Sensing - Kunio Aizaa and Masayuki Aoyama Dept. of Mathematics and Computer
Massachusetts Institute of Technology 6.042J/18.062J, Fall 02: Mathematics for Computer Science Professor Albert Meyer and Dr.
 Massachusetts Institute of Technology 6.042J/18.062J, Fall 02: Mathematics for Computer Science Professor Albert Meyer and Dr. Radhika Nagpal Quiz 1 Appendix Appendix Contents 1 Induction 2 2 Relations
Massachusetts Institute of Technology 6.042J/18.062J, Fall 02: Mathematics for Computer Science Professor Albert Meyer and Dr. Radhika Nagpal Quiz 1 Appendix Appendix Contents 1 Induction 2 2 Relations
Well-Ordering Principle. Axiom: Every nonempty subset of Z + has a least element. That is, if S Z + and S, then S has a smallest element.
 Well-Ordering Principle Axiom: Every nonempty subset of Z + has a least element. That is, if S Z + and S, then S has a smallest element. Well-Ordering Principle Example: Use well-ordering property to prove
Well-Ordering Principle Axiom: Every nonempty subset of Z + has a least element. That is, if S Z + and S, then S has a smallest element. Well-Ordering Principle Example: Use well-ordering property to prove
one two three four five six seven eight nine ten eleven twelve thirteen fourteen fifteen zero oneteen twoteen fiveteen tenteen
 Stacking races game Numbers, ordinal numbers, dates, days of the week, months, times Instructions for teachers Cut up one pack of cards. Divide the class into teams of two to four students and give them
Stacking races game Numbers, ordinal numbers, dates, days of the week, months, times Instructions for teachers Cut up one pack of cards. Divide the class into teams of two to four students and give them
Sugars, such as glucose or fructose are the basic building blocks of more complex carbohydrates. Which of the following
 Name: Score: / Quiz 2 on Lectures 3 &4 Part 1 Sugars, such as glucose or fructose are the basic building blocks of more complex carbohydrates. Which of the following foods is not a significant source of
Name: Score: / Quiz 2 on Lectures 3 &4 Part 1 Sugars, such as glucose or fructose are the basic building blocks of more complex carbohydrates. Which of the following foods is not a significant source of
Using DNA to Solve NP-Complete Problems. Richard J. Lipton y. Princeton University. Princeton, NJ 08540
 Using DNA to Solve NP-Complete Problems Richard J. Lipton y Princeton University Princeton, NJ 08540 rjl@princeton.edu Abstract: We show how to use DNA experiments to solve the famous \SAT" problem of
Using DNA to Solve NP-Complete Problems Richard J. Lipton y Princeton University Princeton, NJ 08540 rjl@princeton.edu Abstract: We show how to use DNA experiments to solve the famous \SAT" problem of
DE58/DC58 LOGIC DESIGN DEC 2014
 Q.2 a. In a base-5 number system, 3 digit representations is used. Find out (i) Number of distinct quantities that can be represented.(ii) Representation of highest decimal number in base-5. Since, r=5
Q.2 a. In a base-5 number system, 3 digit representations is used. Find out (i) Number of distinct quantities that can be represented.(ii) Representation of highest decimal number in base-5. Since, r=5
MATH Dr. Halimah Alshehri Dr. Halimah Alshehri
 MATH 1101 haalshehri@ksu.edu.sa 1 Introduction To Number Systems First Section: Binary System Second Section: Octal Number System Third Section: Hexadecimal System 2 Binary System 3 Binary System The binary
MATH 1101 haalshehri@ksu.edu.sa 1 Introduction To Number Systems First Section: Binary System Second Section: Octal Number System Third Section: Hexadecimal System 2 Binary System 3 Binary System The binary
A class of heuristics for the constrained forest problem
 Discrete Applied Mathematics 154 (2006) 6 14 www.elsevier.com/locate/dam Communication A class of heuristics for the constrained forest problem Michael Laszlo, Sumitra Mukherjee Nova Southeastern University,
Discrete Applied Mathematics 154 (2006) 6 14 www.elsevier.com/locate/dam Communication A class of heuristics for the constrained forest problem Michael Laszlo, Sumitra Mukherjee Nova Southeastern University,
Systems I: Computer Organization and Architecture
 Systems I: Computer Organization and Architecture Lecture 6 - Combinational Logic Introduction A combinational circuit consists of input variables, logic gates, and output variables. The logic gates accept
Systems I: Computer Organization and Architecture Lecture 6 - Combinational Logic Introduction A combinational circuit consists of input variables, logic gates, and output variables. The logic gates accept
Adders - Subtractors
 Adders - Subtractors Lesson Objectives: The objectives of this lesson are to learn about: 1. Half adder circuit. 2. Full adder circuit. 3. Binary parallel adder circuit. 4. Half subtractor circuit. 5.
