Structural Heterogeneity in Populations of the Budding Yeast
|
|
|
- Peregrine Clark
- 5 years ago
- Views:
Transcription
1 JOURNAL OF BACTERIOLOGY, Dec. 1983, p /83/ $02.00/0 Copyright 1983, American Society for Microbiology Vol. 156, No. 3 Structural Heterogeneity in Populations of the Budding Yeast Saccharomyces cerevisiae MARCO VANONI, MARINA VAI, LAURA POPOLO, AND LILIA ALBERGHINA* Sezione di Biochimica Comparata, Dipartimento di Fisiologia e Biochimica Generali, Universitd di Milano, Milan, Italy Received 8 June 1983/Accepted 23 August 1983 Bud scar analysis integrated with mathematical analysis of DNA and protein distributions obtained by flow microfluorometry have been used to analyze the cell cycle of the budding yeast Saccharomyces cerevisiae. In populations of this yeast growing exponentially in batch at 30 C on different carbon and nitrogen sources with duplication times between 75 and 314 min, the budded period is always shorter (approximately 5 to 10 min) than the sum of the S + G2 + M + G1* phases (determined by the Fried analysis of DNA distributions), and parent cells always show a prereplicative unbudded period. The analysis of protein distributions obtained by flow microfluorometry indicates that the protein level per cell required for bud emergence increases at each new generation of parent cells, as observed previously for cell volume. A wide heterogeneity of cell populations derives from this pattern of budding, since older (and less frequent) parent cells have shorter generation times and produce larger (and with shorter cycle times) daughter cells. A possible molecular mechanism for the observed increase with genealogical age of the critical protein level required for bud emergence is discussed. The asymmetrical cell division of Saccharomyces cerevisiae has been extensively investigated by time-lapse cinephotomicrography experiments and by bud scar analysis (6, 13, 18, 19, 20, 29, 30). The results of these experiments may be interpreted by assuming that cells need to attain a critical size to initiate the cycle, a regulatory step that requires the execution of the CDC28 event and commits the cell to DNA replication and cell division (12, 18, 24, 29). Such a critical cell size model implies defined relationships between growth metabolismmore specifically, protein accumulation (23)- and the nuclear division cycle (1, 3). These relations may be more carefully evaluated by using the recently developed flow microfluorometry technique, which allows the experimental determination of the frequency functions of DNA, protein, and RNA contents of individual cells on significant samples (10W to 106 cells) of microbial populations (4, 11). In fact, these functions are determined by the law of growth of the individual cells, the structure of the population, and the variability of the momentary size distribution (33). It has been found that cell sizes at a given age (for instance, at budding or at division) are distributed around a mean value according to a function (the momentary size distribution) that can be well represented by a Gaussian function with a coefficient of variation (CV) generally in the range of 20% (15, 32). Computer programs that take into account the experimental variability may thus make it possible to extract the law of growth of the single cell or the structure of the population from the experimental frequency functions in a statistically significant manner. Relevant information on cell cycle dynamics may thus be obtained on sizable samples of unperturbed populations in balanced exponential growth in shaken liquid cultures, thereby overcoming the uncertainties given by the analysis of the synchronized populations (21, 22) and by the investigation of a small number of cells growing in an unusual environment, as in the time-lapse cinematography experiments. MATERIALS AND METHODS The haploid strain S288c (MATa SUC2 mal gal2 CUP)) of S. cerevisiae, kindly provided by Dr. S. Sora (Istituto di Genetica, Universita di Milano), was used throughout this study. Cells were grown in two types of basal medium: the first (YEP) contained (per liter of medium) 10 g of yeast extract, 20 g of Bacto-Peptone, and 20 g of carbon source (YEP-gluconate medium was buffered to ph 5.8 with orthophosphoric acid), and the second type (YNB) contained (per liter of medium) 6.7 g of yeast nitrogen base (Difco) without amino acids and 40 g of the carbon source. When glutamine (500 mg/liter) was used as the sole nitrogen source, Difco YNB without amino acids and ammoni- 1282
2 VOL. 156, 1983 um sulfate was used. Flasks (750 ml) containing 200 ml of medium were inoculated with fresh yeast cells and incubated in a Dubnoff water bath with shaking at 30 C. Growth was monitored as the increase in cell number, and the rate constant of exponential growth, a (min-') = ln2/t, where T is the duplication time of the culture, was determined by linear regression on a semilogarithmic plot. Determination of cel number and cel volume distributions. The number of cells present in small samples of cell culture diluted with Isoton (Coulter Electronics, Harpenden, United Kingdom) and mildly sonicated was determined by the use of a Coulter model ZBI Counter with a 100-gxm orifice tube. Mean cell volume (V) was estimated from the cell volume distributions obtained with a Coulter Channalyzer C-1000 by use of the equation IV = (Ini - * c/ Xn,) F (1) where ni is the number of cells contained in the c; channel (0 < i < 100) and F is a calibration factor. The equipment was calibrated by the use of 8.9-pLm-diameter latex spheres (Coulter Electronics). Determination of the percentage of budded and of binucleate cels. The percentage of total budded cells was determined by direct microscopic counting on at least 400 cells that had been fixed in 10%o Formalin and mildly sonicated. After staining with 0.2% Calcofluor (a gift from E. Cabib, National Institutes of Health, Bethesda, Md.), at least 1,000 cells were scored under a Leitz Dialux Fluorescence microscope at x 1,000 and assigned to one of the following classes: (i) unbudded daughters; (ii) budded daughters; (iii) unbudded parents; (iv) budded parents. The percentage of binucleated G1* cells, with two nuclei with presynthetic DNA content (cells between mitosis and cell division), was determined after at least 20 min of staining with 1,ug of 4',6'-diamino-2-phenylindole * 2HCl (DAPI) per ml in phosphate-buffered saline (PBS). At least 400 cells were scored for each determination under a Leitz Dialux fluorescence microscope at x 1,000. Deternination of the duration of cell cycle phases, budded period, and parent and daughter generation times. The formulas used to calculate TB, the length of the budded period, which is of appreciable constant duration in both parent and daughter cells (18, 19), TP (the average parent cycle time), and TD (the average daughter cycle time) are as follows. TB = 10g2(1+ FB) * T (2) TP = lo02 (FBIFPB) * T (3) TD = - lg2 (FBIFDB) T (4) where FB is the percentage of budded cells, and FPB and FDB are the fractions of budded parents and daughters, respectively, on the whole population, determined by bud scar analysis. Let FS+G2+M+Gol = Fx and FG2+M+G1* be the fractions of cells between initiation or completion of DNA synthesis, respectively, and cell division, as judged by Fried's deconvolution of DNA histograms obtained by flow cytometry. An estimation of Ty and TG2+M+G1l is given by Ty = log2 (1G+ F) T (5) TG2+M+Gol = 102 (1 + FG2+M+G1*), T (6) CELL CYCLE HETEROGENEITY IN S. CEREVISIAE 1283 By similar reasoning, TG1. is given by TGI- = 1og2 (1 + FG1.) * T (7) where FG1. is the fraction of binucleate cells. By subtracting equation 7 from equation 6, TG2+M is given by TG2+M t1 + FG1*J (8) G2+M ( 1 + FG2+M+G1*. T Similarly, subtraction of equation 6 from equation 5 gives the average duration of the DNA synthetic phase: ( 1 + FE lo21 + FG2+M+Gl*/(9 Because of the unequal mode of division of S. cerevisiae, no meaningful length of Gl phase on the whole population may be calculated from flow cytometric data. Flow cytometric analysis. Samples of cells were mildly sonicated, collected by centrifugation, washed, and suspended in 70% ethanol before analysis. To obtain protein distributions, S. cerevisiae cells were washed once with PBS (ph 7.4) and then were stained with 50,ug of fluorescein isothiocyanate per ml in 0.5 M sodium bicarbonate. After 30 min in ice, cells were recovered and washed three times with PBS (2, 7). The intensity of green fluorescence (>520 nm) was measured on at least 105 cells with a FACS IV fluorescence-activated cell sorter (Becton, Dickinson & Co.) equipped with a 5-W argon ion laser which yielded ca. 400 mw at 488 nm in our conditions. For DNA staining, cells were washed once with PBS, treated with RNase (1 mg/ml) for 90 min at 37 C, washed with PBS, and suspended in 50 mm Tris (ph 7.7) containing 15 mm MgCl2 and 46 mm propidium iodide (23). After at least 20 min of staining in ice, cells were directly examined, and total fluorescence intensity was measured as described above on at least 105 cells. By this procedure, fluorescence is collected by a light detector which transmits an electric pulse whose height is proportional to the fluorescence intensity; under our staining conditions it is also proportional to the protein or DNA content per cell. Each pulse is stored in a multichannel analyzer. In this way, the frequency distribution functions of protein and of DNA content are obtained at the end of the run. Protein and DNA distribution analysis. Protein distribution was analyzed by a computer program developed in our laboratory. First, from the age density function of the population and from the assumptions made on the behavior of the threshold required for budding at the different genealogical ages, the ideal protein density function (i.e., the frequency of cells in a population with a given protein content), which is stable in balanced exponential growth, was calculated as described in the appendix. Then, given the ideal protein density function, the program took into account variability in size for cells at any given age and variability added by measuring methods. Simulations were performed by setting CV1 = CV2 = 0.2 (the "'momentary" CV of each subpopulation of parent and daughter cells, respectively). After normalization, the program compared experimental and computed distributions (see reference 2 and Martegaui et al., Cytom- (9)
3 1284 VANONI ET AL. etry, in press, for a more detailed description). DNA distributions were analyzed by the method of Fried and Mandel (10), which briefly works as follows. (i) The population is separated into compartments, one consisting of Gl cells, another of G2 and M cells, and several of S phase cells which have synthesized different specified fractions of their DNA; (ii) the fluorescence intensity of cells in each compartment is normally distributed with the mean of the G2 + M compartment having a channel location twice that of GI; the S phase compartments have means at intermediate values. The sizes of the compartments, and hence the fraction of cells in each cell cycle phase, are determined by a least-squares fitting procedure involving the observed data, using a computer program implemented on a PDP 11/34 (Digital Equipment Corp.). Chemical determination of protein content. Samples of ca. 5 x 108 cells were collected by centrifugation and were washed once with cold distilled water and three times with cold 5% trichloroacetic acid. The protein content of the cultures was determined as previously described (23) by the standard microbiuret method. P, and Pm were calculated by the equations P. Pp ea(tpt) (10) Pm = Pp ea(tp-tg1*) (11) Pp, the average protein content of a parent cell at the beginning of a cycle, was calculated as in reference 30: =(Tp P - TD + TD etp) (12) where P is the mean protein content per cell. RESULTS Parameters of the nuclear division cycle. DNA distributions were obtained for S. cerevisiae populations in balanced exponential growth on different carbon and nitrogen sources in batch cultures at 30 C. The Fried algorithm (10) was used for the deconvolution of the DNA distributions, and the average duration of various cell phases was calculated as indicated above. The length of the S phase remained constant (approximately 15 min) for duplication times up to 110 min, then it slowly increased as the doubling time of the culture increased (Table 1). The duration of the G2+M phase steadily increased with increasing duplication time. Interestingly, at all growth rates, the budded period was shorter