Implementation of EM algorithm in HMM training. Jens Lagergren
|
|
|
- Kimberly Powers
- 5 years ago
- Views:
Transcription
1 Implementation of EM algorithm in HMM training Jens Lagergren April 22, 2009
2 EM Algorithms: An expectation-maximization (EM) algorithm is used in statistics for finding maximum likelihood estimates of parameters in probabilistic models, where the model depends on hidden variables. EM is a 2 step iterative method. In the first step(e-step) it computes an expectation of the log likelihood with respect to the current estimate of the distribution for the hidden variables. In second step (M-step), it computes the parameters which maximize the expected log likelihood. In each cycle, it uses the aproximations from previous step to aproximate the solution progressively[wikipedia.com]. In BioInformatics, EM is commonly used to generate HMM s from selected genetic sequences. Training Hidden Markov Models using EM HMM profiles are usually trained with a set of sequences that are known to belong to a single family. Once the HMM has been trained and optimized to identify these known members, it is then used to clasify unknown genes. Namely, this requires us to construct a HMM, M, that generates a collection of sequences F with the maximum probability: max Pr[x M] (1) HMMsM But this can be computationally hard. Therefore we break down the problem into following sub-problems Finding the right structure, i.e. determining the number of layers. (We will not consider this sub-problem at the moment) Finding optimal parameters for a given structure. To solve the latter problem we now let A π be the expected number of π transitions. A π be the expected number of visits to π. G π,σ be the expected number of times σ is generated from state π. θ be the set of parameters(hidden variables to be optimized) composed of: e π (σ) is the probability to generate the symbol σ in state π. a π is the transition probability from state π to. We also continue with the notation X i := x 1,..., x i and Π i = π 1,..., π i, from previous notes. 1
3 Expectation Maximization We will use the EM algorithm to optimize the parameters to create the best fitting HMM from a given set of sequences F. From the initial HMM M and the initial parameter set θ we can generate a new optimized parameter set: θ, by iterating this procedure the aproximation will eventually converge with the local maximum. We calculate this new set θ as follows E[A π x, θ] a π = e π(σ) = E[A π X, θ] E[G π,σ X, θ] E[A π X, θ] We can compute each nominator and denominator separately: 1. E[A π X, θ] (denominator) 2. E[G π,σ X, θ] (nominator for e π(σ)) 3. E[A π X, θ] (nominator for a π) 1. Calculating the denominator E[A π X, θ]: Notice that: E[A π X, θ] = E[A π X, θ] Therefore E[A π X, θ] is easily computed once E[A π, X, θ] is resolved in point Calculating the nominator:e[g π,σ X, θ] E[G π,σ X, θ] = 3. Calculating E[A π X, θ] i,x i=σ = i,x i =σ = i,x i =σ P r[π i = π X, θ] P r[π i = π, X θ] P r[x θ] P r[π i 1 = π, π i = π, X θ] P r[x θ] E[A π X, θ] = Pr[π i = π, π i+1 = X, θ] i i = Pr[π i = π, π i+1 =, X θ] Pr[X θ] 2
4 so it is enough to be able to compute Pr[π i = π, π i+1 =, X θ] Pr[π i = π, π i+1 =, X θ] = Pr[π i = π, π i+1 =, X X i, π i = π, θ]pr[x i, π i = π θ] = Pr[x i+1,..., x n, π i+1 = π i = π, θ] Pr[X i, π i = π θ] }{{}}{{} The Markov property! f π (i) = Pr[x i+1,..., x n π i+1 =, π i = π, θ] Pr[π i+1 = π i = π, θ] f π (i) }{{} a π = Pr[x i+2,..., x n π i+1 =, θ] Pr[x i+1 π i+1 = ] a πf π (i) }{{}}{{} b π (i+1) e (x i+1 ) = b π (i + 1)e (x i+1 )a πf π (i)... where 1. b π (i + 1) is the backward variable defined as: b π (i) = Pr[x i+1,..., x n π i = π, θ] which can be computed using dynamic programming in a similar way as f π (i) was computed in lecture f π (i) is the forward variable defined in lecture e (x i+1 ) and a π were already calculated in the last cycle. Stopping the iteration When to stop? Notice that for θ given by a π and e π(σ) either θ are the locally optimal parameters, in which case you may stop the algorithm. or Pr[x θ ] > [x θ], which means a solution has not been reached yet. Time analysis of EM alorithm in HMM Training Problem A full time analysis of the complete EM algorithm would need to analyze the number of iterations required for convergence which may vary in different cases, nevertheless one may estimate the time required to calculate some of its components, Ie: Claim 1. max π1,...,π n Pr[x 1,..., x n, π 0,..., π n ] can be computed in time O( Q 2 l). 3
5 Proof: Let v π (i) = max Pr[x 1,..., x i, π 0,..., π i 1, π i = π] and Then v π (0) = { 1 when π = qstart 0 otherwise. v π (i) π 1,...,π i 1 Pr[x 1,..., x i, π 0,..., π i 1, π i = π] That is, max Pr[X i, Π i 2, π i 1 =, π i = π] max Pr[X i, Π i 2, π i 1 =, π i = π X i 1, Π i 2, π 0, π i 1 = ] Pr[X i 1, Π i 2, π 0, π i 1 = ] max Pr[x i, π i = π π i 1 = π i 1 ] }{{} The Markov property! Pr[X i 1, Π i 2, π 0, π i 1 = ] max Pr[x i, π i = π]pr[π i = π π i 1 = π i 1 ] Pr[X i 1, Π i 2, π 0, π i 1 = ] = Pr[x i, π i = π] max }{{} Pr[π i = π π i 1 = π i 1 ] max Pr[X i 1, Π i 2, π 0, π i 1 = ] }{{} π 1,...,π i 1 e π (x i ) a π }{{} v (i 1) = e π (x i ) max a πv (i 1) v π (i) v (i 1)a πe π (x i ) Since there are Q n subproblems V π (i) and each can be computed in time O( Q ) this gives a recursion that can be computed in time O( Q 2 n). 4
CS1820 Notes. hgupta1, kjline, smechery. April 3-April 5. output: plausible Ancestral Recombination Graph (ARG)
 CS1820 Notes hgupta1, kjline, smechery April 3-April 5 April 3 Notes 1 Minichiello-Durbin Algorithm input: set of sequences output: plausible Ancestral Recombination Graph (ARG) note: the optimal ARG is
CS1820 Notes hgupta1, kjline, smechery April 3-April 5 April 3 Notes 1 Minichiello-Durbin Algorithm input: set of sequences output: plausible Ancestral Recombination Graph (ARG) note: the optimal ARG is
Hidden Markov Models. Three classic HMM problems
 An Introduction to Bioinformatics Algorithms www.bioalgorithms.info Hidden Markov Models Slides revised and adapted to Computational Biology IST 2015/2016 Ana Teresa Freitas Three classic HMM problems
