Project 4/5 - Molecular dynamics part II: advanced study of Lennard-Jones fluids, deadline December 1st (noon)
|
|
|
- Jennifer Summers
- 6 years ago
- Views:
Transcription
1 Format for delivery of report and programs The format of the project is that of a printed file or hand-written report. The programs should also be included with the report. Write only your candidate number on the first page of the report and state clearly that this is your report for the last project of FYS3150/FYS4150, fall The project counts 50% of the final mark. If you have collaborated with one or more fellow students, please state the respective candidate number(s). There will be a box marked FYS3150/FYS4150 at the reception of the Department of Physics (room FV128). We encourage you to work two and two together. Optimal working groups consist of 2-3 students. You can then hand in a common report. Project 4/5 - Molecular dynamics part II: advanced study of Lennard-Jones fluids, deadline December 1st (noon) So far you should have a working MD code that simulates argon atoms starting in an FCC lattice. Your code should use cell lists to improve the speed of calculating forces by only counting atom pairs closer than a cutoff length r cut. Here it is also important that you shift the energy as mentioned in the first project. Before you start working on this project, be sure that the energy is conserved fairly well (fluctuations in the 5th leading digit in total energy) and that you can simulate a few thousand atoms with a decent speed. Note: the vec3 class has been updated to support more of the expected features/operators on vectors. In this project we will improve the previous code by adding essential features such as save/load functions and a Python framework for running more advanced simulations. We will also measure the pressure and the heat capacity. a) Save/load function: An MD simulation typically consists of several steps. First we create the initial state (i.e. an FCC lattice), then we heat the system to some temperature before we let the system equilibrate before the measuring of statistical properties begins. We might want to reuse some of these states for other studies, so we should create a save/load function. Since ASCII text files are slow, we will write/read binary files (see appendix ). Create two functions on the system class as described in the appendix. b) Save statistical properties to files: In the StatisticsSampler class, you should now have functions measuring kinetic energy, potential energy and temperature. Add another function that measures the number density. Also, you should write the measured numbers to file so you easily can plot them with your favorite plotting tool. This is important to be able to plot 1
2 the properties and calculate more advanced properties based on the fluctuations/variance of the statistical properties (the heat capacity can be calculated as a function of fluctuations in the kinetic energy). c) Measure the pressure P : The pressure of an MD system can be calculated as (it is actually for the NV T ensemble, but it works fairly well in NV E as well) P = ρ n k B T + 1 N F i r i, (1) 3V i=1 where ρ n is the number density, k B is the Boltzmann constant, T is the temperature, V is the volume, r i is the position of atom i and F i is the total force on atom i. The brackets mean that the pressure should be computed by taking the average value of the sum over many timesteps. Assuming we are using two particle forces only, show that the above equation can be rewritten as P = ρ n k B T + 1 3V r ij F ij, (2) where r ij is the relative vector between two atoms i and j and F ij is the force between them. Implement this (you will have to compute this inside the force loop, just as the potential energy). d) Initial behaviour on new systems : Create a new system with 4000 atoms ( unit cells) and an initial temperature T = 300 K. Plot the energy (kinetic, potential and total), temperature and the pressure as functions of time and discuss what happens. Why is the temperature decreasing? What happens to the pressure? How many picoseconds does the system need to stabilize? e) Create a (small) Python framework: With your program, you can now do one simulation, save the state and continue the state with a new set med parameters (such as with or without the Berendsen thermostat). Instead of writing a complete simulation inside your main function, you should write a Python framework (a sample framework can be found on github.com/andeplane/simulation-framework-fys3150) so that you more easily can run more complicated simulations. The Python code should run your executable with the set of parameters sent as arguments to the program. This will allow you to quickly set up a set of simulations with a Python script and let it run automatically for days. You can also, by using for example matplotlib, create all the plots within the same script. f) Measure the heat capacity C V : i>j 2
3 The heat capacity can be calculated through the relation Ek 2 E k 2 = 3k ( BT 2 1 3k ) B, (3) 2N 2C V where we recognize the left hand side as the variance of the kinetic energy and C V as the heat capacity. Does this correlate with experimental values? g) P ρ n diagram (optional, but gives an extra 30% on your final score): It is now time to make full use of the Python framework. We will now do a series of numerical experiments. In each of these experiments, you should create a new state with a given density (you can control this with the FCC lattice parameter b), use the Berendsen thermostat to reach a desired temperature, let the system equilibrate before you sample statistics. Before we start, we should figure out a few things first: 1) Find the number density ρ n as a function of the lattice constant b. 2) How long should the thermostat be enabled before we have reached the temperature we want? 3) How long do we need to equilibrate before we can start sample statistics, and why not immediately after the thermostat has been turned off? 4) Since each state is very correlated to the state of the previous timestep we shouldn t measure statistical properties every timestep, but rather every n th timestep. How long should we wait between each measurement? Now create a run script using your Python framework and prepare a series of experiments. We want to plot a P ρ n diagram. This means that you should measure the pressure for different densities for a given temperature T giving you a isotherm. Repeat this for different temperatures so that you you get many such isotherms. Remember that for each simulation, the measured value for the pressure should be averaged over many measurements. At what temperature do we see a change? Does this indicate a phase transition? Does the diagram look like what you might expect from a van der Waals gas? Appendices Reading/writing binary files We want to store the atom data (positions and velocities) in a binary format. Compared to ASCII text format, the file size will be much smaller. Here follows an example code on 3
4 how to read and write binary files. void save ( s t r i n g filename, System system ) { ofstream f i l e ( filename, i o s : : out i o s : : binary ) ; i f (! f i l e. is_open ( ) ) { c e r r << "Could not open f i l e " << f i l e n a m e << ". Aborting! " << endl ; e x i t ( 1 ) ; int numberofphasespacecoordinates = 6 system >atoms ( ). s i z e ( ) ; double phasespace = new double [ numberofphasespacecoordinates ] ; vec3 systemsize = system >systemsize ( ) ; int phasespacecounter = 0 ; for ( int i =0; i <system >num_atoms_local ; i ++) { phasespace [ phasespacecounter++] = system >atoms ( ) [ i ]. p o s i t i o n [ 0 ] ; phasespace [ phasespacecounter++] = system >atoms ( ) [ i ]. p o s i t i o n [ 1 ] ; phasespace [ phasespacecounter++] = system >atoms ( ) [ i ]. p o s i t i o n [ 2 ] ; phasespace [ phasespacecounter++] = system >atoms ( ) [ i ]. v e l o c i t y [ 0 ] ; phasespace [ phasespacecounter++] = system >atoms ( ) [ i ]. v e l o c i t y [ 1 ] ; phasespace [ phasespacecounter++] = system >atoms ( ) [ i ]. v e l o c i t y [ 2 ] ; int numberofatoms = system >atoms ( ). s i z e ( ) ; f i l e. w r i t e ( reinterpret_cast<char >&numberofatoms, sizeof ( int ) ) ; f i l e. w r i t e ( reinterpret_cast<char >(phasespace ), numberofphasespacecoordinates sizeof ( double ) ) ; f i l e. w r i t e ( reinterpret_cast<char >(&systemsize [ 0 ] ), 3 sizeof ( double ) ) ; f i l e. c l o s e ( ) ; delete phasespace ; void load ( s t r i n g filename, System system ) { i f s t r e a m f i l e ( filename, i o s : : in i o s : : binary ) ; i f (! f i l e. is_open ( ) ) { c e r r << "Could not open f i l e " << f i l e n a m e << ". Aborting! " << endl ; e x i t ( 1 ) ; int numberofatoms ; f i l e. read ( reinterpret_cast<char >(&numberofatoms ), sizeof ( int ) ) ; double phasespace = new double [ 6 numberofatoms ] ; 4
5 f i l e. read ( reinterpret_cast<char >(phasespace ), 6 numberofatoms sizeof ( dou vec3 systemsize ; f i l e. read ( reinterpret_cast<char >(&systemsize [ 0 ] ), 3 sizeof ( double ) ) ; system >setsystemsize ( systemsize ) ; int phasespacecounter = 0 ; double mass = ; system >atoms ( ). c l e a r ( ) ; for ( int i =0; i<numberofatoms ; i++) { Atom atom = new Atom( mass ) ; atom >p o s i t i o n = vec3 ( phasespace [ phasespacecounter ++], phasespace [ phasespacecounter ++], phasespace [ phasespacecounter ++]); atom >v e l o c i t y = vec3 ( phasespace [ phasespacecounter ++], phasespace [ phasespacecounter ++], phasespace [ phasespacecounter ++]); system >atoms ( ). push_back ( atom ) ; f i l e. c l o s e ( ) ; delete phasespace ; 5
 A Nobel Prize for Molecular Dynamics and QM/MM What is Classical Molecular Dynamics? Simulation of explicit particles (atoms, ions,... ) Particles interact via relatively simple analytical potential
A Nobel Prize for Molecular Dynamics and QM/MM What is Classical Molecular Dynamics? Simulation of explicit particles (atoms, ions,... ) Particles interact via relatively simple analytical potential
What is Classical Molecular Dynamics?