by 5 to 10 min than the sum of S + G2 + M +G1* (To) phases. This finding indicates that the initiation of DNA replication occurs only a few minutes before the detection of the emergence of the bud. This is not always the case; in a different yeast strain (A364a) in exponential growth or under unbalanced conditions of growth, the two events (budding and onset of DNA replication) are less closely associated; therefore, caution should be taken in using bud emergence as a morphological marker of the entrance into the S phase (23). J. BACTERIOL. TABLE 1. Timing of bud emergence and of initiation of DNA replication T TSj TG2+Ma TG1b T1a TBc a Cell cycle phases were calculated by using equations S to 9 in the text. Percentages of cells ifi S phase and G2 + M + G1* were determined by deconvolution (10) of the DNA profiles obtained by flow cytometry. Values used are means of an analysis conducted on three independent samples taken at different times during exponential growth. The correct G2 + M fraction was obtained by subtracting the fraction of binucleated cells. b TG1* was calculated according to equation 7, using values obtained by DAPI staining of nuclei. c TB was calculated by means of equation 2, using the fraction of budded cells obtained by bud scar analysis. As duplication time increased, mean cell volume (determined with a Coulter Counter Channelyzer) and average protein content (determined with the standard microbiuret method) decreased in a similar manner (Fig. 1A). Two apparent exceptions were seen in cells growing in YEP-gluconate (T = 185) and YNB-ethanol (T = 314), in which mean protein content and volume, respectively, were abnormally high. The increase in volume in cells growing on ethanol could be due to increased vacuolation, a response often seen in yeast cells in suboptimal growth conditions. Progression of the nuclear division cycle appears to depend on the reaching of a critical protein level (Ps) for the onset of DNA replication and on the reaching of a critical protein level for the onset of mitosis (Pm,Pm>Ps) and the elapse of a timer (the minimal G2 phase) for nuclear division (1, 3). The determination of the duration of the nuclear phases allows the calculation of Ps and Pm from the average protein content (1, 18). It has been found that the protein level at the initiation of DNA replication is growth modulated, i.e., it is higher at faster growth rates, as reported by other investigators for volume at bud emergence (16, 18, 30). Also, protein content at mitosis (Pm) depends on the growth rate, but the ratio between the two thresholds remains markedly constant at all growth rates (Fig. 1B and C). Deconvolution of protein distribution functions. Protein distributions of fluorescein isothiocyanate-stained cells taken from exponentially growing populations are easily reproducible. The
4 VOL. 156, ca t 0~~~~~~3 * 30 0~~~ CELL CYCLE HETEROGENEITY IN S. CEREVISIAE E 20 E Unger model, which assumes instead that all ~~~~~~~~~~I> parent cells have the same generation time CL E1.0 ce 0.5 A DUPLICATION TIME (min) FIG. 1. (A) Mean volume (0) and protein content (0) at different duplication times, measured as described in the text. Both sets of data are means of three independent determinations made on samples taken during exponential growth. Standard deviations were always smaller than l1o. The duplication time of the culture, T (min, = In2/a), was determined from the rate constant of growth (a) measured as indicated in the text. (B) Protein content required for the initiation of DNA replication, Ps, plotted as a function of duplication time. (C) Ratio between P, and Pm (the protein content required for mitosis) at different duplication times. Ps and Pm were calculated as described in the text. shape and CV of the distribution depend on the growth conditions and may have a predictive value on the state of the population, foretelling, for instance, the reaching of an exponential growth condition before it can be assessed by counting the cell number (2). Typical protein distributions are shown in Fig. 2 (closed circles). For duplication times up to 200 min, distributions were unimodal, with a CV of approximately 35 to 40%, whereas at longer duplication times, distributions became bimodal. Protein distributions were analyzed with a fitting program that accounted for measurement errors and the variability of the momentary size distribution and made the assumption that the protein content, as well as the cell volume at bud emergence, increases at each new generation of parent cells; hence, older parent cells have shorter generation times (16, 19). Computergenerated histograms fit-with fairly good accuracy-protein distributions in balanced exponential growth (Fig. 2), whereas protein distribution functions from the Hartwell and regardless of their genealogical age, are not satisfactory (2). Protein and DNA distributions are a sensitive test to ascertain the establishment of a condition z w IL cr.~ ~ ~ ~ ~ A PROTEIN CONTENT FIG. 2. Fitting of experimental protein distributions of FITC-stained cells obtained for S. cerevisiae cells exponentially growing in different media. Circles indicate experimental distributions; lines indicate computed distributions. (A) YEP-maltose; T = 139 min; TB = 113 min. (B) YNB-ethanol; T = 314 min; TB = 170 min. Simulations were performed by setting CV1 = CV2 = 0.2, Q = 0.2, and a = 0.5. CV1 and CV2 are the momentary coefficients of variation for each generation of parent and daughter cells, respectively. Q represents the increase of the critical protein level required for bud emergence for parent cells with one bud scar with respect to daughter cells, and a1- Q represents the increase in this value for parent cells with genealogical age i with respect to parent cells with genealogical age i - 1. A
5 1286 VANONI ET AL. of balanced exponential growth, i.e., a growth condition in which "the ages of randomly picked cells are independently distributed according to the stable age distribution of the population" (14). Since the cell age is not easily measured, other cell parameters must be considered. The classical requirement of an exponential increase in cell number is not in itself enough for one to assume that this condition of growth has been established. In fact, S. cerevisiae cells grown on YNB with glutamine as a nitrogen source showed an exponential increase in cell number (Fig. 3a) and a constant fraction of unbudded cells, Fu (Fig. 3b, lane 1). Nevertheless, the increase with time in the fraction of Gl cells, FG1, as judged by the Fried analysis of the DNA histograms (Fig. 3b and 4b, lane 2), and the failure of our computer program to fit protein distributions (Fig. 4a), indicate that the population is not in balanced exponential growth. Thus, protein and DNA distributions strictly related to cell age should be regarded as better indicators of the age structure of yeast cell populations than should cell number or bud emergence. Structure of the population. Since, as shown above, the protein level at bud emergence ap- E LU z -J -J Ui C) B A 108 h 1o 7 1y TIME FU 0 FG FIG. 3. (A) Growth curve for S. cerevisiae cells on YNB-glutamine. (B) Frequency of unbudded cells (Fu) obtained by direct microscopic examination, and frequency of Gl cells (FG1) derived by the Fried analysis (shown in Fig. 4B) in samples (1 to 4) taken at various times during growth, as indicated by arrows in (A). z LUI ILi -J -i LU LUJ LUI B _ X, I 3 E -I4 p. '~~~~~Cl J. BACTERIOL CHANNEL NUMBER FIG. 4. (A) Attempt to fit protein distribution in unbalanced exponential growth. Cells withdrawn at time 2 during growth of Fig. 3A were stained with FITC and analyzed as usual. Simulations were performed by setting CV1 = CV2 = 0.2. Experimentally determined parameters were T = 97 and TB = 72. circles indicate experimental distribution; line indicates computed distribution. (B) Analysis, by means of the Fried algorithm, of the DNA distributions obtained by flow cytometry at times indicated by arrows in Fig. 3A. pears to increase at each new generation, the cell division cycle of S. cerevisiae may be represented as shown in Fig. 5. A population in balanced exponential growth is then composed of subpopulations of-parents P1, P2...Pi and of daughters D1, D2...Di, where i is the genealogical age which, for parents, is indicated by the number of bud scars. The relative frequencies of the different subpopulations and average parent and daughter generation times may be calculated as indicated in the Appendix. The experimental and the calculated frequencies of parent and daughter cells closely agree with the frequencies estimated by bud scar analysis (Table 2). Also, the calculated average cycle times of parent and daughter cells are almost identical to the cycle times estimated by bud 2
6 VOL. 156, (1+8) FIG. 5. Model of cell cycle of different genealogically aged parents (P,) and daughters (D,). All daughter cells begin to bud at a relative protein content of 1, independently of the genealogical age of their parents. On the contrary, the critical protein level required for parent cells to bud increased by an a-' Q factor of each generation and thus had a value of 1 + Q for firstgeneration parents, 1 + Q (1 + a) for second-generation, etc. Since the budded period is of constant duration at each genealogical age, genealogically older parent cells and their daughters are larger at division. Hence, P2 and D2 cells are shown here to begin their cycle on the right of P1 and D1, respectively, to indicate their larger protein content. scar analysis (Table 2). To have a better understanding of the degree of heterogeneity derived from the increase in the threshold protein level for bud emergence with genealogical age, the results shown in Table 3, which shows the cycle time and the frequency of daughters (Dl to D4) and parents (P1 to P4), may be useful. There is a rather sharp decrease in cycle times of older parent and daughter cells with respect to firstgeneration parents and daughters. At all tested growth rates, the percentage of first-generation parents was almost identical to the sum of subsequent generation parents, and the budded CELL CYCLE HETEROGENEITY IN S. CEREVISIAE 1287 period was shorter than the parent cycle time, particularly for the younger and more frequent genealogical ages. This means that after cell division, parent cells are not immediately ready to undertake a new cycle and have to traverse a Gl period, which decreases in length at increasing genealogical age. The increase in cell mass of the parent cells with genealogical age causes an increase in the cell size at birth of the daughters. Accordingly, the cycle times of the daughters decrease with genealogical age (Table 3). DISCUSSION The integration of the well-known bud scar analysis with the mathematical analysis of the DNA and protein distributions, obtained by flow microfluorometry, is the element of novelty of this paper. It allows a deeper understanding of the relations between growth metabolism and the nuclear division cycle in budding yeasts. Flow cytometric techniques are a powerful tool for cell cycle analysis: in fact, small differences in the kinetics of growth of individual cells and slight variations in the structure of the population are easily detected, in a statistically significant manner, by flow microfluorometry in unperturbed populations in balanced exponential growth. Size density functions introduce a process of amplification for the detection of differences in the kinetics of single cell growth: in fact, small differences between linear and exponential growth kinetics over a cell cycle result in a much more pronounced difference between the respective size density functions (33). The analysis of protein distributions of exponentially growing cells may thus be more informative in the assessment of cell growth kinetics than the direct observation of single cells or of perturbed "synchronous" cultures (16, 21, 22). Instead, for instance, the determination of the average cycle times of parent and of daughter cells does TABLE 2. Experimental and theoretical frequency and cycle time of parent and daughter cellsa FP TP TD FD T By bud scar Calculated By bud scar Calculated By bud scar Calculated By bud scar Calculated analysis Cacltd analysis Cacltd analysis Cacltd analysis Cluae a Values of parent and daughter frequency and average cycle times were computed by using experimental a and TB and values of Q = 0.2 for all duplication times tested, except for T = 122, for which we used Q = 0.15 and a = 0.5, which gave the best fit to protein distributions.
