28 Bayesian Mixture Models for Gene Expression and Protein Profiles
|
|
|
- Eustacia Quinn
- 5 years ago
- Views:
Transcription
1 28 Bayesian Mixture Models for Gene Expression and Protein Profiles Michele Guindani, Kim-Anh Do, Peter Müller and Jeff Morris M.D. Anderson Cancer Center 1 Introduction We review the use of semi-parametric mixture models for Bayesian inference in high throughput genomic data. We discuss three specific approaches for microarray data, for protein mass spectrometry experiments, and for SAGE data. For the microarray data and the protein mass spectrometry we assume group comparison experiments, i.e., experiments that seek to identify genes and proteins that are differentially expressed across two biologic conditions of interest. For the SAGE data example we consider inference for a single biologic sample. Several aspects of data analysis for microarray and other high throughput gene and protein expression experiments give rise to mixture models. One important application of mixture models is for flexible modeling of sampling distributions. This is attractive, for example, when the number of genes on a microarray is the relevant sample size, thus allowing flexible semi-parametric representations. Such approaches are discussed, among others, in Broet P (2002), Dahl (2003), or Tadesse et al. (2005). The latter exploit the clustering implicitely defined by the mixture model. Also, see Dahl (2006); Tadesse et al. (2006) in this volume. Related non-bayesian approaches using mixture models have been discussed, for example, in Pan et al. (2002) and McLachlan, Bean, and Peel (2002) KIM: reference?. In this chapter we review three approaches that are typical of this literature. In Section 2 we discuss the use of Dirichlet process mixtures for model based inference about differential gene expression. In Sectin 3 we describe a mixture of Beta model for the mass/charge spectrum in MALDI-TOF mass spectrometry experiments. In Section 4 we introduce a semiparametric mixture of Poisson model for SAGE data. Another important class of applications for mixture models in data analysis for high throughput gene expression data are finite mixtures, with each term in the mixture corresponding to a different condition of interest. A typical example is the model used in Parmigiani et al. (2002) who construct a sampling model for observed gene expression in microarray experiments as a mixture of three terms corresponding to normal, under- and over-expression. Newton et al. 1
2 Density SAMPLE z z z p 0 f 0 + (1 p 0 ) f 1 = f Figure 1: Hypothetical distribution of difference scores for non-differentially expressed (left, f 0 ) and differentially expessed genes (center, f 1 ), and the observed mixture (right, f). (2001) define a Gamma/Gamma hierarchical model with a mixture induced by an indicator for ties between two biologic conditions of interest. In this chapter we only focus on the use of semi-parametric mixtures to represent an unknown sampling model, and will not discuss the other type of mixture models. 2 A Non-parametric Bayesian model for Differential Gene Expression We consider inference for microarray group comparsion experiments. Assume that the data has been summarized as a set of difference scores, z i, i = 1,..., n, for n genes. The difference score z i could be, for example, a two-sample t- statistic for observed fluorescence intensities for gene i in samples under two biologic conditions of interest. See Efron et al. (2001) for a discussion of appropriate data pr-eproscesing. We assume that the set i = 1,..., n of genes is partitioned into a subset of differentially expressed genes and non-differentially expressed genes. Inference proceeds by assuming that for differentially expressed genes, the difference scores z i arise by independent sampling from some unknown distribution f 1 ; for non-differentially expressed genes, z i are independent samples from an unknown distribution f 0. For a reasonable choice of difference scores, the distribution f 0 should be a unimodal distribution centered at zero. The distribution f 1 should be a bimodal distribution with symmetric modes to the left and right of zero corresponding to over- and underexpressed genes. Figure 1 show possible histograms for observed difference scores generated from f 0 and f 1. Of course, the partition into differentially and non-differentially expressed genes is unknown. Thus, instead of samples from f 0 and f 1, we can only work with the sample z i, i = 1,..., n, generated from a mixture of f 0 and f 1. Let p 0 denote the unknown proporition of non-differentially genes. We assume z i iid f(z) = p 0 f 0 (z) + (1 p 0 ) f 1 (z), i = 1,..., n. (1) 2
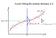
