INPUT DESCRIPTION FOR SQM version 2.0
|
|
|
- Victor Garrison
- 6 years ago
- Views:
Transcription
1 INPUT DESCRIPTION FOR SQM version 2.0 INTRODUCTION SQM is an add-on module for the PQS program which scales force constants to produce a Scaled Quantum Mechanical (SQM) Force Field. This can correct for deficiencies in the calculated (harmonic) force constants, giving a better fit to experimentally observed vibrational fundamentals and infrared (IR) intensities. The method has considerable predictive power and is often of great help in understanding and assigning experimental vibrational spectra. The first ab initio calculations of harmonic force constants were carried out at the Hartree-Fock level of theory. Hartree-Fock theory overestimates harmonic force constants significantly and empirical scaling is needed to produce force constants which can reproduce experimental vibrational frequencies. The scaling compensates for basis set deficiencies, anharmonicity and mostly for the lack of electron correlation. The need for empirical correction diminishes but is not completely eliminated if the quality of the wavefunction improves by adding electron correlation and increasing the size of the basis. The first scaling methods applied to ab initio force constants used several different scale factors to correct for systematic errors in different types of molecular deformations, e.g., stretches, bends and torsions. This procedure requires the transformation of the molecular force field (the Hessian matrix) to chemically meaningful internal coordinates and cannot be applied directly to the calculated frequencies. It is thus less convenient than global scaling using a single scaling factor. Global scaling can be applied directly to the frequencies, the scale factor for frequencies being near 0.9, corresponding to a scale factor of 0.81 for force constants. Because of its simplicity, global scaling became popular, but using multiple scale factors yields much better results as was convincingly demonstrated by Blom and Altona in a series of papers starting in the mid 1970s [1]. Their method forms the basis of the SQM procedure which has been in widespread use for over 20 years [2]. In the original SQM procedure, the molecular geometry was expressed in terms of a full set of nonredundant natural internal coordinates [3]. Natural internals use individual bond displacements as stretching coordinates and localized linear combinations of bond angles and torsions as deformational coordinates. (They are the precursors to the delocalized internal coordinates used in PQS, which are linear combinations of all stretches, bends and torsions in the molecule [4].) On the basis of chemical intuition, the natural internal coordinates of all molecules under consideration are sorted into groups sharing a common scaling factor, and factors for each group are determined by a least-squares fit to experimental vibrational frequencies. Force constants, originally calculated in Cartesian coordinates, are transformed into an internal coordinate representation, and scaling is applied to the elements of the internal force constant matrix (not to the individual vibrational frequencies) according to F ij (scaled) = (s i s j ) 1/2 F ij 1
2 where s i and s j are scaling factors for internal coordinates i and j, respectively. The accuracy obtained by selective scaling in this way is naturally greater than if just a single overall scaling factor were used. Additionally, scaling the force constant matrix also affects the resultant normal modes, and hence the calculated intensities (which are unaffected if only the frequencies are scaled), leading to better agreement with experimental intensities. The SQM procedure has been widely used in the interpretation of vibrational spectra. A further important role is the development of transferable scale factors which can be used to modify calculated force constants and so predict the vibrational spectrum a priori. The SQM module uses a modified scaling procedure involving the scaling of individual valence coordinates [5] (not the linear combinations present in natural internal coordinates). This has immediate advantages in terms of ease of use, as no natural internals need to be generated (a procedure which may fail for complicated molecular topologies), and it simplifies the classification and presorting of the coordinates. In addition, the extra flexibility involved in the scaling of individual prinmitive internals generally leads to an increase in accuracy and to more transferable scale factors. The user is encouraged to view the references provided, especially ref. 5. PROGRAM CAPABILITY AND INPUT SQM capabilities include 1. Scaling a force constant matrix using a set of one or more precalculated scale factors. Eleven optimized scale factors are available for standard organics containing H, C, N, O and Cl for force constants calculated at B3LYP/6-31G*. This is one of the most cost-effective and reliable theoretical methods currently available. The recommended scale factors are [5]: stretch X-X stretch C-Cl stretch C-H stretch N-H stretch O-H bend X-X-X bend X-X-H bend H-C-H bend H-N-H torsion all linear bends all
3 where X denotes a non-hydrogen atom (C, N or O). 2. Adjusting atomic masses to give normal modes, frequencies and IR and potentially Raman and VCD intensities for isotopomers. 3. Optimizing scale factors to give the best least-squares fit to a set of experimental vibrational frequencies. 4. Carrying out a total energy distribution analysis [6] to determine how much a given primitive (stretch, bend or torsion) contributes to a particular normal mode. 5. Determining invariant diagonal force constants for all the stretches, bends and torsions in a molecule 6. Carrying out a full vibrational and thermodynamic analysis based on the scaled force constant matrix. The analysis is the same as that in PQS. INPUT FILE A sample input file for formamide (HCONH 2 ) with all keywords shown is given below: $molecules formamide $scaling stre X X fixed stre C H fixed stre N H bend X X X fixed bend X X H fixed bend H N H tors H X X X fixed tors H X X H fixed $print_ted 3.0 $print_level 4 $max_atoms 10 $end $molecules: file prefix names for <hess>, <deriv>, <aat> and <evib> files SQM needs molecular geometries, force constant (Hessian) matrices, dipole (and possibly quadrupole) derivatives and atomic axial tensors as input. The latter are available following a PQS frequency run in the files jobname.hess, jobname.deriv and jobname.aat - in this example these files are called formamide.hess, formamide.deriv and formamide.aat. These files must exist. (SQM will still function if the <deriv> or <aat> files are missing, but no IR, Raman or VCD intensities will be available.) 3
4 Various molecules can be grouped together (for example to determine the best scale factors for the whole set). In this case all the file prefix names should be given, one per line with no blank lines, following the $molecules keyword. $scaling: scaling parameters <scale type> <atom types> <scale group> <value> <action> <scale type>: one of stre, bend, tors, linc, linp (corresponding to stretches, planar bends, proper torsions, and colinear and coplanar bend, respectively. Out-of-plane bends (outp) are currently not accessible) <atom types>: <scale group>: <value>: <action>: up to four atomic symbols (X for any non-hydrogen atom) depending on the <scale type> scale factors with the same (integer) scale group number will be grouped together during any optimization of the scale factors. The scale group number should start at 1 and must be listed consecutively as shown in the example input initial scale factor value either fixed (for a fixed scale factor) or optimize The default if this field is blank is to optimize the scale factor In the formamide example, there are seven scale factors, with the two different torsions in the molecule grouped together. The scale factors for the N-H stretch and the H-N-H bend will be optimized to give the best least-squares fit to a set of experimental frequencies, keeping all the other scale factors fixed at their initial input values. **IMPORTANT** When applying the scale factors, SQM takes each scale factor in turn in the order they appear in the input file and scales any Hessian element that fits. Care must be taken that scale factors involving the use of "X" - for a general non-hydrogen atom - appear first in the list of each scale type (stre, bend, tors, linc, linp) as, if these appear subsequent to a specific atom type (say a C-C stretch), then the specific atoms will be taken as general non-hydrogen atoms and the wrong scale factor will be applied. Note also that "X" cannot be used in place of a hydrogen atom. Thus if all torsions, say, in a given molecule/set of molecules are to be scaled with the same scale factor, then any torsions involving hydrogen must be specifically provided. Thus tors X X X X alone will not scale any torsions involving hydrogen; you also need to specify both tors H X X X and tors H X X H to scale these torsions. 4
5 The other input options are: $print_ted=<real>: print threshold for total energy distribution analysis. This analysis gives the percentage contribution of each primitive stretch, bend and torsion in the molecule to each normal mode. The default if no value is given is 5.0, i.e., only primitives which contribute 5% or more to a given normal mode will be printed for that mode. $print_level=<integer>: controls the amount of printout (larger integer - more printout). In particular, a value of 4 will print the normal modes. Values higher than 4 will progressively output more and more intermediate quantities constructed during the SQM procedure and are essentially for debug printout. $max_atoms=<integer>: for allocating memory. Should be set to at least the number of atoms in the largest molecule under consideration (the default is 50). $end terminates input (must be present) The <evib> File The <evib> file contains both the molecular geometry and, if scale factors are being optimized, the experimental vibrational frequencies. The formamide.evib file is given below: $coordinates angstrom C O H N H H $point_group cs $nsymop 2 $frequencies J.Raman Spectros. 25 (1994) d $end 5
