Computer exercise INLA
|
|
|
- Clinton Claude Patrick
- 6 years ago
- Views:
Transcription
1 SARMA 2012 Sharp statistical tools Computer exercise INLA Loading R-packages Load the needed packages: > library(inla) > library(plotrix) > library(geor) > library(fields) > library(maps) > library(spatstat) You might need to install INLA > source(" Loading the data As an example we ll study the locations of 3605 trees in a tropical rain forest. The data comes with covariate data giving the elevation (altitude) and slope of elevation in the study region. The data could be analysed as a spatial-point process, but we ll simplify matters and compute the number of trees in each grid cell. The data is then modelled using a Poisson-process with spatially varying mean. > ##Load and examine the data > data(bei) > plot(bei) > ##extract elevation and gradient > elevation <- bei.extra$elev$v > gradient <- bei.extra$grad$v > ##compute counts in each grid-cell > counts <- matrix(0,dim(gradient)[1],dim(gradient)[2]) > ##size of each grid cell > dx <- (bei.extra$elev$xcol[2] - bei.extra$elev$xcol[1])/2 > dy <- (bei.extra$elev$yrow[2] - bei.extra$elev$yrow[1])/2 > ##loop over the grid cells, and count points > for(i in 1:dim(counts)[1]){ Y <- bei.extra$grad$yrow[i] for(j in 1:dim(counts)[2]){ X <- bei.extra$grad$xcol[j] counts[i,j] <- sum( (X-dx)<bei$x & bei$x<=(x+dy) & (Y-dy)<bei$y & bei$y<=(y+dx) ) } } Lets plot the resulting gridded data, > par(mfrow=c(2,2)) > image.plot(elevation) > image.plot(gradient) > image.plot(counts) and collect the data into a suitable data-frame > bei <- data.frame(counts=c(counts), x=rep(bei.extra$elev$xcol, each=length(bei.extra$elev$yrow)), y=rep(bei.extra$elev$yrow, length(bei.extra$elev$xcol)), elevation=c(elevation), gradient=c(gradient))
2 SARMA 2012 Sharp statistical tools 2 Triangulation The first step in creating a Gaussian Markov random field solution to the Stochastic Partial Differential Equation, as described in the lecture notes, is to create a triangular mesh over the observation locations. We could create a mesh for the lattice > mesh.lattice <- inla.mesh.lattice(x=bei.extra$elev$xcol, y=bei.extra$elev$yrow) > mesh.lattice <- inla.mesh.create( lattice=mesh.lattice ) > plot( mesh.lattice ) > summary( mesh.lattice ) Howeverthismesh ismuch denserthanneeded, insteadwe computeamesh with a suitable internal triangulation (max edge length 50) and coarser extension. > loc <- as.matrix(bei[,c("x","y")]) > mesh <- inla.mesh.create.helper(points.domain=loc, offset = c(-0.05, -0.2), n=c(8,16), max.edge = c(25,150)) > summary(mesh) > ##plot > plot(mesh) > ##add the points for reference (note the possible aliasing effect) > points(loc[,1], loc[,2], col="red", pch=19, cex=.1) Precision matrices Given the mesh the next step is to compute the matrices needed to create the precision matrix Q, and the observation matrix needed to link the mesh to our observations locations. > spde <- inla.spde2.matern(mesh, alpha=2) > A <- inla.spde.make.a(mesh, loc) The resulting matrices are sparse. > image( spde$param.inla$m0 ) > image( spde$param.inla$m1 ) > image( spde$param.inla$m2 ) > image(a) > ##or only the first 1000 observations > image(a[1:1000,]) In addition to these we also need an index vector for the latent field > spde.idx <- inla.spde.make.index("spatial", n.mesh=mesh$n) Setting up the model Thenextstepistocreatealistof effectsandaninla-stackobjectcontaining the relevant components of the model. For numerical reasons we select to put the intercept with the field; the second component of the effects contains all possible covariates. The stack object combines the observations, observations matrices, effects, and a tag which can be used to identify components of the posterior. > effects <- list(c(list(intercept=rep(1,mesh$n)), spde.idx), covar=bei[,c("x","y","elevation","gradient")]) > stack.obs <- inla.stack(data=list(y=bei$counts), A=list(A, 1), effects=effects, tag="obs") The stack must contain one observation matrix for each effect. Here the 1 indicates an identity matrix linking the covariates (second element in effects) with the observations. Predictions at unobserved locations can be added by creating a stack without observations, i.e. Y = NA.
3 SARMA 2012 Sharp statistical tools 3 The model The model defined in stack.obs above can be written as y i Po(e zi ) Z = Aw +IBβw N(0,Q)β N ( 0,I 10 2) Thus we have Poissonobservations, y i ; with intensity, z i ; determined by a latent field, w; linked to the observations location through an observation matrix, A; and with a mean consisting of covariates, B; and with a Normal-prior for the regresison coefficients. Strictly speaking the model only defines things up to z i, the type of observations are defined in the call to inla in a few paragraphs. In principal the latent field is expanded to Z = [ A B ][ ] w β [ ( [0 ] [ ] 1 ) w Qspde 0 N, β] 0 0 I 10 2, Priors The default INLA priors can be extracted, and changing the values is straight forward. For the poission observations we have > hyper.obs <- inla.models()$likelihood$poisson$hyper > hyper.obs Which is rather uninteresting. The prior for Gaussian observations has a little more to offer > hyper.gaussian <- inla.models()$likelihood$gaussian$hyper > str(hyper.gaussian) Even the spatial model (the spde-object) comes with a default prior; the range parameter has been chosen based on the coordinates of the mesh. > spde$param.inla$theta.mu > spde$param.inla$theta.q Finally the precision of the regression coefficients, β, can be altered from the predetermined values through the control.fixed option in the call to inla, see. >?control.fixed Running INLA Having defined the model, data structure, initial values, and prior distributions we are ready to estimate the model. We try both a simple, and a more complex mean. In the following the model is defined using the standard notation in R with the names refering to elements in stack.obs. family="poisson" tells INLA that our data should be considered as Poisson observations of the latent field. Other possible observation models are: > names(inla.models()$likelihood) Finally control.family allows us alter the priors of the observation likelihood. DO NOT RUN!!! Estimation of the two models. > res1 <- inla(y ~ -1 + intercept + f(spatial, model=spde), family="poisson", data = inla.stack.data(stack.obs), control.family = list(hyper = hyper.obs), control.predictor = list(a=inla.stack.a(stack.obs),