 What is Classical Molecular Dynamics? Simulation of explicit particles (atoms, ions,... ) Particles interact via relatively simple analytical potential functions Newton s equations of motion are integrated
What is Classical Molecular Dynamics? Simulation of explicit particles (atoms, ions,... ) Particles interact via relatively simple analytical potential functions Newton s equations of motion are integrated
Molecular Dynamics Simulation of Argon
 Molecular Dynamics Simulation of Argon The fundamental work on this problem was done by A. Rahman, Phys. Rev. 136, A405 (1964). It was extended in many important ways by L. Verlet, Phys. Rev. 159, 98 (1967),
Molecular Dynamics Simulation of Argon The fundamental work on this problem was done by A. Rahman, Phys. Rev. 136, A405 (1964). It was extended in many important ways by L. Verlet, Phys. Rev. 159, 98 (1967),
Lab 4: Handout Molecular dynamics.
 1 Lab 4: Handout Molecular dynamics. In this lab, we will be using MOLDY as our molecular dynamics (MD) code. The MOLDY manual is online. It is a well-written manual, and is an excellent resource for learning
1 Lab 4: Handout Molecular dynamics. In this lab, we will be using MOLDY as our molecular dynamics (MD) code. The MOLDY manual is online. It is a well-written manual, and is an excellent resource for learning
Exercise 2: Solvating the Structure Before you continue, follow these steps: Setting up Periodic Boundary Conditions
 Exercise 2: Solvating the Structure HyperChem lets you place a molecular system in a periodic box of water molecules to simulate behavior in aqueous solution, as in a biological system. In this exercise,
Exercise 2: Solvating the Structure HyperChem lets you place a molecular system in a periodic box of water molecules to simulate behavior in aqueous solution, as in a biological system. In this exercise,
Project 5: Molecular Dynamics
 Physics 2300 Spring 2018 Name Lab partner Project 5: Molecular Dynamics If a computer can model three mutually interacting objects, why not model more than three? As you ll soon see, there is little additional
Physics 2300 Spring 2018 Name Lab partner Project 5: Molecular Dynamics If a computer can model three mutually interacting objects, why not model more than three? As you ll soon see, there is little additional
1 Bulk Simulations in a HCP lattice
 1 Bulk Simulations in a HCP lattice 1.1 Introduction This project is a continuation of two previous projects that studied the mechanism of melting in a FCC and BCC lattice. The current project studies
1 Bulk Simulations in a HCP lattice 1.1 Introduction This project is a continuation of two previous projects that studied the mechanism of melting in a FCC and BCC lattice. The current project studies
Worksheet 1: Integrators
 Simulation Methods in Physics I WS 2014/2015 Worksheet 1: Integrators Olaf Lenz Alexander Schlaich Stefan Kesselheim Florian Fahrenberger Peter Košovan October 22, 2014 Institute for Computational Physics,
Simulation Methods in Physics I WS 2014/2015 Worksheet 1: Integrators Olaf Lenz Alexander Schlaich Stefan Kesselheim Florian Fahrenberger Peter Košovan October 22, 2014 Institute for Computational Physics,
Gear methods I + 1/18
 Gear methods I + 1/18 Predictor-corrector type: knowledge of history is used to predict an approximate solution, which is made more accurate in the following step we do not want (otherwise good) methods
Gear methods I + 1/18 Predictor-corrector type: knowledge of history is used to predict an approximate solution, which is made more accurate in the following step we do not want (otherwise good) methods
Molecular Dynamics. A very brief introduction
 Molecular Dynamics A very brief introduction Sander Pronk Dept. of Theoretical Physics KTH Royal Institute of Technology & Science For Life Laboratory Stockholm, Sweden Why computer simulations? Two primary
Molecular Dynamics A very brief introduction Sander Pronk Dept. of Theoretical Physics KTH Royal Institute of Technology & Science For Life Laboratory Stockholm, Sweden Why computer simulations? Two primary
Scientific Computing II
 Scientific Computing II Molecular Dynamics Simulation Michael Bader SCCS Summer Term 2015 Molecular Dynamics Simulation, Summer Term 2015 1 Continuum Mechanics for Fluid Mechanics? Molecular Dynamics the
Scientific Computing II Molecular Dynamics Simulation Michael Bader SCCS Summer Term 2015 Molecular Dynamics Simulation, Summer Term 2015 1 Continuum Mechanics for Fluid Mechanics? Molecular Dynamics the
2D Clausius Clapeyron equation
 2D Clausius Clapeyron equation 1/14 Aim: Verify the Clausius Clapeyron equation by simulations of a 2D model of matter Model: 8-4 type potential ( Lennard-Jones in 2D) u(r) = 1 r 8 1 r 4 Hard attractive
2D Clausius Clapeyron equation 1/14 Aim: Verify the Clausius Clapeyron equation by simulations of a 2D model of matter Model: 8-4 type potential ( Lennard-Jones in 2D) u(r) = 1 r 8 1 r 4 Hard attractive
Tutorial Three: Loops and Conditionals
 Tutorial Three: Loops and Conditionals Imad Pasha Chris Agostino February 18, 2015 1 Introduction In lecture Monday we learned that combinations of conditionals and loops could make our code much more
Tutorial Three: Loops and Conditionals Imad Pasha Chris Agostino February 18, 2015 1 Introduction In lecture Monday we learned that combinations of conditionals and loops could make our code much more
CHEM-UA 652: Thermodynamics and Kinetics
 1 CHEM-UA 652: Thermodynamics and Kinetics Notes for Lecture 4 I. THE ISOTHERMAL-ISOBARIC ENSEMBLE The isothermal-isobaric ensemble is the closest mimic to the conditions under which most experiments are
