Example: The Dishonest Casino. Hidden Markov Models. Question # 1 Evaluation. The dishonest casino model. Question # 3 Learning. Question # 2 Decoding
|
|
|
- Reynold King
- 6 years ago
- Views:
Transcription
1 Example: The Dishonest Casino Hidden Markov Models Durbin and Eddy, chapter 3 Game:. You bet $. You roll 3. Casino player rolls 4. Highest number wins $ The casino has two dice: Fair die P() = P() = P(3) = P(5) = P(6) = /6 Loaded die P() = P() = P(3) = P(5) = /0 P(6) = / Casino player switches between fair and loaded die (not too often, and not for too long) The dishonest casino model Question # Evaluation GIVEN: A sequence of rolls by the casino player FAIR LOADED QUESTION: Prob =.3 x 0-35 P( F) = /6 P( F) = /6 P(3 F) = /6 P(4 F) = /6 P(5 F) = /6 P(6 F) = /6 0. P( L) = /0 P( L) = /0 P(3 L) = /0 P(4 L) = /0 P(5 L) = /0 P(6 L) = / How likely is this sequence, given our model of how the casino works? This is the EVALUATION problem in HMMs Question # Decoding Question # 3 Learning GIVEN: A sequence of rolls by the casino player QUESTION: FAIR LOADED FAIR What portion of the sequence was generated with the fair die, and what portion with the loaded die? GIVEN: A sequence of rolls by the casino player QUESTION: How does the casino player work: How loaded is the loaded die? How fair is the fair die? How often does the casino player change from fair to loaded, and back? This is the LEARNING question in HMMs This is the DECODING question in HMMs
2 The dishonest casino model Definition of a hidden Markov model Alphabet Σ = { b, b,,b M } Set of states Q = {,..., } ( = Q ) Transition probabilities between any two states FAIR LOADED a ij = transition probability from state i to state j a i + + a i =, for all states i P( F) = /6 P( F) = /6 P(3 F) = /6 P(4 F) = /6 P(5 F) = /6 P(6 F) = /6 0. P( L) = /0 P( L) = /0 P(3 L) = /0 P(4 L) = /0 P(5 L) = /0 P(6 L) = / Initial probabilities a 0i a a 0 = Emission probabilities within each state e k (b) = P( x i = b π i = k) e k (b ) + + e k (b M ) = Hidden states and observed sequence An HMM is memory-less At time step t, π t denotes the (hidden) state in the Markov chain x t denotes the symbol emitted in state π t At time step t, the only thing that affects the next state is the current State, π t A path of length N is: An observed sequence of length N is: π, π,, π N x, x,, x N P(π t+ = k whatever happened so far ) = P(π t+ = k π, π,, π t, x, x,, x t ) = P(π t+ = k π t ) A parse of a sequence Given a sequence x = x x N, A parse of x is a sequence of states π = π,, π N Given a sequence x = x,x N and a parse π = π,,π N, How likely is the parse (given our HMM)? P(x, π) = P(x,, x N, π,, π N ) Likelihood of a parse = P(x N, π N x x N-, π,, π N- ) P(x x N-, π,, π N- ) = P(x N, π N π N- ) P(x x N-, π,, π N- ) = = P(x N, π N π N- ) P(x N-, π N- π N- )P(x, π π ) P(x, π ) = P(x N π N ) P(π N π N- ) P(x π ) P(π π ) P(x π ) P(π ) = a 0π a ππ a πn-πn e π (x )e πn (x N ) N =! a# e ( x i" i i i ) # # i=
3 Example: the dishonest casino What is the probability of a sequence of rolls x =,,, 5, 6,,, 6,, 4 and the parse π = Fair, Fair, Fair, Fair, Fair, Fair, Fair, Fair, Fair, Fair? (say initial probs a 0,Fair = ½, a 0,Loaded = ½) ½ P( Fair) P(Fair Fair) P( Fair) P(Fair Fair) P(4 Fair) = ½ (/6) 0 (0.95) 9 = Example: the dishonest casino So, the likelihood the die is fair in all this run is What about π = Loaded, Loaded, Loaded, Loaded, Loaded, Loaded, Loaded, Loaded, Loaded, Loaded? ½ P( Loaded) P(Loaded Loaded) P(4 Loaded) = ½ (/0) 8 (/) (0.9) 9 = Therefore, it is more likely that the die is fair all the way, than loaded all the way Example: the dishonest casino Let the sequence of rolls be: x =, 6, 6, 5, 6,, 6, 6, 3, 6 And let s consider π = F, F,, F P(x, π) = ½ (/6) 0 (0.95) 9 = (same as before) And for π = L, L,, L: P(x, π) = ½ (/0) 4 (/) 6 (0.9) 9 = So, the observed sequence is ~00 times more likely if a loaded die is used Clarification of notation P[ x M ]: The probability that sequence x was generated by the model The model is: architecture (#states, etc) + parameters θ = a ij, e i (.) So, P[x M] is the same as P[ x θ ], and P[ x ], when the architecture, and the parameters, respectively, are implied Similarly, P[ x, π M ], P[ x, π θ ] and P[ x, π ] are the same when the architecture, and the parameters, are implied In the LEARNING problem we write P[ x θ ] to emphasize that we are seeking the θ* that maximizes P[ x θ ] What we know Given a sequence x = x,,x N and a parse π = π,,π N, we know how to compute how likely the parse is: P(x, π). Evaluation What we would like to know GIVEN HMM M, and a sequence x, FIND Prob[ x M ]. Decoding GIVEN HMM M, and a sequence x, FIND the sequence π of states that maximizes P[ x, π M ] 3. Learning GIVEN HMM M, with unspecified transition/emission probs., and a sequence x, FIND parameters θ = (e i (.), a ij ) that maximize P[ x θ ] 3
4 Decoding Problem : Decoding GIVEN x = x x,,x N We want to find π = π,, π N, such that P[ x, π ] is maximized π * = argmax π P[ x, π ] Maximize a 0π e π (x ) a ππ a πn-πn e πn (x N ) Find the best parse of a sequence We can use dynamic programming! Let V k (i) = max {π,, π i-} P[x x i-, π,, π i-, x i, π i = k] = Probability of maximum probability path ending at state π i = k Decoding main idea Inductive assumption: Given V k (i) = max {π,, π i-} P[x x i-, π,, π i-, x i, π i = k] What is V r (i+)? V r (i+) = max {π,, πi} P[ x x i, π,, π i, x i+, π i+ = r ] = max {π,, πi} P(x i+, π i+ = r x x i,π,, π i ) P[x x i, π,, π i ] = max {π,, πi} P(x i+, π i+ = r π i ) P[x x i-, π,, π i-, x i, π i ] = max k [P(x i+, π i+ = r π i = k) max {π,, π i-} P[x x i-,π,,π i-, x i,π i =k]] = max k e r (x i+ ) a kr V k (i) = e r (x i+ ) max k a kr V k (i) The Viterbi Algorithm Input: x = x,,x N V 0 (0) = (0 is the imaginary first position) V k (0) = 0, for all k > 0 V j (i) = e j (x i ) max k a kj V k (i-) Ptr j (i) = argmax k a kj V k (i-) P(x, π*) = max k V k (N) Traceback: π N * = argmax k V k (N) π i- * = Ptr πi (i) The Viterbi Algorithm The Viterbi Algorithm Input: x = x,,x N V 0 (0) = (0 is the imaginary first position) V k (0) = 0, for all k > 0 x i- Traceback: V j (i) = e j (x i ) max k a kj V k (i-) Ptr j (i) = argmax k a kj V k (i-) P(x, π*) = max k a k0 V k (N) (with an end state) π N * = argmax k V k (N) π i- * = Ptr πi (i) V j (i) Similar to aligning a set of states to a sequence Time: O( N) Space: O(N) 4
5 The edit graph for the decoding problem Viterbi Algorithm a practical detail Underflows are a significant problem P[ x,., x i, π,, π i ] = a 0π a ππ,,a πi e π (x ),,e πi (x i ) These numbers become extremely small underflow Solution: Take the logs of all values V r (i) = log e k (x i ) + max k [ V k (i-) + log a kr ] The Decoding Problem is essentially finding a longest path in a directed acyclic graph (DAG) Example Let x be a sequence with a portion that has /6 6 s, followed by a portion with ½ 6 s x = Then, it is not hard to show that the optimal parse is: FFF...F LLL...L Example Observed Sequence: x =,,,6,6 P(x) = Best 8 paths: LLLLL FFFFF FFFLL FFLLL FLLLL FFFFL LLLLF LFFFF Conditional Probabilities given x Position F L Generating a sequence by the model Given a HMM, we can generate a sequence of length n as follows: Problem : Evaluation Finding the probability a sequence is generated by the model. Start at state π according to probability a 0π. Emit letter x according to probability e π (x ) 3. Go to state π according to probability a ππ 4. until emitting x n 0 5
