Basics of Electronic-Structure Theory
|
|
|
- Gervais Barton
- 5 years ago
- Views:
Transcription
1 A guideline through the tutorial For each exercise in this tutorial, a separate directory should be created in your home directory in which the input files are generated and the calculation is started, like e.g. mkdir tutorial1/n2 mkdir tutorial1/si3... For every exercise, a directory is specified at the top of the description which contains reference files for all required input files as well as a reference output to check your results. However, we strongly recommend to use the provided control.in only in case of time problems and instead to try to generate the input files on your own. All references files are highlighted in italics throughout this script. Note: To avoid confusion, do not touch the directories including the reference files! Instead do all the calculations in your home directory with the created subdirectories as described above. In case you need the reference files and edit them, copy them into your home directory. Several scripts are required for differerent tasks. All scripts that are needed for this tutorial can be found in - /usr/local/aimsfiles/scripts Additional programs are installed to visualize results, like the basic geometry of a structure, the vibrational modes, charge density plots, binding curve etc. These programs are molden vmd jmol gdis As plotting tools, you can use xmgrace gnuplot Several editors are provided, such as emacs vi kwrite Basics of Electronic-Structure Theory How to use FHI-aims In this section, the most fundamental aspects of how to run an FHI-aims calculation are covered. For a detailed description please refer to the manual provided with the conference material. Furthermore, we present the most important tools that will be used throughout the tutorials as well as a guideline for the present exercises. Some fundamental aspects about FHI-aims For the tutorials a binary of FHI-aims will be provided on your workstation. A typical calculation is done in a separate directory which contains the two mandatory input files control.in and geometry.in. FHI-aims is then simply called in this directory by - mpirun -np 2 aims.workshop.mpi.x tee calculation.out which starts an mpi calculation on two processors and pipes the main output to the file calculation.out. This file contains the basic information such as the total energy, atomic forces, etc.. Additional output files might be generated according to the specified settings. The basic input files geometry.in contains the initial system geometry, and any information which is specifically tied to certain atoms. All other runtime settings are included in control.in. We note that the input units in both files for energies and lengths are generally ev and Å, respectively. Internally, these are converted to atomic units (Hartree for energies and bohr radii for lengths), using the 2002 National Institute of Standards CODATA recommended values. The order of lines is irrelevant, execept that information specific to a given atom must follow after the line specifying that atom and before any following atom is specified. The control.in typically consists of a general part, where, again, the particular order of lines is unimportant. In addition, this file contains species subtags that are referenced by geometry.in. Within the description of a given species, the order of lines is again unimportant, but all information concerning the same species must follow the initial species tag in one block. Lines beginning with a # symbol are treated as comments, and empty lines are ignored. FHI-aims provides preconstructed default definitions for the important subkeywords associated with different species (chemical elements) from Z=1-102 (H-Md). These can be found in the directory - /usr/local/aimsfiles/species defaults and are built for inclusion into a control.in file after the general part by simple copy-paste. We provide two default levels of accuracy. tight Tightly converged settings, the basis set might still need to be increased for light elements, though. light For fast prerelaxations. Any of these settings can always be further tightened by modifying them by hand in the species section. A summary of the most important convergence settings can be found in the manual in chapter 2.3.
2 Check the output of the atomic calculation. Is Hund s rule fulfilled? The multiplicity (J = 2S + 1) you obtain should be 4. (N.pw-lda.out.ref) Plot the binding energy as a function of the bond distance (using e.g. xmgrace). Redo the upper exercise with the pbe xc functional. Compare the (roughly) obtained equilibrium bond distance and binding energies to the experimental value of 9.8 ev and 1.1 Å [4]. Do you get the expected result for pw-lda and pbe? (N.pbe.out.ref) Now let s check the most important parameter to converge: the size of the basis set. Take the equilibrium bond distance (which should be 1.1 Å) and recalculate the binding energy for the basis sets minimal, tier1, tier2, tier3 using the pw-lda-functional (still using light settings for all other parameters). This is done by simply uncommenting/commenting basis functions near the end of the species defaults. Recalculate the binding energy with the tight settings. Have a closer look at the differences in the parameters, like the grid settings for instance. How much do the results change? To separate the effect of the basis set, compare the results with the light settings using the same basis set as specified in the tight settings. The nitrogen dimer Problem I: Getting started - N 2 Directory: /usr/local/aimsfiles/tutorial1/n2 /usr/local/aimsfiles/tutorial1/n2/reference output Motivation: In this exercise, the basics of FHI-aims will be conveyed, that are the generation of the input files the discussion of the output file the different functionals available in FHI-aims basis set convergence technical parameters to be aware of Generate a simple geometry.in file by hand for a nitrogen dimer with an initial (too short) bond distance of 0.8 Å(geometry.in.N2.ref). Generate a control.in file for nitrogen using light species defaults and the following settings for the physical model and the smearing (control.in.pw-lda.light.ref). For the general part, you can use the provided template (control.in.basic). xc pw-lda spin none occupation type gaussian 0.01 KS method lapack empty states 5 The self-consistency accuracy can be chosen throughout the whole tutorial as sc accuracy rho 1E-5 sc accuracy eev 1E-3 sc accuracy etot 1E-6 sc iter limit 200 Paste the species defaults in the control.in file. /usr/local/aimsfiles/species defaults/light/07 N default Now, run FHI-aims for this bond distance and have a look at the final total energy after self-consistency is achieved. Redo the calculation for the bond distances (1.0, 1.1, 1.2, 1.5 Å) and plot the binding curve (e.g. using xmgrace). Note: Since you are dealing with clusters, you will always consider the total energy and not the back-extrapolated total energy for T->0! To get the binding energy, you need the atomic reference energy. Therefore create a geometry.in filefortheatom.(geometry.in.atom.ref) The atom needs to be calculated spin-polarized. Therefore, add to the general part of the control.in file which does an unconstrained spin calculation. Furthermore, the cutoff radius needs to be enhanced for a single atom since there are no overlapping basis functions of the neighbouring atoms. cut pot
