Covariance decomposition in multivariate analysis
|
|
|
- Regina Sharp
- 5 years ago
- Views:
Transcription
1 Covariance decomposition in multivariate analysis By BEATRIX JONES 1 & MIKE WEST Institute of Statistics and Decision Sciences Duke University, Durham NC {trix,mw}@stat.duke.edu Summary The covariance between two variables in a multivariate Gaussian distribution is decomposed into a sum of path weights for all paths connecting the two variables in an undirected graph. These weights are useful in determining which variables are important in mediating correlation between the two path endpoints. The decomposition arises in undirected Gaussian graphical models and does not require or involve any assumptions of causality. The novel theory underlying this covariance decomposition is derived using basic linear algebra. The resulting weights can be interpreted as involving two components: a measure of the strengths of the relationships between the variables along the path, conditioning on all other variables; and a function of the entropy lost in that conditioning. The decomposition is feasible for very large numbers of variables if the corresponding precision matrix is sparse, a circumstance that arises in currently topical examples such as gene expression studies in functional genomics. Additional computational efficiencies are possible when the undirected graph is derived from an acyclic directed graph. Some key words: Cofactor expansions, Concentration graph, Conditional independence, Covariance Selection, Path analysis 1 From summer the mailing address for the first author will be: Beatrix Jones, Institute of Information and Mathematical Sciences, Massey University, Private Bag , North Shore Mail Centre, Auckland, New Zealand 1
2 1 Introduction Graphical models over undirected graphs (Lauritzen 1996) are increasingly used to exhibit the conditional (in)dependence structures in multivariate distributions, and advances in computational techniques are providing increased access to methods associated with graphical modelling in problems of increasing complexity and dimension (Dobra et al. 2004, Rich et al. 2004). Undirected Gaussian graphical models, where edges in the graph correspond to non-zero entries in the precision matrix, are a special case of particular interest (Dempster 1972, Giudici 1996, Jones et al. 2004). In exploring and aiming to interpret relationships exhibited in such multivariate analyses, we are often faced with questions about how subsets of possibly many variables play roles in - or mediate - observed relationships between pairs of variables. In this communication, we present a novel representation of the covariance between two random variables in a multivariate Gaussian distribution in terms of sums of components related to individual paths between the variables in an underlying graphical model. This decomposition directly defines path weights that contribute to the overall pairwise association, and so highlights - and quantifies - the nature of the roles played by intervening variables along multiple such paths. The covariance decomposition we present is derived using basic linear algebra, relying only on analytical expressions using cofactors for matrix determinants and inverses. The linear algebra literature has used similar approaches to, for instance, provide alternate expressions for the determinant of a matrix (Johnson et al. 1994); however, to our knowledge they have not been used to aid in interpretation of covariance patterns and associations in graphical models. We envision this decomposition to provide an alternative to the venerable results on path coefficients introduced by Wright (1921), which provide covariance decomposition along paths between two variables in a context of directional modelling. Originally developed in the context of quantitative genetics, Wright s results are now used in a variety of fields to examine the relative importance of direct and indirect variate relationships (e.g. sociology, Duncan, 1966, and ecology, Grace & Pugesek, 1998). However, use of the method of path coefficients is predicated on an a priori understanding or hypothesis of directed causal relationships between the variables (Wright, 1934). Application relies on successive marginalisation over variables without descendants, so altering the direction of one or more edges produces a different decomposition, even if the corresponding covariance matrix is unaltered. In contrast, our new theory of covariance decomposition arises directly from the covariance matrix and is therefore not reliant on directionality; consequently, it is suitable for use in exploratory analysis of (roughly Gaussian) multivariate data, where the interest lies in generating insights into the nature of contributions to an estimated or inferred pairwise association from potentially many intervening variables. The implied weight on any single path between two variables can be represented, and interpreted, in terms of the product of two components: one component is a a measure of the strength of the relationships between the variables along the path in the distribution conditional on all other 2
3 variables; the second component is a simple function of the entropy lost in the conditioning. As in Wright s directional path decompositions, these path weights will help prioritize which inter-variate relationships are most worthy of additional investigation. For example, the precision matrix of many gene expression levels may be inferred from an observational study of human patients; the highest weighted path between a pair of genes of interest may then generate insight into underlying biological pathways, or suggest which additional experiments in cell lines or model systems will be most fruitful for elucidating the major relationships between genes in the system. 2 Covariance Decomposition Over Paths Consider an n dimensional multivariate Gaussian distribution with a finite and non-singular covariance matrix Σ. It is convenient to work in terms of the inverse of Σ, the precision matrix Ω. In a graphical model over an undirected graph, the n nodes of the graph represent variables and lack of an edge between any two nodes represents the fact that the corresponding two variables are conditionally independent given all other variables. In the case of a Gaussian distribution, conditional independence of two variables implies, and is implied by, a zero entry in the corresponding element of Ω (Dempster 1972); i.e., the pattern of non-zero off-diagonal elements of the precision matrix is that of the adjacency matrix of the graph. The elements of the covariance matrix Σ can be written in terms of the cofactors Ω ij of Ω: Ω 11 Ω 21 Ω n1 1 Ω 12 Ω 22 Ω n2 Σ = det(ω)..... Ω 1n Ω 2n Ω nn Thus σ xy = Ω xy / det(ω). Recalling that Ω ij is ( 1) i+j times the determinant of the matrix resulting from eliminating the ith row and jth column from Ω, we see that the expression can be further expanded by expressing Ω xy using a cofactor expansion along the column corresponding to the variable x. Let Ω xy,ix be the appropriate power of 1 times the determinant of the matrix where we have eliminated the row corresponding to variable x, the column corresponding to y, and then the row corresponding to variable i and the column corresponding to x. Then σ xy = 1 det(ω) (ω 1xΩ xy,1x + ω 2x Ω xy,2x ω nx Ω xy,2(n 1)x ). Terms in this sum will drop out for cases with ω ix = 0. In the Gaussian case we are considering, this corresponds exactly to cases where there is no edge between variables i and x in the corresponding graphical model. Note that there is no 3
