Statistical Applications in Genetics and Molecular Biology
|
|
|
- Bertha Smith
- 6 years ago
- Views:
Transcription
1 Statistical Applications in Genetics and Molecular Biology Volume 4, Issue Article 10 Prediction of Missing Values in Microarray and Use of Mixed Models to Evaluate the Predictors Guri Feten Trygve Almøy Are H. Aastveit Norwegian University of Life Sciences, guri.feten@umb.no Norwegian University of Life Sciences, trygve.almoy@umb.no Norwegian University of Life Sciences, are.aastveit@umb.no Copyright c 2005 by the authors. All rights reserved. No part of this publication may be reproduced, stored in a retrieval system, or transmitted, in any form or by any means, electronic, mechanical, photocopying, recording, or otherwise, without the prior written permission of the publisher, bepress, which has been given certain exclusive rights by the author. Statistical Applications in Genetics and Molecular Biology is produced by The Berkeley Electronic Press (bepress).
2 Prediction of Missing Values in Microarray and Use of Mixed Models to Evaluate the Predictors Guri Feten, Trygve Almøy, and Are H. Aastveit Abstract Gene expression microarray experiments generate data sets with multiple missing expression values. In some cases, analysis of gene expression requires a complete matrix as input. Either genes with missing values can be removed, or the missing values can be replaced using prediction. We propose six imputation methods. A comparative study of the methods was performed on data from mice and data from the bacterium Enterococcus faecalis, and a linear mixed model was used to test for differences between the methods. The study showed that different methods capability to predict is dependent on the data, hence the ideal choice of method and number of components are different for each data set. For data with correlation structure methods based on K-nearest neighbours seemed to be best, while for data without correlation structure using the average of the gene was to be preferred. We would like to thank two anonymous referees for constructive criticism. This work has been financed by Norwegian University of Life Sciences.
3 Feten et al.: Prediction of Missing Values in Microarray 1 1 Introduction The technology of DNA microarray allows monitoring expression levels for thousands of genes simultaneously. Large-scale gene expression studies have been carried out to study cell cycle (Eisen et al., 1998), tumor tissues (DeRisi et al., 1996), yeast sporulation (Chu et al., 1998), and resequence and mutational analysis (Hacia, 1999). An introduction to the microarray technology can be found in e.g. Nguyen et al. (2002a). The microarray data are characterized by many measured variables (genes) on only a few observations (replications, parallels). Often microarray experiments generate data sets with multiple missing expression values. In some cases, analysis of gene expression requires a complete matrix as input, e.g. hierarchical clustering. There are different reasons for missing expression values. The microarray may contain weak spots. Usually these spots are filtered out. One way to sort out weak spots is to compare the pixels of the spot with the pixels of the background. If the fraction of spot pixels greater than the median of the background pixels is less than a given threshold, the gene expression that corresponds to this spot will be set as missing. Another reason for missing expression values is technical errors during the hybridization. Microarrays are scanned in a microarray scanner, producing either fluorescence intensities or radioactive intensities. The intensities must be higher than a given value. If the intensity is below this threshold, we define the value as missing. A third reason for missing values is dust, scratches, and systematic errors on the slides. Recently comparative studies of three data imputation methods; a singular value decomposition based method, weighted K-nearest neighbours, and row average were presented (Troyanskaya et al., 2001; Hastie et al., 1999). Bø et al. (2004) compared methods that utilize correlations between both genes and arrays based on the least square principle and the method of K-nearest neighbours. Ouyang et al. (2004) proposed an imputation method based on Gaussian mixture clustering and model averaging. All of these papers investigated the methods for different fractions of missing data. Nguyen et al. (2004) investigated how the accuracy of four different prediction methods; mean, K-nearest neighbours, ordinary lest squares regression, and partial least squares regression, is depended on the actual gene expression. In addition to the work that has been done on missing value prediction for microarray data, larger studies have been devoted to similar problems in other fields. Common methods include iterating procedures (Yates, 1933; Healy and Westmacott, 1956), imputing conditional means (Buck, 1960), hot deck imputation (Ernst, 1980), multiple imputation (Rubin, 1987; Rubin and Schenker, 1991), and bootstrap (Efron, 1994). In this paper different methods for replacing missing data have been studied. In addition to previously studied methods; Principal Component Regression (PCR), Partial Least Square Regression (PLSR), weighted Produced by The Berkeley Electronic Press, 2005
4 2 Statistical Applications in Genetics and Molecular Biology Vol. 4 [2005], No. 1, Article 10 K-nearest neighbours (KNN) with genes as neighbours, and gene average, we focused on two methods; Factor Analysis Regression (FAR) and weighted K-nearest neighbours (KNN) with observations as neighbours. Earlier papers on missing values have none or few attempts on comparing the prediction methods statistically. Bø et al. (2004) used paired t-test to compare the methods. Cross Validation Analysis of Variance (CVANOVA) was introduced to compare prediction methods by Indahl and Næs (1998). Based on the idea of CVANOVA, in this paper we have used mixed models to compare the methods. In Section 2 we will present six methods to predict missing values, four based on regression, and two based on K-nearest neighbours. A method based on linear mixed models to compare the prediction methods is also described. Further on, in Section 3, there is a presentation of the data used in the study, followed, in Section 4, by the results of the study. 2 Methods 2.1 Prediction and missing values Suppose we will study the gene expressions of p genes (typically ) on n observations (p n), where several of the genes have missing expressions. Our aim is to predict these missing values. Let y j denote the n 1 vector containing the n gene expressions of gene j. Let X (j) denote the n (p 1) matrix containing the gene expressions of the p 1 other genes. If nothing else is stated, both y j and X (j) are centered by subtracting their column averages. Some methods for predicting missing values are based on the regression approach. Since p is larger than n, common methods as least square or maximum likelihood does not apply. Hence other prediction methods have to be used for the purpose of reducing the predictor space spanned by the (p 1) columns in X (j) to a lower K-dimensional space. This is achieved by constructing K components in the predictor space, where the components optimize a defined object criterion. If observation i has a missing value for gene j, it can generally be predicted by a linear predictor ŷ ij given by ŷ ij = ȳ j + ˆβ t j(x (j)i x (j) ), (1) where ȳ j is the average of the uncentered expression of gene j, x (j)i is a vector of the expressions of all the other genes of the observation with the missing value, and x (j) is a vector containing the average of the uncentered gene expression for each gene except gene j. To simplify the notation we let y = y j, X = X (j), and ˆβ = ˆβ j in the rest of the paper. We will in the following present different methods for computing ˆβ, methods that are
5 Feten et al.: Prediction of Missing Values in Microarray 3 known from other fields to perform well, e.g. chemometrics (Martens and Næs, 1989). The methods predict the missing values by aim of an iterative method based on the idea of the Expectation Maximization (EM) algorithm (Dempster et al., 1977; Wu, 1983). For iteration q, define the matrix W (q) = [ y (q) X ] (q). Since methods based on regression can only be performed on complete matrices, we initially (that is q = 0) set the missing values equal to the average of the non-missing values for each gene. Let q = q + 1, and then for each gene with missing values, compute the vector ˆβ in (1) from y (q 1) and X (q 1). Furthermore, y (q), is produced by replacing the missing values in y with the fitted values from (1), and then update W (q). The computation is repeated until W (q 1) W (q) / W (q 1) is below some threshold, e.g. 10 2, where W = et al. 1999). 2.2 Prediction methods Parametric methods n i=1 p j=1 w2 ij (Hastie In situations where there are more variables than observations, the matrix X t X will be singular, and hence (X t X) 1 does not exist and ordinary least square cannot be used. Hopefully a few linear combinations of the variables will take care of most of the available information in data, and the remaining combinations could be declared as noise and then removed. The regression methods studied in this paper have in common that they express the (p 1)-dimensional vector ˆβ in (1) by ˆβ = R(R t SR) 1 R t s, (2) where S = X t X, s = X t y, and the p K matrix R are specified by the regression methods. The vector ˆβ in (2) can be written as ˆβ = K a k b k. (3) k=1 Geometrically ˆβ is element of a K-dimensional (K p) subspace spanned by the vectors b 1,..., b K. The constants a k and the p 1 vectors b k are specified by the regression methods. The optimal number of components (K) for getting the best prediction, needs to be determined empirically. The topic is discussed by Næs and Helland (1993) and Helland and Almøy (1994). The average The simplest method to predict missing values is to use the average over the uncentered expression values for the associated gene. This is equal to assuming ˆβ = 0. The predictor in (1) is then simplified to ŷ ij = ȳ j. Produced by The Berkeley Electronic Press, 2005