3 Now, open the Si3.prerelax.irc file with molden. molden -m Si3.prerelax.irc Open the file Si3.prerelax.out and copy the final atomic structure into a new geometry.in file as initial geometry for the ensuing post-relaxation. (geometry.in.si3.prerelaxed) Generate a control.in file with the upper specifications but with tight species defaults for Si. (control.in.si3.pbe.tight, control.in.si3.pw-lda.tight) Post-relax the structure for both the pw-lda and the pbe functional and have a look at the corresponding structures using molden. (Si3.postrelax.pbe.out, Si3.postrelax.pw-lda.out) How much does the structure change going from light to tight settings? What are the differences in the structure between the two functionals? estimated cpu time: 5 min Problem III: Vibrational analysis (harmonic) Directory: /usr/local/aimsfiles/tutorial1/si3/vib pw-lda, /usr/local/aimsfiles/tutorial1/si3/vib pbe The harmonic vibrational analysis is done using finite differences by the provided script aims.vibrations.pl. This script has to be called in the working directory which contains the control.in and geometry.in files. Create subdirectories (e.g. vib pw-lda and vib pbe) for the corresponding calculations and copy the corresponding control.in files that you have used for the post-relaxation. (control.in.si3.pbe.tight, control.in.si3.pw-lda.tight) Create the corresponding geometry.in files like in the procedure above by copying and pasting the finalized atomic structure from the output files of the post-relaxation. (geometry.in.si3.postrelaxed.pw-lda, geometry.in.si3.postrelaxed.pbe) The same control.in files from the post-relaxationscan be used. (control.in.si3.pbe.postrelax, control.in.si3.pw-lda.postrelax) In case of a vibrational analysis, the script does always a preceding local structural relaxation to ensure the structure to be in a local minimum. In your case, this should be done in one geometry step since you use the post-relaxed structure as a starting geometry. Invoke the vibrational analysis by aims.vibrations.pl jobname where you should provide a reasonable jobname (e.g. Si3 pw-lda, or Si3 pbe respectively). In the working directory, the main output is jobname.xyz which can be opened with jmol to visualize the frequencies and the corresponding eigenmodes and IR-frequencies. Alternatively, the output file jobname.mol can be used to visualize the vibrational spectrum with molden. AlltheFHI-aims output-files, that have been created, are summarized in the jobname.tar.gz file, which can be opened with tar -xzvf jobname.tar.gz Regarding the interpretation of the eigenvalues in the output Si3 pbe.vib.out, thevalues of the first six are compared with the largest one and should be zero (the three translational and vibrational degrees of freedom). Since SI-units are used at this point (the actual frequencies are given in wavenumbers), these values are very large (> O(20)). Due to numerical uncertainties, the six smallest ones are not exactly zero but several orders of magnitude smaller than the largest one. What do the frequencies tell you about the stability of the isomer? estimated cpu time: 15 min The silicon trimer Problem II: Local structural relaxation Motivation: In this exercise, the basic procedure for a local structural relaxation is presented: First, pre-relax a given initial structure with light settings using the pw-lda functional. Then, further post-relax the structure using tight settings and the desired exchangecorrelation functional. Directory: /usr/local/aimsfiles/tutorial1/si3 Generate a geometry.in file containing a random structure of three silicon atoms using the provided random structure generator create geometry.x: create geometry.x d min D Si 3 > geometry.in (geometry.in.si3 random) which creates a random structure with each atom having coordinates between { D...D} and not being closer to each other than d min. The origin of the coordinate system is set to the centroid of the cluster. Provide reasonable ranges d min=0.5 Å, D=1.0 Å. create geometry.x Si 3 > geometry.in (geometry.in.si3 random) Note: Type 1.0 and not 1 as d min! Generate a control.in file using the following settings (control.in.si3.prerelax) xc pw-lda occupation type gaussian 0.1 spin mix param 0.2 relax geometry bfgs 1d-2 and the light species defaults for Si. For that, paste /usr/local/aimsfiles/species defaults/light/14 Si default into the control.in file. Throughout this tutorial, the self-consistency accuracy for the forces can be set to sc accuracy forces 1E-4 Invoke the FHI-aims calculation and pipe the output to a file, e.g. mpirun -np 2 aims.workshop.mpi.x tee Si3.prerelax.out Si3.prerelax.out Visualize the local structural relaxation with molden. Therefore, you first have to generate a molden file by the provided script create relax movie.pl create relax movie.pl Si3.prerelax.out > Si3.prerelax.irc Si3.prerelax.irc
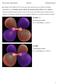
4 Figure 2: Isosurface (0.5 Å 3 ) of the charge density distribution for a silicon trimer with pw-lda. X (+) Si 3,X=Sc,Ti,V,Cr Problem V: Predicting the structure of small molecules Directory: /usr/local/aimsfiles/tutorial1/x(+)si3/xsi3.pw-lda, /usr/local/aimsfiles/tutorial1/x(+)si3/xsi3.pbe, /usr/local/aimsfiles/tutorial1/x(+)si3/x+si3.pw-lda, /usr/local/aimsfiles/tutorial1/x(+)si3/x+si3.pbe Motivation: This exercise gives a simple example of how to determine cluster geometries. In this simple approach, atoms are placed randomly (like in the case of Si 3) and locally relaxed. This is done multiple times, thereby generating cluster geometries without putting any chemical intuition into the starting geometry. The identified ground-state geometries are then analyzed and compared. More sophisticated methods that are important for larger systems like Basin-Hopping [1, 2] or Genetic Algorithms [3] are more or less based upon this principle but differ in the way to generate trial geometries. A structure search is done for the four-atomic systems X (+) Si 3,withX=Sc,Ti,Vand Cr. Each system should be treated both cationic and neutral for both the pw-lda and the pbe functional, so all in all sixteen cases. Note: Each group will only take care of ONE of these cases, where three groups will share one system, thus tripling the amount of trial structures for each case. The distribution of the systems will be coordinated by the tutor. Create a new working directory for your testsystem. With the already known tool create geometry.x, five random structures should be generated like - create geometry.x d min D Si 3 Sc 1 > geometry.in.struct1 which for instance generates a four-atomic geometry containing three silicon atoms and one scandium atom with minimum bond distance d min=0.5 ÅandD=1.0 Å. Generate a control.in file using light species defaults for the corresponding species. Use the specific control.in file settings: (control.in.prerelax.ref) charge 1.0 (for cationic systems) charge 0.0 (for neutral systems) spin mix param 1.0 relax geometry bfgs 1d-2 Problem IV: Charge density plots Figure 1: IR-spectra for the silicon trimer using tight. Directory: /usr/local/aimsfiles/tutorial1/si3/vib pw-lda, /usr/local/aimsfiles/tutorial1/si3/vib pbe The cube files for the charge density plots as well as for the HOMO can be generated as post-processing in the same sub-directories where the vibrational analysis has been done. The restart.dat files can be used to reinitialize the densities by adding to the control.in file. (control.in.si3.pw-lda.cube.out, control.in.si3.pbe.cube.out) restart read only restart.dat output cube total density cube origin cube edge cube edge cube edge output cube eigenstate density i max cube state σ where i max is the index of the HOMO to plot the eigenstate density of the HOMO that you can get by checking the eigenvalue spectrum of the post-relaxed structures. σ denotes the corresponding spin channel (1 ˆ=spin-up, 2 ˆ=spin-down). total density plots the total charge density distribution of the system. Running the calculation again will generate the cube files for the total density and for the HOMO state. Depending upon which spin-channel contains the HOMO, your cube-file of interest is eigenstate density i max spin σ.cube Visualize the cube files with molden or jmol. estimated cpu time: 2 min