6 The $coordinates section contains the geometry in Cartesian coordinates. The format is: atomic symbol X Y Z atomic mass (A8,2X,4F20.14). This is essentially the same format as the PQS <coord> file except that the atomic charge is missing. Coordinates can be given either in bohr or angstrom; the units are specified following the $coordinates string as shown. If no units are specified, the default is bohr. Atomic Masses There are two ways of specifying isotopomers. The atomic mass can be given following the coordinates (as per the above format) or the isotope can be specified as a part of the atomic symbol, e.g., H-2 will use deuterium instead of hydrogen for that atom, C-13 will give carbon 13. If neither of these options is specified, then the isotopically averaged atomic mass will be used. The SQM program has built-in isotopically averaged atomic masses for all elements up to and including Xenon (N=54). There are also up to four individual isotope masses for each of these elements (if an element has more than four isotopes, then the four with the highest percentage abundance are available). The atomic masses for any atoms not included in the above description must be specified in the $coordinates section. Use of Symmetry The inclusion of full point group symmetry is the major difference between SQM version 2.0 and previous versions. Knowing the molecular point group allows appropriate symmetry labels to be given to each vibrational mode and also allows for a proper thermodynamic analysis. The data required are the point group symbol and the number of symmetry operations. $point_group=<string>: character string (maximum four characters) giving the point group symbol, e.g., cs, c2v, td etc. $nsymop=<integer>: the number of symmetry operations in the point group, e.g., 2, 4, 24 etc.. Symmetry information can either be provided in the <evib> file following the geometry (as shown above) or it can be read from a PQS <sym> file, if one exists. Note that if a <sym> file does exist then it will be read and any symmetry information given in the <evib> file will be overwritten. If no symmetry data is given, C 1 symmetry (i.e., no symmetry) will be assumed. If there is symmetry and it is not used, then certain quantities calculated in the thermodynamic analysis will be incorrect. SQM version 2.0 makes use of the same symmetry routines as PQS and we have made a conscious effort to make the output from SQM as close as possible to the corresponding output in PQS. Indeed if all you want to do is an SQM scaling of the force constant matrix using known scale factors, then this can now be done within PQS itself using the newly introduced SQM keyword (see the PQS manual). 6
7 The $frequencies section contains the experimental (or other) frequencies that will be used in the least-squares fit. The format is <frequency> (in cm -1 ) <weight> with each frequency on a separate line. weight=<real>: gives the weight each vibrational frequency is given in the leastsquares fit. Frequencies that are not known accurately can be given a lower weight in the fitting; conversely frequencies that are regarded as being reliable or for which a good agreement is particularly desired can be given a higher weighting. The default if no weight is given is 1000 x inverse experimental frequency (in cm -1 ). If the experimental frequency for a particular vibration is very suspect or if it is not known at all, it should be given a zero weight. The number of fundamentals a molecule has is given - for a non-linear system it is 3N-6, where N is the number of atoms. Quite often there are less than 3N-6 vibrational fundamentals that are reliably known. In this case, the $frequencies section should have one or more lines containing zeroes (which correspond to an unknown frequency with a zero weight in the fit). It may be necessary to vary the position of these zero lines in the input depending on the accuracy of the fit; note also that although frequencies should generally be given in increasing order, it may be that two fairly close values with different symmetries may need to be switched. This can often be easily detected if one mode is strongly IR active whilst the other is only weakly active or IR inactive; the theoretical mode with the large IR intensity should be fit to the experimentally IR active mode. It is not so uncommon to find experimentally assigned bands that you simply cannot fit at all because they have, in fact, been misassigned. The SQM procedure, when used correctly with a good theoretical method (such as B3LYP/6-31G*), usually gives average errors in band positions of around 8 cm -1, and maximum errors of the order of cm -1. If you find maximum errors significantly outside this range, there is a good chance that the experimental assignment is wrong. $end terminates input (must be present) In the formamide example, both the lowest and the highest frequency fundamentals are not known experimentally (hence the two zero lines) and the fundamental assigned experimentally at 841 cm -1 is considered to be unreliable and has been given zero weight in the fit. The source of the experimental data (J.Raman Spectros. 25 (1994) 183) has been given after the $frequencies keyword as a reminder to the user. 7
8 Invariant ( relaxed ) Force Constants Force constants in internal coordinates can be obtained by a suitable transformation of the Cartesian force constant matrix. Diagonal elements of the internal coordinate force constant matrix give individual stretching, bending and torsional force constants. Unfortunately, the values of these internal coordinate force constants depend on which primitive internals have been choosen to describe the molecular geometry. For example, if a given stretch is present in two sets of internal coordinates (which contain different bends and/or torsions), both sets of which can be used to describe a molecular geometry, then the stretching force constant calculated using one set of coordinates will not have the same value in the other set. This fact is perhaps not as widely known as it ought to be. However, if the force constants are defined not as the diagonal elements of the Hessian matrix in internal coordinates, but instead as the inverse of the diagonal elements of the inverse Hessian, then the two stretching force constants in the example above will be the same. In this way force constants in internal coordinates can be defined in an invariant way, independent of the precise choice of coordinates. The inverse Hessian matrix is known as the compliance matrix. For a fairly recent discussion on relaxed force constants see [7]. Invariant force constants defined in this manner can be output from the SQM program by setting $print_level to 4 in the SQM input file. **IMPORTANT** Both invariant force constants and a total energy distribution of the normal modes will only be performed for single molecules, i.e., there must be only one molecule filename specified in the $molecules section. Program Usage sqm.x filename where the input file should be filename.inp and only the file prefix (i.e., no.inp) should be given. Output will be in filename.out and files associated with the molecule(s) specified in the $molecules section in filename.inp, namely molecule.evib, molecule.hess and (optionally) molecule.deriv and molecule.aat must exist. The SQM program produces four temporary files: sqm-temp1, sqm-temp2, sqmtemp3 and sqm-temp4. As these do not have unique names associated with the input only one SQM job should be run in a given directory at any one time. 8
9 REFERENCES [1] (a) C. E. Blom and C. Altona, Mol. Phys. 31 (1976) 1377 (b) C. E. Blom, L. P. Otto and C. Altona, Mol. Phys. 32 (1976) 1137 (c) C. E. Blom and C. Altona, Mol. Phys. 33 (1977) 875; ibid. 34 (1977) 1137 [2] P. Pulay, G. Fogarasi, G. Pongor, J. E. Boggs and A. Vargha, J. Am. Chem. Soc. 105 (1983) 7037 [3] (a) P. Pulay, G. Fogarasi, F. Pang, J. Boggs, J. Am. Chem. Soc. 101 (1979) 2550; (b) G. Fogarasi, X. Zhou, P. W. Taylor, P. Pulay, J. Am. Chem. Soc. 114 (1992) 8191 [4] J. Baker, A. Kessi and B. Delley, J. Chem. Phys. 105 (1996) 192 [5] J. Baker, A. A. Jarzecki and P. Pulay, J. Phys. Chem. A 102 (1998) 1412 [6] P. Pulay and F. Török, Acta. Chim. Acad. Sci. Hung. 47 (1965) 273 [7] J. Baker, J. Chem. Phys., 125 (2006)
INPUT DESCRIPTION FOR SQM version 1.0
 INPUT DESCRIPTION FOR SQM version 1.0 INTRODUCTION SQM is an add-on module for the PQS program which scales force constants to produce a Scaled Quantum Mechanical (SQM) Force Field. This can correct for
INPUT DESCRIPTION FOR SQM version 1.0 INTRODUCTION SQM is an add-on module for the PQS program which scales force constants to produce a Scaled Quantum Mechanical (SQM) Force Field. This can correct for
Example questions for Molecular modelling (Level 4) Dr. Adrian Mulholland
 Example questions for Molecular modelling (Level 4) Dr. Adrian Mulholland 1) Question. Two methods which are widely used for the optimization of molecular geometies are the Steepest descents and Newton-Raphson
Example questions for Molecular modelling (Level 4) Dr. Adrian Mulholland 1) Question. Two methods which are widely used for the optimization of molecular geometies are the Steepest descents and Newton-Raphson
(N.B. In this document the underlining symbol _ is used to denote a blank.).