6 4 Statistical Applications in Genetics and Molecular Biology Vol. 4 [2005], No. 1, Article 10 We call the method average. Principal Component Regression (PCR) In Principal Component Analysis (PCA), we will explain as much of the variance-covariance structure as possible through a few linear combinations of the original explanatory variables (Massy, 1965). The PCR method is equivalent to applying these combinations as explanatory variables, and performing a least square regression on the new variables. This transformation eliminates the collinearity between the variables, and the stability of the regression coefficients is increased, while some bias is introduced. The scores of the principal components are given by Z = XE, where E is the orthogonal matrix whose K columns are the eigenvectors of X t X. The eigenvectors contribution to the expression is quantified by the corresponding eigenvalues. We identify the most significant eigenvectors by sorting them based on their corresponding eigenvalues. The components having small eigenvalues are assumed corresponding to noise. We get ˆβ with PCR if we insert R = E = [e 1,e 2,...,e K ] in equation (2), or equivalently insert a k = et k s l k and b k = e k in (3), where l k is the eigenvalue corresponding to the eigenvector e k. In PCR we therefore use fewer variables, but keep the important information. Partial Least Square Regression (PLSR) This method is well known from chemometrics (Martens and Næs, 1989), and has previously been used in classification based on microarray gene expression data (Nguyen and Rocke, 2002b; Ghosh, 2003). It is shown that PLSR is useful as a predictive modelling regression method in the kind of data where there are many more variables than observations (Næs et al., 1986; Höskuldsson, 1988). We receive ˆβ with PLSR by inserting R = [s 1,s 2,...,s K ] in (2), where the vector s k = X t k y k is the non-normalized eigenvector corresponding to the largest eigenvalue of X t k y ky t k X k, and where X k and y k denote the residual after k components, given by y k = y k 1 X k 1 s k 1 (s t k 1S k 1 s k 1 ) 1 s t k 1s k 1, X k = X k 1 X k 1 s k 1 (s t k 1S k 1 s k 1 ) 1 s t k 1S k 1,
7 Feten et al.: Prediction of Missing Values in Microarray 5 where S k 1 = X t k 1 X k 1, X 1 = X, and y 1 = y. There is no simple expression of the a k s and b k s in (3), the exception is one component, then a 1 = st 1s 1 s 1 S 1 s 1 and b 1 = s 1. Factor Analysis Regression (FAR) In contrast to PCR and PLSR, the FAR method is based on a statistical model. Let y be the expression of a gene with missing values, and let x be a vector containing the expressions of the (p 1) other genes. Let us assume that both x and y simultaneously follow the factor analysis model given by (Lawley and Maxwell, 1973) [ ] y = µ + Γf + ǫ, (4) x where µ (p 1) is a vector of constants, Γ (p r) is a matrix of factor loadings, f (r 1) is a vector of factor scores, and ǫ (p 1) is a vector of specific factors with equal variance. Let f N(0,I r ), ǫ N(0,ψI p ), and Cov(f,ǫ) = 0 (r p), then the covariance matrix of [ y x ] t t is given by [ ] γ ΓΓ t t + ψi p+1 = y γ y + ψ γ t yγ t x Γ x γ y Γ x Γ t. (5) x + ψi p More details regarding the factor model can be found in any textbook in multivariate analysis, e.g. Mardia et al. (1979). If we assume that Γ is a matrix with orthogonal columns, then cumbersome, but straightforward calculation gives the following maximum likelihood estimators of the parameters in (5) p ˆψ = (p r) 1 l k = l r, k=r+1 ˆΓ x = H r (L r l r I r ) 0.5, ˆγ y = (L r l r I r ) 0.5 g r. Here l k is the k th largest eigenvalues of ([y,x] t [y,x]), L r is the diagonal matrix with the r largest eigenvalues, and [ ] g r, H t t r is the p r matrix of the corresponding eigenvectors (Almøy, 1994). Assuming (5) we obtain by straightforward matrix calculation the following maximum likelihood regression vector for β = (Γ x Γ t x + ψi) 1 Γ x γ y = Γ x (Γ t xγ x + ψi) 1 γ y as ˆβ = H r (L r l r I r )L 1 r g r [1 gr(l t r l r I r )L 1 r g r ] 1, or expressed as in (3) where a k = g k (1 l K l 1 k )(1 gt K(L K l K I K )L 1 K g K) 1 and b k = h k, with g k being the k th element of g K, and h k being the k th vector of H K. Produced by The Berkeley Electronic Press, 2005
8 6 Statistical Applications in Genetics and Molecular Biology Vol. 4 [2005], No. 1, Article Non-parametric methods K-Nearest Neighbours (KNN) While all the previously presented methods are parametric, another way to predict missing values is by the application of some non-parametric methods. The most used method is K-nearest neighbours, which predict missing values by imputing them with a weighted average of their neighbours (Troyanskaya et al., 2001). The neighbours can be either observations or genes. We assume that there exist one or several groups of coregulated genes, hence we can use genes as neighbours. To define a neighbour we need a distance measure between the object with the missing values of interest and the neighbours. In this paper we have used Euclidean distance based on all pairs of non-missing data. Different objects have different numbers of complete pairs, hence the sum is scaled up proportionally to the number of complete pairs. When observations are neighbours use X = X, and when genes are neighbours use X = X t. The distance between two rows (i and l) is given by p d il = (x ij c x lj )2, (6) il j C il where p is the number of columns in X, c il is the number of complete pairs of rows i and l, C il is the set of complete pairs of rows i and l, x ij is the value in column j and row i, and x lj is the value in column j and row l. The algorithm of K-nearest neighbours uses imputed values from objects close to the object with the missing value (Little and Rubin, 1987). We have used an extended version where neighbours are weighted down according to increasing distance (Troyanskaya et al., 2001). Consider a row i that has one missing value in column j, the KNN method finds K other rows which have a value present in column j, with value most similar to row i. The missing value in column j and row i can then be replaced by using ˆx ij = x l i (i) + (x lj /d il) l i (1/d il), where x (i) is the average value of the uncentered row i, x lj is the l th value in column j, and d il is the Euclidean distance given in (6) (Little and Rubin, 1987). Note that when we use genes as neighbours, we center the observations instead of the variables. 2.3 Validation The original data set was pre-processed by removing the genes containing missing expression values, yielding complete matrices. To create test data sets we deleted randomly between 0.1% and 20% of the data from the
9 Feten et al.: Prediction of Missing Values in Microarray 7 complete matrix. Each method was then used to recover the introduced missing values for the data set. The predicted values were compared to those in the original data set. We determined the method with optimal number of components for every fraction of data missing. For Principal Component Regression, Partial Least Square Regression, and Factor Analysis Regression different numbers of components were used. For K-nearest neighbours, both the number of neighbouring genes and the number of neighbouring observations optimal for prediction were varied. The ability of the predictor is usually evaluated by its expected square loss, θ 2 jk(m) = E j (y j ŷ jk(m) ) 2, interpreted as the long run average square difference between the uncentered expression of gene j and the predicted expression using k components within method m. Since θ is a complicated function of unknown parameters, it can be estimated by the root mean square error of prediction (RMSEP). The commonly used RMSEP is given by 1 RMSEP = j Q t (y ij ŷ ijk(m) ) 2, j j Q i T j where Q is the set of genes with missing values, t j is the number of missing values in gene j, T j is the set of observations with missing values in gene j. Further on, y ij is the observed uncentered expression of observation i in gene j, and ŷ ijk(m) is the predicted value of observation i in gene j using method m with k components/neighbours. Only if all the genes, or the number of missing values is equal for all genes, RMSEP 2 is an unbiased estimate of the average (over genes) prediction error, θk(m) 2 = 1 θ 2 q jk(m), (7) where q is the number of genes with missing values. Instead we have used a modified version of RMSEP, whose square value is an unbiased estimate of θk(m) 2, given by RMSEP k(m) = 1 1 (y ij ŷ ijk(m) ) q t 2. (8) j j Q i T j j Q Comparison of methods In order to compare the different methods for predicting missing values, the predictors may be compared using ideas from CVANOVA (Cross Validation Analysis of Variance) introduced by Indahl and Næs (1998). This method is based on two-way analysis of variance of prediction results obtained using cross-validation. Produced by The Berkeley Electronic Press, 2005
10 8 Statistical Applications in Genetics and Molecular Biology Vol. 4 [2005], No. 1, Article 10 Here we will generalize this method and base the analyses on the mixed model (y ij ŷ ijk(m) ) 2 = µ + τ i + η j + ξ m + δ k(m) + (ηξ) jm + (ηδ) jk(m) + ǫ ijk(m), (9) where µ is the common average, τ i is the effect of the i th observation, η j is the effect of the j th gene, ξ m is a parameter associated with the m th method, δ k(m) is the effect of the k th component within method m, (ηξ) jm is the effect of the interaction between the j th gene and the m th method. Further on, (ηδ) jk(m) is the effect of the interaction between the j th gene and the k th component within method m, and ǫ ijk(m) is a random error component. In the model we will assume η j N(0,σ 2 η), (ηξ) jm N(0,σ 2 ηξ ), and ǫ ijk(m) N(0,σ 2 ǫ). Since we will only apply our conclusions to the observations considered in the analysis, the effect of observation is fixed. The data were analyzed as a mixed model, and the hypotheses that there is effect of method, component(method), gene, gene*method, and gene*component(method) were tested by PROC GLM in SAS (The SAS System for Windows, release 8.02, SAS Institute Inc). These tests are usually robust against deviations from the normal distribution (Lindman, 1992), and the problem of using the non-normal (y ij ŷ ijk(m) ) 2 should be minimal. We could have used the absolute error of prediction, y ij ŷ ijk(m), but since we evaluate the methods by RMSEP, and RMSEP uses the square error of prediction, we do not. 3 Data To demonstrate the different methods for predicting missing values we have used data from two cdna microarray experiments as examples. The first is gene expression data from a study of lipid metabolism in mice focusing on identifying genes with altered expression. A mouse model with very low High Density Lipoprotein (HDL) cholesterol levels was compared to inbred control mouse. The treatment group consisted of 8 mice where the apolipoprotein (apo) AI gene was knocked out. Apo AI is a gene playing an important role in the HDL metabolism. The control group consisted of 8 inbred normal mice. Gene expressions from 5548 genes were measured, including 200 related to lipid metabolism. We removed all genes containing missing values, yielding 5486 genes. We denote this data set apo AI. (Callow et al., 2000.) The eigenvalues of the covariance matrix of apo AI are presented in Figure 1a. The fact that one of the eigenvalues is large compared to the others is an indication of strong correlation structure between the genes. The second example is gene expression data from a study of bacterial response to stress (not published earlier). The reference samples were normal cells of the bacterium Enterococcus faecalis (denoted V583), and the test samples were cells from the same bacterium, but stressed with nisin.