5 References [1] D.J. Wales and H.A. Scheraga, Science 285, 1368 (1999). [2] D.J. Wales, J.P.K. Doye, M.A. Miller, P.N. Mortenson, and T.R. Walsh, Adv. Chem. Phys. 115, 1 (2000). [3] D.M. Deaven and K.M. Ho, Phys. Rev. Lett. 75, 288 (1995). [4] K. Huber and G. Herzberg, Molecular Spectra and Molecular Structure. IV. Constants of Diatomic Molecules (Van Nostrand, Princeton, 1979). occupation type gaussian 0.2 restart write only restart.dat Give the control.in file an appendix like control.in.prerelax mv control.in control.in.prerelax The provided script combi run.pl which is simply called in the working directory runs all combinations of control.in and geometry.in files and pipes the output to aims.struct1.prerelax for instance. The corresponding restart.dat file is copied to restart.struct1.prerelax.dat so that it can later be used for post-processing. Generate all the relaxation files with create relax movie.pl and have a look at the geometries. Compare results with the partner groups and generate a small table with the energy differences between the different isomers you found. Generate the total density plot and the HOMO eigenstate density plot for the post-relaxed ground-state structure using the restart file. Additionally, have a look at the density of states. Use the following control.in settings (control.in.cube.out.ref) occupation type gaussian 0.05 restart read only restartfile output dos output cube total density cube origin cube edge cube edge cube edge output cube eigenstate density i max cube state σ where again i max corresponds to the index of the HOMO eigenstate and σ to the spin channel (1 ˆ=spin-up, 2 ˆ=spin-down). Use as restartfile the corresponding file created in the prerelaxation, e.g. restart read only restart.struct1.prerelax.dat The density of states is printed to the ASCII-files KS DOS total raw.dat and KS DOS total.dat. The second file contains the dos shifted with the Fermi energy as reference energy while the first one contains the original dos. Printed is the energy range ev using 5000 points and a gaussian smearing of 0.01 ev to obtain sharp peaks for the Kohn-Sham eigenvalues. The electronic smearing is now reduced to 0.05 ev since you start from a reasonable density which facilitates the scf-convergence. Note: In the reference directories of the systems, all structure files structurei.xyz are given in the energetic order of the isomers. Additionally, the corresponding geometry.in.i files are provided that can be used for any kind of post-processing. For the identified ground-state structure, the dos, a cube file of the total density as well as of the HOMO are provided. The energetic differences and spin states are summarized in isomers.dat. Compare your results with those based upon the other xc functional. What are the differences in the geometries and energetic differences? Compare your results with those with a different charge state. Differences? Do the structural motifs or the energetic order change due to the different charge? Plot the dos for both spin channels using the shifted dos. Compare the different results obtained for the individual cases. Try to understand where the overall peaks come from and why they change going to the right in the row of the 3d-metals. Postrelax the ground-state structure with tight settings and a tighter force convergence criterium. (control.in.tight.ref) relax geometry bfgs 1.d-3 Does anything change significantly? Check the stability of the isomer explicitly by doing a vibrational analysis as explained in Problem III. estimated cpu time: 30 min
Before we start: Important setup of your Computer
 Before we start: Important setup of your Computer change directory: cd /afs/ictp/public/shared/smr2475./setup-config.sh logout login again 1 st Tutorial: The Basics of DFT Lydia Nemec and Oliver T. Hofmann
Before we start: Important setup of your Computer change directory: cd /afs/ictp/public/shared/smr2475./setup-config.sh logout login again 1 st Tutorial: The Basics of DFT Lydia Nemec and Oliver T. Hofmann
DFT and beyond: Hands-on Tutorial Workshop Tutorial 1: Basics of Electronic Structure Theory
 DFT and beyond: Hands-on Tutorial Workshop 2011 Tutorial 1: Basics of Electronic Structure Theory V. Atalla, O. T. Hofmann, S. V. Levchenko Theory Department, Fritz-Haber-Institut der MPG Berlin July 13,
DFT and beyond: Hands-on Tutorial Workshop 2011 Tutorial 1: Basics of Electronic Structure Theory V. Atalla, O. T. Hofmann, S. V. Levchenko Theory Department, Fritz-Haber-Institut der MPG Berlin July 13,
Hands-On Tutorial on Ab Initio Molecular Simulations Berlin, July 12 21, 2011
 Hands-On Tutorial on Ab Initio Molecular Simulations Berlin, July 12 21, 2011 Tutorial I: Basics of Electronic-Structure Theory Manuscript for Exercise Problems Prepared by Viktor Atalla, Oliver Hofmann,
Hands-On Tutorial on Ab Initio Molecular Simulations Berlin, July 12 21, 2011 Tutorial I: Basics of Electronic-Structure Theory Manuscript for Exercise Problems Prepared by Viktor Atalla, Oliver Hofmann,
Theoretical Material Science: Electronic structure theory at the computer Exercise 14: Brillouin zone integration
 Theoretical Material Science: Electronic structure theory at the computer Exercise 14: Brillouin zone integration Prepared by Lydia Nemec and Volker Blum Berlin, May 2012 Some rules on expected documentation
Theoretical Material Science: Electronic structure theory at the computer Exercise 14: Brillouin zone integration Prepared by Lydia Nemec and Volker Blum Berlin, May 2012 Some rules on expected documentation
Hands-on Summer School: Electronic Structure Theory for Materials and (Bio)molecules Los Angeles, July 21 - August 1, 2014
 Hands-on Summer School: Electronic Structure Theory for Materials and (Bio)molecules Los Angeles, July 21 - August 1, 2014 Tutorial I: Basics of Electronic-Structure Theory Manuscript for Exercise Problems
Hands-on Summer School: Electronic Structure Theory for Materials and (Bio)molecules Los Angeles, July 21 - August 1, 2014 Tutorial I: Basics of Electronic-Structure Theory Manuscript for Exercise Problems
Theoretical Material Science: Electronic Structure Theory at the Computer
 Theoretical Material Science: Electronic Structure Theory at the Computer Prepared by Somayeh Nafchi, Mohsen Yarmohammadi, Niklas Menzel, and Christian Carbogno Based on an exercise by Björn Bieniek, Volker
Theoretical Material Science: Electronic Structure Theory at the Computer Prepared by Somayeh Nafchi, Mohsen Yarmohammadi, Niklas Menzel, and Christian Carbogno Based on an exercise by Björn Bieniek, Volker
Predicting the Structure of Solids by DFT
 Questions? Hudson Hall 235 or Hudson Hall 1111 Predicting the Structure of Solids by DFT Hands-On Instructions Contents 1 Cohesive Energy for Bulk Phases of Si 11 Setting up the Structures 12 Structure-Dependent
Questions? Hudson Hall 235 or Hudson Hall 1111 Predicting the Structure of Solids by DFT Hands-On Instructions Contents 1 Cohesive Energy for Bulk Phases of Si 11 Setting up the Structures 12 Structure-Dependent
Tutorial I: Basics of Electronic-Structure Theory Manuscript for Exercise Problems
 Hands-on workshop: Density-functional theory and beyond - accuracy, efficiency and reproducibility in computational materials science Berlin, July 31 - August 11, 2017 Tutorial I: Basics of Electronic-Structure
Hands-on workshop: Density-functional theory and beyond - accuracy, efficiency and reproducibility in computational materials science Berlin, July 31 - August 11, 2017 Tutorial I: Basics of Electronic-Structure
Conformational space and energetics of biomolecules: Physical concepts and performance of DFT based methods
 Conformational space and energetics of biomolecules: Physical concepts and performance of DFT based methods Alexandre Tkatchenko, Carsten Baldauf, Matti Ropo Practical Session III / Weekend Project Haber
Conformational space and energetics of biomolecules: Physical concepts and performance of DFT based methods Alexandre Tkatchenko, Carsten Baldauf, Matti Ropo Practical Session III / Weekend Project Haber
Making Electronic Structure Theory work
 Making Electronic Structure Theory work Ralf Gehrke Fritz-Haber-Institut der Max-Planck-Gesellschaft Berlin, 23 th June 2009 Hands-on Tutorial on Ab Initio Molecular Simulations: Toward a First-Principles
Making Electronic Structure Theory work Ralf Gehrke Fritz-Haber-Institut der Max-Planck-Gesellschaft Berlin, 23 th June 2009 Hands-on Tutorial on Ab Initio Molecular Simulations: Toward a First-Principles
ab initio Lattice Vibrations: Calculating the Thermal Expansion Coeffcient Felix Hanke & Martin Fuchs June 30, 2009 This afternoon s plan
 ab initio Lattice Vibrations: Calculating the Thermal Expansion Coeffcient Felix Hanke & Martin Fuchs June 3, 29 This afternoon s plan introductory talk Phonons: harmonic vibrations for solids Phonons:
ab initio Lattice Vibrations: Calculating the Thermal Expansion Coeffcient Felix Hanke & Martin Fuchs June 3, 29 This afternoon s plan introductory talk Phonons: harmonic vibrations for solids Phonons:
Hands-On Tutorial on Ab Initio Molecular Simulations Berlin, July 12 21, Tutorial II: Periodic systems Manuscript for Exercise Problems
 Hands-On Tutorial on Ab Initio Molecular Simulations Berlin, July 12 21, 2011 Tutorial II: Periodic systems Manuscript for Exercise Problems Prepared by Jürgen Wieferink, Lydia Nemec, and Volker Blum.
Hands-On Tutorial on Ab Initio Molecular Simulations Berlin, July 12 21, 2011 Tutorial II: Periodic systems Manuscript for Exercise Problems Prepared by Jürgen Wieferink, Lydia Nemec, and Volker Blum.