4 term with ω xx because it has already been eliminated from Ω in producing Ω xy. The non-zero terms can then be expanded further in terms of adjacent edges. For example, for the first term we can consider paths through node 1: ω 1x Ω xy,1x = ω 1x ω 21 Ω xy,1x, ω 1x ω n1 Ω xy,1x,n1, again letting Ω xy,ix,ji represent the determinant of the matrix where we have successively eliminated the row and column corresponding to variables x and y, i and x, and finally j and i, and also subsuming the signs in this term. Imagine continuing the expansion along adjacent edges until either (1) an edge to y is reached, or (2) there are no more adjacent edges. In the latter case, the determinant of the remaining matrix is zero, and the term does not contribute to σ xy. The first case constitutes a path from x to y; the remaining cofactor term is the determinant of Ω with the variables in the path removed. The contribution of a path thorough variables P = p 1, p 2... p m is then ( 1) m+1 ω p2p 1 ω p3p 2... ω pmp m 1 det(ω P ) det(ω), (1) where Ω P is the precision matrix with only the rows and columns for variables not in P. The fact that relevant power of 1 corresponds to the number of edges in the path can be seen by considering the matrix where x is the first variable listed, y is the mth variable listed, and the intervening variables are listed in the order they are traversed from x to y, with variables not in the path listed after m. For this case, when we eliminate the row corresponding to x (the first) and column corresponding to y (the m th ) we introduce a factor of ( 1) m+1. The rest of the eliminations involve the first row and first column, so the sign is not altered. We see that this argument generalizes to any ordering of the variables by noting that changing the index of a particular variable involves two elementary row operations, a row swap and a column swap, and thus does not alter the relevant determinant. Each ω ij has a sign opposite to that of the associated partial regression coefficients, so the path weight has the same sign as the product of the partial regression coefficients associated with the edges. An alternate representation of (1) is useful in its interpretation. It can be written S V where S = ( 1) m+1 ω p 2p 1 ω p3p 2... ω pmp m 1 det(ω P ) V = det(ω P ) det(ω P ). det(ω) S is the path weight in the reduced system obtained by conditioning on all variables not in the path. Unless the path variables are independent of all other variables in the system, this conditioning will reduce the variance of the path variables. V reflects the increase in the covariance resulting from restoring this 4
5 variance. V = exp( 2 H), where H is the entropy lost in the conditioning: H = 1 ( ) 2 log det(σp P ) det(σ P ) = 1 ( ) 2 log det(ω). det(ω P ) det(ω P ) H is always non-positive, so exp( 2 H) 1. 3 Examples Two examples provide useful illustration of the decomposition. Note that the figures of graphs in these examples are produced using GraphExplore, Wang & Dobra An Illustrative Synthetic Example Consider the following simple four variable example. Variables x, y, a and b have precision matrix given in Table 1, corresponding to the graph in Figure 1. Table 1: Precision matrix for the variables depicted in Figure 1. x a b y x a b y There are three paths from a to b, with weights given in Table 2. These weights indicate that the direct path between a and b does not contribute as much to the correlation as the indirect paths via x and y. The direct path also has sign opposite to the marginal correlation of a and b. We will call paths of this type moderating paths, and paths with the same sign as the marginal correlation mediating paths. We do not use the terms mediating and moderating to imply any causal relationship; any causal relationship compatible with the undirected graph is possible, possibly including unobserved variables. Nevertheless, the breakdown of the path weights implies that study of the mediating variables x and y, rather than just the direct interactions between a and b, are important in understanding the bulk of the correlation between a and b. Now consider the four paths from x to y in this same graph, with weights also given in Table 2. Here we see that both a and b are each utilized in one mediating path, and two moderating paths of smaller magnitude. In such situations it may be more useful to classify variables as mediating or moderating based on the sum 5
6 Figure 1: Undirected graph corresponding to the precision matrix in Table 1 Table 2: Path weights for all paths between a and b, and between x and y. a b path weight x y path weight a b x a y a x b x b y a y b x a b y x b a y Total (σ ab ) Total (σ xy ) of the paths through them, rather examining individual paths. Here a has this sum equal to and b has 0.091, so both are mediating variables. Similarly, one can think of the importance of a variable to the covariance in question as the sum of the path weights for paths through that variable. 3.2 An Illustrative Example from Gene Expression Analysis A larger example arises from exploration of aspects of large-scale graphical models developed for analyses of gene expression data in brain cancer genomics. This utilises a (sparse) acyclic directed graph, and the associated precision matrix, fitted to the gene expression data from Rich et al. (2004). This data consists of 8408 variables, each representing the expression level of a particular gene in tumor tissue. The variables are observed in 41 glioblastoma patients, and have been transformed so that they have roughly a multivariate Gaussian distribution. The graph was fit using the tools provided in Dobra (2004). A subgraph of the system is shown in Figure 2. The directed graph can be seen by considering the edges with direction indicated. The edges without direction were added during the conversion to an undirected graph, which requires linking the parents of each node (called moralisation ). It is appealing to work with the undirected 6
7 graph, because the inferred covariance matrix is consistent with many acyclic directed graphs in addition to the one depicted, and there is no a priori preference among these. Suppose our interest is in whether the expression of gene KIAA0913, which encodes a protein of unknown function, is worth investigating as a potential intermediary between the genes ZBP1 and TGM1. The expression levels for ZBP1 and TGM1 have covariance We will consider all paths between ZBP1 and TGM1. Figure 2: Subgraph of an 8408 node graph representing gene expression in glioblastomas. In working with this large example, two observations led to important efficiencies in accomplishing the computation. First, since Ω is very large compared to the number of variables in the paths, computation of the ratio between det(ω P ) and det(ω) is much more stable if it is performed using the identity ( ) A B det = det(a) det(d CA 1 B), (2) C D with A = Ω P and D = Ω P. Note that A 1 need not be computed; B has relatively few columns, so solving for each column of A 1 B is easier than computation of the full inverse, especially as both matrices are sparse in this example. Computations of the path weights in Table 3 was accomplished in Matlab in less than one minute. Second, the structure of the underlying acyclic directed graph can be used to eliminate the need to consider some paths. While the moralised edges added 7
8 between parents of the same node correspond to non-zero elements in the inverse covariance matrix, they do not represent marginal correlation. When all paths involving the moralised edge could replace that edge with the structures through the common offspring implied by the moralisation, this set of paths through the endpoints of the moralised edge has total weight zero. For example, in Figure 2, the nodes (genes) ZBP1-PAIP1 3-TGM1 and ZBP1-PAIP1 3- LOC TGM1 have path weights of the exactly the same magnitude and opposite sign. These sets of canceling paths can be removed from consideration before computations begin, resulting in major savings. The ZBP1-PAIP1 3- TGM1 path is included in Figure 2 just as an example: if all paths that occurred in canceling sets were included, there would be more than 60 additional paths (and the associated nodes) in the drawing. Table 3: Paths contributing to the covariance of ZBP1 and TGM1. All paths start at ZBP1 and end at TGM1, so these endpoints have been omitted. Path via S V Weight F F2-KCNJ MGC31957-KIAA0913-DIPA 1-KCNJ MGC31957-KIAA0913-DIPA 1-KCNJ12-F Total 0.68 Only four paths (all shown) actually contribute to the covariance. Their path weights are given in Table 3; the path endpoints, ZBP1 and TGM1, are common to all and have been omitted. Note that not all paths involving moralised edges can be disregarded; for example, ZBP1-F2-KCNJ12-TGM1 is an important moderating path. We also see that the paths through KIAA0913 do make an important contribution, accounting for about 40% of the covariance between ZBP1 and TGM1. Finally, Table 3 also gives the factorization of the path weights into S and V. Much of the variance of the path variables is explained by the over 8,000 additional variables in the system, so V has a major impact on the path weights. Acknowledgments Partial support for the research described here was provided by the Keck Foundation through the Keck Center for Neurooncogenomics at Duke University, and the National Science Foundation under grant DMS We would also like to acknowledge Carlos Carvalho for helpful conversations, and Adrian Dobra and Quanli Wang for software for graphical model fitting and visualisation. 8
