For the Degree of MASTER OF SCIENCE. Denton, Texas. August, 1988
|
|
|
- Annice Townsend
- 6 years ago
- Views:
Transcription
1 379 QUANTITATION OF ENDOGENOUS NUCLEOTIDE POOLS IN Pseudomonas aeruginosa TESIS Presented to the Graduate Council of the University of North Texas in Partial Fulfillment of the Requirements For the Degree of MASTER OF SCIENCE By Mohammad Entezampour, B. S. Denton, Texas August, 1988
2 Entezampour, Mohammad, Quantitation of Endogenous Nucleotide Pools in Pseudomonas aeruginosa. Master of Science (Biology),, August, 1988, 54 pp., 7 illustrations, 7 tables, bibliography, 43 titles. Nucleotide pools were extracted and quantified from Pyr+ and Pyr~ strains of P. aerucjinosa. Strains were grown in succinate minimal medium with and without pyrimidines, and nucleotides were extracted using trichloracetic acid (TCA; 6% w/v). The pyrimidine requirement was satisfied by uracil, uridine, cytosine or cytidine. Pyr~ mutants were starved for pyrimidines for two hours before nucleotide levels were measured. This starvation depleted the nucleotide pools which were restored to wild type levels by the addition of pyrimidines to the medium. When the pyrimidine analogue, 6-azauracil, known to inhibit OMP decarboxylase, was added to cultures of the wild type strain, the uridine and cytidine nucleotides were depleted to near zero. Thus, the nucleotide pool levels of Pseudomonas strains can be manipulated.
3 TABLE OF CONTENTS List of Tables.....iv Page List of Illustrations v Chapter I. INTRODUCTION & 1 II. MATERIALS AND METODS... Chemicals and Reagents Bacterial Strains Growth Media and Cultures Extraction of Nucleotides Chromatographic Apparatus Chromatographic Conditions Identification and Quantitation of Nucleotides Calculations 14 III. RESULTS IV. DISCUSSION BIBLIOGRAPY... -~~~~~~ iii
4 LIST OF TABLES Table Page I. Genotypes and Phenotypes of Pyrimidine Auxotrophic Mutants in Bacteria II. III. IV. Identification of the Twelve Nucleotide Standards Used in This Work..... Nucleoside Trighosphate in umol (gram dry weight) from P. aeruginosa PAO-1 Wild Type Strain in the Presence and Absence of the Listed Additions. Nucleoside Triphosphates in umol (gram dry weight) -l from P. aeruginosa pyrb Mutant in the Presence of Uracil, Cytosine and Cytidine V. Nucleoside Tri hosphates in umol (gram dry weight)' from P. aeruginosa pyrf Mutant in the Presence and Absence of Uracil, Cytosine and Cytidine VI. VII. Nucleoside Triyhosphates in umol (gram dry weight) from P putida PPN Wild Type Strain in the Presence of the Listed Additions Nucleoside Triyhosphates in umol (gram dry weight)~ from P putida pyrb Mutants in the Presence of Uracil, Cytosine, and Cytidine iv
5 LIST OF ILLUSTRATIONS Figure Page 1. Pyrimidine nucleotide biosynthetic and salvage pathways in Escherichia coli and Salmonella typhimurium Schematic diagram representing the extraction procedure for ribonucleotides from P. aeruginosa Chromatograph of the 12 ribonucleotide standards Elution profile of the ribonucleotides extracted with 6% (w/v) trichloroacetic acid from P. aeruginosa Growth curves for P. aerucinosa PAO-1 wild type strain in succinate minimal (SM) medium Growth curve for P. aeruginosa pyrb strain in succinate minimal (SM) medium at 37C with and without pyrimidines Growth curve for P. aeruginosa yrf strain in succinate minimal (SM) medium at 37C with and without pyrimidines.. 32 v
6 CAPTER I INTRODUCTION The pyrimidine biosynthetic pathway is universal with the same sequence of reactions (Fig. 1) found for every organism (O'Donovan & Neuhard, 197). The first step in the pathway catalyzed by the enzyme carbamoyl phosphate synthetase (E C ; CPSase) forms carbamoyl phosphate from bicarbonate, the amide group of glutamine and ATP (Anderson & Meister, 1966). Carbamoyl phosphate is used to synthesize both arginine and pyrimidines in approximately equal amounts (Abdel,.etal., 1969). For pyrimidine biosynthesis, carbamoyl phosphate condenses with aspartate to form carbamoyl aspartate and inorganic phosphate (Gerhart & Pardee, 1962). This step is catalyzed by the enzyme aspartate transcarbamoylase (E C ; ATCase). For arginine biosynthesis, ornithine and carbamoyl phosphate combine to form citrulline and inorganic phosphate. This step is catalyzed by ornithine transcarbamoylase (E C ; OTCase) (Isaac & olloway, 1972). The third step in the pyrimidine pathway, catalyzed by dihydroorotase (E C ; DOase), converts carbamoyl aspartate to the cyclic dihydroorotate by the removal of water (Lieberman & Kornberg, 1953). Dihydroorotate dehydrogenase (E C ; DOdehase) produces orotate in the fourth 1
7 2 Fig. 1. Pyrimidine nucleotide biosynthetic and salvage pathways in Escherichia coli and Salmonella typhimurium. Abbreviations are: U, uracil; C, cytosine; UR, uridine; CR, cytidine; OMP, orotidine 5'-monophosphate; UMP, uridine 5'- monophosphate; UDP, uridine 5'-diphosphate; UTP, uridine 5'- triphosphate; CMP, cytidine 5'-monophosphate; CDP, cytidine 5' -diphosphate; CTP, cytidine 5' -triphosphate.
8 . 3 CTP PyIG CD P SALVAGE PATWAYS UTP CMP SALVAGE PATWAYS- de novo PATWAYS - -- N I CR4 f Cwft I- pyr LUP pyrft AOOOOOO N%%6*1ft UR U 44 - UR - U CMP Orotate pyrd Dihydroorotate t c ~.6 - C CELL. MEMBRANE Arginine Argininosuccinate Carbamoyl Aspartate D Citrulline Q Ornithine pyrb ftcase Aspartate Carbamoyl- (Z) pyra Glutamine ATP CO3 3
9 4 fourth step (Lieberman & Kornberg, 1953). Orotate and phosphoribosyl pyrophosphate (PRPP) combine to form the first pyrimidine ribonucleotide, orotidine-5-monophosphate (OMP) (urlbert and Reichard, 1955) in a reaction catalyzed by orotate phosphoribosyl transferase (EC ; OPRTase). Inorganic pyrophosphate is also a product. The final step on uridine-5'-monophosphate (UMP) biosynthesis is the decarboxylation of OMP to form UMP (Lieberman et al., 1955). This sixth step is catalyzed by OMP decarboxylase (EC ; OMP decase). The same workers (Lieberman et al., 1955) showed that the UMP is now converted to UTP in two kinase steps: first by the highly specific UMP klinase (EC ) to form UDP, and then UTP is converted to UTP by the non-specific nucleoside diphosphate kinase (EC ; NDKase). Finally UTP is aminated to produce CTP (Lieberman, 1955) by the enzyme CTP synthetase (EC ; CTPSase). The pathway in Escherichia coli is controlled at three sites at the level of enzyme activity by feedback inhibition and by activation. Specifically (i) CPSase is feedback inhibited by UMP and activated by ornithine (Pierard, 1966); (ii) ATCase is feedback inhibited by CTP and activated by ATP (Gerhart & Pardee, 1964); and (iii) CTP synthetase is inhibited by CTP and activated by UTP (Long & Pardee, 1967). The E. coli pyrimidine pathway is also controlled at the level of enzyme synthesis. When E.
10 5 coli is grown in uracil or other pyrimidines, such as uridine, cytidine, or cytosine, all enzymes of the pathway are repressed. Thus, when E. coli is grown in minimal medium (without uracil) ATCase specific activity is four times greater than when the cells are grown in minimal medium plus uracil (Kelln et a., 1975). Table 1 shows the genotypes of wild type and of mutant strains that lack pyrimidine enzymes in bacteria. All of these enzymes are found in E. coli, Salmonella typhimurium and in Pseudomonas aeruginosa, the organism being studied in this research. owever, there are some major differences in the pyrimidine pathway of Pseudomonas from the pathway in E. coli. For instance it has not been possible to repress the synthesis of the pyrimidine enzymes by growth on exogenous pyrimidines such as uracil, uridine, cytosine or cytidine (Condon et al., 1976). The pathway is controlled by feedback inhibition by UTP rather than by CTP. Indeed UTP is the primary effector of ATCase in P. aeruginosa (Isaac & olloway, 1968; Neumann & Jones, 1964). Mutants exist in pyrb, pyrd, pyre and pyrf of P. aeruginosa pathway. These will be examined in this present study. When Pyr~ mutants of E. coli are grown in uracil and then starved for uracil, the enzymes of the pathway become derepressed over 1-fold (i.e., their specific activity increases over 1-fold). At the same time the concentration of nucleoside triphosphates, UTP and CTP, are depleted and drop to near zero levels
11 6 Table I. Genotypes and Phenotypes of Pyrimidine Auxotrophic Mutants in Bacteria.