4 SARMA 2012 Sharp statistical tools 4 compute=true), control.results=list(return.marginals.random=false, return.marginals.predictor=false), verbose=true) > res2 <- inla(y ~ -1 + intercept + elevation + gradient + x + y + f(spatial, model=spde), family="poisson", data = inla.stack.data(stack.obs), control.family = list(hyper = hyper.obs), control.predictor = list(a=inla.stack.a(stack.obs), compute=true), control.results=list(return.marginals.random=false, return.marginals.predictor=false), verbose=true) Run this instead Load precompute values instead. > load("inlares.rdata") We extract parameters for the latent field using > spde.res1 <- inla.spde2.result(res1, "spatial", spde) > spde.res2 <- inla.spde2.result(res2, "spatial", spde) If the option return.marginals.random=true had been chosen above these structures would also include posteriors for the field-parameters, rather than only summaries. Analysing INLA output Estimated parameters As a first step in analysing the output we compare the estimated regression parameters for the two models, > res1$summary.fixed > res2$summary.fixed and for the hyper-parameters of the latent field. > res1$summary.hyperpar > res2$summary.hyperpar Or the equivalent range and variances > spde.res1$summary.log.range.nominal > spde.res2$summary.log.range.nominal > spde.res1$summary.log.variance.nominal > spde.res2$summary.log.variance.nominal We can also plot the posterior distributions of the hyper-parameters, along with their priors (in red) > par(mfcol=c(2,1)) > theta.mu <- spde$param.inla$theta.mu > theta.q <- spde$param.inla$theta.q > for(i in 1:2){ ##posterior plot(res1$marginals.hyperpar[[i]], type="l", main=names(res1$marginals.hyperpar)[i]) ##prior x <- res1$marginals.hyperpar[[i]][,1] lines(x, dnorm(x, theta.mu[i], sqrt(1/theta.q[i,i])), col="red") ##and for the second model lines(res2$marginals.hyperpar[[i]], col="blue") } Marginals for the regression parameters can be found in
5 SARMA 2012 Sharp statistical tools 5 > str(res2$marginals.fixed) Latent field Lets first plot the observations and the mean component of the second field > par(mfrow=c(2,1)) > image.plot( as.image(bei$counts, x=bei[,c("x","y")]),asp=1 ) > ##compute the mean, and plot > B <- as.matrix( cbind(1,bei[,rownames(res2$summary.fixed)[-1]]) ) > mu <- B %*% res2$summary.fixed[,"mean"] > image.plot( as.image(mu, x=bei[,c("x","y")]),asp=1 ) To plot the GMRF component of the fields we need to define a projector from the field to a suitable grid that can be used for plotting > mesh.projector <- inla.mesh.projector(mesh, dims=c(200,200)) We can now plot the expectation and standard deviation of the field component w > ##study component > str(res1$summary.random) > ##extract mean and sd > EZ.1 <- res1$summary.random$spatial[,"mean"] > EZ.2 <- res2$summary.random$spatial[,"mean"] > sd.1 <- res1$summary.random$spatial[,"sd"] > sd.2 <- res2$summary.random$spatial[,"sd"] > ##plot the fields > par( mfrow=c(2,2) ) inla.mesh.project(mesh.projector, EZ.1), main="ez.1") inla.mesh.project(mesh.projector, EZ.2), main="ez.2") inla.mesh.project(mesh.projector, sd.1), main="sd.1") inla.mesh.project(mesh.projector, sd.2), main="sd.2") > ##add the mesh for reference to one of the plots > plot(mesh,add=true,col="grey") A better alternative is to limit the plotting to the same are as the observations inla.mesh.project(mesh.projector, EZ.1), xlim=range(bei$x), ylim=range(bei$y)) We have now plotted the mean component and the random component of the latent field, we would also like to plot the full latent field. Predictions for the full field are contained in > str(res1$summary.linear.predictor) with indecies for the observed points given by > I <- inla.stack.index(stack.obs, "obs") > str(i) > ##and extract the relevant components > E1 <- res1$summary.linear.predictor[i$data,] > E2 <- res2$summary.linear.predictor[i$data,] Lets plot the field and uncertainties for the two fields
6 SARMA 2012 Sharp statistical tools 6 > par( mfrow=c(3,3) ) > zlim <- range( c(e1[,"mean"],e2[,"mean"]) ) > image.plot( as.image(e1[,"mean"], x=bei[,c("x","y")]),asp=1, zlim=zlim, main="ez first field") > image.plot( as.image(e2[,"mean"], x=bei[,c("x","y")]),asp=1, zlim=zlim, main="ez second field") > image.plot( as.image(bei$counts, x=bei[,c("x","y")]), asp=1, main="observations") > ##just the regression part > image.plot( as.image(rep(res1$summary.fixed[,"mean"],dim(bei)[1]), x=bei[,c("x","y")]), asp=1, zlim=zlim, main="mean first field") > image.plot( as.image(mu, x=bei[,c("x","y")]), asp=1, zlim=zlim, main="mean second field") > plot(0,0,type="n") > ##the uncertainties > zlimv <- range( c(e1[,"sd"],e2[,"sd"]) ) > image.plot( as.image(e1[,"sd"], x=bei[,c("x","y")]),asp=1, zlim=zlimv, main="vz first field") > image.plot( as.image(e2[,"sd"], x=bei[,c("x","y")]),asp=1, zlim=zlimv, main="vz second field") Comparing the covariance to Matérn We could also compare dependence in the SPDE model to those of the Matérn covariance we re approximating. First we compute the precision matrix given the estimated parameters > Q <- inla.spde2.precision(spde, res2$summary.hyperpar[,"mean"]) and study the sparsity > image(q) Inverting the precision matrix gives the covariance matrix, and correlation matrix (this takes several seconds). > S <- as.matrix(solve(q)) > S <- diag(1/sqrt(diag(s))) %*% S %*% diag(1/sqrt(diag(s))) > ##we also need the distance matrix > D <- as.matrix(dist(mesh$loc[,1:2])) We now need the covariance function and variograms for the SPDE model. We obtain the approximate posterior means for the variance τ and range κ. > tau <- exp( spde.res2$summary.log.tau[,"mean"] ) > kappa <- exp( spde.res2$summary.log.kappa[,"mean"] ) We also compute the theoretical Matérn correlation function > d <- seq(0, max(d), len=1e3) > S.theory <- (d*kappa)*besselk(d*kappa,1) > S.theory[1] <- 1 and compare the two > par(mfrow=c(2,1)) > ##using only a few interior points
7 SARMA 2012 Sharp statistical tools 7 > plot(d[,1:10], as.matrix(s[,1:10]), pch=19, cex=.1, xlab="dist", ylab="") > lines(d, S.theory, col=2) > ##or only the edge points > I <- unique(c(mesh$segm$bnd$idx)) > plot(d[,i], as.matrix(s[,i]), pch=19, cex=.1, xlab="dist", ylab="") > lines(d, S.theory, col=2) The rather larger spread around the theoretical function in the second plot is due to edge effects and the mesh size being reasonable large compared to the range. The first plot gives the correlation between 10 central points and the rest of the field, reducing the edge effects. The end!