1 CHEM-UA 652: Thermodynamics and Kinetics Notes for Lecture 4 I. THE ISOTHERMAL-ISOBARIC ENSEMBLE The isothermal-isobaric ensemble is the closest mimic to the conditions under which most experiments are
2D Clausius Clapeyron equation
 2D Clausius Clapeyron equation 1/14 Aim: Verify the Clausius Clapeyron equation by simulations of a 2D model of matter Model: 8-4 type potential ( Lennard-Jones in 2D) u(r) = 1 r 8 1 r 4 Hard attractive
2D Clausius Clapeyron equation 1/14 Aim: Verify the Clausius Clapeyron equation by simulations of a 2D model of matter Model: 8-4 type potential ( Lennard-Jones in 2D) u(r) = 1 r 8 1 r 4 Hard attractive
Homework Assignment 4 Root Finding Algorithms The Maximum Velocity of a Car Due: Friday, February 19, 2010 at 12noon
 ME 2016 A Spring Semester 2010 Computing Techniques 3-0-3 Homework Assignment 4 Root Finding Algorithms The Maximum Velocity of a Car Due: Friday, February 19, 2010 at 12noon Description and Outcomes In
ME 2016 A Spring Semester 2010 Computing Techniques 3-0-3 Homework Assignment 4 Root Finding Algorithms The Maximum Velocity of a Car Due: Friday, February 19, 2010 at 12noon Description and Outcomes In
Multiple time step Monte Carlo
 JOURNAL OF CHEMICAL PHYSICS VOLUME 117, NUMBER 18 8 NOVEMBER 2002 Multiple time step Monte Carlo Balázs Hetényi a) Department of Chemistry, Princeton University, Princeton, NJ 08544 and Department of Chemistry
JOURNAL OF CHEMICAL PHYSICS VOLUME 117, NUMBER 18 8 NOVEMBER 2002 Multiple time step Monte Carlo Balázs Hetényi a) Department of Chemistry, Princeton University, Princeton, NJ 08544 and Department of Chemistry
Hands-on : Model Potential Molecular Dynamics
 Hands-on : Model Potential Molecular Dynamics OUTLINE 0. DL_POLY code introduction 0.a Input files 1. THF solvent molecule 1.a Geometry optimization 1.b NVE/NVT dynamics 2. Liquid THF 2.a Equilibration
Hands-on : Model Potential Molecular Dynamics OUTLINE 0. DL_POLY code introduction 0.a Input files 1. THF solvent molecule 1.a Geometry optimization 1.b NVE/NVT dynamics 2. Liquid THF 2.a Equilibration
MatSci 331 Homework 4 Molecular Dynamics and Monte Carlo: Stress, heat capacity, quantum nuclear effects, and simulated annealing
 MatSci 331 Homework 4 Molecular Dynamics and Monte Carlo: Stress, heat capacity, quantum nuclear effects, and simulated annealing Due Thursday Feb. 21 at 5pm in Durand 110. Evan Reed In this homework,
MatSci 331 Homework 4 Molecular Dynamics and Monte Carlo: Stress, heat capacity, quantum nuclear effects, and simulated annealing Due Thursday Feb. 21 at 5pm in Durand 110. Evan Reed In this homework,
CPMD Tutorial Atosim/RFCT 2009/10
 These exercices were inspired by the CPMD Tutorial of Axel Kohlmeyer http://www.theochem.ruhruni-bochum.de/ axel.kohlmeyer/cpmd-tutor/index.html and by other tutorials. Here is a summary of what we will
These exercices were inspired by the CPMD Tutorial of Axel Kohlmeyer http://www.theochem.ruhruni-bochum.de/ axel.kohlmeyer/cpmd-tutor/index.html and by other tutorials. Here is a summary of what we will
Thermodynamics of Three-phase Equilibrium in Lennard Jones System with a Simplified Equation of State
 23 Bulletin of Research Center for Computing and Multimedia Studies, Hosei University, 28 (2014) Thermodynamics of Three-phase Equilibrium in Lennard Jones System with a Simplified Equation of State Yosuke
23 Bulletin of Research Center for Computing and Multimedia Studies, Hosei University, 28 (2014) Thermodynamics of Three-phase Equilibrium in Lennard Jones System with a Simplified Equation of State Yosuke
A Phase Transition in Ammonium Chloride. CHM 335 TA: David Robinson Office Hour: Wednesday, 11 am
 A Phase Transition in Ammonium Chloride CHM 335 TA: David Robinson Email: Drobinson@chm.uri.edu Office Hour: Wednesday, 11 am Purpose Determine the critical exponent for the order to disorder transition
A Phase Transition in Ammonium Chloride CHM 335 TA: David Robinson Email: Drobinson@chm.uri.edu Office Hour: Wednesday, 11 am Purpose Determine the critical exponent for the order to disorder transition
Molecular Dynamics. What to choose in an integrator The Verlet algorithm Boundary Conditions in Space and time Reading Assignment: F&S Chapter 4
 Molecular Dynamics What to choose in an integrator The Verlet algorithm Boundary Conditions in Space and time Reading Assignment: F&S Chapter 4 MSE485/PHY466/CSE485 1 The Molecular Dynamics (MD) method
Molecular Dynamics What to choose in an integrator The Verlet algorithm Boundary Conditions in Space and time Reading Assignment: F&S Chapter 4 MSE485/PHY466/CSE485 1 The Molecular Dynamics (MD) method
CS Deep Reinforcement Learning HW2: Policy Gradients due September 19th 2018, 11:59 pm
 CS294-112 Deep Reinforcement Learning HW2: Policy Gradients due September 19th 2018, 11:59 pm 1 Introduction The goal of this assignment is to experiment with policy gradient and its variants, including
CS294-112 Deep Reinforcement Learning HW2: Policy Gradients due September 19th 2018, 11:59 pm 1 Introduction The goal of this assignment is to experiment with policy gradient and its variants, including
Lab 1: Empirical Energy Methods Due: 2/14/18
 Lab 1: Empirical Energy Methods Due: 2/14/18 General remarks on scientific scripting Scientific scripting for managing the input and output data is an important component of modern materials computations,
Lab 1: Empirical Energy Methods Due: 2/14/18 General remarks on scientific scripting Scientific scripting for managing the input and output data is an important component of modern materials computations,
Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar.
 Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar. 25, 2002 Molecular Dynamics: Introduction At physiological conditions, the
Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar. 25, 2002 Molecular Dynamics: Introduction At physiological conditions, the
Lab 2: Photon Counting with a Photomultiplier Tube
 Lab 2: Photon Counting with a Photomultiplier Tube 1 Introduction 1.1 Goals In this lab, you will investigate properties of light using a photomultiplier tube (PMT). You will assess the quantitative measurements
Lab 2: Photon Counting with a Photomultiplier Tube 1 Introduction 1.1 Goals In this lab, you will investigate properties of light using a photomultiplier tube (PMT). You will assess the quantitative measurements
Managing Uncertainty
 Managing Uncertainty Bayesian Linear Regression and Kalman Filter December 4, 2017 Objectives The goal of this lab is multiple: 1. First it is a reminder of some central elementary notions of Bayesian
Managing Uncertainty Bayesian Linear Regression and Kalman Filter December 4, 2017 Objectives The goal of this lab is multiple: 1. First it is a reminder of some central elementary notions of Bayesian
CSCE 155N Fall Homework Assignment 2: Stress-Strain Curve. Assigned: September 11, 2012 Due: October 02, 2012
 CSCE 155N Fall 2012 Homework Assignment 2: Stress-Strain Curve Assigned: September 11, 2012 Due: October 02, 2012 Note: This assignment is to be completed individually - collaboration is strictly prohibited.
CSCE 155N Fall 2012 Homework Assignment 2: Stress-Strain Curve Assigned: September 11, 2012 Due: October 02, 2012 Note: This assignment is to be completed individually - collaboration is strictly prohibited.
Homework Assignment 2 Modeling a Drivetrain Model Accuracy Due: Friday, September 16, 2005
 ME 2016 Sections A-B-C-D Fall Semester 2005 Computing Techniques 3-0-3 Homework Assignment 2 Modeling a Drivetrain Model Accuracy Due: Friday, September 16, 2005 Description and Outcomes In this assignment,
ME 2016 Sections A-B-C-D Fall Semester 2005 Computing Techniques 3-0-3 Homework Assignment 2 Modeling a Drivetrain Model Accuracy Due: Friday, September 16, 2005 Description and Outcomes In this assignment,
In this approach, the ground state of the system is found by modeling a diffusion process.
 The Diffusion Monte Carlo (DMC) Method In this approach, the ground state of the system is found by modeling a diffusion process. Diffusion and random walks Consider a random walk on a lattice with spacing
The Diffusion Monte Carlo (DMC) Method In this approach, the ground state of the system is found by modeling a diffusion process. Diffusion and random walks Consider a random walk on a lattice with spacing
4.4 The Calendar program
 4.4. THE CALENDAR PROGRAM 109 4.4 The Calendar program To illustrate the power of functions, in this section we will develop a useful program that allows the user to input a date or a month or a year.
4.4. THE CALENDAR PROGRAM 109 4.4 The Calendar program To illustrate the power of functions, in this section we will develop a useful program that allows the user to input a date or a month or a year.
Atomic modelling of Argon lattice
 Atomic modelling of Argon lattice Fifth project in Computational physics autumn 2015 Candidate number: 8 University of Oslo Abstract A molecular dynamics simulation of argon atoms initially placed in the
Atomic modelling of Argon lattice Fifth project in Computational physics autumn 2015 Candidate number: 8 University of Oslo Abstract A molecular dynamics simulation of argon atoms initially placed in the
Molecular Dynamics Simulations
 Molecular Dynamics Simulations Dr. Kasra Momeni www.knanosys.com Outline LAMMPS AdHiMad Lab 2 What is LAMMPS? LMMPS = Large-scale Atomic/Molecular Massively Parallel Simulator Open-Source (http://lammps.sandia.gov)
Molecular Dynamics Simulations Dr. Kasra Momeni www.knanosys.com Outline LAMMPS AdHiMad Lab 2 What is LAMMPS? LMMPS = Large-scale Atomic/Molecular Massively Parallel Simulator Open-Source (http://lammps.sandia.gov)
MD Thermodynamics. Lecture 12 3/26/18. Harvard SEAS AP 275 Atomistic Modeling of Materials Boris Kozinsky
 MD Thermodynamics Lecture 1 3/6/18 1 Molecular dynamics The force depends on positions only (not velocities) Total energy is conserved (micro canonical evolution) Newton s equations of motion (second order
MD Thermodynamics Lecture 1 3/6/18 1 Molecular dynamics The force depends on positions only (not velocities) Total energy is conserved (micro canonical evolution) Newton s equations of motion (second order
MATHCAD SOLUTIONS TO THE CHEMICAL ENGINEERING PROBLEM SET
 MATHCA SOLUTIONS TO THE CHEMICAL ENGINEERING PROBLEM SET Mathematical Software Session John J. Hwalek, epartment of Chemical Engineering University of Maine, Orono, Me 4469-5737 (hwalek@maine.maine.edu)
MATHCA SOLUTIONS TO THE CHEMICAL ENGINEERING PROBLEM SET Mathematical Software Session John J. Hwalek, epartment of Chemical Engineering University of Maine, Orono, Me 4469-5737 (hwalek@maine.maine.edu)
Department of Chemical Engineering University of California, Santa Barbara Spring Exercise 2. Due: Thursday, 4/19/09
 Department of Chemical Engineering ChE 210D University of California, Santa Barbara Spring 2012 Exercise 2 Due: Thursday, 4/19/09 Objective: To learn how to compile Fortran libraries for Python, and to
Department of Chemical Engineering ChE 210D University of California, Santa Barbara Spring 2012 Exercise 2 Due: Thursday, 4/19/09 Objective: To learn how to compile Fortran libraries for Python, and to
An Extended van der Waals Equation of State Based on Molecular Dynamics Simulation
 J. Comput. Chem. Jpn., Vol. 8, o. 3, pp. 97 14 (9) c 9 Society of Computer Chemistry, Japan An Extended van der Waals Equation of State Based on Molecular Dynamics Simulation Yosuke KATAOKA* and Yuri YAMADA
J. Comput. Chem. Jpn., Vol. 8, o. 3, pp. 97 14 (9) c 9 Society of Computer Chemistry, Japan An Extended van der Waals Equation of State Based on Molecular Dynamics Simulation Yosuke KATAOKA* and Yuri YAMADA
Practical Information
 MA2501 Numerical Methods Spring 2018 Norwegian University of Science and Technology Department of Mathematical Sciences Semester Project Practical Information This project counts for 30 % of the final
MA2501 Numerical Methods Spring 2018 Norwegian University of Science and Technology Department of Mathematical Sciences Semester Project Practical Information This project counts for 30 % of the final
Computation of non-bonded interactions: Part 1
 Computation of non-bonded interactions: Part 1 Computation of non-local (non-bonded) interactions in the potential function E p non-local scales with the number of atoms as N. For the large molecular systems
Computation of non-bonded interactions: Part 1 Computation of non-local (non-bonded) interactions in the potential function E p non-local scales with the number of atoms as N. For the large molecular systems
This Answer/Solution Example is taken from a student s actual homework report. I thank him for permission to use it here. JG
 This Answer/Solution Example is taken from a student s actual homework report. I thank him for permission to use it here. JG Chem 8021 Spring 2005 Project II Calculation of Self-Diffusion Coeffecient of
This Answer/Solution Example is taken from a student s actual homework report. I thank him for permission to use it here. JG Chem 8021 Spring 2005 Project II Calculation of Self-Diffusion Coeffecient of
1 Newton s 2nd and 3rd Laws
 Physics 13 - Winter 2007 Lab 2 Instructions 1 Newton s 2nd and 3rd Laws 1. Work through the tutorial called Newton s Second and Third Laws on pages 31-34 in the UW Tutorials in Introductory Physics workbook.