7 1288 VANONI ET AL. J. BACTERIOL. TABLE 3. Structure of yeast populations growing at different growth rates Parent and daughter cell generation timesa Parent and daughter cell frequencies' T TB Tpl TP2 TP3 TP4 TD1 TD2 TD3 TD4 FP1 FP2 FP3 Fp4 FDI FD2 FD3 FD a Generation times of different genealogically aged parent and daughter cells were calculated by using equations A4 and A5. b Frequencies of different genealogically aged parent and daughter cells were calculated by using equations A12 and A13. not enable one to discriminate between two different age structures, i.e., that derived from the Hartwell and Unger model and that derived from our model, which implies a much larger heterogeneity in both parent and daughter cell populations (Tables 2 and 3). Since the size distribution derived from the classical unequal division model fails to fit protein distributions, because the theoretical distribution is not dispersed enough (2), the increase in volume at budding for genealogically older cells (16) appears to be accompanied by an increase in protein content. The cell division cycle in budding yeast cells appears, therefore, to take place as shown in Fig. 5 and 6. This mnechanism of cell division generates a large variability in populations of S. cerevisiae cells in balanced exponential growth, which then become composed of subpopulations of parents and daughters that have progressively shorter cycle times and that are less frequent at increasing genealogical age (Table 3). This structure of the population accounts for several observations made in budding yeast cell populations that do not find a completely satisfactory interpretation in the classical unequal division model, which yields only two subpopulations, parents and daughters (13, 20), which are briefly described here. Steady states of growth are generated in which the parent cell contributes the mass accumulated during the budded period to its daughter at division and, in the meantime, parent cells increase in size in successive generations, as previously observed (13). Parent cells grow before the next bud initiation at all growth rates. Parents are predicted to spend a longer time as unbudded at slow growth rates than at fast growth rates (Table 3), although in a less marked way than daughters, as observed previously (18, 29). In our experiments, the budded period considerably expanded at longer duplication times, as reported by other authors for cells whose growth rate was changed by limiting nutrient concentration in a chemostat (29). This behavior has been tentatively ascribed to catabolite derepression, although there is no biochemical indication of how a change in metabolism may affect cell cycle progression. However, it could be more satisfactorily interpreted in the frame of the cell size model developed in our laboratory: it holds that cell cycle progression requires the attainment of two threshold protein levels, one controlling the entrance in the S phase and the other controlling mitosis. In fact, in all growth conditions, the protein level at the beginning of mitosis is increased by the same percentage as that of the protein level at the onset of DNA replication, which suggests that both thresholds are nutritionally modulated, but in such a way that their ratio remains constant at all growth rates. The larger part of the budded period increase is observed in the post-replicative phase, although at longer duplication times, the S phase also expands. A more dramatic increase in the S phase has been reported by other authors using a different strain (A364a) and an autoradiographic technique (26), whereas a study using flow microfluorometry to investigate the yeast cell cycle found that the S phase duration remains con- TD1 i TD2' - TD3 TD4 i Tp1 TP2 Tp3 TP4 FIG. 6. Graphic representation of the cycle times of different genealogically aged parents and daughters (Tp. and TD). Heavier lines represent the budded phase, common to both parent and daughter cells and independent of genealogical age; thinner lines indicate the unbudded period, progressively shorter in genealogically older cells.
8 VOL. 156, 1983 stant at all but the lowest growth rates (17, 28). In fact, such differences could be due to the different methods used to analyze DNA histograms, which, especially with large CVs (about 10%), may yield different results, and to the different strain and temperature used. The budded period is found to be shorter by 5 to 10 min than Ty, i.e., DNA replication initiates when cells are still unbudded, or at least appear as unbudded under a light or fluorescence microscope. Such a difference could depend on the different sensitivities of the methods of detection, but it also depends on strain, temperature, and methods employed to alter the growth rate. By the use of low doses of cycloheximide on strain A364a at 24 C, much more pronounced differences have been measured (23). The percentage offirst-generation parents predicted by our model is very close, or even identical, to the sum of subsequent-generation parents (Table 3); therefore, this finding cannot be taken as an indication that the generation time of parents is independent of genealogical age, as previously proposed (5). A trend for larger parent cells to produce larger daughter cells has been observed (13), and it is explained by our model. The cycle times of both parents and daughters as a function of the genealogical age have not been extensively investigated. Uncertainty in comparing results on this aspect of the cell cycle of budding yeast cells is compounded by the fact that in time-lapse experiments, cells that are budding for their first time-as well as cells (in decreasing proportion) that are budding for their second and third time, etc. (13)-are all called first-generation parents. A recent careful study indicated that second-generation daughters have shorter cycle times than first-generation daughters (32; Table 3). The model discussed in this paper holds that there is a cell size control on'budding: it has often been suggested that a cell may do so by monitoring a specific protein whose synthesis is correlated with overall growth and that triggers cell cycle progression when it reaches a critical level (9). The assumption that such an activator protein first titrates an inhibitor, whose presence is suggested by several experimental findings (25, 31), and then activates the cell cycle progression, has been developed in a model (L. Alberghina, E. Martegani, L. Mariani, and G. Bortolan, submitted for publication). Interestingly, this mathematical model predicts for unequally dividing cells an increase in protein content at bud initiation for cells of increasing genealogical age, in agreement with the findings reported in this paper, and thereby suggests a possible molecular mechanism for the increase in the protein threshold with genealogical age. CELL CYCLE HETEROGENEITY IN S. CEREVISIAE 1289 ACKNOWLEDGMENTS We thank G. Frascotti for skillful technical assistance, and E. Martegani and L. Mariani for helpful discussions atid suggestions. This work was partially supported by grant no. CT from the Consiglio Nazionale delle Ricerche, Rome. APPENDIX The model for unequally dividing yeast cell populations used in this paper differs from that originally proposed by Hartwell and Unger (13) in that it makes the further assumption that protein level at bud emergence increases at each new generation, as suggested by the observed increase in cell volume at the onset of budding (16). Basic assumptions and relations may be summarized as follows. (i) Protein growth dynamics are described by exponential functions (8, 27). (ii) A critical protein level Pi, whose value increases with the genealogical age i, is required for bud emergence and for the beginning of DNA replication. More precisely, taken as 1, the critical protein level for daughter cells, i.e., P,O = 1, the protein content at bud emergence for ith-generation parent cells is given by: Pci= 1 + Q(1+ a+...+ a'-') (Al) with Q < 1 and a < 1, where Q represents the increase in value of the critical protein content for parent cells with one bud scar with respect to daughter cells, and a'- Q represents the increase in such level for cells with genealogical age (i) with respect to cells with genealogical age (i-1). (iii) At any given growth rate, the budded period (TB) is appreciably constant for both mother and daughter cells. (iv) At division, the protein synthesized during the budded phase is given to the bud. With these assumptions, the normalized protein contents for parent cells Pi and their daughter cells Di are given by: pi; (T) = 1 + Q(1 + a ai-1) ex(rb -- T) 0 C< T C- TPi (A2) PDi (T) = e'(b O ' T ' TDj (A3) with the generation times for ith generation of parent and daughter TPi and TDi given by: TDj = TB = TB +-lnii 1t (1 -a) +Q(1 -a') -In [1(1)+(e + Q A 4) (AS) more exactly, so that also for very great values of i, TDi 2 TB, the following assumption (v) must be made: Q 2 - etb 1 -a e otb-1 (vi) If the cell population is in balanced steady state of exponential growth, then
9 1290 VANONI ET AL. (vii) there is no cell loss in the population, and (viii) the critical protein level p and therefore the duration of the generation times depend only on genealogical age: the ideal age and protein density functions may thus be derived. The age density function np, (T); ndi (T) of the different subpopulations can be expressed as a function of a normalization factor (A), whose value can be computed to satisfy: TPI TD; 2 [InPi (T) dt + ndi ('r) dt = 1 (A6) and is given by: Ae-U(i *7W ea npi ('r) =[1 + Q(1 + a +.. ail)](ea?b - 1) 0 S T s Tpi (A7) ndi (T) = Ae-(i 1)*TB e*r 0 :C T S TDi (A8) with i = 1,2,3... and Tp1 and TD; as given by equations A4 and A5. Starting from equations A2, A3, A7, and A8, the protein density functions of the various subpopulations can be derived as and are given by n(p) = n [T(p)] * (-dtldp) (A9) Ae - 2) *TB 1 npi (p) = (eatb-1) (AlO) [1 + Q(1+ a +...+a- 2)] s p s[1 + Q(1+ a + ai'-)] eatb Ae-U - 2) atb ndi (P) = p2 (All) [1 + Q(1 + a a' - 2) eatb- 1)] S P S eatb The frequency of each generation of parent and daughter cells is given by: Tpi' Fpi = npi (T) dt / FDi = ndi (T) dt I ITPI npi (r) dr TPi ndi (T) dt Average parent and daughter generation times are thus given by Tp = Tp; * Fpi (A14) TD = I TDi * FDi (A15) The preceding equations have been used in this paper to analyze protein distributions and to determine the structure of yeast cell populations in balanced exponential growth. LITERATURE CITED J. BACTERIOL. 1. Alberghina, L., L. Marlani, and E. Martepnl Analysis of a model of cell cycle in eucaryotes. J. Theor. Biol. 87: Alberghlna, L., L. Mariani, E. Martepni, and M. Vanonl Analysis of protein distribution in budding yeast. Biotechnol. Bioeng. 25: Alberghbina, L., and E. Shuwan Control of growth and of the nuclear division cycle in Neurospora crassa. Microbiol. Rev. 45: Bailey, J. E., J. Fazel.-madJIei, D. N. McQultty, and M. F. Gilbert Measuring microbial population dynamics. Ann. N.Y. Acad. Sci. 326: Carter, B. L. A The control of cell division in Saccharomyces cerevisiae, p In P. C. L. John (ed.), The cell cycle. Cambridge University Press, Cambridge, England. 6. Carter, B. L. A., and M. N. Jagadish Control of cell division in the yeast Saccharomyces cerevisiae cultured at different growth rates. Exp. Cell. Res. 112: Crbsman, H. A., and J. A. Steinkamp Rapid simultaneous measurement of DNA, protein and cell volume in single cells from large mammalian cell populations. J. Cell. Biol. 59: Elliott, S. G., and C. S. McLaughlin Rate of macromolecular synthesis through the cell cycle of the yeast Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. U.S.A. 75: Fantes, P. A., W. D. Grant, R. H. Pritcard, P. E. Sudberry, and A. E. Wbeak The regulation of cell size and the control of mitosis. J. Theor. Biol. S0: Fried, J., and M. Mandel Multi-user system for analysis of data from flow-cytometry. Comput. Programs Biomed Gray, J. W., and P. N. Dean Display and analysis of flow-cytometric data. Annu. Rev. Biophys. Bioeng. 9: Hartwerl, L. H Saccharomyces cerevisiae cell cycle. Bacteriol. Rev. 38: Hartwell, L. H., and M. Unger Unequal division in Saccharomyces cerevisiae and its implications for the control of cell division. J. Cell Biol. 75: Jagers, P Balanced exponential growth: what does it mean and %#hen is it there?, p In A. J. Valleron and P. D. M. McDonald (ed.), Biomathematics and cell kinetics. Elsevier/North-Holland Biomedical Press, Amsterdam. 15. James, T. W., P. Hemond, G. Czer, and R. Bobman Parametric analysis of volume distributions of Schizosaccharomyces pombe and other cells. Exp. Cell Res. 94: Johnston, G. C., C. W. Ehrhardt, A. Lorincz, and B. L. A. Carter Regulation of cell size in the yeast Saccharomyces cerevisiae. J. Bacteriol. 137: Johnston, G. C., R. A. Sger, S. 0. Sharrow, and'm. L. Slater Cell division in the yeast Saccharomyces cerevisiae growing at different rates. J. Gen. Microbiol. 118: Lord, P. G., and A. E. Wbeala Asymmetrical division of Saccharomyces cerevisiae. J. Bacteriol. 142: Lord, P. G., and A. E. Wheals Variability in individual cell cycles of Saccharomyces cerevisiae. J. Cell. Sci. 5& Lord, P. G., and A. E. Wheala Rate of cell cycle initiation of yeast cells when cell size is not a ratedetermining factor. J. Cell Sci. 59: Mltchlson, J. M The biology of cell cycle. Cambridge University Press, Cambridge, England.
