Fanyu Meng Molecular Profiling Proteomics, Merck Research Laboratories, Merck and Co. Inc., Rahway, New Jersey, USA
|
|
|
- Cody Burns
- 5 years ago
- Views:
Transcription
1 Quantitative Analysis of Complex Peptide Mixtures Using FTMS and Differential Mass Spectrometry Fanyu Meng Molecular Profiling Proteomics, Merck Research Laboratories, Merck and Co. Inc., Rahway, New Jersey, USA Matthew C. Wiener, Jeffrey R. Sachs, Chrissina Burns, and Priyanka Verma Applied Computer Science and Mathematics, Merck Research Laboratories, Merck and Co. Inc., Rahway, New Jersey, USA Cloud P. Paweletz, Matthew T. Mazur, Ekaterina G. Deyanova, Nathan A. Yates, and Ronald C. Hendrickson Molecular Profiling Proteomics, Merck Research Laboratories, Merck and Co. Inc., Rahway, New Jersey, USA Label-free LC-MS profiling is a powerful quantitative proteomic method to study relative peptide abundances between two or more biological samples. Here we demonstrate the use of a previously described comparative LC-MS method, differential mass spectrometry (dms), to analyze high-resolution Fourier transform mass spectrometry (FTMS) data for detection and quantification of known peptide differences between two sets of complex mixtures. Six standard peptides were spiked into a processed plasma background at fixed ratios from 1.25:1 to 4:1 to make two sets of samples. The resulting mixtures were analyzed by microcapillary LC-FTMS and dms. dms successfully identified five out of the six peptides as statistically significant differences (p 0.005). In this experiment, the smallest fold change reliably detected by our method was 1.5:1, and the errors of estimated ratios of concentrations were less than 20% for peptides spiked at 1.5:1 to 4:1. We conclude that LC-FTMS coupled with dms is a useful label-free quantitative MS method that can be used to detect subtle yet statistically significant peptide differences in complex protein mixtures, including plasma samples. (J Am Soc Mass Spectrom 2007, 18, ) 2007 American Society for Mass Spectrometry In recent years, quantitative proteomics has been used in a wide range of applications, including biomarker discovery, the analysis of cellular dynamics, and biological pathway studies [1 3]. Many quantitative proteomics techniques use isotopic labeling of proteins, including metabolic labeling [4 6] or chemical labeling strategies [7 9], before mass spectrometry (MS) analysis. Methods that involve direct analyses of complex protein samples without isotope labeling (labelfree MS profiling methods) have also had some success [10 14]. Label-free methods do not require sample mixing or metabolic labeling, meaning that diverse samples, including human tissues or body fluids, can be analyzed. Lastly, label- free methods permit direct comparison of multiple samples across multiple conditions [13], enabling complex experimental designs. Published online October 25, 2006 Address reprint requests to Dr. R. C. Hendrickson, Molecular Profiling Proteomics, Merck Research Laboratories, Merck and Co. Inc., 126 E. Lincoln Ave., RY800-B300, Rahway, NJ 07065, USA. ronald_hendrickson@merck.com These characteristics make label-free mass spectrometry methods appropriate for large scale biomarker studies, e.g., plasma profiling experiments [15]. In label-free LC-MS experiments, a key step is to analyze raw LC-MS data files, and to determine the relative quantities in different samples of the observed ion species. In 2004, we reported a method termed differential mass spectrometry (dms) [14], which can identify statistically significant differences between two or more LC-MS datasets. This method was demonstrated using standard protein digests; a 2-fold change in peptide abundance was found in data collected by an ion trap instrument (LCQ) [14]. In this application note, we describe a well-controlled experiment to evaluate the ability of dms to analyze Fourier transform ion cyclotron resonance mass spectrometry (FT-ICR MS, or FTMS) datasets. Six standard peptides were differentially spiked into processed rat plasma samples at known fixed ratios, and the resulting mixtures were profiled by LC-FTMS to measure the m/z, retention time, and ion intensity of species present in the complex 2007 American Society for Mass Spectrometry. Published by Elsevier Inc. Received June 27, /07/$32.00 Revised September 14, 2006 doi: /j.jasms Accepted September 20, 2006
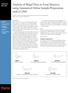
2 J Am Soc Mass Spectrom 2007, 18, DIFFERENTIAL MASS SPECTROMETRY 227 Table 1. Six peptides were spiked into a complex plasma digest at different ratios Peptide (Sigma #) Peptide name Peptide sequence Concentration (fmol/ul) a Targeted ratio (B:A) A9525 Angiotensin-II human DRVYIHPF :1 O3632 Osteocalcin fragments GAPVPYPDPLEPR :1 G3774 Thymopoietin II fragment GEQRKDVYVQLYL :1 C1680 Chromostatin-20 SDEDSDGDRPQASPGLGPGP :1 C5900 Bovine b-casomorphin YPFPGPI :1 C6446 Chromogranin A fragment WSKMDQLAKELTAE :1 a Concentration of each peptide for Mixture A. mixture. dms was used to automatically detect these peptide changes, integrate their selected ion chromatographic (SIC) peaks, and calculate their relative abundance ratios. Based on this experiment, we determined figures of merit for this label-free LC-MS profiling method, including the minimum fold change detectable and the accuracy of the relative ratio calculations. Experimental Preparation of Mixture A (Low Concentration) and Mixture B (High Concentration) Rat plasma sample was purchased from Charles River Laboratory (Wilmington, MA) and N-linked glycopeptides were enriched using the method reported by Zhang et al. [16, 17]. The resulting glycopeptide-enriched plasma pool was used as the constant peptide background in this study. Six commercially available standard peptides (Sigma Aldrich, St. Louis, MO) were spiked into the plasma digests to make Mixture A (lower concentration) and Mixture B (higher concentration) as shown in Table 1. To minimize any variability of sample spiking, three technical replicates were made for Mixture A and Mixture B (designated as samples A1, A2, A3 and B1, B2, B3). LC-FTMS Analysis of Complex Peptide Mixtures An Agilent HP1100 capillary pump (Palo Alto, CA), a Famos autoinjector (Sunnyvale, CA), and a hybrid linear ion trap-ftms Thermo Electron, Bremen, Germany) were used in this study. Samples that were equivalent to 0.1 ul of plasma were loaded onto a trap column (100 um i.d., 2.5 cm, New Objective, Woburn, MA) packed with ProteoPepII C 18 media through the autoinjector. After washing by 100% Solvent A (0.1 M acetic acid in H 2 O) for 5 min at 3 ul/min, the peptides were eluted out through a spraying column (100 um i.d., packed in-house with POROS R2 media) using a 50-min gradient from 100% Solvent A to 90% Solvent B (0.1M acetic acid in 90% acetonitrile and 10% H 2 O) at 1uL/min. Both columns were then equilibrated by 100% Solvent A before next injection. Samples from different conditions were interleaved (e.g., A1, B1, A2, B2, A3 ) to average out any systematic chromatographic and mass spectrometric performance changes over time. Each sample was analyzed four times, generating a total of 24 high-resolution LC-FTMS data files. Key parameters for the mass spectrometer were: AGC 1E 6; maximum injection time 1.0 s; resolving power 50,000; and one FTMS full scan, three data-dependant ion trap MS/MS, and one ion trap full scan were acquired sequentially (the total cycle is 0.9 to 1 s). A nanospray source with no sheath gas was used, and the spraying voltage was 3.0 kv. A commercial data management system, Xcalibur, was used to acquire and analyze LC-FTMS data. This software employs an automatic background reduction algorithm during data acquisition, which reduces most of the background data points before data storage. In this report, signal to background ratios for spiked peptides were all estimated in the following fashion. The signal is calculated by integrating selected ion chromatographic peak area (observed accurate mass, 5 ppm mass range). To estimate the background level, we averaged SIC peak areas for five selected mass ranges (0.5 m/z each) that appeared to be noise at the corresponding elution time. dms Analysis and Ratio Determination Differential mass spectrometry was performed as previously reported [14]. Measured (time, m/z, intensity) points were regularized onto a standard (time, m/z) grid, with spacing of3sintime and 0.01 m/z units [14]. A t-test was performed to produce a single p value describing the extent to which mean intensities differ between the two conditions at each elution time and m/z. Full scan FT data from each of the 12 data files acquired for Mixtures A and B were used to determine the p values. Because real differences are expected to persist in (elution) time, only sufficiently long runs of statistically significant pointwise differences (here, p 0.005) were considered to be differentially expressed features (as defined by the relevant m/z and time range). Here, we required persistence over at least 18 s (seven points including both ends), allowing for a single, less significantly different point within that range. In this study, signals with m/z between 400 and 1500 and retention time between 13 and 50 min, were analyzed;
3 228 MENG ET AL. J Am Soc Mass Spectrom 2007, 18, Figure 1. Base peak chromatograms of four LC-FTMS runs for sample A3. Base peak m/z and retention time are labeled for five selected peaks. These data were acquired at different times over a 39 h period. this range included most of the peptidic ion species. LTQ data were analyzed by dms in a similar fashion, except that p 0.05 and an m/z bin width of 1.0 were used. A peak alignment algorithm was used to compensate for peak retention time shift (retention time shifts of up to 1.5 min were allowed). Briefly, at each binned m/z,we searched the selected ion chromatograms for peaks existing in all samples within a condition. Because chromatography can change slightly from sample to sample, the common peaks will probably not all occur at exactly the same time. For example, a peak at m/z might appear at 26.9 min in the reference sample (which we choose arbitrarily), and at 27.5 min in another sample. This gives rise to a shift that will align the peak with the corresponding peak in a reference sample. In the example above, the time 27.5 would need to be shifted back 0.6 min. We assume that the actual shift required depends only on elution time in a sample and, therefore, is the same for all peaks. Nonetheless, due to estimation issues, different peaks may indicate somewhat different shifts. Smoothing the set of shifts arising from different peaks gives a consensus function to align a particular sample to the reference sample for the appropriate condition. Similarly, peaks existing in the mean data for each condition were used to align the reference samples for different conditions. Applying the within-condition and then the between-condition alignments put all samples into a common time frame for analysis. Because of the high resolving power of FTMS, each peptide can give rise to features representing multiple isotopes at multiple charge states. Based on the mass (as determined by observed charge and m/z) and peak elution time, features believed to arise from the same peptide were automatically grouped. In other controlled experiments, we found that eliminating groups containing only one feature (that is, requiring that at least two isotopes of a differentially expressed peptide to be found for it to be considered for further analysis) substantially reduced the relative frequency of false positives. Therefore, we accepted only groups with at least two features as positive peptide differences in this experiment. For each feature, a relative ratio was automatically determined by dms using mean SIC peak area from each condition. First, dms finds the full peak surrounding the area of statistically significant difference based on the mean SIC from the higher-abundance condition. dms then extracts the peak areas for both conditions and calculates the relative ratio.