6 The Forward Algorithm The Forward Algorithm derivation want to calculate P(x) = probability of x, given the HMM (x = x,,x N ) Sum over all possible ways of generating x: P(x) = Σ all paths π P(x, π) To avoid summing over an exponential number of paths π, define f k (i) = P(x,,x i, π i = k) (the forward probability) the forward probability: f k (i) = P(x x i, π i = k) = Σ ππi- P(x x i-, π,, π i-, π i = k) e k (x i ) = Σ r Σ ππi- P(x x i-, π,, π i-, π i- = r) a rk e k (x i ) = Σ r P(x x i-, π i- = r) a rk e k (x i ) = e k (x i ) Σ r f r (i-) a rk The Forward Algorithm A dynamic programming algorithm: f 0 (0) = ; f k (0) = 0, for all k > 0 f k (i) = e k (x i ) Σ r f r (i-) a rk P(x) = Σ k f k (N) The Forward Algorithm If our model has an end state: f 0 (0) = ; f k (0) = 0, for all k > 0 f k (i) = e k (x i ) Σ r f r (i-) a rk P(x) = Σ k f k (N) a k0 Where, a k0 is the probability that the terminating state is k (usually = a 0k ) Relation between Forward and Viterbi The most probable state VITERBI V 0 (0) = V k (0) = 0, for all k > 0 FORWARD f 0 (0) = f k (0) = 0, for all k > 0 Given a sequence x, what is the most likely state that emitted x i? In other words, we want to compute P(π i = k x) Example: the dishonest casino Say x = V j (i) = e j (x i ) max k V k (i-) a kj P(x, π*) = max k V k (N) f l (i) = e l (x i ) Σ k f k (i-) a kl P(x) = Σ k f k (N) Most likely path: π = FFF However: marked letters more likely to be L 6
7 Motivation for the Backward Algorithm The Backward Algorithm derivation We want to compute P(π i = k x), Define the backward probability: We start by computing P(π i = k, x) = P(x x i, π i = k, x i+ x N ) = P(x x i, π i = k) P(x i+ x N x x i, π i = k) = P(x x i, π i = k) P(x i+ x N π i = k) Then, P(π i = k x) = P(π i = k, x) / P(x) = f k (i) b k (i) / P(x) b k (i) = P(x i+ x N π i = k) = Σ πi+πn P(x i+,x i+,, x N, π i+,, π N π i = k) = Σ r Σ πi+πn P(x i+,x i+,, x N, π i+ = r, π i+,, π N π i = k) = Σ r e r (x i+ ) a kr Σ πi+πn P(x i+,, x N, π i+,, π N π i+ = r) = Σ r e r (x i+ ) a kr b r (i+) The Backward Algorithm A dynamic programming algorithm for b k (i): b k (N) =, for all k b k (i) = Σ r e r (x i+ ) a kr b r (i+) P(x) = Σ r a 0r e r (x ) b r () The Backward Algorithm In case of an end state: b k (N) = a k0, for all k b k (i) = Σ r e r (x i+ ) a kr b r (i+) P(x) = Σ r a 0r e r (x ) b r () Computational Complexity What is the running time, and space required for Forward, and Backward algorithms? Time: O( N) Space: O(N) Useful implementation technique to avoid underflows - Rescaling at each position by multiplying by a constant Define: Recursion: Scaling Choosing scaling factors such that Leads to: Scaling for the backward probabilities: 7
8 Posterior Decoding We can now calculate f k (i) b k (i) P(π i = k x) = P(x) Then, we can ask What is the most likely state at position i of sequence x? Using Posterior Decoding we can now define: = argmax k P(π i = k x) For each state, Posterior Decoding Posterior Decoding gives us a curve of probability of state for each position, given the sequence x That is sometimes more informative than Viterbi path π * Posterior Decoding may give an invalid sequence of states Why? Viterbi vs. posterior decoding Viterbi, Forward, Backward A class takes a multiple choice test. How does the lazy professor construct the answer key? Viterbi approach: use the answers of the best student Posterior decoding: majority vote VITERBI V 0 (0) = V k (0) = 0, for all k > 0 V r (i) = e r (x i ) max k V k (i-) a kr FORWARD f 0 (0) = f k (0) = 0, for all k > 0 f r (i) = e r (x i ) Σ k f k (i-) a kr BACWARD b k (N) = a k0, for all k b r (i) = Σ k e r (x i +) a kr b k (i+) P(x, π*) = max k V k (N) P(x) = Σ k f k (N) a k0 P(x) = Σ k a 0k e k (x ) b k () Methylation & Silencing A modeling Example CpG islands in DNA sequences Methylation Addition of CH 3 in C- nucleotides Silences genes in region CG (denoted CpG) often mutates to TG, when methylated Methylation is inherited during cell division 8
9 CpG Islands CpG Islands CpG nucleotides in the genome are frequently methylated C methyl-c T Methylation often suppressed around genes, promoters CpG islands In CpG islands: CG is more frequent Other dinucleotides (AA, AG, AT) have different frequencies Problem: Detect CpG islands A model of CpG Islands Architecture A model of CpG Islands Transitions How do we estimate parameters of the model? Emission probabilities: /0. Transition probabilities within CpG islands + A C G T A C Established from known CpG islands G T Transition probabilities within non-cpg islands - A C G T A Established from non-cpg islands C G T p A model of CpG Islands Transitions What about transitions between (+) and (-) states? Their probabilities affect Avg. length of CpG island Avg. separation between two CpG islands -p q Length distribution of X region: P[L X = ] = -p P[L X = ] = p(-p) P[L X = k] = p k- (-p) A model of CpG Islands Transitions No reason to favor exiting/entering (+) and (-) regions at a particular nucleotide Estimate average length L CPG of a CpG island: L CPG = /(-p) p = /L CPG For each pair of (+) states: a kr p a kr + For each (+) state k, (-) state r: a kr =(-p)(a 0r- ) Do the same for (-) states A problem with this model: CpG islands don t have a geometric length distribution p -q -q E[L X ] = /(-p) Geometric distribution, with mean /(-p) This is a defect of HMMs a price we pay for ease of analysis & efficient computation 9
10 Using the model Posterior decoding Given a DNA sequence x, The Viterbi algorithm predicts locations of CpG islands Given a nucleotide x i, (say x i = A) The Viterbi parse tells whether x i is in a CpG island in the most likely parse Results of applying posterior decoding to a part of human chromosome Using the Forward/Backward algorithms we can calculate P(x i is in CpG island) = P(π i = A + x) Posterior Decoding can assign locally optimal predictions of CpG islands = argmax k P(π i = k x) Posterior decoding Viterbi decoding Sliding window (size = 00) Sliding widow (size = 00) 0
11 Sliding window (size = 300) Sliding window (size = 400) Sliding window (size = 600) Sliding window (size = 000) Modeling CpG islands with silent states What if a new genome comes? Suppose we just sequenced the porcupine genome We know CpG islands play the same role in this genome However, we have no known CpG islands for porcupines We suspect the frequency and characteristics of CpG islands are quite different in porcupines How do we adjust the parameters in our model? LEARNING
12 Two learning scenarios Estimation when the right answer is known Examples: GIVEN: a genomic region x = x x N where we have good (experimental) annotations of the CpG islands GIVEN: the casino player allows us to observe him one evening as he changes the dice and produces 0,000 rolls Estimation when the right answer is unknown Examples: GIVEN: the porcupine genome; we don t know how frequent are the CpG islands there, neither do we know their composition GIVEN: 0,000 rolls of the casino player, but we don t see when he changes dice GOAL: Update the parameters θ of the model to maximize P(x θ) When the right answer is known Given x = x x N for which π = π π N is known, Define: A kr = # of times k r transition occurs in π E k (b) = # of times state k in π emits b in x The maximum likelihood estimates of the parameters are: A kr E k (b) a kr = e k (b) = Σ i A ki Σ c E k (c) When the right answer is known Intuition: When we know the underlying states, the best estimate is the average frequency of transitions & emissions that occur in the training data Drawback: Given little data, there may be overfitting: P(x θ) is maximized, but θ is unreasonable 0 probabilities VERY BAD Example: Suppose we observe: x = π = F F F F F F F F F F F F F F F L L L L L Then: a FF = 4/5 a FL = /5 a LL = a LF = 0 e F (4) = 0; e L (4) = 0 Pseudocounts Solution for small training sets: Add pseudocounts A kr E k (b) = # times k r state transition occurs + t kr = # times state k emits symbol b in x + t k (b) t kr, t k (b) are pseudocounts representing our prior belief Larger pseudocounts Strong prior belief Small pseudocounts (ε < ): just to avoid 0 probabilities Pseudocounts Pseudocounts A kr + t kr E k (b) + t k (b) a kr = e k (b) = Σ i A ki + t kr Σ c E k (c) + t k (b) Example: dishonest casino We will observe player for one day, 600 rolls Reasonable pseudocounts: t 0F = t 0L = t F0 = t L0 = ; t FL = t LF = t FF = t LL = ; t F () = t F () = = t F (6) = 0 (strong belief fair is fair) t L () = t L () = = t L (6) = 5 (wait and see for loaded) Above numbers arbitrary assigning priors is an art