An Introduction to Bioinformatics Algorithms www.bioalgorithms.info Hidden Markov Models Slides revised and adapted to Computational Biology IST 2015/2016 Ana Teresa Freitas Three classic HMM problems
Computational Genomics and Molecular Biology, Fall
 Computational Genomics and Molecular Biology, Fall 2011 1 HMM Lecture Notes Dannie Durand and Rose Hoberman October 11th 1 Hidden Markov Models In the last few lectures, we have focussed on three problems
Computational Genomics and Molecular Biology, Fall 2011 1 HMM Lecture Notes Dannie Durand and Rose Hoberman October 11th 1 Hidden Markov Models In the last few lectures, we have focussed on three problems
CISC 889 Bioinformatics (Spring 2004) Hidden Markov Models (II)
 CISC 889 Bioinformatics (Spring 24) Hidden Markov Models (II) a. Likelihood: forward algorithm b. Decoding: Viterbi algorithm c. Model building: Baum-Welch algorithm Viterbi training Hidden Markov models
CISC 889 Bioinformatics (Spring 24) Hidden Markov Models (II) a. Likelihood: forward algorithm b. Decoding: Viterbi algorithm c. Model building: Baum-Welch algorithm Viterbi training Hidden Markov models
Note Set 5: Hidden Markov Models
 Note Set 5: Hidden Markov Models Probabilistic Learning: Theory and Algorithms, CS 274A, Winter 2016 1 Hidden Markov Models (HMMs) 1.1 Introduction Consider observed data vectors x t that are d-dimensional
Note Set 5: Hidden Markov Models Probabilistic Learning: Theory and Algorithms, CS 274A, Winter 2016 1 Hidden Markov Models (HMMs) 1.1 Introduction Consider observed data vectors x t that are d-dimensional
6.864: Lecture 5 (September 22nd, 2005) The EM Algorithm
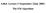 6.864: Lecture 5 (September 22nd, 2005) The EM Algorithm Overview The EM algorithm in general form The EM algorithm for hidden markov models (brute force) The EM algorithm for hidden markov models (dynamic
6.864: Lecture 5 (September 22nd, 2005) The EM Algorithm Overview The EM algorithm in general form The EM algorithm for hidden markov models (brute force) The EM algorithm for hidden markov models (dynamic
HMM: Parameter Estimation
 I529: Machine Learning in Bioinformatics (Spring 2017) HMM: Parameter Estimation Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Content Review HMM: three problems
I529: Machine Learning in Bioinformatics (Spring 2017) HMM: Parameter Estimation Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Content Review HMM: three problems
Parametric Models Part III: Hidden Markov Models
 Parametric Models Part III: Hidden Markov Models Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2014 CS 551, Spring 2014 c 2014, Selim Aksoy (Bilkent
Parametric Models Part III: Hidden Markov Models Selim Aksoy Department of Computer Engineering Bilkent University saksoy@cs.bilkent.edu.tr CS 551, Spring 2014 CS 551, Spring 2014 c 2014, Selim Aksoy (Bilkent
CS6220 Data Mining Techniques Hidden Markov Models, Exponential Families, and the Forward-backward Algorithm
 CS6220 Data Mining Techniques Hidden Markov Models, Exponential Families, and the Forward-backward Algorithm Jan-Willem van de Meent, 19 November 2016 1 Hidden Markov Models A hidden Markov model (HMM)
CS6220 Data Mining Techniques Hidden Markov Models, Exponential Families, and the Forward-backward Algorithm Jan-Willem van de Meent, 19 November 2016 1 Hidden Markov Models A hidden Markov model (HMM)
Hidden Markov Models
 Hidden Markov Models Slides revised and adapted to Bioinformática 55 Engª Biomédica/IST 2005 Ana Teresa Freitas Forward Algorithm For Markov chains we calculate the probability of a sequence, P(x) How
Hidden Markov Models Slides revised and adapted to Bioinformática 55 Engª Biomédica/IST 2005 Ana Teresa Freitas Forward Algorithm For Markov chains we calculate the probability of a sequence, P(x) How
Hidden Markov Models: All the Glorious Gory Details
 Hidden Markov Models: All the Glorious Gory Details Noah A. Smith Department of Computer Science Johns Hopkins University nasmith@cs.jhu.edu 18 October 2004 1 Introduction Hidden Markov models (HMMs, hereafter)
Hidden Markov Models: All the Glorious Gory Details Noah A. Smith Department of Computer Science Johns Hopkins University nasmith@cs.jhu.edu 18 October 2004 1 Introduction Hidden Markov models (HMMs, hereafter)
Stephen Scott.
 1 / 27 sscott@cse.unl.edu 2 / 27 Useful for modeling/making predictions on sequential data E.g., biological sequences, text, series of sounds/spoken words Will return to graphical models that are generative
1 / 27 sscott@cse.unl.edu 2 / 27 Useful for modeling/making predictions on sequential data E.g., biological sequences, text, series of sounds/spoken words Will return to graphical models that are generative
Lecture 4: Hidden Markov Models: An Introduction to Dynamic Decision Making. November 11, 2010
 Hidden Lecture 4: Hidden : An Introduction to Dynamic Decision Making November 11, 2010 Special Meeting 1/26 Markov Model Hidden When a dynamical system is probabilistic it may be determined by the transition
Hidden Lecture 4: Hidden : An Introduction to Dynamic Decision Making November 11, 2010 Special Meeting 1/26 Markov Model Hidden When a dynamical system is probabilistic it may be determined by the transition
Multiscale Systems Engineering Research Group
 Hidden Markov Model Prof. Yan Wang Woodruff School of Mechanical Engineering Georgia Institute of echnology Atlanta, GA 30332, U.S.A. yan.wang@me.gatech.edu Learning Objectives o familiarize the hidden
Hidden Markov Model Prof. Yan Wang Woodruff School of Mechanical Engineering Georgia Institute of echnology Atlanta, GA 30332, U.S.A. yan.wang@me.gatech.edu Learning Objectives o familiarize the hidden
Natural Language Processing Prof. Pushpak Bhattacharyya Department of Computer Science & Engineering, Indian Institute of Technology, Bombay
 Natural Language Processing Prof. Pushpak Bhattacharyya Department of Computer Science & Engineering, Indian Institute of Technology, Bombay Lecture - 21 HMM, Forward and Backward Algorithms, Baum Welch
Natural Language Processing Prof. Pushpak Bhattacharyya Department of Computer Science & Engineering, Indian Institute of Technology, Bombay Lecture - 21 HMM, Forward and Backward Algorithms, Baum Welch