12 7 C ' C *rr4.4 4 ) p - -r4 r- r- r-4 r- r- r- q.4 r- u Q U u z 4 o o o c e o41 c E-.4 o OP Cd W E.. CnM M4P4 p 4 O En 4) (1) cd Oco U) MCd 4) 44 (1 -r4 4-J X r-4 o 4) P 4'-d P to 4) r- O E "Ag P >-'C PL ' co 4 U ) - QOr-4 TI > ON % O - -'P n oo > Cdo En -* 4 Z o r-4 (1) a 4J d fr- 1-4J C4) U Q z p d Mdm >< 4 4-) cd o4j4-)o J4CU O E- 4 r-4 (1) p p p lc -A >- 4-) C 1 4J ' ', P.N P4 z P644m CO 1< -A - P Z P u aaaa
13 8 (Kelln et al., 1975). When uracil is restored to the culture, the UTP and CTP pools expand to wild type levels and the enzymes are no longer derepressed (Kelln et al., 1975). Whey Pyr~ mutants of P aeruginosa are starved for uracil there is a slight increase (2-fold) in ATCase specific activity but it has not been determined if UTP and CTP decreased. In this investigation the levels of UTP and CTP (as well as other nucleotides) are examined in wild type nad in Pyr~ mutants of P. aeruginosa and P. putida under different conditions of growth in order to ascertain the levels of nucleotides. Nucleotides are important compounds for the synthesis of macromolecules in all living cells. Nucleotides are phosphorylated, heterocyclic compounds classified under two major groups known as purines and pyrimidines. Two purine bases, adenine and guanine, and three pyrimidine bases, uracil, thymine and cytosine are the major nitrogenous bases for nucleotide make up. The ribonucleoside triphosphates, uridine triphosphate (UTP), cytidine triphosphate (CTP), adenosine triphosphate (ATP) and guanosine triphosphate (GTP) have different important functions within the cell. These include (a) control of biosynthetic pathways; (b) synthesis of RNA UTP, CTP, ATP and GTP) and DNA (dttp, dctp, datp and dgtp); (c) nucleotides are carried of phosphate and pyrophosphate in several enzymatic reactions involved in the transfer of chemical energy; (d) and serve as coenzyme-like,
14 9 energized carriers of specific types of building block molecules. Several techniques have been developed (Chargeys & Davidson, 1955) for the separation and quantification of nucleic acids and related substances (a) enzymatic analysis where specificity of enzymes is used to determine the presence of specific substances (Strehler & McElroy, 1957) (b) paper electrophoresis where nucleic acids and related compounds, due to a variety of ionizable groups can be separated and their concentrations measured according to differences in net charge of the different species at given p values; (c) planar chromatrography where the stationary phase assumes a planar arrangement in the form of a sheet or thin film and the mobile phase migrates by means of capillary action, separating the various species according to their differential solubilities. Planar chromatography includes thin layer chromatography and paper chromatography; (d) column chromatography where the stationary phase is either a solid packing material, a solid support that is coated with a sorbent latex or gel, any of which can be packed into a tube, a migrating liquid phase serves to dissolve the various substances according to their differential solubilities. Gas chromatography, ion-exchange chromatography and liquid chromatography are three forms of column chromatography. Classical liquid chromatography has been improved by the addition of high pressure on the moving phase (PLC). In recent years, the measurement of
15 1 endogenous ribo- and deoxyribonucleotide concentrations has been revolutionized by the development of PLC (Glaser, 1986). The two most commonly employed separation techniques involve either reverse-phase chromatography on octyldecylsilica resins (offman & Liao, 1977; orvath, Melander & Molnar, 1977) or ion-exchange chromatography on microparticulate anion-exchange resins (artwick & Brown, 1975; McKeag & Brown, 1978). Even though reverse-phase chromatography provides greater sensitivity and more rapid analysis compared to any ion-exchange chromatographic method currently available, it lacks the selectivity necessary for the analysis of the many closely related nucleotides present in cell extracts (Reiss, Zuurendonk, & Veech, 1984). In our laboratory, PLC is employed simply because efficient, rapid and reliable quantitative nucleotide separations are needed. In the present study and in previous studies, Partisil 1 SAX columns were used. These are strong anion-exchange columns which are packed with particles 1um in mean size. Because of the sensitivity of these microparticle packings to various chemicals, the extraction procedure is crucial not only in attaining optimal sensitivity and column efficiency but also in protecting the column for maximal life (artwick & Brown, 1975). It has been found that even trace amounts of perchlorate may bring about rapid deterioration of this kind of column. In addition to this, neutralization of
16 (Table II) adenosine 5'-triphosphate (ATP), adenosine 5' - (GDP), guanosine 5' -monophosphate (GMP), uridine 5' - 11 perchloric acid with tris (hydroxymethyl) aminoethane which is used in a common, rapid, and simple method of preparing extracts for use with some columns, cannot be used with these packings (Brown & Miech, 1972). On the other hand, multiple ether extractions are time-consuming and small amounts of nucleotides may be partitioned into the etherwater interphase causing loss of some nucleotides and lower total nucleotide values on analysis. The extraction procedure described by Khym (Khym, 1975) is the best studied for use with these microparticle, chemically bonded columns. Trichloroacetic acid (TCA) is used to extract the nucleotides from cellular material and is subsequently removed from cell extracts with a Freon-amine solution. The objective of the present investigation was to compare the profiles of 12 acid-soluble ribonucleotides diphosphate (ADP), adenosine 5' -monophosphate (AMP) guanosine 5' -triphosphate (GTP), guanosine 5' -diphosphate triphosphate (UTP), uridine 5' -diphosphate (UDP), uridine 5' -monophosphate (UMP), cytidine 5' -triphosphate (CMP), cytidine 5' -diphosphate (CDP), and cytidine 5' - monophosphate (CMP) during the growth of P. aeruginosa on succinate minimal medium.
17 12 Table II. Identification of the 12 Nucleotide Standards Used in this Work by Their Retention Times (Minutes).
18 13-4-) - Or oin r W r- - NSNS N m* qcs q gc onwwrk- NON S- L E- C-4 L O Fe C. 4-) 4 N NG 4-) rzie l 111 E-4 4-) 4.) - 4J 4- J o 4 - MO -)P 44 d O Cdi4,c Q4 4 4 m )-P Ci4-),4J(,S S Q 4) Wd 4 w 4) (Cl. 4 - U) Q4 o -,, ( cmo 4 En ~ Q 4 U)W4WO M1, 44 M 4 o , 4 O 4., ,, - - Z M 9 ct$-t$,d F -)--Pr: - I 4 I J 4 I I I O I LO LO 1 I I)O UC) o- % %blc (O1 LnOO)LO OO) 4 U (1) ( - M) U)M -U U) E - M)tU Fd - -Fe F Fe- 99Fe d - -) 4d - - () S 5-5 r 1r 4-i-P 4 SQSQ z z - Cc oo-8oc %- CC # -8 M-dy & -Sy&M -'C OS-
19 CAPTER II MATERIALS AND METODS Chemicals and Reagents Nucleotides, trichloroacetic acid (TCA) and tri-noctylamine were purchased from Sigma Chemical Company (St. Louis, Missouri); monobasic ammonium phosphate from Mallinckrodt Inc. (Paris, Kentucky): and 1,1, 2-trichloro 1,2,2-trifluoroethane (Freon) from Eastman-Kodak Company (Rochester, New York). All other chemicals were of analytical grade and were purchased from Fisher Scientific Company (Fair Lawn, New Jersey). Bacterial Strains Pseudomonas aeruginosa PAO and P. putida PPN wild types and their Pyr~ mutants pyrb, pyrd, pyre, and pyrf were obtained from Professor Bruce W. olloway, Department of Genetics, Monash University, Clayton, Victoria, Australia.) The pyrb mutant strain is the only pyrimidine mutant available for P. putida. Growth Media and Cultures Bacterial cells were grown in Basal Minimal Medium (Cohen-Bazire, Sistrom, & Stanier, 1957) containing the following chemical composition in grams per liter of deionized water: Na 2 PO 4, 7.1; K 2 PO 4, 6.8; (N 4 )2So 4, 14
20 (N 4 ) 6 Mo 7 O , 18.5; ZnSO ; FeSO ; MnSO nitrilotriacetic acid, MgSO 4, 28.9; CaCI 2, 6.67; 2, CuSO , Co(NO 3 ) , Na 2 B , (utner's metals '44'.) was used with succinate as the carbon and energy source. Pyr~ mutants were supplemented with uracil at 5 ug/ml. These were periodically checked for purity on basal minimal agar plates. Gram stain tests and other appropriate morphological observations were made. All experiments were started by streaking the organism on basal minimal agar. Cells of P. aeruginosa from a single, isolated colony were inoculated into basal minimal liquid medium and incubated on a rotatory shaker at 37C and agitated at 9 revolutions per minute. P. putida strains were grown at 3C. Overnight cultures was transferred to a fresh basal medium and grown to about 1 KU. Growth was measured by turbidity with a Photoelectric Klett Summerson Colorimeter (Klett Manufacturing Co., New York, N.Y.) using a green filter #54 and recorded as Klett Units (IKU=1 7 /ml). The logarithms of the KUs were plotted against time. Extraction of Nucleotides Volumes of 5 ml of bacterial cultures were harvested at about 1 KU, centrifuged at 4 C at 12, x g for 5 minutes. After decanting the supernatant, nucleotides were extracted from the cell pellet according to the scheme shown in Figure 2. One ml of ice-cold 6% (w/v) trichloroacetic
21 16 Fig. 2. Schematic diagram representing the extraction procedure for ribonucleotides from P. aeruginosa cultures.
22 17 Bacterial Culture Centrifuge at 12, X g for 5 min at 4 C Supernatant Cell pellet Add TCA (6% w/v) Mix by vortexing Allow to stand for 3 min Centrifuge at 12, X g for 15 min at 4 C Pellet F- -+ Supernatant Add Freon-amine, vortex, and stand Bottom Layer Top Aqueous Layer contains nucleotides Filter Store at -2C for analysis.4&"- 444t ilw-
23 18 acid (TCA) was added to the cell pellet, which was then thoroughly mixed for 2 minutes in the vortex mixer. The mixture was allowed to stand at 4C for 3 minutes before centrifuging at 12, x g for 15 minutes. The clear supernatant was then neutralized with ice cold Freon-amine (Khym, 1975) solution (1.6 ml of.7 M tri-n-octylamine in 5 ml of Freon 113). The sample was then mixed on the vortex mixer for 2 minutes then allowed to separate for 15 minutes at 4C. The top, aqueous layer, which contained the nucleotides, was removed, filtered through a.45 um ACRO LCl3 filter (Gelman Sciences, Ann Arbor, Michigan) and frozen at -2C until ready for analysis. Chromatographic Apparatus The PLC equipment (Waters Assoc., Milford, Massachusetts) consisted of two Model 51 pumps, a Model 68 automated gradient controller, a U6K injector, and a Model 481 LC spectrophotometer. Nucleotides were detected by monitoring the column effluent at 254nm with a sensitivity fixed at.5 absorbance units at full scale deflection (AUFS). Separations were performed on a Waters Radial-Pak Partisil SAX cartridge (1 cm x.8 cm) using a Waters radial compressing Z-Module system. Chromatographic Conditions The entire chromatographic system including the column was stored in 5:5 (v/v) filtered PLC grade methanol and
24 19 filtered, double distilled water (2x) when not in use. After priming the pumps, the system was flushed with 5 ml of methanol:water mixture at 3 ml per minute. Next, the system was thoroughly washed with distilled water with the initial flow rate at 3 ml per minute. After 1 minutes the flow rate was increased to 4 ml per minute. When the back pressure of the column dropped to 85 pounds per square inch, the methanol:water mixture was completely washed from the system. Pump A was flushed with starting buffer (filtered ultra pure, 7 mm monobasic ammonium phosphate, p 3.8) followed by pump B which was flushed separately with final buffer (filtered, 25 mm monobasic ammonium phosphate containing 5 mm potassium chloride, p 4.5). The LC spectrophotometer was set at 254 nm and.5 AUFS, and the recorder and automated gradient controller were turned on. An initial program with a linear slope (curve profile #6) of low concentration buffer was run for 1 minutes. A 1- minute reverse gradient of high concentration buffer was run, followed by a 1-minute rest with low concentration buffer. Identification and Quantitation of Nucleotides Nucleotide samples of 2 ul each were prepared from Pseudomonas cells as previously described. These were thawed and injected onto a column of Partisil SAX-1 which consisted of a silica matrix coated with porous
25 2 microparticles of silica of 1 um particle size. The particles contained fixed-charge quaternary nitrogen groups and mobile counterions of 2 PO 4. Nucleotides bind to the quaternary nitrogen group with different affinities because of the functional groups in the bases and the number of phosphates at the C-5' of the sugar. A linear gradient (curve profile #6) of low to high concentration buffer was applied for 2 minutes, followed by an isocratic period of 1 minutes of high concentration buffer. The column was regenerated by washing with 3 ml of 7mM N 4 2 PO 4, ph 3.8 buffer (Pogolotti & Santi, 1982). The flow rate was maintained at 4 ml per minute and all analyses were done at ambient temperature. Peaks were integrated using a Waters Data Module 74 attached to a microprocessor. The sample peaks were identified by comparing their retention times with those of appropriate standards. The concentration of nucleotides in samples was calculated by comparing peak heights to standards (ATP, ADP, AMP, GTP, GDP, GMP, UTP, UDP, UMP, CTP, CDP, and CMP) of known concentration (.1 mm) and expressed as nmoles per 18 cells. The system was terminated by first flushing with distilled water and then with the methanol-water mixture in the same manner as used to start the apparatus.
26 21 Calculations Moles (gram dry weight)~ 1 for all nucleotides were computed as follows: Sa V 1 X C X X St Vi Dw Where Sa = peak height of sample, St = peak height of standard, C = grams compound in standard/molecular weight of compound, V = total volume of extract, Vi = volume of extract injected and Dw = dry weight (Dutta & O'Donovan, 1987).