Table of Contents. Table of Contents Spin-orbit splitting of semiconductor band structures
 Table of Contents Table of Contents Spin-orbit splitting of semiconductor band structures Relavistic effects in Kohn-Sham DFT Silicon band splitting with ATK-DFT LSDA initial guess for the ground state
Table of Contents Table of Contents Spin-orbit splitting of semiconductor band structures Relavistic effects in Kohn-Sham DFT Silicon band splitting with ATK-DFT LSDA initial guess for the ground state
Introduction to Hartree-Fock calculations in Spartan
 EE5 in 2008 Hannes Jónsson Introduction to Hartree-Fock calculations in Spartan In this exercise, you will get to use state of the art software for carrying out calculations of wavefunctions for molecues,
EE5 in 2008 Hannes Jónsson Introduction to Hartree-Fock calculations in Spartan In this exercise, you will get to use state of the art software for carrying out calculations of wavefunctions for molecues,
CPMD Tutorial Atosim/RFCT 2009/10
 These exercices were inspired by the CPMD Tutorial of Axel Kohlmeyer http://www.theochem.ruhruni-bochum.de/ axel.kohlmeyer/cpmd-tutor/index.html and by other tutorials. Here is a summary of what we will
These exercices were inspired by the CPMD Tutorial of Axel Kohlmeyer http://www.theochem.ruhruni-bochum.de/ axel.kohlmeyer/cpmd-tutor/index.html and by other tutorials. Here is a summary of what we will
QUANTUM CHEMISTRY WITH GAUSSIAN : A VERY BRIEF INTRODUCTION (PART 2)
 QUANTUM CHEMISTRY WITH GAUSSIAN : A VERY BRIEF INTRODUCTION (PART 2) TARAS V. POGORELOV AND MIKE HALLOCK SCHOOL OF CHEMICAL SCIENCES, UIUC This tutorial continues introduction to Gaussian [2]. Here we
QUANTUM CHEMISTRY WITH GAUSSIAN : A VERY BRIEF INTRODUCTION (PART 2) TARAS V. POGORELOV AND MIKE HALLOCK SCHOOL OF CHEMICAL SCIENCES, UIUC This tutorial continues introduction to Gaussian [2]. Here we
Hands-on Summer School: Electronic Structure Theory for Materials and (Bio)molecules Los Angeles, July 21 - August 1, 2014
 Hands-on Summer School: Electronic Structure Theory for Materials and (Bio)molecules Los Angeles, July 21 - August 1, 2014 Tutorial 7: Integrating Concepts- Cluster Structure Prediction and Spectroscopy
Hands-on Summer School: Electronic Structure Theory for Materials and (Bio)molecules Los Angeles, July 21 - August 1, 2014 Tutorial 7: Integrating Concepts- Cluster Structure Prediction and Spectroscopy
IFM Chemistry Computational Chemistry 2010, 7.5 hp LAB2. Computer laboratory exercise 1 (LAB2): Quantum chemical calculations
 Computer laboratory exercise 1 (LAB2): Quantum chemical calculations Introduction: The objective of the second computer laboratory exercise is to get acquainted with a program for performing quantum chemical
Computer laboratory exercise 1 (LAB2): Quantum chemical calculations Introduction: The objective of the second computer laboratory exercise is to get acquainted with a program for performing quantum chemical
Pseudo potential exercises
 Pseudo potential exercises Johan M. Carlsson Fritz-Haber-Institut der Max-Planck-Gesellschaft D-14195 Berlin Introduction Castep contains an "on the fly" OTF-pseudo potential generator that can be used
Pseudo potential exercises Johan M. Carlsson Fritz-Haber-Institut der Max-Planck-Gesellschaft D-14195 Berlin Introduction Castep contains an "on the fly" OTF-pseudo potential generator that can be used
FME Modelling course 2011 Tutorial 1 ( )
 FME Modelling course 2011 Tutorial 1 (21.02.2011) Brief introduction to LMTO: To obtain the state, φ nlm (r), of an ELECTRON -and hence its charge density ρ(r) = φ nlm (r) 2 - we must solve Schrödinger's
FME Modelling course 2011 Tutorial 1 (21.02.2011) Brief introduction to LMTO: To obtain the state, φ nlm (r), of an ELECTRON -and hence its charge density ρ(r) = φ nlm (r) 2 - we must solve Schrödinger's
ABC of DFT: Hands-on session 1 Introduction into calculations on molecules
 ABC of DFT: Hands-on session 1 Introduction into calculations on molecules Tutor: Alexej Bagrets Wann? 09.11.2012, 11:30-13:00 Wo? KIT Campus Nord, Flachbau Physik, Geb. 30.22, Computerpool, Raum FE-6
ABC of DFT: Hands-on session 1 Introduction into calculations on molecules Tutor: Alexej Bagrets Wann? 09.11.2012, 11:30-13:00 Wo? KIT Campus Nord, Flachbau Physik, Geb. 30.22, Computerpool, Raum FE-6
Worksheet 2 Properties of fermions and Density Functional Theory
 Simulation Methods in Physics II (SS 2013) Worksheet 2 Properties of fermions and Density Functional Theory Jens Smiatek, Bibek Adhikari and Maria Fyta April 30, 2013 ICP, Uni Stuttgart Important remarks
Simulation Methods in Physics II (SS 2013) Worksheet 2 Properties of fermions and Density Functional Theory Jens Smiatek, Bibek Adhikari and Maria Fyta April 30, 2013 ICP, Uni Stuttgart Important remarks
Computing IR / Raman / VCD / ROA Spectra with CP2k and TRAVIS
 Computing IR / Raman / VCD / ROA Spectra with CP2k and TRAVIS Martin Brehm Martin Luther Universität Halle Wittenberg Martin_Brehm@gmx.de 0. Introduction In this exercise, we will compute the full set
Computing IR / Raman / VCD / ROA Spectra with CP2k and TRAVIS Martin Brehm Martin Luther Universität Halle Wittenberg Martin_Brehm@gmx.de 0. Introduction In this exercise, we will compute the full set
Geometry Optimisation
 Geometry Optimisation Matt Probert Condensed Matter Dynamics Group Department of Physics, University of York, UK http://www.cmt.york.ac.uk/cmd http://www.castep.org Motivation Overview of Talk Background
Geometry Optimisation Matt Probert Condensed Matter Dynamics Group Department of Physics, University of York, UK http://www.cmt.york.ac.uk/cmd http://www.castep.org Motivation Overview of Talk Background
GaAs -- MLWF. Special thanks to Elias Assmann (TU Graz) for the generous help in preparation of this tutorial
 GaAs -- MLWF + + Special thanks to Elias Assmann (TU Graz) for the generous help in preparation of this tutorial YouTube video: https://youtu.be/r4c1yhdh3ge 1. Wien2k SCF Create a tutorial directory, e.g.
GaAs -- MLWF + + Special thanks to Elias Assmann (TU Graz) for the generous help in preparation of this tutorial YouTube video: https://youtu.be/r4c1yhdh3ge 1. Wien2k SCF Create a tutorial directory, e.g.
Excercise : Ammonia Flipping
 Excercise : Ammonia Flipping Rennes, 1. September 2016 Faculty of Physics, AG-CMP, University of Vienna general remarks (1) this excercise consists of 4 steps which unfold if you untar the file ammonia
Excercise : Ammonia Flipping Rennes, 1. September 2016 Faculty of Physics, AG-CMP, University of Vienna general remarks (1) this excercise consists of 4 steps which unfold if you untar the file ammonia
On-the-fly pseudopotential generation in CASTEP
 On-the-fly pseudopotential generation in CASTEP Chris J. Pickard School of Physics and Astronomy, University of St Andrews St Andrews, KY16 9SS, United Kingdom September 13, 2006 A quick tutorial Default
On-the-fly pseudopotential generation in CASTEP Chris J. Pickard School of Physics and Astronomy, University of St Andrews St Andrews, KY16 9SS, United Kingdom September 13, 2006 A quick tutorial Default
Technical Note Calculations of Orbital Overlap Range Function EDR( r ; d) and Overlap Distance D(r )using Multiwfn
 Technical Note Calculations of Orbital Overlap Range Function EDR( r ; d) and Overlap Distance D(r )using Multiwfn Abstract The orbital overlap range function EDR( r; d) (J. Chem. Phys. 2014, 141, 144104)
Technical Note Calculations of Orbital Overlap Range Function EDR( r ; d) and Overlap Distance D(r )using Multiwfn Abstract The orbital overlap range function EDR( r; d) (J. Chem. Phys. 2014, 141, 144104)
Exchange Correlation Functional Investigation of RT-TDDFT on a Sodium Chloride. Dimer. Philip Straughn
 Exchange Correlation Functional Investigation of RT-TDDFT on a Sodium Chloride Dimer Philip Straughn Abstract Charge transfer between Na and Cl ions is an important problem in physical chemistry. However,
Exchange Correlation Functional Investigation of RT-TDDFT on a Sodium Chloride Dimer Philip Straughn Abstract Charge transfer between Na and Cl ions is an important problem in physical chemistry. However,
Homework 5 Computational Chemistry (CBE 60547)
 Homework 5 Computational Chemistry (CBE 60547) Prof. William F. Schneider Due: 1 Not Ar again! As they say, there are many ways to skin a cat! You have computed the wavefunctions of Ar
Homework 5 Computational Chemistry (CBE 60547) Prof. William F. Schneider Due: 1 Not Ar again! As they say, there are many ways to skin a cat! You have computed the wavefunctions of Ar
How to run SIESTA. Introduction to input & output files
 How to run SIESTA Introduction to input & output files Linear-scaling DFT based on Numerical Atomic Orbitals (NAOs) Born-Oppenheimer DFT Pseudopotentials Numerical atomic orbitals relaxations, MD, phonons.
How to run SIESTA Introduction to input & output files Linear-scaling DFT based on Numerical Atomic Orbitals (NAOs) Born-Oppenheimer DFT Pseudopotentials Numerical atomic orbitals relaxations, MD, phonons.