9 References Dempster, A. (1972). Covariance selection. Biometrics 28, Dobra, A. (2004). Hdbcs: Bayesian covariance selection in high dimensions. adobra/hdbcs.html. Dobra, A., Hans, C., Jones, B., Nevins, J. & West, M. (2004). Sparse graphical models for exploring gene expression data. J. Mult. Anal. -,. Duncan, O. (1966). Path analysis: Sociological examples. Am. J. of Sociol. 72, Grace, J. & Pugesek, B. (1998). On the use of path analysis and related procedures for the investigation of ecological problems. Am. Nat. 152, Guidici, P. (1996). Learning in graphical Gaussian models. In Bayesian Statistics 5, J. M. Bernado, J. O. Berger, A. P. Dawid & A. F. M. Smith, eds. Oxford Univeristy Press, pp Johnson, C., Olesky, D., Tsatsomeros, M. & van den Dressche, P. (1994). An identity for the determinant. Linear Multilinear A. 36, Jones, B., Carvalho, C., Dobra, A., Hans, C., Carter, C. & West, M. (2004). Experiments in stochastic computation for high-dimensional graphical models. ISDS Discussion Paper, Duke University (submitted for publication). Lauritzen, S. L. (1996). Graphical Models. Clarendon Press, Oxford. Rich, J., Hans, C., Jones, B., Dobra, A., Dressman, H., Rashed, A., McClendon, R., Nevins, J., Bigner, D. & West, M. (2004). Predictive gene expression profiling and graphical association studies in glioblastoma survival. Submitted for publication. Wang, Q. & Dobra, A. (2004). Graphexplore: A software tool for graph visualization. Wright, S. (1921). Correlation and causation. J. Agr. Res. 20, Wright, S. (1934). The method of path coefficients. Ann. Math. Stat. 5,
Comment on Article by Scutari
 Bayesian Analysis (2013) 8, Number 3, pp. 543 548 Comment on Article by Scutari Hao Wang Scutari s paper studies properties of the distribution of graphs ppgq. This is an interesting angle because it differs
Bayesian Analysis (2013) 8, Number 3, pp. 543 548 Comment on Article by Scutari Hao Wang Scutari s paper studies properties of the distribution of graphs ppgq. This is an interesting angle because it differs
Dynamic Matrix-Variate Graphical Models A Synopsis 1
 Proc. Valencia / ISBA 8th World Meeting on Bayesian Statistics Benidorm (Alicante, Spain), June 1st 6th, 2006 Dynamic Matrix-Variate Graphical Models A Synopsis 1 Carlos M. Carvalho & Mike West ISDS, Duke
Proc. Valencia / ISBA 8th World Meeting on Bayesian Statistics Benidorm (Alicante, Spain), June 1st 6th, 2006 Dynamic Matrix-Variate Graphical Models A Synopsis 1 Carlos M. Carvalho & Mike West ISDS, Duke
Directed acyclic graphs and the use of linear mixed models
 Directed acyclic graphs and the use of linear mixed models Siem H. Heisterkamp 1,2 1 Groningen Bioinformatics Centre, University of Groningen 2 Biostatistics and Research Decision Sciences (BARDS), MSD,
Directed acyclic graphs and the use of linear mixed models Siem H. Heisterkamp 1,2 1 Groningen Bioinformatics Centre, University of Groningen 2 Biostatistics and Research Decision Sciences (BARDS), MSD,
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 3 Linear
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 3 Linear
Computational Genomics
 Computational Genomics http://www.cs.cmu.edu/~02710 Introduction to probability, statistics and algorithms (brief) intro to probability Basic notations Random variable - referring to an element / event
Computational Genomics http://www.cs.cmu.edu/~02710 Introduction to probability, statistics and algorithms (brief) intro to probability Basic notations Random variable - referring to an element / event
Bayesian Analysis of Matrix Normal Graphical Models
 Bayesian Analysis of Matrix Normal Graphical Models By HAO WANG and MIKE WEST Department of Statistical Science, Duke University, Durham, North Carolina 27708-0251, U.S.A. www.stat.duke.edu {hao, mw}@stat.duke.edu
Bayesian Analysis of Matrix Normal Graphical Models By HAO WANG and MIKE WEST Department of Statistical Science, Duke University, Durham, North Carolina 27708-0251, U.S.A. www.stat.duke.edu {hao, mw}@stat.duke.edu
MATH 829: Introduction to Data Mining and Analysis Graphical Models I
 MATH 829: Introduction to Data Mining and Analysis Graphical Models I Dominique Guillot Departments of Mathematical Sciences University of Delaware May 2, 2016 1/12 Independence and conditional independence:
MATH 829: Introduction to Data Mining and Analysis Graphical Models I Dominique Guillot Departments of Mathematical Sciences University of Delaware May 2, 2016 1/12 Independence and conditional independence:
Introduction to Graphical Models
 Introduction to Graphical Models STA 345: Multivariate Analysis Department of Statistical Science Duke University, Durham, NC, USA Robert L. Wolpert 1 Conditional Dependence Two real-valued or vector-valued
Introduction to Graphical Models STA 345: Multivariate Analysis Department of Statistical Science Duke University, Durham, NC, USA Robert L. Wolpert 1 Conditional Dependence Two real-valued or vector-valued
Chris Bishop s PRML Ch. 8: Graphical Models
 Chris Bishop s PRML Ch. 8: Graphical Models January 24, 2008 Introduction Visualize the structure of a probabilistic model Design and motivate new models Insights into the model s properties, in particular
Chris Bishop s PRML Ch. 8: Graphical Models January 24, 2008 Introduction Visualize the structure of a probabilistic model Design and motivate new models Insights into the model s properties, in particular
Evaluating Determinants by Row Reduction
 Evaluating Determinants by Row Reduction MATH 322, Linear Algebra I J. Robert Buchanan Department of Mathematics Spring 2015 Objectives Reduce a matrix to row echelon form and evaluate its determinant.
Evaluating Determinants by Row Reduction MATH 322, Linear Algebra I J. Robert Buchanan Department of Mathematics Spring 2015 Objectives Reduce a matrix to row echelon form and evaluate its determinant.
Learning from Sensor Data: Set II. Behnaam Aazhang J.S. Abercombie Professor Electrical and Computer Engineering Rice University
 Learning from Sensor Data: Set II Behnaam Aazhang J.S. Abercombie Professor Electrical and Computer Engineering Rice University 1 6. Data Representation The approach for learning from data Probabilistic
Learning from Sensor Data: Set II Behnaam Aazhang J.S. Abercombie Professor Electrical and Computer Engineering Rice University 1 6. Data Representation The approach for learning from data Probabilistic
MATH 2050 Assignment 8 Fall [10] 1. Find the determinant by reducing to triangular form for the following matrices.
![MATH 2050 Assignment 8 Fall [10] 1. Find the determinant by reducing to triangular form for the following matrices. MATH 2050 Assignment 8 Fall [10] 1. Find the determinant by reducing to triangular form for the following matrices.](/thumbs/95/123052681.jpg) MATH 2050 Assignment 8 Fall 2016 [10] 1. Find the determinant by reducing to triangular form for the following matrices. 0 1 2 (a) A = 2 1 4. ANS: We perform the Gaussian Elimination on A by the following
MATH 2050 Assignment 8 Fall 2016 [10] 1. Find the determinant by reducing to triangular form for the following matrices. 0 1 2 (a) A = 2 1 4. ANS: We perform the Gaussian Elimination on A by the following
Conditional Independence
 Conditional Independence Sargur Srihari srihari@cedar.buffalo.edu 1 Conditional Independence Topics 1. What is Conditional Independence? Factorization of probability distribution into marginals 2. Why
Conditional Independence Sargur Srihari srihari@cedar.buffalo.edu 1 Conditional Independence Topics 1. What is Conditional Independence? Factorization of probability distribution into marginals 2. Why
Computational Genomics. Systems biology. Putting it together: Data integration using graphical models
 02-710 Computational Genomics Systems biology Putting it together: Data integration using graphical models High throughput data So far in this class we discussed several different types of high throughput