Physics 13 - Winter 2007 Lab 2 Instructions 1 Newton s 2nd and 3rd Laws 1. Work through the tutorial called Newton s Second and Third Laws on pages 31-34 in the UW Tutorials in Introductory Physics workbook.
Lab 1: Handout GULP: an Empirical energy code
 Lab 1: Handout GULP: an Empirical energy code We will be using the GULP code as our energy code. GULP is a program for performing a variety of types of simulations on 3D periodic solids, gas phase clusters,
Lab 1: Handout GULP: an Empirical energy code We will be using the GULP code as our energy code. GULP is a program for performing a variety of types of simulations on 3D periodic solids, gas phase clusters,
Assignment 2 Atomic-Level Molecular Modeling
 Assignment 2 Atomic-Level Molecular Modeling CS/BIOE/CME/BIOPHYS/BIOMEDIN 279 Due: November 3, 2016 at 3:00 PM The goal of this assignment is to understand the biological and computational aspects of macromolecular
Assignment 2 Atomic-Level Molecular Modeling CS/BIOE/CME/BIOPHYS/BIOMEDIN 279 Due: November 3, 2016 at 3:00 PM The goal of this assignment is to understand the biological and computational aspects of macromolecular
Looking at a two binary digit sum shows what we need to extend addition to multiple binary digits.
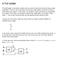 A Full Adder The half-adder is extremely useful until you want to add more that one binary digit quantities. The slow way to develop a two binary digit adders would be to make a truth table and reduce
A Full Adder The half-adder is extremely useful until you want to add more that one binary digit quantities. The slow way to develop a two binary digit adders would be to make a truth table and reduce
Matlab week II. Tuesday and Wednesday: Exercise 7 Matrices with Minchao Wu, submission required. Thursday: Exercise 9 Functions.
 Matlab week II Monday: 10-12 - Vectors and matrices, tutorial with exercises. No tasks to submit. 13-15 Exercise 6, vectors. 1 preparation exercise and 2 tasks to submit. Tuesday and Wednesday: Exercise
Matlab week II Monday: 10-12 - Vectors and matrices, tutorial with exercises. No tasks to submit. 13-15 Exercise 6, vectors. 1 preparation exercise and 2 tasks to submit. Tuesday and Wednesday: Exercise
Econ 1123: Section 2. Review. Binary Regressors. Bivariate. Regression. Omitted Variable Bias
 Contact Information Elena Llaudet Sections are voluntary. My office hours are Thursdays 5pm-7pm in Littauer Mezzanine 34-36 (Note room change) You can email me administrative questions to ellaudet@gmail.com.
Contact Information Elena Llaudet Sections are voluntary. My office hours are Thursdays 5pm-7pm in Littauer Mezzanine 34-36 (Note room change) You can email me administrative questions to ellaudet@gmail.com.
IAP 2006: From nano to macro: Introduction to atomistic modeling techniques and application in a case study of modeling fracture of copper (1.
 IAP 2006: From nano to macro: Introduction to atomistic modeling techniques and application in a case study of modeling fracture of copper (1.978 PDF) http://web.mit.edu/mbuehler/www/teaching/iap2006/intro.htm
IAP 2006: From nano to macro: Introduction to atomistic modeling techniques and application in a case study of modeling fracture of copper (1.978 PDF) http://web.mit.edu/mbuehler/www/teaching/iap2006/intro.htm
Homework 4, Part B: Structured perceptron
 Homework 4, Part B: Structured perceptron CS 585, UMass Amherst, Fall 2016 Overview Due Friday, Oct 28. Get starter code/data from the course website s schedule page. You should submit a zipped directory
Homework 4, Part B: Structured perceptron CS 585, UMass Amherst, Fall 2016 Overview Due Friday, Oct 28. Get starter code/data from the course website s schedule page. You should submit a zipped directory
Temperature and Pressure Controls
 Ensembles Temperature and Pressure Controls 1. (E, V, N) microcanonical (constant energy) 2. (T, V, N) canonical, constant volume 3. (T, P N) constant pressure 4. (T, V, µ) grand canonical #2, 3 or 4 are
Ensembles Temperature and Pressure Controls 1. (E, V, N) microcanonical (constant energy) 2. (T, V, N) canonical, constant volume 3. (T, P N) constant pressure 4. (T, V, µ) grand canonical #2, 3 or 4 are
Problem Set 3 September 25, 2009
 September 25, 2009 General Instructions: 1. You are expected to state all your assumptions and provide step-by-step solutions to the numerical problems. Unless indicated otherwise, the computational problems
September 25, 2009 General Instructions: 1. You are expected to state all your assumptions and provide step-by-step solutions to the numerical problems. Unless indicated otherwise, the computational problems
PHY 6500 Thermal and Statistical Physics - Fall 2017
 PHY 6500 Thermal and Statistical Physics - Fall 2017 Time: M, F 12:30 PM 2:10 PM. From 08/30/17 to 12/19/17 Place: Room 185 Physics Research Building Lecturer: Boris Nadgorny E-mail: nadgorny@physics.wayne.edu
PHY 6500 Thermal and Statistical Physics - Fall 2017 Time: M, F 12:30 PM 2:10 PM. From 08/30/17 to 12/19/17 Place: Room 185 Physics Research Building Lecturer: Boris Nadgorny E-mail: nadgorny@physics.wayne.edu
Analysis of the simulation
 Analysis of the simulation Marcus Elstner and Tomáš Kubař January 7, 2014 Thermodynamic properties time averages of thermodynamic quantites correspond to ensemble averages (ergodic theorem) some quantities
Analysis of the simulation Marcus Elstner and Tomáš Kubař January 7, 2014 Thermodynamic properties time averages of thermodynamic quantites correspond to ensemble averages (ergodic theorem) some quantities
Model-to-Model Interface for Multiscale Materials Modeling
 Graduate Theses and Dissertations Iowa State University Capstones, Theses and Dissertations 2016 Model-to-Model Interface for Multiscale Materials Modeling Perry Antonelli Iowa State University Follow
Graduate Theses and Dissertations Iowa State University Capstones, Theses and Dissertations 2016 Model-to-Model Interface for Multiscale Materials Modeling Perry Antonelli Iowa State University Follow
V = c( 1 x 12 1 x 6 )
 Introduction The Lennard-Jones potential is used relentlessly in chemistry and physics[2][3][4] (Just the top few results from Google Scholar). It is often employed to model the van der Waals effect in
Introduction The Lennard-Jones potential is used relentlessly in chemistry and physics[2][3][4] (Just the top few results from Google Scholar). It is often employed to model the van der Waals effect in
Principles of Equilibrium Statistical Mechanics
 Debashish Chowdhury, Dietrich Stauffer Principles of Equilibrium Statistical Mechanics WILEY-VCH Weinheim New York Chichester Brisbane Singapore Toronto Table of Contents Part I: THERMOSTATICS 1 1 BASIC
Debashish Chowdhury, Dietrich Stauffer Principles of Equilibrium Statistical Mechanics WILEY-VCH Weinheim New York Chichester Brisbane Singapore Toronto Table of Contents Part I: THERMOSTATICS 1 1 BASIC
JASS Modeling and visualization of molecular dynamic processes
 JASS 2009 Konstantin Shefov Modeling and visualization of molecular dynamic processes St Petersburg State University, Physics faculty, Department of Computational Physics Supervisor PhD Stepanova Margarita
JASS 2009 Konstantin Shefov Modeling and visualization of molecular dynamic processes St Petersburg State University, Physics faculty, Department of Computational Physics Supervisor PhD Stepanova Margarita
Chemical reaction networks and diffusion