TUTORIAL 8: PHONONS, LATTICE EXPANSION, AND BAND-GAP RENORMALIZATION
 TUTORIAL 8: PHONONS, LATTICE EXPANSION, AND BAND-GAP RENORMALIZATION 1 INVESTIGATED SYSTEM: Silicon, diamond structure Electronic and 0K properties see W. Huhn, Tutorial 2, Wednesday August 2 2 THE HARMONIC
TUTORIAL 8: PHONONS, LATTICE EXPANSION, AND BAND-GAP RENORMALIZATION 1 INVESTIGATED SYSTEM: Silicon, diamond structure Electronic and 0K properties see W. Huhn, Tutorial 2, Wednesday August 2 2 THE HARMONIC
Example: H 2 O (the car file)
 Example: H 2 O (the car file) As a practical example of DFT methods we calculate the energy and electronic properties of the water molecule. In order to carry out the DFT calculation you will need a set
Example: H 2 O (the car file) As a practical example of DFT methods we calculate the energy and electronic properties of the water molecule. In order to carry out the DFT calculation you will need a set
Computational Material Science Part II-1: introduction. Horng-Tay Jeng ( 鄭弘泰 ) Institute of Physics, Academia Sinica
 Computational Material Science Part II-1: introduction Horng-Tay Jeng ( 鄭弘泰 ) Institute of Physics, Academia Sinica Outline Introduction of Computational Material Science (CMS) Density Functional Theory
Computational Material Science Part II-1: introduction Horng-Tay Jeng ( 鄭弘泰 ) Institute of Physics, Academia Sinica Outline Introduction of Computational Material Science (CMS) Density Functional Theory
Journal of Theoretical Physics
 1 Journal of Theoretical Physics Founded and Edited by M. Apostol 53 (2000) ISSN 1453-4428 Ionization potential for metallic clusters L. C. Cune and M. Apostol Department of Theoretical Physics, Institute
1 Journal of Theoretical Physics Founded and Edited by M. Apostol 53 (2000) ISSN 1453-4428 Ionization potential for metallic clusters L. C. Cune and M. Apostol Department of Theoretical Physics, Institute
QuantumWise. QuantumWise is now part of Synopsys
 Table of Contents Table of Contents NiSi2 Si interface Create the NiSi2/Si device The screening region Increase the length of the central region Set-up the calculation for the undoped device Dope the device
Table of Contents Table of Contents NiSi2 Si interface Create the NiSi2/Si device The screening region Increase the length of the central region Set-up the calculation for the undoped device Dope the device
Lab 1: Handout GULP: an Empirical energy code
 Lab 1: Handout GULP: an Empirical energy code We will be using the GULP code as our energy code. GULP is a program for performing a variety of types of simulations on 3D periodic solids, gas phase clusters,
Lab 1: Handout GULP: an Empirical energy code We will be using the GULP code as our energy code. GULP is a program for performing a variety of types of simulations on 3D periodic solids, gas phase clusters,
DFT EXERCISES. FELIPE CERVANTES SODI January 2006
 DFT EXERCISES FELIPE CERVANTES SODI January 2006 http://www.csanyi.net/wiki/space/dftexercises Dr. Gábor Csányi 1 Hydrogen atom Place a single H atom in the middle of a largish unit cell (start with a
DFT EXERCISES FELIPE CERVANTES SODI January 2006 http://www.csanyi.net/wiki/space/dftexercises Dr. Gábor Csányi 1 Hydrogen atom Place a single H atom in the middle of a largish unit cell (start with a
Ethene. Introduction. The ethene molecule is planar (i.e. all the six atoms lie in the same plane) and has a high degree of symmetry:
 FY1006 Innføring i kvantefysikk og TFY4215 Kjemisk fysikk og kvantemekanikk Spring 2012 Chemical Physics Exercise 1 To be delivered by Friday 27.04.12 Introduction Ethene. Ethylene, C 2 H 4, or ethene,
FY1006 Innføring i kvantefysikk og TFY4215 Kjemisk fysikk og kvantemekanikk Spring 2012 Chemical Physics Exercise 1 To be delivered by Friday 27.04.12 Introduction Ethene. Ethylene, C 2 H 4, or ethene,
Speed-up of ATK compared to
 What s new @ Speed-up of ATK 2008.10 compared to 2008.02 System Speed-up Memory reduction Azafulleroid (molecule, 97 atoms) 1.1 15% 6x6x6 MgO (bulk, 432 atoms, Gamma point) 3.5 38% 6x6x6 MgO (k-point sampling
What s new @ Speed-up of ATK 2008.10 compared to 2008.02 System Speed-up Memory reduction Azafulleroid (molecule, 97 atoms) 1.1 15% 6x6x6 MgO (bulk, 432 atoms, Gamma point) 3.5 38% 6x6x6 MgO (k-point sampling
Introduction to Hartree-Fock calculations using ORCA and Chemcraft
 Introduction to Hartree-Fock calculations using ORCA and Chemcraft In this exercise, you will get to use software for carrying out calculations of wavefunctions for molecules, the ORCA program. While ORCA
Introduction to Hartree-Fock calculations using ORCA and Chemcraft In this exercise, you will get to use software for carrying out calculations of wavefunctions for molecules, the ORCA program. While ORCA
MENA 9520 FME Modelling Tutorial 5 ( )
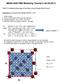 MENA 9520 FME Modelling Tutorial 5 (25.02.2011) Task 5: Understanding type of bonding using charge-density plots Exercise 5.1: Visualizing charge density in Si: 1. mkdir charge 2. Copy a converged CTRL
MENA 9520 FME Modelling Tutorial 5 (25.02.2011) Task 5: Understanding type of bonding using charge-density plots Exercise 5.1: Visualizing charge density in Si: 1. mkdir charge 2. Copy a converged CTRL
COMPUTATIONAL TOOL. Fig. 4.1 Opening screen of w2web
 CHAPTER -4 COMPUTATIONAL TOOL Ph.D. Thesis: J. Maibam CHAPTER: 4 4.1 The WIEN2k code In this work, all the calculations presented are performed using the WIEN2k software package (Blaha et al., 2001). The
CHAPTER -4 COMPUTATIONAL TOOL Ph.D. Thesis: J. Maibam CHAPTER: 4 4.1 The WIEN2k code In this work, all the calculations presented are performed using the WIEN2k software package (Blaha et al., 2001). The
NWChem: Hartree-Fock, Density Functional Theory, Time-Dependent Density Functional Theory
 NWChem: Hartree-Fock, Density Functional Theory, Time-Depent Density Functional Theory Hartree-Fock! Functionality! Input! Wavefunctions! Initial MO vectors! Direct and semidirect algorithms! Convergence,
NWChem: Hartree-Fock, Density Functional Theory, Time-Depent Density Functional Theory Hartree-Fock! Functionality! Input! Wavefunctions! Initial MO vectors! Direct and semidirect algorithms! Convergence,
Module 2: Quantum Espresso Walkthrough
 Module 2: Quantum Espresso Walkthrough Energy and Geometry Optimization of the H 2 Molecule We will be using the PWSCF code for quantum mechanical calculations of extended systems. The PWSCF program is
Module 2: Quantum Espresso Walkthrough Energy and Geometry Optimization of the H 2 Molecule We will be using the PWSCF code for quantum mechanical calculations of extended systems. The PWSCF program is
Tutorial on DFPT and TD-DFPT: calculations of phonons and absorption spectra
 Tutorial on DFPT and TD-DFPT: calculations of phonons and absorption spectra Iurii Timrov SISSA Scuola Internazionale Superiore di Studi Avanzati, Trieste Italy itimrov@sissa.it Computer modelling of materials
Tutorial on DFPT and TD-DFPT: calculations of phonons and absorption spectra Iurii Timrov SISSA Scuola Internazionale Superiore di Studi Avanzati, Trieste Italy itimrov@sissa.it Computer modelling of materials
An Approximate DFT Method: The Density-Functional Tight-Binding (DFTB) Method
 Fakultät für Mathematik und Naturwissenschaften - Lehrstuhl für Physikalische Chemie I / Theoretische Chemie An Approximate DFT Method: The Density-Functional Tight-Binding (DFTB) Method Jan-Ole Joswig
Fakultät für Mathematik und Naturwissenschaften - Lehrstuhl für Physikalische Chemie I / Theoretische Chemie An Approximate DFT Method: The Density-Functional Tight-Binding (DFTB) Method Jan-Ole Joswig
Hands-on Workshop: First-principles simulations of molecules and materials: Berlin, July 13 - July 23, 2015
 Hands-on Workshop: First-principles simulations of molecules and materials: Berlin, July 13 - July 23, 2015 R L e y E N e x Tutorial IV: Phonons, Lattice Expansion, and Band-gap Renormalization Manuscript
Hands-on Workshop: First-principles simulations of molecules and materials: Berlin, July 13 - July 23, 2015 R L e y E N e x Tutorial IV: Phonons, Lattice Expansion, and Band-gap Renormalization Manuscript
QUANTUM CHEMISTRY PROJECT 3: PARTS B AND C
 Chemistry 460 Fall 2017 Dr. Jean M. Standard November 6, 2017 QUANTUM CHEMISTRY PROJECT 3: PARTS B AND C PART B: POTENTIAL CURVE, SPECTROSCOPIC CONSTANTS, AND DISSOCIATION ENERGY OF DIATOMIC HYDROGEN (20