10 VOL. 156, Mthdbson, J. M Changing perspectives in the cell cycle, p In P. C. L. John (ed.), The cell cycle. Cambridge University Press, Cambridge, England. 23. Popolo, L., M. Vanoai, and L. Albergblna Control of the yeast cell cycle by protein synthesis. Exp. Cell Res. 142: PrInge, J. R., and L. H. Hartwell The Saccharomyces cerevisiae cell cycle. Cold Spring Harbor Monogr. Ser., p Pritchard, R. H Control of DNA replication in bacteria, p In I. Molineux and M. Kohivama (ed.), the DNA synthesis, present and future. Plenum Press, New York. 26. RIvIn, C. J., and W. L. Fangman Cell cycle phase expansion in nitrogen-limited cultures of Saccharomyces cerevisiae. J. Cell Biol. 85: Ronnlng, 0. W., E. 0. Pettersen, and P. O. Seglen Protein synthesis and protein degradation through the cell cycle of human NHIK 3025 cells in vitro. Exp. Cell Res. CELL CYCLE HETEROGENEITY IN S. CEREVISIAE : Slater, M. L., S. 0. Sharrow, and J. J. Gart Cell cycle of Saccharomyces cerevisiae in populations growing at different rates. Proc. NatI. Acad. Sci. U.S.A. 74: Tbompson, P. W., and A. E. Wheals Asymmetrical division of Saccharomyces cerevisiae in glucose-limited chemostat culture. J. Gen. Microbiol. 121: Tyson, C. B., P. G. Lord, and A. E. Wheals Dependency of size of Saccharomyces cerevisiae cells on growth rate. J. Bacteriol. 138: Tyson, J., G. Garcia-Herdurgo, and W. Sachsenmaier Control of nuclear division in Phisarum polycephalum. Exp. Cell Res. 119: Wheals, A. E Size control models of Saccharomvces cerevisiae cell population. Mol. Cell. Biol. 2: Wilams, F. M Dynamics of microbial populations, p In B. C. Patten (ed.), System analysis and simulgtion in ecology, vol. 1. Academic Press, Inc. N.Y. Downloaded from on October 29, 2018 by guest
Cell Division in the Yeast Saccharomyces cerevisiae Growing at Different Rates
 Journal of General Microbiology (1980), 118, 479-484. Printed in Great Britain 479 Cell Division in the Yeast Saccharomyces cerevisiae Growing at Different Rates By G. C. JOHNSTON,I* R. A. S. 0. SHARROW3
Journal of General Microbiology (1980), 118, 479-484. Printed in Great Britain 479 Cell Division in the Yeast Saccharomyces cerevisiae Growing at Different Rates By G. C. JOHNSTON,I* R. A. S. 0. SHARROW3
16 The Cell Cycle. Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization
 The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
Analysis and Simulation of Biological Systems
 Analysis and Simulation of Biological Systems Dr. Carlo Cosentino School of Computer and Biomedical Engineering Department of Experimental and Clinical Medicine Università degli Studi Magna Graecia Catanzaro,
Analysis and Simulation of Biological Systems Dr. Carlo Cosentino School of Computer and Biomedical Engineering Department of Experimental and Clinical Medicine Università degli Studi Magna Graecia Catanzaro,
J. Cell Sci. 35, (1979) 41 Printed in Great Britain Company of Biologists Limited 1979
 J. Cell Sci. 35, 41-51 (1979) 41 Printed in Great Britain Company of Biologists Limited 1979 ANALYSIS OF THE SIGNIFICANCE OF A PERIODIC, CELL SIZE-CONTROLLED DOUBLING IN RATES OF MACROMOLECULAR SYNTHESIS
J. Cell Sci. 35, 41-51 (1979) 41 Printed in Great Britain Company of Biologists Limited 1979 ANALYSIS OF THE SIGNIFICANCE OF A PERIODIC, CELL SIZE-CONTROLLED DOUBLING IN RATES OF MACROMOLECULAR SYNTHESIS
Dependency of Size of Saccharomyces cerevisiae Cells on
 JOURNAL OF BACTRIOLOGY, Apr. 1979, p. 9298 Vol. 138, No. 1 00219193/79/040092/07$02.00/0 Dependency of Size of Saccharomyces cerevisiae Cells on Growth Rate CHRISTINA B. TYSON, PTR G. LORD, AND ALAN. WHALS*
JOURNAL OF BACTRIOLOGY, Apr. 1979, p. 9298 Vol. 138, No. 1 00219193/79/040092/07$02.00/0 Dependency of Size of Saccharomyces cerevisiae Cells on Growth Rate CHRISTINA B. TYSON, PTR G. LORD, AND ALAN. WHALS*
J. Cell Sci. 22, (1976) Printed in Great Britain
 J. Cell Sci. 22, 59-65 (1976) Printed in Great Britain THE CONTROL OF MITOSIS IN PHYSARUM POLYCEPHALUM: THE EFFECT OF DELAYING MITOSIS AND EVIDENCE FOR THE OPERATION OF THE CONTROL MECHANISM IN THE ABSENCE
J. Cell Sci. 22, 59-65 (1976) Printed in Great Britain THE CONTROL OF MITOSIS IN PHYSARUM POLYCEPHALUM: THE EFFECT OF DELAYING MITOSIS AND EVIDENCE FOR THE OPERATION OF THE CONTROL MECHANISM IN THE ABSENCE
Tineke Jones Agriculture and Agri-Food Canada Lacombe Research Centre Lacombe, Alberta
 Growth of Escherichia Coli at Chiller Temperatures Tineke Jones Agriculture and Agri-Food Canada Lacombe Research Centre Lacombe, Alberta \ Introduction The responses of mesophilic microorganisms to chiller
Growth of Escherichia Coli at Chiller Temperatures Tineke Jones Agriculture and Agri-Food Canada Lacombe Research Centre Lacombe, Alberta \ Introduction The responses of mesophilic microorganisms to chiller
Flow cytometric analysis of Ploidy level
 Flow cytometric analysis of Ploidy level Centre of Plant Structural and Functional Genomics of the Institute of Experimental Botany AS CR, Olomouc, Czech Republic Riccardo Pasculli Roman Hudec Field application
Flow cytometric analysis of Ploidy level Centre of Plant Structural and Functional Genomics of the Institute of Experimental Botany AS CR, Olomouc, Czech Republic Riccardo Pasculli Roman Hudec Field application
Experiences with the Coulter Counter in Bacteriology1
 Experiences with the Coulter Counter in Bacteriology1 ELLEN M. SWANTON, WILLIAM A. CTJRBY, AND HOWARD E. LIND Sias Laboratories, Brooks Hospital, Brookline, Massachusetts Received for publication May 24,
Experiences with the Coulter Counter in Bacteriology1 ELLEN M. SWANTON, WILLIAM A. CTJRBY, AND HOWARD E. LIND Sias Laboratories, Brooks Hospital, Brookline, Massachusetts Received for publication May 24,
Effect of Oxygen-Supply Rates on Growth
 APPLIED MICROBIOLOGY, Jan., 1965 Vol. 13, No. 1 Copyright 1965 American Society for Microbiology Printed in U.S.A. Effect of Oxygen-Supply Rates on Growth of Escherichia coli II. Comparison of Results
APPLIED MICROBIOLOGY, Jan., 1965 Vol. 13, No. 1 Copyright 1965 American Society for Microbiology Printed in U.S.A. Effect of Oxygen-Supply Rates on Growth of Escherichia coli II. Comparison of Results
NORTHERN ILLINOIS UNIVERSITY. Screening of Chemical Libraries in Search of Inhibitors of Aflatoxin Biosynthesis. A Thesis Submitted to the
 NORTHERN ILLINOIS UNIVERSITY Screening of Chemical Libraries in Search of Inhibitors of Aflatoxin Biosynthesis A Thesis Submitted to the University Honors Program In Partial Fulfillment of the Requirements
NORTHERN ILLINOIS UNIVERSITY Screening of Chemical Libraries in Search of Inhibitors of Aflatoxin Biosynthesis A Thesis Submitted to the University Honors Program In Partial Fulfillment of the Requirements
RayBio CaspGLOW TM Fluorescein Active Caspase-3 Staining Kit
 RayBio CaspGLOW TM Fluorescein Active Caspase-3 Staining Kit User Manual Version 1.0 May 10 th, 2015 RayBio Caspase-3 Fluorometric Assay Kit Protocol (Cat#: 68FLS-Casp3-S) RayBiotech, Inc. We Provide You
RayBio CaspGLOW TM Fluorescein Active Caspase-3 Staining Kit User Manual Version 1.0 May 10 th, 2015 RayBio Caspase-3 Fluorometric Assay Kit Protocol (Cat#: 68FLS-Casp3-S) RayBiotech, Inc. We Provide You
Lecture 10: Cyclins, cyclin kinases and cell division
 Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
In Situ Gelation-Induced Death of Cancer Cells Based on Proteinosomes
 Supporting information for In Situ Gelation-Induced Death of Cancer Cells Based on Proteinosomes Yuting Zhou, Jianmin Song, Lei Wang*, Xuting Xue, Xiaoman Liu, Hui Xie*, and Xin Huang* MIIT Key Laboratory
Supporting information for In Situ Gelation-Induced Death of Cancer Cells Based on Proteinosomes Yuting Zhou, Jianmin Song, Lei Wang*, Xuting Xue, Xiaoman Liu, Hui Xie*, and Xin Huang* MIIT Key Laboratory