Adders - Subtractors Lesson Objectives: The objectives of this lesson are to learn about: 1. Half adder circuit. 2. Full adder circuit. 3. Binary parallel adder circuit. 4. Half subtractor circuit. 5.
Chapter 9: Relations Relations
 Chapter 9: Relations 9.1 - Relations Definition 1 (Relation). Let A and B be sets. A binary relation from A to B is a subset R A B, i.e., R is a set of ordered pairs where the first element from each pair
Chapter 9: Relations 9.1 - Relations Definition 1 (Relation). Let A and B be sets. A binary relation from A to B is a subset R A B, i.e., R is a set of ordered pairs where the first element from each pair
A DNA procedure for solving the shortest path problem q
 Applied Mathematics and Computation 183 (006) 79 84 www.elsevier.com/locate/amc A DNA procedure for solving the shortest path problem q Zhaocai Wang a,c, Dongmei Xiao a,c, Wenxia Li b,c, *,1, Lin He c
Applied Mathematics and Computation 183 (006) 79 84 www.elsevier.com/locate/amc A DNA procedure for solving the shortest path problem q Zhaocai Wang a,c, Dongmei Xiao a,c, Wenxia Li b,c, *,1, Lin He c
17.1 The Halting Problem
 CS125 Lecture 17 Fall 2016 17.1 The Halting Problem Consider the HALTING PROBLEM (HALT TM ): Given a TM M and w, does M halt on input w? Theorem 17.1 HALT TM is undecidable. Suppose HALT TM = { M,w : M
CS125 Lecture 17 Fall 2016 17.1 The Halting Problem Consider the HALTING PROBLEM (HALT TM ): Given a TM M and w, does M halt on input w? Theorem 17.1 HALT TM is undecidable. Suppose HALT TM = { M,w : M
Complete Induction and the Well- Ordering Principle
 Complete Induction and the Well- Ordering Principle Complete Induction as a Rule of Inference In mathematical proofs, complete induction (PCI) is a rule of inference of the form P (a) P (a + 1) P (b) k
Complete Induction and the Well- Ordering Principle Complete Induction as a Rule of Inference In mathematical proofs, complete induction (PCI) is a rule of inference of the form P (a) P (a + 1) P (b) k
THE TEACHER UNDERSTANDS THE REAL NUMBER SYSTEM AND ITS STRUCTURE, OPERATIONS, ALGORITHMS, AND REPRESENTATIONS
 THE REAL NUMBER SYSTEM C O M P E T E N C Y 1 THE TEACHER UNDERSTANDS THE REAL NUMBER SYSTEM AND ITS STRUCTURE, OPERATIONS, ALGORITHMS, AND REPRESENTATIONS This competency section reviews some of the fundamental
THE REAL NUMBER SYSTEM C O M P E T E N C Y 1 THE TEACHER UNDERSTANDS THE REAL NUMBER SYSTEM AND ITS STRUCTURE, OPERATIONS, ALGORITHMS, AND REPRESENTATIONS This competency section reviews some of the fundamental
Randomized Sorting Algorithms Quick sort can be converted to a randomized algorithm by picking the pivot element randomly. In this case we can show th
 CSE 3500 Algorithms and Complexity Fall 2016 Lecture 10: September 29, 2016 Quick sort: Average Run Time In the last lecture we started analyzing the expected run time of quick sort. Let X = k 1, k 2,...,
CSE 3500 Algorithms and Complexity Fall 2016 Lecture 10: September 29, 2016 Quick sort: Average Run Time In the last lecture we started analyzing the expected run time of quick sort. Let X = k 1, k 2,...,
VERIZON PENNSYLVANIA LLC Pittsburgh Pa. P.U.C.-No. 185B
 VERIZON PENNSYLVANIA LLC Pittsburgh Pa. P.U.C.-No. 185B Exchange Areas Preface 2nd Revised Sheet 1 Canceling 1st Revised Sheet 1 The name of Verizon Pennsylvania Inc. has been changed to Verizon Pennsylvania