11 Feten et al.: Prediction of Missing Values in Microarray 9 a) b) Figure 1: The eigenvalues of apo AI (a) and V583 (b). There were five replicates of every gene on the array. Gene expressions from 3245 genes were measured. We removed all genes containing missing values, yielding 2744 genes. The eigenvalues of the covariance matrix of V583 are presented in Figure 1b. All the eigenvalues are approximately equal. This is an indication of weak correlation structure between the genes. 4 Results We are interested in knowing whether the results of the methods are dependent of the fraction of missing values. Figure 2 presents the minimum RMSEP for each method at different fractions of missing values. Notice that a) b) Figure 2: The minimum root mean square error of prediction (RMSEP) for different methods with different fractions of data missing for apo AI (a) and V583 (b). Average ( ), FAR ( ), KNNgene ( ), KNNobs ( ), PCR ( ), and PLSR ( ). there seems to be almost no change in RMSEP with increasing number of missing values, except when the fraction of missing values increases from 0.1% to 1%. The differences between Average and the other methods are constant over different fractions of missing values, except 0.1%. From Figure 2a the non-parametric methods (KNN) seem to give best results, with KNN with genes as neighbours as the superior. The method FAR gave approximately the same results as PCR, and was also close to PLSR. All the regression methods gave better results than averaging over the gene. Regarding Figure 2b approximately equal results were obtained by all methods. Produced by The Berkeley Electronic Press, 2005
12 10 Statistical Applications in Genetics and Molecular Biology Vol. 4 [2005], No. 1, Article 10 The number of components leading to the minimum RMSEP (Figure 2), are given in Table 1. For the regression methods the number of components apo AI Method 0.1% 1% 5% 10% 15% 20% PCR PLSR FAR KNNobs KNNgene V583 Method 0.1% 1% 5% 10% 15% 20% PCR PLSR * * FAR KNNobs KNNgene Table 1: Number of components for each method for each fraction of missing values. When 15% and 20% of the values in V583 are missing R t SR in (2) becomes singular for PLSR. at the minimum RMSEP seems to be independent of the fraction of missing values, that means the fraction of missing values is of little importance for the decision of number of components. For the non-parametric methods (KNN) the number of components seems to change with increasing fraction of missing values. Note that minimum RMSEP is obtained by different number of components for the two data sets. Figure 3 shows the behaviour of the methods at different numbers of components/neighbours when 1% of the data are missing. The results of a) b) Figure 3: The RMSEP for different methods with 1% of the data missing for apo AI (a) and V583 (b). Average ( ), FAR ( ), KNNgene (- - -), KNNobs ( - ), PCR ( - - ), and PLSR ( ). the other fractions are not shown here, but these results are almost identical. For V583 we achieved the best results using gene average, while the opposite was the case for apo AI. In the latter case few components for the regression methods gave better results than a large number of components. The methods obtained their minimum with 1-2 components (Table 1). On the
13 Feten et al.: Prediction of Missing Values in Microarray 11 contrary, using KNN the number of neighbours has almost no relevance as long as we use enough neighbours. The results in this paper are based on one realization of the data set at each fraction of missing values. To study the effect of which values are missing, five different realizations of the data set of 1% missing values for apo AI were done (results not shown). The methods and the number of components were ranked in the same way in all the realizations, hence the choice of method seems independent of the data randomly chosen as missing. There were a slight difference in the RMSEP for the different realizations (the standard deviation of RMSEP was approximately 0.012). The standard deviation of RMSEP will decrease with increasing number of missing values (the standard deviation of RMSEP was approximately for 20% missing values), since RMSEP will be based on a higher fraction of the data and different realizations may have more of the missing values in common. To decide if the methods gave significantly different error of prediction, i.e. there exists an optimal method with an optimal number of components, we used the model in equation (9) on responses received from FAR, PCR, PLSR and KNN with observations as neighbours. This model was also used to decide if the optimal method and the optimal number of components were different for each gene. Table 2 presents the p-values for different tests based on this model. (Since square error is probably non-normal, an Source p-value (apo AI) p-value (V583) method < component(method) < < gene < < gene*method < < gene*component(method) > < Table 2: Table of p-values corresponding to the tests based on model (9). analysis based on absolute error was carried out. However the p-values were almost identical (results not shown).) From Table 2 it is clear that both method and component within method were significant. There was also a significant effect of gene and of the interaction between gene and method, which means that the optimal method for one gene is not necessarily the optimal method for another gene. In apo AI we could not prove any effect of interaction between gene and number of components within each method, this is due to large variance inside the genes. In V583 we found this interaction to have significant effect. Produced by The Berkeley Electronic Press, 2005
14 12 Statistical Applications in Genetics and Molecular Biology Vol. 4 [2005], No. 1, Article 10 5 Discussion We have studied different methods for replacing missing values. In addition to previously studied methods; PCR, PLSR, KNN with genes as neighbours, and gene average, we focused on two methods; FAR and KNN with observations as neighbours. In addition we applied a method based on linear mixed models to compare the prediction methods. This study considers situations where data are missing completely at random, i.e. the mechanism responsible for the missing values is not influenced by the values of the variables. Another situation, beyond the scope of this paper, is when data are missing at random, i.e. the missing values are independent of the gene expression, but dependent of some other variables, e.g. the corresponding background value. The methods studied in this paper have been applied to two different data sets, one with relatively strong correlation structure between the genes and one with weak correlation structure between the genes. The strength of the correlation is reflected by the proportions between the eigenvalues of the covariance matrix. If there is strong correlation structure, one or several of the eigenvalues are relatively large. In the case of weak correlation structure, all eigenvalues are approximately equal. In the latter case the average is assumed to be the best predictor. For data with stronger correlation structure, the regression methods or the non-parametric methods should be applied, since they use the information among genes that lies in the correlation. The non-parametric methods (KNN) seem to give best results, with KNN with genes as neighbours as the superior. This can be explained by the assumption that there exist groups of coregulated genes, hence using genes from the same group as neighbours we achieve good prediction. Intuitively we might assume that an increased error of prediction (defined as θ in (7)), and hence RMSEP, should follow from an increasing number of missing values, due to the fact that we have less information. If this is not the case, it indicates that the error of prediction differs for some genes. When 0.1% of the values were missing, approximately 1.5% (apo AI) and 0.5% (V583) of the genes had missing values, and the estimate of the prediction error was sensitive to which genes had missing values due to the effect of gene. When more genes had missing values, the estimate was more robust, and it stabilized. If the prediction ability among the methods for different fractions of missing values was non constant, it would be an indication of an interaction effect between gene and method. In this paper we have studied the ability of different methods to predict missing values in gene expression matrices. However, the final interest is not the predicted values themselves, but rather how they influence on the further analysis. The optimal prediction method is the method that gives similar conclusions as we would obtain by analyzing the original data. Another criteria than RMSEP could have been better suited for this purpose. To improve the prediction for further analysis of the data,
15 Feten et al.: Prediction of Missing Values in Microarray 13 the accuracy of the methods could have been investigated over the range of the expression values (Nguyen et al., 2004). However, this is beyond the scope of this paper. Our final goal is to predict the gene expression values originally missing in the gene expression matrix. Those genes are never used in the calibration of the model, since they were removed to receive a complete matrix. The significant effect of gene indicates that there exists no general error of prediction for all the genes, hence it is hard to estimate the error of prediction for the genes that originally have missing values. In situations with significant effect of the interaction between gene and number of components within method, there exists no number of components/neighbours optimal for prediction of every gene. When predicting the genes that originally missed values, we do not know what method is best for those genes. With no interaction, we assume that the optimal number of components in the calibration is also the best on the genes originally left out. Our study shows that the optimal prediction method and number of components heavily depend on the gene expression matrix, hence there is no general advice for all situations. The first step should always be an investigation of the eigenvalues of the covariance matrix, for the purpose of achieving some knowledge of the correlation structure, and thereby the most suitable prediction methods. Weak correlation structure indicates that average is the optimal method, while stronger correlation structure indicates that regression methods and KNN are better choices. Stronger correlation structure requires a smaller number of components than weaker correlation structure. The best method for each situation can be found by first removing all genes with missing values, yielding a complete matrix. Further on, one removes from the complete matrix approximately the same fraction of data originally missing, and tries out different methods and number of components on the new matrix. The mixed model proposed in (9) can then be used to test which methods and number of components that are best for each gene, and if the differences are significant. A new realization of the data set should be used to estimate the corresponding error of prediction. Finally one finds the combination of methods and components that gives the best prediction and uses it on the original matrix. References [1] Almøy, T. (1994). Prediction methods with many possible explanatory variables. PhD Thesis, Department of Mathematics, University of Oslo, Norway. [2] Buck, S.F. (1960). A method of estimation of missing values in multivariate data suitable for use with an electronic computer. Journal of the Royal Statistical Society Series B - Statistical Methodology, 22, Produced by The Berkeley Electronic Press, 2005