02-710 Computational Genomics Systems biology Putting it together: Data integration using graphical models High throughput data So far in this class we discussed several different types of high throughput
Algebra & Trig. I. For example, the system. x y 2 z. may be represented by the augmented matrix
 Algebra & Trig. I 8.1 Matrix Solutions to Linear Systems A matrix is a rectangular array of elements. o An array is a systematic arrangement of numbers or symbols in rows and columns. Matrices (the plural
Algebra & Trig. I 8.1 Matrix Solutions to Linear Systems A matrix is a rectangular array of elements. o An array is a systematic arrangement of numbers or symbols in rows and columns. Matrices (the plural
Learning in Bayesian Networks
 Learning in Bayesian Networks Florian Markowetz Max-Planck-Institute for Molecular Genetics Computational Molecular Biology Berlin Berlin: 20.06.2002 1 Overview 1. Bayesian Networks Stochastic Networks
Learning in Bayesian Networks Florian Markowetz Max-Planck-Institute for Molecular Genetics Computational Molecular Biology Berlin Berlin: 20.06.2002 1 Overview 1. Bayesian Networks Stochastic Networks
CPSC 540: Machine Learning
 CPSC 540: Machine Learning Undirected Graphical Models Mark Schmidt University of British Columbia Winter 2016 Admin Assignment 3: 2 late days to hand it in today, Thursday is final day. Assignment 4:
CPSC 540: Machine Learning Undirected Graphical Models Mark Schmidt University of British Columbia Winter 2016 Admin Assignment 3: 2 late days to hand it in today, Thursday is final day. Assignment 4:
A MCMC Approach for Learning the Structure of Gaussian Acyclic Directed Mixed Graphs
 A MCMC Approach for Learning the Structure of Gaussian Acyclic Directed Mixed Graphs Ricardo Silva Abstract Graphical models are widely used to encode conditional independence constraints and causal assumptions,
A MCMC Approach for Learning the Structure of Gaussian Acyclic Directed Mixed Graphs Ricardo Silva Abstract Graphical models are widely used to encode conditional independence constraints and causal assumptions,
Introduction to Probabilistic Graphical Models
 Introduction to Probabilistic Graphical Models Sargur Srihari srihari@cedar.buffalo.edu 1 Topics 1. What are probabilistic graphical models (PGMs) 2. Use of PGMs Engineering and AI 3. Directionality in
Introduction to Probabilistic Graphical Models Sargur Srihari srihari@cedar.buffalo.edu 1 Topics 1. What are probabilistic graphical models (PGMs) 2. Use of PGMs Engineering and AI 3. Directionality in
Instrumental Sets. Carlos Brito. 1 Introduction. 2 The Identification Problem
 17 Instrumental Sets Carlos Brito 1 Introduction The research of Judea Pearl in the area of causality has been very much acclaimed. Here we highlight his contributions for the use of graphical languages
17 Instrumental Sets Carlos Brito 1 Introduction The research of Judea Pearl in the area of causality has been very much acclaimed. Here we highlight his contributions for the use of graphical languages
Math 240 Calculus III
 The Calculus III Summer 2015, Session II Wednesday, July 8, 2015 Agenda 1. of the determinant 2. determinants 3. of determinants What is the determinant? Yesterday: Ax = b has a unique solution when A
The Calculus III Summer 2015, Session II Wednesday, July 8, 2015 Agenda 1. of the determinant 2. determinants 3. of determinants What is the determinant? Yesterday: Ax = b has a unique solution when A
Chapter 17: Undirected Graphical Models
 Chapter 17: Undirected Graphical Models The Elements of Statistical Learning Biaobin Jiang Department of Biological Sciences Purdue University bjiang@purdue.edu October 30, 2014 Biaobin Jiang (Purdue)
Chapter 17: Undirected Graphical Models The Elements of Statistical Learning Biaobin Jiang Department of Biological Sciences Purdue University bjiang@purdue.edu October 30, 2014 Biaobin Jiang (Purdue)
Undirected Graphical Models
 Undirected Graphical Models 1 Conditional Independence Graphs Let G = (V, E) be an undirected graph with vertex set V and edge set E, and let A, B, and C be subsets of vertices. We say that C separates
Undirected Graphical Models 1 Conditional Independence Graphs Let G = (V, E) be an undirected graph with vertex set V and edge set E, and let A, B, and C be subsets of vertices. We say that C separates
1 Data Arrays and Decompositions
 1 Data Arrays and Decompositions 1.1 Variance Matrices and Eigenstructure Consider a p p positive definite and symmetric matrix V - a model parameter or a sample variance matrix. The eigenstructure is
1 Data Arrays and Decompositions 1.1 Variance Matrices and Eigenstructure Consider a p p positive definite and symmetric matrix V - a model parameter or a sample variance matrix. The eigenstructure is
Undirected Graphical Models
 Outline Hong Chang Institute of Computing Technology, Chinese Academy of Sciences Machine Learning Methods (Fall 2012) Outline Outline I 1 Introduction 2 Properties Properties 3 Generative vs. Conditional
Outline Hong Chang Institute of Computing Technology, Chinese Academy of Sciences Machine Learning Methods (Fall 2012) Outline Outline I 1 Introduction 2 Properties Properties 3 Generative vs. Conditional
1 Counting spanning trees: A determinantal formula
 Math 374 Matrix Tree Theorem Counting spanning trees: A determinantal formula Recall that a spanning tree of a graph G is a subgraph T so that T is a tree and V (G) = V (T ) Question How many distinct
Math 374 Matrix Tree Theorem Counting spanning trees: A determinantal formula Recall that a spanning tree of a graph G is a subgraph T so that T is a tree and V (G) = V (T ) Question How many distinct
10708 Graphical Models: Homework 2
 10708 Graphical Models: Homework 2 Due Monday, March 18, beginning of class Feburary 27, 2013 Instructions: There are five questions (one for extra credit) on this assignment. There is a problem involves
10708 Graphical Models: Homework 2 Due Monday, March 18, beginning of class Feburary 27, 2013 Instructions: There are five questions (one for extra credit) on this assignment. There is a problem involves
Machine Learning, Fall 2009: Midterm
 10-601 Machine Learning, Fall 009: Midterm Monday, November nd hours 1. Personal info: Name: Andrew account: E-mail address:. You are permitted two pages of notes and a calculator. Please turn off all
10-601 Machine Learning, Fall 009: Midterm Monday, November nd hours 1. Personal info: Name: Andrew account: E-mail address:. You are permitted two pages of notes and a calculator. Please turn off all
Decomposable and Directed Graphical Gaussian Models
 Decomposable Decomposable and Directed Graphical Gaussian Models Graphical Models and Inference, Lecture 13, Michaelmas Term 2009 November 26, 2009 Decomposable Definition Basic properties Wishart density
Decomposable Decomposable and Directed Graphical Gaussian Models Graphical Models and Inference, Lecture 13, Michaelmas Term 2009 November 26, 2009 Decomposable Definition Basic properties Wishart density
MATH Topics in Applied Mathematics Lecture 12: Evaluation of determinants. Cross product.
 MATH 311-504 Topics in Applied Mathematics Lecture 12: Evaluation of determinants. Cross product. Determinant is a scalar assigned to each square matrix. Notation. The determinant of a matrix A = (a ij
MATH 311-504 Topics in Applied Mathematics Lecture 12: Evaluation of determinants. Cross product. Determinant is a scalar assigned to each square matrix. Notation. The determinant of a matrix A = (a ij
Graphical Models and Independence Models
 Graphical Models and Independence Models Yunshu Liu ASPITRG Research Group 2014-03-04 References: [1]. Steffen Lauritzen, Graphical Models, Oxford University Press, 1996 [2]. Christopher M. Bishop, Pattern
Graphical Models and Independence Models Yunshu Liu ASPITRG Research Group 2014-03-04 References: [1]. Steffen Lauritzen, Graphical Models, Oxford University Press, 1996 [2]. Christopher M. Bishop, Pattern
4. Determinants.
 4. Determinants 4.1. Determinants; Cofactor Expansion Determinants of 2 2 and 3 3 Matrices 2 2 determinant 4.1. Determinants; Cofactor Expansion Determinants of 2 2 and 3 3 Matrices 3 3 determinant 4.1.
4. Determinants 4.1. Determinants; Cofactor Expansion Determinants of 2 2 and 3 3 Matrices 2 2 determinant 4.1. Determinants; Cofactor Expansion Determinants of 2 2 and 3 3 Matrices 3 3 determinant 4.1.