13 When the right answer is unknown We don t know the actual A kr, E k (b) Idea: Initialize the model Compute A kr, E k (b) Update the parameters of the model, based on A kr, E k (b) Repeat until convergence Two algorithms: Baum-Welch, Viterbi training When the right answer is unknown Given x = x x N for which the true π = π π N is unknown, EXPECTATION MAXIMIZATION (EM). Pick model parameters θ. Estimate A kr, E k (b) (Expectation) 3. Update θ according to A kr, E k (b) (Maximization) 4. Repeat & 3, until convergence To estimate A kr : Estimating new parameters At each position of sequence x, find probability that transition k r is used: P(π i = k, π i+ = r, x x N ) Q P(π i = k, π i+ = r x) = = P(x) P(x) Q = P(x x i, π i = k, π i+ = r, x i+ x N ) = P(π i+ = r, x i+ x N π i = k) P(x x i, π i = k) = P(π i+ = r, x i+ x i+ x N π i = k) f k (i) = P(x i+ x N π i+ = r) P(x i+ π i+ = r) P(π i+ = r π i = k) f k (i) = b r (i+) e r (x i+ ) a kr f k (i) f k (i) a kr e r (x i+ ) b r (i+) So: P(π i = k, π i+ = r x, θ) = P(x θ) Estimating new parameters f k (i) a kr e r (x i+ ) b r (i+) P(π i = k, π i+ = r x, θ) = P(x θ) f k (i) x x i- k x i a kr e r (x i+ ) Summing over all positions gives the expected number of times the transition is used: f k (i) a kr e r (x i+ ) b r (i+) A kr = Σ i P(π i = k, π i+ = r x, θ) = Σ i P(x θ) r x i+ b r (i+) x i+ x N Emission probabilities: Estimating new parameters Σ {i xi = b} f k(i) b k (i) E k (b) = P(x θ) When you have multiple sequences: sum over them. The Baum-Welch Algorithm Pick the best-guess for model parameters (or random). Forward. Backward 3. Calculate A kr, E k (b) + pseudocounts 4. Calculate new model parameters a kr, e k (b) 5. Calculate new log-likelihood P(x θ) Likelihood guaranteed to increase (EM) Repeat until P(x θ) does not change much 3
14 Time Complexity: The Baum-Welch Algorithm # iterations O( N) Guaranteed to increase the log likelihood P(x θ) Not guaranteed to find globally best parameters: Converges to local optimum Too many parameters / too large model: Overfitting Alternative: Viterbi Training Same. Perform Viterbi, to find π *. Calculate A kr, E k (b) according to π * + pseudocounts 3. Calculate the new parameters a kr, e k (b) Until convergence Comments: Convergence is guaranteed Why? Does not maximize P(x θ) Maximizes P(x θ, π * ) In general, worse performance than Baum-Welch Higher-order HMMs How do we model memory of more than one time step? HMM variants First order HMM: P(π i+ = r π i = k) (a kr ) nd order HMM: P(π i+ = r π i = k, π i - = j) (a jkr ) A second order HMM with states is equivalent to a first order HMM with states a HT (prev = H) a HT (prev = T) state HH a HHT a HTH state HT state H a TH (prev = H) a TH (prev = T) state T a THH state TH a THT a TTH a HTT state TT Note that not all transitions are allowed! Modeling the Duration of States Solution : Chain several states Length distribution of region X: E[L X ] = /(-p) Geometric distribution, with mean /(-p) This is a significant disadvantage of HMMs Several solutions exist for modeling different length distributions Disadvantage: Still very inflexible L X = C + geometric with mean /(-p) 4
15 Solution : Negative binomial distribution Example: genes in prokaryotes n copies of state X: EasyGene: a prokaryotic gene-finder (Larsen TS, rogh A) Suppose the duration in X is m, where During first m turns, exactly n arrows to next state are followed In the m th turn, an arrow to next state is followed m P(L X = m) = n ( p) n p m-n Codons are modeled with 3 looped triplets of states (negative binomial with n = 3) Solution 3: Duration modeling Upon entering a state:. Choose duration d, according to probability distribution. Generate d letters according to emission probs 3. Take a transition to next state according to transition probs Disadvantage: Increase in complexity: Time: O(D ) Space: O(D) where D = maximum duration of state 5
Hidden Markov Models. x 1 x 2 x 3 x K
 Hidden Markov Models 1 1 1 1 2 2 2 2 K K K K x 1 x 2 x 3 x K HiSeq X & NextSeq Viterbi, Forward, Backward VITERBI FORWARD BACKWARD Initialization: V 0 (0) = 1 V k (0) = 0, for all k > 0 Initialization:
Hidden Markov Models 1 1 1 1 2 2 2 2 K K K K x 1 x 2 x 3 x K HiSeq X & NextSeq Viterbi, Forward, Backward VITERBI FORWARD BACKWARD Initialization: V 0 (0) = 1 V k (0) = 0, for all k > 0 Initialization:
Hidden Markov Models. x 1 x 2 x 3 x K
 Hidden Markov Models 1 1 1 1 2 2 2 2 K K K K x 1 x 2 x 3 x K Viterbi, Forward, Backward VITERBI FORWARD BACKWARD Initialization: V 0 (0) = 1 V k (0) = 0, for all k > 0 Initialization: f 0 (0) = 1 f k (0)
Hidden Markov Models 1 1 1 1 2 2 2 2 K K K K x 1 x 2 x 3 x K Viterbi, Forward, Backward VITERBI FORWARD BACKWARD Initialization: V 0 (0) = 1 V k (0) = 0, for all k > 0 Initialization: f 0 (0) = 1 f k (0)
Hidden Markov Models and Applica2ons. Spring 2017 February 21,23, 2017
 Hidden Markov Models and Applica2ons Spring 2017 February 21,23, 2017 Gene finding in prokaryotes Reading frames A protein is coded by groups of three nucleo2des (codons): ACGTACGTACGTACGT ACG-TAC-GTA-CGT-ACG-T
Hidden Markov Models and Applica2ons Spring 2017 February 21,23, 2017 Gene finding in prokaryotes Reading frames A protein is coded by groups of three nucleo2des (codons): ACGTACGTACGTACGT ACG-TAC-GTA-CGT-ACG-T
Hidden Markov Models for biological sequence analysis
 Hidden Markov Models for biological sequence analysis Master in Bioinformatics UPF 2017-2018 http://comprna.upf.edu/courses/master_agb/ Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA
Hidden Markov Models for biological sequence analysis Master in Bioinformatics UPF 2017-2018 http://comprna.upf.edu/courses/master_agb/ Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA
CSCE 471/871 Lecture 3: Markov Chains and
 and and 1 / 26 sscott@cse.unl.edu 2 / 26 Outline and chains models (s) Formal definition Finding most probable state path (Viterbi algorithm) Forward and backward algorithms State sequence known State
and and 1 / 26 sscott@cse.unl.edu 2 / 26 Outline and chains models (s) Formal definition Finding most probable state path (Viterbi algorithm) Forward and backward algorithms State sequence known State
HIDDEN MARKOV MODELS
 HIDDEN MARKOV MODELS Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm Forward-Backward Algorithm Profile HMMs HMM Parameter Estimation Viterbi training Baum-Welch algorithm
HIDDEN MARKOV MODELS Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm Forward-Backward Algorithm Profile HMMs HMM Parameter Estimation Viterbi training Baum-Welch algorithm
An Introduction to Bioinformatics Algorithms Hidden Markov Models
 Hidden Markov Models Hidden Markov Models Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm Forward-Backward Algorithm Profile HMMs HMM Parameter Estimation Viterbi training
Hidden Markov Models Hidden Markov Models Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm Forward-Backward Algorithm Profile HMMs HMM Parameter Estimation Viterbi training
Hidden Markov Models. Ivan Gesteira Costa Filho IZKF Research Group Bioinformatics RWTH Aachen Adapted from:
 Hidden Markov Models Ivan Gesteira Costa Filho IZKF Research Group Bioinformatics RWTH Aachen Adapted from: www.ioalgorithms.info Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm
Hidden Markov Models Ivan Gesteira Costa Filho IZKF Research Group Bioinformatics RWTH Aachen Adapted from: www.ioalgorithms.info Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm
An Introduction to Bioinformatics Algorithms Hidden Markov Models
 Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
CISC 889 Bioinformatics (Spring 2004) Hidden Markov Models (II)
 CISC 889 Bioinformatics (Spring 24) Hidden Markov Models (II) a. Likelihood: forward algorithm b. Decoding: Viterbi algorithm c. Model building: Baum-Welch algorithm Viterbi training Hidden Markov models
CISC 889 Bioinformatics (Spring 24) Hidden Markov Models (II) a. Likelihood: forward algorithm b. Decoding: Viterbi algorithm c. Model building: Baum-Welch algorithm Viterbi training Hidden Markov models
Hidden Markov Models
 Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Stephen Scott.