 FYTN05 Computer Assignment 2 Chemical reaction networks and diffusion Supervisor: Adriaan Merlevede Office: K336-337, E-mail: adriaan@thep.lu.se 1 Introduction This exercise focuses on understanding and
FYTN05 Computer Assignment 2 Chemical reaction networks and diffusion Supervisor: Adriaan Merlevede Office: K336-337, E-mail: adriaan@thep.lu.se 1 Introduction This exercise focuses on understanding and
Computersimulation in der Materialwissenschaft
 Computersimulation in der Materialwissenschaft WS 2014-2015 Diplom Werkstoffwissenschaft + andere Interessenten Arezoo Dianat und Leonardo Medrano Institut für Werkstoffwissenschaft Professur Werkstoffwissenschaft
Computersimulation in der Materialwissenschaft WS 2014-2015 Diplom Werkstoffwissenschaft + andere Interessenten Arezoo Dianat und Leonardo Medrano Institut für Werkstoffwissenschaft Professur Werkstoffwissenschaft
CS Homework 3. October 15, 2009
 CS 294 - Homework 3 October 15, 2009 If you have questions, contact Alexandre Bouchard (bouchard@cs.berkeley.edu) for part 1 and Alex Simma (asimma@eecs.berkeley.edu) for part 2. Also check the class website
CS 294 - Homework 3 October 15, 2009 If you have questions, contact Alexandre Bouchard (bouchard@cs.berkeley.edu) for part 1 and Alex Simma (asimma@eecs.berkeley.edu) for part 2. Also check the class website
( 3, 5) and zeros of 2 and 8.
 FOM 11 T26 QUADRATIC FUNCTIONS IN VERTEX FORM - 2 1 DETERMINING QUADRATIC FUNCTIONS IN VERTEX FORM I) THE VERTEX FORM OF A QUADRATIC FUNCTION (PARABOLA) IS. To write a quadratic function in vertex form
FOM 11 T26 QUADRATIC FUNCTIONS IN VERTEX FORM - 2 1 DETERMINING QUADRATIC FUNCTIONS IN VERTEX FORM I) THE VERTEX FORM OF A QUADRATIC FUNCTION (PARABOLA) IS. To write a quadratic function in vertex form
Appendix 4 Weather. Weather Providers
 Appendix 4 Weather Using weather data in your automation solution can have many benefits. Without weather data, your home automation happens regardless of environmental conditions. Some things you can
Appendix 4 Weather Using weather data in your automation solution can have many benefits. Without weather data, your home automation happens regardless of environmental conditions. Some things you can
Lecture 7: Kinetic Theory of Gases, Part 2. ! = mn v x
 Lecture 7: Kinetic Theory of Gases, Part 2 Last lecture, we began to explore the behavior of an ideal gas in terms of the molecules in it We found that the pressure of the gas was: P = N 2 mv x,i! = mn
Lecture 7: Kinetic Theory of Gases, Part 2 Last lecture, we began to explore the behavior of an ideal gas in terms of the molecules in it We found that the pressure of the gas was: P = N 2 mv x,i! = mn
Written Exam Linear and Integer Programming (DM554)
 Written Exam Linear and Integer Programming (DM554) Department of Mathematics and Computer Science University of Southern Denmark Monday, June 22, 2015, 10:00 14:00, Festsalen, Niels Bohr Allé 1 The exam
Written Exam Linear and Integer Programming (DM554) Department of Mathematics and Computer Science University of Southern Denmark Monday, June 22, 2015, 10:00 14:00, Festsalen, Niels Bohr Allé 1 The exam
CS 1124Media computation. Oct 6, 2008 Steve Harrison
 CS 1124Media computation Oct 6, 2008 Steve Harrison Today Midterm Introduction to working with sound HW 5 - Faster and Faster Today Midterm Introduction to working with sound HW 5 - Faster and Faster Mid
CS 1124Media computation Oct 6, 2008 Steve Harrison Today Midterm Introduction to working with sound HW 5 - Faster and Faster Today Midterm Introduction to working with sound HW 5 - Faster and Faster Mid
Written Exam Linear and Integer Programming (DM545)
 Written Exam Linear and Integer Programming (DM545) Department of Mathematics and Computer Science University of Southern Denmark Monday, June 22, 2015, 10:00 14:00, Festsalen, Niels Bohr Allé 1 The exam
Written Exam Linear and Integer Programming (DM545) Department of Mathematics and Computer Science University of Southern Denmark Monday, June 22, 2015, 10:00 14:00, Festsalen, Niels Bohr Allé 1 The exam
Conservation of Mechanical Energy Activity Purpose
 Conservation of Mechanical Energy Activity Purpose During the lab, students will become familiar with solving a problem involving the conservation of potential and kinetic energy. A cart is attached to
Conservation of Mechanical Energy Activity Purpose During the lab, students will become familiar with solving a problem involving the conservation of potential and kinetic energy. A cart is attached to
DOAS measurements of Atmospheric Species
 Practical Environmental Measurement Techniques: DOAS measurements of Atmospheric Species Last change of document: April 14, 2014 Supervisor: Dr. Andreas Richter, room U2090, tel 62103, and Dr. Folkard
Practical Environmental Measurement Techniques: DOAS measurements of Atmospheric Species Last change of document: April 14, 2014 Supervisor: Dr. Andreas Richter, room U2090, tel 62103, and Dr. Folkard
MAT Mathematics in Today's World
 MAT 1000 Mathematics in Today's World Last Time We discussed the four rules that govern probabilities: 1. Probabilities are numbers between 0 and 1 2. The probability an event does not occur is 1 minus
MAT 1000 Mathematics in Today's World Last Time We discussed the four rules that govern probabilities: 1. Probabilities are numbers between 0 and 1 2. The probability an event does not occur is 1 minus
Computer Simulation of Shock Waves in Condensed Matter. Matthew R. Farrow 2 November 2007
 Computer Simulation of Shock Waves in Condensed Matter Matthew R. Farrow 2 November 2007 Outline of talk Shock wave theory Results Conclusion Computer simulation of shock waves Shock Wave Theory Shock
Computer Simulation of Shock Waves in Condensed Matter Matthew R. Farrow 2 November 2007 Outline of talk Shock wave theory Results Conclusion Computer simulation of shock waves Shock Wave Theory Shock
HONORS CHEMISTRY Bonding Unit Summative Assessment
 Assignment overview For the bonding unit final assessment, you will create a story called A Tale of Four Electrons that incorporates concepts from the unit. The story will consist of four parts: Life before
Assignment overview For the bonding unit final assessment, you will create a story called A Tale of Four Electrons that incorporates concepts from the unit. The story will consist of four parts: Life before
VPython Class 2: Functions, Fields, and the dreaded ˆr
 Physics 212E Classical and Modern Physics Spring 2016 1. Introduction VPython Class 2: Functions, Fields, and the dreaded ˆr In our study of electric fields, we start with a single point charge source
Physics 212E Classical and Modern Physics Spring 2016 1. Introduction VPython Class 2: Functions, Fields, and the dreaded ˆr In our study of electric fields, we start with a single point charge source
An (incomplete) Introduction to FreeFem++
 An (incomplete) Introduction to FreeFem++ Nathaniel Mays Wheeling Jesuit University November 12, 2011 Nathaniel Mays (Wheeling Jesuit University)An (incomplete) Introduction to FreeFem++ November 12, 2011
An (incomplete) Introduction to FreeFem++ Nathaniel Mays Wheeling Jesuit University November 12, 2011 Nathaniel Mays (Wheeling Jesuit University)An (incomplete) Introduction to FreeFem++ November 12, 2011
Tutorial 1: Lennard-Jones Liquid ESPResSo Basics
 Tutorial 1: Lennard-Jones Liquid ESPResSo Basics October 10, 2016 Contents 1 Introduction 2 2 Background 2.1 The Lennard-Jones potential......................... 2.2 Units.......................................