1 What is a hidden Markov model?
 1 What is a hidden Markov model? Consider a Markov chain {X k }, where k is a non-negative integer. Suppose {X k } embedded in signals corrupted by some noise. Indeed, {X k } is hidden due to noise and
1 What is a hidden Markov model? Consider a Markov chain {X k }, where k is a non-negative integer. Suppose {X k } embedded in signals corrupted by some noise. Indeed, {X k } is hidden due to noise and
11.3 Decoding Algorithm
 11.3 Decoding Algorithm 393 For convenience, we have introduced π 0 and π n+1 as the fictitious initial and terminal states begin and end. This model defines the probability P(x π) for a given sequence
11.3 Decoding Algorithm 393 For convenience, we have introduced π 0 and π n+1 as the fictitious initial and terminal states begin and end. This model defines the probability P(x π) for a given sequence
Data Mining in Bioinformatics HMM
 Data Mining in Bioinformatics HMM Microarray Problem: Major Objective n Major Objective: Discover a comprehensive theory of life s organization at the molecular level 2 1 Data Mining in Bioinformatics
Data Mining in Bioinformatics HMM Microarray Problem: Major Objective n Major Objective: Discover a comprehensive theory of life s organization at the molecular level 2 1 Data Mining in Bioinformatics
MACHINE LEARNING 2 UGM,HMMS Lecture 7
 LOREM I P S U M Royal Institute of Technology MACHINE LEARNING 2 UGM,HMMS Lecture 7 THIS LECTURE DGM semantics UGM De-noising HMMs Applications (interesting probabilities) DP for generation probability
LOREM I P S U M Royal Institute of Technology MACHINE LEARNING 2 UGM,HMMS Lecture 7 THIS LECTURE DGM semantics UGM De-noising HMMs Applications (interesting probabilities) DP for generation probability
STA 414/2104: Machine Learning
 STA 414/2104: Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistics! rsalakhu@cs.toronto.edu! http://www.cs.toronto.edu/~rsalakhu/ Lecture 9 Sequential Data So far
STA 414/2104: Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistics! rsalakhu@cs.toronto.edu! http://www.cs.toronto.edu/~rsalakhu/ Lecture 9 Sequential Data So far
Weighted Finite-State Transducers in Computational Biology
 Weighted Finite-State Transducers in Computational Biology Mehryar Mohri Courant Institute of Mathematical Sciences mohri@cims.nyu.edu Joint work with Corinna Cortes (Google Research). 1 This Tutorial
Weighted Finite-State Transducers in Computational Biology Mehryar Mohri Courant Institute of Mathematical Sciences mohri@cims.nyu.edu Joint work with Corinna Cortes (Google Research). 1 This Tutorial
Dynamic Approaches: The Hidden Markov Model
 Dynamic Approaches: The Hidden Markov Model Davide Bacciu Dipartimento di Informatica Università di Pisa bacciu@di.unipi.it Machine Learning: Neural Networks and Advanced Models (AA2) Inference as Message
Dynamic Approaches: The Hidden Markov Model Davide Bacciu Dipartimento di Informatica Università di Pisa bacciu@di.unipi.it Machine Learning: Neural Networks and Advanced Models (AA2) Inference as Message
O 3 O 4 O 5. q 3. q 4. Transition
 Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Steven L. Scott. Presented by Ahmet Engin Ural
 Steven L. Scott Presented by Ahmet Engin Ural Overview of HMM Evaluating likelihoods The Likelihood Recursion The Forward-Backward Recursion Sampling HMM DG and FB samplers Autocovariance of samplers Some
Steven L. Scott Presented by Ahmet Engin Ural Overview of HMM Evaluating likelihoods The Likelihood Recursion The Forward-Backward Recursion Sampling HMM DG and FB samplers Autocovariance of samplers Some
CSCE 478/878 Lecture 9: Hidden. Markov. Models. Stephen Scott. Introduction. Outline. Markov. Chains. Hidden Markov Models. CSCE 478/878 Lecture 9:
 Useful for modeling/making predictions on sequential data E.g., biological sequences, text, series of sounds/spoken words Will return to graphical models that are generative sscott@cse.unl.edu 1 / 27 2
Useful for modeling/making predictions on sequential data E.g., biological sequences, text, series of sounds/spoken words Will return to graphical models that are generative sscott@cse.unl.edu 1 / 27 2
Pair Hidden Markov Models
 Pair Hidden Markov Models Scribe: Rishi Bedi Lecturer: Serafim Batzoglou January 29, 2015 1 Recap of HMMs alphabet: Σ = {b 1,...b M } set of states: Q = {1,..., K} transition probabilities: A = [a ij ]
Pair Hidden Markov Models Scribe: Rishi Bedi Lecturer: Serafim Batzoglou January 29, 2015 1 Recap of HMMs alphabet: Σ = {b 1,...b M } set of states: Q = {1,..., K} transition probabilities: A = [a ij ]
An Introduction to Bioinformatics Algorithms Hidden Markov Models
 Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
University of Cambridge. MPhil in Computer Speech Text & Internet Technology. Module: Speech Processing II. Lecture 2: Hidden Markov Models I
 University of Cambridge MPhil in Computer Speech Text & Internet Technology Module: Speech Processing II Lecture 2: Hidden Markov Models I o o o o o 1 2 3 4 T 1 b 2 () a 12 2 a 3 a 4 5 34 a 23 b () b ()
University of Cambridge MPhil in Computer Speech Text & Internet Technology Module: Speech Processing II Lecture 2: Hidden Markov Models I o o o o o 1 2 3 4 T 1 b 2 () a 12 2 a 3 a 4 5 34 a 23 b () b ()
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 11 Project
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 11 Project
Hidden Markov Models Part 2: Algorithms
 Hidden Markov Models Part 2: Algorithms CSE 6363 Machine Learning Vassilis Athitsos Computer Science and Engineering Department University of Texas at Arlington 1 Hidden Markov Model An HMM consists of:
Hidden Markov Models Part 2: Algorithms CSE 6363 Machine Learning Vassilis Athitsos Computer Science and Engineering Department University of Texas at Arlington 1 Hidden Markov Model An HMM consists of:
10. Hidden Markov Models (HMM) for Speech Processing. (some slides taken from Glass and Zue course)
 10. Hidden Markov Models (HMM) for Speech Processing (some slides taken from Glass and Zue course) Definition of an HMM The HMM are powerful statistical methods to characterize the observed samples of
10. Hidden Markov Models (HMM) for Speech Processing (some slides taken from Glass and Zue course) Definition of an HMM The HMM are powerful statistical methods to characterize the observed samples of
STATS 306B: Unsupervised Learning Spring Lecture 5 April 14
 STATS 306B: Unsupervised Learning Spring 2014 Lecture 5 April 14 Lecturer: Lester Mackey Scribe: Brian Do and Robin Jia 5.1 Discrete Hidden Markov Models 5.1.1 Recap In the last lecture, we introduced
STATS 306B: Unsupervised Learning Spring 2014 Lecture 5 April 14 Lecturer: Lester Mackey Scribe: Brian Do and Robin Jia 5.1 Discrete Hidden Markov Models 5.1.1 Recap In the last lecture, we introduced