 1 / 27 sscott@cse.unl.edu 2 / 27 Useful for modeling/making predictions on sequential data E.g., biological sequences, text, series of sounds/spoken words Will return to graphical models that are generative
1 / 27 sscott@cse.unl.edu 2 / 27 Useful for modeling/making predictions on sequential data E.g., biological sequences, text, series of sounds/spoken words Will return to graphical models that are generative
Hidden Markov Models. Three classic HMM problems
 An Introduction to Bioinformatics Algorithms www.bioalgorithms.info Hidden Markov Models Slides revised and adapted to Computational Biology IST 2015/2016 Ana Teresa Freitas Three classic HMM problems
An Introduction to Bioinformatics Algorithms www.bioalgorithms.info Hidden Markov Models Slides revised and adapted to Computational Biology IST 2015/2016 Ana Teresa Freitas Three classic HMM problems
6.047/6.878/HST.507 Computational Biology: Genomes, Networks, Evolution. Lecture 05. Hidden Markov Models Part II
 6.047/6.878/HST.507 Computational Biology: Genomes, Networks, Evolution Lecture 05 Hidden Markov Models Part II 1 2 Module 1: Aligning and modeling genomes Module 1: Computational foundations Dynamic programming:
6.047/6.878/HST.507 Computational Biology: Genomes, Networks, Evolution Lecture 05 Hidden Markov Models Part II 1 2 Module 1: Aligning and modeling genomes Module 1: Computational foundations Dynamic programming:
Hidden Markov Models
 Hidden Markov Models Slides revised and adapted to Bioinformática 55 Engª Biomédica/IST 2005 Ana Teresa Freitas Forward Algorithm For Markov chains we calculate the probability of a sequence, P(x) How
Hidden Markov Models Slides revised and adapted to Bioinformática 55 Engª Biomédica/IST 2005 Ana Teresa Freitas Forward Algorithm For Markov chains we calculate the probability of a sequence, P(x) How
Hidden Markov Models
 Hidden Markov Models Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm Forward-Backward Algorithm Profile HMMs HMM Parameter Estimation Viterbi training Baum-Welch algorithm
Hidden Markov Models Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm Forward-Backward Algorithm Profile HMMs HMM Parameter Estimation Viterbi training Baum-Welch algorithm
Hidden Markov Models. x 1 x 2 x 3 x N
 Hidden Markov Models 1 1 1 1 K K K K x 1 x x 3 x N Example: The dishonest casino A casino has two dice: Fair die P(1) = P() = P(3) = P(4) = P(5) = P(6) = 1/6 Loaded die P(1) = P() = P(3) = P(4) = P(5)
Hidden Markov Models 1 1 1 1 K K K K x 1 x x 3 x N Example: The dishonest casino A casino has two dice: Fair die P(1) = P() = P(3) = P(4) = P(5) = P(6) = 1/6 Loaded die P(1) = P() = P(3) = P(4) = P(5)
Hidden Markov Models for biological sequence analysis I
 Hidden Markov Models for biological sequence analysis I Master in Bioinformatics UPF 2014-2015 Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA Barcelona, Spain Example: CpG Islands
Hidden Markov Models for biological sequence analysis I Master in Bioinformatics UPF 2014-2015 Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA Barcelona, Spain Example: CpG Islands
Hidden Markov Models. By Parisa Abedi. Slides courtesy: Eric Xing
 Hidden Markov Models By Parisa Abedi Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed data Sequential (non i.i.d.) data Time-series data E.g. Speech
Hidden Markov Models By Parisa Abedi Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed data Sequential (non i.i.d.) data Time-series data E.g. Speech
CSCE 478/878 Lecture 9: Hidden. Markov. Models. Stephen Scott. Introduction. Outline. Markov. Chains. Hidden Markov Models. CSCE 478/878 Lecture 9:
 Useful for modeling/making predictions on sequential data E.g., biological sequences, text, series of sounds/spoken words Will return to graphical models that are generative sscott@cse.unl.edu 1 / 27 2
Useful for modeling/making predictions on sequential data E.g., biological sequences, text, series of sounds/spoken words Will return to graphical models that are generative sscott@cse.unl.edu 1 / 27 2
Hidden Markov Models
 Hidden Markov Models Slides revised and adapted to Bioinformática 55 Engª Biomédica/IST 2005 Ana Teresa Freitas CG-Islands Given 4 nucleotides: probability of occurrence is ~ 1/4. Thus, probability of
Hidden Markov Models Slides revised and adapted to Bioinformática 55 Engª Biomédica/IST 2005 Ana Teresa Freitas CG-Islands Given 4 nucleotides: probability of occurrence is ~ 1/4. Thus, probability of
Introduction to Machine Learning CMU-10701
 Introduction to Machine Learning CMU-10701 Hidden Markov Models Barnabás Póczos & Aarti Singh Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed
Introduction to Machine Learning CMU-10701 Hidden Markov Models Barnabás Póczos & Aarti Singh Slides courtesy: Eric Xing i.i.d to sequential data So far we assumed independent, identically distributed
Hidden Markov Models. Aarti Singh Slides courtesy: Eric Xing. Machine Learning / Nov 8, 2010
 Hidden Markov Models Aarti Singh Slides courtesy: Eric Xing Machine Learning 10-701/15-781 Nov 8, 2010 i.i.d to sequential data So far we assumed independent, identically distributed data Sequential data
Hidden Markov Models Aarti Singh Slides courtesy: Eric Xing Machine Learning 10-701/15-781 Nov 8, 2010 i.i.d to sequential data So far we assumed independent, identically distributed data Sequential data
Markov Chains and Hidden Markov Models. COMP 571 Luay Nakhleh, Rice University
 Markov Chains and Hidden Markov Models COMP 571 Luay Nakhleh, Rice University Markov Chains and Hidden Markov Models Modeling the statistical properties of biological sequences and distinguishing regions
Markov Chains and Hidden Markov Models COMP 571 Luay Nakhleh, Rice University Markov Chains and Hidden Markov Models Modeling the statistical properties of biological sequences and distinguishing regions
Hidden Markov Models. Main source: Durbin et al., Biological Sequence Alignment (Cambridge, 98)
 Hidden Markov Models Main source: Durbin et al., Biological Sequence Alignment (Cambridge, 98) 1 The occasionally dishonest casino A P A (1) = P A (2) = = 1/6 P A->B = P B->A = 1/10 B P B (1)=0.1... P
Hidden Markov Models Main source: Durbin et al., Biological Sequence Alignment (Cambridge, 98) 1 The occasionally dishonest casino A P A (1) = P A (2) = = 1/6 P A->B = P B->A = 1/10 B P B (1)=0.1... P
Hidden Markov Models (I)
 GLOBEX Bioinformatics (Summer 2015) Hidden Markov Models (I) a. The model b. The decoding: Viterbi algorithm Hidden Markov models A Markov chain of states At each state, there are a set of possible observables
GLOBEX Bioinformatics (Summer 2015) Hidden Markov Models (I) a. The model b. The decoding: Viterbi algorithm Hidden Markov models A Markov chain of states At each state, there are a set of possible observables
Lecture 9. Intro to Hidden Markov Models (finish up)
 Lecture 9 Intro to Hidden Markov Models (finish up) Review Structure Number of states Q 1.. Q N M output symbols Parameters: Transition probability matrix a ij Emission probabilities b i (a), which is
Lecture 9 Intro to Hidden Markov Models (finish up) Review Structure Number of states Q 1.. Q N M output symbols Parameters: Transition probability matrix a ij Emission probabilities b i (a), which is
Hidden Markov Models 1
 Hidden Markov Models Dinucleotide Frequency Consider all 2-mers in a sequence {AA,AC,AG,AT,CA,CC,CG,CT,GA,GC,GG,GT,TA,TC,TG,TT} Given 4 nucleotides: each with a probability of occurrence of. 4 Thus, one
Hidden Markov Models Dinucleotide Frequency Consider all 2-mers in a sequence {AA,AC,AG,AT,CA,CC,CG,CT,GA,GC,GG,GT,TA,TC,TG,TT} Given 4 nucleotides: each with a probability of occurrence of. 4 Thus, one
Hidden Markov Models. Ron Shamir, CG 08
 Hidden Markov Models 1 Dr Richard Durbin is a graduate in mathematics from Cambridge University and one of the founder members of the Sanger Institute. He has also held carried out research at the Laboratory
Hidden Markov Models 1 Dr Richard Durbin is a graduate in mathematics from Cambridge University and one of the founder members of the Sanger Institute. He has also held carried out research at the Laboratory
Hidden Markov Models. based on chapters from the book Durbin, Eddy, Krogh and Mitchison Biological Sequence Analysis via Shamir s lecture notes
 Hidden Markov Models based on chapters from the book Durbin, Eddy, Krogh and Mitchison Biological Sequence Analysis via Shamir s lecture notes music recognition deal with variations in - actual sound -