3 The main goal of inference in the two group comparison microarray experiment can be formally described as the deconvolution of (1). We introduce a latent indicator variables r i {0, 1} to rewrite (1) equivalently as a hierarchical model p(z i r i = j) = f j (z i ) P r(r i = 0) = p 0 (2) The latent variable r i can be interpreted as indicator for gene i being differentially expressed. Efron et al. (2001) propose cleverly chosen point estimates for p 0, f 0 and f 1 and report the implied inference for r i. To develop the point estimate they introduce an additional set of difference scores, z i, i = n + 1,..., 2n. The additional difference scores are generated using the same original data, but deliberately computing difference scores for samples under the same biologic conditions. Thus, z i f 0 (z i ), i = n + 1,..., 2n for this additional null sample. In Do et al. (2005) we propose a model-based semiparametric Bayesian approach to inference in this problem. We recognize f 0, f 1 and p 0 as unknown quantities and proceed by defining a suitable prior probability model. Probability models for unknown functions, including distributions such as f 0 and f 1 in this problem, are known as non-parametric Bayesian models. See, for example Müller and Quintana (2004) for a recent review of non-parametric Bayesian inference. The term non-parametric is a misnomer, as the random functions are infinite dimensional parameters. However, the name is traditionally used because implied posterior inference closely resembles inference under classical non-parametric methods. In chosing a prior probability model for f 0 and f 1 we face two competing aims. On one hand we wish to generalize traditional parametric models, like a normal sampling model. On the other hand we want to retain as much computational simplicity as possible. This leads us to use a mixture of normal model, with a non-parametric prior on the mixing measure. Inference under this model is almost as straightforward as under a simple normal model, yet, subject to some technical constraints, the mixture of normal model can approximate arbitrary sampling distributions. As probabilty model for the mixing measure we use a Dirichlet process (DP) prior (Ferguson, 1973; Antoniak, 1974). For reasons of computational simplicity and ease of interpretation the DP prior is one of the most widely used non-parametric Bayes models. The DP model has two parameters, a base measure and a total mass parameter. We write G DP (G, M) to indicate that G has a DP prior with a base measure G and total mass M. The base measure has the interpretation as mean measure, in fact E(G) = G. The total mass parameter can be interpreted as a precision parameter. The larger M, the closer the random G will be to G. In sumary, we assume the following model. Let N(z; m, s) denote a normal distribution for the random variable z, with moments (m, s). We define a 3
4 probability model for the random distributions f 0 and f 1 as: f j (z) = N(z; µ, σ) dg j (µ) G j DP (G j, M). (3) One of the critical properties of the DP prior is that a DP generated random measure is almost surely discrete. Thus the integral in (3) is simply a sum over all point masses in G j. Mixture models with respect to a DP, such as (3), are known as mixture of DP (MDP) models and are widely used in non-parametric Bayesian inference. See, for example, MacEachern and Müller (2000) for a review of such models. We complete the model given by the likelihood (2) and prior (3) with a hyperprior on the base measures G j. We assume G 0 = N(0, τ 2 ) with a conjugate inverse Gamma hyperprior on τ 2, and G 1 = 1 2 N( b, τ 2 ) N(b, τ 2 ) with a conjugate normal hyperprior on b. Finally, we assume a Beta prior for p 0, p 0 Be(α, β). The hyperparameters α, β, M are fixed. Inference in the proposed model is implemented by Markov chain Monte Carlo (MCMC) simulation. See Do et al. (2005) for a detailed description of the posterior MCMC algorithm. A direct implication of the model (2) and (3) is that the marginal posterior probability of differential expression, P r(r i = 1 data), is the same for all genes with equal difference score z i. Thus posterior inference can be summarized as a function P r(r i = 1 z i = z, data). Starting with model (2), a straightforward use of Bayes theorem shows P r(r i = 0 z i = z, f 0, f 1, p 0 ) = p 0 f 0 (z)/[p 0 f 0 (z) + (1 p 0 ) f 1 (z)]. }{{} f(z) Let P 1 = p 0 f 0 /f. Then the posterior expectation P 1 = E(P 1 data) is exactly the desired marginal posterior probability of differential expression, P 1 = P r(r i = 1 z i = z, data). Figure 2 shows posterior inference for a simulation experiment. The figure shows the simulation truth, the reported posterior mean curve P 1 (z), and pointwise posterior credible intervals for P 1 (z). 3 A Mixture of Beta Model for MALDI-TOF Data Matrix assisted laser disorption - time of flight (MALDI-TOF) experiments allow to simultaneously measure abundance for a large number of proteins. Details of the experimental setup are described, for example, in Baggerly et al. (2003). See also Baggerly et al. (2006), in this volume. Briefly, the biological sample for which we wish to determine protein abundance is fixed in a matrix. A laser beam is used to break free and ionize individual protein molecules. The experiment is arranged such that ionized proteins are exposed to an electric field that accelerates molecules along a flight tube. On the other end of the flight 4