Tutorial 1: Lennard-Jones Liquid ESPResSo Basics October 10, 2016 Contents 1 Introduction 2 2 Background 2.1 The Lennard-Jones potential......................... 2.2 Units.......................................
FIT100 Spring 01. Project 2. Astrological Toys
 FIT100 Spring 01 Project 2 Astrological Toys In this project you will write a series of Windows applications that look up and display astrological signs and dates. The applications that will make up the
FIT100 Spring 01 Project 2 Astrological Toys In this project you will write a series of Windows applications that look up and display astrological signs and dates. The applications that will make up the
6. How Functions Work and Are Accessed. Topics: Modules and Functions More on Importing Call Frames
 6. How Functions Work and Are Accessed Topics: Modules and Functions More on Importing Call Frames Let s Talk About Modules What Are They? M1.py A module is a.py file that contains Python code The name
6. How Functions Work and Are Accessed Topics: Modules and Functions More on Importing Call Frames Let s Talk About Modules What Are They? M1.py A module is a.py file that contains Python code The name
EXPERIMENT : Thermal Equilibrium. Topics of investigation: Temperature, heat, heat capacity, entropy, irreversibility
 EXPERIMENT 2000.06.1: Thermal Equilibrium Topics of investigation: Temperature, heat, heat capacity, entropy, irreversibility Read about this topic in: Serway, Chs. 16, 17, 18; C&J Chs 13, 14, 15; also,
EXPERIMENT 2000.06.1: Thermal Equilibrium Topics of investigation: Temperature, heat, heat capacity, entropy, irreversibility Read about this topic in: Serway, Chs. 16, 17, 18; C&J Chs 13, 14, 15; also,
CSE 140 Spring 2017: Final Solutions (Total 50 Points)
 CSE 140 Spring 2017: Final Solutions (Total 50 Points) 1. (Boolean Algebra) Prove the following Boolean theorem using Boolean laws only, i.e. no theorem is allowed for the proof. State the name of the
CSE 140 Spring 2017: Final Solutions (Total 50 Points) 1. (Boolean Algebra) Prove the following Boolean theorem using Boolean laws only, i.e. no theorem is allowed for the proof. State the name of the
Introduction to Simulation - Lectures 17, 18. Molecular Dynamics. Nicolas Hadjiconstantinou
 Introduction to Simulation - Lectures 17, 18 Molecular Dynamics Nicolas Hadjiconstantinou Molecular Dynamics Molecular dynamics is a technique for computing the equilibrium and non-equilibrium properties
Introduction to Simulation - Lectures 17, 18 Molecular Dynamics Nicolas Hadjiconstantinou Molecular Dynamics Molecular dynamics is a technique for computing the equilibrium and non-equilibrium properties
Pre-yield non-affine fluctuations and a hidden critical point in strained crystals
 Supplementary Information for: Pre-yield non-affine fluctuations and a hidden critical point in strained crystals Tamoghna Das, a,b Saswati Ganguly, b Surajit Sengupta c and Madan Rao d a Collective Interactions
Supplementary Information for: Pre-yield non-affine fluctuations and a hidden critical point in strained crystals Tamoghna Das, a,b Saswati Ganguly, b Surajit Sengupta c and Madan Rao d a Collective Interactions
Introduction to functions
 Introduction to functions Comp Sci 1570 Introduction to C++ Outline 1 2 Functions A function is a reusable sequence of s designed to do a particular job. In C++, a function is a group of s that is given
Introduction to functions Comp Sci 1570 Introduction to C++ Outline 1 2 Functions A function is a reusable sequence of s designed to do a particular job. In C++, a function is a group of s that is given
Conservation of Mechanical Energy Activity Purpose
 Conservation of Mechanical Energy Activity Purpose During the lab, students will become familiar with solving a problem involving the conservation of potential and kinetic energy. A cart is attached to
Conservation of Mechanical Energy Activity Purpose During the lab, students will become familiar with solving a problem involving the conservation of potential and kinetic energy. A cart is attached to
the energy of motion!