Hidden Markov Models based on chapters from the book Durbin, Eddy, Krogh and Mitchison Biological Sequence Analysis via Shamir s lecture notes music recognition deal with variations in - actual sound -
11.3 Decoding Algorithm
 11.3 Decoding Algorithm 393 For convenience, we have introduced π 0 and π n+1 as the fictitious initial and terminal states begin and end. This model defines the probability P(x π) for a given sequence
11.3 Decoding Algorithm 393 For convenience, we have introduced π 0 and π n+1 as the fictitious initial and terminal states begin and end. This model defines the probability P(x π) for a given sequence
Hidden Markov Models. Hosein Mohimani GHC7717
 Hidden Markov Models Hosein Mohimani GHC7717 hoseinm@andrew.cmu.edu Fair et Casino Problem Dealer flips a coin and player bets on outcome Dealer use either a fair coin (head and tail equally likely) or
Hidden Markov Models Hosein Mohimani GHC7717 hoseinm@andrew.cmu.edu Fair et Casino Problem Dealer flips a coin and player bets on outcome Dealer use either a fair coin (head and tail equally likely) or
Introduction to Hidden Markov Models for Gene Prediction ECE-S690
 Introduction to Hidden Markov Models for Gene Prediction ECE-S690 Outline Markov Models The Hidden Part How can we use this for gene prediction? Learning Models Want to recognize patterns (e.g. sequence
Introduction to Hidden Markov Models for Gene Prediction ECE-S690 Outline Markov Models The Hidden Part How can we use this for gene prediction? Learning Models Want to recognize patterns (e.g. sequence
Advanced Data Science
 Advanced Data Science Dr. Kira Radinsky Slides Adapted from Tom M. Mitchell Agenda Topics Covered: Time series data Markov Models Hidden Markov Models Dynamic Bayes Nets Additional Reading: Bishop: Chapter
Advanced Data Science Dr. Kira Radinsky Slides Adapted from Tom M. Mitchell Agenda Topics Covered: Time series data Markov Models Hidden Markov Models Dynamic Bayes Nets Additional Reading: Bishop: Chapter
Markov Chains and Hidden Markov Models. = stochastic, generative models
 Markov Chains and Hidden Markov Models = stochastic, generative models (Drawing heavily from Durbin et al., Biological Sequence Analysis) BCH339N Systems Biology / Bioinformatics Spring 2016 Edward Marcotte,
Markov Chains and Hidden Markov Models = stochastic, generative models (Drawing heavily from Durbin et al., Biological Sequence Analysis) BCH339N Systems Biology / Bioinformatics Spring 2016 Edward Marcotte,
O 3 O 4 O 5. q 3. q 4. Transition
 Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Hidden Markov Models
 s Ben Langmead Department of Computer Science Please sign guestbook (www.langmead-lab.org/teaching-materials) to tell me briefly how you are using the slides. For original Keynote files, email me (ben.langmead@gmail.com).
s Ben Langmead Department of Computer Science Please sign guestbook (www.langmead-lab.org/teaching-materials) to tell me briefly how you are using the slides. For original Keynote files, email me (ben.langmead@gmail.com).
6 Markov Chains and Hidden Markov Models
 6 Markov Chains and Hidden Markov Models (This chapter 1 is primarily based on Durbin et al., chapter 3, [DEKM98] and the overview article by Rabiner [Rab89] on HMMs.) Why probabilistic models? In problems
6 Markov Chains and Hidden Markov Models (This chapter 1 is primarily based on Durbin et al., chapter 3, [DEKM98] and the overview article by Rabiner [Rab89] on HMMs.) Why probabilistic models? In problems
1/22/13. Example: CpG Island. Question 2: Finding CpG Islands
 I529: Machine Learning in Bioinformatics (Spring 203 Hidden Markov Models Yuzhen Ye School of Informatics and Computing Indiana Univerty, Bloomington Spring 203 Outline Review of Markov chain & CpG island
I529: Machine Learning in Bioinformatics (Spring 203 Hidden Markov Models Yuzhen Ye School of Informatics and Computing Indiana Univerty, Bloomington Spring 203 Outline Review of Markov chain & CpG island
Basic math for biology
 Basic math for biology Lei Li Florida State University, Feb 6, 2002 The EM algorithm: setup Parametric models: {P θ }. Data: full data (Y, X); partial data Y. Missing data: X. Likelihood and maximum likelihood
Basic math for biology Lei Li Florida State University, Feb 6, 2002 The EM algorithm: setup Parametric models: {P θ }. Data: full data (Y, X); partial data Y. Missing data: X. Likelihood and maximum likelihood
CS711008Z Algorithm Design and Analysis
 .. Lecture 6. Hidden Markov model and Viterbi s decoding algorithm Institute of Computing Technology Chinese Academy of Sciences, Beijing, China . Outline The occasionally dishonest casino: an example
.. Lecture 6. Hidden Markov model and Viterbi s decoding algorithm Institute of Computing Technology Chinese Academy of Sciences, Beijing, China . Outline The occasionally dishonest casino: an example
ROBI POLIKAR. ECE 402/504 Lecture Hidden Markov Models IGNAL PROCESSING & PATTERN RECOGNITION ROWAN UNIVERSITY
 BIOINFORMATICS Lecture 11-12 Hidden Markov Models ROBI POLIKAR 2011, All Rights Reserved, Robi Polikar. IGNAL PROCESSING & PATTERN RECOGNITION LABORATORY @ ROWAN UNIVERSITY These lecture notes are prepared
BIOINFORMATICS Lecture 11-12 Hidden Markov Models ROBI POLIKAR 2011, All Rights Reserved, Robi Polikar. IGNAL PROCESSING & PATTERN RECOGNITION LABORATORY @ ROWAN UNIVERSITY These lecture notes are prepared
Computational Biology Lecture #3: Probability and Statistics. Bud Mishra Professor of Computer Science, Mathematics, & Cell Biology Sept
 Computational Biology Lecture #3: Probability and Statistics Bud Mishra Professor of Computer Science, Mathematics, & Cell Biology Sept 26 2005 L2-1 Basic Probabilities L2-2 1 Random Variables L2-3 Examples
Computational Biology Lecture #3: Probability and Statistics Bud Mishra Professor of Computer Science, Mathematics, & Cell Biology Sept 26 2005 L2-1 Basic Probabilities L2-2 1 Random Variables L2-3 Examples
Hidden Markov Models. music recognition. deal with variations in - pitch - timing - timbre 2
 Hidden Markov Models based on chapters from the book Durbin, Eddy, Krogh and Mitchison Biological Sequence Analysis Shamir s lecture notes and Rabiner s tutorial on HMM 1 music recognition deal with variations
Hidden Markov Models based on chapters from the book Durbin, Eddy, Krogh and Mitchison Biological Sequence Analysis Shamir s lecture notes and Rabiner s tutorial on HMM 1 music recognition deal with variations
Lecture 7 Sequence analysis. Hidden Markov Models
 Lecture 7 Sequence analysis. Hidden Markov Models Nicolas Lartillot may 2012 Nicolas Lartillot (Universite de Montréal) BIN6009 may 2012 1 / 60 1 Motivation 2 Examples of Hidden Markov models 3 Hidden
Lecture 7 Sequence analysis. Hidden Markov Models Nicolas Lartillot may 2012 Nicolas Lartillot (Universite de Montréal) BIN6009 may 2012 1 / 60 1 Motivation 2 Examples of Hidden Markov models 3 Hidden
HMMs and biological sequence analysis
 HMMs and biological sequence analysis Hidden Markov Model A Markov chain is a sequence of random variables X 1, X 2, X 3,... That has the property that the value of the current state depends only on the
HMMs and biological sequence analysis Hidden Markov Model A Markov chain is a sequence of random variables X 1, X 2, X 3,... That has the property that the value of the current state depends only on the
What s an HMM? Extraction with Finite State Machines e.g. Hidden Markov Models (HMMs) Hidden Markov Models (HMMs) for Information Extraction
 Hidden Markov Models (HMMs) for Information Extraction Daniel S. Weld CSE 454 Extraction with Finite State Machines e.g. Hidden Markov Models (HMMs) standard sequence model in genomics, speech, NLP, What
Hidden Markov Models (HMMs) for Information Extraction Daniel S. Weld CSE 454 Extraction with Finite State Machines e.g. Hidden Markov Models (HMMs) standard sequence model in genomics, speech, NLP, What
Plan for today. ! Part 1: (Hidden) Markov models. ! Part 2: String matching and read mapping
 Plan for today! Part 1: (Hidden) Markov models! Part 2: String matching and read mapping! 2.1 Exact algorithms! 2.2 Heuristic methods for approximate search (Hidden) Markov models Why consider probabilistics
Plan for today! Part 1: (Hidden) Markov models! Part 2: String matching and read mapping! 2.1 Exact algorithms! 2.2 Heuristic methods for approximate search (Hidden) Markov models Why consider probabilistics
Hidden Markov Models The three basic HMM problems (note: change in notation) Mitch Marcus CSE 391
 Hidden Markov Models The three basic HMM problems (note: change in notation) Mitch Marcus CSE 391 Parameters of an HMM States: A set of states S=s 1, s n Transition probabilities: A= a 1,1, a 1,2,, a n,n