4 J Am Soc Mass Spectrom 2007, 18, DIFFERENTIAL MASS SPECTROMETRY 229 Figure 2. Fifteen full scan FT mass spectra summed over retention time range of min to 21.01min. Only peaks between m/z are shown. In Total, 35 isotopic distributions can be observed in this region. Three low-intensity isotopic distributions around m/z 575 are shown in the inset (note change in scale); triangles, squares, and circles represent different sets of isotopic distribution. Results and Discussion Reproducible LC-FTMS Analysis of Complex Peptide Mixtures The processed plasma background samples used in this study contain thousands of different ion species. Four base-peak chromatograms for sample A3 are shown in Figure 1. We used one-dimensional capillary LC to separate this mixture into 50 chromatographic peaks, each containing many coeluting ion species. Figure 2 shows a small region of a typical averaged FTMS mass spectrum (20.65 to min, 15 scans, m/z 450 to 600 is shown), which contains 35 isotopic distributions. Three isotopic distributions can be observed between m/z and m/z (Figure 2, inset). A doubly-charged ion with m/z of (circle), and a triply-charged ion measured at m/z of (square) are readily resolved by FTMS, and another doubly-charged ion with m/z of (triangle) is also observed in this 3.5 m/z window. It is not uncommon to observe multiple components with similar m/z coeluting when analyzing complex mixtures like plasma. Importantly, the high resolving power of FTMS [18] makes it possible to simultaneously detect and quantify these three individual peptides rather than mistakenly integrate the signals as a single species. Key experimental requirements in our label-free LC- FTMS method are reproducible chromatography and mass spectrometry performances. We checked the chromatographic reproducibility of our LC-FTMS platform by monitoring the retention time shift of selected peaks. Figure 1 shows base-peak chromatograms for four LC-MS analyses of sample A3 acquired at different time points over the span of 39 h, and the m/z values and retention time for five randomly selected abundant peaks are labeled. As shown, the overall base peak chromatograms are very similar, but the retention times for these five peaks still have shift of 0.2 min. Based on our experience, larger experiments that involve more LC-MS analyses and run over multiple weeks can give a wider range of retention time shift using the nanoflow LC systems we employ (e.g., up to 1.2 min for 200 samples [19]. We therefore find it necessary to perform retention time correction (i.e., alignment) before performing dms analysis. We have found that retention time shift of less than 1.5 min can be compensated for by our alignment algorithm. We have found that retention time shifts are usually less than 1.5 min. Further, we investigated the reproducibility of MS signal intensity by calculating the coefficients of variance (CVs) for observed ion species. For example, an endogenous peptide from the rodent plasma matrix has a triply charged ion with m/z of (mono) and an observed retention time of 25.0 min. MS/MS analysis reveals the peptide as a haptoglobin peptide, R.ATDLKDWVQETM#AK.N (data not shown). SICs for the monoisotopic peak of this peptide (3, m/z , 5 ppm) for all 24 LC-FTMS runs were manually integrated using peak integration software in Xcalibur, and the CV for peak areas was calculated as 17.5%. Additionally, we calculated CVs for 5000 selected features (excluding those that we believe are from the spikes) using the automatic peak integration tool in dms, and the mean and median CVs were 27 and 25%, respectively. Peak intensity variability is inherent in label-free LC-MS experiments, and this variability can arise from multiple sources, including biochemical sample processing (enzymatic digestion, reductive al-
5 230 MENG ET AL. J Am Soc Mass Spectrom 2007, 18, Figure 3. (a) Selected ion chromatogram for a feature detected by dms, which corresponds to the monoisotopic peak of Angiotensin-II peptide (2 ); solid black line, averaged SIC for B samples; solid red line, averaged SIC for A samples; dashed red and black lines, one standard error about the mean for the A and B samples, respectively. The FTMS scans in the region between dotted lines were statistically different (t-test, p 0.005) as detected by dms. (b) SIC for individual samples. B samples are shown in the upper panel, and A samples are shown in the lower panel. Red portions are the region of detected significant difference. kylation, etc.) and LC-MS profiling (e.g., variation in sample injection or electrospray condition). Had we compared data from an individual LC-MS acquisition for each condition (Mixture A and Mixture B) and used a subtraction method to look for differences in abundance, almost all of the observed ion species would have been found to have different abundances. The majority of these differences could be considered false positives because they arise due to variability in peak intensity measurements rather than from the differentially-spiked peptides in this experiment. dms [14] minimizes these false positives because it considers only peptide changes that are statistically significant between two or more conditions. In other experiments, biological variability among different subjects as well as technical variability in sample preparation and MS measurement must be taken into consideration in the statistical analysis of the MS signals. Detection of Differentially Spiked Peptides by dms To identify the differentially spiked peptides, we performed dms analysis (see the Experimental section) and detected 70 features as statistically significant differences between Mixture A (N 12 samples) and B (N 12 samples) at a pointwise p-value of These features fall into 16 groups, each containing features believed to arise from a single peptide. An exemplary feature for the monoisotopic peak of Angiotensin-II (2 ) is shown in Figure 3a, where red and black solid lines show the averaged SIC for A samples and B samples, respectively. For the mass range between m/z of and , signal intensities in the time bins from 20.3 to 20.9 min (Figure 3a, between blue dotted vertical lines) were statistically different between Mixtures A andb(p 0.005). Therefore, this signal was considered a differentially-expressed feature, and the SICs of this feature for each individual sample are shown in Figure 3b. For the Angiotensin-II peptide, two additional features from 2 charge state (m/z at 20.5 min and m/z at 20.5 min) and one from 1 charge state (m/z at 20.5 min) were found to be differentially expressed between the two mixtures, and all these features were grouped together (Table 2). Overall, dms detected five of the six spiked peptides but missed the thymopoietin II fragment peptide, which was spiked in at a target ratio of 1.25:1 (Table 2). Fourteen of the 16 detected groups of features can be attributed to spiked standards (as they were observed in a separate LC-FTMS analysis of a sample with only the spikes). These fourteen groups can be assigned as spiked peptides (five groups, as shown in Table 2), their adduction forms (such as oxidation or sodium adduction, six groups), or other unknown impurity ion species in the spikes (three groups). The other two groups were from plasma peptide background and they were not expected to be different. These two groups were less statistically significant than the others (ranked 14th and 16th among the 16 groups). In dms analyses performed with a less stringent pointwise p-value threshold of 0.05, allowing more results to be classified as significantly different, features corresponding to the five spiked peptides remain at the top of the list, with relatively low p values. A manual check of the raw data in Xcalibur showed that thymopoietin II was actually present with ratio of the mean intensities nearly equal to 1:1, so it is not surprising that it was not detected even using the less stringent point wise p value. A power analysis showed that with the sample size (N 12) and p value used here, and assuming constant
6 J Am Soc Mass Spectrom 2007, 18, DIFFERENTIAL MASS SPECTROMETRY 231 Table 2. Five spiked peptides detected by dms and their relative ratios m/z Charge Mean area (B) b Mean area (A) b Signal to Ratio background ratio c Ratio (FT) d (LT) e Ratio (FT, Xcal) f Relative error, FT (%) g Chromostatin (2.00:1) a 2.26 / E E E E E E E E E E E E E E E E E E E E 03 Angiotensin-II (2.00:1) 1.91 / E E E E E E E E 03 Chromogranin A (4.00:1) 3.31 / E E E E E E E E E E E E E E E E E E 03 Osteocalcin (1.50:1) 1.51 / E E E E E E E E E E E E Bovine b casomorphin (1.25:1) 1.64 / E E E E E E a Spiked peptide name; numbers in the parenthesis are their respective theoretical ratio. b Mean peak areas are determined automatically by dms using mean SIC peak across each condition (12 A or B samples in this study), in arbitrary units. c Signal to background ratio for selected ion chromatographic peaks for A sample (see Experimental section for details). d Ratio determined by dms (Mean Area B divided by Mean Area A) for each detected feature using FTMS data. Averaged ratio across all isotopes for each peptide and the standard deviation of the measurements are displayed in bold. e Ratio determined by dms using LTQ data. Only four peptides were found by dms as significant differences. f Ratio is manually determined using Xcalibur software on monoisotopic peaks in FTMS data, and averaged ratios are showed in bold (see text for details). g Relatively error of the averaged ratio for each peptide as determined by dms, e.g., error for chromostatin is calculated as ( )/ %, where 2.26 is the experimental ratio, and 2.00 is the theoretical (intended) ratio. CV of 25% regardless of signal intensity, we would expect to detect a 1.5-fold difference about 90% of the time, and a 1.25-fold difference only about 20% of the time. This is consistent with what we observed in this study. Taken together, we estimate that the minimum fold change that can be reliably detected with our current experimental and computational techniques is 1.5:1. For experiments in which it is desired to detect smaller fold changes, we will probably have to increase the sample size (increase N), improve our measurement reproducibility (i.e., lower the CV), or use a less stringent pointwise p value (accepting the risk of more false positives). Relative Quantitation of Differentially Spiked Peptides For each detected feature, dms automatically integrated the mean SIC peaks for Mixtures A and B (Table 2). We found that only SIC peaks above certain abundance level can be clearly defined and accurately integrated. In our LC-FTMS system, this minimum signal requirement is mean peak area of 150,000 arbitrary units. Taking this into consideration, relative abundance ratios were determined for all isotopes of the five detected peptides that meet the minimum signal level requirement (Table 2).