Hidden Markov Model. Ying Wu. Electrical Engineering and Computer Science Northwestern University Evanston, IL 60208
 Hidden Markov Model Ying Wu Electrical Engineering and Computer Science Northwestern University Evanston, IL 60208 http://www.eecs.northwestern.edu/~yingwu 1/19 Outline Example: Hidden Coin Tossing Hidden
Hidden Markov Model Ying Wu Electrical Engineering and Computer Science Northwestern University Evanston, IL 60208 http://www.eecs.northwestern.edu/~yingwu 1/19 Outline Example: Hidden Coin Tossing Hidden
Hidden Markov Models. Ivan Gesteira Costa Filho IZKF Research Group Bioinformatics RWTH Aachen Adapted from:
 Hidden Markov Models Ivan Gesteira Costa Filho IZKF Research Group Bioinformatics RWTH Aachen Adapted from: www.ioalgorithms.info Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm
Hidden Markov Models Ivan Gesteira Costa Filho IZKF Research Group Bioinformatics RWTH Aachen Adapted from: www.ioalgorithms.info Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm
Hidden Markov Models. based on chapters from the book Durbin, Eddy, Krogh and Mitchison Biological Sequence Analysis via Shamir s lecture notes
 Hidden Markov Models based on chapters from the book Durbin, Eddy, Krogh and Mitchison Biological Sequence Analysis via Shamir s lecture notes music recognition deal with variations in - actual sound -
Hidden Markov Models based on chapters from the book Durbin, Eddy, Krogh and Mitchison Biological Sequence Analysis via Shamir s lecture notes music recognition deal with variations in - actual sound -
VL Algorithmen und Datenstrukturen für Bioinformatik ( ) WS15/2016 Woche 16
 VL Algorithmen und Datenstrukturen für Bioinformatik (19400001) WS15/2016 Woche 16 Tim Conrad AG Medical Bioinformatics Institut für Mathematik & Informatik, Freie Universität Berlin Based on slides by
VL Algorithmen und Datenstrukturen für Bioinformatik (19400001) WS15/2016 Woche 16 Tim Conrad AG Medical Bioinformatics Institut für Mathematik & Informatik, Freie Universität Berlin Based on slides by
Basic math for biology
 Basic math for biology Lei Li Florida State University, Feb 6, 2002 The EM algorithm: setup Parametric models: {P θ }. Data: full data (Y, X); partial data Y. Missing data: X. Likelihood and maximum likelihood
Basic math for biology Lei Li Florida State University, Feb 6, 2002 The EM algorithm: setup Parametric models: {P θ }. Data: full data (Y, X); partial data Y. Missing data: X. Likelihood and maximum likelihood
Automatic Speech Recognition (CS753)
 Automatic Speech Recognition (CS753) Lecture 6: Hidden Markov Models (Part II) Instructor: Preethi Jyothi Aug 10, 2017 Recall: Computing Likelihood Problem 1 (Likelihood): Given an HMM l =(A, B) and an
Automatic Speech Recognition (CS753) Lecture 6: Hidden Markov Models (Part II) Instructor: Preethi Jyothi Aug 10, 2017 Recall: Computing Likelihood Problem 1 (Likelihood): Given an HMM l =(A, B) and an
Applications of Hidden Markov Models
 18.417 Introduction to Computational Molecular Biology Lecture 18: November 9, 2004 Scribe: Chris Peikert Lecturer: Ross Lippert Editor: Chris Peikert Applications of Hidden Markov Models Review of Notation
18.417 Introduction to Computational Molecular Biology Lecture 18: November 9, 2004 Scribe: Chris Peikert Lecturer: Ross Lippert Editor: Chris Peikert Applications of Hidden Markov Models Review of Notation
Multiplex network inference
 (using hidden Markov models) University of Cambridge Bioinformatics Group Meeting 11 February 2016 Words of warning Disclaimer These slides have been produced by combining & translating two of my previous
(using hidden Markov models) University of Cambridge Bioinformatics Group Meeting 11 February 2016 Words of warning Disclaimer These slides have been produced by combining & translating two of my previous
Speech Recognition Lecture 8: Expectation-Maximization Algorithm, Hidden Markov Models.
 Speech Recognition Lecture 8: Expectation-Maximization Algorithm, Hidden Markov Models. Mehryar Mohri Courant Institute and Google Research mohri@cims.nyu.com This Lecture Expectation-Maximization (EM)
Speech Recognition Lecture 8: Expectation-Maximization Algorithm, Hidden Markov Models. Mehryar Mohri Courant Institute and Google Research mohri@cims.nyu.com This Lecture Expectation-Maximization (EM)
The Expectation-Maximization Algorithm
 1/29 EM & Latent Variable Models Gaussian Mixture Models EM Theory The Expectation-Maximization Algorithm Mihaela van der Schaar Department of Engineering Science University of Oxford MLE for Latent Variable
1/29 EM & Latent Variable Models Gaussian Mixture Models EM Theory The Expectation-Maximization Algorithm Mihaela van der Schaar Department of Engineering Science University of Oxford MLE for Latent Variable
Hidden Markov Models The three basic HMM problems (note: change in notation) Mitch Marcus CSE 391
 Hidden Markov Models The three basic HMM problems (note: change in notation) Mitch Marcus CSE 391 Parameters of an HMM States: A set of states S=s 1, s n Transition probabilities: A= a 1,1, a 1,2,, a n,n
Hidden Markov Models The three basic HMM problems (note: change in notation) Mitch Marcus CSE 391 Parameters of an HMM States: A set of states S=s 1, s n Transition probabilities: A= a 1,1, a 1,2,, a n,n
STOCHASTIC MODELING OF ENVIRONMENTAL TIME SERIES. Richard W. Katz LECTURE 5
 STOCHASTIC MODELING OF ENVIRONMENTAL TIME SERIES Richard W Katz LECTURE 5 (1) Hidden Markov Models: Applications (2) Hidden Markov Models: Viterbi Algorithm (3) Non-Homogeneous Hidden Markov Model (1)
STOCHASTIC MODELING OF ENVIRONMENTAL TIME SERIES Richard W Katz LECTURE 5 (1) Hidden Markov Models: Applications (2) Hidden Markov Models: Viterbi Algorithm (3) Non-Homogeneous Hidden Markov Model (1)
Lecture 9. Intro to Hidden Markov Models (finish up)
 Lecture 9 Intro to Hidden Markov Models (finish up) Review Structure Number of states Q 1.. Q N M output symbols Parameters: Transition probability matrix a ij Emission probabilities b i (a), which is
Lecture 9 Intro to Hidden Markov Models (finish up) Review Structure Number of states Q 1.. Q N M output symbols Parameters: Transition probability matrix a ij Emission probabilities b i (a), which is
L23: hidden Markov models
 L23: hidden Markov models Discrete Markov processes Hidden Markov models Forward and Backward procedures The Viterbi algorithm This lecture is based on [Rabiner and Juang, 1993] Introduction to Speech
L23: hidden Markov models Discrete Markov processes Hidden Markov models Forward and Backward procedures The Viterbi algorithm This lecture is based on [Rabiner and Juang, 1993] Introduction to Speech
HMM : Viterbi algorithm - a toy example
 MM : Viterbi algorithm - a toy example 0.6 et's consider the following simple MM. This model is composed of 2 states, (high GC content) and (low GC content). We can for example consider that state characterizes