Hidden Markov Models The three basic HMM problems (note: change in notation) Mitch Marcus CSE 391 Parameters of an HMM States: A set of states S=s 1, s n Transition probabilities: A= a 1,1, a 1,2,, a n,n
R. Durbin, S. Eddy, A. Krogh, G. Mitchison: Biological sequence analysis. Cambridge University Press, ISBN (Chapter 3)
 9 Markov chains and Hidden Markov Models We will discuss: Markov chains Hidden Markov Models (HMMs) lgorithms: Viterbi, forward, backward, posterior decoding Profile HMMs Baum-Welch algorithm This chapter
9 Markov chains and Hidden Markov Models We will discuss: Markov chains Hidden Markov Models (HMMs) lgorithms: Viterbi, forward, backward, posterior decoding Profile HMMs Baum-Welch algorithm This chapter
Today s Lecture: HMMs
 Today s Lecture: HMMs Definitions Examples Probability calculations WDAG Dynamic programming algorithms: Forward Viterbi Parameter estimation Viterbi training 1 Hidden Markov Models Probability models
Today s Lecture: HMMs Definitions Examples Probability calculations WDAG Dynamic programming algorithms: Forward Viterbi Parameter estimation Viterbi training 1 Hidden Markov Models Probability models
Lecture 4: Hidden Markov Models: An Introduction to Dynamic Decision Making. November 11, 2010
 Hidden Lecture 4: Hidden : An Introduction to Dynamic Decision Making November 11, 2010 Special Meeting 1/26 Markov Model Hidden When a dynamical system is probabilistic it may be determined by the transition
Hidden Lecture 4: Hidden : An Introduction to Dynamic Decision Making November 11, 2010 Special Meeting 1/26 Markov Model Hidden When a dynamical system is probabilistic it may be determined by the transition
Genome 373: Hidden Markov Models II. Doug Fowler
 Genome 373: Hidden Markov Models II Doug Fowler Review From Hidden Markov Models I What does a Markov model describe? Review From Hidden Markov Models I A T A Markov model describes a random process of
Genome 373: Hidden Markov Models II Doug Fowler Review From Hidden Markov Models I What does a Markov model describe? Review From Hidden Markov Models I A T A Markov model describes a random process of
Pairwise alignment using HMMs
 Pairwise alignment using HMMs The states of an HMM fulfill the Markov property: probability of transition depends only on the last state. CpG islands and casino example: HMMs emit sequence of symbols (nucleotides
Pairwise alignment using HMMs The states of an HMM fulfill the Markov property: probability of transition depends only on the last state. CpG islands and casino example: HMMs emit sequence of symbols (nucleotides
Soundex distance metric
 Text Algorithms (4AP) Lecture: Time warping and sound Jaak Vilo 008 fall Jaak Vilo MTAT.03.90 Text Algorithms Soundex distance metric Soundex is a coarse phonetic indexing scheme, widely used in genealogy.
Text Algorithms (4AP) Lecture: Time warping and sound Jaak Vilo 008 fall Jaak Vilo MTAT.03.90 Text Algorithms Soundex distance metric Soundex is a coarse phonetic indexing scheme, widely used in genealogy.
Markov chains and Hidden Markov Models
 Discrete Math for Bioinformatics WS 10/11:, b A. Bockmar/K. Reinert, 7. November 2011, 10:24 2001 Markov chains and Hidden Markov Models We will discuss: Hidden Markov Models (HMMs) Algorithms: Viterbi,
Discrete Math for Bioinformatics WS 10/11:, b A. Bockmar/K. Reinert, 7. November 2011, 10:24 2001 Markov chains and Hidden Markov Models We will discuss: Hidden Markov Models (HMMs) Algorithms: Viterbi,
Multiple Sequence Alignment using Profile HMM
 Multiple Sequence Alignment using Profile HMM. based on Chapter 5 and Section 6.5 from Biological Sequence Analysis by R. Durbin et al., 1998 Acknowledgements: M.Sc. students Beatrice Miron, Oana Răţoi,
Multiple Sequence Alignment using Profile HMM. based on Chapter 5 and Section 6.5 from Biological Sequence Analysis by R. Durbin et al., 1998 Acknowledgements: M.Sc. students Beatrice Miron, Oana Răţoi,
Pair Hidden Markov Models
 Pair Hidden Markov Models Scribe: Rishi Bedi Lecturer: Serafim Batzoglou January 29, 2015 1 Recap of HMMs alphabet: Σ = {b 1,...b M } set of states: Q = {1,..., K} transition probabilities: A = [a ij ]
Pair Hidden Markov Models Scribe: Rishi Bedi Lecturer: Serafim Batzoglou January 29, 2015 1 Recap of HMMs alphabet: Σ = {b 1,...b M } set of states: Q = {1,..., K} transition probabilities: A = [a ij ]
Bioinformatics: Biology X
 Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA Model Building/Checking, Reverse Engineering, Causality Outline 1 Where (or of what) one cannot speak, one must pass over in silence.
Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA Model Building/Checking, Reverse Engineering, Causality Outline 1 Where (or of what) one cannot speak, one must pass over in silence.
Lecture 5: December 13, 2001
 Algorithms for Molecular Biology Fall Semester, 2001 Lecture 5: December 13, 2001 Lecturer: Ron Shamir Scribe: Roi Yehoshua and Oren Danewitz 1 5.1 Hidden Markov Models 5.1.1 Preface: CpG islands CpG is
Algorithms for Molecular Biology Fall Semester, 2001 Lecture 5: December 13, 2001 Lecturer: Ron Shamir Scribe: Roi Yehoshua and Oren Danewitz 1 5.1 Hidden Markov Models 5.1.1 Preface: CpG islands CpG is
MACHINE LEARNING 2 UGM,HMMS Lecture 7
 LOREM I P S U M Royal Institute of Technology MACHINE LEARNING 2 UGM,HMMS Lecture 7 THIS LECTURE DGM semantics UGM De-noising HMMs Applications (interesting probabilities) DP for generation probability
LOREM I P S U M Royal Institute of Technology MACHINE LEARNING 2 UGM,HMMS Lecture 7 THIS LECTURE DGM semantics UGM De-noising HMMs Applications (interesting probabilities) DP for generation probability
HMM: Parameter Estimation
 I529: Machine Learning in Bioinformatics (Spring 2017) HMM: Parameter Estimation Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Content Review HMM: three problems
I529: Machine Learning in Bioinformatics (Spring 2017) HMM: Parameter Estimation Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Content Review HMM: three problems
Computational Genomics and Molecular Biology, Fall
 Computational Genomics and Molecular Biology, Fall 2011 1 HMM Lecture Notes Dannie Durand and Rose Hoberman October 11th 1 Hidden Markov Models In the last few lectures, we have focussed on three problems
Computational Genomics and Molecular Biology, Fall 2011 1 HMM Lecture Notes Dannie Durand and Rose Hoberman October 11th 1 Hidden Markov Models In the last few lectures, we have focussed on three problems
Hidden Markov Model. Ying Wu. Electrical Engineering and Computer Science Northwestern University Evanston, IL 60208
 Hidden Markov Model Ying Wu Electrical Engineering and Computer Science Northwestern University Evanston, IL 60208 http://www.eecs.northwestern.edu/~yingwu 1/19 Outline Example: Hidden Coin Tossing Hidden
Hidden Markov Model Ying Wu Electrical Engineering and Computer Science Northwestern University Evanston, IL 60208 http://www.eecs.northwestern.edu/~yingwu 1/19 Outline Example: Hidden Coin Tossing Hidden
Grundlagen der Bioinformatik, SS 09, D. Huson, June 16, S. Durbin, S. Eddy, A. Krogh and G. Mitchison, Biological Sequence
 rundlagen der Bioinformatik, SS 09,. Huson, June 16, 2009 81 7 Markov chains and Hidden Markov Models We will discuss: Markov chains Hidden Markov Models (HMMs) Profile HMMs his chapter is based on: nalysis,
rundlagen der Bioinformatik, SS 09,. Huson, June 16, 2009 81 7 Markov chains and Hidden Markov Models We will discuss: Markov chains Hidden Markov Models (HMMs) Profile HMMs his chapter is based on: nalysis,
Hidden Markov Models
 Andrea Passerini passerini@disi.unitn.it Statistical relational learning The aim Modeling temporal sequences Model signals which vary over time (e.g. speech) Two alternatives: deterministic models directly
Andrea Passerini passerini@disi.unitn.it Statistical relational learning The aim Modeling temporal sequences Model signals which vary over time (e.g. speech) Two alternatives: deterministic models directly
COMS 4771 Probabilistic Reasoning via Graphical Models. Nakul Verma
 COMS 4771 Probabilistic Reasoning via Graphical Models Nakul Verma Last time Dimensionality Reduction Linear vs non-linear Dimensionality Reduction Principal Component Analysis (PCA) Non-linear methods
COMS 4771 Probabilistic Reasoning via Graphical Models Nakul Verma Last time Dimensionality Reduction Linear vs non-linear Dimensionality Reduction Principal Component Analysis (PCA) Non-linear methods
Hidden Markov Models,99,100! Markov, here I come!