7 232 MENG ET AL. J Am Soc Mass Spectrom 2007, 18, For the five detected peptide differences, peak area ratios were consistent for different isotopes and charge states of each peptide (Table 2). For example, the calculated ratios for different isotopes of Chromostatin peptide ranged from 2.08 to 2.42, and the overall unweighted averaged ratio was 2.26, with standard deviation of 0.10 (CV 5%). In terms of the absolute value, the bovine b-casomorphin peptide (target ratio 1.25:1) had the biggest relative error at 31%, while all the other peptides had relative errors of less than 20%. Along with the fact that thymopoietin II fragment, also spiked at 1.25:1, was not detected as differentially expressed, and the error of 31% for bovine b-casomorphin suggests that the minimum detectable fold change in our LC-FTMS and dms platform was 1.5:1. Overall, our method can measure relative ratios with less than 20% errors for these small and subtle (1.5:1 to 4:1) peptide differences in the complex matrix. For comparison purposes, we also analyzed the acquired low-resolution LTQ data. Six charge states arising from four of the spiked peptides were found as statistically significant differences. As in the FT data, the expected results, along with a few adduction forms, were the highest ranked results. The minimum detectable fold change was higher using LTQ data than using FT data: osteocalcin (targeted ratio of 1.50:1) was not found by dms using the LTQ data. For the spiked-in peptides that were detected, the ratios calculated using LTQ data were similar to those calculated using FT data (column Ratio LT in Table 2). However, the FTMS data shows isotopic distributions and charge state information. In more complex mixtures, we would expect a larger number of overlapping distributions like the ones in Figure 2, inset, where using the higher-resolution data would be necessary for accurate quantitation. This study was designed to test our ability to detect and accurately measure small differences with ratios ranging from 1.25:1 to 4:1. In this case, it is reasonable to assume comparable or linear response for both the lower and higher abundant species. However, it is important to note that when quantifying large differences with fold changes greater that 10:1, the mass spectrometer response may deviate from linearity and introduce systematic errors into the measured ratio. Errors of Ratio Measurement Introduced by dms Peak Integration Many factors can contribute to errors in ratio determination, including sample preparation (such as pipetting errors), LC/MS profiling, and dms peak integration. We focus our attention mainly on the error introduced by dms peak integration algorithm. To evaluate these errors, we manually integrated SIC peak areas for all monoisotopic peaks of peptide spikes. The relative abundance ratios were calculated and listed in Table 2 (Column Ratio Xcal ). These ratios represent the ratio raw LC-MS data could achieve, and comparison between averaged ratio for each peptide (Table 2, Column Ratio Xcal, bold) and its ratio determined by dms revealed discrepancies from as low as 1% up to 10% (i.e., for chromogranin A). This suggests that up to 10% ratio determination error can come from the automated peak integration of dms. These results suggest that improvements can be achieved by optimizing the automatic chromatographic peak area integration in dms (work in progress). Conclusions We have demonstrated that automatic analysis of LC- FTMS data by dms is a powerful tool to analyze complex protein mixtures and detect small peptide changes between two sets of complex samples. dms considers not only differences between conditions, but also accounts for variability within each condition, so that statistically significant differences can be efficiently found and studied. The minimum detectable fold change using our current experimental and computational systems is 1.5:1, and the error of ratio measurements for these subtle changes (1.5:1 to 4:1) at relatively low signal strength (signal to background ratio 2-44) was less than 20%. Although we designed this experiment to test the ability of dms to analyze relatively small signals, similar minimum fold change detectable and quantification accuracy have been observed for ion species with high signal levels (signal to background of up to 250) [20]. Therefore, this method is applicable for abundant signals as well as less abundant ones, which is important for analyses of complex protein samples with large dynamic ranges, e.g., plasma samples. Unlike many other quantitative proteomic analysis methods [9, 10], dms finds signals that are statistically different without relying on the identification of their amino acid sequences. In our laboratory, we find this workflow particularly useful for experiments designed to detect quantitative changes in biological systems because it separates protein identification efforts from the detection of differences. Indeed, in some cases, we have performed experiments where we detect thousands of peptide ions in the mass spectrometer but observed no significant quantitative change between conditions. In these cases we can quickly conclude that our comparative LC-MS profiling does not currently have sufficient sensitivity to detect the quantitative changes that may be present in the experiment, our platform variability is too large to detect the fold change that is present, or there are no statistically significant quantitative changes in the biological experiment. In such cases, we no longer need to perform protein ID, and efforts are then focused to redesign the experiment or explore an alternate experimental system or hypothesis. For features found to be differentially expressed, tandem mass spectra for features can be acquired in a targeted fashion by running the samples again. Because only selected peptides are targeted, the MS/MS data acquisition and data analysis efficiency will be im-
8 J Am Soc Mass Spectrom 2007, 18, DIFFERENTIAL MASS SPECTROMETRY 233 proved. Many relatively slower fragmentation techniques such as electron capture dissociation (ECD) [21] and infrared multiphoton dissociation (IRMPD) [22] can be considered, and multiple MS/MS scans can be averaged. These techniques can improve MS/MS data quality, which may lead to more confident and efficient protein identifications. Lastly, we used peptides in this study, but it is expected that these techniques would be equally applicable to small molecules such as human metabolites. This label-free relative quantitation method could play an important role in many proteomics, metabolomics, and drug metabolism studies. Acknowledgments The authors thank Dr. Haihong Zhou for helpful comments on the manuscript and Drs. Alan Sachs and Stephen Friend for their support. References 1. Aebersold, R.; Mann, M. Mass Spectrometry-Based Proteomics. Nature 2003, 422(6928), Ideker, T.; Thorsson, V.; Ranish, J. A.; Christmas, R.; Buhler, J.; Eng, J. K.; Bumgarner, R.; Goodlett, D. R.; Aebersold, R.; Hood, L. Integrated Genomic and Proteomic Analyses of a Systematically Perturbed Metabolic Network. Science 2001, 292(5518), Kratchmarova, I.; Blagoev, B.; Haack-Sorensen, M.; Kassem, M.; Mann, M. Mechanism of Divergent Growth Factor Effects in Mesenchymal Stem Cell Differentiation. Science 2005, 308(5727), Ballif, B. A.; Roux, P. P.; Gerber, S. A.; MacKeigan, J. P.; Blenis, J.; Gygi, S. P. Quantitative Phosphorylation Profiling of the ERK/p90 Ribosomal S6 Kinase-Signaling Cassette and Its Targets, the Tuberous Sclerosis Tumor Suppressors. Proc. Natl. Acad. Sci. U.S.A. 2005, 102(3), Krijgsveld, J.; Ketting, R. F.; Mahmoudi, T.; Johansen, J.; Artal-Sanz, M.; Verrijzer, C. P.; Plasterk, R. H.; Heck, A. J. Metabolic Labeling of C. elegans and D. melanogaster for Quantitative Proteomics. Nat. Biotechnol. 2003, 21(8), Wu, C. C.; MacCoss, M. J.; Howell, K. E.; Matthews, D. E.; Yates, J. R. III. Metabolic Labeling of Mammalian Organisms with Stable Isotopes for Quantitative Proteomic Analysis. Anal. Chem. 2004, 76(17), Gygi, S. P.; Rist, B.; Gerber, S. A.; Turecek, F.; Gelb, M. H.; Aebersold, R. Quantitative Analysis of Complex Protein Mixtures Using Isotope- Coded Affinity Tags. Nat. Biotechnol. 1999, 17(10), Ross, P. L.; Huang, Y. N.; Marchese, J. N.; Williamson, B.; Parker, K.; Hattan, S.; Khainovski, N.; Pillai, S.; Dey, S.; Daniels, S.; Purkayastha, S.; Juhasz, P.; Martin, S.; Bartlet-Jones, M.; He, F.; Jacobson, A.; Pappin, D. J. Multiplexed Protein Quantitation in Saccharomyces cerevisiae Using Amine-Reactive Isobaric Tagging Reagents. Mol. Cell. Proteom. 2004, 3(12), Yao, X.; Freas, A.; Ramirez, J.; Demirev, P. A.; Fenselau, C. Proteolytic 18O Labeling for Comparative Proteomics: Model Studies with Two Serotypes of Adenovirus. Anal. Chem. 2001, 73(13), Bondarenko, P. V.; Chelius, D.; Shaler, T. A. Identification and Relative Quantitation of Protein Mixtures by Enzymatic Digestion Followed by Capillary Reversed-Phase Liquid Chromatography-Tandem Mass Spectrometry. Anal. Chem. 2002, 74(18), Chelius, D.; Bondarenko, P. V. Quantitative Profiling of Proteins in Complex Mixtures Using Liquid Chromatography and Mass Spectrometry. J. Proteome Res. 2002, 1(4), Old, W. M.; Meyer-Arendt, K.; Aveline-Wolf, L.; Pierce, K. G.; Mendoza, A.; Sevinsky, J. R.; Resing, K. A.; Ahn, N. G. Comparison of Label-Free Methods for Quantifying Human Proteins by Shotgun Proteomics. Mol. Cell. Proteom. 2005, 4(10), Wang, W.; Zhou, H.; Lin, H.; Roy, S.; Shaler, T. A.; Hill, L. R.; Norton, S.; Kumar, P.; Anderle, M.; Becker, C. H. Quantification of Proteins and Metabolites by Mass Spectrometry Without Isotopic Labeling or Spiked Standards. Anal. Chem. 2003, 75(18), Wiener, M. C.; Sachs, J. R.; Deyanova, E. G.; Yates, N. A. Differential Mass Spectrometry: A Label-Free LC-MS Method for Finding Significant Differences in Complex Peptide and Protein Mixtures. Anal. Chem. 2004, 76(20), Anderson, N. L.; Anderson, N. G. The Human Plasma Proteome: History, Character, and Diagnostic Prospects. Mol. Cell. Proteom. 2002, 1(11), Zhang, H.; Li, X. J.; Martin, D. B.; Aebersold, R. Identification and Quantification of N-Linked Glycoproteins Using Hydrazide Chemistry, Stable Isotope Labeling, and Mass Spectrometry. Nat. Biotechnol. 2003, 21(6), Zhang, H.; Yi, E. C.; Li, X. J.; Mallick, P.; Kelly-Spratt, K. S.; Masselon, C. D.; Camp, D. G.; Smith, R. D.; Kemp, C. J.; Aebersold, R. High Throughput Quantitative Analysis of Serum Proteins Using Glycopeptide Capture and Liquid Chromatography Mass Spectrometry. Mol. Cell. Proteom. 2005, 4(2), Marshall, A. G.; Hendrickson, C. L.; Jackson, G. S. Fourier Transform Ion Cyclotron Resonance Mass Spectrometry: A Primer. Mass Spectrom. Rev. 1998, 17(1), Deyanova E. G.; Zhao, X.; Meng, F.; Mazur, M. T.; Paweletz, C.; Lee, A. Y. H.; Mistry, J.; Wiener, M. C.; Sachs, J. R.; Yates, N. A.; Hendrickson, R. C. Analysis of an FTMS Based Platform for Plasma Profiling and Comparative Proteomics. Proceedings of the 53rd ASMS Conference on Mass Spectrometry and Allied Topics; San Antonio, TX, June Meng, F.; Wiener, M. C.; Sachs, J. R.; Zhao, X.; Paweletz, C.; Lee, A. Y. H.; Deyanova, E. G.; Mistry, J.; Mazur, M. T.; Yates, N. A.; Hendrickson, R. C. Quantitative Protein Profiling Using FTMS and Differential Mass Spectrometry. Proceedings of the 53rd ASMS Conference on Mass Spectrometry and Applied Topics; San Antonio, TX, June Little, D. P.; Speir, J. P.; Senko, M. W.; O Connor, P. B.; McLafferty, F. W. Infrared Multiphoton Dissociation of Large Multiply Charged Ions for Biomolecule Sequencing. Anal. Chem. 1994, 66(18), Zubarev, R. A.; Horn, D. M.; Fridriksson, E. K.; Kelleher, N. L.; Kruger, N. A.; Lewis, M. A.; Carpenter, B. K.; McLafferty, F. W. Electron Capture Dissociation for Structural Characterization of Multiply Charged Protein Cations. Anal. Chem. 2000, 72(3),
Mass spectrometry-based proteomics has become
 FOCUS: THE ORBITRAP Computational Principles of Determining and Improving Mass Precision and Accuracy for Proteome Measurements in an Orbitrap Jürgen Cox and Matthias Mann Proteomics and Signal Transduction,
FOCUS: THE ORBITRAP Computational Principles of Determining and Improving Mass Precision and Accuracy for Proteome Measurements in an Orbitrap Jürgen Cox and Matthias Mann Proteomics and Signal Transduction,