5 P(differentially expressed Z) P(differentially expressed Z) Z Z Figure 2: P 1 (z): posterior mean probability of differential expression as a function of the observed difference score z (solid black line). The left figure conditions on the full data, z i, i = 1,..., 2n, including the null data. The right figure does not make use of the null data, conditioning only on z i, i = 1,..., n. The dark grey shaded band shows the central 50% posterior density interval. Light grey shows a 75% posterior interval The dark and light grey shaded areas are very narrow and can hardly be distinguished from the posterior mean curve. tube, molecules hit a detector that records a histogram of number of molecules that hit over time. Assumig that all ionized molecules carry a unit charge, the time of flight is deterministically related to the molecule mass. The histogram of detector events over time can therefore be be changed to a histogram of detector events over protein masses. Allowing for multiple charges, the mass scale is replaced by a scale of mass/charge ratios. The histogram of detector events is known as mass/charge spectrum. Figure 3 shows typical spectra. plot of one spec plot of other spec Figure 3: Spectra for a normal samples (left) and a tumor samples (right), on grid of size I = 60, 000. (plots are big; i have removed them temp ly -- simply two spectra) Ideally, each protein that is present in the original probe should correspond to a peak in the spectrum. Because of the random initial velocities when proteins are ionized by the laser impact we would expect to see peaks rather than sharp lines even in an idealized experiment. Many additional artifacts of the experiment add to the idealized description, leading to an additional baseline that adds to the protein peaks. See the data shown in 3. Assume we observe spectra for experiments k = 1,..., K. Let y k (m i ) denote 5
6 the recorded count for sample k at mass/charge grid point m i, and let f k (m i ) denote the assumed underlying cleaned spectra corresponding to detected proteins only. The desired inference about the unknown protein abundance in the original probes can be formalized as (i) removing noise and baseline from the observed spectra y ki to impute p k ; (ii) finding peaks in p k ; and (iii) reporting the relative sizes of these peaks. The relative size of the peaks corresponds to the relative abundance of the corresponding protein in the probe. If samples are collected under different biologic conditions we need additional inference about different versus equal abundance of different proteins. In Müller et al. (2005) we develop a non-parametric Bayes model to allow such inference. Based on the above stylized description of the experiment we consider y k as the empirical histogram of detector events. We represent it as a mixture of a baseline B k corresponding to detector noise, protein fragments, etc., and a cleaned spectrum f k. p k (m) = p 0k B k (m) + (1 p 0k ) f k (m). The spectrum f k is a sum of peaks, with each detected protein contributing a peak centered at its mass/charge value. The experimental arrangement implies a finite support for f k. Motivated by nonparametric models for random distributions on a finite support developed in Petrone (1999) and Robert and Rousseau (2002) we use a mixture of Beta distributions to define the random distribution f k. The location for each Beta kernel is interpreted as the mass/charge ratio of the protein giving rise to this peak. To facilitate later interpretation, we use a non-standard parametriztion of the Beta distribution. We write Be(x; ɛ, α) for a Beta kernel for the random variable x, with mean and standard deviation ɛ and α (with appropriate constraints on α). Let x denote the biologic condition of sample k. We assume a two group comparison, i.e., x {0, 1}. Then f k (m) = J w xj Beta(m; ɛ j, α j ). (4) j=1 In words, the k-th spectrum is a mixture of Beta kernels, corresponding to J distinct proteins with mass/charge values ɛ j. The relative weight w xj, i.e., relative abundance of protein j, is assumed the same for all samples under the same biologic condition. For reasons of technical convenience we chose an identical mixture of Beta prior for the baseline B k. Different hyperparameters reflect the fact that the baseline is much smoother than f k and we expect fewer terms in the mixture. B k (m) = J k j=1 v kj Beta(m; η kj, β kj ). The sizes of the mixtures are random. We use truncated Poisson priors for J and J k, k = 1,..., K. Baseline B k and mean spectrum f k are combined to define the random distribution of mass/charge ratios p k = p 0k B k + (1 p 0k ) f k. Following the idealized description of the experimental setup, the sampling model is random sampling from p k. Let y k = (y ki, i = 1,..., I) denote the empirical spectrum for the k-th sample over the grid of mass/charge values. Typically I is a large, 6