 CAMVIB CAMVIB originated in early 1980 as a program to transform cartesian force constants (generated by the HONDO ab initio quantum chemistry package) to internal force constants. It was subsequently
CAMVIB CAMVIB originated in early 1980 as a program to transform cartesian force constants (generated by the HONDO ab initio quantum chemistry package) to internal force constants. It was subsequently
IFM Chemistry Computational Chemistry 2010, 7.5 hp LAB2. Computer laboratory exercise 1 (LAB2): Quantum chemical calculations
 Computer laboratory exercise 1 (LAB2): Quantum chemical calculations Introduction: The objective of the second computer laboratory exercise is to get acquainted with a program for performing quantum chemical
Computer laboratory exercise 1 (LAB2): Quantum chemical calculations Introduction: The objective of the second computer laboratory exercise is to get acquainted with a program for performing quantum chemical
THE VIBRATIONAL SPECTRUM OF A POLYATOMIC MOLECULE (Revised 4/7/2004)
 INTRODUCTION THE VIBRATIONAL SPECTRUM OF A POLYATOMIC MOLECULE (Revised 4/7/2004) The vibrational motion of a molecule is quantized and the resulting energy level spacings give rise to transitions in the
INTRODUCTION THE VIBRATIONAL SPECTRUM OF A POLYATOMIC MOLECULE (Revised 4/7/2004) The vibrational motion of a molecule is quantized and the resulting energy level spacings give rise to transitions in the
DENSITY FUNCTIONAL THEORY STUDIES ON IR SPECTRA OF THE TRIPHENYLENE DERIVATIVES. A SCALED QUANTUM MECHANICAL FORCE FIELD APPROACH
 Vol. 98 (2000) ACTA PHYSICA POLONICA A No. 5 Proceedings of the International Conference "Condensed Matter Physics", Jaszowiec 2000 DENSITY FUNCTIONAL THEORY STUDIES ON IR SPECTRA OF THE TRIPHENYLENE DERIVATIVES.
Vol. 98 (2000) ACTA PHYSICA POLONICA A No. 5 Proceedings of the International Conference "Condensed Matter Physics", Jaszowiec 2000 DENSITY FUNCTIONAL THEORY STUDIES ON IR SPECTRA OF THE TRIPHENYLENE DERIVATIVES.
Chemistry 416 Spectroscopy Fall Semester 1997 Dr. Rainer Glaser
 Chemistry 416 Spectroscopy Fall Semester 1997 Dr. Rainer Glaser Third 1-Hour Examination Vibrational Spectroscopy Monday, November 24, 1997, 8:40-9:30 Name: Answer Key Question 1 (Combination) 25 Question
Chemistry 416 Spectroscopy Fall Semester 1997 Dr. Rainer Glaser Third 1-Hour Examination Vibrational Spectroscopy Monday, November 24, 1997, 8:40-9:30 Name: Answer Key Question 1 (Combination) 25 Question
DMDW: A set of tools to calculate Debye-Waller factors and other related quantities using dynamical matrices.
 DMDW: A set of tools to calculate Debye-Waller factors and other related quantities using dynamical matrices. DMDW is a set of tools developed to calculate Debye-Waller (DW) factors and other related quantities
DMDW: A set of tools to calculate Debye-Waller factors and other related quantities using dynamical matrices. DMDW is a set of tools developed to calculate Debye-Waller (DW) factors and other related quantities
Symmetry: Translation and Rotation
 Symmetry: Translation and Rotation The sixth column of the C 2v character table indicates the symmetry species for translation along (T) and rotation about (R) the Cartesian axes. y y y C 2 F v (x) T x
Symmetry: Translation and Rotation The sixth column of the C 2v character table indicates the symmetry species for translation along (T) and rotation about (R) the Cartesian axes. y y y C 2 F v (x) T x
THE VIBRATIONAL SPECTRA OF A POLYATOMIC MOLECULE (Revised 3/27/2006)
 THE VIBRATIONAL SPECTRA OF A POLYATOMIC MOLECULE (Revised 3/27/2006) 1) INTRODUCTION The vibrational motion of a molecule is quantized and the resulting energy level spacings give rise to transitions in
THE VIBRATIONAL SPECTRA OF A POLYATOMIC MOLECULE (Revised 3/27/2006) 1) INTRODUCTION The vibrational motion of a molecule is quantized and the resulting energy level spacings give rise to transitions in
Description of the MULFIT program February 1998 by G. G. Ferenczy, C. A. Reynolds and P. J. Winn modified by A. J.
 Description of the MULFIT program February 1998 by G. G. Ferenczy, C. A. Reynolds and P. J. Winn modified by A. J. Stone, 2006 2009 1 Introduction This program is the realisation of a method for calculating
Description of the MULFIT program February 1998 by G. G. Ferenczy, C. A. Reynolds and P. J. Winn modified by A. J. Stone, 2006 2009 1 Introduction This program is the realisation of a method for calculating
ARTICLES. Normal-mode analysis without the Hessian: A driven molecular-dynamics approach
 JOURNAL OF CHEMICAL PHYSICS VOLUME 119, NUMBER 2 8 JULY 2003 ARTICLES Normal-mode analysis without the Hessian: A driven molecular-dynamics approach Joel M. Bowman, a) Xiubin Zhang, and Alex Brown Cherry
JOURNAL OF CHEMICAL PHYSICS VOLUME 119, NUMBER 2 8 JULY 2003 ARTICLES Normal-mode analysis without the Hessian: A driven molecular-dynamics approach Joel M. Bowman, a) Xiubin Zhang, and Alex Brown Cherry
Calculating Vibrational Spectra from Molecular Dynamics
 Calculating Vibrational Spectra from Molecular Dynamics A Simulating a Trajectory with Wannier Centers To calculate IR spectra from Molecular Dynamics, it is necessary to have dipole information for the
Calculating Vibrational Spectra from Molecular Dynamics A Simulating a Trajectory with Wannier Centers To calculate IR spectra from Molecular Dynamics, it is necessary to have dipole information for the
Chemistry 4560/5560 Molecular Modeling Fall 2014
 Final Exam Name:. User s guide: 1. Read questions carefully and make sure you understand them before answering (if not, ask). 2. Answer only the question that is asked, not a different question. 3. Unless
Final Exam Name:. User s guide: 1. Read questions carefully and make sure you understand them before answering (if not, ask). 2. Answer only the question that is asked, not a different question. 3. Unless
Chemistry 543--Final Exam--Keiderling May 5, pm SES
 Chemistry 543--Final Exam--Keiderling May 5,1992 -- 1-5pm -- 174 SES Please answer all questions in the answer book provided. Make sure your name is clearly indicated and that the answers are clearly numbered,
Chemistry 543--Final Exam--Keiderling May 5,1992 -- 1-5pm -- 174 SES Please answer all questions in the answer book provided. Make sure your name is clearly indicated and that the answers are clearly numbered,
MO Calculation for a Diatomic Molecule. /4 0 ) i=1 j>i (1/r ij )
 MO Calculation for a Diatomic Molecule Introduction The properties of any molecular system can in principle be found by looking at the solutions to the corresponding time independent Schrodinger equation
MO Calculation for a Diatomic Molecule Introduction The properties of any molecular system can in principle be found by looking at the solutions to the corresponding time independent Schrodinger equation
QUANTUM CHEMISTRY WITH GAUSSIAN : A VERY BRIEF INTRODUCTION (PART 2)
 QUANTUM CHEMISTRY WITH GAUSSIAN : A VERY BRIEF INTRODUCTION (PART 2) TARAS V. POGORELOV AND MIKE HALLOCK SCHOOL OF CHEMICAL SCIENCES, UIUC This tutorial continues introduction to Gaussian [2]. Here we
QUANTUM CHEMISTRY WITH GAUSSIAN : A VERY BRIEF INTRODUCTION (PART 2) TARAS V. POGORELOV AND MIKE HALLOCK SCHOOL OF CHEMICAL SCIENCES, UIUC This tutorial continues introduction to Gaussian [2]. Here we
Renner-Teller Effect in Tetra-Atomic Molecules
 Groupe de Chimie Théorique du MSME Renner-Teller Effect in Tetra-Atomic Molecules Laurent Jutier, G. Dhont, H. Khalil and C. Léonard jutier@univ-mlv.fr (non linear) Outline General Presentation Structure
Groupe de Chimie Théorique du MSME Renner-Teller Effect in Tetra-Atomic Molecules Laurent Jutier, G. Dhont, H. Khalil and C. Léonard jutier@univ-mlv.fr (non linear) Outline General Presentation Structure
Transition states and reaction paths
 Transition states and reaction paths Lab 4 Theoretical background Transition state A transition structure is the molecular configuration that separates reactants and products. In a system with a single