Chemistry 460 Fall 2017 Dr. Jean M. Standard November 6, 2017 QUANTUM CHEMISTRY PROJECT 3: PARTS B AND C PART B: POTENTIAL CURVE, SPECTROSCOPIC CONSTANTS, AND DISSOCIATION ENERGY OF DIATOMIC HYDROGEN (20
DFT calculation of pressure induced phase transition in silicon
 DFT calculation of pressure induced phase transition in silicon Michael Scherbela scherbela@student.tugraz.at 2015-12-07 Contents 1 Introduction 2 2 Structure of the input file 2 3 Calculation of phase
DFT calculation of pressure induced phase transition in silicon Michael Scherbela scherbela@student.tugraz.at 2015-12-07 Contents 1 Introduction 2 2 Structure of the input file 2 3 Calculation of phase
An ASE Calculator for OpenMX: providing automation of running OpenMX
 .dat.out.c An ASE Calculator for OpenMX: providing automation of running OpenMX Charles Johnson * and Jaejun Yu Seoul National University *Current Address: Imperial College London 2016.11.24 2nd OpenMX
.dat.out.c An ASE Calculator for OpenMX: providing automation of running OpenMX Charles Johnson * and Jaejun Yu Seoul National University *Current Address: Imperial College London 2016.11.24 2nd OpenMX
A self-interaction free non-local correlation functional derived from exact criteria
 A self-interaction free non-local correlation functional derived from exact criteria Tobias Schmidt and Stephan Kümmel Theoretical Physics IV University of Bayreuth Germany - International Workshop DFT
A self-interaction free non-local correlation functional derived from exact criteria Tobias Schmidt and Stephan Kümmel Theoretical Physics IV University of Bayreuth Germany - International Workshop DFT
Exercise 2: Solvating the Structure Before you continue, follow these steps: Setting up Periodic Boundary Conditions
 Exercise 2: Solvating the Structure HyperChem lets you place a molecular system in a periodic box of water molecules to simulate behavior in aqueous solution, as in a biological system. In this exercise,
Exercise 2: Solvating the Structure HyperChem lets you place a molecular system in a periodic box of water molecules to simulate behavior in aqueous solution, as in a biological system. In this exercise,
Quantum ESPRESSO. PWSCF: first steps
 Quantum ESPRESSO PWSCF: first steps What can I learn in this tutorial? What can I learn in this tutorial? How to run PWscf (pw.x) in self-consistent mode for Silicon How to get the band structure of Silicon
Quantum ESPRESSO PWSCF: first steps What can I learn in this tutorial? What can I learn in this tutorial? How to run PWscf (pw.x) in self-consistent mode for Silicon How to get the band structure of Silicon
Prerequisites for reliable modeling with first-principles methods. P. Kratzer Fritz-Haber-Institut der MPG D Berlin-Dahlem, Germany
 Prerequisites for reliable modeling with first-principles methods P. Kratzer Fritz-Haber-Institut der MPG D-14195 Berlin-Dahlem, Germany Prerequisites for modeling (I) Issues to consider when applying
Prerequisites for reliable modeling with first-principles methods P. Kratzer Fritz-Haber-Institut der MPG D-14195 Berlin-Dahlem, Germany Prerequisites for modeling (I) Issues to consider when applying
The Electronic Structure of Dye- Sensitized TiO 2 Clusters from Many- Body Perturbation Theory
 The Electronic Structure of Dye- Sensitized TiO 2 Clusters from Many- Body Perturbation Theory Noa Marom Center for Computational Materials Institute for Computational Engineering and Sciences The University
The Electronic Structure of Dye- Sensitized TiO 2 Clusters from Many- Body Perturbation Theory Noa Marom Center for Computational Materials Institute for Computational Engineering and Sciences The University
TUTORIAL 6: PHONONS, LATTICE EXPANSION, AND BAND-GAP RENORMALIZATION
 Hands-On Tutorial Workshop, July 29 th 2014 TUTORIAL 6: PHONONS, LATTICE EXPANSION, AND BAND-GAP RENORMALIZATION Christian Carbogno & Manuel Schöttler Fritz-Haber-Institut der Max-Planck-Gesellschaft,
Hands-On Tutorial Workshop, July 29 th 2014 TUTORIAL 6: PHONONS, LATTICE EXPANSION, AND BAND-GAP RENORMALIZATION Christian Carbogno & Manuel Schöttler Fritz-Haber-Institut der Max-Planck-Gesellschaft,
ABC of DFT: Hands-on session 4 Molecular vibrations
 ABC of DFT: Hands-on session 4 Molecular vibrations Tutor: Alexej Bagrets Wann? 29.11.2012, 11:30-13:00 Wo? KIT Campus Süd, Flachbau Physik, Geb. 30.22, Computerpool, Raum FE-6 1 ABC of DFT, Hands-on session
ABC of DFT: Hands-on session 4 Molecular vibrations Tutor: Alexej Bagrets Wann? 29.11.2012, 11:30-13:00 Wo? KIT Campus Süd, Flachbau Physik, Geb. 30.22, Computerpool, Raum FE-6 1 ABC of DFT, Hands-on session
Transition states and reaction paths
 Transition states and reaction paths Lab 4 Theoretical background Transition state A transition structure is the molecular configuration that separates reactants and products. In a system with a single
Transition states and reaction paths Lab 4 Theoretical background Transition state A transition structure is the molecular configuration that separates reactants and products. In a system with a single
Outline. Introduction: graphene. Adsorption on graphene: - Chemisorption - Physisorption. Summary
 Outline Introduction: graphene Adsorption on graphene: - Chemisorption - Physisorption Summary 1 Electronic band structure: Electronic properties K Γ M v F = 10 6 ms -1 = c/300 massless Dirac particles!
Outline Introduction: graphene Adsorption on graphene: - Chemisorption - Physisorption Summary 1 Electronic band structure: Electronic properties K Γ M v F = 10 6 ms -1 = c/300 massless Dirac particles!
DFT / SIESTA algorithms
 DFT / SIESTA algorithms Javier Junquera José M. Soler References http://siesta.icmab.es Documentation Tutorials Atomic units e = m e = =1 atomic mass unit = m e atomic length unit = 1 Bohr = 0.5292 Ang
DFT / SIESTA algorithms Javier Junquera José M. Soler References http://siesta.icmab.es Documentation Tutorials Atomic units e = m e = =1 atomic mass unit = m e atomic length unit = 1 Bohr = 0.5292 Ang
CHEM 498Q / CHEM 630Q: Molecular Modeling of Proteins TUTORIAL #3a: Empirical force fields
 Department of Chemistry and Biochemistry, Concordia University! page 1 of 6 CHEM 498Q / CHEM 630Q: Molecular Modeling of Proteins TUTORIAL #3a: Empirical force fields INTRODUCTION The goal of this tutorial
Department of Chemistry and Biochemistry, Concordia University! page 1 of 6 CHEM 498Q / CHEM 630Q: Molecular Modeling of Proteins TUTORIAL #3a: Empirical force fields INTRODUCTION The goal of this tutorial
References. Documentation Manuals Tutorials Publications
 References http://siesta.icmab.es Documentation Manuals Tutorials Publications Atomic units e = m e = =1 atomic mass unit = m e atomic length unit = 1 Bohr = 0.5292 Ang atomic energy unit = 1 Hartree =
References http://siesta.icmab.es Documentation Manuals Tutorials Publications Atomic units e = m e = =1 atomic mass unit = m e atomic length unit = 1 Bohr = 0.5292 Ang atomic energy unit = 1 Hartree =
Periodic Trends in Properties of Homonuclear
 Chapter 8 Periodic Trends in Properties of Homonuclear Diatomic Molecules Up to now, we have discussed various physical properties of nanostructures, namely, two-dimensional - graphene-like structures:
Chapter 8 Periodic Trends in Properties of Homonuclear Diatomic Molecules Up to now, we have discussed various physical properties of nanostructures, namely, two-dimensional - graphene-like structures:
Tutorial 1. Atoms, molecules, and bulk systems. Computational Materials Physics, Faculty of Physics, University Vienna, Vienna, Austria
 Tutorial 1 Atoms, molecules, and bulk systems Computational Materials Physics, Faculty of Physics, University Vienna, Vienna, Austria I. The INCAR file II. The POSCAR file III. The KPOINTS file IV. The
Tutorial 1 Atoms, molecules, and bulk systems Computational Materials Physics, Faculty of Physics, University Vienna, Vienna, Austria I. The INCAR file II. The POSCAR file III. The KPOINTS file IV. The
Pseudopotentials for hybrid density functionals and SCAN
 Pseudopotentials for hybrid density functionals and SCAN Jing Yang, Liang Z. Tan, Julian Gebhardt, and Andrew M. Rappe Department of Chemistry University of Pennsylvania Why do we need pseudopotentials?