 What are the molecules of matter doing all the time?! Heat and Temperature! Notes! All matter is composed of continually jiggling atoms or molecules! The jiggling is! If something is vibrating, what kind
What are the molecules of matter doing all the time?! Heat and Temperature! Notes! All matter is composed of continually jiggling atoms or molecules! The jiggling is! If something is vibrating, what kind
PC Control / Touch Control
 PC Control / Touch Control Version 6.0 / 5.840.0150 New Features Manual 8.840.8007EN Metrohm AG CH-9101 Herisau Switzerland Phone +41 71 353 85 85 Fax +41 71 353 89 01 info@metrohm.com www.metrohm.com
PC Control / Touch Control Version 6.0 / 5.840.0150 New Features Manual 8.840.8007EN Metrohm AG CH-9101 Herisau Switzerland Phone +41 71 353 85 85 Fax +41 71 353 89 01 info@metrohm.com www.metrohm.com
9.33 Ab initio Molecular Dynamics Simulations
 9.33 Ab initio Molecular Dynamics Simulations 1 9.33 Ab initio Molecular Dynamics Simulations Recently, it was decided to include an ab initio molecular dynamics (AIMD) module in ORCA. 1 As a plethora
9.33 Ab initio Molecular Dynamics Simulations 1 9.33 Ab initio Molecular Dynamics Simulations Recently, it was decided to include an ab initio molecular dynamics (AIMD) module in ORCA. 1 As a plethora
CSE546 Machine Learning, Autumn 2016: Homework 1
 CSE546 Machine Learning, Autumn 2016: Homework 1 Due: Friday, October 14 th, 5pm 0 Policies Writing: Please submit your HW as a typed pdf document (not handwritten). It is encouraged you latex all your
CSE546 Machine Learning, Autumn 2016: Homework 1 Due: Friday, October 14 th, 5pm 0 Policies Writing: Please submit your HW as a typed pdf document (not handwritten). It is encouraged you latex all your
Advanced Molecular Molecular Dynamics
 Advanced Molecular Molecular Dynamics Technical details May 12, 2014 Integration of harmonic oscillator r m period = 2 k k and the temperature T determine the sampling of x (here T is related with v 0
Advanced Molecular Molecular Dynamics Technical details May 12, 2014 Integration of harmonic oscillator r m period = 2 k k and the temperature T determine the sampling of x (here T is related with v 0
Department of Chemical Engineering University of California, Santa Barbara Spring Exercise 3. Due: Thursday, 5/3/12
 Department of Chemical Engineering ChE 210D University of California, Santa Barbara Spring 2012 Exercise 3 Due: Thursday, 5/3/12 Objective: To learn how to write & compile Fortran libraries for Python,
Department of Chemical Engineering ChE 210D University of California, Santa Barbara Spring 2012 Exercise 3 Due: Thursday, 5/3/12 Objective: To learn how to write & compile Fortran libraries for Python,
MATLAB Ordinary Differential Equation (ODE) solver for a simple example 1. Introduction
 MATLAB Ordinary Differential Equation (ODE) solver for a simple example 1. Introduction Differential equations are a convenient way to express mathematically a change of a dependent variable (e.g. concentration
MATLAB Ordinary Differential Equation (ODE) solver for a simple example 1. Introduction Differential equations are a convenient way to express mathematically a change of a dependent variable (e.g. concentration
ICCP Project 2 - Advanced Monte Carlo Methods Choose one of the three options below
 ICCP Project 2 - Advanced Monte Carlo Methods Choose one of the three options below Introduction In statistical physics Monte Carlo methods are considered to have started in the Manhattan project (1940
ICCP Project 2 - Advanced Monte Carlo Methods Choose one of the three options below Introduction In statistical physics Monte Carlo methods are considered to have started in the Manhattan project (1940
CPSC 531 Systems Modeling and Simulation FINAL EXAM
 CPSC 531 Systems Modeling and Simulation FINAL EXAM Department of Computer Science University of Calgary Professor: Carey Williamson December 21, 2017 This is a CLOSED BOOK exam. Textbooks, notes, laptops,
CPSC 531 Systems Modeling and Simulation FINAL EXAM Department of Computer Science University of Calgary Professor: Carey Williamson December 21, 2017 This is a CLOSED BOOK exam. Textbooks, notes, laptops,
Introduction to model potential Molecular Dynamics A3hourcourseatICTP
 Introduction to model potential Molecular Dynamics A3hourcourseatICTP Alessandro Mattoni 1 1 Istituto Officina dei Materiali CNR-IOM Unità di Cagliari SLACS ICTP School on numerical methods for energy,
Introduction to model potential Molecular Dynamics A3hourcourseatICTP Alessandro Mattoni 1 1 Istituto Officina dei Materiali CNR-IOM Unità di Cagliari SLACS ICTP School on numerical methods for energy,
E = <ij> N 2. (0.3) (so that k BT
 Phys 4 Project solutions. November 4, 7 Note the model and units Quantities The D Ising model has the total energy E written in terms of the spins σ i = ± site i, E = σ i σ j (.) where < ij > indicates
Phys 4 Project solutions. November 4, 7 Note the model and units Quantities The D Ising model has the total energy E written in terms of the spins σ i = ± site i, E = σ i σ j (.) where < ij > indicates
Assignment 6: A/D-Converters and PID Controllers
 Assignment 6: A/D-Converters and PID Controllers 1DT056: Programming Embedded Systems Uppsala University March 4th, 2012 You can achieve a maximum number of 20 points in this assignment. 12 out of the
Assignment 6: A/D-Converters and PID Controllers 1DT056: Programming Embedded Systems Uppsala University March 4th, 2012 You can achieve a maximum number of 20 points in this assignment. 12 out of the
Graded Project #1. Part 1. Explicit Runge Kutta methods. Goals Differential Equations FMN130 Gustaf Söderlind and Carmen Arévalo
 2008-11-07 Graded Project #1 Differential Equations FMN130 Gustaf Söderlind and Carmen Arévalo This homework is due to be handed in on Wednesday 12 November 2008 before 13:00 in the post box of the numerical
2008-11-07 Graded Project #1 Differential Equations FMN130 Gustaf Söderlind and Carmen Arévalo This homework is due to be handed in on Wednesday 12 November 2008 before 13:00 in the post box of the numerical
Molecular computer experiment
 Molecular computer experiment 1/13 Also simulation or pseudoexperiment REAL EXPERIMENT Record everything in a lab notebook Build the experimental apparatus (from parts) Purchase chemicals, synthetise if
Molecular computer experiment 1/13 Also simulation or pseudoexperiment REAL EXPERIMENT Record everything in a lab notebook Build the experimental apparatus (from parts) Purchase chemicals, synthetise if
2008 Biowerkzeug Ltd.
 2008 Biowerkzeug Ltd. 1 Contents Summary...3 1 Simulation...4 1.1 Setup...4 1.2 Output...4 2 Settings...5 3 Analysis...9 3.1 Setup...9 3.2 Input options...9 3.3 Descriptions...10 Please note that we cannot
2008 Biowerkzeug Ltd. 1 Contents Summary...3 1 Simulation...4 1.1 Setup...4 1.2 Output...4 2 Settings...5 3 Analysis...9 3.1 Setup...9 3.2 Input options...9 3.3 Descriptions...10 Please note that we cannot
CS1210 Lecture 23 March 8, 2019
 CS1210 Lecture 23 March 8, 2019 HW5 due today In-discussion exams next week Optional homework assignment next week can be used to replace a score from among HW 1 3. Will be posted some time before Monday
CS1210 Lecture 23 March 8, 2019 HW5 due today In-discussion exams next week Optional homework assignment next week can be used to replace a score from among HW 1 3. Will be posted some time before Monday
Chemical reaction networks and diffusion
 FYTN05 Fall 2012 Computer Assignment 2 Chemical reaction networks and diffusion Supervisor: Erica Manesso (Office: K217, Phone: +46 46 22 29232, E-mail: erica.manesso@thep.lu.se) Deadline: October 23,
FYTN05 Fall 2012 Computer Assignment 2 Chemical reaction networks and diffusion Supervisor: Erica Manesso (Office: K217, Phone: +46 46 22 29232, E-mail: erica.manesso@thep.lu.se) Deadline: October 23,
From Atoms to Materials: Predictive Theory and Simulations
 From Atoms to Materials: Predictive Theory and Simulations Week 3 Lecture 4 Potentials for metals and semiconductors Ale Strachan strachan@purdue.edu School of Materials Engineering & Birck anotechnology
From Atoms to Materials: Predictive Theory and Simulations Week 3 Lecture 4 Potentials for metals and semiconductors Ale Strachan strachan@purdue.edu School of Materials Engineering & Birck anotechnology