Quantitation of a target protein in crude samples using targeted peptide quantification by Mass Spectrometry
 Quantitation of a target protein in crude samples using targeted peptide quantification by Mass Spectrometry Jon Hao, Rong Ye, and Mason Tao Poochon Scientific, Frederick, Maryland 21701 Abstract Background:
Quantitation of a target protein in crude samples using targeted peptide quantification by Mass Spectrometry Jon Hao, Rong Ye, and Mason Tao Poochon Scientific, Frederick, Maryland 21701 Abstract Background:
High-Throughput Protein Quantitation Using Multiple Reaction Monitoring
 High-Throughput Protein Quantitation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine Miller, Joe Roark, Norton Kitagawa and Keith Waddell Agilent Technologies, Inc. Santa
High-Throughput Protein Quantitation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine Miller, Joe Roark, Norton Kitagawa and Keith Waddell Agilent Technologies, Inc. Santa
Electrospray ionization mass spectrometry (ESI-
 Automated Charge State Determination of Complex Isotope-Resolved Mass Spectra by Peak-Target Fourier Transform Li Chen a and Yee Leng Yap b a Bioinformatics Institute, 30 Biopolis Street, Singapore b Davos
Automated Charge State Determination of Complex Isotope-Resolved Mass Spectra by Peak-Target Fourier Transform Li Chen a and Yee Leng Yap b a Bioinformatics Institute, 30 Biopolis Street, Singapore b Davos
Modeling Mass Spectrometry-Based Protein Analysis
 Chapter 8 Jan Eriksson and David Fenyö Abstract The success of mass spectrometry based proteomics depends on efficient methods for data analysis. These methods require a detailed understanding of the information
Chapter 8 Jan Eriksson and David Fenyö Abstract The success of mass spectrometry based proteomics depends on efficient methods for data analysis. These methods require a detailed understanding of the information
Improved 6- Plex TMT Quantification Throughput Using a Linear Ion Trap HCD MS 3 Scan Jane M. Liu, 1,2 * Michael J. Sweredoski, 2 Sonja Hess 2 *
 Improved 6- Plex TMT Quantification Throughput Using a Linear Ion Trap HCD MS 3 Scan Jane M. Liu, 1,2 * Michael J. Sweredoski, 2 Sonja Hess 2 * 1 Department of Chemistry, Pomona College, Claremont, California
Improved 6- Plex TMT Quantification Throughput Using a Linear Ion Trap HCD MS 3 Scan Jane M. Liu, 1,2 * Michael J. Sweredoski, 2 Sonja Hess 2 * 1 Department of Chemistry, Pomona College, Claremont, California
Identification of proteins by enzyme digestion, mass
 Method for Screening Peptide Fragment Ion Mass Spectra Prior to Database Searching Roger E. Moore, Mary K. Young, and Terry D. Lee Beckman Research Institute of the City of Hope, Duarte, California, USA
Method for Screening Peptide Fragment Ion Mass Spectra Prior to Database Searching Roger E. Moore, Mary K. Young, and Terry D. Lee Beckman Research Institute of the City of Hope, Duarte, California, USA
Protein Quantitation II: Multiple Reaction Monitoring. Kelly Ruggles New York University
 Protein Quantitation II: Multiple Reaction Monitoring Kelly Ruggles kelly@fenyolab.org New York University Traditional Affinity-based proteomics Use antibodies to quantify proteins Western Blot RPPA Immunohistochemistry
Protein Quantitation II: Multiple Reaction Monitoring Kelly Ruggles kelly@fenyolab.org New York University Traditional Affinity-based proteomics Use antibodies to quantify proteins Western Blot RPPA Immunohistochemistry
Protein Quantitation II: Multiple Reaction Monitoring. Kelly Ruggles New York University
 Protein Quantitation II: Multiple Reaction Monitoring Kelly Ruggles kelly@fenyolab.org New York University Traditional Affinity-based proteomics Use antibodies to quantify proteins Western Blot Immunohistochemistry
Protein Quantitation II: Multiple Reaction Monitoring Kelly Ruggles kelly@fenyolab.org New York University Traditional Affinity-based proteomics Use antibodies to quantify proteins Western Blot Immunohistochemistry
Tandem mass spectra were extracted from the Xcalibur data system format. (.RAW) and charge state assignment was performed using in house software
 Supplementary Methods Software Interpretation of Tandem mass spectra Tandem mass spectra were extracted from the Xcalibur data system format (.RAW) and charge state assignment was performed using in house
Supplementary Methods Software Interpretation of Tandem mass spectra Tandem mass spectra were extracted from the Xcalibur data system format (.RAW) and charge state assignment was performed using in house
Agilent MassHunter Profinder: Solving the Challenge of Isotopologue Extraction for Qualitative Flux Analysis
 Agilent MassHunter Profinder: Solving the Challenge of Isotopologue Extraction for Qualitative Flux Analysis Technical Overview Introduction Metabolomics studies measure the relative abundance of metabolites
Agilent MassHunter Profinder: Solving the Challenge of Isotopologue Extraction for Qualitative Flux Analysis Technical Overview Introduction Metabolomics studies measure the relative abundance of metabolites
HR/AM Targeted Peptide Quantification on a Q Exactive MS: A Unique Combination of High Selectivity, High Sensitivity, and High Throughput
 HR/AM Targeted Peptide Quantification on a Q Exactive MS: A Unique Combination of High Selectivity, High Sensitivity, and High Throughput Yi Zhang 1, Zhiqi Hao 1, Markus Kellmann 2 and Andreas FR. Huhmer
HR/AM Targeted Peptide Quantification on a Q Exactive MS: A Unique Combination of High Selectivity, High Sensitivity, and High Throughput Yi Zhang 1, Zhiqi Hao 1, Markus Kellmann 2 and Andreas FR. Huhmer
Designed for Accuracy. Innovation with Integrity. High resolution quantitative proteomics LC-MS
 Designed for Accuracy High resolution quantitative proteomics Innovation with Integrity LC-MS Setting New Standards in Accuracy The development of mass spectrometry based proteomics approaches has dramatically
Designed for Accuracy High resolution quantitative proteomics Innovation with Integrity LC-MS Setting New Standards in Accuracy The development of mass spectrometry based proteomics approaches has dramatically
High-Field Orbitrap Creating new possibilities
 Thermo Scientific Orbitrap Elite Hybrid Mass Spectrometer High-Field Orbitrap Creating new possibilities Ultrahigh resolution Faster scanning Higher sensitivity Complementary fragmentation The highest
Thermo Scientific Orbitrap Elite Hybrid Mass Spectrometer High-Field Orbitrap Creating new possibilities Ultrahigh resolution Faster scanning Higher sensitivity Complementary fragmentation The highest
A Software Suite for the Generation and Comparison of Peptide Arrays from Sets. of Data Collected by Liquid Chromatography-Mass Spectrometry
 MCP Papers in Press. Published on July 26, 2005 as Manuscript M500141-MCP200 A Software Suite for the Generation and Comparison of Peptide Arrays from Sets of Data Collected by Liquid Chromatography-Mass
MCP Papers in Press. Published on July 26, 2005 as Manuscript M500141-MCP200 A Software Suite for the Generation and Comparison of Peptide Arrays from Sets of Data Collected by Liquid Chromatography-Mass
Analysis of Labeled and Non-Labeled Proteomic Data Using Progenesis QI for Proteomics
 Analysis of Labeled and Non-Labeled Proteomic Data Using Progenesis QI for Proteomics Lee Gethings, Gushinder Atwal, Martin Palmer, Chris Hughes, Hans Vissers, and James Langridge Waters Corporation, Wilmslow,
Analysis of Labeled and Non-Labeled Proteomic Data Using Progenesis QI for Proteomics Lee Gethings, Gushinder Atwal, Martin Palmer, Chris Hughes, Hans Vissers, and James Langridge Waters Corporation, Wilmslow,
Towards the Prediction of Protein Abundance from Tandem Mass Spectrometry Data
 Towards the Prediction of Protein Abundance from Tandem Mass Spectrometry Data Anthony J Bonner Han Liu Abstract This paper addresses a central problem of Proteomics: estimating the amounts of each of
Towards the Prediction of Protein Abundance from Tandem Mass Spectrometry Data Anthony J Bonner Han Liu Abstract This paper addresses a central problem of Proteomics: estimating the amounts of each of
Data pre-processing in liquid chromatography mass spectrometry-based proteomics
 BIOINFORMATICS ORIGINAL PAPER Vol. 21 no. 21 25, pages 454 459 doi:1.193/bioinformatics/bti66 Data and text mining Data pre-processing in liquid chromatography mass spectrometry-based proteomics Xiang
BIOINFORMATICS ORIGINAL PAPER Vol. 21 no. 21 25, pages 454 459 doi:1.193/bioinformatics/bti66 Data and text mining Data pre-processing in liquid chromatography mass spectrometry-based proteomics Xiang
MasSPIKE (Mass SPectrum Interpretation and Kernel Extraction) for Biological Samples p.1
 MasSPIKE (Mass SPectrum Interpretation and Kernel Extraction) for Biological Samples Parminder Kaur Bogdan Budnik, Konstantin Aizikov and Peter B. O Connor, Department of Electrical and Computer Engineering,
MasSPIKE (Mass SPectrum Interpretation and Kernel Extraction) for Biological Samples Parminder Kaur Bogdan Budnik, Konstantin Aizikov and Peter B. O Connor, Department of Electrical and Computer Engineering,
Improved Throughput and Reproducibility for Targeted Protein Quantification Using a New High-Performance Triple Quadrupole Mass Spectrometer
 Improved Throughput and Reproducibility for Targeted Protein Quantification Using a New High-Performance Triple Quadrupole Mass Spectrometer Reiko Kiyonami, Mary Blackburn, Andreas FR Hühme: Thermo Fisher
Improved Throughput and Reproducibility for Targeted Protein Quantification Using a New High-Performance Triple Quadrupole Mass Spectrometer Reiko Kiyonami, Mary Blackburn, Andreas FR Hühme: Thermo Fisher
Chemical Labeling Strategy for Generation of Internal Standards for Targeted Quantitative Proteomics
 Chemical Labeling Strategy for Generation of Internal Standards for Targeted Quantitative Proteomics mtraq Reagents Triplex Christie Hunter, Brian Williamson, Marjorie Minkoff AB SCIEX, USA The utility
Chemical Labeling Strategy for Generation of Internal Standards for Targeted Quantitative Proteomics mtraq Reagents Triplex Christie Hunter, Brian Williamson, Marjorie Minkoff AB SCIEX, USA The utility
Analytical determination of testosterone in human serum using an Agilent Ultivo Triple Quadrupole LC/MS
 Application Note Clinical Research Analytical determination of testosterone in human serum using an Agilent Ultivo Triple Quadrupole LC/MS Authors Yanan Yang 1, Victor Mandragon 2, and Peter Stone 1 1
Application Note Clinical Research Analytical determination of testosterone in human serum using an Agilent Ultivo Triple Quadrupole LC/MS Authors Yanan Yang 1, Victor Mandragon 2, and Peter Stone 1 1
Methods for proteome analysis of obesity (Adipose tissue)
 Methods for proteome analysis of obesity (Adipose tissue) I. Sample preparation and liquid chromatography-tandem mass spectrometric analysis Instruments, softwares, and materials AB SCIEX Triple TOF 5600
Methods for proteome analysis of obesity (Adipose tissue) I. Sample preparation and liquid chromatography-tandem mass spectrometric analysis Instruments, softwares, and materials AB SCIEX Triple TOF 5600
Guide to Peptide Quantitation. Agilent clinical research
 Guide to Peptide Quantitation Agilent clinical research Peptide Quantitation for the Clinical Research Laboratory Peptide quantitation is rapidly growing in clinical research as scientists are translating
Guide to Peptide Quantitation Agilent clinical research Peptide Quantitation for the Clinical Research Laboratory Peptide quantitation is rapidly growing in clinical research as scientists are translating
Thermo Scientific LTQ Orbitrap Velos Hybrid FT Mass Spectrometer
 IET International Equipment Trading Ltd. www.ietltd.com Proudly serving laboratories worldwide since 1979 CALL +847.913.0777 for Refurbished & Certified Lab Equipment Thermo Scientific LTQ Orbitrap Velos
IET International Equipment Trading Ltd. www.ietltd.com Proudly serving laboratories worldwide since 1979 CALL +847.913.0777 for Refurbished & Certified Lab Equipment Thermo Scientific LTQ Orbitrap Velos
Peptide Targeted Quantification By High Resolution Mass Spectrometry A Paradigm Shift? Zhiqi Hao Thermo Fisher Scientific San Jose, CA
 Peptide Targeted Quantification By High Resolution Mass Spectrometry A Paradigm Shift? Zhiqi Hao Thermo Fisher Scientific San Jose, CA Proteomics is Turning Quantitative Hmmm.. Which ones are my targets?