Effects of dependence in high-dimensional multiple testing problems. Kyung In Kim and Mark van de Wiel
 Effects of dependence in high-dimensional multiple testing problems Kyung In Kim and Mark van de Wiel Department of Mathematics, Vrije Universiteit Amsterdam. Contents 1. High-dimensional multiple testing
Effects of dependence in high-dimensional multiple testing problems Kyung In Kim and Mark van de Wiel Department of Mathematics, Vrije Universiteit Amsterdam. Contents 1. High-dimensional multiple testing
Bayesian networks (1) Lirong Xia
 Bayesian networks (1) Lirong Xia Random variables and joint distributions Ø A random variable is a variable with a domain Random variables: capital letters, e.g. W, D, L values: small letters, e.g. w,
Bayesian networks (1) Lirong Xia Random variables and joint distributions Ø A random variable is a variable with a domain Random variables: capital letters, e.g. W, D, L values: small letters, e.g. w,
Directed Graphical Models
 CS 2750: Machine Learning Directed Graphical Models Prof. Adriana Kovashka University of Pittsburgh March 28, 2017 Graphical Models If no assumption of independence is made, must estimate an exponential
CS 2750: Machine Learning Directed Graphical Models Prof. Adriana Kovashka University of Pittsburgh March 28, 2017 Graphical Models If no assumption of independence is made, must estimate an exponential
Lemma 8: Suppose the N by N matrix A has the following block upper triangular form:
 17 4 Determinants and the Inverse of a Square Matrix In this section, we are going to use our knowledge of determinants and their properties to derive an explicit formula for the inverse of a square matrix
17 4 Determinants and the Inverse of a Square Matrix In this section, we are going to use our knowledge of determinants and their properties to derive an explicit formula for the inverse of a square matrix
THE VINE COPULA METHOD FOR REPRESENTING HIGH DIMENSIONAL DEPENDENT DISTRIBUTIONS: APPLICATION TO CONTINUOUS BELIEF NETS
 Proceedings of the 00 Winter Simulation Conference E. Yücesan, C.-H. Chen, J. L. Snowdon, and J. M. Charnes, eds. THE VINE COPULA METHOD FOR REPRESENTING HIGH DIMENSIONAL DEPENDENT DISTRIBUTIONS: APPLICATION
Proceedings of the 00 Winter Simulation Conference E. Yücesan, C.-H. Chen, J. L. Snowdon, and J. M. Charnes, eds. THE VINE COPULA METHOD FOR REPRESENTING HIGH DIMENSIONAL DEPENDENT DISTRIBUTIONS: APPLICATION
1. what conditional independencies are implied by the graph. 2. whether these independecies correspond to the probability distribution
 NETWORK ANALYSIS Lourens Waldorp PROBABILITY AND GRAPHS The objective is to obtain a correspondence between the intuitive pictures (graphs) of variables of interest and the probability distributions of
NETWORK ANALYSIS Lourens Waldorp PROBABILITY AND GRAPHS The objective is to obtain a correspondence between the intuitive pictures (graphs) of variables of interest and the probability distributions of
Markovian Combination of Decomposable Model Structures: MCMoSt
 Markovian Combination 1/45 London Math Society Durham Symposium on Mathematical Aspects of Graphical Models. June 30 - July, 2008 Markovian Combination of Decomposable Model Structures: MCMoSt Sung-Ho
Markovian Combination 1/45 London Math Society Durham Symposium on Mathematical Aspects of Graphical Models. June 30 - July, 2008 Markovian Combination of Decomposable Model Structures: MCMoSt Sung-Ho
Based on slides by Richard Zemel
 CSC 412/2506 Winter 2018 Probabilistic Learning and Reasoning Lecture 3: Directed Graphical Models and Latent Variables Based on slides by Richard Zemel Learning outcomes What aspects of a model can we
CSC 412/2506 Winter 2018 Probabilistic Learning and Reasoning Lecture 3: Directed Graphical Models and Latent Variables Based on slides by Richard Zemel Learning outcomes What aspects of a model can we
Bayesian (conditionally) conjugate inference for discrete data models. Jon Forster (University of Southampton)
 Bayesian (conditionally) conjugate inference for discrete data models Jon Forster (University of Southampton) with Mark Grigsby (Procter and Gamble?) Emily Webb (Institute of Cancer Research) Table 1:
Bayesian (conditionally) conjugate inference for discrete data models Jon Forster (University of Southampton) with Mark Grigsby (Procter and Gamble?) Emily Webb (Institute of Cancer Research) Table 1:
CSC 412 (Lecture 4): Undirected Graphical Models
 CSC 412 (Lecture 4): Undirected Graphical Models Raquel Urtasun University of Toronto Feb 2, 2016 R Urtasun (UofT) CSC 412 Feb 2, 2016 1 / 37 Today Undirected Graphical Models: Semantics of the graph:
CSC 412 (Lecture 4): Undirected Graphical Models Raquel Urtasun University of Toronto Feb 2, 2016 R Urtasun (UofT) CSC 412 Feb 2, 2016 1 / 37 Today Undirected Graphical Models: Semantics of the graph:
Probabilistic Graphical Models
 School of Computer Science Probabilistic Graphical Models Gaussian graphical models and Ising models: modeling networks Eric Xing Lecture 0, February 7, 04 Reading: See class website Eric Xing @ CMU, 005-04
School of Computer Science Probabilistic Graphical Models Gaussian graphical models and Ising models: modeling networks Eric Xing Lecture 0, February 7, 04 Reading: See class website Eric Xing @ CMU, 005-04
Sparse Permutation Invariant Covariance Estimation: Motivation, Background and Key Results
 Sparse Permutation Invariant Covariance Estimation: Motivation, Background and Key Results David Prince Biostat 572 dprince3@uw.edu April 19, 2012 David Prince (UW) SPICE April 19, 2012 1 / 11 Electronic
Sparse Permutation Invariant Covariance Estimation: Motivation, Background and Key Results David Prince Biostat 572 dprince3@uw.edu April 19, 2012 David Prince (UW) SPICE April 19, 2012 1 / 11 Electronic
Review of Basic Concepts in Linear Algebra
 Review of Basic Concepts in Linear Algebra Grady B Wright Department of Mathematics Boise State University September 7, 2017 Math 565 Linear Algebra Review September 7, 2017 1 / 40 Numerical Linear Algebra
Review of Basic Concepts in Linear Algebra Grady B Wright Department of Mathematics Boise State University September 7, 2017 Math 565 Linear Algebra Review September 7, 2017 1 / 40 Numerical Linear Algebra
Gaussian Graphical Models and Graphical Lasso
 ELE 538B: Sparsity, Structure and Inference Gaussian Graphical Models and Graphical Lasso Yuxin Chen Princeton University, Spring 2017 Multivariate Gaussians Consider a random vector x N (0, Σ) with pdf
ELE 538B: Sparsity, Structure and Inference Gaussian Graphical Models and Graphical Lasso Yuxin Chen Princeton University, Spring 2017 Multivariate Gaussians Consider a random vector x N (0, Σ) with pdf
Lecture 4 October 18th
 Directed and undirected graphical models Fall 2017 Lecture 4 October 18th Lecturer: Guillaume Obozinski Scribe: In this lecture, we will assume that all random variables are discrete, to keep notations
Directed and undirected graphical models Fall 2017 Lecture 4 October 18th Lecturer: Guillaume Obozinski Scribe: In this lecture, we will assume that all random variables are discrete, to keep notations
Applying Bayesian networks in the game of Minesweeper
 Applying Bayesian networks in the game of Minesweeper Marta Vomlelová Faculty of Mathematics and Physics Charles University in Prague http://kti.mff.cuni.cz/~marta/ Jiří Vomlel Institute of Information
Applying Bayesian networks in the game of Minesweeper Marta Vomlelová Faculty of Mathematics and Physics Charles University in Prague http://kti.mff.cuni.cz/~marta/ Jiří Vomlel Institute of Information
Bayesian Inference of Multiple Gaussian Graphical Models
 Bayesian Inference of Multiple Gaussian Graphical Models Christine Peterson,, Francesco Stingo, and Marina Vannucci February 18, 2014 Abstract In this paper, we propose a Bayesian approach to inference
Bayesian Inference of Multiple Gaussian Graphical Models Christine Peterson,, Francesco Stingo, and Marina Vannucci February 18, 2014 Abstract In this paper, we propose a Bayesian approach to inference
Causal Effect Identification in Alternative Acyclic Directed Mixed Graphs
 Proceedings of Machine Learning Research vol 73:21-32, 2017 AMBN 2017 Causal Effect Identification in Alternative Acyclic Directed Mixed Graphs Jose M. Peña Linköping University Linköping (Sweden) jose.m.pena@liu.se