VERIZON PENNSYLVANIA LLC Pittsburgh Pa. P.U.C.-No. 185B Exchange Areas Preface 2nd Revised Sheet 1 Canceling 1st Revised Sheet 1 The name of Verizon Pennsylvania Inc. has been changed to Verizon Pennsylvania
Mathematics for Health and Physical Sciences
 1 Mathematics for Health and Physical Sciences Collection edited by: Wendy Lightheart Content authors: Wendy Lightheart, OpenStax, Wade Ellis, Denny Burzynski, Jan Clayton, and John Redden Online:
1 Mathematics for Health and Physical Sciences Collection edited by: Wendy Lightheart Content authors: Wendy Lightheart, OpenStax, Wade Ellis, Denny Burzynski, Jan Clayton, and John Redden Online:
Computer Science 324 Computer Architecture Mount Holyoke College Fall Topic Notes: Digital Logic
 Computer Science 324 Computer Architecture Mount Holyoke College Fall 2007 Topic Notes: Digital Logic Our goal for the next few weeks is to paint a a reasonably complete picture of how we can go from transistor
Computer Science 324 Computer Architecture Mount Holyoke College Fall 2007 Topic Notes: Digital Logic Our goal for the next few weeks is to paint a a reasonably complete picture of how we can go from transistor
1 Some loose ends from last time
 Cornell University, Fall 2010 CS 6820: Algorithms Lecture notes: Kruskal s and Borůvka s MST algorithms September 20, 2010 1 Some loose ends from last time 1.1 A lemma concerning greedy algorithms and
Cornell University, Fall 2010 CS 6820: Algorithms Lecture notes: Kruskal s and Borůvka s MST algorithms September 20, 2010 1 Some loose ends from last time 1.1 A lemma concerning greedy algorithms and
CS 7220 Computational Complexity and Algorithm Analysis
 CS 7220 Computational Complexity and Algorithm Analysis Spring 2016 Section 7: Computability Part IV Undecidability Pascal Hitzler Data Semantics Laboratory Wright State University, Dayton, OH http://www.pascal-hitzler.de
CS 7220 Computational Complexity and Algorithm Analysis Spring 2016 Section 7: Computability Part IV Undecidability Pascal Hitzler Data Semantics Laboratory Wright State University, Dayton, OH http://www.pascal-hitzler.de
Turing Machines. The Language Hierarchy. Context-Free Languages. Regular Languages. Courtesy Costas Busch - RPI 1
 Turing Machines a n b n c The anguage Hierarchy n? ww? Context-Free anguages a n b n egular anguages a * a *b* ww Courtesy Costas Busch - PI a n b n c n Turing Machines anguages accepted by Turing Machines
Turing Machines a n b n c The anguage Hierarchy n? ww? Context-Free anguages a n b n egular anguages a * a *b* ww Courtesy Costas Busch - PI a n b n c n Turing Machines anguages accepted by Turing Machines
DNA, Chromosomes, and Genes
 N, hromosomes, and Genes 1 You have most likely already learned about deoxyribonucleic acid (N), chromosomes, and genes. You have learned that all three of these substances have something to do with heredity
N, hromosomes, and Genes 1 You have most likely already learned about deoxyribonucleic acid (N), chromosomes, and genes. You have learned that all three of these substances have something to do with heredity
Introduction to Binary Convolutional Codes [1]
![Introduction to Binary Convolutional Codes [1] Introduction to Binary Convolutional Codes [1]](/thumbs/81/82925569.jpg) Introduction to Binary Convolutional Codes [1] Yunghsiang S. Han Graduate Institute of Communication Engineering, National Taipei University Taiwan E-mail: yshan@mail.ntpu.edu.tw Y. S. Han Introduction