Peptide Targeted Quantification By High Resolution Mass Spectrometry A Paradigm Shift? Zhiqi Hao Thermo Fisher Scientific San Jose, CA Proteomics is Turning Quantitative Hmmm.. Which ones are my targets?
The Power of LC MALDI: Identification of Proteins by LC MALDI MS/MS Using the Applied Biosystems 4700 Proteomics Analyzer with TOF/TOF Optics
 APPLICATION NOTE TOF MS The Power of LC MALDI: Identification of Proteins by LC MALDI MS/MS Using the Applied Biosystems 4700 Proteomics Analyzer with TOF/TOF Optics Purpose The Applied Biosystems 4700
APPLICATION NOTE TOF MS The Power of LC MALDI: Identification of Proteins by LC MALDI MS/MS Using the Applied Biosystems 4700 Proteomics Analyzer with TOF/TOF Optics Purpose The Applied Biosystems 4700
WADA Technical Document TD2003IDCR
 IDENTIFICATION CRITERIA FOR QUALITATIVE ASSAYS INCORPORATING CHROMATOGRAPHY AND MASS SPECTROMETRY The appropriate analytical characteristics must be documented for a particular assay. The Laboratory must
IDENTIFICATION CRITERIA FOR QUALITATIVE ASSAYS INCORPORATING CHROMATOGRAPHY AND MASS SPECTROMETRY The appropriate analytical characteristics must be documented for a particular assay. The Laboratory must
Aplicació de la proteòmica a la cerca de Biomarcadors proteics Barcelona, 08 de Juny 2010
 Aplicació de la proteòmica a la cerca de Biomarcadors proteics Barcelona, 8 de Juny 21 Eliandre de Oliveira Plataforma de Proteòmica Parc Científic de Barcelona Protein Chemistry Proteomics Hypothesis-free
Aplicació de la proteòmica a la cerca de Biomarcadors proteics Barcelona, 8 de Juny 21 Eliandre de Oliveira Plataforma de Proteòmica Parc Científic de Barcelona Protein Chemistry Proteomics Hypothesis-free
Rapid and Accurate Forensics Analysis using High Resolution All Ions MS/MS
 Rapid and Accurate Forensics Analysis using High Resolution All Ions MS/MS Application Note Forensic Toxicology Authors Martin Josefsson, and Markus Roman National Board of Forensic Medicine Linköping,
Rapid and Accurate Forensics Analysis using High Resolution All Ions MS/MS Application Note Forensic Toxicology Authors Martin Josefsson, and Markus Roman National Board of Forensic Medicine Linköping,
Accelerating the Metabolite Identification Process Using High Resolution Q-TOF Data and Mass-MetaSite Software
 Accelerating the Metabolite Identification Process Using High Resolution Q-TOF Data and Mass-MetaSite Software Application ote Drug discovery and development: Metabolite Identifi cation Authors Yuqin Dai,
Accelerating the Metabolite Identification Process Using High Resolution Q-TOF Data and Mass-MetaSite Software Application ote Drug discovery and development: Metabolite Identifi cation Authors Yuqin Dai,
Analysis of Peptide MS/MS Spectra from Large-Scale Proteomics Experiments Using Spectrum Libraries
 Anal. Chem. 2006, 78, 5678-5684 Analysis of Peptide MS/MS Spectra from Large-Scale Proteomics Experiments Using Spectrum Libraries Barbara E. Frewen, Gennifer E. Merrihew, Christine C. Wu, William Stafford
Anal. Chem. 2006, 78, 5678-5684 Analysis of Peptide MS/MS Spectra from Large-Scale Proteomics Experiments Using Spectrum Libraries Barbara E. Frewen, Gennifer E. Merrihew, Christine C. Wu, William Stafford
Choosing the metabolomics platform
 GBS 748 Choosing the metabolomics platform Stephen Barnes, PhD 4 7117; sbarnes@uab.edu So, I have my samples what s next? You ve collected your samples and you may have extracted them Protein precipitation
GBS 748 Choosing the metabolomics platform Stephen Barnes, PhD 4 7117; sbarnes@uab.edu So, I have my samples what s next? You ve collected your samples and you may have extracted them Protein precipitation
LECTURE-11. Hybrid MS Configurations HANDOUT. As discussed in our previous lecture, mass spectrometry is by far the most versatile
 LECTURE-11 Hybrid MS Configurations HANDOUT PREAMBLE As discussed in our previous lecture, mass spectrometry is by far the most versatile technique used in proteomics. We had also discussed some of the
LECTURE-11 Hybrid MS Configurations HANDOUT PREAMBLE As discussed in our previous lecture, mass spectrometry is by far the most versatile technique used in proteomics. We had also discussed some of the
Mass Spectrometry. Hyphenated Techniques GC-MS LC-MS and MS-MS
 Mass Spectrometry Hyphenated Techniques GC-MS LC-MS and MS-MS Reasons for Using Chromatography with MS Mixture analysis by MS alone is difficult Fragmentation from ionization (EI or CI) Fragments from
Mass Spectrometry Hyphenated Techniques GC-MS LC-MS and MS-MS Reasons for Using Chromatography with MS Mixture analysis by MS alone is difficult Fragmentation from ionization (EI or CI) Fragments from
Relative quantification using TMT11plex on a modified Q Exactive HF mass spectrometer
 POSTER NOTE 6558 Relative quantification using TMT11plex on a modified mass spectrometer Authors Tabiwang N. Arrey, 1 Rosa Viner, 2 Ryan D. Bomgarden, 3 Eugen Damoc, 1 Markus Kellmann, 1 Thomas Moehring,
POSTER NOTE 6558 Relative quantification using TMT11plex on a modified mass spectrometer Authors Tabiwang N. Arrey, 1 Rosa Viner, 2 Ryan D. Bomgarden, 3 Eugen Damoc, 1 Markus Kellmann, 1 Thomas Moehring,
Making Sense of Differences in LCMS Data: Integrated Tools
 Making Sense of Differences in LCMS Data: Integrated Tools David A. Weil Agilent Technologies MassHunter Overview Page 1 March 2008 How Clean is our Water?... Page 2 Chemical Residue Analysis.... From
Making Sense of Differences in LCMS Data: Integrated Tools David A. Weil Agilent Technologies MassHunter Overview Page 1 March 2008 How Clean is our Water?... Page 2 Chemical Residue Analysis.... From
MassHunter TOF/QTOF Users Meeting
 MassHunter TOF/QTOF Users Meeting 1 Qualitative Analysis Workflows Workflows in Qualitative Analysis allow the user to only see and work with the areas and dialog boxes they need for their specific tasks
MassHunter TOF/QTOF Users Meeting 1 Qualitative Analysis Workflows Workflows in Qualitative Analysis allow the user to only see and work with the areas and dialog boxes they need for their specific tasks
HR/AM Targeted Peptide Quantitation on a Q Exactive MS: A Unique Combination of High Selectivity, Sensitivity and Throughput
 Application Note: 554 HR/AM Targeted Peptide Quantitation on a Q Exactive MS: A Unique Combination of High Selectivity, Sensitivity and Throughput Yi Zhang 1, Zhiqi Hao 1, Markus Kellmann 2 and Andreas
Application Note: 554 HR/AM Targeted Peptide Quantitation on a Q Exactive MS: A Unique Combination of High Selectivity, Sensitivity and Throughput Yi Zhang 1, Zhiqi Hao 1, Markus Kellmann 2 and Andreas
NPTEL VIDEO COURSE PROTEOMICS PROF. SANJEEVA SRIVASTAVA
 LECTURE-25 Quantitative proteomics: itraq and TMT TRANSCRIPT Welcome to the proteomics course. Today we will talk about quantitative proteomics and discuss about itraq and TMT techniques. The quantitative
LECTURE-25 Quantitative proteomics: itraq and TMT TRANSCRIPT Welcome to the proteomics course. Today we will talk about quantitative proteomics and discuss about itraq and TMT techniques. The quantitative
Translational Biomarker Core
 Translational Biomarker Core Instrumentation Thermo Scientific TSQ Quantum Triple Quadrupole Mass Spectrometers. There are two TSQ Quantum Ultra AM instruments available in the TBC. The TSQ Quantum Ultra
Translational Biomarker Core Instrumentation Thermo Scientific TSQ Quantum Triple Quadrupole Mass Spectrometers. There are two TSQ Quantum Ultra AM instruments available in the TBC. The TSQ Quantum Ultra
Isotopic-Labeling and Mass Spectrometry-Based Quantitative Proteomics
 Isotopic-Labeling and Mass Spectrometry-Based Quantitative Proteomics Xiao-jun Li, Ph.D. Current address: Homestead Clinical Day 4 October 19, 2006 Protein Quantification LC-MS/MS Data XLink mzxml file
Isotopic-Labeling and Mass Spectrometry-Based Quantitative Proteomics Xiao-jun Li, Ph.D. Current address: Homestead Clinical Day 4 October 19, 2006 Protein Quantification LC-MS/MS Data XLink mzxml file
Targeted protein quantification
 Targeted Quantitative Proteomics Targeted protein quantification with high-resolution, accurate-mass MS Highly selective Very sensitive Complex samples HR/AM A more complete quantitative proteomics picture
Targeted Quantitative Proteomics Targeted protein quantification with high-resolution, accurate-mass MS Highly selective Very sensitive Complex samples HR/AM A more complete quantitative proteomics picture
Atomic masses. Atomic masses of elements. Atomic masses of isotopes. Nominal and exact atomic masses. Example: CO, N 2 ja C 2 H 4
 High-Resolution Mass spectrometry (HR-MS, HRAM-MS) (FT mass spectrometry) MS that enables identifying elemental compositions (empirical formulas) from accurate m/z data 9.05.2017 1 Atomic masses (atomic
High-Resolution Mass spectrometry (HR-MS, HRAM-MS) (FT mass spectrometry) MS that enables identifying elemental compositions (empirical formulas) from accurate m/z data 9.05.2017 1 Atomic masses (atomic
6 x 5 Ways to Ensure Your LC-MS/MS is Healthy
 6 x 5 Ways to Ensure Your LC-MS/MS is Healthy (Also known as - Tracking Performance with the 6 x 5 LC-MS/MS Peptide Reference Mixture) Mike Rosenblatt, Ph.D. Group Leader Mass Spec Reagents 215. We monitor
6 x 5 Ways to Ensure Your LC-MS/MS is Healthy (Also known as - Tracking Performance with the 6 x 5 LC-MS/MS Peptide Reference Mixture) Mike Rosenblatt, Ph.D. Group Leader Mass Spec Reagents 215. We monitor
Self-assembling covalent organic frameworks functionalized. magnetic graphene hydrophilic biocomposite as an ultrasensitive
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2017 Electronic Supporting Information for: Self-assembling covalent organic frameworks functionalized