16 14 Statistical Applications in Genetics and Molecular Biology Vol. 4 [2005], No. 1, Article 10 [3] Bø, T.H., Dysvik, B., and Jonassen, I. (2004). LSImpute: accurate estimation of missing values in microarray data with least squares methods. Nucleic Acids Research, 32, e34. [4] Callow, M.J., Dudoit, S., Gong, E.L., Speed, T.P., and Rubin, E.M. (2000). Microarray expression profiling identifies genes with altered expression in HDL deficient mice. Genome Research, 10, [5] Chu, S., DeRisi, J., Eisen, M., Mulholland, J., Botstein, D., Brown, P.O., and Herskowitz, I. (1998). The transcriptional program of sporulation in budding yeast. Science, 282, [6] Dempster, A.P, Laird, N.M, and Rubin, D.B. (1977). Maximum likelihood from incomplete data via the EM algorithm (with Discussion). Journal of the Royal Statistical Society Series B - Statistical Methodology, 39, no. 1, [7] DeRisi, J., Penland, L., Brown, P.O., Bittner, M.L., et al. (1996). Use of a cdna microarray to analyse gene expression patterns in human cancer. Nature Genetics, 14, [8] Efron, B. (1994). Missing data, imputation and the bootstrap. Journal of the American Statistical Association, 89, [9] Eisen, M.B., Spellman, P.T., Brown, P.O., and Botstein, D. (1998). Cluster analysis and display of genome-wide expression patterns. Proceedings of the National Academy of Sciences of the United States of America, 95, [10] Ernst, L.R. (1980). Variance of the estimated mean for several imputation procedures. American Statistical Association, Proceedings of the Survey Research Methods Section, [11] Ghosh, D. (2003). Penalized discriminant methods for the classification of tumors from gene expression data. Biometrics, 59, [12] Hacia, J.G. (1999). Resequencing and mutational analysis using oligonucleotide microarrays. Nature Genetics, 21, [13] Hastie, T., Tibshirani, R., Sherlock, G., Eisen, M., Brown, P., and Botstein, D. (1999). Imputing missing data for gene expression arrays. Technical report, Stanford Statistics Department. [14] Healy, M.J.R., and Westmacott, M. (1956). Missing values in experiments analyzed on automatic computers. Applied Statistics - Journal of the Royal Statistical Society Series C, 5,
17 Feten et al.: Prediction of Missing Values in Microarray 15 [15] Helland, I.S., and Almøy, T. (1994). Comparison of prediction methods when only a few components are relevant. Journal of the American Statistical Association, 89, [16] Höskuldsson, A. (1988). PLS regression methods. Journal of Chemometrics, 2, [17] Indahl, U.G., and Næs, T. (1998). Evaluation of alternative spectral feature extraction methods of textural images for multivariate modeling. Journal of Chemometrics, 12, [18] Lawley, D.N. and Maxwell, A.E. (1973). Regression and factor analysis. Biometrica, 60, [19] Lindman, H.R. (1992). Analysis of Variance in Experimental Design. Springer-Verlag, New York. [20] Little, R.J.A. and Rubin, D.B (1987). Statistical Analysis with Missing Data. Wiley, New York. [21] Mardia, K.V., Kent, J.T. and Bibby, J.M. (1979). Multivariate Analysis. Academic Press, London. [22] Martens, H. and Næs, T. (1989). Multivariate Calibration. John Wiley & Sons, New York. [23] Massy, W.F. (1965). Principal components regression in exploratory statistical research. Journal of the American Statistical Association, 60, [24] Nguyen, D.V., Arpat, A.B., Wang, N., and Carroll, R.J. (2002a). DNA microarray experiments: biological and technological aspects. Biometrics, 58, [25] Nguyen, D.V., and Rocke, D.M. (2002b). Tumor classification by partial least squares using microarray gene expression data. Bioinformatics, 18, [26] Nguyen, D.V., Wang, N., and Carroll, R.J. (2004). Evaluation of missing value estimation for microarray data. Journal of Data Science, 2, [27] Næs, T., and Helland, I.S. (1993). Relevant components in regression. Scandinavian Journal of Statistics, 21, [28] Næs, T., Irgens, C., and Martens, H. (1986). Comparisons of linear statistical methods for calibration of NIR instruments. Applied Statistics - Journal of the Royal Statistical Society Series C, 35, Produced by The Berkeley Electronic Press, 2005
18 16 Statistical Applications in Genetics and Molecular Biology Vol. 4 [2005], No. 1, Article 10 [29] Ouyang, M., Welsh, W.J., and Georgopoulos, P. (2004). Gaussian mixture clustering and imputation of microarray data. Bioinformatics, 20, [30] Rubin, D.B. (1987). Multiple Imputation for Nonresponse in Surveys. Wiley, New York. [31] Rubin, D.B., and Schenker, N. (1991). Multiple imputation in healthcare databases: an overview and some applications. Statistics in Medicine, 10, [32] Troyanskaya, O., Cantor, M., Sherlock, G., Brown, P., Hastie, T., Tibshirani, R., Botstein, D., and Altman, R.B. (2001). Missing value estimation methods for DNA microarrays. Bioinformatics, 17, [33] Wu, C.F.J. (1983). On the convergence of the EM algorithm. Annals of Statistics, 11, [34] Yates, F. (1933). The analysis of replicated experiments when the field results are incomplete. Empire Journal of Experimental Agriculture, 1,
Regression Model In The Analysis Of Micro Array Data-Gene Expression Detection
 Jamal Fathima.J.I 1 and P.Venkatesan 1. Research Scholar -Department of statistics National Institute For Research In Tuberculosis, Indian Council For Medical Research,Chennai,India,.Department of statistics
Jamal Fathima.J.I 1 and P.Venkatesan 1. Research Scholar -Department of statistics National Institute For Research In Tuberculosis, Indian Council For Medical Research,Chennai,India,.Department of statistics
International Journal of Pure and Applied Mathematics Volume 19 No , A NOTE ON BETWEEN-GROUP PCA
 International Journal of Pure and Applied Mathematics Volume 19 No. 3 2005, 359-366 A NOTE ON BETWEEN-GROUP PCA Anne-Laure Boulesteix Department of Statistics University of Munich Akademiestrasse 1, Munich,
International Journal of Pure and Applied Mathematics Volume 19 No. 3 2005, 359-366 A NOTE ON BETWEEN-GROUP PCA Anne-Laure Boulesteix Department of Statistics University of Munich Akademiestrasse 1, Munich,
Principal component analysis (PCA) for clustering gene expression data
 Principal component analysis (PCA) for clustering gene expression data Ka Yee Yeung Walter L. Ruzzo Bioinformatics, v17 #9 (2001) pp 763-774 1 Outline of talk Background and motivation Design of our empirical
Principal component analysis (PCA) for clustering gene expression data Ka Yee Yeung Walter L. Ruzzo Bioinformatics, v17 #9 (2001) pp 763-774 1 Outline of talk Background and motivation Design of our empirical
Topographic Independent Component Analysis of Gene Expression Time Series Data
 Topographic Independent Component Analysis of Gene Expression Time Series Data Sookjeong Kim and Seungjin Choi Department of Computer Science Pohang University of Science and Technology San 31 Hyoja-dong,
Topographic Independent Component Analysis of Gene Expression Time Series Data Sookjeong Kim and Seungjin Choi Department of Computer Science Pohang University of Science and Technology San 31 Hyoja-dong,
Lecture 5: November 19, Minimizing the maximum intracluster distance
 Analysis of DNA Chips and Gene Networks Spring Semester, 2009 Lecture 5: November 19, 2009 Lecturer: Ron Shamir Scribe: Renana Meller 5.1 Minimizing the maximum intracluster distance 5.1.1 Introduction
Analysis of DNA Chips and Gene Networks Spring Semester, 2009 Lecture 5: November 19, 2009 Lecturer: Ron Shamir Scribe: Renana Meller 5.1 Minimizing the maximum intracluster distance 5.1.1 Introduction
Analyzing Microarray Time course Genome wide Data
 OR 779 Functional Data Analysis Course Project Analyzing Microarray Time course Genome wide Data Presented by Xin Zhao April 29, 2002 Cornell University Overview 1. Introduction Biological Background Biological
OR 779 Functional Data Analysis Course Project Analyzing Microarray Time course Genome wide Data Presented by Xin Zhao April 29, 2002 Cornell University Overview 1. Introduction Biological Background Biological
EXTENDING PARTIAL LEAST SQUARES REGRESSION
 EXTENDING PARTIAL LEAST SQUARES REGRESSION ATHANASSIOS KONDYLIS UNIVERSITY OF NEUCHÂTEL 1 Outline Multivariate Calibration in Chemometrics PLS regression (PLSR) and the PLS1 algorithm PLS1 from a statistical
EXTENDING PARTIAL LEAST SQUARES REGRESSION ATHANASSIOS KONDYLIS UNIVERSITY OF NEUCHÂTEL 1 Outline Multivariate Calibration in Chemometrics PLS regression (PLSR) and the PLS1 algorithm PLS1 from a statistical
Fractal functional regression for classification of gene expression data by wavelets
 Fractal functional regression for classification of gene expression data by wavelets Margarita María Rincón 1 and María Dolores Ruiz-Medina 2 1 University of Granada Campus Fuente Nueva 18071 Granada,
Fractal functional regression for classification of gene expression data by wavelets Margarita María Rincón 1 and María Dolores Ruiz-Medina 2 1 University of Granada Campus Fuente Nueva 18071 Granada,
CS281 Section 4: Factor Analysis and PCA
 CS81 Section 4: Factor Analysis and PCA Scott Linderman At this point we have seen a variety of machine learning models, with a particular emphasis on models for supervised learning. In particular, we
CS81 Section 4: Factor Analysis and PCA Scott Linderman At this point we have seen a variety of machine learning models, with a particular emphasis on models for supervised learning. In particular, we
Degenerate Expectation-Maximization Algorithm for Local Dimension Reduction
 Degenerate Expectation-Maximization Algorithm for Local Dimension Reduction Xiaodong Lin 1 and Yu Zhu 2 1 Statistical and Applied Mathematical Science Institute, RTP, NC, 27709 USA University of Cincinnati,
Degenerate Expectation-Maximization Algorithm for Local Dimension Reduction Xiaodong Lin 1 and Yu Zhu 2 1 Statistical and Applied Mathematical Science Institute, RTP, NC, 27709 USA University of Cincinnati,