27 CAPTER III RESULTS The purpose of this research was to ascertain if the nucleoside mono-, di-, and triphosphates could be altered in Pseudomonas strains. To find the answer to this question, wild type P. aeruginosa, strain PAO-l, and two P. aeruginosa PAO-1 mutant derivatives namely yrb and pyrf, were employed. Strains were grown in succinate minimal medium in the presence and absence of uracil, uridine, cytosine and cytidine. In addition, the wild type strain was grown in the presence of 6-azauracil, a pyrimidine analogue known to inhibit the enzyme OMP decarboxylase in E. coli thereby starving the cells for pyrimidines. All P. aeruginosa cultures were grown under identical conditions on a rotary shaker water bath at 37 C. Bases and nucleosides were added at the rate of 5 ug/ml and pyrimidine auxotrophs were starved for pyrimidines for two hours at which time samples were taken. Typically an experiment was carried out as follows: 1 to 2 ml of succinate minimal medium were inoculated with the culture to be tested in the presence (at 5 ug/ml) or absence of appropriate additions. The cells were grown to a density of 1 Klett units (1 9 /ml). Fifty ml samples of culture were then harvested and treated with 6%(w/v) trichloroacetic acid to extract the nucleotides as 22
28 23 Fig. 3. Chromatogram of the 12 ribonucleotide standards. The numbers refer to the compounds listed in Table II.
29 24 co. C cz 2 D 3 4 t[[ imue ( nu.s Time (minutes)
30 25 Fig. 4. Elution profile of ribonucleotides extracted with 6% (w/v) trichloroacetic acid from P. aeruginosa. The numbers refer to the compounds listed in Table II.
31 26 II (T Cu C C LL Time (minutes)
32 27 Fig. 5. Growth curves for P. aeruginosa PAO-1 wild type strain in succinate minimal (SM) medium at 37 C with and without the noted additions. 6-Azauracil was added to the growing culture in SM medium at 9 KU. Symbols and abbreviations are: SM+CAA+U, casamino acid + uracil (), SM+U, uracil (a), SM+C, cytosine (A), SM+CR, Cytidine, (U), SM+UR, uridine, (a) SM+AzaU, 6-azauracil (A).
33 28 S.12 A Time (min)
34 29 Fig. 6. Growth curve for P. aeruginosa pyrb strain in the succinate minimal (SM) medium at 37 with and without pyrimidines. When the cultures reached 1 KU they were starved for pyrimidines for two hours. Symbols and abbreviations are as in Fig. 5.
35 + 3 a I Time (min)
36 31 Fig. 7. Growth curve for P. aeruinosa pyrf strain in the succinate minimal (SM) medium at 37 C with and without pyrimidines. When the cultures reached 1 KU they were starved for pyrimidines for two hours. Symbols and abbreviations are as in Fig. 6.
37 32 C Time (min)
38 33 as described in Materials and Methods section. Nucleotides were quantitated by PLC. Growth curves for the P. aeruginosa wild type PAO-1 strain are shown in Fig. 5 where the log of the Klett Units (KU) is plotted against time. As can be seen from Fig. 5, the PAO-l strain grows equally well on succinate minimal medium and on all supplements. When 6-azauracil was added (denoted by the arrow in Fig. 5) growth ceased. After one hour in 6-azauracil a sample was taken for PLC analysis of nucleotides. Growth curves for the P. aeruginosa pyrb strain are shown in Fig. 6. A previous calculation indicated the amount of exogenous pyrimidine base or nucleoside that was needed for the auxutroph to reach 1 KU. This is seen in Fig. 6 for uracil and uridine, which grow exponentially to about 1 KU before running out of pyrimidine nutrient. After two hours of such starvation, samples were taken. Growth on cytidine was poor as reflected by the slower growth rate as seen in Fig. 6. Growth curves for P. aeruginosa pyrf are shown in Fig. 7. In this figure the results are identical to those seen in Fig. 6. Once again growth on cytidine was poor. Ribonucleoside mono-, di-, and triphosphates were measured for each of the growth conditions described in Figures 5, 6, and 7. Profiles of the 12 ribonucleoside standards, UMP, CMP, AMP, GMP, UDP, CDP, ADP, GDP, UTP,
39 34 CTP, ATP AND GTP are shown in Fig. 3. Since growth is reflected best by the concentration of triphosphate, in this thesis only the nucleoside triphosphate levels are shown. Table III shows the nucleoside triphosphates for the different conditions of growth of the wild type P. aeruginosa strain PAO-l. As can be seen from Table II the overall levels of the triphosphates in P. aeruginosa are similar to those reported earlier for Salmonella typhimurium by Kelln et al., When uracil or uridine was added to the growth cultures, the UTP level increased two-fold from 5.23 umol/(gram dry weight)~ 1 to 1. umol(gram dry weight)-1. One extraordinary result was obtained as reflected by the column-headed 6-azauracil in Table III. When this pyrimidine analogue was added to a growing culture of PAO-1 wild type, growth ceased immediately as reflected by the growth curve in Fig. 5 (bottom line) but it appears from TAble III that the nucleoside triphosphates increased dramatically. The levels seen for UTP and CTP are unusually high. These findings will be detailed in the Discussion. Table IV gives the results of the nucleoside triphosphates found for the P. aeruginosa pyrb mutant. It can be noted from Table IV that when the pyrb mutant is starved for pyrimidines, the UTP and CTP levels decreased to near zero very rapidly. The possible interconversions of Ma, 'tmd4mm- -ll-, - -, ,
40 35 TABLE III. Nucleoside triphosphates in umole (gram dry weight) from P. aeruginosa PAO-1 wild type strain in the presence and absence of the listed additions. Supplements were added to succinate minimal (SM) medium at the rate of 5 ug/ml. 6-Azauracil was added to a growing culture at 9 Klett Units and sampled after one hour.
41 N Z ZQ 4 32 ZQ C) >4 u os os Co ' c)c% 4 C.- I 4 co co C 5 ' Pa wc/ 4 OP z W4 F ' o. En U) 4 ' M r 4 -t-4 I 4 - r- LO. LC). -4 WQ4 t z )i z M 41 A pq ++ o ' 9 c 'C ' LO LC' rn 4 E-4 po p - E- pe
42 37 Tabje IV. Nucleoside triphosphates in umol (gram dry weight) from P. aeruginosa pyrb mutant in the presence and absence of uracil, cytosine and cytidine. Supplements were added to succinate minimal (SM) medium at the rate of 5 ug/ml and starvation (no supplement added) for pyrimidines was for two hours.
43 >4 - - N 4 ro 2 w Fz: Fe -r-$ N r" N r U] IwoI G U E-4 Ln. co LO.- E m ) M ' 1 r) o ILA co. N 4 E9 (1)A 4 to z w z a l o ' ' D F e C',. r- CD N ' E- E- E- E-
44 39 the nucleobases and nucleosides are presented in the Discussion. In Table V another P. aeruginosa Pyr~ strain, namely a pyrf mutant (Fig. 7) was examined. Results seen in Table V are similar to those of Table IV except that it was difficult to elevate the UTP level by exogenously added pyrimidines in the pyrf strains. In order to compare the results from P. aeruginosa PAO-1 with another Pseudomonas strain, the nucleotides were also quantitated from wild type P., putida PPN. The results are shown in Table VI. Table VI shows that the nucleoside triphosphates of the wild type P. putida are lower in general than those of P. aeruginosa. In Table VII the nucleoside triphosphate levels are shown for a P. putida pyrb mutant. Results for this pyrb mutant of P. putida shows that uracil is the best (and only) exogenous pyrimidine that can increase the levels of UTP as reflected in the UTP level on addition of exogenous uracil (14.1 umol) (gram dry weight)~ 1, cytosine (4.4 umol) and cytidine (3.6 umol).
45 4 Tabje V. Nucleoside triphosphates in umol (gram dry weight)~ from P. aeruginosa pyrf mutant in the presence and absence of uracil, cytosine and cytidine. Supplements were added to succinate minimal (SM) medium at the rate of 5 ug/ml and starvation (no supplement added) for pyrimidines was two hours.
46 C ec h E-4 :z U4 >1 O- N E- 4-) Cu - o o k (N z z >4 > 4 E- -) C) O LO N 4~ CO ~ 1-11 E-4 m, > M Z z E-4 (1 W o o CNJ Co -) 9' r4 N C4 N c4 o q.oc y ei 43 EP E-q E-4 E- :D u 4
47 42 Table VI. Nucleoside triphosphates in umol (gram dry weight)~ from P. putida PPN wild type strain in the presence of the listed additions. Supplements were added to succinate minimal (SM) medium at the rate of 5 ug/ml. Growth was at 3C for _. putida strains.
48 J E-4 W Er >1 cc z E 4J >4 U) ~ E rzoz Q>4rzw E-4 P- z~ 41) >4 uc - U) u Co o o L u~c c4i..j. - > E ZE-4u :Z ) ok 1: 4) - C) ' f- 14 N c4 ci co o Z 4 m C)D N N + U <-4 <C/) - E-4 E 4 E
49 - 44 Tabje VII. Nucleoside triphosphates in umol (gram dry weight) from P. putida DxrB mutants in the presence of uracil, cytosine, or cytidine. Growth was at 3 C.