Proceedings of Machine Learning Research vol 73:21-32, 2017 AMBN 2017 Causal Effect Identification in Alternative Acyclic Directed Mixed Graphs Jose M. Peña Linköping University Linköping (Sweden) jose.m.pena@liu.se
Markov properties for directed graphs
 Graphical Models, Lecture 7, Michaelmas Term 2009 November 2, 2009 Definitions Structural relations among Markov properties Factorization G = (V, E) simple undirected graph; σ Say σ satisfies (P) the pairwise
Graphical Models, Lecture 7, Michaelmas Term 2009 November 2, 2009 Definitions Structural relations among Markov properties Factorization G = (V, E) simple undirected graph; σ Say σ satisfies (P) the pairwise
Graphical models and causality: Directed acyclic graphs (DAGs) and conditional (in)dependence
 Graphical models and causality: Directed acyclic graphs (DAGs) and conditional (in)dependence General overview Introduction Directed acyclic graphs (DAGs) and conditional independence DAGs and causal effects
Graphical models and causality: Directed acyclic graphs (DAGs) and conditional (in)dependence General overview Introduction Directed acyclic graphs (DAGs) and conditional independence DAGs and causal effects
SC705: Advanced Statistics Instructor: Natasha Sarkisian Class notes: Introduction to Structural Equation Modeling (SEM)
 SC705: Advanced Statistics Instructor: Natasha Sarkisian Class notes: Introduction to Structural Equation Modeling (SEM) SEM is a family of statistical techniques which builds upon multiple regression,
SC705: Advanced Statistics Instructor: Natasha Sarkisian Class notes: Introduction to Structural Equation Modeling (SEM) SEM is a family of statistical techniques which builds upon multiple regression,
Probabilistic Graphical Models
 2016 Robert Nowak Probabilistic Graphical Models 1 Introduction We have focused mainly on linear models for signals, in particular the subspace model x = Uθ, where U is a n k matrix and θ R k is a vector
2016 Robert Nowak Probabilistic Graphical Models 1 Introduction We have focused mainly on linear models for signals, in particular the subspace model x = Uθ, where U is a n k matrix and θ R k is a vector
On the adjacency matrix of a block graph
 On the adjacency matrix of a block graph R. B. Bapat Stat-Math Unit Indian Statistical Institute, Delhi 7-SJSS Marg, New Delhi 110 016, India. email: rbb@isid.ac.in Souvik Roy Economics and Planning Unit
On the adjacency matrix of a block graph R. B. Bapat Stat-Math Unit Indian Statistical Institute, Delhi 7-SJSS Marg, New Delhi 110 016, India. email: rbb@isid.ac.in Souvik Roy Economics and Planning Unit
Methods for Solving Linear Systems Part 2
 Methods for Solving Linear Systems Part 2 We have studied the properties of matrices and found out that there are more ways that we can solve Linear Systems. In Section 7.3, we learned that we can use
Methods for Solving Linear Systems Part 2 We have studied the properties of matrices and found out that there are more ways that we can solve Linear Systems. In Section 7.3, we learned that we can use
Chapter 16. Structured Probabilistic Models for Deep Learning
 Peng et al.: Deep Learning and Practice 1 Chapter 16 Structured Probabilistic Models for Deep Learning Peng et al.: Deep Learning and Practice 2 Structured Probabilistic Models way of using graphs to describe
Peng et al.: Deep Learning and Practice 1 Chapter 16 Structured Probabilistic Models for Deep Learning Peng et al.: Deep Learning and Practice 2 Structured Probabilistic Models way of using graphs to describe
Causal Discovery by Computer
 Causal Discovery by Computer Clark Glymour Carnegie Mellon University 1 Outline 1. A century of mistakes about causation and discovery: 1. Fisher 2. Yule 3. Spearman/Thurstone 2. Search for causes is statistical
Causal Discovery by Computer Clark Glymour Carnegie Mellon University 1 Outline 1. A century of mistakes about causation and discovery: 1. Fisher 2. Yule 3. Spearman/Thurstone 2. Search for causes is statistical
Probabilistic Graphical Networks: Definitions and Basic Results
 This document gives a cursory overview of Probabilistic Graphical Networks. The material has been gleaned from different sources. I make no claim to original authorship of this material. Bayesian Graphical
This document gives a cursory overview of Probabilistic Graphical Networks. The material has been gleaned from different sources. I make no claim to original authorship of this material. Bayesian Graphical
MAC Module 3 Determinants. Learning Objectives. Upon completing this module, you should be able to:
 MAC 2 Module Determinants Learning Objectives Upon completing this module, you should be able to:. Determine the minor, cofactor, and adjoint of a matrix. 2. Evaluate the determinant of a matrix by cofactor
MAC 2 Module Determinants Learning Objectives Upon completing this module, you should be able to:. Determine the minor, cofactor, and adjoint of a matrix. 2. Evaluate the determinant of a matrix by cofactor
Using Graphs to Describe Model Structure. Sargur N. Srihari
 Using Graphs to Describe Model Structure Sargur N. srihari@cedar.buffalo.edu 1 Topics in Structured PGMs for Deep Learning 0. Overview 1. Challenge of Unstructured Modeling 2. Using graphs to describe
Using Graphs to Describe Model Structure Sargur N. srihari@cedar.buffalo.edu 1 Topics in Structured PGMs for Deep Learning 0. Overview 1. Challenge of Unstructured Modeling 2. Using graphs to describe
Multivariate Gaussians. Sargur Srihari
 Multivariate Gaussians Sargur srihari@cedar.buffalo.edu 1 Topics 1. Multivariate Gaussian: Basic Parameterization 2. Covariance and Information Form 3. Operations on Gaussians 4. Independencies in Gaussians
Multivariate Gaussians Sargur srihari@cedar.buffalo.edu 1 Topics 1. Multivariate Gaussian: Basic Parameterization 2. Covariance and Information Form 3. Operations on Gaussians 4. Independencies in Gaussians
Linear Algebra: Lecture Notes. Dr Rachel Quinlan School of Mathematics, Statistics and Applied Mathematics NUI Galway
 Linear Algebra: Lecture Notes Dr Rachel Quinlan School of Mathematics, Statistics and Applied Mathematics NUI Galway November 6, 23 Contents Systems of Linear Equations 2 Introduction 2 2 Elementary Row
Linear Algebra: Lecture Notes Dr Rachel Quinlan School of Mathematics, Statistics and Applied Mathematics NUI Galway November 6, 23 Contents Systems of Linear Equations 2 Introduction 2 2 Elementary Row
Independencies. Undirected Graphical Models 2: Independencies. Independencies (Markov networks) Independencies (Bayesian Networks)
 (Bayesian Networks) Undirected Graphical Models 2: Use d-separation to read off independencies in a Bayesian network Takes a bit of effort! 1 2 (Markov networks) Use separation to determine independencies
(Bayesian Networks) Undirected Graphical Models 2: Use d-separation to read off independencies in a Bayesian network Takes a bit of effort! 1 2 (Markov networks) Use separation to determine independencies
Singular Value Decomposition and Principal Component Analysis (PCA) I
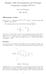 Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Bayes methods for categorical data. April 25, 2017
 Bayes methods for categorical data April 25, 2017 Motivation for joint probability models Increasing interest in high-dimensional data in broad applications Focus may be on prediction, variable selection,
Bayes methods for categorical data April 25, 2017 Motivation for joint probability models Increasing interest in high-dimensional data in broad applications Focus may be on prediction, variable selection,
9 Graphical modelling of dynamic relationships in multivariate time series
 9 Graphical modelling of dynamic relationships in multivariate time series Michael Eichler Institut für Angewandte Mathematik Universität Heidelberg Germany SUMMARY The identification and analysis of interactions
9 Graphical modelling of dynamic relationships in multivariate time series Michael Eichler Institut für Angewandte Mathematik Universität Heidelberg Germany SUMMARY The identification and analysis of interactions
Basic Concepts in Linear Algebra
 Basic Concepts in Linear Algebra Grady B Wright Department of Mathematics Boise State University February 2, 2015 Grady B Wright Linear Algebra Basics February 2, 2015 1 / 39 Numerical Linear Algebra Linear