Spatial modelling using INLA
 Spatial modelling using INLA August 30, 2012 Bayesian Research and Analysis Group Queensland University of Technology Log Gaussian Cox process Hierarchical Poisson process λ(s) = exp(z(s)) for Z(s) a Gaussian
Spatial modelling using INLA August 30, 2012 Bayesian Research and Analysis Group Queensland University of Technology Log Gaussian Cox process Hierarchical Poisson process λ(s) = exp(z(s)) for Z(s) a Gaussian
Journal of Statistical Software
 JSS Journal of Statistical Software January 2015, Volume 63, Issue 19. http://www.jstatsoft.org/ Bayesian Spatial Modelling with R-INLA Finn Lindgren University of Bath Håvard Rue Norwegian University
JSS Journal of Statistical Software January 2015, Volume 63, Issue 19. http://www.jstatsoft.org/ Bayesian Spatial Modelling with R-INLA Finn Lindgren University of Bath Håvard Rue Norwegian University
Spacetime models in R-INLA. Elias T. Krainski
 Spacetime models in R-INLA Elias T. Krainski 2 Outline Separable space-time models Infant mortality in Paraná PM-10 concentration in Piemonte, Italy 3 Multivariate dynamic regression model y t : n observations
Spacetime models in R-INLA Elias T. Krainski 2 Outline Separable space-time models Infant mortality in Paraná PM-10 concentration in Piemonte, Italy 3 Multivariate dynamic regression model y t : n observations
Statistics in Environmental Research (BUC Workshop Series) IV Problem sheet 1
 Statistics in Environmental Research (BUC Workshop Series) IV Problem sheet 1 Website: http://www.stat.ubc.ca/~gavin/stepibooknewstyle/course_bath.html Spatial modelling with INLA - Part 1 The aims of
Statistics in Environmental Research (BUC Workshop Series) IV Problem sheet 1 Website: http://www.stat.ubc.ca/~gavin/stepibooknewstyle/course_bath.html Spatial modelling with INLA - Part 1 The aims of
Chapter 4 - Fundamentals of spatial processes Lecture notes
 Chapter 4 - Fundamentals of spatial processes Lecture notes Geir Storvik January 21, 2013 STK4150 - Intro 2 Spatial processes Typically correlation between nearby sites Mostly positive correlation Negative
Chapter 4 - Fundamentals of spatial processes Lecture notes Geir Storvik January 21, 2013 STK4150 - Intro 2 Spatial processes Typically correlation between nearby sites Mostly positive correlation Negative
Chapter 4 - Spatial processes R packages and software Lecture notes
 TK4150 - Intro 1 Chapter 4 - Spatial processes R packages and software Lecture notes Odd Kolbjørnsen and Geir Storvik February 13, 2017 STK4150 - Intro 2 Last time General set up for Kriging type problems
TK4150 - Intro 1 Chapter 4 - Spatial processes R packages and software Lecture notes Odd Kolbjørnsen and Geir Storvik February 13, 2017 STK4150 - Intro 2 Last time General set up for Kriging type problems
R-INLA. Sam Clifford Su-Yun Kang Jeff Hsieh. 30 August Bayesian Research and Analysis Group 1 / 14
 1 / 14 R-INLA Sam Clifford Su-Yun Kang Jeff Hsieh Bayesian Research and Analysis Group 30 August 2012 What is R-INLA? R package for Bayesian computation Integrated Nested Laplace Approximation MCMC free
1 / 14 R-INLA Sam Clifford Su-Yun Kang Jeff Hsieh Bayesian Research and Analysis Group 30 August 2012 What is R-INLA? R package for Bayesian computation Integrated Nested Laplace Approximation MCMC free
A short introduction to INLA and R-INLA
 A short introduction to INLA and R-INLA Integrated Nested Laplace Approximation Thomas Opitz, BioSP, INRA Avignon Workshop: Theory and practice of INLA and SPDE November 7, 2018 2/21 Plan for this talk
A short introduction to INLA and R-INLA Integrated Nested Laplace Approximation Thomas Opitz, BioSP, INRA Avignon Workshop: Theory and practice of INLA and SPDE November 7, 2018 2/21 Plan for this talk
Bayesian computation using INLA. 5: Spatial Markovian models The SPDE approach
 Bayesian computation using INLA 5: Spatial Markovian models The SPDE approach Outline Spatial models Gaussian random fields Low-dimensional methods Spatial Matérn fields Example: Continuous vs Discrete
Bayesian computation using INLA 5: Spatial Markovian models The SPDE approach Outline Spatial models Gaussian random fields Low-dimensional methods Spatial Matérn fields Example: Continuous vs Discrete
Gaussian Processes 1. Schedule
 1 Schedule 17 Jan: Gaussian processes (Jo Eidsvik) 24 Jan: Hands-on project on Gaussian processes (Team effort, work in groups) 31 Jan: Latent Gaussian models and INLA (Jo Eidsvik) 7 Feb: Hands-on project
1 Schedule 17 Jan: Gaussian processes (Jo Eidsvik) 24 Jan: Hands-on project on Gaussian processes (Team effort, work in groups) 31 Jan: Latent Gaussian models and INLA (Jo Eidsvik) 7 Feb: Hands-on project
INLA for Spatial Statistics
 INLA for Spatial Statistics 5. Log-Gaussian Co Processes Daniel Simpson Department of Mathematical Sciences University of Bath Outline spatial point processes fitting log Gaussian Co processes eamples
INLA for Spatial Statistics 5. Log-Gaussian Co Processes Daniel Simpson Department of Mathematical Sciences University of Bath Outline spatial point processes fitting log Gaussian Co processes eamples
Machine learning: Hypothesis testing. Anders Hildeman
 Location of trees 0 Observed trees 50 100 150 200 250 300 350 400 450 500 0 100 200 300 400 500 600 700 800 900 1000 Figur: Observed points pattern of the tree specie Beilschmiedia pendula. Location of
Location of trees 0 Observed trees 50 100 150 200 250 300 350 400 450 500 0 100 200 300 400 500 600 700 800 900 1000 Figur: Observed points pattern of the tree specie Beilschmiedia pendula. Location of
Data are collected along transect lines, with dense data along. Spatial modelling using GMRFs. 200m. Today? Spatial modelling using GMRFs
 Centre for Mathematical Sciences Lund University Engineering geology Lund University Results A non-stationary extension The model Estimation Gaussian Markov random fields Basics Approximating Mate rn covariances
Centre for Mathematical Sciences Lund University Engineering geology Lund University Results A non-stationary extension The model Estimation Gaussian Markov random fields Basics Approximating Mate rn covariances
Spatial Statistics with Image Analysis. Outline. A Statistical Approach. Johan Lindström 1. Lund October 6, 2016
 Spatial Statistics Spatial Examples More Spatial Statistics with Image Analysis Johan Lindström 1 1 Mathematical Statistics Centre for Mathematical Sciences Lund University Lund October 6, 2016 Johan Lindström
Spatial Statistics Spatial Examples More Spatial Statistics with Image Analysis Johan Lindström 1 1 Mathematical Statistics Centre for Mathematical Sciences Lund University Lund October 6, 2016 Johan Lindström
Computation fundamentals of discrete GMRF representations of continuous domain spatial models
 Computation fundamentals of discrete GMRF representations of continuous domain spatial models Finn Lindgren September 23 2015 v0.2.2 Abstract The fundamental formulas and algorithms for Bayesian spatial
Computation fundamentals of discrete GMRF representations of continuous domain spatial models Finn Lindgren September 23 2015 v0.2.2 Abstract The fundamental formulas and algorithms for Bayesian spatial
How to solve the stochastic partial differential equation that gives a Matérn random field using the finite element method
 arxiv:1803.03765v1 [stat.co] 10 Mar 2018 How to solve the stochastic partial differential equation that gives a Matérn random field using the finite element method Haakon Bakka bakka@r-inla.org March 13,