50 IM6M!MMMMilidiinNIRMIMEM E M araniassamment er.asm S SE- FQO >1 OU S E-4 E- E-4 u8 ' u: 4-) u % AD ) LO n LO L O NC COI W AE-1 q E-4 4> 4 to E-4 M PC e-p\ (N O ro rio E- 4 E-4 E-4 E-P : D u 4
51 CAPTER IV DISCUSSION Nucleotides serve two general functions in the cell. They play key roles in regulating metabolic reactions and they act as precursors for the synthesis of the nucleic acids. Metabolic pools of nucleotides and ribonucleic acids are constantly being synthesized and degraded in bacteria. The purine and pyrimidine nucleotides are continuously broken down to free bases, ribose and deoxyribose-l phosphates. These free bases, ribose and deoxyribose-lphosphates are required for salvage pathways to rebuild nucleotides (Fig. 1). Nucleotides can be synthesized do novo from glucose and it is known that every organism has a functional de novo pathway for purine and pyrimidine nucleotides. In the past 3 years, purine and pyrimidine metabolism in microorganisms has been studied extensively (Moat & Friedman, 196; O'Donovan & Neuhard, 197). Since intracellular purine and pyrimidine nucleotide pools control the individual biosynthetic pathways producing the substrates for RNA, accurate measurement of the endogenous nucleoside triphosphates is very important. PLC is both rapid and sensitive and provides the high degree of resolution necessary to detect and quantitate 46
52 47 compounds in complex biological samples. As a research tool, PLC has been successfully applied to such problems as detecting nucleotides (Krstulovic e al., 1979) in tissue extracts and determining alterations in blood nucleotides and metabolite levels in human disease states (artwick et al., 1979: Payne eta l., 1981). PLC has been modified successfully to measure ribo- and deoxyribonucleotides in bacteria (Dutta & O'Donovan, 1987). But very little effort (Chen et.a., 1977) has been made in developing extraction procedures for the detection and quantitation of low levels of nucleotide pools in microorganisms (Dutta, 1986). We have used PLC to answer the question regarding the concentration and stability of nucleotide pools in the Pseudomonas. It has been established for several members of the enteric bacteria, notably for E. coli and S. typhimurium, that nucleotide pools can be extracted, separated and accurately quantified. Moreover, it has been shown by Kelln et al., (1975) that exogenously added bases and nucleosides (refer to top right of Fig. 1) elevate the nucleoside triphosphates such that these triphosphates can act as feedback inhibitors and repressing metabolites in controlling the -(purine and) pyrimidine biosynthetic pathways. It has also been established by West (1986; 1987), which nucleic acid bases and nucleosides can satisfy the requirement of Pseudomonas Pyr~ mutants. owever to date there has been no systematic study on the nucleotide v m- MINwo
53 48 pool levels in the Pseudomonas. That is the purpose of the present research and its results are summarized below. When wild type P. aeruginosa was grown in succinate minimal medium its nucleoside triphosphate levels (Tables III-VII) measured in umoles (gram dry weight)~ 1 were as follows: UTP, 5.23 (3.42); CTP, 5.48 (2.21); ATP, 12.1 (1.3) and GTP, 4.53 (3.84). For comparison purposes the values from Salmonella typhimurium wild type strain grown in glucose minimal medium are given in parentheses (Kelln et.al., 1975). The following seven points are made with respect to the data shown in Tables III-VII. 1) The ribonucleoside mono-, di-, and triphosphates are present in P. aeruginosa at about the same level as found in different members of the enteric bacteria as compared above. (Only the triphosphates, and not the mono-, and diphosphates, are shown). 2) Addition of exogenous bases and nucleosides altered the appropriate triphosphates e.g. addition of uracil increased the UTP pool from 5.23 umole (gram dry weight)~ 1 to 1. umoles (gram dry weight)~ 1 (Table III Line 1.) Similar results were seen for the UTP pool in both the pyrb (Table IV, Line 1 plus uracil) and the pyrf (Table V, Line 1 plus uracil). 3) Why Pyr~ mutants of P. aerucinosa were starved for the pyrimidines, uracil or cytosine, the UTP and CTP pools
54 49 dropped to zero within two minutes. Refer to Table IV pyrb UTP, 11.3 umoles (gram dry weight)~i (+uracil);. umoles (-uracil), Table V pyrf 14.8 umoles (gram dry weight)~ 1 (+cytosine);. umoles (-cytosine). 4) Cytidine satisfies the pyrimidine requirement poorly as reflected by the minor change in UTP levels on the addition of exogenous cytidine. Pyr~ mutants of P. aeruginosa and P. putida do not grown on cytidine (J. Williams, unpublished results). 5) When a Pyr~ mutant of Pseudomonas was starved for pyrimidines, the ATP level increased dramatically. A similar result has been reported for other enteric bacteria (O'Donovan & Neuhard, 197). 6) All nucleoside triphosphates of wild type P. putida strain PPN are lower than the corresponding levels in P. aeruginosa PAO-1 strain. Compare Table III to Table VI. 7) When 6-azauracil, a pyrimidine analogue, known to inhibit OMP decarboxylase (andschumacher, 196) was added to growing culture of a P. aeruginosa, growth ceased immediately even though the UTP, CTP, and ATP pools increased dramatically. Since 6-azauracil is converted to 6-azaUMP by bacterial extracts (andschumacher, 196) and the 6-azaUMP is a competitive inhibitor of OMP decarboxylase it follows that the pyrimidine pathway is effectively shut off. In such cases (e.cg., starvation for pyrimidines) the ATP pool expands while the UTP and CTP pools decrease
55 5 simultaneously. One possible explanation for the high levels of UTP and CTP is that the 6-azaUMP is metabolized further and converted to 6-azaUTP and 6-azaCTP. These triphosphate analogues would register as ribotriphosphates and possibly give the result seen. There is one report (Brockman & Anderson, 1963) which suggested that Pseudomonas species can anabolize 6-azaUMP to the triphosphate level. In summary the question posed at the start of this thesis is now answered. It is possible to alter nucleoside triphosphate pools in the pseudomonads. Indeed, the ribonucleoside mono-, di-, and triphosphates are as sensitive to manipulation in P. aeruginosa and P. putida as they are in the enteric bacteria. ANON
56 BIBLIOGRAPY Abd-El-Al, A. D., Kessler, D.P., & Ingraham, J. L. (1969). Arginine-auxotrophic phenotype resulting from a mutation in pyra gene of Escherichia coli. Journal of Bacteriology 97, Anderson, P. M. & Meister, A. (1969). Control of Escherichia coli carbamyl phosphate synthetase by purine and pyrimidine nucleotides. Biochemistry 5, Assenza, S. P. & Brown, P. R. (1983). Reversed-phase retention of nucleic acid components. Separation and Purification Methods 12, Boulieu, R. & Bory, C. (1985). igh performance liquid chromatographic method for the analysis of purine and pyrimidine bases, ribonucleosides in biological fluids. Journal of Chromatography 399, Brown, P. K., & Miech, R. P. (1972). Comparison of cell extraction procedures for use with high pressure liquid chromatography, Analytical Chemistry 44, 172. Brown, P. R. (1973). igh Pressure Liquid Chromatography: Biomedical and Biochemical Applications. New York: Academic Press. Brockman, R. W. & Anderson, E. P. (1963). Pyrimidine analogues. p In Metabolic Inhibitors, pp Edited by R. M. ochster and J.. Quastels. New York: Academic Press. Chargeys, E. & Davidson, J. N. (1955). The Nucleic Acids, New York: Academic Press, Inc., Volume 1. Chen, S. C., Brown, P. R., & Rosie, D.. (1977). Extraction procedures for use prior to PLC nucleotide analysis using microparticle chemically bonded packings. Journal of Chromatographic Sciences 15, Cohen-Bazire, C., Sistrom, W. R. & Stanier, R. Y. (1957). Kinetic studies of pigment synthesis by nonsulfur purple bacteria. Journal of Cellular and Comparative Physiology 49,
57 52 Colowick, S. P. & Kaplan, N.. (eds), (1957). Methods in Enzymology, k, Academic Press, Inc., New York. Dutta, P. K. (1986). "Radial Compression igh Performance Liquid Chromatography as a Tool for the Measurement of Endogenous Nucleotides in Bacteria," Doctoral Dissertation, North Texas State University, Denton, Texas, Dutta, P. K. & O'Donovan, G. A. (1987). Separation and quantitation of bacterial ribonucleoside triphosphate extracted with trifluroacetic and by anion exchange PLC. Journal of Chromatography, 385, Gerhart, J. C. & Pardee, A. B. (1962). The enzymology of control by feedback inhibition. Journal of Biological Chemistry 237, Gerhart, J. C. & Pardee, A. B. (1964). Aspartate transcarbamylase, an enzyme designed for feedback inhibition. Federation Proceedings 23, Glaser, V. (1986). Chromatography: Primer and Current Practice, Bio-Technology 4, artwick, R. A., Assenza, S. P., & Brown, P. R. (1979). Identification and quantitation of nucleosides, bases and other UV-absorbing compounds in serum, using reversed-phase high-performance liquid chromatography. Journal of Chromatography 186, artwick, R. A. & Brown, P. R. (1975). The performance of microparticle chemically bonded anion exchange resins in analysis of nucleotides. Journal of Cromatography 112, 651, 662. offman, N. E. & Liao, J. C. (1977). Reverse phase high performance liquid chromatography separations of nucleotides in the presence of solvophobic ions, Analytical Chemistry 49, orvath, C., Melander, W., & Molnar, 1. (1977). Liquid chromatography of ionogenic substances with nonpolar stationary phase. Analytical Chemistry 49, urlbert, R. B. & P. Reichard. (1955). The conversion of orotic acid to uridine nucleotides in vitro. Acta Chemica Scandinavia 9,
58 53 Isaac, J.. & olloway, B. W. (1968). Control of pyrimidine biosynthesis in Pseudomonas aeruginosa. Journal of Bacteriology 96, Isaac, J.. & olloway, B. W. (1972). Control of arginine biosynthesis in Pseudomonas aeruginosa. Journal of General Microbiology 73, Karl, D. M. (1978). Occurrence and ecological significance of GTP in the ocean and in microbial cells. Applied and Environmental Microbiology. 36, Kelln, R. A., Kinahan, J. J., Foltermann, K. F., & O'Donovan, G. A. (1975). Pyrimidine biosynthetic enzymes of Salmonella typhimurium, repressed specifically by growth in the presence of cytidine. Journal of Bacteriology 124, Khym, J. X. (1975). An analytical system for the rapid separation of tissue nucleotide at low pressures on conventional anion exchangers. Clinical Chemistry 21, Krstulovic, A. M., artwick, R. A., & Brown, P. R. (1979). Reversed-phase liquid chromatographic separation of 3',5'-cyclic ribonucleotides. Clinical Chemistry 25, Leung,. B. & Schramm, V. L. (198). Adenylate degradation in E. coli. Journal of Biological Chemistry 255, Lieberman, I. (1955). Enzymatic amination of uridine triphosphate to cytidine triphosphate. Journal of the American Chemical Society 77, Lieberman, I., Kornberg, A. & Simms, E. S. (1955). Enzymatic synthesis of pyrimidine nucleotides. Orotidine-5' -phosphate and uridine-5' -phosphate. Journal of Biological Chemistry 215, Long, C. W. & Pardee, A. B. (1976). Cytidine triphosphate synthetase of Escherchia coli, B. I. Purification and kinetics. Journal of Biological Chemistry 242, McKeag, M. & Brown, P. R. (1978). Modification of high pressure liquid chromatographic nucleotide analysis. Journal of Chromatography 152,
59 54 Moat, A. G. & Friedman,. (196). The biosynthesis and interconversion of purines and their derivatives. Bacteriological Reviews 24, Neumann, J. & Jones, M. E. (1964). End product inhibition of asparate transcarbamylase in various species. Archives of Biochemistry and Biophysics 14, O'Donovan, G. A. & Neuhard, J. (197). Pyrimidine metabolism in microorganisms. Bacteriological Reviews 34, Payne, S. M. & Ames, B. N. (1982). A procedure for rapid extraction and high-presure liquid chromatographic separation of nucleotides and other small molecules from bacterial cells. Analytical Biochemistry 123, Pierard, A. (1966). Control of the activity of Escherichia coli carbamyl phosphate synthetase by antagonistic allesteric effectors. Science 154, Pogolotti, A. L. Jr. & Santi, D. L. (1982). igh pressure liquid chromatography-ultraviolet analysis of intracellular nucleotides. Analytical Biochemistry 126, Reiss, P. D., Zuurendonk, P. F., & Veech, R. L. (1984). Measurement of tissue purine, pyrimidine, and other nucleotides by radial compression high presure liquid chromatography. Analytical Biochemistry 14, Strehler, B. L. & McElroy, W. D. (1975). Assay of adenosine triphosphate. Methods of Enzymology 3,
Prerequisites Properties of allosteric enzymes. Basic mechanisms involving regulation of metabolic pathways.
 Case 16 Allosteric Regulation of ATCase Focus concept An enzyme involved in nucleotide synthesis is subject to regulation by a variety of combinations of nucleotides. Prerequisites Properties of allosteric
Case 16 Allosteric Regulation of ATCase Focus concept An enzyme involved in nucleotide synthesis is subject to regulation by a variety of combinations of nucleotides. Prerequisites Properties of allosteric
Analyze Nucleotides, Nucleosides, Purine, and Pyrimidine Bases Simultaneously with the Ultra IBD Column
 pharmaceutical #9 Applications note Analyze Nucleotides, Nucleosides, Purine, and Pyrimidine Bases Simultaneously with the Ultra IBD Column Mixtures of nucleotides, nucleosides, and their respective purine
pharmaceutical #9 Applications note Analyze Nucleotides, Nucleosides, Purine, and Pyrimidine Bases Simultaneously with the Ultra IBD Column Mixtures of nucleotides, nucleosides, and their respective purine
Chem Lecture 10 Lipid, Amino Acid, and Nucleotide Metabolism Part III: Nucleotide Metabolism
 Chem 352 - Lecture 10 Lipid, Amino Acid, and Nucleotide Metabolism Part III: Nucleotide Metabolism Lipid Metabolism Chem 352, Lecture 10, Part I: Lipid Metabolism 2 Lipid Metabolism Question: Draw a general
Chem 352 - Lecture 10 Lipid, Amino Acid, and Nucleotide Metabolism Part III: Nucleotide Metabolism Lipid Metabolism Chem 352, Lecture 10, Part I: Lipid Metabolism 2 Lipid Metabolism Question: Draw a general
ins 7 AND ENTERIC BACTERIA Presented to the Graduate Council of the University of North Texas in Partial Fulfillment of the Requirements
 ins 7 PYRIMIDINE NUCLEOSIDE METABOLISM IN PSEUDOMONADS AND ENTERIC BACTERIA THESIS Presented to the Graduate Council of the University of North Texas in Partial Fulfillment of the Requirements For the
ins 7 PYRIMIDINE NUCLEOSIDE METABOLISM IN PSEUDOMONADS AND ENTERIC BACTERIA THESIS Presented to the Graduate Council of the University of North Texas in Partial Fulfillment of the Requirements For the
The body has three primary lines of defense against changes in hydrogen ion concentration in the body fluids.