Transition states and reaction paths Lab 4 Theoretical background Transition state A transition structure is the molecular configuration that separates reactants and products. In a system with a single
Example: H 2 O (the car file)
 Example: H 2 O (the car file) As a practical example of DFT methods we calculate the energy and electronic properties of the water molecule. In order to carry out the DFT calculation you will need a set
Example: H 2 O (the car file) As a practical example of DFT methods we calculate the energy and electronic properties of the water molecule. In order to carry out the DFT calculation you will need a set
Jaguar DFT Optimizations and Transition State Searches
 Jaguar DFT Optimizations and Transition State Searches Density Functional Theory (DFT) is a quantum mechanical (QM) method that gives results superior to Hartree Fock (HF) in less computational time. A
Jaguar DFT Optimizations and Transition State Searches Density Functional Theory (DFT) is a quantum mechanical (QM) method that gives results superior to Hartree Fock (HF) in less computational time. A
Vibrations of Carbon Dioxide and Carbon Disulfide
 Vibrations of Carbon Dioxide and Carbon Disulfide Purpose Vibration frequencies of CO 2 and CS 2 will be measured by Raman and Infrared spectroscopy. The spectra show effects of normal mode symmetries
Vibrations of Carbon Dioxide and Carbon Disulfide Purpose Vibration frequencies of CO 2 and CS 2 will be measured by Raman and Infrared spectroscopy. The spectra show effects of normal mode symmetries
Vibrational frequencies in solids: tools and tricks
 Vibrational frequencies in solids: tools and tricks Roberto Dovesi Gruppo di Chimica Teorica Università di Torino Torino, 4-9 September 2016 This morning 3 lectures: R. Dovesi Generalities on vibrations
Vibrational frequencies in solids: tools and tricks Roberto Dovesi Gruppo di Chimica Teorica Università di Torino Torino, 4-9 September 2016 This morning 3 lectures: R. Dovesi Generalities on vibrations
Using Molecular Dynamics to Compute Properties CHEM 430
 Using Molecular Dynamics to Compute Properties CHEM 43 Heat Capacity and Energy Fluctuations Running an MD Simulation Equilibration Phase Before data-collection and results can be analyzed the system
Using Molecular Dynamics to Compute Properties CHEM 43 Heat Capacity and Energy Fluctuations Running an MD Simulation Equilibration Phase Before data-collection and results can be analyzed the system
16.1 Molecular Vibrations
 16.1 Molecular Vibrations molecular degrees of freedom are used to predict the number of vibrational modes vibrations occur as coordinated movement among many nuclei the harmonic oscillator approximation
16.1 Molecular Vibrations molecular degrees of freedom are used to predict the number of vibrational modes vibrations occur as coordinated movement among many nuclei the harmonic oscillator approximation
Figure 1: Transition State, Saddle Point, Reaction Pathway
 Computational Chemistry Workshops West Ridge Research Building-UAF Campus 9:00am-4:00pm, Room 009 Electronic Structure - July 19-21, 2016 Molecular Dynamics - July 26-28, 2016 Potential Energy Surfaces
Computational Chemistry Workshops West Ridge Research Building-UAF Campus 9:00am-4:00pm, Room 009 Electronic Structure - July 19-21, 2016 Molecular Dynamics - July 26-28, 2016 Potential Energy Surfaces
Spectroscopy in Inorganic Chemistry. Vibration and Rotation Spectroscopy
 Spectroscopy in Inorganic Chemistry Vibrational energy levels in a diatomic molecule f = k r r V = ½kX 2 Force constant r Displacement from equilibrium point 2 X= r=r-r eq V = ½kX 2 Fundamental Vibrational
Spectroscopy in Inorganic Chemistry Vibrational energy levels in a diatomic molecule f = k r r V = ½kX 2 Force constant r Displacement from equilibrium point 2 X= r=r-r eq V = ½kX 2 Fundamental Vibrational
DFT calculations of NMR indirect spin spin coupling constants
 DFT calculations of NMR indirect spin spin coupling constants Dalton program system Program capabilities Density functional theory Kohn Sham theory LDA, GGA and hybrid theories Indirect NMR spin spin coupling
DFT calculations of NMR indirect spin spin coupling constants Dalton program system Program capabilities Density functional theory Kohn Sham theory LDA, GGA and hybrid theories Indirect NMR spin spin coupling
Chemistry 431. NC State University. Lecture 17. Vibrational Spectroscopy
 Chemistry 43 Lecture 7 Vibrational Spectroscopy NC State University The Dipole Moment Expansion The permanent dipole moment of a molecule oscillates about an equilibrium value as the molecule vibrates.
Chemistry 43 Lecture 7 Vibrational Spectroscopy NC State University The Dipole Moment Expansion The permanent dipole moment of a molecule oscillates about an equilibrium value as the molecule vibrates.
This is called a singlet or spin singlet, because the so called multiplicity, or number of possible orientations of the total spin, which is
 9. Open shell systems The derivation of Hartree-Fock equations (Chapter 7) was done for a special case of a closed shell systems. Closed shell means that each MO is occupied by two electrons with the opposite
9. Open shell systems The derivation of Hartree-Fock equations (Chapter 7) was done for a special case of a closed shell systems. Closed shell means that each MO is occupied by two electrons with the opposite
V( x) = V( 0) + dv. V( x) = 1 2
 Spectroscopy 1: rotational and vibrational spectra The vibrations of diatomic molecules Molecular vibrations Consider a typical potential energy curve for a diatomic molecule. In regions close to R e (at
Spectroscopy 1: rotational and vibrational spectra The vibrations of diatomic molecules Molecular vibrations Consider a typical potential energy curve for a diatomic molecule. In regions close to R e (at
Geometry optimization in redundant internal coordinates
 Geometry optimization in redundant internal coordinates P. Pulay and G. Fogarasi a ) Department a/chemistry and Biochemistry, The University 0/ Arkansas, Fayetteville, Arkansas 72701 (Received 25 October
Geometry optimization in redundant internal coordinates P. Pulay and G. Fogarasi a ) Department a/chemistry and Biochemistry, The University 0/ Arkansas, Fayetteville, Arkansas 72701 (Received 25 October
Computational chemistry with GAMESS: a very brief overview with examples
 Computational chemistry with GAMESS: a very brief overview with examples PHY-6120 Molecular Physics (Spring 2015), UConn Phys. Dept. Feb 17 th 2015 H = ħ2 2μ i Intro: V(R) for diatomic molecules + k Z
Computational chemistry with GAMESS: a very brief overview with examples PHY-6120 Molecular Physics (Spring 2015), UConn Phys. Dept. Feb 17 th 2015 H = ħ2 2μ i Intro: V(R) for diatomic molecules + k Z
An Introduction to Quantum Chemistry and Potential Energy Surfaces. Benjamin G. Levine
 An Introduction to Quantum Chemistry and Potential Energy Surfaces Benjamin G. Levine This Week s Lecture Potential energy surfaces What are they? What are they good for? How do we use them to solve chemical
An Introduction to Quantum Chemistry and Potential Energy Surfaces Benjamin G. Levine This Week s Lecture Potential energy surfaces What are they? What are they good for? How do we use them to solve chemical
Lecture 8. Assumed knowledge
 Chemistry 2 Lecture 8 IR Spectroscopy of Polyatomic Molecles Assumed knowledge There are 3N 6 vibrations in a non linear molecule and 3N 5 vibrations in a linear molecule. Only modes that lead to a change
Chemistry 2 Lecture 8 IR Spectroscopy of Polyatomic Molecles Assumed knowledge There are 3N 6 vibrations in a non linear molecule and 3N 5 vibrations in a linear molecule. Only modes that lead to a change
The successful wavefunction can be written as a determinant: # 1 (2) # 2 (2) Electrons. This can be generalized to our 2N-electron wavefunction:
 T2. CNDO to AM1: The Semiempirical Molecular Orbital Models The discussion in sections T2.1 T2.3 applies also to ab initio molecular orbital calculations. T2.1 Slater Determinants Consider the general
T2. CNDO to AM1: The Semiempirical Molecular Orbital Models The discussion in sections T2.1 T2.3 applies also to ab initio molecular orbital calculations. T2.1 Slater Determinants Consider the general
Semi-Empirical MO Methods
 Semi-Empirical MO Methods the high cost of ab initio MO calculations is largely due to the many integrals that need to be calculated (esp. two electron integrals) semi-empirical MO methods start with the
Semi-Empirical MO Methods the high cost of ab initio MO calculations is largely due to the many integrals that need to be calculated (esp. two electron integrals) semi-empirical MO methods start with the
NPA/NBO-Analysis. Examples POP =
 NPA/NBO-Analysis Examples POP = NBO Requests a full Natural Bond Orbital analysis, using NBO version 3 NPA Requests just the Natural Population Analysis phase of NBO. NBORead Requests a full NBO analysis,
NPA/NBO-Analysis Examples POP = NBO Requests a full Natural Bond Orbital analysis, using NBO version 3 NPA Requests just the Natural Population Analysis phase of NBO. NBORead Requests a full NBO analysis,
Quantum chemistry and vibrational spectra
 Chapter 3 Quantum chemistry and vibrational spectra This chapter presents the quantum chemical results for the systems studied in this work, FHF (Section 3.) and OHF (Section 3.3). These triatomic anions
Chapter 3 Quantum chemistry and vibrational spectra This chapter presents the quantum chemical results for the systems studied in this work, FHF (Section 3.) and OHF (Section 3.3). These triatomic anions
Exercise 1: Structure and dipole moment of a small molecule
 Introduction to computational chemistry Exercise 1: Structure and dipole moment of a small molecule Vesa Hänninen 1 Introduction In this exercise the equilibrium structure and the dipole moment of a small
Introduction to computational chemistry Exercise 1: Structure and dipole moment of a small molecule Vesa Hänninen 1 Introduction In this exercise the equilibrium structure and the dipole moment of a small
Literature values: ΔH f, gas = % error Source: ΔH f, solid = % error. For comparison, your experimental value was ΔH f = phase:
 1 Molecular Calculations Lab: Some guideline given at the bottom of page 3. 1. Use the semi-empirical AM1 method to calculate ΔH f for the compound you used in the heat of combustion experiment. Be sure
1 Molecular Calculations Lab: Some guideline given at the bottom of page 3. 1. Use the semi-empirical AM1 method to calculate ΔH f for the compound you used in the heat of combustion experiment. Be sure
Session 1. Introduction to Computational Chemistry. Computational (chemistry education) and/or (Computational chemistry) education
 Session 1 Introduction to Computational Chemistry 1 Introduction to Computational Chemistry Computational (chemistry education) and/or (Computational chemistry) education First one: Use computational tools
Session 1 Introduction to Computational Chemistry 1 Introduction to Computational Chemistry Computational (chemistry education) and/or (Computational chemistry) education First one: Use computational tools
Introduction to Vibrational Spectroscopy
 Introduction to Vibrational Spectroscopy Harmonic oscillators The classical harmonic oscillator The uantum mechanical harmonic oscillator Harmonic approximations in molecular vibrations Vibrational spectroscopy
Introduction to Vibrational Spectroscopy Harmonic oscillators The classical harmonic oscillator The uantum mechanical harmonic oscillator Harmonic approximations in molecular vibrations Vibrational spectroscopy
Probing the Origins of Intermolecular Vibrational and Relaxational Dynamics in Organic Solids with CP2K
 Probing the Origins of Intermolecular Vibrational and Relaxational Dynamics in Organic Solids with CP2K Michael Ruggiero Department of Chemical Engineering and Biotechnology, University of Cambridge CP2K
Probing the Origins of Intermolecular Vibrational and Relaxational Dynamics in Organic Solids with CP2K Michael Ruggiero Department of Chemical Engineering and Biotechnology, University of Cambridge CP2K
NMR and IR spectra & vibrational analysis
 Lab 5: NMR and IR spectra & vibrational analysis A brief theoretical background 1 Some of the available chemical quantum methods for calculating NMR chemical shifts are based on the Hartree-Fock self-consistent
Lab 5: NMR and IR spectra & vibrational analysis A brief theoretical background 1 Some of the available chemical quantum methods for calculating NMR chemical shifts are based on the Hartree-Fock self-consistent
Computational Chemistry. An Introduction to Molecular Dynamic Simulations
 Computational Chemistry An Introduction to Molecular Dynamic Simulations Computational chemistry simulates chemical structures and reactions numerically, based in full or in part on the fundamental laws
Computational Chemistry An Introduction to Molecular Dynamic Simulations Computational chemistry simulates chemical structures and reactions numerically, based in full or in part on the fundamental laws
J.Phys. & Theo.Chem.I.A.U. Iran M.Monajjemi et al. Vol.4, No.1, Spring 2007
 Journal of Physical & Theoretical Chemistry Islamic Azad University of Iran 4 (1) (27) Science and Research Campus ISSN: 1735-2126 AB Initio Calculations and IR Studies of Tautometric forms of Uracil and
Journal of Physical & Theoretical Chemistry Islamic Azad University of Iran 4 (1) (27) Science and Research Campus ISSN: 1735-2126 AB Initio Calculations and IR Studies of Tautometric forms of Uracil and
Beyond the Hartree-Fock Approximation: Configuration Interaction
 Beyond the Hartree-Fock Approximation: Configuration Interaction The Hartree-Fock (HF) method uses a single determinant (single electronic configuration) description of the electronic wavefunction. For
Beyond the Hartree-Fock Approximation: Configuration Interaction The Hartree-Fock (HF) method uses a single determinant (single electronic configuration) description of the electronic wavefunction. For
Algebraic Study of Stretching and Bending Modes in Linear Tetra-atomic Molecules: HCCCl
 The African Review of Physics (2013) 8:0016 99 Algebraic Study of Stretching and Bending Modes in Linear Tetra-atomic Molecules: HCCCl Kamal Ziadi * Department of Chemistry, Faculty of Science, University
The African Review of Physics (2013) 8:0016 99 Algebraic Study of Stretching and Bending Modes in Linear Tetra-atomic Molecules: HCCCl Kamal Ziadi * Department of Chemistry, Faculty of Science, University
Appendix D Simulating Spectroscopic Bands Using Gaussian and PGopher
 429 Appendix D Simulating Spectroscopic Bands Using Gaussian and PGopher This appendix contains methods for using Gaussian 09 121 and PGopher 120 to simulate vibrational and electronic bands of molecules.
429 Appendix D Simulating Spectroscopic Bands Using Gaussian and PGopher This appendix contains methods for using Gaussian 09 121 and PGopher 120 to simulate vibrational and electronic bands of molecules.
T6.2 Molecular Mechanics
 T6.2 Molecular Mechanics We have seen that Benson group additivities are capable of giving heats of formation of molecules with accuracies comparable to those of the best ab initio procedures. However,
T6.2 Molecular Mechanics We have seen that Benson group additivities are capable of giving heats of formation of molecules with accuracies comparable to those of the best ab initio procedures. However,
Degrees of Freedom and Vibrational Modes
 Degrees of Freedom and Vibrational Modes 1. Every atom in a molecule can move in three possible directions relative to a Cartesian coordinate, so for a molecule of n atoms there are 3n degrees of freedom.
Degrees of Freedom and Vibrational Modes 1. Every atom in a molecule can move in three possible directions relative to a Cartesian coordinate, so for a molecule of n atoms there are 3n degrees of freedom.