Bioconductor Project Working Papers
 Bioconductor Project Working Papers Bioconductor Project Year 2004 Paper 6 Error models for microarray intensities Wolfgang Huber Anja von Heydebreck Martin Vingron Department of Molecular Genome Analysis,
Bioconductor Project Working Papers Bioconductor Project Year 2004 Paper 6 Error models for microarray intensities Wolfgang Huber Anja von Heydebreck Martin Vingron Department of Molecular Genome Analysis,
Explaining Correlations by Plotting Orthogonal Contrasts
 Explaining Correlations by Plotting Orthogonal Contrasts Øyvind Langsrud MATFORSK, Norwegian Food Research Institute. www.matforsk.no/ola/ To appear in The American Statistician www.amstat.org/publications/tas/
Explaining Correlations by Plotting Orthogonal Contrasts Øyvind Langsrud MATFORSK, Norwegian Food Research Institute. www.matforsk.no/ola/ To appear in The American Statistician www.amstat.org/publications/tas/
Cluster Analysis of Gene Expression Microarray Data. BIOL 495S/ CS 490B/ MATH 490B/ STAT 490B Introduction to Bioinformatics April 8, 2002
 Cluster Analysis of Gene Expression Microarray Data BIOL 495S/ CS 490B/ MATH 490B/ STAT 490B Introduction to Bioinformatics April 8, 2002 1 Data representations Data are relative measurements log 2 ( red
Cluster Analysis of Gene Expression Microarray Data BIOL 495S/ CS 490B/ MATH 490B/ STAT 490B Introduction to Bioinformatics April 8, 2002 1 Data representations Data are relative measurements log 2 ( red
Alternative implementations of Monte Carlo EM algorithms for likelihood inferences
 Genet. Sel. Evol. 33 001) 443 45 443 INRA, EDP Sciences, 001 Alternative implementations of Monte Carlo EM algorithms for likelihood inferences Louis Alberto GARCÍA-CORTÉS a, Daniel SORENSEN b, Note a
Genet. Sel. Evol. 33 001) 443 45 443 INRA, EDP Sciences, 001 Alternative implementations of Monte Carlo EM algorithms for likelihood inferences Louis Alberto GARCÍA-CORTÉS a, Daniel SORENSEN b, Note a
A Case Study -- Chu et al. The Transcriptional Program of Sporulation in Budding Yeast. What is Sporulation? CSE 527, W.L. Ruzzo 1
 A Case Study -- Chu et al. An interesting early microarray paper My goals Show arrays used in a real experiment Show where computation is important Start looking at analysis techniques The Transcriptional
A Case Study -- Chu et al. An interesting early microarray paper My goals Show arrays used in a real experiment Show where computation is important Start looking at analysis techniques The Transcriptional
From dummy regression to prior probabilities in PLS-DA
 JOURNAL OF CHEMOMETRICS J. Chemometrics (2007) Published online in Wiley InterScience (www.interscience.wiley.com).1061 From dummy regression to prior probabilities in PLS-DA Ulf G. Indahl 1,3, Harald
JOURNAL OF CHEMOMETRICS J. Chemometrics (2007) Published online in Wiley InterScience (www.interscience.wiley.com).1061 From dummy regression to prior probabilities in PLS-DA Ulf G. Indahl 1,3, Harald
Clustering by Mixture Models. General background on clustering Example method: k-means Mixture model based clustering Model estimation
 Clustering by Mixture Models General bacground on clustering Example method: -means Mixture model based clustering Model estimation 1 Clustering A basic tool in data mining/pattern recognition: Divide
Clustering by Mixture Models General bacground on clustering Example method: -means Mixture model based clustering Model estimation 1 Clustering A basic tool in data mining/pattern recognition: Divide
MS-E2112 Multivariate Statistical Analysis (5cr) Lecture 8: Canonical Correlation Analysis
 MS-E2112 Multivariate Statistical (5cr) Lecture 8: Contents Canonical correlation analysis involves partition of variables into two vectors x and y. The aim is to find linear combinations α T x and β
MS-E2112 Multivariate Statistical (5cr) Lecture 8: Contents Canonical correlation analysis involves partition of variables into two vectors x and y. The aim is to find linear combinations α T x and β
Fuzzy Clustering of Gene Expression Data
 Fuzzy Clustering of Gene Data Matthias E. Futschik and Nikola K. Kasabov Department of Information Science, University of Otago P.O. Box 56, Dunedin, New Zealand email: mfutschik@infoscience.otago.ac.nz,
Fuzzy Clustering of Gene Data Matthias E. Futschik and Nikola K. Kasabov Department of Information Science, University of Otago P.O. Box 56, Dunedin, New Zealand email: mfutschik@infoscience.otago.ac.nz,
Uncertainty quantification and visualization for functional random variables
 Uncertainty quantification and visualization for functional random variables MascotNum Workshop 2014 S. Nanty 1,3 C. Helbert 2 A. Marrel 1 N. Pérot 1 C. Prieur 3 1 CEA, DEN/DER/SESI/LSMR, F-13108, Saint-Paul-lez-Durance,
Uncertainty quantification and visualization for functional random variables MascotNum Workshop 2014 S. Nanty 1,3 C. Helbert 2 A. Marrel 1 N. Pérot 1 C. Prieur 3 1 CEA, DEN/DER/SESI/LSMR, F-13108, Saint-Paul-lez-Durance,
Modeling Gene Expression from Microarray Expression Data with State-Space Equations. F.X. Wu, W.J. Zhang, and A.J. Kusalik
 Modeling Gene Expression from Microarray Expression Data with State-Space Equations FX Wu, WJ Zhang, and AJ Kusalik Pacific Symposium on Biocomputing 9:581-592(2004) MODELING GENE EXPRESSION FROM MICROARRAY
Modeling Gene Expression from Microarray Expression Data with State-Space Equations FX Wu, WJ Zhang, and AJ Kusalik Pacific Symposium on Biocomputing 9:581-592(2004) MODELING GENE EXPRESSION FROM MICROARRAY
Forecasting 1 to h steps ahead using partial least squares
 Forecasting 1 to h steps ahead using partial least squares Philip Hans Franses Econometric Institute, Erasmus University Rotterdam November 10, 2006 Econometric Institute Report 2006-47 I thank Dick van
Forecasting 1 to h steps ahead using partial least squares Philip Hans Franses Econometric Institute, Erasmus University Rotterdam November 10, 2006 Econometric Institute Report 2006-47 I thank Dick van
Missing Value Estimation for Time Series Microarray Data Using Linear Dynamical Systems Modeling
 22nd International Conference on Advanced Information Networking and Applications - Workshops Missing Value Estimation for Time Series Microarray Data Using Linear Dynamical Systems Modeling Connie Phong
22nd International Conference on Advanced Information Networking and Applications - Workshops Missing Value Estimation for Time Series Microarray Data Using Linear Dynamical Systems Modeling Connie Phong
Performance Comparison of K-Means and Expectation Maximization with Gaussian Mixture Models for Clustering EE6540 Final Project
 Performance Comparison of K-Means and Expectation Maximization with Gaussian Mixture Models for Clustering EE6540 Final Project Devin Cornell & Sushruth Sastry May 2015 1 Abstract In this article, we explore
Performance Comparison of K-Means and Expectation Maximization with Gaussian Mixture Models for Clustering EE6540 Final Project Devin Cornell & Sushruth Sastry May 2015 1 Abstract In this article, we explore
CMSC858P Supervised Learning Methods
 CMSC858P Supervised Learning Methods Hector Corrada Bravo March, 2010 Introduction Today we discuss the classification setting in detail. Our setting is that we observe for each subject i a set of p predictors
CMSC858P Supervised Learning Methods Hector Corrada Bravo March, 2010 Introduction Today we discuss the classification setting in detail. Our setting is that we observe for each subject i a set of p predictors
Accounting for measurement uncertainties in industrial data analysis
 Accounting for measurement uncertainties in industrial data analysis Marco S. Reis * ; Pedro M. Saraiva GEPSI-PSE Group, Department of Chemical Engineering, University of Coimbra Pólo II Pinhal de Marrocos,
Accounting for measurement uncertainties in industrial data analysis Marco S. Reis * ; Pedro M. Saraiva GEPSI-PSE Group, Department of Chemical Engineering, University of Coimbra Pólo II Pinhal de Marrocos,
December 20, MAA704, Multivariate analysis. Christopher Engström. Multivariate. analysis. Principal component analysis
 .. December 20, 2013 Todays lecture. (PCA) (PLS-R) (LDA) . (PCA) is a method often used to reduce the dimension of a large dataset to one of a more manageble size. The new dataset can then be used to make
.. December 20, 2013 Todays lecture. (PCA) (PLS-R) (LDA) . (PCA) is a method often used to reduce the dimension of a large dataset to one of a more manageble size. The new dataset can then be used to make
Restricted Maximum Likelihood in Linear Regression and Linear Mixed-Effects Model
 Restricted Maximum Likelihood in Linear Regression and Linear Mixed-Effects Model Xiuming Zhang zhangxiuming@u.nus.edu A*STAR-NUS Clinical Imaging Research Center October, 015 Summary This report derives
Restricted Maximum Likelihood in Linear Regression and Linear Mixed-Effects Model Xiuming Zhang zhangxiuming@u.nus.edu A*STAR-NUS Clinical Imaging Research Center October, 015 Summary This report derives
Data Mining Techniques
 Data Mining Techniques CS 622 - Section 2 - Spring 27 Pre-final Review Jan-Willem van de Meent Feedback Feedback https://goo.gl/er7eo8 (also posted on Piazza) Also, please fill out your TRACE evaluations!
Data Mining Techniques CS 622 - Section 2 - Spring 27 Pre-final Review Jan-Willem van de Meent Feedback Feedback https://goo.gl/er7eo8 (also posted on Piazza) Also, please fill out your TRACE evaluations!
Tree-Dependent Components of Gene Expression Data for Clustering
 Tree-Dependent Components of Gene Expression Data for Clustering Jong Kyoung Kim and Seungjin Choi Department of Computer Science Pohang University of Science and Technology San 31 Hyoja-dong, Nam-gu Pohang
Tree-Dependent Components of Gene Expression Data for Clustering Jong Kyoung Kim and Seungjin Choi Department of Computer Science Pohang University of Science and Technology San 31 Hyoja-dong, Nam-gu Pohang
STATISTICAL LEARNING SYSTEMS
 STATISTICAL LEARNING SYSTEMS LECTURE 8: UNSUPERVISED LEARNING: FINDING STRUCTURE IN DATA Institute of Computer Science, Polish Academy of Sciences Ph. D. Program 2013/2014 Principal Component Analysis
STATISTICAL LEARNING SYSTEMS LECTURE 8: UNSUPERVISED LEARNING: FINDING STRUCTURE IN DATA Institute of Computer Science, Polish Academy of Sciences Ph. D. Program 2013/2014 Principal Component Analysis
A Bias Correction for the Minimum Error Rate in Cross-validation
 A Bias Correction for the Minimum Error Rate in Cross-validation Ryan J. Tibshirani Robert Tibshirani Abstract Tuning parameters in supervised learning problems are often estimated by cross-validation.
A Bias Correction for the Minimum Error Rate in Cross-validation Ryan J. Tibshirani Robert Tibshirani Abstract Tuning parameters in supervised learning problems are often estimated by cross-validation.