 ph and Nucleic acids Hydrogen Ion (H+) concentration is precisely regulated. The H+ concentration in the extracellular fluid is maintained at a very low level, averaging 0.00000004Eq/L. normal variations
ph and Nucleic acids Hydrogen Ion (H+) concentration is precisely regulated. The H+ concentration in the extracellular fluid is maintained at a very low level, averaging 0.00000004Eq/L. normal variations
Hillel K. Brandes and David S. Bell Supelco, Liquid Separations, Bellefonte PA T GIH
 Hillel K. Brandes and David S. Bell Supelco, Liquid Separations, Bellefonte PA 16823 T403163 GIH Abstract 2 ucleotides are ubiquitous in cells, with major functions including the control of cellular energetics,
Hillel K. Brandes and David S. Bell Supelco, Liquid Separations, Bellefonte PA 16823 T403163 GIH Abstract 2 ucleotides are ubiquitous in cells, with major functions including the control of cellular energetics,
Review of Lecture 1. Be able to identify the cell components for bacterial, animal, and plant cells and know their functions Properties of water
 Review of Lecture 1 Be able to identify the cell components for bacterial, animal, and plant cells and know their functions Properties of water Bulk properties Atomic properties Weak acids and bases Acid
Review of Lecture 1 Be able to identify the cell components for bacterial, animal, and plant cells and know their functions Properties of water Bulk properties Atomic properties Weak acids and bases Acid
Convenient Synthesis of Nucleoside 5 -Triphosphates for RNA Transcription. Supplemental Materials
 Supplementary Material (ESI) for Chemical Communications This journal is The Royal Society of Chemistry 2010 Convenient Synthesis of ucleoside 5 -Triphosphates for RA Transcription Julianne Caton-Williams,
Supplementary Material (ESI) for Chemical Communications This journal is The Royal Society of Chemistry 2010 Convenient Synthesis of ucleoside 5 -Triphosphates for RA Transcription Julianne Caton-Williams,
CHAPTER 8. An Introduction to Metabolism
 CHAPTER 8 An Introduction to Metabolism WHAT YOU NEED TO KNOW: Examples of endergonic and exergonic reactions. The key role of ATP in energy coupling. That enzymes work by lowering the energy of activation.
CHAPTER 8 An Introduction to Metabolism WHAT YOU NEED TO KNOW: Examples of endergonic and exergonic reactions. The key role of ATP in energy coupling. That enzymes work by lowering the energy of activation.
MET ABOLISM: Str uctur e and Metabolism of Nucleotide USMAN SUMO FRIEND T AMBUNAN ARLI ADIT YA PARIKESIT RIDO UT OMO
 MET ABLISM: Str uctur e and Metabolism of ucleotide USMA SUM FRIED T AMBUA ARLI ADIT YA PARIKESIT RID UT M ucleotide Function Building blocks for DA and RA Intracellular source of energy - Adenosine tr
MET ABLISM: Str uctur e and Metabolism of ucleotide USMA SUM FRIED T AMBUA ARLI ADIT YA PARIKESIT RID UT M ucleotide Function Building blocks for DA and RA Intracellular source of energy - Adenosine tr
 Diffusion of Some Nucleotides into Title irradiated and Gamma-Irradiated Eug on Physical, Chemical and Biologica Radiation, VII) Author(s) Matsuoka, Saburo Citation Bulletin of the Institute for Chemi
Diffusion of Some Nucleotides into Title irradiated and Gamma-Irradiated Eug on Physical, Chemical and Biologica Radiation, VII) Author(s) Matsuoka, Saburo Citation Bulletin of the Institute for Chemi
Transcarbamylase in Salmonella typhimurium
 JOURNAL oi BACTERIOLOGY, Apr. 1972, p. 66-70 Copyright i 1972 American Society for Microbiology Vol. 110, No. 1 Printed in U.S.A. Structural Genes for Ornithine Transcarbamylase in Salmonella typhimurium
JOURNAL oi BACTERIOLOGY, Apr. 1972, p. 66-70 Copyright i 1972 American Society for Microbiology Vol. 110, No. 1 Printed in U.S.A. Structural Genes for Ornithine Transcarbamylase in Salmonella typhimurium
Chapter 6- An Introduction to Metabolism*
 Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
CHAPTER 15 Metabolism: Basic Concepts and Design
 CHAPTER 15 Metabolism: Basic Concepts and Design Chapter 15 An overview of Metabolism Metabolism is the sum of cellular reactions - Metabolism the entire network of chemical reactions carried out by living
CHAPTER 15 Metabolism: Basic Concepts and Design Chapter 15 An overview of Metabolism Metabolism is the sum of cellular reactions - Metabolism the entire network of chemical reactions carried out by living
Ch. 2 BASIC CHEMISTRY. Copyright 2010 Pearson Education, Inc.
 Ch. 2 BASIC CHEMISTRY Matter and Composition of Matter Definition: Anything that has mass and occupies space Matter is made up of elements An element cannot be broken down by ordinary chemical means Atoms
Ch. 2 BASIC CHEMISTRY Matter and Composition of Matter Definition: Anything that has mass and occupies space Matter is made up of elements An element cannot be broken down by ordinary chemical means Atoms
Chapter 2: Chemical Basis of Life
 Chapter 2: Chemical Basis of Life Chemistry is the scientific study of the composition of matter and how composition changes. In order to understand human physiological processes, it is important to understand
Chapter 2: Chemical Basis of Life Chemistry is the scientific study of the composition of matter and how composition changes. In order to understand human physiological processes, it is important to understand
Outline. Metabolism: Energy and Enzymes. Forms of Energy. Chapter 6
 Metabolism: Energy and Enzymes Chapter 6 Forms of Energy Outline Laws of Thermodynamics Metabolic Reactions ATP Metabolic Pathways Energy of Activation Enzymes Photosynthesis Cellular Respiration 1 2 Forms
Metabolism: Energy and Enzymes Chapter 6 Forms of Energy Outline Laws of Thermodynamics Metabolic Reactions ATP Metabolic Pathways Energy of Activation Enzymes Photosynthesis Cellular Respiration 1 2 Forms
2) Matter composed of a single type of atom is known as a(n) 2) A) element. B) mineral. C) electron. D) compound. E) molecule.
 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Which of the following is a particle found in the nucleus of an atom and that has no electrical
MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Which of the following is a particle found in the nucleus of an atom and that has no electrical
Dr. Nafith Abu Tarboush
 8 Dr. Nafith Abu Tarboush June 30 th 2013 Ahmad Ayyat Nucleic Acids: Molecules that carries information for growth and production of cells, and they are Polymers of "Nucleotides" (the monomers).01 Nucleotide
8 Dr. Nafith Abu Tarboush June 30 th 2013 Ahmad Ayyat Nucleic Acids: Molecules that carries information for growth and production of cells, and they are Polymers of "Nucleotides" (the monomers).01 Nucleotide
2/25/2013. Electronic Configurations
 1 2 3 4 5 Chapter 2 Chemical Principles The Structure of Atoms Chemistry is the study of interactions between atoms and molecules The atom is the smallest unit of matter that enters into chemical reactions
1 2 3 4 5 Chapter 2 Chemical Principles The Structure of Atoms Chemistry is the study of interactions between atoms and molecules The atom is the smallest unit of matter that enters into chemical reactions
HPLC. GRATE Chromatography Lab Course. Dr. Johannes Ranke. September 2003
 HPLC GRATE Chromatography Lab Course Dr. Johannes Ranke Organisation The groups Start at 9:00 am End at 18:00 pm at the latest Friday, 19th we will finish at 2:00 pm Thursday, 11th: Lecture at 08:15 am
HPLC GRATE Chromatography Lab Course Dr. Johannes Ranke Organisation The groups Start at 9:00 am End at 18:00 pm at the latest Friday, 19th we will finish at 2:00 pm Thursday, 11th: Lecture at 08:15 am
Enzymatic Assay of HYPOXANTHINE-GUANINE PHOSPHORIBOSYL TRANSFERASE (EC )
 PRINCIPLE: PRPP + Guanine HGPRT > GMP + Pyrophosphate Mg ++ GMP + ATP GK > GDP + ADP ADP + PEP PK > ATP + Pyruvate GDP + PEP PK > GTP + Pyruvate 2 Pyruvate + 2 ß-NADH LDH > 2 Lactate + 2 ß-NAD Abbreviations
PRINCIPLE: PRPP + Guanine HGPRT > GMP + Pyrophosphate Mg ++ GMP + ATP GK > GDP + ADP ADP + PEP PK > ATP + Pyruvate GDP + PEP PK > GTP + Pyruvate 2 Pyruvate + 2 ß-NADH LDH > 2 Lactate + 2 ß-NAD Abbreviations
Microbiology with Diseases by Taxonomy, 5e (Bauman) Chapter 2 The Chemistry of Microbiology. 2.1 Multiple Choice Questions
 Microbiology with Diseases by Taxonomy, 5e (Bauman) Chapter 2 The Chemistry of Microbiology 2.1 Multiple Choice Questions 1) Which of the following does not contribute significantly to the mass of an atom?
Microbiology with Diseases by Taxonomy, 5e (Bauman) Chapter 2 The Chemistry of Microbiology 2.1 Multiple Choice Questions 1) Which of the following does not contribute significantly to the mass of an atom?
A pentose bisphosphate pathway for nucleoside degradation in Archaea. Engineering, Kyoto University, Katsura, Nishikyo-ku, Kyoto , Japan.
 SUPPLEMENTARY INFORMATION A pentose bisphosphate pathway for nucleoside degradation in Archaea Riku Aono 1,, Takaaki Sato 1,, Tadayuki Imanaka, and Haruyuki Atomi 1, * 7 8 9 10 11 1 Department of Synthetic
SUPPLEMENTARY INFORMATION A pentose bisphosphate pathway for nucleoside degradation in Archaea Riku Aono 1,, Takaaki Sato 1,, Tadayuki Imanaka, and Haruyuki Atomi 1, * 7 8 9 10 11 1 Department of Synthetic
The biomolecules of terrestrial life
 Functional groups in biomolecules Groups of atoms that are responsible for the chemical properties of biomolecules The biomolecules of terrestrial life Planets and Astrobiology (2017-2018) G. Vladilo 1
Functional groups in biomolecules Groups of atoms that are responsible for the chemical properties of biomolecules The biomolecules of terrestrial life Planets and Astrobiology (2017-2018) G. Vladilo 1
BIOLOGY 10/11/2014. An Introduction to Metabolism. Outline. Overview: The Energy of Life
 8 An Introduction to Metabolism CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson Outline I. Forms of Energy II. Laws of Thermodynamics III. Energy and metabolism IV. ATP V. Enzymes
8 An Introduction to Metabolism CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson Outline I. Forms of Energy II. Laws of Thermodynamics III. Energy and metabolism IV. ATP V. Enzymes
Metabolism and Enzymes
 Energy Basics Metabolism and Enzymes Chapter 5 Pgs. 77 86 Chapter 8 Pgs. 142 162 Energy is the capacity to cause change, and is required to do work. Very difficult to define quantity. Two types of energy:
Energy Basics Metabolism and Enzymes Chapter 5 Pgs. 77 86 Chapter 8 Pgs. 142 162 Energy is the capacity to cause change, and is required to do work. Very difficult to define quantity. Two types of energy:
Supporting Information. Chemo-enzymatic Synthesis of Isotopically Labeled Nicotinamide Ribose
 Electronic Supplementary Material (ESI) for Organic & Biomolecular Chemistry. This journal is The Royal Society of Chemistry 2018 Supporting Information Chemo-enzymatic Synthesis of Isotopically Labeled
Electronic Supplementary Material (ESI) for Organic & Biomolecular Chemistry. This journal is The Royal Society of Chemistry 2018 Supporting Information Chemo-enzymatic Synthesis of Isotopically Labeled
Chemistry 5.07SC Biological Chemistry I Fall Semester, 2013
 Chemistry 5.07SC Biological Chemistry I Fall Semester, 2013 Lecture 10. Biochemical Transformations II. Phosphoryl transfer and the kinetics and thermodynamics of energy currency in the cell: ATP and GTP.