arxiv:1803.03765v1 [stat.co] 10 Mar 2018 How to solve the stochastic partial differential equation that gives a Matérn random field using the finite element method Haakon Bakka bakka@r-inla.org March 13,
Lecture 9: Predictive Inference
 Lecture 9: Predictive Inference There are (at least) three levels at which we can make predictions with a regression model: we can give a single best guess about what Y will be when X = x, a point prediction;
Lecture 9: Predictive Inference There are (at least) three levels at which we can make predictions with a regression model: we can give a single best guess about what Y will be when X = x, a point prediction;
Multi-resolution models for large data sets
 Multi-resolution models for large data sets Douglas Nychka, National Center for Atmospheric Research National Science Foundation NORDSTAT, Umeå, June, 2012 Credits Steve Sain, NCAR Tia LeRud, UC Davis
Multi-resolution models for large data sets Douglas Nychka, National Center for Atmospheric Research National Science Foundation NORDSTAT, Umeå, June, 2012 Credits Steve Sain, NCAR Tia LeRud, UC Davis
Lecture 9: Predictive Inference for the Simple Linear Model
 See updates and corrections at http://www.stat.cmu.edu/~cshalizi/mreg/ Lecture 9: Predictive Inference for the Simple Linear Model 36-401, Fall 2015, Section B 29 September 2015 Contents 1 Confidence intervals
See updates and corrections at http://www.stat.cmu.edu/~cshalizi/mreg/ Lecture 9: Predictive Inference for the Simple Linear Model 36-401, Fall 2015, Section B 29 September 2015 Contents 1 Confidence intervals
Spatial Statistics with Image Analysis. Lecture L08. Computer exercise 3. Lecture 8. Johan Lindström. November 25, 2016
 C3 Repetition Creating Q Spectral Non-grid Spatial Statistics with Image Analysis Lecture 8 Johan Lindström November 25, 216 Johan Lindström - johanl@maths.lth.se FMSN2/MASM25L8 1/39 Lecture L8 C3 Repetition
C3 Repetition Creating Q Spectral Non-grid Spatial Statistics with Image Analysis Lecture 8 Johan Lindström November 25, 216 Johan Lindström - johanl@maths.lth.se FMSN2/MASM25L8 1/39 Lecture L8 C3 Repetition
Generalized Linear Models in R
 Generalized Linear Models in R NO ORDER Kenneth K. Lopiano, Garvesh Raskutti, Dan Yang last modified 28 4 2013 1 Outline 1. Background and preliminaries 2. Data manipulation and exercises 3. Data structures
Generalized Linear Models in R NO ORDER Kenneth K. Lopiano, Garvesh Raskutti, Dan Yang last modified 28 4 2013 1 Outline 1. Background and preliminaries 2. Data manipulation and exercises 3. Data structures
A tutorial in spatial and spatio-temporal models with R-INLA
 A tutorial in spatial and spatio-temporal models with R-INLA Marta Blangiardo 1, Michela Cameletti 2, Gianluca Baio 3,4, Håvard Rue 5 1 MRC-HPA Centre for Environment and Health, Department of Epidemiology
A tutorial in spatial and spatio-temporal models with R-INLA Marta Blangiardo 1, Michela Cameletti 2, Gianluca Baio 3,4, Håvard Rue 5 1 MRC-HPA Centre for Environment and Health, Department of Epidemiology
Bayesian Inference for Regression Parameters
 Bayesian Inference for Regression Parameters 1 Bayesian inference for simple linear regression parameters follows the usual pattern for all Bayesian analyses: 1. Form a prior distribution over all unknown
Bayesian Inference for Regression Parameters 1 Bayesian inference for simple linear regression parameters follows the usual pattern for all Bayesian analyses: 1. Form a prior distribution over all unknown
Package horseshoe. November 8, 2016
 Title Implementation of the Horseshoe Prior Version 0.1.0 Package horseshoe November 8, 2016 Description Contains functions for applying the horseshoe prior to highdimensional linear regression, yielding
Title Implementation of the Horseshoe Prior Version 0.1.0 Package horseshoe November 8, 2016 Description Contains functions for applying the horseshoe prior to highdimensional linear regression, yielding
2015 SISG Bayesian Statistics for Genetics R Notes: Generalized Linear Modeling
 2015 SISG Bayesian Statistics for Genetics R Notes: Generalized Linear Modeling Jon Wakefield Departments of Statistics and Biostatistics, University of Washington 2015-07-24 Case control example We analyze
2015 SISG Bayesian Statistics for Genetics R Notes: Generalized Linear Modeling Jon Wakefield Departments of Statistics and Biostatistics, University of Washington 2015-07-24 Case control example We analyze
On Gaussian Process Models for High-Dimensional Geostatistical Datasets
 On Gaussian Process Models for High-Dimensional Geostatistical Datasets Sudipto Banerjee Joint work with Abhirup Datta, Andrew O. Finley and Alan E. Gelfand University of California, Los Angeles, USA May
On Gaussian Process Models for High-Dimensional Geostatistical Datasets Sudipto Banerjee Joint work with Abhirup Datta, Andrew O. Finley and Alan E. Gelfand University of California, Los Angeles, USA May
Explore the data. Anja Bråthen Kristoffersen
 Explore the data Anja Bråthen Kristoffersen density 0.2 0.4 0.6 0.8 Probability distributions Can be either discrete or continuous (uniform, bernoulli, normal, etc) Defined by a density function, p(x)
Explore the data Anja Bråthen Kristoffersen density 0.2 0.4 0.6 0.8 Probability distributions Can be either discrete or continuous (uniform, bernoulli, normal, etc) Defined by a density function, p(x)
-A wild house sparrow population case study
 Bayesian animal model using Integrated Nested Laplace Approximations -A wild house sparrow population case study Anna M. Holand Ingelin Steinsland Animal model workshop with application to ecology at Oulu
Bayesian animal model using Integrated Nested Laplace Approximations -A wild house sparrow population case study Anna M. Holand Ingelin Steinsland Animal model workshop with application to ecology at Oulu
Lecture: Gaussian Process Regression. STAT 6474 Instructor: Hongxiao Zhu
 Lecture: Gaussian Process Regression STAT 6474 Instructor: Hongxiao Zhu Motivation Reference: Marc Deisenroth s tutorial on Robot Learning. 2 Fast Learning for Autonomous Robots with Gaussian Processes
Lecture: Gaussian Process Regression STAT 6474 Instructor: Hongxiao Zhu Motivation Reference: Marc Deisenroth s tutorial on Robot Learning. 2 Fast Learning for Autonomous Robots with Gaussian Processes
Statistical Learning
 Statistical Learning Supervised learning Assume: Estimate: quantity of interest function predictors to get: error Such that: For prediction and/or inference Model fit vs. Model stability (Bias variance
Statistical Learning Supervised learning Assume: Estimate: quantity of interest function predictors to get: error Such that: For prediction and/or inference Model fit vs. Model stability (Bias variance
1 Bayesian Linear Regression (BLR)
 Statistical Techniques in Robotics (STR, S15) Lecture#10 (Wednesday, February 11) Lecturer: Byron Boots Gaussian Properties, Bayesian Linear Regression 1 Bayesian Linear Regression (BLR) In linear regression,
Statistical Techniques in Robotics (STR, S15) Lecture#10 (Wednesday, February 11) Lecturer: Byron Boots Gaussian Properties, Bayesian Linear Regression 1 Bayesian Linear Regression (BLR) In linear regression,
Variance. Standard deviation VAR = = value. Unbiased SD = SD = 10/23/2011. Functional Connectivity Correlation and Regression.