STATISTICAL ANALYSIS WITH MISSING DATA
 STATISTICAL ANALYSIS WITH MISSING DATA SECOND EDITION Roderick J.A. Little & Donald B. Rubin WILEY SERIES IN PROBABILITY AND STATISTICS Statistical Analysis with Missing Data Second Edition WILEY SERIES
STATISTICAL ANALYSIS WITH MISSING DATA SECOND EDITION Roderick J.A. Little & Donald B. Rubin WILEY SERIES IN PROBABILITY AND STATISTICS Statistical Analysis with Missing Data Second Edition WILEY SERIES
Introduction to Machine Learning. PCA and Spectral Clustering. Introduction to Machine Learning, Slides: Eran Halperin
 1 Introduction to Machine Learning PCA and Spectral Clustering Introduction to Machine Learning, 2013-14 Slides: Eran Halperin Singular Value Decomposition (SVD) The singular value decomposition (SVD)
1 Introduction to Machine Learning PCA and Spectral Clustering Introduction to Machine Learning, 2013-14 Slides: Eran Halperin Singular Value Decomposition (SVD) The singular value decomposition (SVD)
ECE 521. Lecture 11 (not on midterm material) 13 February K-means clustering, Dimensionality reduction
 ECE 521 Lecture 11 (not on midterm material) 13 February 2017 K-means clustering, Dimensionality reduction With thanks to Ruslan Salakhutdinov for an earlier version of the slides Overview K-means clustering
ECE 521 Lecture 11 (not on midterm material) 13 February 2017 K-means clustering, Dimensionality reduction With thanks to Ruslan Salakhutdinov for an earlier version of the slides Overview K-means clustering
Lecture 6: Methods for high-dimensional problems
 Lecture 6: Methods for high-dimensional problems Hector Corrada Bravo and Rafael A. Irizarry March, 2010 In this Section we will discuss methods where data lies on high-dimensional spaces. In particular,
Lecture 6: Methods for high-dimensional problems Hector Corrada Bravo and Rafael A. Irizarry March, 2010 In this Section we will discuss methods where data lies on high-dimensional spaces. In particular,
Drift Reduction For Metal-Oxide Sensor Arrays Using Canonical Correlation Regression And Partial Least Squares
 Drift Reduction For Metal-Oxide Sensor Arrays Using Canonical Correlation Regression And Partial Least Squares R Gutierrez-Osuna Computer Science Department, Wright State University, Dayton, OH 45435,
Drift Reduction For Metal-Oxide Sensor Arrays Using Canonical Correlation Regression And Partial Least Squares R Gutierrez-Osuna Computer Science Department, Wright State University, Dayton, OH 45435,
Streamlining Missing Data Analysis by Aggregating Multiple Imputations at the Data Level
 Streamlining Missing Data Analysis by Aggregating Multiple Imputations at the Data Level A Monte Carlo Simulation to Test the Tenability of the SuperMatrix Approach Kyle M Lang Quantitative Psychology
Streamlining Missing Data Analysis by Aggregating Multiple Imputations at the Data Level A Monte Carlo Simulation to Test the Tenability of the SuperMatrix Approach Kyle M Lang Quantitative Psychology
L11: Pattern recognition principles
 L11: Pattern recognition principles Bayesian decision theory Statistical classifiers Dimensionality reduction Clustering This lecture is partly based on [Huang, Acero and Hon, 2001, ch. 4] Introduction
L11: Pattern recognition principles Bayesian decision theory Statistical classifiers Dimensionality reduction Clustering This lecture is partly based on [Huang, Acero and Hon, 2001, ch. 4] Introduction
Structure in Data. A major objective in data analysis is to identify interesting features or structure in the data.
 Structure in Data A major objective in data analysis is to identify interesting features or structure in the data. The graphical methods are very useful in discovering structure. There are basically two
Structure in Data A major objective in data analysis is to identify interesting features or structure in the data. The graphical methods are very useful in discovering structure. There are basically two
6.047 / Computational Biology: Genomes, Networks, Evolution Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Simultaneous variable selection and class fusion for high-dimensional linear discriminant analysis
 Biostatistics (2010), 11, 4, pp. 599 608 doi:10.1093/biostatistics/kxq023 Advance Access publication on May 26, 2010 Simultaneous variable selection and class fusion for high-dimensional linear discriminant
Biostatistics (2010), 11, 4, pp. 599 608 doi:10.1093/biostatistics/kxq023 Advance Access publication on May 26, 2010 Simultaneous variable selection and class fusion for high-dimensional linear discriminant
The prediction of house price
 000 001 002 003 004 005 006 007 008 009 010 011 012 013 014 015 016 017 018 019 020 021 022 023 024 025 026 027 028 029 030 031 032 033 034 035 036 037 038 039 040 041 042 043 044 045 046 047 048 049 050
000 001 002 003 004 005 006 007 008 009 010 011 012 013 014 015 016 017 018 019 020 021 022 023 024 025 026 027 028 029 030 031 032 033 034 035 036 037 038 039 040 041 042 043 044 045 046 047 048 049 050
Single gene analysis of differential expression
 Single gene analysis of differential expression Giorgio Valentini DSI Dipartimento di Scienze dell Informazione Università degli Studi di Milano valentini@dsi.unimi.it Comparing two conditions Each condition
Single gene analysis of differential expression Giorgio Valentini DSI Dipartimento di Scienze dell Informazione Università degli Studi di Milano valentini@dsi.unimi.it Comparing two conditions Each condition
SPOTTED cdna MICROARRAYS
 SPOTTED cdna MICROARRAYS Spot size: 50um - 150um SPOTTED cdna MICROARRAYS Compare the genetic expression in two samples of cells PRINT cdna from one gene on each spot SAMPLES cdna labelled red/green e.g.
SPOTTED cdna MICROARRAYS Spot size: 50um - 150um SPOTTED cdna MICROARRAYS Compare the genetic expression in two samples of cells PRINT cdna from one gene on each spot SAMPLES cdna labelled red/green e.g.
Principal Component Analysis, A Powerful Scoring Technique
 Principal Component Analysis, A Powerful Scoring Technique George C. J. Fernandez, University of Nevada - Reno, Reno NV 89557 ABSTRACT Data mining is a collection of analytical techniques to uncover new
Principal Component Analysis, A Powerful Scoring Technique George C. J. Fernandez, University of Nevada - Reno, Reno NV 89557 ABSTRACT Data mining is a collection of analytical techniques to uncover new
FACTOR ANALYSIS AS MATRIX DECOMPOSITION 1. INTRODUCTION
 FACTOR ANALYSIS AS MATRIX DECOMPOSITION JAN DE LEEUW ABSTRACT. Meet the abstract. This is the abstract. 1. INTRODUCTION Suppose we have n measurements on each of taking m variables. Collect these measurements
FACTOR ANALYSIS AS MATRIX DECOMPOSITION JAN DE LEEUW ABSTRACT. Meet the abstract. This is the abstract. 1. INTRODUCTION Suppose we have n measurements on each of taking m variables. Collect these measurements
10-810: Advanced Algorithms and Models for Computational Biology. Optimal leaf ordering and classification
 10-810: Advanced Algorithms and Models for Computational Biology Optimal leaf ordering and classification Hierarchical clustering As we mentioned, its one of the most popular methods for clustering gene
10-810: Advanced Algorithms and Models for Computational Biology Optimal leaf ordering and classification Hierarchical clustering As we mentioned, its one of the most popular methods for clustering gene
1 Data Arrays and Decompositions
 1 Data Arrays and Decompositions 1.1 Variance Matrices and Eigenstructure Consider a p p positive definite and symmetric matrix V - a model parameter or a sample variance matrix. The eigenstructure is
1 Data Arrays and Decompositions 1.1 Variance Matrices and Eigenstructure Consider a p p positive definite and symmetric matrix V - a model parameter or a sample variance matrix. The eigenstructure is
Estimating the parameters of hidden binomial trials by the EM algorithm
 Hacettepe Journal of Mathematics and Statistics Volume 43 (5) (2014), 885 890 Estimating the parameters of hidden binomial trials by the EM algorithm Degang Zhu Received 02 : 09 : 2013 : Accepted 02 :
Hacettepe Journal of Mathematics and Statistics Volume 43 (5) (2014), 885 890 Estimating the parameters of hidden binomial trials by the EM algorithm Degang Zhu Received 02 : 09 : 2013 : Accepted 02 :
Experimental Design and Data Analysis for Biologists
 Experimental Design and Data Analysis for Biologists Gerry P. Quinn Monash University Michael J. Keough University of Melbourne CAMBRIDGE UNIVERSITY PRESS Contents Preface page xv I I Introduction 1 1.1
Experimental Design and Data Analysis for Biologists Gerry P. Quinn Monash University Michael J. Keough University of Melbourne CAMBRIDGE UNIVERSITY PRESS Contents Preface page xv I I Introduction 1 1.1
Factor Analysis (10/2/13)
 STA561: Probabilistic machine learning Factor Analysis (10/2/13) Lecturer: Barbara Engelhardt Scribes: Li Zhu, Fan Li, Ni Guan Factor Analysis Factor analysis is related to the mixture models we have studied.
STA561: Probabilistic machine learning Factor Analysis (10/2/13) Lecturer: Barbara Engelhardt Scribes: Li Zhu, Fan Li, Ni Guan Factor Analysis Factor analysis is related to the mixture models we have studied.
Data Mining. Dimensionality reduction. Hamid Beigy. Sharif University of Technology. Fall 1395
 Data Mining Dimensionality reduction Hamid Beigy Sharif University of Technology Fall 1395 Hamid Beigy (Sharif University of Technology) Data Mining Fall 1395 1 / 42 Outline 1 Introduction 2 Feature selection
Data Mining Dimensionality reduction Hamid Beigy Sharif University of Technology Fall 1395 Hamid Beigy (Sharif University of Technology) Data Mining Fall 1395 1 / 42 Outline 1 Introduction 2 Feature selection
ECE 661: Homework 10 Fall 2014
 ECE 661: Homework 10 Fall 2014 This homework consists of the following two parts: (1) Face recognition with PCA and LDA for dimensionality reduction and the nearest-neighborhood rule for classification;
ECE 661: Homework 10 Fall 2014 This homework consists of the following two parts: (1) Face recognition with PCA and LDA for dimensionality reduction and the nearest-neighborhood rule for classification;
Dimensional reduction of clustered data sets
 Dimensional reduction of clustered data sets Guido Sanguinetti 5th February 2007 Abstract We present a novel probabilistic latent variable model to perform linear dimensional reduction on data sets which
Dimensional reduction of clustered data sets Guido Sanguinetti 5th February 2007 Abstract We present a novel probabilistic latent variable model to perform linear dimensional reduction on data sets which
c Springer, Reprinted with permission.