Electrical Sensing Zone Particle Analyzer for Measuring Germination of Fungal Spores in the Presence of Other Particles'
 APPUED MicRoBImoLY, July 1967, p. 935-639 Vol. 15, No. 4 Copyright 1967 American Society for Microbiology Printed bi U.S.A. Electrical Sensing Zone Particle Analyzer for Measuring Germination of Fungal
APPUED MicRoBImoLY, July 1967, p. 935-639 Vol. 15, No. 4 Copyright 1967 American Society for Microbiology Printed bi U.S.A. Electrical Sensing Zone Particle Analyzer for Measuring Germination of Fungal
Introduction to Biological Modeling
 Introduction to Biological odeling Lecture : odeling dynamics Sept. 9, 010 Steve Andrews Brent lab, Basic Sciences Division, FHCRC Last week Why model biology? Example: E. coli chemotaxis Typical modeling
Introduction to Biological odeling Lecture : odeling dynamics Sept. 9, 010 Steve Andrews Brent lab, Basic Sciences Division, FHCRC Last week Why model biology? Example: E. coli chemotaxis Typical modeling
EFFECT OF ph AND AMMONIUM IONS ON THE PERMEABILITY
 EFFECT OF ph AND AMMONIUM IONS ON THE PERMEABILITY OF BACILLUS PASTEURII W. R. WILEY AND J. L. STOKES Department of Bacteriology and Public Health, Washington State University, Pullman, Washington ABSTRACT
EFFECT OF ph AND AMMONIUM IONS ON THE PERMEABILITY OF BACILLUS PASTEURII W. R. WILEY AND J. L. STOKES Department of Bacteriology and Public Health, Washington State University, Pullman, Washington ABSTRACT
Supporting Online Material. On-Chip Dielectrophoretic Co-Assembly of Live Cells and. Particles into Responsive Biomaterials
 Supporting Online Material On-Chip Dielectrophoretic Co-Assembly of Live Cells and Particles into esponsive Biomaterials Shalini Gupta, ossitza G. Alargova, Peter K. Kilpatrick and Orlin D. Velev* Description
Supporting Online Material On-Chip Dielectrophoretic Co-Assembly of Live Cells and Particles into esponsive Biomaterials Shalini Gupta, ossitza G. Alargova, Peter K. Kilpatrick and Orlin D. Velev* Description
The Effect of Temperature on the Growth Rate of Saccharomyces cerevisiae
 The Effect of Temperature on the Growth Rate of Saccharomyces cerevisiae Tim Cheung, Jessica Ko, Jae Lee, and Manpreet Takhi Abstract: We conducted our experiment to determine whether increasing incubation
The Effect of Temperature on the Growth Rate of Saccharomyces cerevisiae Tim Cheung, Jessica Ko, Jae Lee, and Manpreet Takhi Abstract: We conducted our experiment to determine whether increasing incubation
7.06 Problem Set #4, Spring 2005
 7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
Production of Recombinant Annexin V from plasmid pet12a-papi
 Tait Research Laboratory Page 1 of 5 Principle Production of Recombinant Annexin V from plasmid pet12a-papi Annexin V is expressed cytoplasmically in BL21(DE3) E. coli (Novagen) with the pet vector system
Tait Research Laboratory Page 1 of 5 Principle Production of Recombinant Annexin V from plasmid pet12a-papi Annexin V is expressed cytoplasmically in BL21(DE3) E. coli (Novagen) with the pet vector system
Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc
 OPTIMIZATION OF IMMUNOBLOT PROTOCOL 121 Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc Jacqueline Bjornton and John Wheeler Faculty Sponsor: Anne
OPTIMIZATION OF IMMUNOBLOT PROTOCOL 121 Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc Jacqueline Bjornton and John Wheeler Faculty Sponsor: Anne
Germination of Individual Bacillus subtilis Spores with Alterations in the GerD and SpoVA Proteins, Which Are Important in Spore Germination
 JOURNAL OF BACTERIOLOGY, May 2011, p. 2301 2311 Vol. 193, No. 9 0021-9193/11/$12.00 doi:10.1128/jb.00122-11 Copyright 2011, American Society for Microbiology. All Rights Reserved. Germination of Individual
JOURNAL OF BACTERIOLOGY, May 2011, p. 2301 2311 Vol. 193, No. 9 0021-9193/11/$12.00 doi:10.1128/jb.00122-11 Copyright 2011, American Society for Microbiology. All Rights Reserved. Germination of Individual
Quantification of Protein Half-Lives in the Budding Yeast Proteome
 Supporting Methods Quantification of Protein Half-Lives in the Budding Yeast Proteome 1 Cell Growth and Cycloheximide Treatment Three parallel cultures (17 ml) of each TAP-tagged strain were grown in separate
Supporting Methods Quantification of Protein Half-Lives in the Budding Yeast Proteome 1 Cell Growth and Cycloheximide Treatment Three parallel cultures (17 ml) of each TAP-tagged strain were grown in separate
CONTROL OF CELL SIZE AND CYCLE TIME IN SCHIZOSACCHAROMYCES POMBE
 J. Cell Set. 24, 51-67 (1977) 51 Printed in Great Britain CONTROL OF CELL SIZE AND CYCLE TIME IN SCHIZOSACCHAROMYCES POMBE P. A. FANTES Department of Zoology, University of Edinburgh, West Mains Road,
J. Cell Set. 24, 51-67 (1977) 51 Printed in Great Britain CONTROL OF CELL SIZE AND CYCLE TIME IN SCHIZOSACCHAROMYCES POMBE P. A. FANTES Department of Zoology, University of Edinburgh, West Mains Road,
Bottom-up Optimization of SERS Hot Spots. Supplementary Information
 Bottom-up Optimization of SERS Hot Spots Laura Fabris, * Department of Materials Science and Engineering, Institute for Advanced Materials Devices ad Nanotechnology, Rutgers, The State University of New
Bottom-up Optimization of SERS Hot Spots Laura Fabris, * Department of Materials Science and Engineering, Institute for Advanced Materials Devices ad Nanotechnology, Rutgers, The State University of New
Distribution of Cell Size in Growing Cultures of Bacteria and the Applicability of the Collins-Richmond Principle
 J. gen. Microbial. (1966), 45, 409-417 Printed in Great Britain 409 Distribution of Cell Size in Growing Cultures of Bacteria and the Applicability of the Collins-Richmond Principle BY A. L. KOCH Departments
J. gen. Microbial. (1966), 45, 409-417 Printed in Great Britain 409 Distribution of Cell Size in Growing Cultures of Bacteria and the Applicability of the Collins-Richmond Principle BY A. L. KOCH Departments
CYTOLOGICAL CHANGES IN AGING BACTERIAL CULTURES
 CYTOLOGICAL CHANGES IN AGING BACTERIAL CULTURES B. R. CHATTERJEE AND ROBERT P. WILLIAMS Department of Microbiology, Baylor University College of Medicine, Houston, Texas Received for publication March
CYTOLOGICAL CHANGES IN AGING BACTERIAL CULTURES B. R. CHATTERJEE AND ROBERT P. WILLIAMS Department of Microbiology, Baylor University College of Medicine, Houston, Texas Received for publication March
ATPase/GTPase ELIPA BIOCHEM KIT
 ATPase/GTPase ELIPA BIOCHEM KIT Cat # BK051/BK052 ORDERING INFORMATION To order by phone: (303) - 322-2254 To order by Fax: (303) - 322-2257 Customer Service cserve@cytoskeleton.com Technical assistance:
ATPase/GTPase ELIPA BIOCHEM KIT Cat # BK051/BK052 ORDERING INFORMATION To order by phone: (303) - 322-2254 To order by Fax: (303) - 322-2257 Customer Service cserve@cytoskeleton.com Technical assistance:
Synchronization of Cell Division in Microorganisms by Percoll
 JOURNAL OF BACTERIOLOGY, Oct. 198, p. 17-21 21-9193/8/1-17/5$2./ Vol. 144, No. 1 Synchronization of Cell Division in Microorganisms by Percoll Gradients RINA D. DWEK, LIAT H. KOBRIN, NILI GROSSMAN, AND
JOURNAL OF BACTERIOLOGY, Oct. 198, p. 17-21 21-9193/8/1-17/5$2./ Vol. 144, No. 1 Synchronization of Cell Division in Microorganisms by Percoll Gradients RINA D. DWEK, LIAT H. KOBRIN, NILI GROSSMAN, AND
HYDROGEN. technique. uptake/co2 uptake, which according to equation (1) should equal 4, has
 184 BA CTERIOLOG Y: H. A. BARKER PROC. N. A. S. STUDIES ON THE METHANE FERMENTATION. VI. THE IN- FLUENCE OF CARBON DIOXIDE CONCENTRATION ON THE RATE OF CARBON DIOXIDE REDUCTION BY MOLECULAR HYDROGEN By
184 BA CTERIOLOG Y: H. A. BARKER PROC. N. A. S. STUDIES ON THE METHANE FERMENTATION. VI. THE IN- FLUENCE OF CARBON DIOXIDE CONCENTRATION ON THE RATE OF CARBON DIOXIDE REDUCTION BY MOLECULAR HYDROGEN By
(diploid) -- P6S (haploid) -* P6SP (diploid).