MM : Viterbi algorithm - a toy example 0.6 et's consider the following simple MM. This model is composed of 2 states, (high GC content) and (low GC content). We can for example consider that state characterizes
Hidden Markov Models
 CS769 Spring 2010 Advanced Natural Language Processing Hidden Markov Models Lecturer: Xiaojin Zhu jerryzhu@cs.wisc.edu 1 Part-of-Speech Tagging The goal of Part-of-Speech (POS) tagging is to label each
CS769 Spring 2010 Advanced Natural Language Processing Hidden Markov Models Lecturer: Xiaojin Zhu jerryzhu@cs.wisc.edu 1 Part-of-Speech Tagging The goal of Part-of-Speech (POS) tagging is to label each
Computer Intensive Methods in Mathematical Statistics
 Computer Intensive Methods in Mathematical Statistics Department of mathematics johawes@kth.se Lecture 16 Advanced topics in computational statistics 18 May 2017 Computer Intensive Methods (1) Plan of
Computer Intensive Methods in Mathematical Statistics Department of mathematics johawes@kth.se Lecture 16 Advanced topics in computational statistics 18 May 2017 Computer Intensive Methods (1) Plan of
Hidden Markov Models
 Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Markov chains and Hidden Markov Models
 Discrete Math for Bioinformatics WS 10/11:, b A. Bockmar/K. Reinert, 7. November 2011, 10:24 2001 Markov chains and Hidden Markov Models We will discuss: Hidden Markov Models (HMMs) Algorithms: Viterbi,
Discrete Math for Bioinformatics WS 10/11:, b A. Bockmar/K. Reinert, 7. November 2011, 10:24 2001 Markov chains and Hidden Markov Models We will discuss: Hidden Markov Models (HMMs) Algorithms: Viterbi,
Hidden Markov Models. By Parisa Abedi. Slides courtesy: Eric Xing
 Hidden Markov Models By Parisa Abedi Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed data Sequential (non i.i.d.) data Time-series data E.g. Speech
Hidden Markov Models By Parisa Abedi Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed data Sequential (non i.i.d.) data Time-series data E.g. Speech
Giri Narasimhan. CAP 5510: Introduction to Bioinformatics. ECS 254; Phone: x3748
 CAP 5510: Introduction to Bioinformatics Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs07.html 2/14/07 CAP5510 1 CpG Islands Regions in DNA sequences with increased
CAP 5510: Introduction to Bioinformatics Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs07.html 2/14/07 CAP5510 1 CpG Islands Regions in DNA sequences with increased
Plan for today. ! Part 1: (Hidden) Markov models. ! Part 2: String matching and read mapping
 Plan for today! Part 1: (Hidden) Markov models! Part 2: String matching and read mapping! 2.1 Exact algorithms! 2.2 Heuristic methods for approximate search (Hidden) Markov models Why consider probabilistics
Plan for today! Part 1: (Hidden) Markov models! Part 2: String matching and read mapping! 2.1 Exact algorithms! 2.2 Heuristic methods for approximate search (Hidden) Markov models Why consider probabilistics
Markov Chains and Hidden Markov Models
 Chapter 1 Markov Chains and Hidden Markov Models In this chapter, we will introduce the concept of Markov chains, and show how Markov chains can be used to model signals using structures such as hidden
Chapter 1 Markov Chains and Hidden Markov Models In this chapter, we will introduce the concept of Markov chains, and show how Markov chains can be used to model signals using structures such as hidden
Linear Dynamical Systems
 Linear Dynamical Systems Sargur N. srihari@cedar.buffalo.edu Machine Learning Course: http://www.cedar.buffalo.edu/~srihari/cse574/index.html Two Models Described by Same Graph Latent variables Observations
Linear Dynamical Systems Sargur N. srihari@cedar.buffalo.edu Machine Learning Course: http://www.cedar.buffalo.edu/~srihari/cse574/index.html Two Models Described by Same Graph Latent variables Observations
Hidden Markov Models: Maxing and Summing
 Hidden Markov Models: Maxing and Summing Introduction to Natural Language Processing Computer Science 585 Fall 2009 University of Massachusetts Amherst David Smith 1 Markov vs. Hidden Markov Models Fed
Hidden Markov Models: Maxing and Summing Introduction to Natural Language Processing Computer Science 585 Fall 2009 University of Massachusetts Amherst David Smith 1 Markov vs. Hidden Markov Models Fed
Markov Chains and Hidden Markov Models. COMP 571 Luay Nakhleh, Rice University
 Markov Chains and Hidden Markov Models COMP 571 Luay Nakhleh, Rice University Markov Chains and Hidden Markov Models Modeling the statistical properties of biological sequences and distinguishing regions
Markov Chains and Hidden Markov Models COMP 571 Luay Nakhleh, Rice University Markov Chains and Hidden Markov Models Modeling the statistical properties of biological sequences and distinguishing regions
Lecture 21: Spectral Learning for Graphical Models
 10-708: Probabilistic Graphical Models 10-708, Spring 2016 Lecture 21: Spectral Learning for Graphical Models Lecturer: Eric P. Xing Scribes: Maruan Al-Shedivat, Wei-Cheng Chang, Frederick Liu 1 Motivation
10-708: Probabilistic Graphical Models 10-708, Spring 2016 Lecture 21: Spectral Learning for Graphical Models Lecturer: Eric P. Xing Scribes: Maruan Al-Shedivat, Wei-Cheng Chang, Frederick Liu 1 Motivation
Today s Lecture: HMMs
 Today s Lecture: HMMs Definitions Examples Probability calculations WDAG Dynamic programming algorithms: Forward Viterbi Parameter estimation Viterbi training 1 Hidden Markov Models Probability models
Today s Lecture: HMMs Definitions Examples Probability calculations WDAG Dynamic programming algorithms: Forward Viterbi Parameter estimation Viterbi training 1 Hidden Markov Models Probability models
Hidden Markov Models and Gaussian Mixture Models
 Hidden Markov Models and Gaussian Mixture Models Hiroshi Shimodaira and Steve Renals Automatic Speech Recognition ASR Lectures 4&5 23&27 January 2014 ASR Lectures 4&5 Hidden Markov Models and Gaussian
Hidden Markov Models and Gaussian Mixture Models Hiroshi Shimodaira and Steve Renals Automatic Speech Recognition ASR Lectures 4&5 23&27 January 2014 ASR Lectures 4&5 Hidden Markov Models and Gaussian
Statistical Methods for NLP
 Statistical Methods for NLP Sequence Models Joakim Nivre Uppsala University Department of Linguistics and Philology joakim.nivre@lingfil.uu.se Statistical Methods for NLP 1(21) Introduction Structured
Statistical Methods for NLP Sequence Models Joakim Nivre Uppsala University Department of Linguistics and Philology joakim.nivre@lingfil.uu.se Statistical Methods for NLP 1(21) Introduction Structured
Hidden Markov Models. Aarti Singh Slides courtesy: Eric Xing. Machine Learning / Nov 8, 2010
 Hidden Markov Models Aarti Singh Slides courtesy: Eric Xing Machine Learning 10-701/15-781 Nov 8, 2010 i.i.d to sequential data So far we assumed independent, identically distributed data Sequential data