Basic Concepts in Linear Algebra Grady B Wright Department of Mathematics Boise State University February 2, 2015 Grady B Wright Linear Algebra Basics February 2, 2015 1 / 39 Numerical Linear Algebra Linear
Properties of optimizations used in penalized Gaussian likelihood inverse covariance matrix estimation
 Properties of optimizations used in penalized Gaussian likelihood inverse covariance matrix estimation Adam J. Rothman School of Statistics University of Minnesota October 8, 2014, joint work with Liliana
Properties of optimizations used in penalized Gaussian likelihood inverse covariance matrix estimation Adam J. Rothman School of Statistics University of Minnesota October 8, 2014, joint work with Liliana
Data Mining 2018 Bayesian Networks (1)
 Data Mining 2018 Bayesian Networks (1) Ad Feelders Universiteit Utrecht Ad Feelders ( Universiteit Utrecht ) Data Mining 1 / 49 Do you like noodles? Do you like noodles? Race Gender Yes No Black Male 10
Data Mining 2018 Bayesian Networks (1) Ad Feelders Universiteit Utrecht Ad Feelders ( Universiteit Utrecht ) Data Mining 1 / 49 Do you like noodles? Do you like noodles? Race Gender Yes No Black Male 10
Elementary maths for GMT
 Elementary maths for GMT Linear Algebra Part 2: Matrices, Elimination and Determinant m n matrices The system of m linear equations in n variables x 1, x 2,, x n a 11 x 1 + a 12 x 2 + + a 1n x n = b 1
Elementary maths for GMT Linear Algebra Part 2: Matrices, Elimination and Determinant m n matrices The system of m linear equations in n variables x 1, x 2,, x n a 11 x 1 + a 12 x 2 + + a 1n x n = b 1
Expression Data Exploration: Association, Patterns, Factors & Regression Modelling
 Expression Data Exploration: Association, Patterns, Factors & Regression Modelling Exploring gene expression data Scale factors, median chip correlation on gene subsets for crude data quality investigation
Expression Data Exploration: Association, Patterns, Factors & Regression Modelling Exploring gene expression data Scale factors, median chip correlation on gene subsets for crude data quality investigation
Sparse Covariance Selection using Semidefinite Programming
 Sparse Covariance Selection using Semidefinite Programming A. d Aspremont ORFE, Princeton University Joint work with O. Banerjee, L. El Ghaoui & G. Natsoulis, U.C. Berkeley & Iconix Pharmaceuticals Support
Sparse Covariance Selection using Semidefinite Programming A. d Aspremont ORFE, Princeton University Joint work with O. Banerjee, L. El Ghaoui & G. Natsoulis, U.C. Berkeley & Iconix Pharmaceuticals Support
Gaussian Models
 Gaussian Models ddebarr@uw.edu 2016-04-28 Agenda Introduction Gaussian Discriminant Analysis Inference Linear Gaussian Systems The Wishart Distribution Inferring Parameters Introduction Gaussian Density
Gaussian Models ddebarr@uw.edu 2016-04-28 Agenda Introduction Gaussian Discriminant Analysis Inference Linear Gaussian Systems The Wishart Distribution Inferring Parameters Introduction Gaussian Density
Factor Analysis (10/2/13)
 STA561: Probabilistic machine learning Factor Analysis (10/2/13) Lecturer: Barbara Engelhardt Scribes: Li Zhu, Fan Li, Ni Guan Factor Analysis Factor analysis is related to the mixture models we have studied.
STA561: Probabilistic machine learning Factor Analysis (10/2/13) Lecturer: Barbara Engelhardt Scribes: Li Zhu, Fan Li, Ni Guan Factor Analysis Factor analysis is related to the mixture models we have studied.
Markov properties for graphical time series models
 Markov properties for graphical time series models Michael Eichler Universität Heidelberg Abstract This paper deals with the Markov properties of a new class of graphical time series models which focus
Markov properties for graphical time series models Michael Eichler Universität Heidelberg Abstract This paper deals with the Markov properties of a new class of graphical time series models which focus
Gaussian Elimination and Back Substitution
 Jim Lambers MAT 610 Summer Session 2009-10 Lecture 4 Notes These notes correspond to Sections 31 and 32 in the text Gaussian Elimination and Back Substitution The basic idea behind methods for solving
Jim Lambers MAT 610 Summer Session 2009-10 Lecture 4 Notes These notes correspond to Sections 31 and 32 in the text Gaussian Elimination and Back Substitution The basic idea behind methods for solving
Lab 2 Worksheet. Problems. Problem 1: Geometry and Linear Equations
 Lab 2 Worksheet Problems Problem : Geometry and Linear Equations Linear algebra is, first and foremost, the study of systems of linear equations. You are going to encounter linear systems frequently in
Lab 2 Worksheet Problems Problem : Geometry and Linear Equations Linear algebra is, first and foremost, the study of systems of linear equations. You are going to encounter linear systems frequently in
Gaussian Processes. Le Song. Machine Learning II: Advanced Topics CSE 8803ML, Spring 2012
 Gaussian Processes Le Song Machine Learning II: Advanced Topics CSE 8803ML, Spring 01 Pictorial view of embedding distribution Transform the entire distribution to expected features Feature space Feature
Gaussian Processes Le Song Machine Learning II: Advanced Topics CSE 8803ML, Spring 01 Pictorial view of embedding distribution Transform the entire distribution to expected features Feature space Feature
Lecture 5: Bayesian Network
 Lecture 5: Bayesian Network Topics of this lecture What is a Bayesian network? A simple example Formal definition of BN A slightly difficult example Learning of BN An example of learning Important topics
Lecture 5: Bayesian Network Topics of this lecture What is a Bayesian network? A simple example Formal definition of BN A slightly difficult example Learning of BN An example of learning Important topics
The Singular Acyclic Matrices of Even Order with a P-Set of Maximum Size
 Filomat 30:13 (016), 3403 3409 DOI 1098/FIL1613403D Published by Faculty of Sciences and Mathematics, University of Niš, Serbia Available at: http://wwwpmfniacrs/filomat The Singular Acyclic Matrices of
Filomat 30:13 (016), 3403 3409 DOI 1098/FIL1613403D Published by Faculty of Sciences and Mathematics, University of Niš, Serbia Available at: http://wwwpmfniacrs/filomat The Singular Acyclic Matrices of
Predicting causal effects in large-scale systems from observational data
 nature methods Predicting causal effects in large-scale systems from observational data Marloes H Maathuis 1, Diego Colombo 1, Markus Kalisch 1 & Peter Bühlmann 1,2 Supplementary figures and text: Supplementary
nature methods Predicting causal effects in large-scale systems from observational data Marloes H Maathuis 1, Diego Colombo 1, Markus Kalisch 1 & Peter Bühlmann 1,2 Supplementary figures and text: Supplementary
ENGR-1100 Introduction to Engineering Analysis. Lecture 21. Lecture outline
 ENGR-1100 Introduction to Engineering Analysis Lecture 21 Lecture outline Procedure (algorithm) for finding the inverse of invertible matrix. Investigate the system of linear equation and invertibility
ENGR-1100 Introduction to Engineering Analysis Lecture 21 Lecture outline Procedure (algorithm) for finding the inverse of invertible matrix. Investigate the system of linear equation and invertibility
A Practical Approach to Inferring Large Graphical Models from Sparse Microarray Data
 A Practical Approach to Inferring Large Graphical Models from Sparse Microarray Data Juliane Schäfer Department of Statistics, University of Munich Workshop: Practical Analysis of Gene Expression Data
A Practical Approach to Inferring Large Graphical Models from Sparse Microarray Data Juliane Schäfer Department of Statistics, University of Munich Workshop: Practical Analysis of Gene Expression Data
Chapter 2:Determinants. Section 2.1: Determinants by cofactor expansion
 Chapter 2:Determinants Section 2.1: Determinants by cofactor expansion [ ] a b Recall: The 2 2 matrix is invertible if ad bc 0. The c d ([ ]) a b function f = ad bc is called the determinant and it associates
Chapter 2:Determinants Section 2.1: Determinants by cofactor expansion [ ] a b Recall: The 2 2 matrix is invertible if ad bc 0. The c d ([ ]) a b function f = ad bc is called the determinant and it associates
ECON 186 Class Notes: Linear Algebra
 ECON 86 Class Notes: Linear Algebra Jijian Fan Jijian Fan ECON 86 / 27 Singularity and Rank As discussed previously, squareness is a necessary condition for a matrix to be nonsingular (have an inverse).