 EFFECT OF HAPLOID OR DIPLOID CONSTITUTION OF ESCHERICHIA COLI ON FORMIC HYDROGENLYASE ACTIVITY RONALD H. OLSEN AND JAMES E. OGG Department of Microbiology and Pathology, Colorado State University, Fort
EFFECT OF HAPLOID OR DIPLOID CONSTITUTION OF ESCHERICHIA COLI ON FORMIC HYDROGENLYASE ACTIVITY RONALD H. OLSEN AND JAMES E. OGG Department of Microbiology and Pathology, Colorado State University, Fort
NucView TM 488 Caspase-3 Assay Kit for Live Cells
 NucView TM 488 Caspase-3 Assay Kit for Live Cells Catalog Number: 30029 (100-500 assays) Contact Information Address: Biotium, Inc. 3423 Investment Blvd. Suite 8 Hayward, CA 94545 USA Telephone: (510)
NucView TM 488 Caspase-3 Assay Kit for Live Cells Catalog Number: 30029 (100-500 assays) Contact Information Address: Biotium, Inc. 3423 Investment Blvd. Suite 8 Hayward, CA 94545 USA Telephone: (510)
Plant Molecular and Cellular Biology Lecture 8: Mechanisms of Cell Cycle Control and DNA Synthesis Gary Peter
 Plant Molecular and Cellular Biology Lecture 8: Mechanisms of Cell Cycle Control and DNA Synthesis Gary Peter 9/10/2008 1 Learning Objectives Explain why a cell cycle was selected for during evolution
Plant Molecular and Cellular Biology Lecture 8: Mechanisms of Cell Cycle Control and DNA Synthesis Gary Peter 9/10/2008 1 Learning Objectives Explain why a cell cycle was selected for during evolution
Multiple Septation in Variants of Bacillus cereus
 JOURNAL OF BACTERIOLOGY, Nov., 1965 Copyright @ 1965 American Society for Microbiology Vol. 90, No. 5 Printed in U.S.A. Multiple Septation in Variants of Bacillus cereus C. C. REMSEN AND D. G. LUNDGREN
JOURNAL OF BACTERIOLOGY, Nov., 1965 Copyright @ 1965 American Society for Microbiology Vol. 90, No. 5 Printed in U.S.A. Multiple Septation in Variants of Bacillus cereus C. C. REMSEN AND D. G. LUNDGREN
Specific Induction of ea2 Transport Activity in MATa Cells of Saccharomyces cerevisiae by a Mating Pheromone, a Factor*
 THE JOURNAL OF BIOLOGICAL CHEMISTRY 1985 by The American Society of Biological Chemists, Inc. Vol. 26, No. 19, Issue of September 5, pp. 1482-1486,1985 Printed in U.S.A. Specific Induction of ea2 Transport
THE JOURNAL OF BIOLOGICAL CHEMISTRY 1985 by The American Society of Biological Chemists, Inc. Vol. 26, No. 19, Issue of September 5, pp. 1482-1486,1985 Printed in U.S.A. Specific Induction of ea2 Transport
*D? part ment of Microbiology and Biochemistry, Slovak Technical Bratislava 1
 Biosynthesis of Chloramphenicol. IVa*. Isolation and Some Properties of 3DeoxyDarabmoheptulosonate 7phosphate Synthetase (E. C. 4. 1. 2. 15) of Streptomyces sp. 3022a *B. ŠKÁRKA,** bd. W. S. WESTLAKE and
Biosynthesis of Chloramphenicol. IVa*. Isolation and Some Properties of 3DeoxyDarabmoheptulosonate 7phosphate Synthetase (E. C. 4. 1. 2. 15) of Streptomyces sp. 3022a *B. ŠKÁRKA,** bd. W. S. WESTLAKE and
09/07/16 12/07/16: 14/07/16:
 09/07/16 Transformation of DH5 alpha with the InterLab transformation protocol : DNA constructions were dried, so we resuspended them into 100µL nuclease-free water. We took 5µL of the product for 25µL
09/07/16 Transformation of DH5 alpha with the InterLab transformation protocol : DNA constructions were dried, so we resuspended them into 100µL nuclease-free water. We took 5µL of the product for 25µL
Helical Macrofiber Formation in Bacillus subtilis: Inhibition by Penicillin G
 JOURNAL OF BACTERIOLOGY, June 1984, p. 1182-1187 0021-9193/84/061182-06$02.00/0 Copyright C 1984, American Society for Microbiology Vol. 158, No. 3 Helical Macrofiber Formation in Bacillus subtilis: Inhibition
JOURNAL OF BACTERIOLOGY, June 1984, p. 1182-1187 0021-9193/84/061182-06$02.00/0 Copyright C 1984, American Society for Microbiology Vol. 158, No. 3 Helical Macrofiber Formation in Bacillus subtilis: Inhibition
NAD + /NADH Assay [Colorimetric]
![NAD + /NADH Assay [Colorimetric] NAD + /NADH Assay [Colorimetric]](/thumbs/91/105496846.jpg) G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name NAD + /NADH Assay [Colorimetric] (Cat. #786 1539, 786 1540) think proteins! think G-Biosciences
G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name NAD + /NADH Assay [Colorimetric] (Cat. #786 1539, 786 1540) think proteins! think G-Biosciences
Measuring Mitochondrial Membrane Potential with JC-1 Using the Cellometer Vision Image Cytometer
 Measuring Mitochondrial Membrane Potential with JC-1 Using the Cellometer Vision Image Cytometer Nexcelom Bioscience LLC. 360 Merrimack Street, Building 9 Lawrence, MA 01843 T: 978.327.5340 F: 978.327.5341
Measuring Mitochondrial Membrane Potential with JC-1 Using the Cellometer Vision Image Cytometer Nexcelom Bioscience LLC. 360 Merrimack Street, Building 9 Lawrence, MA 01843 T: 978.327.5340 F: 978.327.5341
Cell Cycle Control in the Fission Yeast Schizosaccharomyces pombe
 Journal of Generul Microbiology ( 1989, 131, 2 123-2 127. Printed in Great Britain 2123 Cell Cycle Control in the Fission Yeast Schizosaccharomyces pombe The Tenth Fleming Lecture ByPAUL NURSE Imperial
Journal of Generul Microbiology ( 1989, 131, 2 123-2 127. Printed in Great Britain 2123 Cell Cycle Control in the Fission Yeast Schizosaccharomyces pombe The Tenth Fleming Lecture ByPAUL NURSE Imperial
Coulter Counters. range can be selectively counted. overlap appreciably, they can then be counted. and threshold settings can be chosen so
 APPUED MICROBILOGY, JUlY 1973, p. 9-13 Copyright 0 1973 American Society for Microbiology Vol. 26, No. 1 Printed in U.S.A. Differential Counting in Mixed Cultures with Coulter Counters J. F. DRAKE AND
APPUED MICROBILOGY, JUlY 1973, p. 9-13 Copyright 0 1973 American Society for Microbiology Vol. 26, No. 1 Printed in U.S.A. Differential Counting in Mixed Cultures with Coulter Counters J. F. DRAKE AND
Supplementary Information for. Single-cell dynamics of the chromosome replication and cell division cycles in mycobacteria
 Supplementary Information for Single-cell dynamics of the chromosome replication and cell division cycles in mycobacteria Isabella Santi 1 *, Neeraj Dhar 1, Djenet Bousbaine 1, Yuichi Wakamoto, John D.
Supplementary Information for Single-cell dynamics of the chromosome replication and cell division cycles in mycobacteria Isabella Santi 1 *, Neeraj Dhar 1, Djenet Bousbaine 1, Yuichi Wakamoto, John D.
Application Note: A TD-700 Laboratory Fluorometer Method for Alkaline Phosphatase Fluorescence
 1. INTRODUCTION Because of their critical functions in eukaryotic cells, methods for measuring protein phosphatases were established at least as early as 1953 1. In 1965 Fernley and Walker 2 decribed the
1. INTRODUCTION Because of their critical functions in eukaryotic cells, methods for measuring protein phosphatases were established at least as early as 1953 1. In 1965 Fernley and Walker 2 decribed the
Microbiology: An Introduction, 12e (Tortora) Chapter 2 Chemical Principles. 2.1 Multiple Choice Questions
 Microbiology An Introduction 12th Edition Tortora TEST BANK Full download at: https://testbankreal.com/download/microbiology-an-introduction-12thedition-tortora-test-bank/ Microbiology An Introduction
Microbiology An Introduction 12th Edition Tortora TEST BANK Full download at: https://testbankreal.com/download/microbiology-an-introduction-12thedition-tortora-test-bank/ Microbiology An Introduction
Supplementary materials Quantitative assessment of ribosome drop-off in E. coli
 Supplementary materials Quantitative assessment of ribosome drop-off in E. coli Celine Sin, Davide Chiarugi, Angelo Valleriani 1 Downstream Analysis Supplementary Figure 1: Illustration of the core steps
Supplementary materials Quantitative assessment of ribosome drop-off in E. coli Celine Sin, Davide Chiarugi, Angelo Valleriani 1 Downstream Analysis Supplementary Figure 1: Illustration of the core steps
Studies on Basidiospore Development in Schizophyllum commune
 Journal of General Microbiology (1976), 96,49-41 3 Printed in Great Britain 49 Studies on Basidiospore Development in Schizophyllum commune By SUSAN K. BROMBERG" AND MARVIN N. SCHWALB Department of Microbiology,
Journal of General Microbiology (1976), 96,49-41 3 Printed in Great Britain 49 Studies on Basidiospore Development in Schizophyllum commune By SUSAN K. BROMBERG" AND MARVIN N. SCHWALB Department of Microbiology,
A Simple Fluorescein Derived Colorimetric and Fluorescent off - on Sensor For The Detection of Hypochlorite
 Electronic Supplementary Material (ESI) for Analytical Methods. This journal is The Royal Society of Chemistry 2018 Supporting information A Simple Fluorescein Derived Colorimetric and Fluorescent off
Electronic Supplementary Material (ESI) for Analytical Methods. This journal is The Royal Society of Chemistry 2018 Supporting information A Simple Fluorescein Derived Colorimetric and Fluorescent off
Eppendorf Plate Deepwell 96 and 384: RecoverMax
 Applications Note 145 March 2007 Eppendorf Plate Deepwell 96 and 384: RecoverMax Investigation into the impact of an optimized well design on resuspension properties, sample losses and contamination effects
Applications Note 145 March 2007 Eppendorf Plate Deepwell 96 and 384: RecoverMax Investigation into the impact of an optimized well design on resuspension properties, sample losses and contamination effects
RELATIONSHIP OF CELL WALL STAINING TO GRAM DIFFERENTIATION'
 RELATONSHP OF CELL WALL STANNG TO GRAM DFFERENTATON' J. W. BARTHOLOMEW AND HAROLD FNKELSTEN Department of Bacteriology, University of Southern California, Los Angeles, California Received for publication
RELATONSHP OF CELL WALL STANNG TO GRAM DFFERENTATON' J. W. BARTHOLOMEW AND HAROLD FNKELSTEN Department of Bacteriology, University of Southern California, Los Angeles, California Received for publication
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06072 SUPPLEMENTARY INFORMATION Molecular noise and size control: origins of variability in the budding yeast cell cycle. Stefano Di Talia 1,2, Jan M. Skotheim 2, James M. Bean 1,*,
doi: 10.1038/nature06072 SUPPLEMENTARY INFORMATION Molecular noise and size control: origins of variability in the budding yeast cell cycle. Stefano Di Talia 1,2, Jan M. Skotheim 2, James M. Bean 1,*,
Solutions. Solution: A solution is homogeneous liquid mixture of two or more substances.
 Solutions Objectives: 1. Learn the various methods of expressing concentrations of solutions. 2. Learn to make percent and molar solutions from solids, liquids, and stock solutions. 3. Learn the various
Solutions Objectives: 1. Learn the various methods of expressing concentrations of solutions. 2. Learn to make percent and molar solutions from solids, liquids, and stock solutions. 3. Learn the various
Magnetic Janus Nanorods for Efficient Capture, Separation. and Elimination of Bacteria
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2016 Magnetic Janus Nanorods for Efficient Capture, Separation and Elimination of Bacteria Zhi-min
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2016 Magnetic Janus Nanorods for Efficient Capture, Separation and Elimination of Bacteria Zhi-min
Prof. Fahd M. Nasr. Lebanese university Faculty of sciences I Department of Natural Sciences.