Hidden Markov Models Aarti Singh Slides courtesy: Eric Xing Machine Learning 10-701/15-781 Nov 8, 2010 i.i.d to sequential data So far we assumed independent, identically distributed data Sequential data
CSCE 471/871 Lecture 3: Markov Chains and
 and and 1 / 26 sscott@cse.unl.edu 2 / 26 Outline and chains models (s) Formal definition Finding most probable state path (Viterbi algorithm) Forward and backward algorithms State sequence known State
and and 1 / 26 sscott@cse.unl.edu 2 / 26 Outline and chains models (s) Formal definition Finding most probable state path (Viterbi algorithm) Forward and backward algorithms State sequence known State
order is number of previous outputs
 Markov Models Lecture : Markov and Hidden Markov Models PSfrag Use past replacements as state. Next output depends on previous output(s): y t = f[y t, y t,...] order is number of previous outputs y t y
Markov Models Lecture : Markov and Hidden Markov Models PSfrag Use past replacements as state. Next output depends on previous output(s): y t = f[y t, y t,...] order is number of previous outputs y t y
p(d θ ) l(θ ) 1.2 x x x
 p(d θ ).2 x 0-7 0.8 x 0-7 0.4 x 0-7 l(θ ) -20-40 -60-80 -00 2 3 4 5 6 7 θ ˆ 2 3 4 5 6 7 θ ˆ 2 3 4 5 6 7 θ θ x FIGURE 3.. The top graph shows several training points in one dimension, known or assumed to
p(d θ ).2 x 0-7 0.8 x 0-7 0.4 x 0-7 l(θ ) -20-40 -60-80 -00 2 3 4 5 6 7 θ ˆ 2 3 4 5 6 7 θ ˆ 2 3 4 5 6 7 θ θ x FIGURE 3.. The top graph shows several training points in one dimension, known or assumed to
HMM : Viterbi algorithm - a toy example
 MM : Viterbi algorithm - a toy example 0.5 0.5 0.5 A 0.2 A 0.3 0.5 0.6 C 0.3 C 0.2 G 0.3 0.4 G 0.2 T 0.2 T 0.3 et's consider the following simple MM. This model is composed of 2 states, (high GC content)
MM : Viterbi algorithm - a toy example 0.5 0.5 0.5 A 0.2 A 0.3 0.5 0.6 C 0.3 C 0.2 G 0.3 0.4 G 0.2 T 0.2 T 0.3 et's consider the following simple MM. This model is composed of 2 states, (high GC content)
Statistical Methods for NLP
 Statistical Methods for NLP Information Extraction, Hidden Markov Models Sameer Maskey Week 5, Oct 3, 2012 *many slides provided by Bhuvana Ramabhadran, Stanley Chen, Michael Picheny Speech Recognition
Statistical Methods for NLP Information Extraction, Hidden Markov Models Sameer Maskey Week 5, Oct 3, 2012 *many slides provided by Bhuvana Ramabhadran, Stanley Chen, Michael Picheny Speech Recognition
Hidden Markov Models. Hosein Mohimani GHC7717
 Hidden Markov Models Hosein Mohimani GHC7717 hoseinm@andrew.cmu.edu Fair et Casino Problem Dealer flips a coin and player bets on outcome Dealer use either a fair coin (head and tail equally likely) or
Hidden Markov Models Hosein Mohimani GHC7717 hoseinm@andrew.cmu.edu Fair et Casino Problem Dealer flips a coin and player bets on outcome Dealer use either a fair coin (head and tail equally likely) or
Bayesian Networks: Construction, Inference, Learning and Causal Interpretation. Volker Tresp Summer 2014
 Bayesian Networks: Construction, Inference, Learning and Causal Interpretation Volker Tresp Summer 2014 1 Introduction So far we were mostly concerned with supervised learning: we predicted one or several
Bayesian Networks: Construction, Inference, Learning and Causal Interpretation Volker Tresp Summer 2014 1 Introduction So far we were mostly concerned with supervised learning: we predicted one or several
Statistical Pattern Recognition
 Statistical Pattern Recognition Expectation Maximization (EM) and Mixture Models Hamid R. Rabiee Jafar Muhammadi, Mohammad J. Hosseini Spring 2014 http://ce.sharif.edu/courses/92-93/2/ce725-2 Agenda Expectation-maximization
Statistical Pattern Recognition Expectation Maximization (EM) and Mixture Models Hamid R. Rabiee Jafar Muhammadi, Mohammad J. Hosseini Spring 2014 http://ce.sharif.edu/courses/92-93/2/ce725-2 Agenda Expectation-maximization
Hidden Markov Models (I)
 GLOBEX Bioinformatics (Summer 2015) Hidden Markov Models (I) a. The model b. The decoding: Viterbi algorithm Hidden Markov models A Markov chain of states At each state, there are a set of possible observables
GLOBEX Bioinformatics (Summer 2015) Hidden Markov Models (I) a. The model b. The decoding: Viterbi algorithm Hidden Markov models A Markov chain of states At each state, there are a set of possible observables
Assignments for lecture Bioinformatics III WS 03/04. Assignment 5, return until Dec 16, 2003, 11 am. Your name: Matrikelnummer: Fachrichtung:
 Assignments for lecture Bioinformatics III WS 03/04 Assignment 5, return until Dec 16, 2003, 11 am Your name: Matrikelnummer: Fachrichtung: Please direct questions to: Jörg Niggemann, tel. 302-64167, email:
Assignments for lecture Bioinformatics III WS 03/04 Assignment 5, return until Dec 16, 2003, 11 am Your name: Matrikelnummer: Fachrichtung: Please direct questions to: Jörg Niggemann, tel. 302-64167, email:
Hidden Markov Models for biological sequence analysis
 Hidden Markov Models for biological sequence analysis Master in Bioinformatics UPF 2017-2018 http://comprna.upf.edu/courses/master_agb/ Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA
Hidden Markov Models for biological sequence analysis Master in Bioinformatics UPF 2017-2018 http://comprna.upf.edu/courses/master_agb/ Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA
Machine Learning & Data Mining Caltech CS/CNS/EE 155 Hidden Markov Models Last Updated: Feb 7th, 2017
 1 Introduction Let x = (x 1,..., x M ) denote a sequence (e.g. a sequence of words), and let y = (y 1,..., y M ) denote a corresponding hidden sequence that we believe explains or influences x somehow
1 Introduction Let x = (x 1,..., x M ) denote a sequence (e.g. a sequence of words), and let y = (y 1,..., y M ) denote a corresponding hidden sequence that we believe explains or influences x somehow
Introduction to Machine Learning CMU-10701
 Introduction to Machine Learning CMU-10701 Hidden Markov Models Barnabás Póczos & Aarti Singh Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed
Introduction to Machine Learning CMU-10701 Hidden Markov Models Barnabás Póczos & Aarti Singh Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed
Probabilistic Graphical Models Homework 2: Due February 24, 2014 at 4 pm
 Probabilistic Graphical Models 10-708 Homework 2: Due February 24, 2014 at 4 pm Directions. This homework assignment covers the material presented in Lectures 4-8. You must complete all four problems to
Probabilistic Graphical Models 10-708 Homework 2: Due February 24, 2014 at 4 pm Directions. This homework assignment covers the material presented in Lectures 4-8. You must complete all four problems to
Machine Learning and Bayesian Inference. Unsupervised learning. Can we find regularity in data without the aid of labels?