Pseudopotentials for hybrid density functionals and SCAN Jing Yang, Liang Z. Tan, Julian Gebhardt, and Andrew M. Rappe Department of Chemistry University of Pennsylvania Why do we need pseudopotentials?
CO 2 molecule. Morse Potential One of the potentials used to simulate chemical bond is a Morse potential of the following form: O C O
 CO 2 molecule The aim of this project is a numerical analysis of adsorption spectra of CO2 molecule simulated by a double Morse potential function. In the project you should achieve following tasks: 1.
CO 2 molecule The aim of this project is a numerical analysis of adsorption spectra of CO2 molecule simulated by a double Morse potential function. In the project you should achieve following tasks: 1.
Density functional theory and beyond: Computational materials science for real materials Los Angeles, July 21 August 1, 2014
 Density functional theory and beyond: Computational materials science for real materials Los Angeles, July 21 August 1, 2014 R L e y E N e x Tutorial VI: Phonons, Lattice Expansion, and Band-gap Renormalization
Density functional theory and beyond: Computational materials science for real materials Los Angeles, July 21 August 1, 2014 R L e y E N e x Tutorial VI: Phonons, Lattice Expansion, and Band-gap Renormalization
Calculating Vibrational Spectra from Molecular Dynamics
 Calculating Vibrational Spectra from Molecular Dynamics A Simulating a Trajectory with Wannier Centers To calculate IR spectra from Molecular Dynamics, it is necessary to have dipole information for the
Calculating Vibrational Spectra from Molecular Dynamics A Simulating a Trajectory with Wannier Centers To calculate IR spectra from Molecular Dynamics, it is necessary to have dipole information for the
Supporting Information for Ultra-narrow metallic armchair graphene nanoribbons
 Supporting Information for Ultra-narrow metallic armchair graphene nanoribbons Supplementary Figure 1 Ribbon length statistics. Distribution of the ribbon lengths and the fraction of kinked ribbons for
Supporting Information for Ultra-narrow metallic armchair graphene nanoribbons Supplementary Figure 1 Ribbon length statistics. Distribution of the ribbon lengths and the fraction of kinked ribbons for
DENSITY FUNCTIONAL THEORY FOR NON-THEORISTS JOHN P. PERDEW DEPARTMENTS OF PHYSICS AND CHEMISTRY TEMPLE UNIVERSITY
 DENSITY FUNCTIONAL THEORY FOR NON-THEORISTS JOHN P. PERDEW DEPARTMENTS OF PHYSICS AND CHEMISTRY TEMPLE UNIVERSITY A TUTORIAL FOR PHYSICAL SCIENTISTS WHO MAY OR MAY NOT HATE EQUATIONS AND PROOFS REFERENCES
DENSITY FUNCTIONAL THEORY FOR NON-THEORISTS JOHN P. PERDEW DEPARTMENTS OF PHYSICS AND CHEMISTRY TEMPLE UNIVERSITY A TUTORIAL FOR PHYSICAL SCIENTISTS WHO MAY OR MAY NOT HATE EQUATIONS AND PROOFS REFERENCES
Key concepts in Density Functional Theory (II)
 Kohn-Sham scheme and band structures European Theoretical Spectroscopy Facility (ETSF) CNRS - Laboratoire des Solides Irradiés Ecole Polytechnique, Palaiseau - France Present Address: LPMCN Université
Kohn-Sham scheme and band structures European Theoretical Spectroscopy Facility (ETSF) CNRS - Laboratoire des Solides Irradiés Ecole Polytechnique, Palaiseau - France Present Address: LPMCN Université
Chemistry 483 Lecture Topics Fall 2009
 Chemistry 483 Lecture Topics Fall 2009 Text PHYSICAL CHEMISTRY A Molecular Approach McQuarrie and Simon A. Background (M&S,Chapter 1) Blackbody Radiation Photoelectric effect DeBroglie Wavelength Atomic
Chemistry 483 Lecture Topics Fall 2009 Text PHYSICAL CHEMISTRY A Molecular Approach McQuarrie and Simon A. Background (M&S,Chapter 1) Blackbody Radiation Photoelectric effect DeBroglie Wavelength Atomic
Density functional theory and beyond: Computational material science for real materials Trieste, August 06 15, 2013
 Density functional theory and beyond: Computational material science for real materials Trieste, August 06 15, 2013 Tutorial V: Theoretical Spectroscopy and Electronic Excitations Manuscript for Exercise
Density functional theory and beyond: Computational material science for real materials Trieste, August 06 15, 2013 Tutorial V: Theoretical Spectroscopy and Electronic Excitations Manuscript for Exercise
Due: since the calculation takes longer than before, we ll make it due on 02/05/2016, Friday
 Homework 3 Due: since the calculation takes longer than before, we ll make it due on 02/05/2016, Friday Email to: jqian@caltech.edu Introduction In this assignment, you will be using a commercial periodic
Homework 3 Due: since the calculation takes longer than before, we ll make it due on 02/05/2016, Friday Email to: jqian@caltech.edu Introduction In this assignment, you will be using a commercial periodic
Introduction to DFTB. Marcus Elstner. July 28, 2006
 Introduction to DFTB Marcus Elstner July 28, 2006 I. Non-selfconsistent solution of the KS equations DFT can treat up to 100 atoms in routine applications, sometimes even more and about several ps in MD
Introduction to DFTB Marcus Elstner July 28, 2006 I. Non-selfconsistent solution of the KS equations DFT can treat up to 100 atoms in routine applications, sometimes even more and about several ps in MD
Benzene Dimer: dispersion forces and electronic correlation
 Benzene Dimer: dispersion forces and electronic correlation Introduction The Benzene dimer is an ideal example of a system bound by π-π interaction, which is in several cases present in many biologically
Benzene Dimer: dispersion forces and electronic correlation Introduction The Benzene dimer is an ideal example of a system bound by π-π interaction, which is in several cases present in many biologically
DFT with Hybrid Functionals
 DFT with Hybrid Functionals Sanliang Ling University College London 4th CP2K Tutorial, 31st August 4th September 2015, Zurich What are hybrid functionals? Hybrid functionals: mixing non-local Hartree-Fock
DFT with Hybrid Functionals Sanliang Ling University College London 4th CP2K Tutorial, 31st August 4th September 2015, Zurich What are hybrid functionals? Hybrid functionals: mixing non-local Hartree-Fock
Analysis of the ultrafast dynamics of the silver trimer upon photodetachment
 J. Phys. B: At. Mol. Opt. Phys. 29 (1996) L545 L549. Printed in the UK LETTER TO THE EDITOR Analysis of the ultrafast dynamics of the silver trimer upon photodetachment H O Jeschke, M E Garcia and K H
J. Phys. B: At. Mol. Opt. Phys. 29 (1996) L545 L549. Printed in the UK LETTER TO THE EDITOR Analysis of the ultrafast dynamics of the silver trimer upon photodetachment H O Jeschke, M E Garcia and K H
1 Construction of norm-conserving semi-local pseudopotentials for Si
 1 Construction of norm-conserving semi-local pseudopotentials for Si As discussed in class, it is desirable to replace the effective interaction of the valence electrons with the ionic core, i.e. nucleus
1 Construction of norm-conserving semi-local pseudopotentials for Si As discussed in class, it is desirable to replace the effective interaction of the valence electrons with the ionic core, i.e. nucleus
The Nature of the Interlayer Interaction in Bulk. and Few-Layer Phosphorus
 Supporting Information for: The Nature of the Interlayer Interaction in Bulk and Few-Layer Phosphorus L. Shulenburger, A.D. Baczewski, Z. Zhu, J. Guan, and D. Tománek, Sandia National Laboratories, Albuquerque,
Supporting Information for: The Nature of the Interlayer Interaction in Bulk and Few-Layer Phosphorus L. Shulenburger, A.D. Baczewski, Z. Zhu, J. Guan, and D. Tománek, Sandia National Laboratories, Albuquerque,
Quantum ESPRESSO. Input and Output description
 Quantum ESPRESSO Input and Output description Where can I find useful information about Quantum ESPRESSO? Where can I find useful information about Quantum ESPRESSO? prompt > cd $espresso_dir/doc; ls *.html
Quantum ESPRESSO Input and Output description Where can I find useful information about Quantum ESPRESSO? Where can I find useful information about Quantum ESPRESSO? prompt > cd $espresso_dir/doc; ls *.html
OPENATOM for GW calculations
 OPENATOM for GW calculations by OPENATOM developers 1 Introduction The GW method is one of the most accurate ab initio methods for the prediction of electronic band structures. Despite its power, the GW
OPENATOM for GW calculations by OPENATOM developers 1 Introduction The GW method is one of the most accurate ab initio methods for the prediction of electronic band structures. Despite its power, the GW