Chemistry 5.07SC Biological Chemistry I Fall Semester, 2013 Lecture 10. Biochemical Transformations II. Phosphoryl transfer and the kinetics and thermodynamics of energy currency in the cell: ATP and GTP.
Chapter Two: The Chemistry of Biology. The molecules of life make up the structure of cells Chemistry of biological molecule
 Chapter Two: The Chemistry of Biology The molecules of life make up the structure of cells Chemistry of biological molecule Atoms and Elements: Atoms: The basic units of all matter, containing three major
Chapter Two: The Chemistry of Biology The molecules of life make up the structure of cells Chemistry of biological molecule Atoms and Elements: Atoms: The basic units of all matter, containing three major
Full file at
 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Which of the following is an uncharged particle found in the nucleus of 1) an atom and which has
MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Which of the following is an uncharged particle found in the nucleus of 1) an atom and which has
Flow of Energy. Flow of Energy. Energy and Metabolism. Chapter 6
 Energy and Metabolism Chapter 6 Flow of Energy Energy: the capacity to do work -kinetic energy: the energy of motion -potential energy: stored energy Energy can take many forms: mechanical electric current
Energy and Metabolism Chapter 6 Flow of Energy Energy: the capacity to do work -kinetic energy: the energy of motion -potential energy: stored energy Energy can take many forms: mechanical electric current
Metabolism: Energy and Enzymes. February 24 th, 2012
 Metabolism: Energy and Enzymes February 24 th, 2012 1 Outline Forms of Energy Laws of Thermodynamics Metabolic Reactions ATP Metabolic Pathways Energy of Activation Enzymes Photosynthesis Cellular Respiration
Metabolism: Energy and Enzymes February 24 th, 2012 1 Outline Forms of Energy Laws of Thermodynamics Metabolic Reactions ATP Metabolic Pathways Energy of Activation Enzymes Photosynthesis Cellular Respiration
Energy Transformation, Cellular Energy & Enzymes (Outline)
 Energy Transformation, Cellular Energy & Enzymes (Outline) Energy conversions and recycling of matter in the ecosystem. Forms of energy: potential and kinetic energy The two laws of thermodynamic and definitions
Energy Transformation, Cellular Energy & Enzymes (Outline) Energy conversions and recycling of matter in the ecosystem. Forms of energy: potential and kinetic energy The two laws of thermodynamic and definitions
2: CHEMICAL COMPOSITION OF THE BODY
 1 2: CHEMICAL COMPOSITION OF THE BODY Although most students of human physiology have had at least some chemistry, this chapter serves very well as a review and as a glossary of chemical terms. In particular,
1 2: CHEMICAL COMPOSITION OF THE BODY Although most students of human physiology have had at least some chemistry, this chapter serves very well as a review and as a glossary of chemical terms. In particular,
Chapter 2! Chapter 2 Chemistry. The Chemical Level of Organization! SECTION 2-1! Atoms are the basic particles of matter! Subatomic Particles!
 Chapter 2 The Chemical Level of Organization SECTION 2-1 Atoms are the basic particles of matter Note: Although we will not cover the first parts of these notes during lecture, you are responsible for
Chapter 2 The Chemical Level of Organization SECTION 2-1 Atoms are the basic particles of matter Note: Although we will not cover the first parts of these notes during lecture, you are responsible for
Chapter 002 The Chemistry of Biology
 Chapter 002 The Chemistry of Biology Multiple Choice Questions 1. Anything that occupies space and has mass is called A. Atomic B. Living C. Matter D. Energy E. Space 2. The electrons of an atom are A.
Chapter 002 The Chemistry of Biology Multiple Choice Questions 1. Anything that occupies space and has mass is called A. Atomic B. Living C. Matter D. Energy E. Space 2. The electrons of an atom are A.
This is an example of cellular respiration, which can be used to make beer and wine using different metabolic pathways For these reasons we call this
 Chapter 6 Carvings from ancient Egypt show barley being crushed and mixed with water (left) and then put into closed vessels (centre) where airless conditions are suitable for the production of alcohol
Chapter 6 Carvings from ancient Egypt show barley being crushed and mixed with water (left) and then put into closed vessels (centre) where airless conditions are suitable for the production of alcohol
Chapter 2: The Chemical Level Of Organization
 Chapter 2: The Chemical Level Of Organization With this chapter we begin our march through the body s levels of organization (recall the top of 10 th Martini Figure 1-1). Here in Chapter 2 we will discuss
Chapter 2: The Chemical Level Of Organization With this chapter we begin our march through the body s levels of organization (recall the top of 10 th Martini Figure 1-1). Here in Chapter 2 we will discuss
NAD metabolome analysis in cultured human cells using 1H NMR spectroscopy
 NAD metabolome analysis in cultured human cells using 1H NMR spectroscopy Konstantin Shabalin 1,2, *, Kirill Nerinovski 3,4, Alexandr Yakimov 2,3, Veronika Kulikova 1,3, Mikhail Khodorkovskiy 3, Mathias
NAD metabolome analysis in cultured human cells using 1H NMR spectroscopy Konstantin Shabalin 1,2, *, Kirill Nerinovski 3,4, Alexandr Yakimov 2,3, Veronika Kulikova 1,3, Mikhail Khodorkovskiy 3, Mathias
Chapter 2. Chemical Principles
 Chapter 2 Chemical Principles Insert Fig CO 2 The Structure of Atoms Chemistry is the study of interactions between atoms and molecules The atom is the smallest unit of matter that enters into chemical
Chapter 2 Chemical Principles Insert Fig CO 2 The Structure of Atoms Chemistry is the study of interactions between atoms and molecules The atom is the smallest unit of matter that enters into chemical
Bio10 Cell and Molecular Lecture Notes SRJC
 Basic Chemistry Atoms Smallest particles that retain properties of an element Made up of subatomic particles: Protons (+) Electrons (-) Neutrons (no charge) Isotopes Atoms of an element with different
Basic Chemistry Atoms Smallest particles that retain properties of an element Made up of subatomic particles: Protons (+) Electrons (-) Neutrons (no charge) Isotopes Atoms of an element with different
NUCLEOSIDES AND NUCLEOTIDES OF THE COLD ACID-SOLUBLE PORTION OF THE BLOOD OF STEERS 1
 NUCLEOSIDES AND NUCLEOTIDES OF THE COLD ACID-SOLUBLE PORTION OF THE BLOOD OF STEERS 1 Robert C. Smith and Connie M. Stricker Auburn University 2, Auburn, Alabama 36830 SUMMARY The ultraviolet absorption
NUCLEOSIDES AND NUCLEOTIDES OF THE COLD ACID-SOLUBLE PORTION OF THE BLOOD OF STEERS 1 Robert C. Smith and Connie M. Stricker Auburn University 2, Auburn, Alabama 36830 SUMMARY The ultraviolet absorption
Chapter content. Reference
 Chapter 7 HPLC Instrumental Analysis Rezaul Karim Environmental Science and Technology Jessore University of Science and Technology Chapter content Liquid Chromatography (LC); Scope; Principles Instrumentation;
Chapter 7 HPLC Instrumental Analysis Rezaul Karim Environmental Science and Technology Jessore University of Science and Technology Chapter content Liquid Chromatography (LC); Scope; Principles Instrumentation;
Regulation of Pyrimidine Biosynthetic Gene Expression in Bacteria: Repression without Repressors
 MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, June 2008, p. 266 300 Vol. 72, No. 2 1092-2172/08/$08.00 0 doi:10.1128/mmbr.00001-08 Copyright 2008, American Society for Microbiology. All Rights Reserved.
MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, June 2008, p. 266 300 Vol. 72, No. 2 1092-2172/08/$08.00 0 doi:10.1128/mmbr.00001-08 Copyright 2008, American Society for Microbiology. All Rights Reserved.
The Chemical Level of Organization
 PowerPoint Lecture Slides prepared by Meg Flemming Austin Community College C H A P T E R 2 The Chemical Level of Organization Chapter 2 Learning Outcomes 2-1 2-2 2-3 2-4 Describe an atom and how atomic
PowerPoint Lecture Slides prepared by Meg Flemming Austin Community College C H A P T E R 2 The Chemical Level of Organization Chapter 2 Learning Outcomes 2-1 2-2 2-3 2-4 Describe an atom and how atomic
Basic Chemistry. Chapter 2 BIOL1000 Dr. Mohamad H. Termos
 Basic Chemistry Chapter 2 BIOL1000 Dr. Mohamad H. Termos Chapter 2 Objectives Following this chapter, you should be able to describe: - Atoms, molecules, and ions - Composition and properties - Types of
Basic Chemistry Chapter 2 BIOL1000 Dr. Mohamad H. Termos Chapter 2 Objectives Following this chapter, you should be able to describe: - Atoms, molecules, and ions - Composition and properties - Types of
9/25/2011. Outline. Overview: The Energy of Life. I. Forms of Energy II. Laws of Thermodynamics III. Energy and metabolism IV. ATP V.
 Chapter 8 Introduction to Metabolism Outline I. Forms of Energy II. Laws of Thermodynamics III. Energy and metabolism IV. ATP V. Enzymes Overview: The Energy of Life Figure 8.1 The living cell is a miniature
Chapter 8 Introduction to Metabolism Outline I. Forms of Energy II. Laws of Thermodynamics III. Energy and metabolism IV. ATP V. Enzymes Overview: The Energy of Life Figure 8.1 The living cell is a miniature
EFFECT OF ph AND AMMONIUM IONS ON THE PERMEABILITY
 EFFECT OF ph AND AMMONIUM IONS ON THE PERMEABILITY OF BACILLUS PASTEURII W. R. WILEY AND J. L. STOKES Department of Bacteriology and Public Health, Washington State University, Pullman, Washington ABSTRACT
EFFECT OF ph AND AMMONIUM IONS ON THE PERMEABILITY OF BACILLUS PASTEURII W. R. WILEY AND J. L. STOKES Department of Bacteriology and Public Health, Washington State University, Pullman, Washington ABSTRACT
Uptake of Adenosine 5'-Monophosphate by Escherichia coli
 JOURNAL OF BACTERIOLoGY, Feb. 1975, p. 41-45 Copyright 1975 American Society for Microbiology Vol. 11, No. Printed in U.S.A. Uptake of Adenosine 5'-Monophosphate by Escherichia coli EZRA YAGIL* AND IFOR
JOURNAL OF BACTERIOLoGY, Feb. 1975, p. 41-45 Copyright 1975 American Society for Microbiology Vol. 11, No. Printed in U.S.A. Uptake of Adenosine 5'-Monophosphate by Escherichia coli EZRA YAGIL* AND IFOR
Chemical Principles and Biomolecules (Chapter 2) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Eastern Campus
 Chemical Principles and Biomolecules (Chapter 2) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Eastern Campus Primary Source for figures and content: Tortora, G.J. Microbiology
Chemical Principles and Biomolecules (Chapter 2) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Eastern Campus Primary Source for figures and content: Tortora, G.J. Microbiology
Chapter 2 The Chemistry of Biology. Dr. Ramos BIO 370
 Chapter 2 The Chemistry of Biology Dr. Ramos BIO 370 2 Atoms, Bonds, and Molecules Matter - all materials that occupy space and have mass Matter is composed of atoms. Atom simplest form of matter not divisible
Chapter 2 The Chemistry of Biology Dr. Ramos BIO 370 2 Atoms, Bonds, and Molecules Matter - all materials that occupy space and have mass Matter is composed of atoms. Atom simplest form of matter not divisible
Nucleotide metabolism
 Nucleotide metabolism Roles of nuclotides energy storage (central role of ) nucleotide units for RNA, DNA synthesis components of cofactors (NAD, FAD) intracellular second messengers (camp, cgmp) synthesis
Nucleotide metabolism Roles of nuclotides energy storage (central role of ) nucleotide units for RNA, DNA synthesis components of cofactors (NAD, FAD) intracellular second messengers (camp, cgmp) synthesis
Chemical Principles. PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R
 PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R 2 Chemical Principles The Structure of Atoms Learning Objective 2-1 Describe the structure of
PowerPoint Lecture Presentations prepared by Bradley W. Christian, McLennan Community College C H A P T E R 2 Chemical Principles The Structure of Atoms Learning Objective 2-1 Describe the structure of
CHROMATOGRAPHY. The term "chromatography" is derived from the original use of this method for separating yellow and green plant pigments.