 Machine Learning and Bayesian Inference Dr Sean Holden Computer Laboratory, Room FC6 Telephone extension 6372 Email: sbh11@cl.cam.ac.uk www.cl.cam.ac.uk/ sbh11/ Unsupervised learning Can we find regularity
Machine Learning and Bayesian Inference Dr Sean Holden Computer Laboratory, Room FC6 Telephone extension 6372 Email: sbh11@cl.cam.ac.uk www.cl.cam.ac.uk/ sbh11/ Unsupervised learning Can we find regularity
State Space and Hidden Markov Models
 State Space and Hidden Markov Models Kunsch H.R. State Space and Hidden Markov Models. ETH- Zurich Zurich; Aliaksandr Hubin Oslo 2014 Contents 1. Introduction 2. Markov Chains 3. Hidden Markov and State
State Space and Hidden Markov Models Kunsch H.R. State Space and Hidden Markov Models. ETH- Zurich Zurich; Aliaksandr Hubin Oslo 2014 Contents 1. Introduction 2. Markov Chains 3. Hidden Markov and State
Conditional Random Field
 Introduction Linear-Chain General Specific Implementations Conclusions Corso di Elaborazione del Linguaggio Naturale Pisa, May, 2011 Introduction Linear-Chain General Specific Implementations Conclusions
Introduction Linear-Chain General Specific Implementations Conclusions Corso di Elaborazione del Linguaggio Naturale Pisa, May, 2011 Introduction Linear-Chain General Specific Implementations Conclusions
Hidden Markov Models. Terminology, Representation and Basic Problems
 Hidden Markov Models Terminology, Representation and Basic Problems Data analysis? Machine learning? In bioinformatics, we analyze a lot of (sequential) data (biological sequences) to learn unknown parameters
Hidden Markov Models Terminology, Representation and Basic Problems Data analysis? Machine learning? In bioinformatics, we analyze a lot of (sequential) data (biological sequences) to learn unknown parameters
Infering the Number of State Clusters in Hidden Markov Model and its Extension
 Infering the Number of State Clusters in Hidden Markov Model and its Extension Xugang Ye Department of Applied Mathematics and Statistics, Johns Hopkins University Elements of a Hidden Markov Model (HMM)
Infering the Number of State Clusters in Hidden Markov Model and its Extension Xugang Ye Department of Applied Mathematics and Statistics, Johns Hopkins University Elements of a Hidden Markov Model (HMM)
Page 1. References. Hidden Markov models and multiple sequence alignment. Markov chains. Probability review. Example. Markovian sequence
 Page Hidden Markov models and multiple sequence alignment Russ B Altman BMI 4 CS 74 Some slides borrowed from Scott C Schmidler (BMI graduate student) References Bioinformatics Classic: Krogh et al (994)
Page Hidden Markov models and multiple sequence alignment Russ B Altman BMI 4 CS 74 Some slides borrowed from Scott C Schmidler (BMI graduate student) References Bioinformatics Classic: Krogh et al (994)
Example: The Dishonest Casino. Hidden Markov Models. Question # 1 Evaluation. The dishonest casino model. Question # 3 Learning. Question # 2 Decoding
 Example: The Dishonest Casino Hidden Markov Models Durbin and Eddy, chapter 3 Game:. You bet $. You roll 3. Casino player rolls 4. Highest number wins $ The casino has two dice: Fair die P() = P() = P(3)
Example: The Dishonest Casino Hidden Markov Models Durbin and Eddy, chapter 3 Game:. You bet $. You roll 3. Casino player rolls 4. Highest number wins $ The casino has two dice: Fair die P() = P() = P(3)
Graphical Models for Collaborative Filtering
 Graphical Models for Collaborative Filtering Le Song Machine Learning II: Advanced Topics CSE 8803ML, Spring 2012 Sequence modeling HMM, Kalman Filter, etc.: Similarity: the same graphical model topology,
Graphical Models for Collaborative Filtering Le Song Machine Learning II: Advanced Topics CSE 8803ML, Spring 2012 Sequence modeling HMM, Kalman Filter, etc.: Similarity: the same graphical model topology,
Hidden Markov Models. Terminology and Basic Algorithms
 Hidden Markov Models Terminology and Basic Algorithms What is machine learning? From http://en.wikipedia.org/wiki/machine_learning Machine learning, a branch of artificial intelligence, is about the construction
Hidden Markov Models Terminology and Basic Algorithms What is machine learning? From http://en.wikipedia.org/wiki/machine_learning Machine learning, a branch of artificial intelligence, is about the construction
Statistical NLP: Hidden Markov Models. Updated 12/15
 Statistical NLP: Hidden Markov Models Updated 12/15 Markov Models Markov models are statistical tools that are useful for NLP because they can be used for part-of-speech-tagging applications Their first
Statistical NLP: Hidden Markov Models Updated 12/15 Markov Models Markov models are statistical tools that are useful for NLP because they can be used for part-of-speech-tagging applications Their first
Shankar Shivappa University of California, San Diego April 26, CSE 254 Seminar in learning algorithms
 Recognition of Visual Speech Elements Using Adaptively Boosted Hidden Markov Models. Say Wei Foo, Yong Lian, Liang Dong. IEEE Transactions on Circuits and Systems for Video Technology, May 2004. Shankar
Recognition of Visual Speech Elements Using Adaptively Boosted Hidden Markov Models. Say Wei Foo, Yong Lian, Liang Dong. IEEE Transactions on Circuits and Systems for Video Technology, May 2004. Shankar
Basic Text Analysis. Hidden Markov Models. Joakim Nivre. Uppsala University Department of Linguistics and Philology
 Basic Text Analysis Hidden Markov Models Joakim Nivre Uppsala University Department of Linguistics and Philology joakimnivre@lingfiluuse Basic Text Analysis 1(33) Hidden Markov Models Markov models are