 Hidden Markov Models,99,100! Markov, here I come! 16.410/413 Principles of Autonomy and Decision-Making Pedro Santana (psantana@mit.edu) October 7 th, 2015. Based on material by Brian Williams and Emilio
Hidden Markov Models,99,100! Markov, here I come! 16.410/413 Principles of Autonomy and Decision-Making Pedro Santana (psantana@mit.edu) October 7 th, 2015. Based on material by Brian Williams and Emilio
6.864: Lecture 5 (September 22nd, 2005) The EM Algorithm
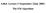 6.864: Lecture 5 (September 22nd, 2005) The EM Algorithm Overview The EM algorithm in general form The EM algorithm for hidden markov models (brute force) The EM algorithm for hidden markov models (dynamic
6.864: Lecture 5 (September 22nd, 2005) The EM Algorithm Overview The EM algorithm in general form The EM algorithm for hidden markov models (brute force) The EM algorithm for hidden markov models (dynamic
Data Mining in Bioinformatics HMM
 Data Mining in Bioinformatics HMM Microarray Problem: Major Objective n Major Objective: Discover a comprehensive theory of life s organization at the molecular level 2 1 Data Mining in Bioinformatics
Data Mining in Bioinformatics HMM Microarray Problem: Major Objective n Major Objective: Discover a comprehensive theory of life s organization at the molecular level 2 1 Data Mining in Bioinformatics
Hidden Markov Models (HMMs) November 14, 2017
 Hidden Markov Models (HMMs) November 14, 2017 inferring a hidden truth 1) You hear a static-filled radio transmission. how can you determine what did the sender intended to say? 2) You know that genes
Hidden Markov Models (HMMs) November 14, 2017 inferring a hidden truth 1) You hear a static-filled radio transmission. how can you determine what did the sender intended to say? 2) You know that genes
Grundlagen der Bioinformatik, SS 08, D. Huson, June 16, S. Durbin, S. Eddy, A. Krogh and G. Mitchison, Biological Sequence
 rundlagen der Bioinformatik, SS 08,. Huson, June 16, 2008 89 8 Markov chains and Hidden Markov Models We will discuss: Markov chains Hidden Markov Models (HMMs) Profile HMMs his chapter is based on: nalysis,
rundlagen der Bioinformatik, SS 08,. Huson, June 16, 2008 89 8 Markov chains and Hidden Markov Models We will discuss: Markov chains Hidden Markov Models (HMMs) Profile HMMs his chapter is based on: nalysis,
Info 2950, Lecture 25
 Info 2950, Lecture 25 4 May 2017 Prob Set 8: due 11 May (end of classes) 4 3.5 2.2 7.4.8 5.5 1.5 0.5 6.3 Consider the long term behavior of a Markov chain: is there some set of probabilities v i for being
Info 2950, Lecture 25 4 May 2017 Prob Set 8: due 11 May (end of classes) 4 3.5 2.2 7.4.8 5.5 1.5 0.5 6.3 Consider the long term behavior of a Markov chain: is there some set of probabilities v i for being
STA 414/2104: Machine Learning
 STA 414/2104: Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistics! rsalakhu@cs.toronto.edu! http://www.cs.toronto.edu/~rsalakhu/ Lecture 9 Sequential Data So far
STA 414/2104: Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistics! rsalakhu@cs.toronto.edu! http://www.cs.toronto.edu/~rsalakhu/ Lecture 9 Sequential Data So far
2 : Directed GMs: Bayesian Networks
 10-708: Probabilistic Graphical Models 10-708, Spring 2017 2 : Directed GMs: Bayesian Networks Lecturer: Eric P. Xing Scribes: Jayanth Koushik, Hiroaki Hayashi, Christian Perez Topic: Directed GMs 1 Types
10-708: Probabilistic Graphical Models 10-708, Spring 2017 2 : Directed GMs: Bayesian Networks Lecturer: Eric P. Xing Scribes: Jayanth Koushik, Hiroaki Hayashi, Christian Perez Topic: Directed GMs 1 Types
Chapter 4: Hidden Markov Models
 Chapter 4: Hidden Markov Models 4.1 Introduction to HMM Prof. Yechiam Yemini (YY) Computer Science Department Columbia University Overview Markov models of sequence structures Introduction to Hidden Markov
Chapter 4: Hidden Markov Models 4.1 Introduction to HMM Prof. Yechiam Yemini (YY) Computer Science Department Columbia University Overview Markov models of sequence structures Introduction to Hidden Markov
Statistical NLP: Hidden Markov Models. Updated 12/15
 Statistical NLP: Hidden Markov Models Updated 12/15 Markov Models Markov models are statistical tools that are useful for NLP because they can be used for part-of-speech-tagging applications Their first
Statistical NLP: Hidden Markov Models Updated 12/15 Markov Models Markov models are statistical tools that are useful for NLP because they can be used for part-of-speech-tagging applications Their first
Giri Narasimhan. CAP 5510: Introduction to Bioinformatics. ECS 254; Phone: x3748
 CAP 5510: Introduction to Bioinformatics Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs07.html 2/14/07 CAP5510 1 CpG Islands Regions in DNA sequences with increased
CAP 5510: Introduction to Bioinformatics Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs07.html 2/14/07 CAP5510 1 CpG Islands Regions in DNA sequences with increased
Hidden Markov Models Part 2: Algorithms
 Hidden Markov Models Part 2: Algorithms CSE 6363 Machine Learning Vassilis Athitsos Computer Science and Engineering Department University of Texas at Arlington 1 Hidden Markov Model An HMM consists of:
Hidden Markov Models Part 2: Algorithms CSE 6363 Machine Learning Vassilis Athitsos Computer Science and Engineering Department University of Texas at Arlington 1 Hidden Markov Model An HMM consists of:
CS 136a Lecture 7 Speech Recognition Architecture: Training models with the Forward backward algorithm
 + September13, 2016 Professor Meteer CS 136a Lecture 7 Speech Recognition Architecture: Training models with the Forward backward algorithm Thanks to Dan Jurafsky for these slides + ASR components n Feature
+ September13, 2016 Professor Meteer CS 136a Lecture 7 Speech Recognition Architecture: Training models with the Forward backward algorithm Thanks to Dan Jurafsky for these slides + ASR components n Feature
Page 1. References. Hidden Markov models and multiple sequence alignment. Markov chains. Probability review. Example. Markovian sequence
 Page Hidden Markov models and multiple sequence alignment Russ B Altman BMI 4 CS 74 Some slides borrowed from Scott C Schmidler (BMI graduate student) References Bioinformatics Classic: Krogh et al (994)
Page Hidden Markov models and multiple sequence alignment Russ B Altman BMI 4 CS 74 Some slides borrowed from Scott C Schmidler (BMI graduate student) References Bioinformatics Classic: Krogh et al (994)
The Computational Problem. We are given a sequence of DNA and we wish to know which subsequence or concatenation of subsequences constitutes a gene.