ECON 86 Class Notes: Linear Algebra Jijian Fan Jijian Fan ECON 86 / 27 Singularity and Rank As discussed previously, squareness is a necessary condition for a matrix to be nonsingular (have an inverse).
CS 2750: Machine Learning. Bayesian Networks. Prof. Adriana Kovashka University of Pittsburgh March 14, 2016
 CS 2750: Machine Learning Bayesian Networks Prof. Adriana Kovashka University of Pittsburgh March 14, 2016 Plan for today and next week Today and next time: Bayesian networks (Bishop Sec. 8.1) Conditional
CS 2750: Machine Learning Bayesian Networks Prof. Adriana Kovashka University of Pittsburgh March 14, 2016 Plan for today and next week Today and next time: Bayesian networks (Bishop Sec. 8.1) Conditional
sum of squared error.
 IT 131 MATHEMATCS FOR SCIENCE LECTURE NOTE 6 LEAST SQUARES REGRESSION ANALYSIS and DETERMINANT OF A MATRIX Source: Larson, Edwards, Falvo (2009): Elementary Linear Algebra, Sixth Edition You will now look
IT 131 MATHEMATCS FOR SCIENCE LECTURE NOTE 6 LEAST SQUARES REGRESSION ANALYSIS and DETERMINANT OF A MATRIX Source: Larson, Edwards, Falvo (2009): Elementary Linear Algebra, Sixth Edition You will now look
ENGR-1100 Introduction to Engineering Analysis. Lecture 21
 ENGR-1100 Introduction to Engineering Analysis Lecture 21 Lecture outline Procedure (algorithm) for finding the inverse of invertible matrix. Investigate the system of linear equation and invertibility
ENGR-1100 Introduction to Engineering Analysis Lecture 21 Lecture outline Procedure (algorithm) for finding the inverse of invertible matrix. Investigate the system of linear equation and invertibility
A structural Markov property for decomposable graph laws that allows control of clique intersections
 Biometrika (2017), 00, 0,pp. 1 11 Printed in Great Britain doi: 10.1093/biomet/asx072 A structural Markov property for decomposable graph laws that allows control of clique intersections BY PETER J. GREEN
Biometrika (2017), 00, 0,pp. 1 11 Printed in Great Britain doi: 10.1093/biomet/asx072 A structural Markov property for decomposable graph laws that allows control of clique intersections BY PETER J. GREEN
LS.1 Review of Linear Algebra
 LS. LINEAR SYSTEMS LS.1 Review of Linear Algebra In these notes, we will investigate a way of handling a linear system of ODE s directly, instead of using elimination to reduce it to a single higher-order
LS. LINEAR SYSTEMS LS.1 Review of Linear Algebra In these notes, we will investigate a way of handling a linear system of ODE s directly, instead of using elimination to reduce it to a single higher-order
On the occurrence times of componentwise maxima and bias in likelihood inference for multivariate max-stable distributions
 On the occurrence times of componentwise maxima and bias in likelihood inference for multivariate max-stable distributions J. L. Wadsworth Department of Mathematics and Statistics, Fylde College, Lancaster
On the occurrence times of componentwise maxima and bias in likelihood inference for multivariate max-stable distributions J. L. Wadsworth Department of Mathematics and Statistics, Fylde College, Lancaster
Mixtures of Gaussians with Sparse Structure
 Mixtures of Gaussians with Sparse Structure Costas Boulis 1 Abstract When fitting a mixture of Gaussians to training data there are usually two choices for the type of Gaussians used. Either diagonal or
Mixtures of Gaussians with Sparse Structure Costas Boulis 1 Abstract When fitting a mixture of Gaussians to training data there are usually two choices for the type of Gaussians used. Either diagonal or
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 7 Approximate
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 7 Approximate
Graphical Models 359
 8 Graphical Models Probabilities play a central role in modern pattern recognition. We have seen in Chapter 1 that probability theory can be expressed in terms of two simple equations corresponding to
8 Graphical Models Probabilities play a central role in modern pattern recognition. We have seen in Chapter 1 that probability theory can be expressed in terms of two simple equations corresponding to
Learning Marginal AMP Chain Graphs under Faithfulness
 Learning Marginal AMP Chain Graphs under Faithfulness Jose M. Peña ADIT, IDA, Linköping University, SE-58183 Linköping, Sweden jose.m.pena@liu.se Abstract. Marginal AMP chain graphs are a recently introduced
Learning Marginal AMP Chain Graphs under Faithfulness Jose M. Peña ADIT, IDA, Linköping University, SE-58183 Linköping, Sweden jose.m.pena@liu.se Abstract. Marginal AMP chain graphs are a recently introduced
Graphical Models and Kernel Methods
 Graphical Models and Kernel Methods Jerry Zhu Department of Computer Sciences University of Wisconsin Madison, USA MLSS June 17, 2014 1 / 123 Outline Graphical Models Probabilistic Inference Directed vs.
Graphical Models and Kernel Methods Jerry Zhu Department of Computer Sciences University of Wisconsin Madison, USA MLSS June 17, 2014 1 / 123 Outline Graphical Models Probabilistic Inference Directed vs.
LINEAR SYSTEMS, MATRICES, AND VECTORS
 ELEMENTARY LINEAR ALGEBRA WORKBOOK CREATED BY SHANNON MARTIN MYERS LINEAR SYSTEMS, MATRICES, AND VECTORS Now that I ve been teaching Linear Algebra for a few years, I thought it would be great to integrate
ELEMENTARY LINEAR ALGEBRA WORKBOOK CREATED BY SHANNON MARTIN MYERS LINEAR SYSTEMS, MATRICES, AND VECTORS Now that I ve been teaching Linear Algebra for a few years, I thought it would be great to integrate
Faithfulness of Probability Distributions and Graphs
 Journal of Machine Learning Research 18 (2017) 1-29 Submitted 5/17; Revised 11/17; Published 12/17 Faithfulness of Probability Distributions and Graphs Kayvan Sadeghi Statistical Laboratory University
Journal of Machine Learning Research 18 (2017) 1-29 Submitted 5/17; Revised 11/17; Published 12/17 Faithfulness of Probability Distributions and Graphs Kayvan Sadeghi Statistical Laboratory University
4.1 Notation and probability review
 Directed and undirected graphical models Fall 2015 Lecture 4 October 21st Lecturer: Simon Lacoste-Julien Scribe: Jaime Roquero, JieYing Wu 4.1 Notation and probability review 4.1.1 Notations Let us recall
Directed and undirected graphical models Fall 2015 Lecture 4 October 21st Lecturer: Simon Lacoste-Julien Scribe: Jaime Roquero, JieYing Wu 4.1 Notation and probability review 4.1.1 Notations Let us recall