Table of Contents. Table of Contents DFT-1/2 and DFT-PPS density functional methods for electronic structure calculations
 Table of Contents Table of Contents DFT-1/2 and DFT-PPS density functional methods for electronic structure calculations DFT-1/2 methods InP bandstructure using PBE-1/2 III-V type semiconductor band gaps
Table of Contents Table of Contents DFT-1/2 and DFT-PPS density functional methods for electronic structure calculations DFT-1/2 methods InP bandstructure using PBE-1/2 III-V type semiconductor band gaps
Part III: Theoretical Surface Science Adsorption at Surfaces
 Technische Universität München Part III: Theoretical Surface Science Adsorption at Surfaces Karsten Reuter Lecture course: Solid State Theory Adsorption at surfaces (T,p) Phase II Phase I Corrosion Growth
Technische Universität München Part III: Theoretical Surface Science Adsorption at Surfaces Karsten Reuter Lecture course: Solid State Theory Adsorption at surfaces (T,p) Phase II Phase I Corrosion Growth
Chemistry 881 Lecture Topics Fall 2001
 Chemistry 881 Lecture Topics Fall 2001 Texts PHYSICAL CHEMISTRY A Molecular Approach McQuarrie and Simon MATHEMATICS for PHYSICAL CHEMISTRY, Mortimer i. Mathematics Review (M, Chapters 1,2,3 & 4; M&S,
Chemistry 881 Lecture Topics Fall 2001 Texts PHYSICAL CHEMISTRY A Molecular Approach McQuarrie and Simon MATHEMATICS for PHYSICAL CHEMISTRY, Mortimer i. Mathematics Review (M, Chapters 1,2,3 & 4; M&S,
Structural optimizations
 Structural optimizations Guido Fratesi (Università di Milano) Urbana, August 2006 Acetylene molecule ( ) We want to use pw.x to find the optimized geometry of acetylene. Let us suppose we have the following
Structural optimizations Guido Fratesi (Università di Milano) Urbana, August 2006 Acetylene molecule ( ) We want to use pw.x to find the optimized geometry of acetylene. Let us suppose we have the following
PHI: Open quantum dynamics of multi-molecule systems
 University of Illinois at Urbana-Champaign Beckman Institute for Advanced Science and Technology Theoretical and Computational Biophysics Group PHI: Open quantum dynamics of multi-molecule systems Johan
University of Illinois at Urbana-Champaign Beckman Institute for Advanced Science and Technology Theoretical and Computational Biophysics Group PHI: Open quantum dynamics of multi-molecule systems Johan
Vibrational Spectroscopy Practical 2:
 Vibrational Spectroscopy Practical 2: The aims of today's session are to introduce you to running larger-scale CASTEP jobs on supercomputer clusters, creation of input files for and setup of CASTEP jobs,
Vibrational Spectroscopy Practical 2: The aims of today's session are to introduce you to running larger-scale CASTEP jobs on supercomputer clusters, creation of input files for and setup of CASTEP jobs,
LUMO + 1 LUMO. Tómas Arnar Guðmundsson Report 2 Reikniefnafræði G
 Q1: Display all the MOs for N2 in your report and classify each one of them as bonding, antibonding or non-bonding, and say whether the symmetry of the orbital is σ or π. Sketch a molecular orbital diagram
Q1: Display all the MOs for N2 in your report and classify each one of them as bonding, antibonding or non-bonding, and say whether the symmetry of the orbital is σ or π. Sketch a molecular orbital diagram
Introduction to numerical projects
 Introduction to numerical projects Here follows a brief recipe and recommendation on how to write a report for each project. Give a short description of the nature of the problem and the eventual numerical
Introduction to numerical projects Here follows a brief recipe and recommendation on how to write a report for each project. Give a short description of the nature of the problem and the eventual numerical
Supplementary information Silver (I) as DNA glue: Ag + - mediated guanine pairing revealed by removing Watson- Crick constraints
 Supplementary information Silver (I) as DNA glue: Ag + - mediated guanine pairing revealed by removing Watson- Crick constraints Steven M. Swasey [b], Leonardo Espinosa Leal [c], Olga Lopez- Acevedo [c],
Supplementary information Silver (I) as DNA glue: Ag + - mediated guanine pairing revealed by removing Watson- Crick constraints Steven M. Swasey [b], Leonardo Espinosa Leal [c], Olga Lopez- Acevedo [c],
Table of Contents. Table of Contents Opening a band gap in silicene and bilayer graphene with an electric field
 Table of Contents Table of Contents Opening a band gap in silicene and bilayer graphene with an electric field Bilayer graphene Building a bilayer graphene structure Calculation and analysis Silicene Optimizing
Table of Contents Table of Contents Opening a band gap in silicene and bilayer graphene with an electric field Bilayer graphene Building a bilayer graphene structure Calculation and analysis Silicene Optimizing
Explaining the apparent arbitrariness of the LDA-1/2 self-energy. correction method applied to purely covalent systems
 Explaining the apparent arbitrariness of the LDA-1/2 self-energy correction method applied to purely covalent systems Kan-Hao Xue, 1,2 Leonardo R. C. Fonseca, 3 and Xiang-Shui Miao 1,2 1 School of Optical
Explaining the apparent arbitrariness of the LDA-1/2 self-energy correction method applied to purely covalent systems Kan-Hao Xue, 1,2 Leonardo R. C. Fonseca, 3 and Xiang-Shui Miao 1,2 1 School of Optical
Lab 3: Handout Quantum-ESPRESSO: a first principles code, part 2.
 1 Lab 3: Handout Quantum-ESPRESSO: a first principles code, part 2. In this lab, we will be using Quantum-ESPRESSO as our first-principles code again. In problem 1, we will compare energy between allotropes
1 Lab 3: Handout Quantum-ESPRESSO: a first principles code, part 2. In this lab, we will be using Quantum-ESPRESSO as our first-principles code again. In problem 1, we will compare energy between allotropes
4 Post-Hartree Fock Methods: MPn and Configuration Interaction
 4 Post-Hartree Fock Methods: MPn and Configuration Interaction In the limit of a complete basis, the Hartree-Fock (HF) energy in the complete basis set limit (ECBS HF ) yields an upper boundary to the
4 Post-Hartree Fock Methods: MPn and Configuration Interaction In the limit of a complete basis, the Hartree-Fock (HF) energy in the complete basis set limit (ECBS HF ) yields an upper boundary to the
Table S2. Pseudopotentials PBE 5.2 applied in the calculations using VASP
 Supporting Information for Understanding the Adsorption of CuPc and ZnPc on Noble Metal Surfaces by Combining Quantum-Mechanical Modelling and Photoelectron Spectroscopy 1. Used vdw Coefficients PBE-vdW
Supporting Information for Understanding the Adsorption of CuPc and ZnPc on Noble Metal Surfaces by Combining Quantum-Mechanical Modelling and Photoelectron Spectroscopy 1. Used vdw Coefficients PBE-vdW
Physical Chemistry Laboratory II (CHEM 337) EXPT 9 3: Vibronic Spectrum of Iodine (I2)
 Physical Chemistry Laboratory II (CHEM 337) EXPT 9 3: Vibronic Spectrum of Iodine (I2) Obtaining fundamental information about the nature of molecular structure is one of the interesting aspects of molecular
Physical Chemistry Laboratory II (CHEM 337) EXPT 9 3: Vibronic Spectrum of Iodine (I2) Obtaining fundamental information about the nature of molecular structure is one of the interesting aspects of molecular
CHE3935. Lecture 4 Quantum Mechanical Simulation Methods Continued
 CHE3935 Lecture 4 Quantum Mechanical Simulation Methods Continued 1 OUTLINE Review Introduction to CPMD MD and ensembles The functionals of density functional theory Return to ab initio methods Binding
CHE3935 Lecture 4 Quantum Mechanical Simulation Methods Continued 1 OUTLINE Review Introduction to CPMD MD and ensembles The functionals of density functional theory Return to ab initio methods Binding
DMDW: A set of tools to calculate Debye-Waller factors and other related quantities using dynamical matrices.
 DMDW: A set of tools to calculate Debye-Waller factors and other related quantities using dynamical matrices. DMDW is a set of tools developed to calculate Debye-Waller (DW) factors and other related quantities
DMDW: A set of tools to calculate Debye-Waller factors and other related quantities using dynamical matrices. DMDW is a set of tools developed to calculate Debye-Waller (DW) factors and other related quantities