Basic Text Analysis Hidden Markov Models Joakim Nivre Uppsala University Department of Linguistics and Philology joakimnivre@lingfiluuse Basic Text Analysis 1(33) Hidden Markov Models Markov models are
9 Forward-backward algorithm, sum-product on factor graphs
 Massachusetts Institute of Technology Department of Electrical Engineering and Computer Science 6.438 Algorithms For Inference Fall 2014 9 Forward-backward algorithm, sum-product on factor graphs The previous
Massachusetts Institute of Technology Department of Electrical Engineering and Computer Science 6.438 Algorithms For Inference Fall 2014 9 Forward-backward algorithm, sum-product on factor graphs The previous
Advanced Data Science
 Advanced Data Science Dr. Kira Radinsky Slides Adapted from Tom M. Mitchell Agenda Topics Covered: Time series data Markov Models Hidden Markov Models Dynamic Bayes Nets Additional Reading: Bishop: Chapter
Advanced Data Science Dr. Kira Radinsky Slides Adapted from Tom M. Mitchell Agenda Topics Covered: Time series data Markov Models Hidden Markov Models Dynamic Bayes Nets Additional Reading: Bishop: Chapter
Sequence labeling. Taking collective a set of interrelated instances x 1,, x T and jointly labeling them
 HMM, MEMM and CRF 40-957 Special opics in Artificial Intelligence: Probabilistic Graphical Models Sharif University of echnology Soleymani Spring 2014 Sequence labeling aking collective a set of interrelated
HMM, MEMM and CRF 40-957 Special opics in Artificial Intelligence: Probabilistic Graphical Models Sharif University of echnology Soleymani Spring 2014 Sequence labeling aking collective a set of interrelated
Human Mobility Pattern Prediction Algorithm using Mobile Device Location and Time Data
 Human Mobility Pattern Prediction Algorithm using Mobile Device Location and Time Data 0. Notations Myungjun Choi, Yonghyun Ro, Han Lee N = number of states in the model T = length of observation sequence
Human Mobility Pattern Prediction Algorithm using Mobile Device Location and Time Data 0. Notations Myungjun Choi, Yonghyun Ro, Han Lee N = number of states in the model T = length of observation sequence
Hidden Markov Models
 Hidden Markov Models Lecture Notes Speech Communication 2, SS 2004 Erhard Rank/Franz Pernkopf Signal Processing and Speech Communication Laboratory Graz University of Technology Inffeldgasse 16c, A-8010
Hidden Markov Models Lecture Notes Speech Communication 2, SS 2004 Erhard Rank/Franz Pernkopf Signal Processing and Speech Communication Laboratory Graz University of Technology Inffeldgasse 16c, A-8010
We Live in Exciting Times. CSCI-567: Machine Learning (Spring 2019) Outline. Outline. ACM (an international computing research society) has named
 We Live in Exciting Times ACM (an international computing research society) has named CSCI-567: Machine Learning (Spring 2019) Prof. Victor Adamchik U of Southern California Apr. 2, 2019 Yoshua Bengio,
We Live in Exciting Times ACM (an international computing research society) has named CSCI-567: Machine Learning (Spring 2019) Prof. Victor Adamchik U of Southern California Apr. 2, 2019 Yoshua Bengio,
Recall: Modeling Time Series. CSE 586, Spring 2015 Computer Vision II. Hidden Markov Model and Kalman Filter. Modeling Time Series
 Recall: Modeling Time Series CSE 586, Spring 2015 Computer Vision II Hidden Markov Model and Kalman Filter State-Space Model: You have a Markov chain of latent (unobserved) states Each state generates
Recall: Modeling Time Series CSE 586, Spring 2015 Computer Vision II Hidden Markov Model and Kalman Filter State-Space Model: You have a Markov chain of latent (unobserved) states Each state generates
Unit 16: Hidden Markov Models
 Computational Statistics with Application to Bioinformatics Prof. William H. Press Spring Term, 2008 The University of Texas at Austin Unit 16: Hidden Markov Models The University of Texas at Austin, CS
Computational Statistics with Application to Bioinformatics Prof. William H. Press Spring Term, 2008 The University of Texas at Austin Unit 16: Hidden Markov Models The University of Texas at Austin, CS
Introduction to Computational Biology Lecture # 14: MCMC - Markov Chain Monte Carlo
 Introduction to Computational Biology Lecture # 14: MCMC - Markov Chain Monte Carlo Assaf Weiner Tuesday, March 13, 2007 1 Introduction Today we will return to the motif finding problem, in lecture 10
Introduction to Computational Biology Lecture # 14: MCMC - Markov Chain Monte Carlo Assaf Weiner Tuesday, March 13, 2007 1 Introduction Today we will return to the motif finding problem, in lecture 10
LECTURER: BURCU CAN Spring
 LECTURER: BURCU CAN 2017-2018 Spring Regular Language Hidden Markov Model (HMM) Context Free Language Context Sensitive Language Probabilistic Context Free Grammar (PCFG) Unrestricted Language PCFGs can
LECTURER: BURCU CAN 2017-2018 Spring Regular Language Hidden Markov Model (HMM) Context Free Language Context Sensitive Language Probabilistic Context Free Grammar (PCFG) Unrestricted Language PCFGs can
Statistical Sequence Recognition and Training: An Introduction to HMMs
 Statistical Sequence Recognition and Training: An Introduction to HMMs EECS 225D Nikki Mirghafori nikki@icsi.berkeley.edu March 7, 2005 Credit: many of the HMM slides have been borrowed and adapted, with
Statistical Sequence Recognition and Training: An Introduction to HMMs EECS 225D Nikki Mirghafori nikki@icsi.berkeley.edu March 7, 2005 Credit: many of the HMM slides have been borrowed and adapted, with