 Prof. Fahd M. Nasr Lebanese university Faculty of sciences I Department of Natural Sciences fnasr@ul.edu.lb B3206 Microbial Genetics Eukaryotic M. G. The yeast Saccharomyces cerevisiae as a genetic model
Prof. Fahd M. Nasr Lebanese university Faculty of sciences I Department of Natural Sciences fnasr@ul.edu.lb B3206 Microbial Genetics Eukaryotic M. G. The yeast Saccharomyces cerevisiae as a genetic model
affected by the ph of the medium, the dependence of the bacteriostasis by dyes
 THE BACTERICIDAL AND BACTERIOSTATIC ACTION OF CRYSTAL VIOLET C. E. HOFFMANN AND OTTO RAHN Bacteriological Laboratory, New York State College of Agriculture, Cornell University, Ithaca, N. Y. Received for
THE BACTERICIDAL AND BACTERIOSTATIC ACTION OF CRYSTAL VIOLET C. E. HOFFMANN AND OTTO RAHN Bacteriological Laboratory, New York State College of Agriculture, Cornell University, Ithaca, N. Y. Received for
ANALYSIS OF MICROBIAL COMPETITION
 ANALYSIS OF MICROBIAL COMPETITION Eric Pomper Microbiology 9 Pittsburgh Central Catholic High School Grade 9 Introduction Escherichia coli (E. coli) and Saccharomyces cerevisiae (Yeast) were grown together
ANALYSIS OF MICROBIAL COMPETITION Eric Pomper Microbiology 9 Pittsburgh Central Catholic High School Grade 9 Introduction Escherichia coli (E. coli) and Saccharomyces cerevisiae (Yeast) were grown together
Chemistry 119: Experiment 6. Sampling and Analysis of a Solid Drain Cleaner
 Chemistry 119: Experiment 6 Sampling and Analysis of a Solid Drain Cleaner An important factor in any analysis is the collection of the sample. How this is done depends upon the use to which the analytical
Chemistry 119: Experiment 6 Sampling and Analysis of a Solid Drain Cleaner An important factor in any analysis is the collection of the sample. How this is done depends upon the use to which the analytical
arxiv:physics/ v1 [physics.bio-ph] 30 Sep 2002
![arxiv:physics/ v1 [physics.bio-ph] 30 Sep 2002 arxiv:physics/ v1 [physics.bio-ph] 30 Sep 2002](/thumbs/73/68632814.jpg) Zipf s Law in Gene Expression Chikara Furusawa Center for Developmental Biology, The Institute of Physical arxiv:physics/0209103v1 [physics.bio-ph] 30 Sep 2002 and Chemical Research (RIKEN), Kobe 650-0047,
Zipf s Law in Gene Expression Chikara Furusawa Center for Developmental Biology, The Institute of Physical arxiv:physics/0209103v1 [physics.bio-ph] 30 Sep 2002 and Chemical Research (RIKEN), Kobe 650-0047,
Last 4 Digits of USC ID:
 Chemistry 05 B Practice Exam Dr. Jessica Parr First Letter of last Name PLEASE PRINT YOUR NAME IN BLOCK LETTERS Name: Last 4 Digits of USC ID: Lab TA s Name: Question Points Score Grader 8 2 4 3 9 4 0
Chemistry 05 B Practice Exam Dr. Jessica Parr First Letter of last Name PLEASE PRINT YOUR NAME IN BLOCK LETTERS Name: Last 4 Digits of USC ID: Lab TA s Name: Question Points Score Grader 8 2 4 3 9 4 0
RayBio Nickel Magnetic Particles
 RayBio Nickel Magnetic Particles Catalog #: 801-108 User Manual Last revised January 4 th, 2017 Caution: Extraordinarily useful information enclosed ISO 1348 Certified 3607 Parkway Lane, Suite 100 Norcross,
RayBio Nickel Magnetic Particles Catalog #: 801-108 User Manual Last revised January 4 th, 2017 Caution: Extraordinarily useful information enclosed ISO 1348 Certified 3607 Parkway Lane, Suite 100 Norcross,
Use of Agilent Feature Extraction Software (v8.1) QC Report to Evaluate Microarray Performance
 Use of Agilent Feature Extraction Software (v8.1) QC Report to Evaluate Microarray Performance Anthea Dokidis Glenda Delenstarr Abstract The performance of the Agilent microarray system can now be evaluated
Use of Agilent Feature Extraction Software (v8.1) QC Report to Evaluate Microarray Performance Anthea Dokidis Glenda Delenstarr Abstract The performance of the Agilent microarray system can now be evaluated
Axenic Growth of Dictyostelium discoideum Wild-type NC-4 Cells and its Relation to Endocytotic Ability
 Journal of General Microbiology (1983), 129, 2467-2473. Printed in Great Britain 2467 Axenic Growth of Dictyostelium discoideum Wild-type NC-4 Cells and its Relation to Endocytotic Ability By YASUO MAEDAT
Journal of General Microbiology (1983), 129, 2467-2473. Printed in Great Britain 2467 Axenic Growth of Dictyostelium discoideum Wild-type NC-4 Cells and its Relation to Endocytotic Ability By YASUO MAEDAT
Inheritance of Capsule and the Manner of Cell-Wall Formation in Bacillus anthracis
 J. gen. Microbiol. (1965), 39, 423427 With 2 plates Printed in Great Britain 423 Inheritance of Capsule and the Manner of Cell-Wall Formation in Bacillus anthracis BY G. G. MEYNELL AND A. M. LAWN Guinness-Lister
J. gen. Microbiol. (1965), 39, 423427 With 2 plates Printed in Great Britain 423 Inheritance of Capsule and the Manner of Cell-Wall Formation in Bacillus anthracis BY G. G. MEYNELL AND A. M. LAWN Guinness-Lister
Cell cycle regulation in the budding yeast
 Cell cycle regulation in the budding yeast Bởi: TS. Nguyen Cuong Introduction The cell cycle is the sequence of events by which a growing cell duplicates all its components and then divides into two daughter
Cell cycle regulation in the budding yeast Bởi: TS. Nguyen Cuong Introduction The cell cycle is the sequence of events by which a growing cell duplicates all its components and then divides into two daughter
A Stochastic Model Correctly Predicts Changes in Budding Yeast Cell Cycle Dynamics upon Periodic Expression of CLN2
 A Stochastic Model Correctly Predicts Changes in Budding Yeast Cell Cycle Dynamics upon Periodic Expression of CLN2 Cihan Oguz 1 *, Alida Palmisano 1,2, Teeraphan Laomettachit 3, Layne T. Watson 2,5,6,
A Stochastic Model Correctly Predicts Changes in Budding Yeast Cell Cycle Dynamics upon Periodic Expression of CLN2 Cihan Oguz 1 *, Alida Palmisano 1,2, Teeraphan Laomettachit 3, Layne T. Watson 2,5,6,
nutrients growth & division repellants movement
 Network Dynamics and Cell Physiology John J. Tyson Department of Biological Sciences & Virginia Bioinformatics Institute Outline 1. Cell Signaling: Physiology 2. Cell Signaling: Molecular Biology 3. Chemical
Network Dynamics and Cell Physiology John J. Tyson Department of Biological Sciences & Virginia Bioinformatics Institute Outline 1. Cell Signaling: Physiology 2. Cell Signaling: Molecular Biology 3. Chemical
Application of Micro-Flow Imaging (MFI TM ) to The Analysis of Particles in Parenteral Fluids. October 2006 Ottawa, Canada
 Application of Micro-Flow Imaging (MFI TM ) to The Analysis of Particles in Parenteral Fluids October 26 Ottawa, Canada Summary The introduction of a growing number of targeted protein-based drug formulations
Application of Micro-Flow Imaging (MFI TM ) to The Analysis of Particles in Parenteral Fluids October 26 Ottawa, Canada Summary The introduction of a growing number of targeted protein-based drug formulations
Thermal Injury and Recovery of Salmonella typhimurium and Its Effect on
 APPLIED MICROBIOLOGY, Sept. 1969, p. 332-336 Copyright @ 1969 American Society for Microbiology Vol. 18, No. 3 Printed in U.S.A. Thermal Injury and Recovery of Salmonella typhimurium and Its Effect on
APPLIED MICROBIOLOGY, Sept. 1969, p. 332-336 Copyright @ 1969 American Society for Microbiology Vol. 18, No. 3 Printed in U.S.A. Thermal Injury and Recovery of Salmonella typhimurium and Its Effect on
Automated Determination of Serum Alkaline Phosphatase Using a Modified. H. Keay and J. A. Trew*
 Automated Determination of Serum Alkaline Phosphatase Using a Modified Bodansky Technic H. Keay and J. A. Trew* An automated procedure for the measurement of alkaline phosphatase activity by a modification
Automated Determination of Serum Alkaline Phosphatase Using a Modified Bodansky Technic H. Keay and J. A. Trew* An automated procedure for the measurement of alkaline phosphatase activity by a modification
MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1)
 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) 1) Which of the following statements about the atom A) It has 12 neutrons in its nucleus. B) It
MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) 1) Which of the following statements about the atom A) It has 12 neutrons in its nucleus. B) It
The Influence of Magnesium on Cell Division
 480 WEBB, M. (1951). J. gen. Mimobiol. 5, 480-484. The Influence of Magnesium on Cell Division 4. The Specificity of Magnesium BY M. WEBB Chemistry Department, The University, Edgbaston, Birmingham 15,
480 WEBB, M. (1951). J. gen. Mimobiol. 5, 480-484. The Influence of Magnesium on Cell Division 4. The Specificity of Magnesium BY M. WEBB Chemistry Department, The University, Edgbaston, Birmingham 15,
(From the May Inctitute /or Medical Researck and Department of Physiology, Uni~ersgty of Cincinnati Medical School, Cincinnati)
 Published Online: 20 March, 1955 Supp Info: http://doi.org/10.1085/jgp.38.4.425 Downloaded from jgp.rupress.org on November 19, 2018 STUDIES IN CELL PERMEABILITY THE UPTAKE O~ PYRUVATE BY YEAST* BY E.
Published Online: 20 March, 1955 Supp Info: http://doi.org/10.1085/jgp.38.4.425 Downloaded from jgp.rupress.org on November 19, 2018 STUDIES IN CELL PERMEABILITY THE UPTAKE O~ PYRUVATE BY YEAST* BY E.
Reactor Operation Without Feedback Effects
 22.05 Reactor Physics - Part Twenty-Six Reactor Operation Without Feedback Effects 1. Reference Material: See pp. 363-368 of the article, Light Water Reactor Control Systems, in Wiley Encyclopedia of Electrical
22.05 Reactor Physics - Part Twenty-Six Reactor Operation Without Feedback Effects 1. Reference Material: See pp. 363-368 of the article, Light Water Reactor Control Systems, in Wiley Encyclopedia of Electrical
7.06 Cell Biology EXAM #3 April 21, 2005
 7.06 Cell Biology EXAM #3 April 21, 2005 This is an open book exam, and you are allowed access to books, a calculator, and notes but not computers or any other types of electronic devices. Please write
7.06 Cell Biology EXAM #3 April 21, 2005 This is an open book exam, and you are allowed access to books, a calculator, and notes but not computers or any other types of electronic devices. Please write
Electron Microscopic Studies on Mode of Action of Polymyxin
 JOURNAL OF BACrERIOLOGY, Jan. 1969, p. 448452 Vol. 97, No. I Copyright 1969 American Society for Microbiology Printed In U.S.A. Electron Microscopic Studies on Mode of Action of Polymyxin M. KOIKE, K.
JOURNAL OF BACrERIOLOGY, Jan. 1969, p. 448452 Vol. 97, No. I Copyright 1969 American Society for Microbiology Printed In U.S.A. Electron Microscopic Studies on Mode of Action of Polymyxin M. KOIKE, K.
MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. Figure 2.1
 Exam Name MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. Figure 2.1 1) Which compound in Figure 2.1 is an ester? 1) A) a b c d e Answer: D 2) A scientist
Exam Name MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. Figure 2.1 1) Which compound in Figure 2.1 is an ester? 1) A) a b c d e Answer: D 2) A scientist
Biological Pathways Representation by Petri Nets and extension
 Biological Pathways Representation by and extensions December 6, 2006 Biological Pathways Representation by and extension 1 The cell Pathways 2 Definitions 3 4 Biological Pathways Representation by and
Biological Pathways Representation by and extensions December 6, 2006 Biological Pathways Representation by and extension 1 The cell Pathways 2 Definitions 3 4 Biological Pathways Representation by and
Nitric Oxide Synthase Ultrasensitive Colorimetric Assay
 Package Insert Nitric Oxide Synthase Ultrasensitive Colorimetric Assay 96 Wells For Research Use Only v. 2.0 09.20.17 Eagle Biosciences, Inc. 20A NW Blvd., Suite 112, Nashua, NH 03063 Phone: 866-419-2019
Package Insert Nitric Oxide Synthase Ultrasensitive Colorimetric Assay 96 Wells For Research Use Only v. 2.0 09.20.17 Eagle Biosciences, Inc. 20A NW Blvd., Suite 112, Nashua, NH 03063 Phone: 866-419-2019
For the rapid, sensitive and accurate detection of active Caspase 9 in living cells
 ab65615 Caspase 9 (active) FITC Staining Kit Instructions for Use For the rapid, sensitive and accurate detection of active Caspase 9 in living cells This product is for research use only and is not intended
ab65615 Caspase 9 (active) FITC Staining Kit Instructions for Use For the rapid, sensitive and accurate detection of active Caspase 9 in living cells This product is for research use only and is not intended
RayBio Human IDUA ELISA Kit
 RayBio Human IDUA ELISA Kit Catalog #: ELH-IDUA User Manual Last revised April 15, 2016 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross, GA
RayBio Human IDUA ELISA Kit Catalog #: ELH-IDUA User Manual Last revised April 15, 2016 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100 Norcross, GA
Dominance of Old End Growth is Inherited in Fission Yeast
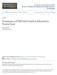 University of Tennessee, Knoxville Trace: Tennessee Research and Creative Exchange University of Tennessee Honors Thesis Projects University of Tennessee Honors Program 5-2015 Dominance of Old End Growth
University of Tennessee, Knoxville Trace: Tennessee Research and Creative Exchange University of Tennessee Honors Thesis Projects University of Tennessee Honors Program 5-2015 Dominance of Old End Growth
Bioinformatics 3. V18 Kinetic Motifs. Fri, Jan 8, 2016
 Bioinformatics 3 V18 Kinetic Motifs Fri, Jan 8, 2016 Modelling of Signalling Pathways Curr. Op. Cell Biol. 15 (2003) 221 1) How do the magnitudes of signal output and signal duration depend on the kinetic
Bioinformatics 3 V18 Kinetic Motifs Fri, Jan 8, 2016 Modelling of Signalling Pathways Curr. Op. Cell Biol. 15 (2003) 221 1) How do the magnitudes of signal output and signal duration depend on the kinetic
RayBio Rat TNF-alpha ELISA Kit (For Lysates)
 RayBio Rat TNF-alpha ELISA Kit (For Lysates) Catalog #: ELR-TNFa-CL User Manual Last revised April 15, 2016 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite
RayBio Rat TNF-alpha ELISA Kit (For Lysates) Catalog #: ELR-TNFa-CL User Manual Last revised April 15, 2016 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite
NOVABEADS FOOD 1 DNA KIT
 NOVABEADS FOOD 1 DNA KIT NOVABEADS FOOD DNA KIT is the new generation tool in molecular biology techniques and allows DNA isolations from highly processed food products. The method is based on the use
NOVABEADS FOOD 1 DNA KIT NOVABEADS FOOD DNA KIT is the new generation tool in molecular biology techniques and allows DNA isolations from highly processed food products. The method is based on the use
5.1 Cell Division and the Cell Cycle
 5.1 Cell Division and the Cell Cycle Lesson Objectives Contrast cell division in prokaryotes and eukaryotes. Identify the phases of the eukaryotic cell cycle. Explain how the cell cycle is controlled.
5.1 Cell Division and the Cell Cycle Lesson Objectives Contrast cell division in prokaryotes and eukaryotes. Identify the phases of the eukaryotic cell cycle. Explain how the cell cycle is controlled.
Symbol Definition Value Range (Experimental) Time of the end of residual growth 10 h 10
 Table 1. Definitions and experimental values of parameters Symbol Definition Value ange (Experimental) T I Time of the end of residual growth 10 h 10 T D D D ate s s G max V max c-fed c-fed Delay in Death
Table 1. Definitions and experimental values of parameters Symbol Definition Value ange (Experimental) T I Time of the end of residual growth 10 h 10 T D D D ate s s G max V max c-fed c-fed Delay in Death
bacteriologist has not sufficient chemical training or the time to
 THE VAN SLYKE METHOD FOR THE DETERMINATION OF AMINO-ACID NITROGEN AS APPLIED TO THE STUDY OF BACTERIAL CULTURES R. W. LAMSON From the Department of Bacteriology and Immunity, Harvard Medical School Received
THE VAN SLYKE METHOD FOR THE DETERMINATION OF AMINO-ACID NITROGEN AS APPLIED TO THE STUDY OF BACTERIAL CULTURES R. W. LAMSON From the Department of Bacteriology and Immunity, Harvard Medical School Received
ACCELERATE ITS BIOCHEMICAL PROCESSES WHICH WERE SLOWED DOWN BY MITOSIS. THE LENGTH OF THE G1 PHASE CREATES THE DIFFERENCE BETWEEN FAST DIVIDING
 CHAPTER 1: OVERVIEW OF THE CELL CYCLE THE THREE STAGES OF INTERPHASE: INTERPHASE BEFORE A CELL CAN ENTER CELL DIVISION, IT NEEDS TO PREPARE ITSELF BY REPLICATING ITS GENETIC INFORMATION AND ALL OF THE
CHAPTER 1: OVERVIEW OF THE CELL CYCLE THE THREE STAGES OF INTERPHASE: INTERPHASE BEFORE A CELL CAN ENTER CELL DIVISION, IT NEEDS TO PREPARE ITSELF BY REPLICATING ITS GENETIC INFORMATION AND ALL OF THE
MARK SCHEME for the October/November 2012 series 9700 BIOLOGY
 CAMBRIDGE INTERNATIONAL EXAMINATIONS GCE Advanced Subsidiary Level and GCE Advanced Level MARK SCHEME for the October/November 2012 series 9700 BIOLOGY 9700/51 Paper 5 (Planning, Analysis and Evaluation),
CAMBRIDGE INTERNATIONAL EXAMINATIONS GCE Advanced Subsidiary Level and GCE Advanced Level MARK SCHEME for the October/November 2012 series 9700 BIOLOGY 9700/51 Paper 5 (Planning, Analysis and Evaluation),
LABORATORY OF ELEMENTARY BIOPHYSICS
 LABORATORY OF ELEMENTARY BIOPHYSICS Experimental exercises for III year of the First cycle studies Field: Applications of physics in biology and medicine Specialization: Molecular Biophysics Fluorescence
LABORATORY OF ELEMENTARY BIOPHYSICS Experimental exercises for III year of the First cycle studies Field: Applications of physics in biology and medicine Specialization: Molecular Biophysics Fluorescence
The Effects of the Carbon Source on Glutamate Dehydrogenase Activities in Aspergillus nidulans
 Joiirnal of General Microbiology ( I 974), 81, I 65-1 70 Printed in Gseat Britnin The Effects of the Carbon Source on Glutamate Dehydrogenase Activities in Aspergillus nidulans By M. J. HYNES Department
Joiirnal of General Microbiology ( I 974), 81, I 65-1 70 Printed in Gseat Britnin The Effects of the Carbon Source on Glutamate Dehydrogenase Activities in Aspergillus nidulans By M. J. HYNES Department
CELL CYCLE AND DIFFERENTIATION
 CELL CYCLE AND DIFFERENTIATION Dewajani Purnomosari Department of Histology and Cell Biology Faculty of Medicine Universitas Gadjah Mada d.purnomosari@ugm.ac.id WHAT IS CELL CYCLE? 09/12/14 d.purnomosari@ugm.ac.id
CELL CYCLE AND DIFFERENTIATION Dewajani Purnomosari Department of Histology and Cell Biology Faculty of Medicine Universitas Gadjah Mada d.purnomosari@ugm.ac.id WHAT IS CELL CYCLE? 09/12/14 d.purnomosari@ugm.ac.id
Enzyme Catalysis Lab
 AP Biology Name: Enzyme Catalysis Lab Objectives In this laboratory, you will observe the role of an enzyme (catalase) in conversion of hydrogen peroxide (H 2 O 2 ) to water and oxygen determine the rate
AP Biology Name: Enzyme Catalysis Lab Objectives In this laboratory, you will observe the role of an enzyme (catalase) in conversion of hydrogen peroxide (H 2 O 2 ) to water and oxygen determine the rate
Initiation of nuclear reactions under laser irradiation of Au nanoparticles in the aqueous solution of Uranium salt. A.V. Simakin and G.A.
 Initiation of nuclear reactions under laser irradiation of Au nanoparticles in the aqueous solution of Uranium salt A.V. Simakin and G.A. Shafeev Wave Research Center of A.M. Prokhorov General Physics
Initiation of nuclear reactions under laser irradiation of Au nanoparticles in the aqueous solution of Uranium salt A.V. Simakin and G.A. Shafeev Wave Research Center of A.M. Prokhorov General Physics
RayBio Human Thyroid Peroxidase ELISA Kit
 RayBio Human Thyroid Peroxidase ELISA Kit Catalog #: ELH-TPX User Manual Last revised July 6, 2017 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100
RayBio Human Thyroid Peroxidase ELISA Kit Catalog #: ELH-TPX User Manual Last revised July 6, 2017 Caution: Extraordinarily useful information enclosed ISO 13485 Certified 3607 Parkway Lane, Suite 100
Enzymatic Assay of PROTEIN KINASE C
 PRINCIPLE: Histone +? 32 P-ATP Protein Kinase > [ 32 P]-Phosphorylated Histone + ADP Abbreviations used:? 32 P-ATP = Adenosine 5'-Triphosphate? 32 P-labelled ADP = Adenosine 5'-Diphosphate CONDITIONS:
PRINCIPLE: Histone +? 32 P-ATP Protein Kinase > [ 32 P]-Phosphorylated Histone + ADP Abbreviations used:? 32 P-ATP = Adenosine 5'-Triphosphate? 32 P-labelled ADP = Adenosine 5'-Diphosphate CONDITIONS:
SDS-PAGE (Sodium Dodecyl Sulfate Polyacrylamide Gel Electrophoresis):
 SDS-PAGE (Sodium Dodecyl Sulfate Polyacrylamide Gel Electrophoresis): Aim: SDS-PAGE (Sodium Dodecyl Sulfate Polyacrylamide Gel Electrophoresis) is one of the common methods used in the molecular biology
SDS-PAGE (Sodium Dodecyl Sulfate Polyacrylamide Gel Electrophoresis): Aim: SDS-PAGE (Sodium Dodecyl Sulfate Polyacrylamide Gel Electrophoresis) is one of the common methods used in the molecular biology
AP Biology Summer Assignment
 AP Biology Summer Assignment 2017-18 Students must complete this assignment by the first week of school. The first exam, which will be the first week of school, will cover the information in this packet.
AP Biology Summer Assignment 2017-18 Students must complete this assignment by the first week of school. The first exam, which will be the first week of school, will cover the information in this packet.
Cell-Cycle Analysis of Fission Yeast Cells by Flow Cytometry
 Cell-Cycle Analysis of Fission Yeast Cells by Flow Cytometry Jon Halvor Jonsrud Knutsen 1, Idun Dale Rein 2, Christiane Rothe 1,3, Trond Stokke 2, Beáta Grallert 1,Erik Boye 1,3 * 1 Department of Cell
Cell-Cycle Analysis of Fission Yeast Cells by Flow Cytometry Jon Halvor Jonsrud Knutsen 1, Idun Dale Rein 2, Christiane Rothe 1,3, Trond Stokke 2, Beáta Grallert 1,Erik Boye 1,3 * 1 Department of Cell