Introduction to Binary Convolutional Codes [1] Yunghsiang S. Han Graduate Institute of Communication Engineering, National Taipei University Taiwan E-mail: yshan@mail.ntpu.edu.tw Y. S. Han Introduction
10.1 Radical Expressions and Functions Math 51 Professor Busken
 0. Radical Expressions and Functions Math 5 Professor Busken Objectives Find square roots without a calculator Simplify expressions of the form n a n Evaluate radical functions and find the domain of radical
0. Radical Expressions and Functions Math 5 Professor Busken Objectives Find square roots without a calculator Simplify expressions of the form n a n Evaluate radical functions and find the domain of radical
CS6901: review of Theory of Computation and Algorithms
 CS6901: review of Theory of Computation and Algorithms Any mechanically (automatically) discretely computation of problem solving contains at least three components: - problem description - computational
CS6901: review of Theory of Computation and Algorithms Any mechanically (automatically) discretely computation of problem solving contains at least three components: - problem description - computational
This article appeared in a journal published by Elsevier. The attached copy is furnished to the author for internal non-commercial research and
 This article appeared in a journal published by Elsevier. The attached copy is furnished to the author for internal non-commercial research and education use, including for instruction at the authors institution
This article appeared in a journal published by Elsevier. The attached copy is furnished to the author for internal non-commercial research and education use, including for instruction at the authors institution
Uncertain Quadratic Minimum Spanning Tree Problem
 Uncertain Quadratic Minimum Spanning Tree Problem Jian Zhou Xing He Ke Wang School of Management Shanghai University Shanghai 200444 China Email: zhou_jian hexing ke@shu.edu.cn Abstract The quadratic minimum
Uncertain Quadratic Minimum Spanning Tree Problem Jian Zhou Xing He Ke Wang School of Management Shanghai University Shanghai 200444 China Email: zhou_jian hexing ke@shu.edu.cn Abstract The quadratic minimum
Gray Codes and Overlap Cycles for Restricted Weight Words
 Gray Codes and Overlap Cycles for Restricted Weight Words Victoria Horan Air Force Research Laboratory Information Directorate Glenn Hurlbert School of Mathematical and Statistical Sciences Arizona State
Gray Codes and Overlap Cycles for Restricted Weight Words Victoria Horan Air Force Research Laboratory Information Directorate Glenn Hurlbert School of Mathematical and Statistical Sciences Arizona State
On the Multivariate Nakagami-m Distribution With Exponential Correlation
 1240 IEEE TRANSACTIONS ON COMMUNICATIONS, VOL. 51, NO. 8, AUGUST 2003 On the Multivariate Nakagami-m Distribution With Exponential Correlation George K. Karagiannidis, Member, IEEE, Dimitris A. Zogas,
1240 IEEE TRANSACTIONS ON COMMUNICATIONS, VOL. 51, NO. 8, AUGUST 2003 On the Multivariate Nakagami-m Distribution With Exponential Correlation George K. Karagiannidis, Member, IEEE, Dimitris A. Zogas,
convert a two s complement number back into a recognizable magnitude.