 10/3/011 Functional Connectivity Correlation and Regression Variance VAR = Standard deviation Standard deviation SD = Unbiased SD = 1 10/3/011 Standard error Confidence interval SE = CI = = t value for
10/3/011 Functional Connectivity Correlation and Regression Variance VAR = Standard deviation Standard deviation SD = Unbiased SD = 1 10/3/011 Standard error Confidence interval SE = CI = = t value for
Lecture 19. Spatial GLM + Point Reference Spatial Data. Colin Rundel 04/03/2017
 Lecture 19 Spatial GLM + Point Reference Spatial Data Colin Rundel 04/03/2017 1 Spatial GLM Models 2 Scottish Lip Cancer Data Observed Expected 60 N 59 N 58 N 57 N 56 N value 80 60 40 20 0 55 N 8 W 6 W
Lecture 19 Spatial GLM + Point Reference Spatial Data Colin Rundel 04/03/2017 1 Spatial GLM Models 2 Scottish Lip Cancer Data Observed Expected 60 N 59 N 58 N 57 N 56 N value 80 60 40 20 0 55 N 8 W 6 W
Fast algorithms that utilise the sparsity of Q exist (c-package ( Q 1. GMRFlib) Seattle February 3, 2009
 GMRF:s Results References Some Theory for and Applications of Gaussian Markov Random Fields Johan Lindström 1 David Bolin 1 Finn Lindgren 1 Håvard Rue 2 1 Centre for Mathematical Sciences Lund University
GMRF:s Results References Some Theory for and Applications of Gaussian Markov Random Fields Johan Lindström 1 David Bolin 1 Finn Lindgren 1 Håvard Rue 2 1 Centre for Mathematical Sciences Lund University
Bayesian Gaussian Process Regression
 Bayesian Gaussian Process Regression STAT8810, Fall 2017 M.T. Pratola October 7, 2017 Today Bayesian Gaussian Process Regression Bayesian GP Regression Recall we had observations from our expensive simulator,
Bayesian Gaussian Process Regression STAT8810, Fall 2017 M.T. Pratola October 7, 2017 Today Bayesian Gaussian Process Regression Bayesian GP Regression Recall we had observations from our expensive simulator,
Explore the data. Anja Bråthen Kristoffersen Biomedical Research Group
 Explore the data Anja Bråthen Kristoffersen Biomedical Research Group density 0.2 0.4 0.6 0.8 Probability distributions Can be either discrete or continuous (uniform, bernoulli, normal, etc) Defined by
Explore the data Anja Bråthen Kristoffersen Biomedical Research Group density 0.2 0.4 0.6 0.8 Probability distributions Can be either discrete or continuous (uniform, bernoulli, normal, etc) Defined by
0.1 gamma.mixed: Mixed effects gamma regression
 0. gamma.mixed: Mixed effects gamma regression Use generalized multi-level linear regression if you have covariates that are grouped according to one or more classification factors. Gamma regression models
0. gamma.mixed: Mixed effects gamma regression Use generalized multi-level linear regression if you have covariates that are grouped according to one or more classification factors. Gamma regression models
A general mixed model approach for spatio-temporal regression data
 A general mixed model approach for spatio-temporal regression data Thomas Kneib, Ludwig Fahrmeir & Stefan Lang Department of Statistics, Ludwig-Maximilians-University Munich 1. Spatio-temporal regression
A general mixed model approach for spatio-temporal regression data Thomas Kneib, Ludwig Fahrmeir & Stefan Lang Department of Statistics, Ludwig-Maximilians-University Munich 1. Spatio-temporal regression
Lecture 2: Poisson point processes: properties and statistical inference
 Lecture 2: Poisson point processes: properties and statistical inference Jean-François Coeurjolly http://www-ljk.imag.fr/membres/jean-francois.coeurjolly/ 1 / 20 Definition, properties and simulation Statistical
Lecture 2: Poisson point processes: properties and statistical inference Jean-François Coeurjolly http://www-ljk.imag.fr/membres/jean-francois.coeurjolly/ 1 / 20 Definition, properties and simulation Statistical
Markov random fields. The Markov property
 Markov random fields The Markov property Discrete time: (X k X k!1,x k!2,... = (X k X k!1 A time symmetric version: (X k! X!k = (X k X k!1,x k+1 A more general version: Let A be a set of indices >k, B
Markov random fields The Markov property Discrete time: (X k X k!1,x k!2,... = (X k X k!1 A time symmetric version: (X k! X!k = (X k X k!1,x k+1 A more general version: Let A be a set of indices >k, B
Spatial smoothing over complex domain
 Spatial smoothing over complex domain Alessandro Ottavi NTNU Department of Mathematical Science August 16-20 2010 Outline Spatial smoothing Complex domain The SPDE approch Manifold Some tests Spatial smoothing
Spatial smoothing over complex domain Alessandro Ottavi NTNU Department of Mathematical Science August 16-20 2010 Outline Spatial smoothing Complex domain The SPDE approch Manifold Some tests Spatial smoothing
Big Data in Environmental Research
 Big Data in Environmental Research Gavin Shaddick University of Bath 7 th July 2016 1/ 121 OUTLINE 10:00-11:00 Introduction to Spatio-Temporal Modelling 14:30-15:30 Bayesian Inference 17:00-18:00 Examples
Big Data in Environmental Research Gavin Shaddick University of Bath 7 th July 2016 1/ 121 OUTLINE 10:00-11:00 Introduction to Spatio-Temporal Modelling 14:30-15:30 Bayesian Inference 17:00-18:00 Examples
Bayesian Dynamic Modeling for Space-time Data in R
 Bayesian Dynamic Modeling for Space-time Data in R Andrew O. Finley and Sudipto Banerjee September 5, 2014 We make use of several libraries in the following example session, including: ˆ library(fields)
Bayesian Dynamic Modeling for Space-time Data in R Andrew O. Finley and Sudipto Banerjee September 5, 2014 We make use of several libraries in the following example session, including: ˆ library(fields)
Introduction to Bayes and non-bayes spatial statistics
 Introduction to Bayes and non-bayes spatial statistics Gabriel Huerta Department of Mathematics and Statistics University of New Mexico http://www.stat.unm.edu/ ghuerta/georcode.txt General Concepts Spatial
Introduction to Bayes and non-bayes spatial statistics Gabriel Huerta Department of Mathematics and Statistics University of New Mexico http://www.stat.unm.edu/ ghuerta/georcode.txt General Concepts Spatial
WinBUGS : part 2. Bruno Boulanger Jonathan Jaeger Astrid Jullion Philippe Lambert. Gabriele, living with rheumatoid arthritis
 WinBUGS : part 2 Bruno Boulanger Jonathan Jaeger Astrid Jullion Philippe Lambert Gabriele, living with rheumatoid arthritis Agenda 2! Hierarchical model: linear regression example! R2WinBUGS Linear Regression
WinBUGS : part 2 Bruno Boulanger Jonathan Jaeger Astrid Jullion Philippe Lambert Gabriele, living with rheumatoid arthritis Agenda 2! Hierarchical model: linear regression example! R2WinBUGS Linear Regression
Pattern Recognition and Machine Learning
 Christopher M. Bishop Pattern Recognition and Machine Learning ÖSpri inger Contents Preface Mathematical notation Contents vii xi xiii 1 Introduction 1 1.1 Example: Polynomial Curve Fitting 4 1.2 Probability
Christopher M. Bishop Pattern Recognition and Machine Learning ÖSpri inger Contents Preface Mathematical notation Contents vii xi xiii 1 Introduction 1 1.1 Example: Polynomial Curve Fitting 4 1.2 Probability
Fast approximations for the Expected Value of Partial Perfect Information using R-INLA
 Fast approximations for the Expected Value of Partial Perfect Information using R-INLA Anna Heath 1 1 Department of Statistical Science, University College London 22 May 2015 Outline 1 Health Economic
Fast approximations for the Expected Value of Partial Perfect Information using R-INLA Anna Heath 1 1 Department of Statistical Science, University College London 22 May 2015 Outline 1 Health Economic
Centre for Biodiversity Dynamics, Department of Biology, NTNU, NO-7491 Trondheim, Norway,
 Animal models and Integrated Nested Laplace Approximations Anna Marie Holand,1, Ingelin Steinsland, Sara Martino,, Henrik Jensen Centre for Biodiversity Dynamics, Department of Biology, NTNU, NO-7491 Trondheim,
Animal models and Integrated Nested Laplace Approximations Anna Marie Holand,1, Ingelin Steinsland, Sara Martino,, Henrik Jensen Centre for Biodiversity Dynamics, Department of Biology, NTNU, NO-7491 Trondheim,
Hierarchical Modeling for Univariate Spatial Data
 Hierarchical Modeling for Univariate Spatial Data Geography 890, Hierarchical Bayesian Models for Environmental Spatial Data Analysis February 15, 2011 1 Spatial Domain 2 Geography 890 Spatial Domain This