 GENE FINDING The Computational Problem We are given a sequence of DNA and we wish to know which subsequence or concatenation of subsequences constitutes a gene. The Computational Problem Confounding Realities:
GENE FINDING The Computational Problem We are given a sequence of DNA and we wish to know which subsequence or concatenation of subsequences constitutes a gene. The Computational Problem Confounding Realities:
Hidden Markov Methods. Algorithms and Implementation
 Hidden Markov Methods. Algorithms and Implementation Final Project Report. MATH 127. Nasser M. Abbasi Course taken during Fall 2002 page compiled on July 2, 2015 at 12:08am Contents 1 Example HMM 5 2 Forward
Hidden Markov Methods. Algorithms and Implementation Final Project Report. MATH 127. Nasser M. Abbasi Course taken during Fall 2002 page compiled on July 2, 2015 at 12:08am Contents 1 Example HMM 5 2 Forward
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 11 Project
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 11 Project
2 : Directed GMs: Bayesian Networks
 10-708: Probabilistic Graphical Models, Spring 2015 2 : Directed GMs: Bayesian Networks Lecturer: Eric P. Xing Scribes: Yi Cheng, Cong Lu 1 Notation Here the notations used in this course are defined:
10-708: Probabilistic Graphical Models, Spring 2015 2 : Directed GMs: Bayesian Networks Lecturer: Eric P. Xing Scribes: Yi Cheng, Cong Lu 1 Notation Here the notations used in this course are defined:
CS839: Probabilistic Graphical Models. Lecture 7: Learning Fully Observed BNs. Theo Rekatsinas
 CS839: Probabilistic Graphical Models Lecture 7: Learning Fully Observed BNs Theo Rekatsinas 1 Exponential family: a basic building block For a numeric random variable X p(x ) =h(x)exp T T (x) A( ) = 1
CS839: Probabilistic Graphical Models Lecture 7: Learning Fully Observed BNs Theo Rekatsinas 1 Exponential family: a basic building block For a numeric random variable X p(x ) =h(x)exp T T (x) A( ) = 1
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 07: profile Hidden Markov Model http://bibiserv.techfak.uni-bielefeld.de/sadr2/databasesearch/hmmer/profilehmm.gif Slides adapted from Dr. Shaojie Zhang
EECS730: Introduction to Bioinformatics Lecture 07: profile Hidden Markov Model http://bibiserv.techfak.uni-bielefeld.de/sadr2/databasesearch/hmmer/profilehmm.gif Slides adapted from Dr. Shaojie Zhang
Hidden Markov Models: All the Glorious Gory Details
 Hidden Markov Models: All the Glorious Gory Details Noah A. Smith Department of Computer Science Johns Hopkins University nasmith@cs.jhu.edu 18 October 2004 1 Introduction Hidden Markov models (HMMs, hereafter)
Hidden Markov Models: All the Glorious Gory Details Noah A. Smith Department of Computer Science Johns Hopkins University nasmith@cs.jhu.edu 18 October 2004 1 Introduction Hidden Markov models (HMMs, hereafter)
Comparative Gene Finding. BMI/CS 776 Spring 2015 Colin Dewey
 Comparative Gene Finding BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2015 Colin Dewey cdewey@biostat.wisc.edu Goals for Lecture the key concepts to understand are the following: using related genomes
Comparative Gene Finding BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2015 Colin Dewey cdewey@biostat.wisc.edu Goals for Lecture the key concepts to understand are the following: using related genomes
V. RANDOM VARIABLES, PROBABILITY DISTRIBUTIONS, EXPECTED VALUE
 V. RANDOM VARIABLES, PROBABILITY DISTRIBUTIONS, EXPECTED VALUE A game of chance featured at an amusement park is played as follows: You pay $ to play. A penny a nickel are flipped. You win $ if either
V. RANDOM VARIABLES, PROBABILITY DISTRIBUTIONS, EXPECTED VALUE A game of chance featured at an amusement park is played as follows: You pay $ to play. A penny a nickel are flipped. You win $ if either
Markov Models & DNA Sequence Evolution
 7.91 / 7.36 / BE.490 Lecture #5 Mar. 9, 2004 Markov Models & DNA Sequence Evolution Chris Burge Review of Markov & HMM Models for DNA Markov Models for splice sites Hidden Markov Models - looking under
7.91 / 7.36 / BE.490 Lecture #5 Mar. 9, 2004 Markov Models & DNA Sequence Evolution Chris Burge Review of Markov & HMM Models for DNA Markov Models for splice sites Hidden Markov Models - looking under
Lecture 12: Algorithms for HMMs
 Lecture 12: Algorithms for HMMs Nathan Schneider (some slides from Sharon Goldwater; thanks to Jonathan May for bug fixes) ENLP 26 February 2018 Recap: tagging POS tagging is a sequence labelling task.
Lecture 12: Algorithms for HMMs Nathan Schneider (some slides from Sharon Goldwater; thanks to Jonathan May for bug fixes) ENLP 26 February 2018 Recap: tagging POS tagging is a sequence labelling task.
Brief Introduction of Machine Learning Techniques for Content Analysis
 1 Brief Introduction of Machine Learning Techniques for Content Analysis Wei-Ta Chu 2008/11/20 Outline 2 Overview Gaussian Mixture Model (GMM) Hidden Markov Model (HMM) Support Vector Machine (SVM) Overview
1 Brief Introduction of Machine Learning Techniques for Content Analysis Wei-Ta Chu 2008/11/20 Outline 2 Overview Gaussian Mixture Model (GMM) Hidden Markov Model (HMM) Support Vector Machine (SVM) Overview
Evolutionary Models. Evolutionary Models
 Edit Operators In standard pairwise alignment, what are the allowed edit operators that transform one sequence into the other? Describe how each of these edit operations are represented on a sequence alignment
Edit Operators In standard pairwise alignment, what are the allowed edit operators that transform one sequence into the other? Describe how each of these edit operations are represented on a sequence alignment
We Live in Exciting Times. CSCI-567: Machine Learning (Spring 2019) Outline. Outline. ACM (an international computing research society) has named
 We Live in Exciting Times ACM (an international computing research society) has named CSCI-567: Machine Learning (Spring 2019) Prof. Victor Adamchik U of Southern California Apr. 2, 2019 Yoshua Bengio,
We Live in Exciting Times ACM (an international computing research society) has named CSCI-567: Machine Learning (Spring 2019) Prof. Victor Adamchik U of Southern California Apr. 2, 2019 Yoshua Bengio,
p(d θ ) l(θ ) 1.2 x x x
 p(d θ ).2 x 0-7 0.8 x 0-7 0.4 x 0-7 l(θ ) -20-40 -60-80 -00 2 3 4 5 6 7 θ ˆ 2 3 4 5 6 7 θ ˆ 2 3 4 5 6 7 θ θ x FIGURE 3.. The top graph shows several training points in one dimension, known or assumed to
p(d θ ).2 x 0-7 0.8 x 0-7 0.4 x 0-7 l(θ ) -20-40 -60-80 -00 2 3 4 5 6 7 θ ˆ 2 3 4 5 6 7 θ ˆ 2 3 4 5 6 7 θ θ x FIGURE 3.. The top graph shows several training points in one dimension, known or assumed to
Lecture 12: Algorithms for HMMs
 Lecture 12: Algorithms for HMMs Nathan Schneider (some slides from Sharon Goldwater; thanks to Jonathan May for bug fixes) ENLP 17 October 2016 updated 9 September 2017 Recap: tagging POS tagging is a
Lecture 12: Algorithms for HMMs Nathan Schneider (some slides from Sharon Goldwater; thanks to Jonathan May for bug fixes) ENLP 17 October 2016 updated 9 September 2017 Recap: tagging POS tagging is a
Math 350: An exploration of HMMs through doodles.
 Math 350: An exploration of HMMs through doodles. Joshua Little (407673) 19 December 2012 1 Background 1.1 Hidden Markov models. Markov chains (MCs) work well for modelling discrete-time processes, or
Math 350: An exploration of HMMs through doodles. Joshua Little (407673) 19 December 2012 1 Background 1.1 Hidden Markov models. Markov chains (MCs) work well for modelling discrete-time processes, or
VL Algorithmen und Datenstrukturen für Bioinformatik ( ) WS15/2016 Woche 16
 VL Algorithmen und Datenstrukturen für Bioinformatik (19400001) WS15/2016 Woche 16 Tim Conrad AG Medical Bioinformatics Institut für Mathematik & Informatik, Freie Universität Berlin Based on slides by
VL Algorithmen und Datenstrukturen für Bioinformatik (19400001) WS15/2016 Woche 16 Tim Conrad AG Medical Bioinformatics Institut für Mathematik & Informatik, Freie Universität Berlin Based on slides by
1 : Introduction. 1 Course Overview. 2 Notation. 3 Representing Multivariate Distributions : Probabilistic Graphical Models , Spring 2014
 10-708: Probabilistic Graphical Models 10-708, Spring 2014 1 : Introduction Lecturer: Eric P. Xing Scribes: Daniel Silva and Calvin McCarter 1 Course Overview In this lecture we introduce the concept of
10-708: Probabilistic Graphical Models 10-708, Spring 2014 1 : Introduction Lecturer: Eric P. Xing Scribes: Daniel Silva and Calvin McCarter 1 Course Overview In this lecture we introduce the concept of