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2017 Electronic Supporting Information for: Self-assembling covalent organic frameworks functionalized
Mass Spectrometry and Proteomics - Lecture 5 - Matthias Trost Newcastle University
 Mass Spectrometry and Proteomics - Lecture 5 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk Previously Proteomics Sample prep 144 Lecture 5 Quantitation techniques Search Algorithms Proteomics
Mass Spectrometry and Proteomics - Lecture 5 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk Previously Proteomics Sample prep 144 Lecture 5 Quantitation techniques Search Algorithms Proteomics
HOWTO, example workflow and data files. (Version )
 HOWTO, example workflow and data files. (Version 20 09 2017) 1 Introduction: SugarQb is a collection of software tools (Nodes) which enable the automated identification of intact glycopeptides from HCD
HOWTO, example workflow and data files. (Version 20 09 2017) 1 Introduction: SugarQb is a collection of software tools (Nodes) which enable the automated identification of intact glycopeptides from HCD
Key Words Q Exactive, Accela, MetQuest, Mass Frontier, Drug Discovery
 Metabolite Stability Screening and Hotspot Metabolite Identification by Combining High-Resolution, Accurate-Mass Nonselective and Selective Fragmentation Tim Stratton, Caroline Ding, Yingying Huang, Dan
Metabolite Stability Screening and Hotspot Metabolite Identification by Combining High-Resolution, Accurate-Mass Nonselective and Selective Fragmentation Tim Stratton, Caroline Ding, Yingying Huang, Dan
Speed Improvement of an FTICR Mass Spectra Analysis Program by Simple Modifications
 Speed Improvement of an FTICR Mass Spectra Analysis Program Bull. Korean Chem. Soc. 2009, Vol. 30, No. 9 2061 Speed Improvement of an FTICR Mass Spectra Analysis Program by Simple Modifications Sang Hyun
Speed Improvement of an FTICR Mass Spectra Analysis Program Bull. Korean Chem. Soc. 2009, Vol. 30, No. 9 2061 Speed Improvement of an FTICR Mass Spectra Analysis Program by Simple Modifications Sang Hyun
All Ions MS/MS: Targeted Screening and Quantitation Using Agilent TOF and Q-TOF LC/MS Systems
 All Ions MS/MS: Targeted Screening and Quantitation Using Agilent TOF and Q-TOF LC/MS Systems Technical Overview Introduction All Ions MS/MS is a technique that is available for Agilent high resolution
All Ions MS/MS: Targeted Screening and Quantitation Using Agilent TOF and Q-TOF LC/MS Systems Technical Overview Introduction All Ions MS/MS is a technique that is available for Agilent high resolution
Multiple Fragmentation Methods for Small Molecule Characterization on a Dual Pressure Linear Ion Trap Orbitrap Hybrid Mass Spectrometer
 Application ote: 54 Multiple Fragmentation Methods for Small Molecule Characterization on a Dual Pressure Linear Ion Trap rbitrap Hybrid Mass Spectrometer Kate Comstock, Yingying Huang; Thermo Fisher Scientific,
Application ote: 54 Multiple Fragmentation Methods for Small Molecule Characterization on a Dual Pressure Linear Ion Trap rbitrap Hybrid Mass Spectrometer Kate Comstock, Yingying Huang; Thermo Fisher Scientific,
for the Novice Mass Spectrometry (^>, John Greaves and John Roboz yc**' CRC Press J Taylor & Francis Group Boca Raton London New York
 Mass Spectrometry for the Novice John Greaves and John Roboz (^>, yc**' CRC Press J Taylor & Francis Group Boca Raton London New York CRC Press is an imprint of the Taylor & Francis Croup, an informa business
Mass Spectrometry for the Novice John Greaves and John Roboz (^>, yc**' CRC Press J Taylor & Francis Group Boca Raton London New York CRC Press is an imprint of the Taylor & Francis Croup, an informa business
Quantitative analysis of small molecules in biological samples. Jeevan Prasain, Ph.D. Department of Pharmacology & Toxicology, UAB.
 Quantitative analysis of small molecules in biological samples 100 Jeevan Prasain, Ph.D. Department of Pharmacology & Toxicology, UAB % 0 300 500 700 900 1100 1300 1500 1700 m/z Class Overview Introduction
Quantitative analysis of small molecules in biological samples 100 Jeevan Prasain, Ph.D. Department of Pharmacology & Toxicology, UAB % 0 300 500 700 900 1100 1300 1500 1700 m/z Class Overview Introduction
Multi-residue analysis of pesticides by GC-HRMS
 An Executive Summary Multi-residue analysis of pesticides by GC-HRMS Dr. Hans Mol is senior scientist at RIKILT- Wageningen UR Introduction Regulatory authorities throughout the world set and enforce strict
An Executive Summary Multi-residue analysis of pesticides by GC-HRMS Dr. Hans Mol is senior scientist at RIKILT- Wageningen UR Introduction Regulatory authorities throughout the world set and enforce strict
Achieve confident synthesis control with the Thermo Scientific ISQ EC single quadrupole mass spectrometer
 APPLICATION NOTE 72385 Achieve confident synthesis control with the Thermo Scientific ISQ EC single quadrupole mass spectrometer Authors Stephan Meding, Katherine Lovejoy, Martin Ruehl Thermo Fisher Scientific,
APPLICATION NOTE 72385 Achieve confident synthesis control with the Thermo Scientific ISQ EC single quadrupole mass spectrometer Authors Stephan Meding, Katherine Lovejoy, Martin Ruehl Thermo Fisher Scientific,
WADA Technical Document TD2015IDCR
 MINIMUM CRITERIA FOR CHROMATOGRAPHIC-MASS SPECTROMETRIC CONFIRMATION OF THE IDENTITY OF ANALYTES FOR DOPING CONTROL PURPOSES. The ability of a method to identify an analyte is a function of the entire
MINIMUM CRITERIA FOR CHROMATOGRAPHIC-MASS SPECTROMETRIC CONFIRMATION OF THE IDENTITY OF ANALYTES FOR DOPING CONTROL PURPOSES. The ability of a method to identify an analyte is a function of the entire
Increasing Speed of UHPLC-MS Analysis Using Single-stage Orbitrap Mass Spectrometer
 Increasing Speed of UHPLC-MS Analysis Using Single-stage Orbitrap Mass Spectrometer Olaf Scheibner and Maciej Bromirski Thermo Fisher Scientific, Bremen, Germany Overview Purpose: Improve the performance
Increasing Speed of UHPLC-MS Analysis Using Single-stage Orbitrap Mass Spectrometer Olaf Scheibner and Maciej Bromirski Thermo Fisher Scientific, Bremen, Germany Overview Purpose: Improve the performance
profileanalysis Innovation with Integrity Quickly pinpointing and identifying potential biomarkers in Proteomics and Metabolomics research
 profileanalysis Quickly pinpointing and identifying potential biomarkers in Proteomics and Metabolomics research Innovation with Integrity Omics Research Biomarker Discovery Made Easy by ProfileAnalysis
profileanalysis Quickly pinpointing and identifying potential biomarkers in Proteomics and Metabolomics research Innovation with Integrity Omics Research Biomarker Discovery Made Easy by ProfileAnalysis
DIA-Umpire: comprehensive computational framework for data independent acquisition proteomics
 DIA-Umpire: comprehensive computational framework for data independent acquisition proteomics Chih-Chiang Tsou 1,2, Dmitry Avtonomov 2, Brett Larsen 3, Monika Tucholska 3, Hyungwon Choi 4 Anne-Claude Gingras
DIA-Umpire: comprehensive computational framework for data independent acquisition proteomics Chih-Chiang Tsou 1,2, Dmitry Avtonomov 2, Brett Larsen 3, Monika Tucholska 3, Hyungwon Choi 4 Anne-Claude Gingras
Application Note. Edgar Naegele. Abstract
 Fast identification of main drug metabolites by quadrupole time-of-flight LC/MS Measuring accurate MS and MS/MS data with the Agilent 651 Q-TOF LC/MS and identification of main meta-bolites by comparison
Fast identification of main drug metabolites by quadrupole time-of-flight LC/MS Measuring accurate MS and MS/MS data with the Agilent 651 Q-TOF LC/MS and identification of main meta-bolites by comparison
Application Note LCMS-112 A Fully Automated Two-Step Procedure for Quality Control of Synthetic Peptides
 Application Note LCMS-112 A Fully Automated Two-Step Procedure for Quality Control of Synthetic Peptides Abstract Here we describe a two-step QC procedure for synthetic peptides. In the first step, the
Application Note LCMS-112 A Fully Automated Two-Step Procedure for Quality Control of Synthetic Peptides Abstract Here we describe a two-step QC procedure for synthetic peptides. In the first step, the
Supporting Information
 Supporting Information Highly Efficient Ionization of Phosphopeptides at Low ph by Desorption Electrospray Ionization Mass Spectrometry Ning Pan, a, b Pengyuan Liu, b Weidong Cui, c Bo Tang, a Jingmin
Supporting Information Highly Efficient Ionization of Phosphopeptides at Low ph by Desorption Electrospray Ionization Mass Spectrometry Ning Pan, a, b Pengyuan Liu, b Weidong Cui, c Bo Tang, a Jingmin
Maximizing Triple Quadrupole Mass Spectrometry Productivity with the Agilent StreamSelect LC/MS System
 Maximizing Triple Quadrupole Mass Spectrometry Productivity with the Agilent StreamSelect LC/MS System Application Note Authors Kevin McCann, Sameer Nene, Doug McIntyre, Edmond Neo, Dennis Nagtalon, and
Maximizing Triple Quadrupole Mass Spectrometry Productivity with the Agilent StreamSelect LC/MS System Application Note Authors Kevin McCann, Sameer Nene, Doug McIntyre, Edmond Neo, Dennis Nagtalon, and
Plasma Metanephrines and 3-Methoxytyramine by LC/MS/MS Using Agilent SimpliQ WCX SPE, 1290 Infi nity LC, and 6460 Triple Quadrupole LC/MS
 Plasma Metanephrines and 3-Methoxytyramine by LC/MS/MS Using Agilent SimpliQ WCX SPE, 129 Infi nity LC, and 646 Triple Quadrupole LC/MS Application Note Clinical Research Authors Linda Côté and Christophe
Plasma Metanephrines and 3-Methoxytyramine by LC/MS/MS Using Agilent SimpliQ WCX SPE, 129 Infi nity LC, and 646 Triple Quadrupole LC/MS Application Note Clinical Research Authors Linda Côté and Christophe
Reagents. Affinity Tag (Biotin) Acid Cleavage Site. Figure 1. Cleavable ICAT Reagent Structure.