Advanced Electronic Structure Theory Density functional theory. Dr Fred Manby
 Advanced Electronic Structure Theory Density functional theory Dr Fred Manby fred.manby@bris.ac.uk http://www.chm.bris.ac.uk/pt/manby/ 6 Strengths of DFT DFT is one of many theories used by (computational)
Advanced Electronic Structure Theory Density functional theory Dr Fred Manby fred.manby@bris.ac.uk http://www.chm.bris.ac.uk/pt/manby/ 6 Strengths of DFT DFT is one of many theories used by (computational)
4. Constraints and Hydrogen Atoms
 4. Constraints and ydrogen Atoms 4.1 Constraints versus restraints In crystal structure refinement, there is an important distinction between a constraint and a restraint. A constraint is an exact mathematical
4. Constraints and ydrogen Atoms 4.1 Constraints versus restraints In crystal structure refinement, there is an important distinction between a constraint and a restraint. A constraint is an exact mathematical
Computational and spectroscopic investigation of 7-azaindole: Solvation and intermolecular interactions
 Computational and spectroscopic investigation of 7-azaindole: Solvation and intermolecular interactions Michael Kamrath, Krista Cruse, Nathan Erickson, Molly Beernink Abstract We report results of an experimental
Computational and spectroscopic investigation of 7-azaindole: Solvation and intermolecular interactions Michael Kamrath, Krista Cruse, Nathan Erickson, Molly Beernink Abstract We report results of an experimental
Chemistry 2. Assumed knowledge
 Chemistry 2 Lecture 8 IR Spectroscopy of Polyatomic Molecles Assumed knowledge There are 3N 6 vibrations in a non linear molecule and 3N 5 vibrations in a linear molecule. Only modes that lead to a change
Chemistry 2 Lecture 8 IR Spectroscopy of Polyatomic Molecles Assumed knowledge There are 3N 6 vibrations in a non linear molecule and 3N 5 vibrations in a linear molecule. Only modes that lead to a change
Introduction to Molecular Vibrations and Infrared Spectroscopy
 hemistry 362 Spring 2017 Dr. Jean M. Standard February 15, 2017 Introduction to Molecular Vibrations and Infrared Spectroscopy Vibrational Modes For a molecule with N atoms, the number of vibrational modes
hemistry 362 Spring 2017 Dr. Jean M. Standard February 15, 2017 Introduction to Molecular Vibrations and Infrared Spectroscopy Vibrational Modes For a molecule with N atoms, the number of vibrational modes
Introduction to computational chemistry Exercise I: Structure and electronic energy of a small molecule. Vesa Hänninen
 Introduction to computational chemistry Exercise I: Structure and electronic energy of a small molecule Vesa Hänninen 1 Introduction In this exercise the equilibrium structure and the electronic energy
Introduction to computational chemistry Exercise I: Structure and electronic energy of a small molecule Vesa Hänninen 1 Introduction In this exercise the equilibrium structure and the electronic energy
Infrared spectra of small biomolecules from first-principle molecular dynamics simulations and effective normal mode analysis
 Infrared spectra of small biomolecules from first-principle molecular dynamics simulations and effective normal mode analysis R. Vuilleumier, M.-P. Gaigeot and D. Borgis Département de chimie, Ecole Normale
Infrared spectra of small biomolecules from first-principle molecular dynamics simulations and effective normal mode analysis R. Vuilleumier, M.-P. Gaigeot and D. Borgis Département de chimie, Ecole Normale
The Effect of Model Internal Flexibility Upon NEMD Simulations of Viscosity
 Draft: September 29, 1999 The Effect of Model Internal Flexibility Upon NEMD Simulations of Viscosity N. G. Fuller 1 and R. L. Rowley 1,2 Abstract The influence of model flexibility upon simulated viscosity
Draft: September 29, 1999 The Effect of Model Internal Flexibility Upon NEMD Simulations of Viscosity N. G. Fuller 1 and R. L. Rowley 1,2 Abstract The influence of model flexibility upon simulated viscosity
Vibrational Spectroscopy
 Vibrational Spectroscopy In this part of the course we will look at the kind of spectroscopy which uses light to excite the motion of atoms. The forces required to move atoms are smaller than those required
Vibrational Spectroscopy In this part of the course we will look at the kind of spectroscopy which uses light to excite the motion of atoms. The forces required to move atoms are smaller than those required
Physical Chemistry Lab II CHEM 4644 Spring 2011 Final Exam 5 questions at 3 points each equals 15 total points possible.
 Physical Chemistry Lab II Name: KEY CHEM 4644 Spring 2011 Final Exam 5 questions at 3 points each equals 15 total points possible. Constants: c = 3.00 10 8 m/s h = 6.63 10-34 J s 1 Hartree = 4.36 10-18
Physical Chemistry Lab II Name: KEY CHEM 4644 Spring 2011 Final Exam 5 questions at 3 points each equals 15 total points possible. Constants: c = 3.00 10 8 m/s h = 6.63 10-34 J s 1 Hartree = 4.36 10-18
Chapter 6 Vibrational Spectroscopy
 Chapter 6 Vibrational Spectroscopy As with other applications of symmetry and group theory, these techniques reach their greatest utility when applied to the analysis of relatively small molecules in either
Chapter 6 Vibrational Spectroscopy As with other applications of symmetry and group theory, these techniques reach their greatest utility when applied to the analysis of relatively small molecules in either
Chem 344 Final Exam Tuesday, Dec. 11, 2007, 3-?? PM
 Chem 344 Final Exam Tuesday, Dec. 11, 2007, 3-?? PM Closed book exam, only pencils and calculators permitted. You may bring and use one 8 1/2 x 11" paper with anything on it. No Computers. Put all of your
Chem 344 Final Exam Tuesday, Dec. 11, 2007, 3-?? PM Closed book exam, only pencils and calculators permitted. You may bring and use one 8 1/2 x 11" paper with anything on it. No Computers. Put all of your
Ethene. Introduction. The ethene molecule is planar (i.e. all the six atoms lie in the same plane) and has a high degree of symmetry:
 FY1006 Innføring i kvantefysikk og TFY4215 Kjemisk fysikk og kvantemekanikk Spring 2012 Chemical Physics Exercise 1 To be delivered by Friday 27.04.12 Introduction Ethene. Ethylene, C 2 H 4, or ethene,
FY1006 Innføring i kvantefysikk og TFY4215 Kjemisk fysikk og kvantemekanikk Spring 2012 Chemical Physics Exercise 1 To be delivered by Friday 27.04.12 Introduction Ethene. Ethylene, C 2 H 4, or ethene,
Free Energy Change and Activation Barrier for a Menshutkin Reaction Including Effects of the Solvent
 Proposed Exercise for the Physical Chemistry Section of the Teaching with Cache Workbook: Free Energy Change and Activation Barrier for a Menshutkin Reaction Including Effects of the Solvent Contributed
Proposed Exercise for the Physical Chemistry Section of the Teaching with Cache Workbook: Free Energy Change and Activation Barrier for a Menshutkin Reaction Including Effects of the Solvent Contributed
Computer Algebraic Tools for Studying the Symmetry Properties of Molecules and Clusters. Katya Rykhlinskaya, University of Kassel
 Computer Algebraic Tools for Studying the Symmetry Properties of Molecules and Clusters Katya Rykhlinskaya, University of Kassel 02. 06. 2005 Computational techniques in the theoretical investigations
Computer Algebraic Tools for Studying the Symmetry Properties of Molecules and Clusters Katya Rykhlinskaya, University of Kassel 02. 06. 2005 Computational techniques in the theoretical investigations
Excited States Calculations for Protonated PAHs
 52 Chapter 3 Excited States Calculations for Protonated PAHs 3.1 Introduction Protonated PAHs are closed shell ions. Their electronic structure should therefore be similar to that of neutral PAHs, but
52 Chapter 3 Excited States Calculations for Protonated PAHs 3.1 Introduction Protonated PAHs are closed shell ions. Their electronic structure should therefore be similar to that of neutral PAHs, but
VIBRATION-ROTATION SPECTRUM OF CO
 Rice University Physics 332 VIBRATION-ROTATION SPECTRUM OF CO I. INTRODUCTION...2 II. THEORETICAL CONSIDERATIONS...3 III. MEASUREMENTS...8 IV. ANALYSIS...9 April 2011 I. Introduction Optical spectroscopy
Rice University Physics 332 VIBRATION-ROTATION SPECTRUM OF CO I. INTRODUCTION...2 II. THEORETICAL CONSIDERATIONS...3 III. MEASUREMENTS...8 IV. ANALYSIS...9 April 2011 I. Introduction Optical spectroscopy
Symmetry. Chemistry 481(01) Spring 2017 Instructor: Dr. Upali Siriwardane Office: CTH 311 Phone Office Hours:
 Chemistry 481(01) Spring 2017 Instructor: Dr. Upali Siriwardane e-mail: upali@latech.edu Office: CT 311 Phone 257-4941 Office ours: M,W 8:00-9:00 & 11:00-12:00 am; Tu,Th, F 9:30-11:30 a.m. April 4, 2017:
Chemistry 481(01) Spring 2017 Instructor: Dr. Upali Siriwardane e-mail: upali@latech.edu Office: CT 311 Phone 257-4941 Office ours: M,W 8:00-9:00 & 11:00-12:00 am; Tu,Th, F 9:30-11:30 a.m. April 4, 2017:
Wolfgang Demtroder. Molecular Physics. Theoretical Principles and Experimental Methods WILEY- VCH. WILEY-VCH Verlag GmbH & Co.