 CHROMATOGRAPHY The term "chromatography" is derived from the original use of this method for separating yellow and green plant pigments. THEORY OF CHROMATOGRAPHY: Separation of two sample components in
CHROMATOGRAPHY The term "chromatography" is derived from the original use of this method for separating yellow and green plant pigments. THEORY OF CHROMATOGRAPHY: Separation of two sample components in
An Introduction to Metabolism
 An Introduction to Metabolism I. All of an organism=s chemical reactions taken together is called metabolism. A. Metabolic pathways begin with a specific molecule, which is then altered in a series of
An Introduction to Metabolism I. All of an organism=s chemical reactions taken together is called metabolism. A. Metabolic pathways begin with a specific molecule, which is then altered in a series of
Production of Recombinant Annexin V from plasmid pet12a-papi
 Tait Research Laboratory Page 1 of 5 Principle Production of Recombinant Annexin V from plasmid pet12a-papi Annexin V is expressed cytoplasmically in BL21(DE3) E. coli (Novagen) with the pet vector system
Tait Research Laboratory Page 1 of 5 Principle Production of Recombinant Annexin V from plasmid pet12a-papi Annexin V is expressed cytoplasmically in BL21(DE3) E. coli (Novagen) with the pet vector system
UNIT 2 CHEMISTRY. Atomic Structure: Ionic Bond: Covalent Bond: Hydrogen Bond:
 UNIT 2 CHEMISTRY Atomic Structure: Ionic Bond: Hydrogen Bond: Covalent Bond: 1 Carbohydrates: >energy yield- >elements- >monomers- >functions- >examples- >misc- Lipids: Proteins: Nucleic Acids: I. Energy
UNIT 2 CHEMISTRY Atomic Structure: Ionic Bond: Hydrogen Bond: Covalent Bond: 1 Carbohydrates: >energy yield- >elements- >monomers- >functions- >examples- >misc- Lipids: Proteins: Nucleic Acids: I. Energy
Introduction to Chromatographic Separations
 Introduction to Chromatographic Separations Analysis of complex samples usually involves previous separation prior to compound determination. Two main separation methods instrumentation are available:
Introduction to Chromatographic Separations Analysis of complex samples usually involves previous separation prior to compound determination. Two main separation methods instrumentation are available:
Full file at Essentials of Anatomy & Physiology (Martini/ Bartholomew) Chapter 2 The Chemical Level of Organization
 Essentials of Anatomy & Physiology (Martini/ Bartholomew) Chapter 2 The Chemical Level of Organization Multiple Choice 1) An unstable isotope that emits subatomic particles spontaneously is called A) a
Essentials of Anatomy & Physiology (Martini/ Bartholomew) Chapter 2 The Chemical Level of Organization Multiple Choice 1) An unstable isotope that emits subatomic particles spontaneously is called A) a
W2. Chemical structures of protein and DNA
 W2. Chemical structures of protein and DNA Copyright Kang, Lin-Woo, Ph.D. Professor Department of Biological Sciences Konkuk University Seoul, Korea Lectures prepared by Christine L. Case The Structure
W2. Chemical structures of protein and DNA Copyright Kang, Lin-Woo, Ph.D. Professor Department of Biological Sciences Konkuk University Seoul, Korea Lectures prepared by Christine L. Case The Structure
An Introduction to Metabolism
 Chapter 8 1 An Introduction to Metabolism PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 8 1 An Introduction to Metabolism PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
 Radiation Damage of Purines and Pyr Titleon Physical, Chemical and Biologica Radiation, IX) Author(s) Matsuoka, Saburo Citation Bulletin of the Institute for Chemi University (1968), 46(1): 1-6 Issue Date
Radiation Damage of Purines and Pyr Titleon Physical, Chemical and Biologica Radiation, IX) Author(s) Matsuoka, Saburo Citation Bulletin of the Institute for Chemi University (1968), 46(1): 1-6 Issue Date
- BIOENERGETICS - DR. A. TARAB DEPT. OF BIOCHEMISTRY HKMU
 - BIOENERGETICS - DR. A. TARAB DEPT. OF BIOCHEMISTRY HKMU Bioenergetics the field of biochemistry concerned with the transfer and use of energy by biological system BIOLOGICAL IMPORTANCE: Suitable fuel
- BIOENERGETICS - DR. A. TARAB DEPT. OF BIOCHEMISTRY HKMU Bioenergetics the field of biochemistry concerned with the transfer and use of energy by biological system BIOLOGICAL IMPORTANCE: Suitable fuel
cgmp ELISA Kit (Direct Competitive) Based on Monoclonal Anti-cGMP Antibody
 (FOR RESEARCH USE ONLY. DO NOT USE IT IN CLINICAL DIAGNOSIS!) cgmp ELISA Kit (Direct Competitive) Based on Monoclonal Anti-cGMP Antibody Catalog No: E-EL-DS02 96T This manual must be read attentively and
(FOR RESEARCH USE ONLY. DO NOT USE IT IN CLINICAL DIAGNOSIS!) cgmp ELISA Kit (Direct Competitive) Based on Monoclonal Anti-cGMP Antibody Catalog No: E-EL-DS02 96T This manual must be read attentively and
Chapter 6. Ground Rules Of Metabolism
 Chapter 6 Ground Rules Of Metabolism Alcohol Dehydrogenase An enzyme Breaks down ethanol and other toxic alcohols Allows humans to drink Metabolism Is the totality of an organism s chemical reactions Arises
Chapter 6 Ground Rules Of Metabolism Alcohol Dehydrogenase An enzyme Breaks down ethanol and other toxic alcohols Allows humans to drink Metabolism Is the totality of an organism s chemical reactions Arises
NAD + /NADH Assay [Colorimetric]
![NAD + /NADH Assay [Colorimetric] NAD + /NADH Assay [Colorimetric]](/thumbs/91/105496846.jpg) G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name NAD + /NADH Assay [Colorimetric] (Cat. #786 1539, 786 1540) think proteins! think G-Biosciences
G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name NAD + /NADH Assay [Colorimetric] (Cat. #786 1539, 786 1540) think proteins! think G-Biosciences
Microbiology: A Systems Approach, 2 nd ed. Chapter 2: The Chemistry of Biology
 Microbiology: A Systems Approach, 2 nd ed. Chapter 2: The Chemistry of Biology 2.1 Atoms, Bonds, and Molecules: Fundamental Building Blocks Matter: anything that occupies space and has mass Can be liquid,
Microbiology: A Systems Approach, 2 nd ed. Chapter 2: The Chemistry of Biology 2.1 Atoms, Bonds, and Molecules: Fundamental Building Blocks Matter: anything that occupies space and has mass Can be liquid,
2017 Ebneshahidi. Dr. Ali Ebneshahidi
 Dr. Ali Ebneshahidi A. Introduction Chemistry science that deals with the composition of substances and the changes that take place in their composition. Organic chemistry chemistry that deals with organic
Dr. Ali Ebneshahidi A. Introduction Chemistry science that deals with the composition of substances and the changes that take place in their composition. Organic chemistry chemistry that deals with organic
Enzymatic Assay of CASEIN KINASE II
 PRINCIPLE: Casein +?- 32 P-ATP Casein Kinase II > [ 32 P]-Phosphorylated Casein + ADP CONDITIONS: T = 37 C, ph 7.5 METHOD: Radiolabelled Stop Reaction REAGENTS: A. 100 mm HEPES Buffer, ph 7.5 at 37 C (Enzyme
PRINCIPLE: Casein +?- 32 P-ATP Casein Kinase II > [ 32 P]-Phosphorylated Casein + ADP CONDITIONS: T = 37 C, ph 7.5 METHOD: Radiolabelled Stop Reaction REAGENTS: A. 100 mm HEPES Buffer, ph 7.5 at 37 C (Enzyme
3.2 ATP: Energy Currency of the Cell 141
 : Energy urrency of the ell Thousands of reactions take place in living cells. Many reactions require the addition of for the assembly of complex molecules from simple reactants. These reactions include
: Energy urrency of the ell Thousands of reactions take place in living cells. Many reactions require the addition of for the assembly of complex molecules from simple reactants. These reactions include
Chapter 2: The Chemical Basis of Life
 Chapter 2: The Chemical Basis of Life I. Basic Chemistry A. Matter, Mass, and Weight 1. All living and nonliving things are composed of 2. represents the amount of matter. 3. is caused by the gravitational
Chapter 2: The Chemical Basis of Life I. Basic Chemistry A. Matter, Mass, and Weight 1. All living and nonliving things are composed of 2. represents the amount of matter. 3. is caused by the gravitational
Biomolecules. Energetics in biology. Biomolecules inside the cell
 Biomolecules Energetics in biology Biomolecules inside the cell Energetics in biology The production of energy, its storage, and its use are central to the economy of the cell. Energy may be defined as
Biomolecules Energetics in biology Biomolecules inside the cell Energetics in biology The production of energy, its storage, and its use are central to the economy of the cell. Energy may be defined as
Characterization of Pyrimidine-Repressible and Arginine- Repressible Carbamyl Phosphate Synthetases from Bacillus
 JOURNAL OF BACTERIOLOGY, Jan. 1979, p. 82-91 0021-9193/79/01-0082/10$02.00/0 Vol. 137, No. 1 Characterization of Pyrimidine-Repressible and Arginine- Repressible Carbamyl Phosphate Synthetases from Bacillus
JOURNAL OF BACTERIOLOGY, Jan. 1979, p. 82-91 0021-9193/79/01-0082/10$02.00/0 Vol. 137, No. 1 Characterization of Pyrimidine-Repressible and Arginine- Repressible Carbamyl Phosphate Synthetases from Bacillus
SECTION J, BIOCHEMISTRY Growth Stimulation of Escherichia coli by 2-Thiouracil
 289 SECTION J, BIOCHEMISTRY Growth Stimulation of Escherichia coli by 2-Thiouracil PAUL T. CARDEILHAC and ERNEST M. HODNETT, Oklahoma State University, Stillwater 2-Thiouracil has been extensively tested
289 SECTION J, BIOCHEMISTRY Growth Stimulation of Escherichia coli by 2-Thiouracil PAUL T. CARDEILHAC and ERNEST M. HODNETT, Oklahoma State University, Stillwater 2-Thiouracil has been extensively tested
AN INTRODUCTION TO METABOLISM. Metabolism, Energy, and Life
 AN INTRODUCTION TO METABOLISM Metabolism, Energy, and Life 1. The chemistry of life is organized into metabolic pathways 2. Organisms transform energy 3. The energy transformations of life are subject
AN INTRODUCTION TO METABOLISM Metabolism, Energy, and Life 1. The chemistry of life is organized into metabolic pathways 2. Organisms transform energy 3. The energy transformations of life are subject
Hole s Human Anatomy and Physiology Eleventh Edition. Chapter 2
 Hole s Human Anatomy and Physiology Eleventh Edition Shier Butler Lewis Chapter 2 1 Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. CHAPTER 2 CHEMICAL BASIS OF
Hole s Human Anatomy and Physiology Eleventh Edition Shier Butler Lewis Chapter 2 1 Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. CHAPTER 2 CHEMICAL BASIS OF
Advanced Cell Biology. Lecture 6