 DATA SHEET Protein Expression Analysis Reagents Background The ultimate goal of proteomics is to identify and quantify proteins that are relevant to a given biological state; and to unearth networks of
DATA SHEET Protein Expression Analysis Reagents Background The ultimate goal of proteomics is to identify and quantify proteins that are relevant to a given biological state; and to unearth networks of
(Refer Slide Time 00:09) (Refer Slide Time 00:13)
 (Refer Slide Time 00:09) Mass Spectrometry Based Proteomics Professor Sanjeeva Srivastava Department of Biosciences and Bioengineering Indian Institute of Technology, Bombay Mod 02 Lecture Number 09 (Refer
(Refer Slide Time 00:09) Mass Spectrometry Based Proteomics Professor Sanjeeva Srivastava Department of Biosciences and Bioengineering Indian Institute of Technology, Bombay Mod 02 Lecture Number 09 (Refer
MS-based proteomics to investigate proteins and their modifications
 MS-based proteomics to investigate proteins and their modifications Francis Impens VIB Proteomics Core October th 217 Overview Mass spectrometry-based proteomics: general workflow Identification of protein
MS-based proteomics to investigate proteins and their modifications Francis Impens VIB Proteomics Core October th 217 Overview Mass spectrometry-based proteomics: general workflow Identification of protein
An Effective Workflow for Impurity Analysis Incorporating High Quality HRAM LCMS & MSMS with Intelligent Automated Data Mining
 An Effective Workflow for Impurity Analysis Incorporating High Quality HRAM LCMS & MSMS with Intelligent Automated Data Mining Dave Weil, Ph.D. and Jim Lau, Ph.D. Typical Method Conditions: 1260 UHPLC
An Effective Workflow for Impurity Analysis Incorporating High Quality HRAM LCMS & MSMS with Intelligent Automated Data Mining Dave Weil, Ph.D. and Jim Lau, Ph.D. Typical Method Conditions: 1260 UHPLC
Department of Chemistry
 Scanning electron micrographs of silica colloidal crystals on the same size scale for (A) 490 ± 10 nm particles and (B) 145 ± 3 nm particles. Department of Chemistry Published in: Angela R. Soemo; Mary
Scanning electron micrographs of silica colloidal crystals on the same size scale for (A) 490 ± 10 nm particles and (B) 145 ± 3 nm particles. Department of Chemistry Published in: Angela R. Soemo; Mary
Chemistry Instrumental Analysis Lecture 37. Chem 4631
 Chemistry 4631 Instrumental Analysis Lecture 37 Most analytes separated by HPLC are thermally stable and non-volatile (liquids) (unlike in GC) so not ionized easily by EI or CI techniques. MS must be at
Chemistry 4631 Instrumental Analysis Lecture 37 Most analytes separated by HPLC are thermally stable and non-volatile (liquids) (unlike in GC) so not ionized easily by EI or CI techniques. MS must be at
Computational Methods for Mass Spectrometry Proteomics
 Computational Methods for Mass Spectrometry Proteomics Eidhammer, Ingvar ISBN-13: 9780470512975 Table of Contents Preface. Acknowledgements. 1 Protein, Proteome, and Proteomics. 1.1 Primary goals for studying
Computational Methods for Mass Spectrometry Proteomics Eidhammer, Ingvar ISBN-13: 9780470512975 Table of Contents Preface. Acknowledgements. 1 Protein, Proteome, and Proteomics. 1.1 Primary goals for studying
A Quadrupole-Orbitrap Hybrid Mass Spectrometer Offers Highest Benchtop Performance for In-Depth Analysis of Complex Proteomes
 A Quadrupole-Orbitrap Hybrid Mass Spectrometer Offers Highest Benchtop Performance for In-Depth Analysis of Complex Proteomes Zhiqi Hao 1, Yi Zhang 1, Shannon Eliuk 1, Justin Blethrow 1, Dave Horn 1, Vlad
A Quadrupole-Orbitrap Hybrid Mass Spectrometer Offers Highest Benchtop Performance for In-Depth Analysis of Complex Proteomes Zhiqi Hao 1, Yi Zhang 1, Shannon Eliuk 1, Justin Blethrow 1, Dave Horn 1, Vlad
LC-MS Based Metabolomics
 LC-MS Based Metabolomics Analysing the METABOLOME 1. Metabolite Extraction 2. Metabolite detection (with or without separation) 3. Data analysis Metabolite Detection GC-MS: Naturally volatile or made volatile
LC-MS Based Metabolomics Analysing the METABOLOME 1. Metabolite Extraction 2. Metabolite detection (with or without separation) 3. Data analysis Metabolite Detection GC-MS: Naturally volatile or made volatile
PC235: 2008 Lecture 5: Quantitation. Arnold Falick
 PC235: 2008 Lecture 5: Quantitation Arnold Falick falickam@berkeley.edu Summary What you will learn from this lecture: There are many methods to perform quantitation using mass spectrometry (any method
PC235: 2008 Lecture 5: Quantitation Arnold Falick falickam@berkeley.edu Summary What you will learn from this lecture: There are many methods to perform quantitation using mass spectrometry (any method
Mass Selective Ejection by Axial Resonant Excitation from a Linear Ion Trap
 Mass Selective Ejection by Axial Resonant Excitation from a Linear Ion Trap Yuichiro Hashimoto, Hideki Hasegawa, Takashi Baba, and Izumi Waki Central Research Laboratory, Hitachi, Ld., Tokyo, Japan We
Mass Selective Ejection by Axial Resonant Excitation from a Linear Ion Trap Yuichiro Hashimoto, Hideki Hasegawa, Takashi Baba, and Izumi Waki Central Research Laboratory, Hitachi, Ld., Tokyo, Japan We
LC/MS/MS qua ntitation of β-estradiol 17-acetate using an Agilent 6460 Triple Quadrupole LC/MS working in ESI negative ion mode
 LC/MS/MS qua ntitation of β-estradiol 17-acetate using an Agilent 6460 Triple Quadrupole LC/MS working in ESI negative ion mode Application Note Authors Siji Joseph Agilent Technologies India Pvt. Ltd.
LC/MS/MS qua ntitation of β-estradiol 17-acetate using an Agilent 6460 Triple Quadrupole LC/MS working in ESI negative ion mode Application Note Authors Siji Joseph Agilent Technologies India Pvt. Ltd.