 Zhijian Yuan and Erkki Oja. A FastICA Algorithm for Non-negative Independent Component Analysis. In Puntonet, Carlos G.; Prieto, Alberto (Eds.), Proceedings of the Fifth International Symposium on Independent
Zhijian Yuan and Erkki Oja. A FastICA Algorithm for Non-negative Independent Component Analysis. In Puntonet, Carlos G.; Prieto, Alberto (Eds.), Proceedings of the Fifth International Symposium on Independent
Dimension reduction, PCA & eigenanalysis Based in part on slides from textbook, slides of Susan Holmes. October 3, Statistics 202: Data Mining
 Dimension reduction, PCA & eigenanalysis Based in part on slides from textbook, slides of Susan Holmes October 3, 2012 1 / 1 Combinations of features Given a data matrix X n p with p fairly large, it can
Dimension reduction, PCA & eigenanalysis Based in part on slides from textbook, slides of Susan Holmes October 3, 2012 1 / 1 Combinations of features Given a data matrix X n p with p fairly large, it can
ReducedPCR/PLSRmodelsbysubspaceprojections
 ReducedPCR/PLSRmodelsbysubspaceprojections Rolf Ergon Telemark University College P.O.Box 2, N-9 Porsgrunn, Norway e-mail: rolf.ergon@hit.no Published in Chemometrics and Intelligent Laboratory Systems
ReducedPCR/PLSRmodelsbysubspaceprojections Rolf Ergon Telemark University College P.O.Box 2, N-9 Porsgrunn, Norway e-mail: rolf.ergon@hit.no Published in Chemometrics and Intelligent Laboratory Systems
Expression arrays, normalization, and error models
 1 Epression arrays, normalization, and error models There are a number of different array technologies available for measuring mrna transcript levels in cell populations, from spotted cdna arrays to in
1 Epression arrays, normalization, and error models There are a number of different array technologies available for measuring mrna transcript levels in cell populations, from spotted cdna arrays to in
Singular Value Decomposition and Principal Component Analysis (PCA) I
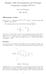 Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Cross model validation and multiple testing in latent variable models
 Cross model validation and multiple testing in latent variable models Frank Westad GE Healthcare Oslo, Norway 2nd European User Meeting on Multivariate Analysis Como, June 22, 2006 Outline Introduction
Cross model validation and multiple testing in latent variable models Frank Westad GE Healthcare Oslo, Norway 2nd European User Meeting on Multivariate Analysis Como, June 22, 2006 Outline Introduction
MATH 829: Introduction to Data Mining and Analysis Principal component analysis
 1/11 MATH 829: Introduction to Data Mining and Analysis Principal component analysis Dominique Guillot Departments of Mathematical Sciences University of Delaware April 4, 2016 Motivation 2/11 High-dimensional
1/11 MATH 829: Introduction to Data Mining and Analysis Principal component analysis Dominique Guillot Departments of Mathematical Sciences University of Delaware April 4, 2016 Motivation 2/11 High-dimensional
Basics of Multivariate Modelling and Data Analysis
 Basics of Multivariate Modelling and Data Analysis Kurt-Erik Häggblom 6. Principal component analysis (PCA) 6.1 Overview 6.2 Essentials of PCA 6.3 Numerical calculation of PCs 6.4 Effects of data preprocessing
Basics of Multivariate Modelling and Data Analysis Kurt-Erik Häggblom 6. Principal component analysis (PCA) 6.1 Overview 6.2 Essentials of PCA 6.3 Numerical calculation of PCs 6.4 Effects of data preprocessing
Linear Models 1. Isfahan University of Technology Fall Semester, 2014
 Linear Models 1 Isfahan University of Technology Fall Semester, 2014 References: [1] G. A. F., Seber and A. J. Lee (2003). Linear Regression Analysis (2nd ed.). Hoboken, NJ: Wiley. [2] A. C. Rencher and
Linear Models 1 Isfahan University of Technology Fall Semester, 2014 References: [1] G. A. F., Seber and A. J. Lee (2003). Linear Regression Analysis (2nd ed.). Hoboken, NJ: Wiley. [2] A. C. Rencher and
Fractional Hot Deck Imputation for Robust Inference Under Item Nonresponse in Survey Sampling
 Fractional Hot Deck Imputation for Robust Inference Under Item Nonresponse in Survey Sampling Jae-Kwang Kim 1 Iowa State University June 26, 2013 1 Joint work with Shu Yang Introduction 1 Introduction
Fractional Hot Deck Imputation for Robust Inference Under Item Nonresponse in Survey Sampling Jae-Kwang Kim 1 Iowa State University June 26, 2013 1 Joint work with Shu Yang Introduction 1 Introduction
Local Polynomial Wavelet Regression with Missing at Random
 Applied Mathematical Sciences, Vol. 6, 2012, no. 57, 2805-2819 Local Polynomial Wavelet Regression with Missing at Random Alsaidi M. Altaher School of Mathematical Sciences Universiti Sains Malaysia 11800
Applied Mathematical Sciences, Vol. 6, 2012, no. 57, 2805-2819 Local Polynomial Wavelet Regression with Missing at Random Alsaidi M. Altaher School of Mathematical Sciences Universiti Sains Malaysia 11800
Shrinkage regression
 Shrinkage regression Rolf Sundberg Volume 4, pp 1994 1998 in Encyclopedia of Environmetrics (ISBN 0471 899976) Edited by Abdel H. El-Shaarawi and Walter W. Piegorsch John Wiley & Sons, Ltd, Chichester,
Shrinkage regression Rolf Sundberg Volume 4, pp 1994 1998 in Encyclopedia of Environmetrics (ISBN 0471 899976) Edited by Abdel H. El-Shaarawi and Walter W. Piegorsch John Wiley & Sons, Ltd, Chichester,
Feature selection and classifier performance in computer-aided diagnosis: The effect of finite sample size
 Feature selection and classifier performance in computer-aided diagnosis: The effect of finite sample size Berkman Sahiner, a) Heang-Ping Chan, Nicholas Petrick, Robert F. Wagner, b) and Lubomir Hadjiiski
Feature selection and classifier performance in computer-aided diagnosis: The effect of finite sample size Berkman Sahiner, a) Heang-Ping Chan, Nicholas Petrick, Robert F. Wagner, b) and Lubomir Hadjiiski
Regression, Ridge Regression, Lasso
 Regression, Ridge Regression, Lasso Fabio G. Cozman - fgcozman@usp.br October 2, 2018 A general definition Regression studies the relationship between a response variable Y and covariates X 1,..., X n.
Regression, Ridge Regression, Lasso Fabio G. Cozman - fgcozman@usp.br October 2, 2018 A general definition Regression studies the relationship between a response variable Y and covariates X 1,..., X n.
PERFORMANCE OF THE EM ALGORITHM ON THE IDENTIFICATION OF A MIXTURE OF WAT- SON DISTRIBUTIONS DEFINED ON THE HYPER- SPHERE
 REVSTAT Statistical Journal Volume 4, Number 2, June 2006, 111 130 PERFORMANCE OF THE EM ALGORITHM ON THE IDENTIFICATION OF A MIXTURE OF WAT- SON DISTRIBUTIONS DEFINED ON THE HYPER- SPHERE Authors: Adelaide
REVSTAT Statistical Journal Volume 4, Number 2, June 2006, 111 130 PERFORMANCE OF THE EM ALGORITHM ON THE IDENTIFICATION OF A MIXTURE OF WAT- SON DISTRIBUTIONS DEFINED ON THE HYPER- SPHERE Authors: Adelaide
Regularized Discriminant Analysis. Part I. Linear and Quadratic Discriminant Analysis. Discriminant Analysis. Example. Example. Class distribution
 Part I 09.06.2006 Discriminant Analysis The purpose of discriminant analysis is to assign objects to one of several (K) groups based on a set of measurements X = (X 1, X 2,..., X p ) which are obtained
Part I 09.06.2006 Discriminant Analysis The purpose of discriminant analysis is to assign objects to one of several (K) groups based on a set of measurements X = (X 1, X 2,..., X p ) which are obtained
Expression Data Exploration: Association, Patterns, Factors & Regression Modelling
 Expression Data Exploration: Association, Patterns, Factors & Regression Modelling Exploring gene expression data Scale factors, median chip correlation on gene subsets for crude data quality investigation
Expression Data Exploration: Association, Patterns, Factors & Regression Modelling Exploring gene expression data Scale factors, median chip correlation on gene subsets for crude data quality investigation
Multiple Change-Point Detection and Analysis of Chromosome Copy Number Variations
 Multiple Change-Point Detection and Analysis of Chromosome Copy Number Variations Yale School of Public Health Joint work with Ning Hao, Yue S. Niu presented @Tsinghua University Outline 1 The Problem
Multiple Change-Point Detection and Analysis of Chromosome Copy Number Variations Yale School of Public Health Joint work with Ning Hao, Yue S. Niu presented @Tsinghua University Outline 1 The Problem
Sparse Linear Models (10/7/13)
 STA56: Probabilistic machine learning Sparse Linear Models (0/7/) Lecturer: Barbara Engelhardt Scribes: Jiaji Huang, Xin Jiang, Albert Oh Sparsity Sparsity has been a hot topic in statistics and machine
STA56: Probabilistic machine learning Sparse Linear Models (0/7/) Lecturer: Barbara Engelhardt Scribes: Jiaji Huang, Xin Jiang, Albert Oh Sparsity Sparsity has been a hot topic in statistics and machine
Principal Components Analysis (PCA) and Singular Value Decomposition (SVD) with applications to Microarrays
 Principal Components Analysis (PCA) and Singular Value Decomposition (SVD) with applications to Microarrays Prof. Tesler Math 283 Fall 2015 Prof. Tesler Principal Components Analysis Math 283 / Fall 2015
Principal Components Analysis (PCA) and Singular Value Decomposition (SVD) with applications to Microarrays Prof. Tesler Math 283 Fall 2015 Prof. Tesler Principal Components Analysis Math 283 / Fall 2015
Improving the identification of differentially expressed genes in cdna microarray experiments
 Improving the identification of differentially expressed genes in cdna microarray experiments Databionics Research Group University of Marburg 33 Marburg, Germany Alfred Ultsch Abstract. The identification
Improving the identification of differentially expressed genes in cdna microarray experiments Databionics Research Group University of Marburg 33 Marburg, Germany Alfred Ultsch Abstract. The identification
Lecture 1: Systems of linear equations and their solutions
 Lecture 1: Systems of linear equations and their solutions Course overview Topics to be covered this semester: Systems of linear equations and Gaussian elimination: Solving linear equations and applications
Lecture 1: Systems of linear equations and their solutions Course overview Topics to be covered this semester: Systems of linear equations and Gaussian elimination: Solving linear equations and applications
MIXTURE OF EXPERTS ARCHITECTURES FOR NEURAL NETWORKS AS A SPECIAL CASE OF CONDITIONAL EXPECTATION FORMULA
 MIXTURE OF EXPERTS ARCHITECTURES FOR NEURAL NETWORKS AS A SPECIAL CASE OF CONDITIONAL EXPECTATION FORMULA Jiří Grim Department of Pattern Recognition Institute of Information Theory and Automation Academy
MIXTURE OF EXPERTS ARCHITECTURES FOR NEURAL NETWORKS AS A SPECIAL CASE OF CONDITIONAL EXPECTATION FORMULA Jiří Grim Department of Pattern Recognition Institute of Information Theory and Automation Academy
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
Permutation-invariant regularization of large covariance matrices. Liza Levina
 Liza Levina Permutation-invariant covariance regularization 1/42 Permutation-invariant regularization of large covariance matrices Liza Levina Department of Statistics University of Michigan Joint work
Liza Levina Permutation-invariant covariance regularization 1/42 Permutation-invariant regularization of large covariance matrices Liza Levina Department of Statistics University of Michigan Joint work
Genetic Networks. Korbinian Strimmer. Seminar: Statistical Analysis of RNA-Seq Data 19 June IMISE, Universität Leipzig
 Genetic Networks Korbinian Strimmer IMISE, Universität Leipzig Seminar: Statistical Analysis of RNA-Seq Data 19 June 2012 Korbinian Strimmer, RNA-Seq Networks, 19/6/2012 1 Paper G. I. Allen and Z. Liu.