 Advanced Cell Biology. Lecture 6 Alexey Shipunov Minot State University January 23, 2013 Shipunov (MSU) Advanced Cell Biology. Lecture 6 January 23, 2013 1 / 48 Outline Questions and answers Nucleic acids
Advanced Cell Biology. Lecture 6 Alexey Shipunov Minot State University January 23, 2013 Shipunov (MSU) Advanced Cell Biology. Lecture 6 January 23, 2013 1 / 48 Outline Questions and answers Nucleic acids
UNIT 2 CHEMISTRY. Atomic Structure: Ionic Bond: Covalent Bond: Hydrogen Bond:
 UNIT 2 CHEMISTRY Atomic Structure: Ionic Bond: Hydrogen Bond: Covalent Bond: 1 Carbohydrates: >energy yield- >elements- >monomers- >functions- >examples- >misc- Lipids: Proteins: Nucleic Acids: I. Energy
UNIT 2 CHEMISTRY Atomic Structure: Ionic Bond: Hydrogen Bond: Covalent Bond: 1 Carbohydrates: >energy yield- >elements- >monomers- >functions- >examples- >misc- Lipids: Proteins: Nucleic Acids: I. Energy
An Introduction to Metabolism
 Chapter 8 An Introduction to Metabolism Dr. Wendy Sera Houston Community College Biology 1406 Key Concepts in Chapter 8 1. An organism s metabolism transforms matter and energy, subject to the laws of
Chapter 8 An Introduction to Metabolism Dr. Wendy Sera Houston Community College Biology 1406 Key Concepts in Chapter 8 1. An organism s metabolism transforms matter and energy, subject to the laws of
Biology. Chapter 2 Notes
 Biology Chapter 2 Notes Section 1: Nature of Matter Objectives: 1) Differentiate between atoms and elements 2) Analyze how compounds are formed 3) Distinguish between covalent bonds, hydrogen bonds and
Biology Chapter 2 Notes Section 1: Nature of Matter Objectives: 1) Differentiate between atoms and elements 2) Analyze how compounds are formed 3) Distinguish between covalent bonds, hydrogen bonds and
CHAPTER CHROMATOGRAPHIC METHODS OF SEPARATIONS
 Islamic University in Madinah Department of Chemistry CHAPTER - ----- CHROMATOGRAPHIC METHODS OF SEPARATIONS Prepared By Dr. Khalid Ahmad Shadid Chemistry Department Islamic University in Madinah TRADITIONAL
Islamic University in Madinah Department of Chemistry CHAPTER - ----- CHROMATOGRAPHIC METHODS OF SEPARATIONS Prepared By Dr. Khalid Ahmad Shadid Chemistry Department Islamic University in Madinah TRADITIONAL
An Introduction to Metabolism
 An Introduction to Metabolism The living cell is a microscopic factory where life s giant processes can be performed: -sugars to amino acids to proteins and vise versa -reactions to dismantle polymers
An Introduction to Metabolism The living cell is a microscopic factory where life s giant processes can be performed: -sugars to amino acids to proteins and vise versa -reactions to dismantle polymers
Chapter 8: An Introduction to Metabolism. 1. Energy & Chemical Reactions 2. ATP 3. Enzymes & Metabolic Pathways
 Chapter 8: An Introduction to Metabolism 1. Energy & Chemical Reactions 2. ATP 3. Enzymes & Metabolic Pathways 1. Energy & Chemical Reactions 2 Basic Forms of Energy Kinetic Energy (KE) energy in motion
Chapter 8: An Introduction to Metabolism 1. Energy & Chemical Reactions 2. ATP 3. Enzymes & Metabolic Pathways 1. Energy & Chemical Reactions 2 Basic Forms of Energy Kinetic Energy (KE) energy in motion
Cell and Molecular Biology
 Cell and Molecular Biology (3000719): academic year 2013 Content & Objective :Cell Chemistry and Biosynthesis 3 rd Edition, 1994, pp. 41-88. 4 th Edition, 2002, pp. 47-127. 5 th Edition, 2008, pp. 45-124.
Cell and Molecular Biology (3000719): academic year 2013 Content & Objective :Cell Chemistry and Biosynthesis 3 rd Edition, 1994, pp. 41-88. 4 th Edition, 2002, pp. 47-127. 5 th Edition, 2008, pp. 45-124.
An Introduction to Metabolism
 CAMPBELL BIOLOGY IN FOCUS Urry Cain Wasserman Minorsky Jackson Reece 6 An Introduction to Metabolism Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge Overview: The Energy of Life The
CAMPBELL BIOLOGY IN FOCUS Urry Cain Wasserman Minorsky Jackson Reece 6 An Introduction to Metabolism Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge Overview: The Energy of Life The
Laboratory Guide to Biochemistry, Enzymology, and Protein Physical Chemistry. A Study of Aspartate Transcarbamylase
 Laboratory Guide to Biochemistry, Enzymology, and Protein Physical Chemistry A Study of Aspartate Transcarbamylase Laboratory Guide to Biochemistry, Enzymology, and Protein Physical Chemistry A Study of
Laboratory Guide to Biochemistry, Enzymology, and Protein Physical Chemistry A Study of Aspartate Transcarbamylase Laboratory Guide to Biochemistry, Enzymology, and Protein Physical Chemistry A Study of
Chapter 2. The Structure of Atoms. The Structure of Atoms. The Structure of Atoms
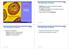 1 The Structure of Atoms 2 Chapter 2 Chemical Principles Chemistry is the study of interactions between atoms and molecules The atom is the smallest unit of matter that enters into chemical reactions Atoms
1 The Structure of Atoms 2 Chapter 2 Chemical Principles Chemistry is the study of interactions between atoms and molecules The atom is the smallest unit of matter that enters into chemical reactions Atoms
Foundations in Microbiology Seventh Edition
 Lecture PowerPoint to accompany Foundations in Microbiology Seventh Edition Talaro Chapter 2 The Chemistry of Biology Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
Lecture PowerPoint to accompany Foundations in Microbiology Seventh Edition Talaro Chapter 2 The Chemistry of Biology Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
https://www.chemicool.com/definition/chromatography.html
 CHROMATOGRAPHY 1 Chromatography - a physical method of mixture separation in which the components to be separated are distributed between two phases, one of which is stationary (stationary phase) while
CHROMATOGRAPHY 1 Chromatography - a physical method of mixture separation in which the components to be separated are distributed between two phases, one of which is stationary (stationary phase) while
CypExpress 3A4 Catalyzed Conversion of Testosterone (TE) to 6β- Hydroxytestosterone (HT)
 TM CASE STUDY CypExpress 3A4 Catalyzed Conversion of Testosterone (TE) to 6β- Hydroxytestosterone (HT) Shuvendu Das, 1 Enrique Martinez, 2 and Mani Subramanian 1 1 Center for Biocatalysis and Bioprocessing,
TM CASE STUDY CypExpress 3A4 Catalyzed Conversion of Testosterone (TE) to 6β- Hydroxytestosterone (HT) Shuvendu Das, 1 Enrique Martinez, 2 and Mani Subramanian 1 1 Center for Biocatalysis and Bioprocessing,
Chromatographic Methods of Analysis Section - 4 : Ion Exchange Chrom. Prof. Tarek A. Fayed
 Chromatographic Methods of Analysis Section - 4 : Ion Exchange Chrom. Prof. Tarek A. Fayed Ion Exchange Chromatography (IEC) In this type of chromatography, the solid stationary phase )organic resin) is
Chromatographic Methods of Analysis Section - 4 : Ion Exchange Chrom. Prof. Tarek A. Fayed Ion Exchange Chromatography (IEC) In this type of chromatography, the solid stationary phase )organic resin) is
Objectives INTRODUCTION TO METABOLISM. Metabolism. Catabolic Pathways. Anabolic Pathways 3/6/2011. How to Read a Chemical Equation
 Objectives INTRODUCTION TO METABOLISM. Chapter 8 Metabolism, Energy, and Life Explain the role of catabolic and anabolic pathways in cell metabolism Distinguish between kinetic and potential energy Distinguish
Objectives INTRODUCTION TO METABOLISM. Chapter 8 Metabolism, Energy, and Life Explain the role of catabolic and anabolic pathways in cell metabolism Distinguish between kinetic and potential energy Distinguish
Chemical Principles. 2-1 Describe the structure of an atom and its relation to the physical properties of elements. 6 C differ from.
 CHAPTER 2 Chemical Principles Learning Objectives 2-1 Describe the structure of an atom and its relation to the physical properties of elements. 2-2 Define ionic bond, covalent bond, hydrogen bond, molecular
CHAPTER 2 Chemical Principles Learning Objectives 2-1 Describe the structure of an atom and its relation to the physical properties of elements. 2-2 Define ionic bond, covalent bond, hydrogen bond, molecular
Human Biology. The Chemistry of Living Things. Concepts and Current Issues. All Matter Consists of Elements Made of Atoms
 2 The Chemistry of Living Things PowerPoint Lecture Slide Presentation Robert J. Sullivan, Marist College Michael D. Johnson Human Biology Concepts and Current Issues THIRD EDITION Copyright 2006 Pearson
2 The Chemistry of Living Things PowerPoint Lecture Slide Presentation Robert J. Sullivan, Marist College Michael D. Johnson Human Biology Concepts and Current Issues THIRD EDITION Copyright 2006 Pearson
camp Direct Immunoassay Kit
 camp Direct Immunoassay Kit Catalog Number KA0886 100 assays Version: 05 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 General Information... 4 Materials
camp Direct Immunoassay Kit Catalog Number KA0886 100 assays Version: 05 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 General Information... 4 Materials
Biology Slide 1 of 20
 Biology 1 of 20 8-1 Energy and Life 2 of 20 8-1 Energy and Life Autotrophs and Heterotrophs Where do plants get the energy they need to produce food? Living things need energy to survive. This energy comes
Biology 1 of 20 8-1 Energy and Life 2 of 20 8-1 Energy and Life Autotrophs and Heterotrophs Where do plants get the energy they need to produce food? Living things need energy to survive. This energy comes
1/23/2012. Atoms. Atoms Atoms - Electron Shells. Chapter 2 Outline. Planetary Models of Elements Chemical Bonds
 Chapter 2 Outline Atoms Chemical Bonds Acids, Bases and the p Scale Organic Molecules Carbohydrates Lipids Proteins Nucleic Acids Are smallest units of the chemical elements Composed of protons, neutrons
Chapter 2 Outline Atoms Chemical Bonds Acids, Bases and the p Scale Organic Molecules Carbohydrates Lipids Proteins Nucleic Acids Are smallest units of the chemical elements Composed of protons, neutrons
Energy Transformation and Metabolism (Outline)
 Energy Transformation and Metabolism (Outline) - Definitions & Laws of Thermodynamics - Overview of energy flow ecosystem - Biochemical processes: Anabolic/endergonic & Catabolic/exergonic - Chemical reactions
Energy Transformation and Metabolism (Outline) - Definitions & Laws of Thermodynamics - Overview of energy flow ecosystem - Biochemical processes: Anabolic/endergonic & Catabolic/exergonic - Chemical reactions
An Introduction to Metabolism
 LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 8 An Introduction to Metabolism
LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 8 An Introduction to Metabolism