Hierarchical Modeling for Univariate Spatial Data Geography 890, Hierarchical Bayesian Models for Environmental Spatial Data Analysis February 15, 2011 1 Spatial Domain 2 Geography 890 Spatial Domain This
ES-2 Lecture: More Least-squares Fitting. Spring 2017
 ES-2 Lecture: More Least-squares Fitting Spring 2017 Outline Quick review of least-squares line fitting (also called `linear regression ) How can we find the best-fit line? (Brute-force method is not efficient)
ES-2 Lecture: More Least-squares Fitting Spring 2017 Outline Quick review of least-squares line fitting (also called `linear regression ) How can we find the best-fit line? (Brute-force method is not efficient)
Generalized Linear Models for Non-Normal Data
 Generalized Linear Models for Non-Normal Data Today s Class: 3 parts of a generalized model Models for binary outcomes Complications for generalized multivariate or multilevel models SPLH 861: Lecture
Generalized Linear Models for Non-Normal Data Today s Class: 3 parts of a generalized model Models for binary outcomes Complications for generalized multivariate or multilevel models SPLH 861: Lecture
Statistics & Data Sciences: First Year Prelim Exam May 2018
 Statistics & Data Sciences: First Year Prelim Exam May 2018 Instructions: 1. Do not turn this page until instructed to do so. 2. Start each new question on a new sheet of paper. 3. This is a closed book
Statistics & Data Sciences: First Year Prelim Exam May 2018 Instructions: 1. Do not turn this page until instructed to do so. 2. Start each new question on a new sheet of paper. 3. This is a closed book
Summary STK 4150/9150
 STK4150 - Intro 1 Summary STK 4150/9150 Odd Kolbjørnsen May 22 2017 Scope You are expected to know and be able to use basic concepts introduced in the book. You knowledge is expected to be larger than
STK4150 - Intro 1 Summary STK 4150/9150 Odd Kolbjørnsen May 22 2017 Scope You are expected to know and be able to use basic concepts introduced in the book. You knowledge is expected to be larger than
georglm : a package for generalised linear spatial models introductory session
 georglm : a package for generalised linear spatial models introductory session Ole F. Christensen & Paulo J. Ribeiro Jr. Last update: October 18, 2017 The objective of this page is to introduce the reader
georglm : a package for generalised linear spatial models introductory session Ole F. Christensen & Paulo J. Ribeiro Jr. Last update: October 18, 2017 The objective of this page is to introduce the reader
Technical Vignette 5: Understanding intrinsic Gaussian Markov random field spatial models, including intrinsic conditional autoregressive models
 Technical Vignette 5: Understanding intrinsic Gaussian Markov random field spatial models, including intrinsic conditional autoregressive models Christopher Paciorek, Department of Statistics, University
Technical Vignette 5: Understanding intrinsic Gaussian Markov random field spatial models, including intrinsic conditional autoregressive models Christopher Paciorek, Department of Statistics, University
Political Science 6000: Beginnings and Mini Math Boot Camp
 Political Science 6000: Beginnings and Mini Math Boot Camp January 20, 2010 First things first Syllabus This is the most important course you will take. 1. You need to understand these concepts in order
Political Science 6000: Beginnings and Mini Math Boot Camp January 20, 2010 First things first Syllabus This is the most important course you will take. 1. You need to understand these concepts in order
Log Gaussian Cox Processes. Chi Group Meeting February 23, 2016
 Log Gaussian Cox Processes Chi Group Meeting February 23, 2016 Outline Typical motivating application Introduction to LGCP model Brief overview of inference Applications in my work just getting started
Log Gaussian Cox Processes Chi Group Meeting February 23, 2016 Outline Typical motivating application Introduction to LGCP model Brief overview of inference Applications in my work just getting started
Modeling Data with Linear Combinations of Basis Functions. Read Chapter 3 in the text by Bishop
 Modeling Data with Linear Combinations of Basis Functions Read Chapter 3 in the text by Bishop A Type of Supervised Learning Problem We want to model data (x 1, t 1 ),..., (x N, t N ), where x i is a vector
Modeling Data with Linear Combinations of Basis Functions Read Chapter 3 in the text by Bishop A Type of Supervised Learning Problem We want to model data (x 1, t 1 ),..., (x N, t N ), where x i is a vector
CPSC 340 Assignment 4 (due November 17 ATE)
 CPSC 340 Assignment 4 due November 7 ATE) Multi-Class Logistic The function example multiclass loads a multi-class classification datasetwith y i {,, 3, 4, 5} and fits a one-vs-all classification model
CPSC 340 Assignment 4 due November 7 ATE) Multi-Class Logistic The function example multiclass loads a multi-class classification datasetwith y i {,, 3, 4, 5} and fits a one-vs-all classification model
Linear Regression. Data Model. β, σ 2. Process Model. ,V β. ,s 2. s 1. Parameter Model
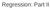 Regression: Part II Linear Regression y~n X, 2 X Y Data Model β, σ 2 Process Model Β 0,V β s 1,s 2 Parameter Model Assumptions of Linear Model Homoskedasticity No error in X variables Error in Y variables
Regression: Part II Linear Regression y~n X, 2 X Y Data Model β, σ 2 Process Model Β 0,V β s 1,s 2 Parameter Model Assumptions of Linear Model Homoskedasticity No error in X variables Error in Y variables
Multiple Linear Regression (solutions to exercises)
 Chapter 6 1 Chapter 6 Multiple Linear Regression (solutions to exercises) Chapter 6 CONTENTS 2 Contents 6 Multiple Linear Regression (solutions to exercises) 1 6.1 Nitrate concentration..........................
Chapter 6 1 Chapter 6 Multiple Linear Regression (solutions to exercises) Chapter 6 CONTENTS 2 Contents 6 Multiple Linear Regression (solutions to exercises) 1 6.1 Nitrate concentration..........................
STA414/2104 Statistical Methods for Machine Learning II
 STA414/2104 Statistical Methods for Machine Learning II Murat A. Erdogdu & David Duvenaud Department of Computer Science Department of Statistical Sciences Lecture 3 Slide credits: Russ Salakhutdinov Announcements
STA414/2104 Statistical Methods for Machine Learning II Murat A. Erdogdu & David Duvenaud Department of Computer Science Department of Statistical Sciences Lecture 3 Slide credits: Russ Salakhutdinov Announcements
Experiment 1: Linear Regression
 Experiment 1: Linear Regression August 27, 2018 1 Description This first exercise will give you practice with linear regression. These exercises have been extensively tested with Matlab, but they should
Experiment 1: Linear Regression August 27, 2018 1 Description This first exercise will give you practice with linear regression. These exercises have been extensively tested with Matlab, but they should
Default Priors and Effcient Posterior Computation in Bayesian
 Default Priors and Effcient Posterior Computation in Bayesian Factor Analysis January 16, 2010 Presented by Eric Wang, Duke University Background and Motivation A Brief Review of Parameter Expansion Literature
Default Priors and Effcient Posterior Computation in Bayesian Factor Analysis January 16, 2010 Presented by Eric Wang, Duke University Background and Motivation A Brief Review of Parameter Expansion Literature
The evdbayes Package
 The evdbayes Package April 19, 2006 Version 1.0-5 Date 2006-18-04 Title Bayesian Analysis in Extreme Theory Author Alec Stephenson and Mathieu Ribatet. Maintainer Mathieu Ribatet
The evdbayes Package April 19, 2006 Version 1.0-5 Date 2006-18-04 Title Bayesian Analysis in Extreme Theory Author Alec Stephenson and Mathieu Ribatet. Maintainer Mathieu Ribatet
Data Assimilation Research Testbed Tutorial
 Data Assimilation Research Testbed Tutorial Section 3: Hierarchical Group Filters and Localization Version 2.: September, 26 Anderson: Ensemble Tutorial 9//6 Ways to deal with regression sampling error:
Data Assimilation Research Testbed Tutorial Section 3: Hierarchical Group Filters and Localization Version 2.: September, 26 Anderson: Ensemble Tutorial 9//6 Ways to deal with regression sampling error:
x. Figure 1: Examples of univariate Gaussian pdfs N (x; µ, σ 2 ).