 1 INTRODUCTION The previous lesson introduced binary and hexadecimal numbers. In this lesson we look at simple arithmetic operations using these number systems. In particular, we examine the problem of
1 INTRODUCTION The previous lesson introduced binary and hexadecimal numbers. In this lesson we look at simple arithmetic operations using these number systems. In particular, we examine the problem of
IN this paper, we consider the capacity of sticky channels, a
 72 IEEE TRANSACTIONS ON INFORMATION THEORY, VOL. 54, NO. 1, JANUARY 2008 Capacity Bounds for Sticky Channels Michael Mitzenmacher, Member, IEEE Abstract The capacity of sticky channels, a subclass of insertion
72 IEEE TRANSACTIONS ON INFORMATION THEORY, VOL. 54, NO. 1, JANUARY 2008 Capacity Bounds for Sticky Channels Michael Mitzenmacher, Member, IEEE Abstract The capacity of sticky channels, a subclass of insertion
THE TEACHER UNDERSTANDS THE REAL NUMBER SYSTEM AND ITS STRUCTURE, OPERATIONS, ALGORITHMS, AND REPRESENTATIONS
 The real number SySTeM C O M P E T E N C Y 1 THE TEACHER UNDERSTANDS THE REAL NUMBER SYSTEM AND ITS STRUCTURE, OPERATIONS, ALGORITHMS, AND REPRESENTATIONS This competency section reviews some of the fundamental
The real number SySTeM C O M P E T E N C Y 1 THE TEACHER UNDERSTANDS THE REAL NUMBER SYSTEM AND ITS STRUCTURE, OPERATIONS, ALGORITHMS, AND REPRESENTATIONS This competency section reviews some of the fundamental
Part 8 Working with Nucleic Acids
 Part 8 Working with Nucleic Acids http://cbm.msoe.edu/newwebsite/learntomodel Introduction Most Protein Databank files loaded into the CBM's Jmol Design Environment include protein structures and small
Part 8 Working with Nucleic Acids http://cbm.msoe.edu/newwebsite/learntomodel Introduction Most Protein Databank files loaded into the CBM's Jmol Design Environment include protein structures and small
Subquadratic space complexity multiplier for a class of binary fields using Toeplitz matrix approach
 Subquadratic space complexity multiplier for a class of binary fields using Toeplitz matrix approach M A Hasan 1 and C Negre 2 1 ECE Department and CACR, University of Waterloo, Ontario, Canada 2 Team
Subquadratic space complexity multiplier for a class of binary fields using Toeplitz matrix approach M A Hasan 1 and C Negre 2 1 ECE Department and CACR, University of Waterloo, Ontario, Canada 2 Team
We want to show P (n) is true for all integers
 Generalized Induction Proof: Let P (n) be the proposition 1 + 2 + 2 2 + + 2 n = 2 n+1 1. We want to show P (n) is true for all integers n 0. Generalized Induction Example: Use generalized induction to
Generalized Induction Proof: Let P (n) be the proposition 1 + 2 + 2 2 + + 2 n = 2 n+1 1. We want to show P (n) is true for all integers n 0. Generalized Induction Example: Use generalized induction to
Performance Evaluation of Signed-Digit Architecture for Weighted-to-Residue and Residue-to-Weighted Number Converters with Moduli Set (2 n 1, 2 n,
 Regular Paper Performance Evaluation of Signed-Digit Architecture for Weighted-to-Residue and Residue-to-Weighted Number Converters with Moduli Set (2 n 1, 2 n, 2 n +1) Shuangching Chen and Shugang Wei
Regular Paper Performance Evaluation of Signed-Digit Architecture for Weighted-to-Residue and Residue-to-Weighted Number Converters with Moduli Set (2 n 1, 2 n, 2 n +1) Shuangching Chen and Shugang Wei