 Wolfgang Demtroder Molecular Physics Theoretical Principles and Experimental Methods WILEY- VCH WILEY-VCH Verlag GmbH & Co. KGaA v Preface xiii 1 Introduction 1 1.1 Short Historical Overview 2 1.2 Molecular
Wolfgang Demtroder Molecular Physics Theoretical Principles and Experimental Methods WILEY- VCH WILEY-VCH Verlag GmbH & Co. KGaA v Preface xiii 1 Introduction 1 1.1 Short Historical Overview 2 1.2 Molecular
LECTURE 3 DIRECT PRODUCTS AND SPECTROSCOPIC SELECTION RULES
 SYMMETRY II. J. M. GOICOECHEA. LECTURE 3 1 LECTURE 3 DIRECT PRODUCTS AND SPECTROSCOPIC SELECTION RULES 3.1 Direct products and many electron states Consider the problem of deciding upon the symmetry of
SYMMETRY II. J. M. GOICOECHEA. LECTURE 3 1 LECTURE 3 DIRECT PRODUCTS AND SPECTROSCOPIC SELECTION RULES 3.1 Direct products and many electron states Consider the problem of deciding upon the symmetry of
Rotational Raman Spectroscopy
 Rotational Raman Spectroscopy If EM radiation falls upon an atom or molecule, it may be absorbed if the energy of the radiation corresponds to the separation of two energy levels of the atoms or molecules.
Rotational Raman Spectroscopy If EM radiation falls upon an atom or molecule, it may be absorbed if the energy of the radiation corresponds to the separation of two energy levels of the atoms or molecules.
Symmetrical: implies the species possesses a number of indistinguishable configurations.
 Chapter 3 - Molecular Symmetry Symmetry helps us understand molecular structure, some chemical properties, and characteristics of physical properties (spectroscopy) used with group theory to predict vibrational
Chapter 3 - Molecular Symmetry Symmetry helps us understand molecular structure, some chemical properties, and characteristics of physical properties (spectroscopy) used with group theory to predict vibrational
Hints on Using the Orca Program
 Computational Chemistry Workshops West Ridge Research Building-UAF Campus 9:00am-4:00pm, Room 009 Electronic Structure - July 19-21, 2016 Molecular Dynamics - July 26-28, 2016 Hints on Using the Orca Program
Computational Chemistry Workshops West Ridge Research Building-UAF Campus 9:00am-4:00pm, Room 009 Electronic Structure - July 19-21, 2016 Molecular Dynamics - July 26-28, 2016 Hints on Using the Orca Program
Chem 673, Problem Set 5 Due Thursday, November 29, 2007
 Chem 673, Problem Set 5 Due Thursday, November 29, 2007 (1) Trigonal prismatic coordination is fairly common in solid-state inorganic chemistry. In most cases the geometry of the trigonal prism is such
Chem 673, Problem Set 5 Due Thursday, November 29, 2007 (1) Trigonal prismatic coordination is fairly common in solid-state inorganic chemistry. In most cases the geometry of the trigonal prism is such
Chem 673, Problem Set 5 Due Tuesday, December 2, 2008
 Chem 673, Problem Set 5 Due Tuesday, December 2, 2008 (1) (a) Trigonal bipyramidal (tbp) coordination is fairly common. Calculate the group overlaps of the appropriate SALCs for a tbp with the 5 d-orbitals
Chem 673, Problem Set 5 Due Tuesday, December 2, 2008 (1) (a) Trigonal bipyramidal (tbp) coordination is fairly common. Calculate the group overlaps of the appropriate SALCs for a tbp with the 5 d-orbitals
5.80 Small-Molecule Spectroscopy and Dynamics
 MIT OpenCourseWare http://ocw.mit.edu 5.80 Small-Molecule Spectroscopy and Dynamics Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms. 5.76 Lecture
MIT OpenCourseWare http://ocw.mit.edu 5.80 Small-Molecule Spectroscopy and Dynamics Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms. 5.76 Lecture
Gustavus Adolphus College. Lab #5: Computational Chemistry
 CHE 372 Gustavus Adolphus College Lab #5: Computational Chemistry Introduction In this investigation we will apply the techniques of computational chemistry to several of the molecular systems that we
CHE 372 Gustavus Adolphus College Lab #5: Computational Chemistry Introduction In this investigation we will apply the techniques of computational chemistry to several of the molecular systems that we
XYZ file format Protein Data Bank (pdb) file format Z - matrix
 Chemistry block (exercise 1) In this exercise, students will be introduced how to preform simple quantum chemical calculations. Input files for Gaussian09. Output file structure. Geometry optimization,
Chemistry block (exercise 1) In this exercise, students will be introduced how to preform simple quantum chemical calculations. Input files for Gaussian09. Output file structure. Geometry optimization,
Organic Spectra Infra Red Spectroscopy H. D. Roth. THEORY and INTERPRETATION of ORGANIC SPECTRA H. D. Roth. Infra Red Spectroscopy
 rganic Spectra Infra Red Spectroscopy. D. Roth TERY and INTERPRETATIN of RGANI SPETRA. D. Roth Infra Red Spectroscopy Infrared spectroscopy (IR) is an analytical technique concerned with molecular vibrations
rganic Spectra Infra Red Spectroscopy. D. Roth TERY and INTERPRETATIN of RGANI SPETRA. D. Roth Infra Red Spectroscopy Infrared spectroscopy (IR) is an analytical technique concerned with molecular vibrations
PAPER No. : 8 (PHYSICAL SPECTROSCOPY) MODULE No. : 5 (TRANSITION PROBABILITIES AND TRANSITION DIPOLE MOMENT. OVERVIEW OF SELECTION RULES)
 Subject Chemistry Paper No and Title Module No and Title Module Tag 8 and Physical Spectroscopy 5 and Transition probabilities and transition dipole moment, Overview of selection rules CHE_P8_M5 TABLE
Subject Chemistry Paper No and Title Module No and Title Module Tag 8 and Physical Spectroscopy 5 and Transition probabilities and transition dipole moment, Overview of selection rules CHE_P8_M5 TABLE
(2) Read each statement carefully and pick the one that is incorrect in its information.
 Organic Chemistry - Problem Drill 17: IR and Mass Spectra No. 1 of 10 1. Which statement about infrared spectroscopy is incorrect? (A) IR spectroscopy is a method of structure determination based on the
Organic Chemistry - Problem Drill 17: IR and Mass Spectra No. 1 of 10 1. Which statement about infrared spectroscopy is incorrect? (A) IR spectroscopy is a method of structure determination based on the
All measurement has a limit of precision and accuracy, and this must be taken into account when evaluating experimental results.