Analysis of Illegal Dyes in Food Matrices using Automated Online Sample Preparation with LC/MS
 Application Note: 56 Analysis of Illegal Dyes in Food Matrices using Automated Online Sample Preparation with LC/MS Yang Shi, Catherine Lafontaine, Matthew Berube, John Fink, François Espourteille Thermo
Application Note: 56 Analysis of Illegal Dyes in Food Matrices using Automated Online Sample Preparation with LC/MS Yang Shi, Catherine Lafontaine, Matthew Berube, John Fink, François Espourteille Thermo
A Strategy for an Unknown Screening Approach on Environmental Samples Using HRAM Mass Spectrometry
 A Strategy for an Unknown Screening Approach on Environmental Samples Using HRAM Mass Spectrometry Olaf Scheibner, 1 Patrizia van Baar, 2 Florian Wode, 2 Uwe Dünnbier, 2 Kristi Akervik, 3 Jamie Humphrie,
A Strategy for an Unknown Screening Approach on Environmental Samples Using HRAM Mass Spectrometry Olaf Scheibner, 1 Patrizia van Baar, 2 Florian Wode, 2 Uwe Dünnbier, 2 Kristi Akervik, 3 Jamie Humphrie,
Chapter 5. Complexation of Tholins by 18-crown-6:
 5-1 Chapter 5. Complexation of Tholins by 18-crown-6: Identification of Primary Amines 5.1. Introduction Electrospray ionization (ESI) is an excellent technique for the ionization of complex mixtures,
5-1 Chapter 5. Complexation of Tholins by 18-crown-6: Identification of Primary Amines 5.1. Introduction Electrospray ionization (ESI) is an excellent technique for the ionization of complex mixtures,
A Strategy for an Unknown Screening Approach on Environmental Samples using HRAM Mass Spectrometry
 A Strategy for an Unknown Screening Approach on Environmental Samples using HRAM Mass Spectrometry O. Scheibner, 1 P. van Baar, 2 F. Wode, 2 U. Dünnbier, 2 K. Akervik, 3 J. Humphries, 3 M. Bromirski 1
A Strategy for an Unknown Screening Approach on Environmental Samples using HRAM Mass Spectrometry O. Scheibner, 1 P. van Baar, 2 F. Wode, 2 U. Dünnbier, 2 K. Akervik, 3 J. Humphries, 3 M. Bromirski 1
The Pitfalls of Peaklist Generation Software Performance on Database Searches
 Proceedings of the 56th ASMS Conference on Mass Spectrometry and Allied Topics, Denver, CO, June 1-5, 2008 The Pitfalls of Peaklist Generation Software Performance on Database Searches Aenoch J. Lynn,
Proceedings of the 56th ASMS Conference on Mass Spectrometry and Allied Topics, Denver, CO, June 1-5, 2008 The Pitfalls of Peaklist Generation Software Performance on Database Searches Aenoch J. Lynn,
Bruker Daltonics. EASY-nLC. Tailored HPLC for nano-lc-ms Proteomics. Nano-HPLC. think forward
 Bruker Daltonics EASY-nLC Tailored HPLC for nano-lc-ms Proteomics think forward Nano-HPLC World-Class Performance with a Small Footprint Bruker Daltonics presents a nano-lc system, perfectly integrated
Bruker Daltonics EASY-nLC Tailored HPLC for nano-lc-ms Proteomics think forward Nano-HPLC World-Class Performance with a Small Footprint Bruker Daltonics presents a nano-lc system, perfectly integrated
Plasma-free Metanephrines Quantitation with Automated Online Sample Preparation and a Liquid Chromatography-Tandem Mass Spectrometry Method
 Plasma-free Metanephrines Quantitation with Automated Online Sample Preparation and a Liquid Chromatography-Tandem Mass Spectrometry Method Xiang He and Marta Kozak ThermoFisher Scientific, San Jose, CA,
Plasma-free Metanephrines Quantitation with Automated Online Sample Preparation and a Liquid Chromatography-Tandem Mass Spectrometry Method Xiang He and Marta Kozak ThermoFisher Scientific, San Jose, CA,
A Rapid Approach to the Confirmation of Drug Metabolites in Preclinical and Clinical Bioanalysis Studies
 A Rapid Approach to the Confirmation of Drug Metabolites in Preclinical and Clinical Bioanalysis Studies APPLICATION BENEFITS Regulatory guidelines and recommendations place a greater emphasis on the detection
A Rapid Approach to the Confirmation of Drug Metabolites in Preclinical and Clinical Bioanalysis Studies APPLICATION BENEFITS Regulatory guidelines and recommendations place a greater emphasis on the detection
Analyst Software. Peptide and Protein Quantitation Tutorial
 This document is provided to customers who have purchased AB Sciex equipment to use in the operation of such AB Sciex equipment. This document is copyright protected and any reproduction of this document
This document is provided to customers who have purchased AB Sciex equipment to use in the operation of such AB Sciex equipment. This document is copyright protected and any reproduction of this document
Chem 250 Unit 1 Proteomics by Mass Spectrometry
 Chem 250 Unit 1 Proteomics by Mass Spectrometry Article #1 Quantitative MS for proteomics: teaching a new dog old tricks. MacCoss MJ, Matthews DE., Anal Chem. 2005 Aug 1;77(15):294A-302A. 1. Synopsis 1.1.
Chem 250 Unit 1 Proteomics by Mass Spectrometry Article #1 Quantitative MS for proteomics: teaching a new dog old tricks. MacCoss MJ, Matthews DE., Anal Chem. 2005 Aug 1;77(15):294A-302A. 1. Synopsis 1.1.
Identification and Characterization of an Isolated Impurity Fraction: Analysis of an Unknown Degradant Found in Quetiapine Fumarate
 Identification and Characterization of an Isolated Impurity Fraction: Analysis of an Unknown Degradant Found in Quetiapine Fumarate Michael D. Jones, Xiang Jin Song, Robert S. Plumb, Peter J. Lee, and
Identification and Characterization of an Isolated Impurity Fraction: Analysis of an Unknown Degradant Found in Quetiapine Fumarate Michael D. Jones, Xiang Jin Song, Robert S. Plumb, Peter J. Lee, and
Quan/Qual Analyses. Unmatched Confidence for. Thermo Scientific Q Exactive Orbitrap LC-MS/MS System. Identify Quantify Confirm
 26.1645 165.91 33.267 Thermo Scientific Q Exactive rbitrap LC-MS/MS System Unmatched Confidence for Quan/Qual Analyses Identify Quantify Confirm Quanfirmation \ kwän-f r-'m -shän\ n 1 : the ability to
26.1645 165.91 33.267 Thermo Scientific Q Exactive rbitrap LC-MS/MS System Unmatched Confidence for Quan/Qual Analyses Identify Quantify Confirm Quanfirmation \ kwän-f r-'m -shän\ n 1 : the ability to
Matrix-assisted laser desorption ionization (MALDI) and
 pubs.acs.org/ac A Simple Method for Quantification of Peptides and Proteins by Matrix-Assisted Laser Desorption Ionization Mass Spectrometry Kyung Man Park, Yong Jin Bae, Sung Hee Ahn, and Myung Soo Kim*,
pubs.acs.org/ac A Simple Method for Quantification of Peptides and Proteins by Matrix-Assisted Laser Desorption Ionization Mass Spectrometry Kyung Man Park, Yong Jin Bae, Sung Hee Ahn, and Myung Soo Kim*,
Sponsored document from Journal of the American Society for Mass Spectrometry
 Sponsored document from Journal of the American Society for Mass Spectrometry Activated Ion Electron Capture Dissociation (AI ECD) of Proteins: Synchronization of Infrared and Electron Irradiation with
Sponsored document from Journal of the American Society for Mass Spectrometry Activated Ion Electron Capture Dissociation (AI ECD) of Proteins: Synchronization of Infrared and Electron Irradiation with
SEAMLESS INTEGRATION OF MASS DETECTION INTO THE UV CHROMATOGRAPHIC WORKFLOW
 SEAMLESS INTEGRATION OF MASS DETECTION INTO THE UV CHROMATOGRAPHIC WORKFLOW Paula Hong, John Van Antwerp, and Patricia McConville Waters Corporation, Milford, MA, USA Historically UV detection has been
SEAMLESS INTEGRATION OF MASS DETECTION INTO THE UV CHROMATOGRAPHIC WORKFLOW Paula Hong, John Van Antwerp, and Patricia McConville Waters Corporation, Milford, MA, USA Historically UV detection has been
Accurate, High-Throughput Protein Identification Using the Q TRAP LC/MS/MS System and Pro ID Software
 www.ietltd.com Proudly serving laboratories worldwide since 1979 CALL +1.847.913.0777 for Refurbished & Certified Lab Equipment ABI Q Trap Pro LC/MS/MS Accurate, High-Throughput Protein Identification
www.ietltd.com Proudly serving laboratories worldwide since 1979 CALL +1.847.913.0777 for Refurbished & Certified Lab Equipment ABI Q Trap Pro LC/MS/MS Accurate, High-Throughput Protein Identification
Confirmation of In Vitro Nefazodone Metabolites using the Superior Fragmentation of the QTRAP 5500 LC/MS/MS System
 Confirmation of In Vitro Nefazodone Metabolites using the Superior Fragmentation of the QTRAP 5500 LC/MS/MS System Claire Bramwell-German, Elliott Jones and Daniel Lebre AB SCIEX, Foster City, California
Confirmation of In Vitro Nefazodone Metabolites using the Superior Fragmentation of the QTRAP 5500 LC/MS/MS System Claire Bramwell-German, Elliott Jones and Daniel Lebre AB SCIEX, Foster City, California
Agilent All Ions MS/MS
 Agilent All Ions MS/MS Workflow Overview A Determine fragment ions for LC/MS Quant method B Develop final Quant method Develop LC/MS Qualitative Analysis method Process data with Find by Formula Build
Agilent All Ions MS/MS Workflow Overview A Determine fragment ions for LC/MS Quant method B Develop final Quant method Develop LC/MS Qualitative Analysis method Process data with Find by Formula Build
Amine specific Labeling Reagents for Multiplexed Relative and Absolute Protein Quantitation
 Product Bulletin itraq Reagents itraq Reagents Amine specific Labeling Reagents for Multiplexed Relative and Absolute Protein Quantitation Background Proteomics research includes the characterization of
Product Bulletin itraq Reagents itraq Reagents Amine specific Labeling Reagents for Multiplexed Relative and Absolute Protein Quantitation Background Proteomics research includes the characterization of
Quantitation of High Resolution MS Data Using UNIFI: Acquiring and Processing Full Scan or Tof-MRM (Targeted HRMS) Datasets for Quantitative Assays
 : Acquiring and Processing Full Scan or Tof-MRM (Targeted HRMS) Datasets for Quantitative Assays Mark Wrona, Jayne Kirk, and Yun Alelyunas Waters Corporation, Milford, MA, USA APPLICATION BENEFITS Ability
: Acquiring and Processing Full Scan or Tof-MRM (Targeted HRMS) Datasets for Quantitative Assays Mark Wrona, Jayne Kirk, and Yun Alelyunas Waters Corporation, Milford, MA, USA APPLICATION BENEFITS Ability
BST 226 Statistical Methods for Bioinformatics David M. Rocke. January 22, 2014 BST 226 Statistical Methods for Bioinformatics 1
 BST 226 Statistical Methods for Bioinformatics David M. Rocke January 22, 2014 BST 226 Statistical Methods for Bioinformatics 1 Mass Spectrometry Mass spectrometry (mass spec, MS) comprises a set of instrumental
BST 226 Statistical Methods for Bioinformatics David M. Rocke January 22, 2014 BST 226 Statistical Methods for Bioinformatics 1 Mass Spectrometry Mass spectrometry (mass spec, MS) comprises a set of instrumental
BIOINF 4120 Bioinformatics 2 - Structures and Systems - Oliver Kohlbacher Summer Systems Biology Exp. Methods
 BIOINF 4120 Bioinformatics 2 - Structures and Systems - Oliver Kohlbacher Summer 2013 14. Systems Biology Exp. Methods Overview Transcriptomics Basics of microarrays Comparative analysis Interactomics:
BIOINF 4120 Bioinformatics 2 - Structures and Systems - Oliver Kohlbacher Summer 2013 14. Systems Biology Exp. Methods Overview Transcriptomics Basics of microarrays Comparative analysis Interactomics:
ALIGNMENT OF LC-MS DATA USING PEPTIDE FEATURES. A Thesis XINCHENG TANG
 ALIGNMENT OF LC-MS DATA USING PEPTIDE FEATURES A Thesis by XINCHENG TANG Submitted to the Office of Graduate Studies of Texas A&M University in partial fulfillment of the requirements for the degree of
ALIGNMENT OF LC-MS DATA USING PEPTIDE FEATURES A Thesis by XINCHENG TANG Submitted to the Office of Graduate Studies of Texas A&M University in partial fulfillment of the requirements for the degree of