Genetic Networks Korbinian Strimmer IMISE, Universität Leipzig Seminar: Statistical Analysis of RNA-Seq Data 19 June 2012 Korbinian Strimmer, RNA-Seq Networks, 19/6/2012 1 Paper G. I. Allen and Z. Liu.
Abstract. comment reviews reports deposited research refereed research interactions information
 http://genomebiology.com/2002/3/5/research/0022.1 Research How many replicates of arrays are required to detect gene expression changes in microarray experiments? A mixture model approach Wei Pan*, Jizhen
http://genomebiology.com/2002/3/5/research/0022.1 Research How many replicates of arrays are required to detect gene expression changes in microarray experiments? A mixture model approach Wei Pan*, Jizhen
Lecture 16: Small Sample Size Problems (Covariance Estimation) Many thanks to Carlos Thomaz who authored the original version of these slides
 Lecture 16: Small Sample Size Problems (Covariance Estimation) Many thanks to Carlos Thomaz who authored the original version of these slides Intelligent Data Analysis and Probabilistic Inference Lecture
Lecture 16: Small Sample Size Problems (Covariance Estimation) Many thanks to Carlos Thomaz who authored the original version of these slides Intelligent Data Analysis and Probabilistic Inference Lecture
Determining the number of components in mixture models for hierarchical data
 Determining the number of components in mixture models for hierarchical data Olga Lukočienė 1 and Jeroen K. Vermunt 2 1 Department of Methodology and Statistics, Tilburg University, P.O. Box 90153, 5000
Determining the number of components in mixture models for hierarchical data Olga Lukočienė 1 and Jeroen K. Vermunt 2 1 Department of Methodology and Statistics, Tilburg University, P.O. Box 90153, 5000
FAST CROSS-VALIDATION IN ROBUST PCA
 COMPSTAT 2004 Symposium c Physica-Verlag/Springer 2004 FAST CROSS-VALIDATION IN ROBUST PCA Sanne Engelen, Mia Hubert Key words: Cross-Validation, Robustness, fast algorithm COMPSTAT 2004 section: Partial
COMPSTAT 2004 Symposium c Physica-Verlag/Springer 2004 FAST CROSS-VALIDATION IN ROBUST PCA Sanne Engelen, Mia Hubert Key words: Cross-Validation, Robustness, fast algorithm COMPSTAT 2004 section: Partial
Generalized Elastic Net Regression
 Abstract Generalized Elastic Net Regression Geoffroy MOURET Jean-Jules BRAULT Vahid PARTOVINIA This work presents a variation of the elastic net penalization method. We propose applying a combined l 1
Abstract Generalized Elastic Net Regression Geoffroy MOURET Jean-Jules BRAULT Vahid PARTOVINIA This work presents a variation of the elastic net penalization method. We propose applying a combined l 1
14 Singular Value Decomposition
 14 Singular Value Decomposition For any high-dimensional data analysis, one s first thought should often be: can I use an SVD? The singular value decomposition is an invaluable analysis tool for dealing
14 Singular Value Decomposition For any high-dimensional data analysis, one s first thought should often be: can I use an SVD? The singular value decomposition is an invaluable analysis tool for dealing
2.3. Clustering or vector quantization 57
 Multivariate Statistics non-negative matrix factorisation and sparse dictionary learning The PCA decomposition is by construction optimal solution to argmin A R n q,h R q p X AH 2 2 under constraint :
Multivariate Statistics non-negative matrix factorisation and sparse dictionary learning The PCA decomposition is by construction optimal solution to argmin A R n q,h R q p X AH 2 2 under constraint :
Biochip informatics-(i)
 Biochip informatics-(i) : biochip normalization & differential expression Ju Han Kim, M.D., Ph.D. SNUBI: SNUBiomedical Informatics http://www.snubi snubi.org/ Biochip Informatics - (I) Biochip basics Preprocessing
Biochip informatics-(i) : biochip normalization & differential expression Ju Han Kim, M.D., Ph.D. SNUBI: SNUBiomedical Informatics http://www.snubi snubi.org/ Biochip Informatics - (I) Biochip basics Preprocessing
A Multivariate Two-Sample Mean Test for Small Sample Size and Missing Data
 A Multivariate Two-Sample Mean Test for Small Sample Size and Missing Data Yujun Wu, Marc G. Genton, 1 and Leonard A. Stefanski 2 Department of Biostatistics, School of Public Health, University of Medicine
A Multivariate Two-Sample Mean Test for Small Sample Size and Missing Data Yujun Wu, Marc G. Genton, 1 and Leonard A. Stefanski 2 Department of Biostatistics, School of Public Health, University of Medicine
Factor Analysis. Robert L. Wolpert Department of Statistical Science Duke University, Durham, NC, USA
 Factor Analysis Robert L. Wolpert Department of Statistical Science Duke University, Durham, NC, USA 1 Factor Models The multivariate regression model Y = XB +U expresses each row Y i R p as a linear combination
Factor Analysis Robert L. Wolpert Department of Statistical Science Duke University, Durham, NC, USA 1 Factor Models The multivariate regression model Y = XB +U expresses each row Y i R p as a linear combination
Statistical Applications in Genetics and Molecular Biology
 Statistical Applications in Genetics and Molecular Biology Volume 5, Issue 1 2006 Article 28 A Two-Step Multiple Comparison Procedure for a Large Number of Tests and Multiple Treatments Hongmei Jiang Rebecca
Statistical Applications in Genetics and Molecular Biology Volume 5, Issue 1 2006 Article 28 A Two-Step Multiple Comparison Procedure for a Large Number of Tests and Multiple Treatments Hongmei Jiang Rebecca
Sparse statistical modelling
 Sparse statistical modelling Tom Bartlett Sparse statistical modelling Tom Bartlett 1 / 28 Introduction A sparse statistical model is one having only a small number of nonzero parameters or weights. [1]
Sparse statistical modelling Tom Bartlett Sparse statistical modelling Tom Bartlett 1 / 28 Introduction A sparse statistical model is one having only a small number of nonzero parameters or weights. [1]
Bi-level feature selection with applications to genetic association
 Bi-level feature selection with applications to genetic association studies October 15, 2008 Motivation In many applications, biological features possess a grouping structure Categorical variables may
Bi-level feature selection with applications to genetic association studies October 15, 2008 Motivation In many applications, biological features possess a grouping structure Categorical variables may
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
MULTICHANNEL SIGNAL PROCESSING USING SPATIAL RANK COVARIANCE MATRICES
 MULTICHANNEL SIGNAL PROCESSING USING SPATIAL RANK COVARIANCE MATRICES S. Visuri 1 H. Oja V. Koivunen 1 1 Signal Processing Lab. Dept. of Statistics Tampere Univ. of Technology University of Jyväskylä P.O.
MULTICHANNEL SIGNAL PROCESSING USING SPATIAL RANK COVARIANCE MATRICES S. Visuri 1 H. Oja V. Koivunen 1 1 Signal Processing Lab. Dept. of Statistics Tampere Univ. of Technology University of Jyväskylä P.O.
Effective Linear Discriminant Analysis for High Dimensional, Low Sample Size Data
 Effective Linear Discriant Analysis for High Dimensional, Low Sample Size Data Zhihua Qiao, Lan Zhou and Jianhua Z. Huang Abstract In the so-called high dimensional, low sample size (HDLSS) settings, LDA
Effective Linear Discriant Analysis for High Dimensional, Low Sample Size Data Zhihua Qiao, Lan Zhou and Jianhua Z. Huang Abstract In the so-called high dimensional, low sample size (HDLSS) settings, LDA
Low-Level Analysis of High- Density Oligonucleotide Microarray Data
 Low-Level Analysis of High- Density Oligonucleotide Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley UC Berkeley Feb 23, 2004 Outline
Low-Level Analysis of High- Density Oligonucleotide Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley UC Berkeley Feb 23, 2004 Outline
Chemometrics. Matti Hotokka Physical chemistry Åbo Akademi University
 Chemometrics Matti Hotokka Physical chemistry Åbo Akademi University Linear regression Experiment Consider spectrophotometry as an example Beer-Lamberts law: A = cå Experiment Make three known references
Chemometrics Matti Hotokka Physical chemistry Åbo Akademi University Linear regression Experiment Consider spectrophotometry as an example Beer-Lamberts law: A = cå Experiment Make three known references
TESTING FOR EQUAL DISTRIBUTIONS IN HIGH DIMENSION
 TESTING FOR EQUAL DISTRIBUTIONS IN HIGH DIMENSION Gábor J. Székely Bowling Green State University Maria L. Rizzo Ohio University October 30, 2004 Abstract We propose a new nonparametric test for equality
TESTING FOR EQUAL DISTRIBUTIONS IN HIGH DIMENSION Gábor J. Székely Bowling Green State University Maria L. Rizzo Ohio University October 30, 2004 Abstract We propose a new nonparametric test for equality