 Chapter 11: Measurement and data processing and analysis 11.1 Uncertainty and error in measurement and results All measurement has a limit of precision and accuracy, and this must be taken into account
Chapter 11: Measurement and data processing and analysis 11.1 Uncertainty and error in measurement and results All measurement has a limit of precision and accuracy, and this must be taken into account
Chapter 6 Answers to Problems
 Chapter 6 Answers to Problems 6.1 (a) NH 3 C3v E 2C3 3 v 4 1 2 3 0 1 12 0 2 3n = 3A 1 A 2 4E trans = A 1 E rot = A 2 E = 2A 2E = 4 frequencies 3n-6 1 Infrared 4 (2A 1 2E) Raman 4 (2A 1 2E) Polarized 2
Chapter 6 Answers to Problems 6.1 (a) NH 3 C3v E 2C3 3 v 4 1 2 3 0 1 12 0 2 3n = 3A 1 A 2 4E trans = A 1 E rot = A 2 E = 2A 2E = 4 frequencies 3n-6 1 Infrared 4 (2A 1 2E) Raman 4 (2A 1 2E) Polarized 2
MRCI calculations in MOLPRO
 1 MRCI calculations in MOLPRO Molpro is a software package written in Fortran and maintained by H.J. Werner and P.J. Knowles. It is often used for performing sophisticated electronic structure calculations,
1 MRCI calculations in MOLPRO Molpro is a software package written in Fortran and maintained by H.J. Werner and P.J. Knowles. It is often used for performing sophisticated electronic structure calculations,
AB INITIO CALCULATIONS AND VIBRATIONAL SPECTROSCOPIC STUDIES OF 2-CHLORO-6- METHOXYPYRIDINE
 AB INITIO CALCULATIONS AND VIBRATIONAL SPECTROSCOPIC STUDIES OF 2-CHLORO-6- METHOXYPYRIDINE L.Usha Kumari a, M.Fathima Beegum a, B.Harikumar b, Hema Tresa Varghese c, C.Yohannan Panicker a* a Department
AB INITIO CALCULATIONS AND VIBRATIONAL SPECTROSCOPIC STUDIES OF 2-CHLORO-6- METHOXYPYRIDINE L.Usha Kumari a, M.Fathima Beegum a, B.Harikumar b, Hema Tresa Varghese c, C.Yohannan Panicker a* a Department
ABC of DFT: Hands-on session 4 Molecular vibrations
 ABC of DFT: Hands-on session 4 Molecular vibrations Tutor: Alexej Bagrets Wann? 29.11.2012, 11:30-13:00 Wo? KIT Campus Süd, Flachbau Physik, Geb. 30.22, Computerpool, Raum FE-6 1 ABC of DFT, Hands-on session
ABC of DFT: Hands-on session 4 Molecular vibrations Tutor: Alexej Bagrets Wann? 29.11.2012, 11:30-13:00 Wo? KIT Campus Süd, Flachbau Physik, Geb. 30.22, Computerpool, Raum FE-6 1 ABC of DFT, Hands-on session
Chemistry 416 Spectroscopy Fall Semester 1993 Dr. Rainer Glaser
 Chemistry 416 Spectroscopy Fall Semester 1993 Dr. Rainer Glaser Second 1-Hour Examination IR/Raman Spectroscopy Monday, October 4, 1993, 8:40-9:30 Name: Question 1 35 Question 2 40 Question 3 25 Total
Chemistry 416 Spectroscopy Fall Semester 1993 Dr. Rainer Glaser Second 1-Hour Examination IR/Raman Spectroscopy Monday, October 4, 1993, 8:40-9:30 Name: Question 1 35 Question 2 40 Question 3 25 Total
Tables for Group Theory
 Tables for Group Theory By P. W. ATKINS, M. S. CHILD, and C. S. G. PHILLIPS This provides the essential tables (character tables, direct products, descent in symmetry and subgroups) required for those
Tables for Group Theory By P. W. ATKINS, M. S. CHILD, and C. S. G. PHILLIPS This provides the essential tables (character tables, direct products, descent in symmetry and subgroups) required for those
QUANTUM CHEMISTRY PROJECT 3: PARTS B AND C
 Chemistry 460 Fall 2017 Dr. Jean M. Standard November 6, 2017 QUANTUM CHEMISTRY PROJECT 3: PARTS B AND C PART B: POTENTIAL CURVE, SPECTROSCOPIC CONSTANTS, AND DISSOCIATION ENERGY OF DIATOMIC HYDROGEN (20
Chemistry 460 Fall 2017 Dr. Jean M. Standard November 6, 2017 QUANTUM CHEMISTRY PROJECT 3: PARTS B AND C PART B: POTENTIAL CURVE, SPECTROSCOPIC CONSTANTS, AND DISSOCIATION ENERGY OF DIATOMIC HYDROGEN (20
Concept of a basis. Based on this treatment we can assign the basis to one of the irreducible representations of the point group.
 Concept of a basis A basis refers to a type of function that is transformed by the symmetry operations of a point group. Examples include the spherical harmonics, vectors, internal coordinates (e..g bonds,
Concept of a basis A basis refers to a type of function that is transformed by the symmetry operations of a point group. Examples include the spherical harmonics, vectors, internal coordinates (e..g bonds,
Exercise 2: Solvating the Structure Before you continue, follow these steps: Setting up Periodic Boundary Conditions
 Exercise 2: Solvating the Structure HyperChem lets you place a molecular system in a periodic box of water molecules to simulate behavior in aqueous solution, as in a biological system. In this exercise,
Exercise 2: Solvating the Structure HyperChem lets you place a molecular system in a periodic box of water molecules to simulate behavior in aqueous solution, as in a biological system. In this exercise,
Problem Set 5 Solutions
 Chemistry 362 Dr Jean M Standard Problem Set 5 Solutions ow many vibrational modes do the following molecules or ions possess? [int: Drawing Lewis structures may be useful in some cases] In all of the
Chemistry 362 Dr Jean M Standard Problem Set 5 Solutions ow many vibrational modes do the following molecules or ions possess? [int: Drawing Lewis structures may be useful in some cases] In all of the
A COMPUTERIZED PROGRAM FOR FINDING THE SYMMETRIES OF THE MOLECULAR NORMAL MODES OF VIBRATION
 Journal of Optoelectronics and Advanced Materials Vol. 5, No. 2, June 23, p. 479-491 A COMPUTERIZED PROGRAM FOR FINDING THE SYMMETRIES OF THE MOLECULAR NORMAL MODES OF VIBRATION Ath. Trutia * University
Journal of Optoelectronics and Advanced Materials Vol. 5, No. 2, June 23, p. 479-491 A COMPUTERIZED PROGRAM FOR FINDING THE SYMMETRIES OF THE MOLECULAR NORMAL MODES OF VIBRATION Ath. Trutia * University
Molecular Symmetry. Symmetry is relevant to: spectroscopy, chirality, polarity, Group Theory, Molecular Orbitals
 Molecular Symmetry Symmetry is relevant to: spectroscopy, chirality, polarity, Group Theory, Molecular Orbitals - A molecule has a symmetry element if it is unchanged by a particular symmetry operation
Molecular Symmetry Symmetry is relevant to: spectroscopy, chirality, polarity, Group Theory, Molecular Orbitals - A molecule has a symmetry element if it is unchanged by a particular symmetry operation
INFRARED ABSORPTION SPECTROSCOPY. References: See relevant sections in undergraduate text. Learn from your instructor how to use the spectrometer.
 INFRARED ABSORPTION SPECTROSCOPY References: See relevant sections in undergraduate text Background: Learn from your instructor how to use the spectrometer. Know definitions of the following and their
INFRARED ABSORPTION SPECTROSCOPY References: See relevant sections in undergraduate text Background: Learn from your instructor how to use the spectrometer. Know definitions of the following and their
2. Infrared spectroscopy
 2. Infrared spectroscopy 2-1Theoretical principles An important tool of the organic chemist is Infrared Spectroscopy, or IR. IR spectra are acquired on a special instrument, called an IR spectrometer.
2. Infrared spectroscopy 2-1Theoretical principles An important tool of the organic chemist is Infrared Spectroscopy, or IR. IR spectra are acquired on a special instrument, called an IR spectrometer.
CHEM-UA 127: Advanced General Chemistry I
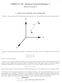 1 CHEM-UA 127: Advanced General Chemistry I Notes for Lecture 6 I MOLECULAR GEOMETRY AND COORDINATES Consider a diatomic molecule AB Imagine fixing this molecule at a very specific spatial location, as
1 CHEM-UA 127: Advanced General Chemistry I Notes for Lecture 6 I MOLECULAR GEOMETRY AND COORDINATES Consider a diatomic molecule AB Imagine fixing this molecule at a very specific spatial location, as
The Theory of HPLC. Quantitative and Qualitative HPLC
 The Theory of HPLC Quantitative and Qualitative HPLC i Wherever you see this symbol, it is important to access the on-line course as there is interactive material that cannot be fully shown in this reference
The Theory of HPLC Quantitative and Qualitative HPLC i Wherever you see this symbol, it is important to access the on-line course as there is interactive material that cannot be fully shown in this reference