 .8.6 µ =, σ = 1 µ = 1, σ = 1 / µ =, σ =.. 3 1 1 3 x Figure 1: Examples of univariate Gaussian pdfs N (x; µ, σ ). The Gaussian distribution Probably the most-important distribution in all of statistics
.8.6 µ =, σ = 1 µ = 1, σ = 1 / µ =, σ =.. 3 1 1 3 x Figure 1: Examples of univariate Gaussian pdfs N (x; µ, σ ). The Gaussian distribution Probably the most-important distribution in all of statistics
The Expectation-Maximization Algorithm
 1/29 EM & Latent Variable Models Gaussian Mixture Models EM Theory The Expectation-Maximization Algorithm Mihaela van der Schaar Department of Engineering Science University of Oxford MLE for Latent Variable
1/29 EM & Latent Variable Models Gaussian Mixture Models EM Theory The Expectation-Maximization Algorithm Mihaela van der Schaar Department of Engineering Science University of Oxford MLE for Latent Variable
Introduction to Statistics and R
 Introduction to Statistics and R Mayo-Illinois Computational Genomics Workshop (2018) Ruoqing Zhu, Ph.D. Department of Statistics, UIUC rqzhu@illinois.edu June 18, 2018 Abstract This document is a supplimentary
Introduction to Statistics and R Mayo-Illinois Computational Genomics Workshop (2018) Ruoqing Zhu, Ph.D. Department of Statistics, UIUC rqzhu@illinois.edu June 18, 2018 Abstract This document is a supplimentary
Linear Regression (9/11/13)
 STA561: Probabilistic machine learning Linear Regression (9/11/13) Lecturer: Barbara Engelhardt Scribes: Zachary Abzug, Mike Gloudemans, Zhuosheng Gu, Zhao Song 1 Why use linear regression? Figure 1: Scatter
STA561: Probabilistic machine learning Linear Regression (9/11/13) Lecturer: Barbara Engelhardt Scribes: Zachary Abzug, Mike Gloudemans, Zhuosheng Gu, Zhao Song 1 Why use linear regression? Figure 1: Scatter
MCMC 2: Lecture 3 SIR models - more topics. Phil O Neill Theo Kypraios School of Mathematical Sciences University of Nottingham
 MCMC 2: Lecture 3 SIR models - more topics Phil O Neill Theo Kypraios School of Mathematical Sciences University of Nottingham Contents 1. What can be estimated? 2. Reparameterisation 3. Marginalisation
MCMC 2: Lecture 3 SIR models - more topics Phil O Neill Theo Kypraios School of Mathematical Sciences University of Nottingham Contents 1. What can be estimated? 2. Reparameterisation 3. Marginalisation
The Bernstein Mechansim
 The Bernstein Mechansim diffpriv team July 18, 2017 Abstract This vignette presents a short tutorial on the application of the generic Bernstein mechanism DPMechBernstein for differentially-private release
The Bernstein Mechansim diffpriv team July 18, 2017 Abstract This vignette presents a short tutorial on the application of the generic Bernstein mechanism DPMechBernstein for differentially-private release
Business Statistics. Lecture 10: Course Review
 Business Statistics Lecture 10: Course Review 1 Descriptive Statistics for Continuous Data Numerical Summaries Location: mean, median Spread or variability: variance, standard deviation, range, percentiles,
Business Statistics Lecture 10: Course Review 1 Descriptive Statistics for Continuous Data Numerical Summaries Location: mean, median Spread or variability: variance, standard deviation, range, percentiles,
Nonparameteric Regression:
 Nonparameteric Regression: Nadaraya-Watson Kernel Regression & Gaussian Process Regression Seungjin Choi Department of Computer Science and Engineering Pohang University of Science and Technology 77 Cheongam-ro,
Nonparameteric Regression: Nadaraya-Watson Kernel Regression & Gaussian Process Regression Seungjin Choi Department of Computer Science and Engineering Pohang University of Science and Technology 77 Cheongam-ro,
Factor Analysis (10/2/13)
 STA561: Probabilistic machine learning Factor Analysis (10/2/13) Lecturer: Barbara Engelhardt Scribes: Li Zhu, Fan Li, Ni Guan Factor Analysis Factor analysis is related to the mixture models we have studied.
STA561: Probabilistic machine learning Factor Analysis (10/2/13) Lecturer: Barbara Engelhardt Scribes: Li Zhu, Fan Li, Ni Guan Factor Analysis Factor analysis is related to the mixture models we have studied.
Non-Parametric Bayes
 Non-Parametric Bayes Mark Schmidt UBC Machine Learning Reading Group January 2016 Current Hot Topics in Machine Learning Bayesian learning includes: Gaussian processes. Approximate inference. Bayesian
Non-Parametric Bayes Mark Schmidt UBC Machine Learning Reading Group January 2016 Current Hot Topics in Machine Learning Bayesian learning includes: Gaussian processes. Approximate inference. Bayesian
Beyond MCMC in fitting complex Bayesian models: The INLA method
 Beyond MCMC in fitting complex Bayesian models: The INLA method Valeska Andreozzi Centre of Statistics and Applications of Lisbon University (valeska.andreozzi at fc.ul.pt) European Congress of Epidemiology
Beyond MCMC in fitting complex Bayesian models: The INLA method Valeska Andreozzi Centre of Statistics and Applications of Lisbon University (valeska.andreozzi at fc.ul.pt) European Congress of Epidemiology
Nonstationary spatial process modeling Part II Paul D. Sampson --- Catherine Calder Univ of Washington --- Ohio State University
 Nonstationary spatial process modeling Part II Paul D. Sampson --- Catherine Calder Univ of Washington --- Ohio State University this presentation derived from that presented at the Pan-American Advanced
Nonstationary spatial process modeling Part II Paul D. Sampson --- Catherine Calder Univ of Washington --- Ohio State University this presentation derived from that presented at the Pan-American Advanced
Cheng Soon Ong & Christian Walder. Canberra February June 2018
 Cheng Soon Ong & Christian Walder Research Group and College of Engineering and Computer Science Canberra February June 2018 Outlines Overview Introduction Linear Algebra Probability Linear Regression
Cheng Soon Ong & Christian Walder Research Group and College of Engineering and Computer Science Canberra February June 2018 Outlines Overview Introduction Linear Algebra Probability Linear Regression
Part 6: Multivariate Normal and Linear Models
 Part 6: Multivariate Normal and Linear Models 1 Multiple measurements Up until now all of our statistical models have been univariate models models for a single measurement on each member of a sample of
Part 6: Multivariate Normal and Linear Models 1 Multiple measurements Up until now all of our statistical models have been univariate models models for a single measurement on each member of a sample of
cor(dataset$measurement1, dataset$measurement2, method= pearson ) cor.test(datavector1, datavector2, method= pearson )
 Tutorial 7: Correlation and Regression Correlation Used to test whether two variables are linearly associated. A correlation coefficient (r) indicates the strength and direction of the association. A correlation
Tutorial 7: Correlation and Regression Correlation Used to test whether two variables are linearly associated. A correlation coefficient (r) indicates the strength and direction of the association. A correlation
Dependence. Practitioner Course: Portfolio Optimization. John Dodson. September 10, Dependence. John Dodson. Outline.
 Practitioner Course: Portfolio Optimization September 10, 2008 Before we define dependence, it is useful to define Random variables X and Y are independent iff For all x, y. In particular, F (X,Y ) (x,
Practitioner Course: Portfolio Optimization September 10, 2008 Before we define dependence, it is useful to define Random variables X and Y are independent iff For all x, y. In particular, F (X,Y ) (x,
1 gamma.mixed: Mixed effects gamma regression
 gamma.mixed: Mixed effects gamma regression Use generalized multi-level linear regression if you have covariates that are grouped according to one or more classification factors. Gamma regression models
gamma.mixed: Mixed effects gamma regression Use generalized multi-level linear regression if you have covariates that are grouped according to one or more classification factors. Gamma regression models
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 11 Project
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 11 Project
Community Health Needs Assessment through Spatial Regression Modeling
 Community Health Needs Assessment through Spatial Regression Modeling Glen D. Johnson, PhD CUNY School of Public Health glen.johnson@lehman.cuny.edu Objectives: Assess community needs with respect to particular
Community Health Needs Assessment through Spatial Regression Modeling Glen D. Johnson, PhD CUNY School of Public Health glen.johnson@lehman.cuny.edu Objectives: Assess community needs with respect to particular
The Bayesian approach to inverse problems
 The Bayesian approach to inverse problems Youssef Marzouk Department of Aeronautics and Astronautics Center for Computational Engineering Massachusetts Institute of Technology ymarz@mit.edu, http://uqgroup.mit.edu
The Bayesian approach to inverse problems Youssef Marzouk Department of Aeronautics and Astronautics Center for Computational Engineering Massachusetts Institute of Technology ymarz@mit.edu, http://uqgroup.mit.edu
CS Homework 3. October 15, 2009
 CS 294 - Homework 3 October 15, 2009 If you have questions, contact Alexandre Bouchard (bouchard@cs.berkeley.edu) for part 1 and Alex Simma (asimma@eecs.berkeley.edu) for part 2. Also check the class website
CS 294 - Homework 3 October 15, 2009 If you have questions, contact Alexandre Bouchard (bouchard@cs.berkeley.edu) for part 1 and Alex Simma (asimma@eecs.berkeley.edu) for part 2. Also check the class website
4 Elementary matrices, continued
 4 Elementary matrices, continued We have identified 3 types of row operations and their corresponding elementary matrices. If you check the previous examples, you ll find that these matrices are constructed
4 Elementary matrices, continued We have identified 3 types of row operations and their corresponding elementary matrices. If you check the previous examples, you ll find that these matrices are constructed
Probabilistic Graphical Models
 Probabilistic Graphical Models Brown University CSCI 2950-P, Spring 2013 Prof. Erik Sudderth Lecture 12: Gaussian Belief Propagation, State Space Models and Kalman Filters Guest Kalman Filter Lecture by
Probabilistic Graphical Models Brown University CSCI 2950-P, Spring 2013 Prof. Erik Sudderth Lecture 12: Gaussian Belief Propagation, State Space Models and Kalman Filters Guest Kalman Filter Lecture by
Bayesian Graphical Models
 Graphical Models and Inference, Lecture 16, Michaelmas Term 2009 December 4, 2009 Parameter θ, data X = x, likelihood L(θ x) p(x θ). Express knowledge about θ through prior distribution π on θ. Inference
Graphical Models and Inference, Lecture 16, Michaelmas Term 2009 December 4, 2009 Parameter θ, data X = x, likelihood L(θ x) p(x θ). Express knowledge about θ through prior distribution π on θ. Inference
REN R 690 Lab Geostatistics Lab
 REN R 690 Lab Geostatistics Lab The objective of this lab is to try out some basic geostatistical tools with R. Geostatistics is used primarily in the resource and environmental areas for estimation, uncertainty
REN R 690 Lab Geostatistics Lab The objective of this lab is to try out some basic geostatistical tools with R. Geostatistics is used primarily in the resource and environmental areas for estimation, uncertainty
Luke B Smith and Brian J Reich North Carolina State University May 21, 2013
 BSquare: An R package for Bayesian simultaneous quantile regression Luke B Smith and Brian J Reich North Carolina State University May 21, 2013 BSquare in an R package to conduct Bayesian quantile regression
BSquare: An R package for Bayesian simultaneous quantile regression Luke B Smith and Brian J Reich North Carolina State University May 21, 2013 BSquare in an R package to conduct Bayesian quantile regression
2017 SISG Bayesian Statistics for Genetics R Notes: Generalized Linear Modeling
 2017 SISG Bayesian Statistics for Genetics R Notes: Generalized Linear Modeling Jon Wakefield Departments of Statistics and Biostatistics, University of Washington 2017-09-05 Overview In this set of notes
2017 SISG Bayesian Statistics for Genetics R Notes: Generalized Linear Modeling Jon Wakefield Departments of Statistics and Biostatistics, University of Washington 2017-09-05 Overview In this set of notes
Computer Vision Group Prof. Daniel Cremers. 2. Regression (cont.)
 Prof. Daniel Cremers 2. Regression (cont.) Regression with MLE (Rep.) Assume that y is affected by Gaussian noise : t = f(x, w)+ where Thus, we have p(t x, w, )=N (t; f(x, w), 2 ) 2 Maximum A-Posteriori
Prof. Daniel Cremers 2. Regression (cont.) Regression with MLE (Rep.) Assume that y is affected by Gaussian noise : t = f(x, w)+ where Thus, we have p(t x, w, )=N (t; f(x, w), 2 ) 2 Maximum A-Posteriori
Stochastic vs Deterministic Pre-stack Inversion Methods. Brian Russell
 Stochastic vs Deterministic Pre-stack Inversion Methods Brian Russell Introduction Seismic reservoir analysis techniques utilize the fact that seismic amplitudes contain information about the geological
Stochastic vs Deterministic Pre-stack Inversion Methods Brian Russell Introduction Seismic reservoir analysis techniques utilize the fact that seismic amplitudes contain information about the geological
Chapter 1. Modeling Basics
 Chapter 1. Modeling Basics What is a model? Model equation and probability distribution Types of model effects Writing models in matrix form Summary 1 What is a statistical model? A model is a mathematical
Chapter 1. Modeling Basics What is a model? Model equation and probability distribution Types of model effects Writing models in matrix form Summary 1 What is a statistical model? A model is a mathematical
coenocliner: a coenocline simulation package for R
 coenocliner: a coenocline simulation package for R Gavin L. Simpson Institute of Environmental Change and Society University of Regina Abstract This vignette provides an introduction to, and user-guide
coenocliner: a coenocline simulation package for R Gavin L. Simpson Institute of Environmental Change and Society University of Regina Abstract This vignette provides an introduction to, and user-guide
Metric Predicted Variable on Two Groups
 Metric Predicted Variable on Two Groups Tim Frasier Copyright Tim Frasier This work is licensed under the Creative Commons Attribution 4.0 International license. Click here for more information. Goals
Metric Predicted Variable on Two Groups Tim Frasier Copyright Tim Frasier This work is licensed under the Creative Commons Attribution 4.0 International license. Click here for more information. Goals
Multivariate Gaussian Random Fields with SPDEs
 Multivariate Gaussian Random Fields with SPDEs Xiangping Hu Daniel Simpson, Finn Lindgren and Håvard Rue Department of Mathematics, University of Oslo PASI, 214 Outline The Matérn covariance function and
Multivariate Gaussian Random Fields with SPDEs Xiangping Hu Daniel Simpson, Finn Lindgren and Håvard Rue Department of Mathematics, University of Oslo PASI, 214 Outline The Matérn covariance function and
CSC2515 Assignment #2
 CSC2515 Assignment #2 Due: Nov.4, 2pm at the START of class Worth: 18% Late assignments not accepted. 1 Pseudo-Bayesian Linear Regression (3%) In this question you will dabble in Bayesian statistics and
CSC2515 Assignment #2 Due: Nov.4, 2pm at the START of class Worth: 18% Late assignments not accepted. 1 Pseudo-Bayesian Linear Regression (3%) In this question you will dabble in Bayesian statistics and