7 say 60,000, defining a very fine grid. Let θ = (J, J k, w xj, v kj, ɛ j, α j, η i, β i, x = 0, 1, j = 1,..., J, k = 1,..., K, i = 1,..., J k ) denote the parameter vector. The likelihood is I log p(y k θ) = y ki log p k (m i ). (5) i=1 Instead of the random sampling model (5) many authors use a regression likelihood, assuming normal residuals, y ki N(p k (m i ), σ 2 ). Little changes in the following discussion if we were to replace (5) by this regression likelihood. The model is completed with a prior for the Beta parameters and the weights. For the weights w xj we use a hierarchical prior with indicators λ j for ties λ j = I(w 0j = w 1j ). Posterior inference on the λ j and the locations ɛ j summarizes the desired inference on proteins that are differentially expressed across the two groups of samples. Implementation of posterior inference requires MCMC over a varying dimension parameter space, as the dimension of the parameter space depends on the sizes J and J k of the mixtures. We use reversible jump MCMC (RJMCMC) as proposed in Green (1995) and, specifically for mixture models, in Richardson and Green (1997). See Müller et al. (2005) for a detailed description of the MCMC algorithm. A minor complication arises in reporting and summarizing posterior inference about distinct proteins and their mass/charge ratios. The mixture f k only includes exchangeable indices j, leading to the complication that the Beta kernel corresponding to a given protein might have different indices at different iterations of the posterior MCMC simulation. In other words, the protein identity is not part of the probability model. To report posterior inference on the mean abundance of a given protein requires additional post-processing to match Beta kernels that correspond to the same protein across iterations. We use a reasonable ad-hoc rule. Any two peaks j and h with a difference in masses below a certain threshold are counted as arising from the same protein. Specifically, we use the condition ɛ j ɛ h < 0.5α j to match peaks. Here j indexes the peak that was imputed in an earlier MCMC iteration than the peak h. The problem of reporting inference related to the terms in a mixture is known as the label switching problem (C. C. Holmes and Stephens, 2005). Figure 4 summarizes estimated masses and abundance of detected proteins. 4 A Semi-Parametric Mixture Model for SAGE Data Consider data from a SAGE (Serial Analysis of Gene Expression) experiment. See Baggerly et al. (2006) for a description of the experimental setup, and the 7
8 WEIGHT NON DIF DIFF Z Figure 4: Posterior mean abundance of detected proteins. All peaks with posterior probability of differential expression greater 50% are marked as solid dots, with a line combining w 0j = E(w 0j data) and w 1j. nature of the data. We consider inference for data from one biologic sample. Let y i, i = 1,..., k, denote observed tag frequencies for k distinct transcripts. Let n = y i denote the total number of recorded transcripts, and let π i denote the unknown true abundance of the i-th transcript in the probe. For large y i, the empirical frequency π i = y i /n is an appropriate point estimate for π i. The associated uncertainty, formalized as variance of the maximum likelihood estimator or as posterior standard deviation in a suitable model, is negligible. However, for scarce tags, the empirical frequency includes considerable sampling variability, and more elaborate estimates are desireable. Also, when the data includes samples across different biologic conditions, the inference goal might not be restricted to estimating the transcript frequencies. For discrimination and classification additional inference about differences in transcript frequencies, and related probability statements are required. In addition to inference on π i for a specific tag i, one might be interested in the distribution of tag frequencies across different transcripts. This can be achieved by model-based posterior inference. Morris et al. (2003) introduce an approach that is based on a hierarchical model with a mixture of two Dirichlet distributions as population distribution prior for the π i. See Morris et al. (2006) in this volume for a review of this approach. Building on this model, we introduce a semi-parametric Bayesian mixture model, replacing the two-component mixture of Dirichlet distributions by an unknown random measure, with a nonparametric Bayesian prior model. For the following model construction it is convenient not to condition on n. In other words, instead of assuming that the set of observed counts arise as a multinomial sample with cell frequencies π i, we assume that, conditional on 8
9 hyperparameters, the counts y i arise as indpendent samples from some appropriate probability model. Specifically, we assume that the counts y i are sampled from a mixture of Poisson model. Let Poi(x; λ) denote a Poisson distribution for the random variable x with parameter λ. We assume y i Poi(y i ; λ) dg(λ), i = 1,..., n, independently conditional on G. We specify a prior distribution for the mixture model by assuming a nonparametric prior on the mixing measure, chosing a DP prior as in (3), G DP (G, M). (6) The mixture model can alternatively be written as a hierarchical model y i λ i Poi(λ i ) with λ i G. (7) The discrete nature of the DP random measure G implies a positive probability for ties among the λ i. We denote with L the number of distinct values. A minor complication arises from the fact that y i = 0 is not observed; it is censored. Let k 0 denote the number of tags with non-zero count, i.e., the number of tags recorded in a SAGE library as shown in Baggerly et al. (2006). One could augment the model to include inference on k, k k 0. Alternatively, we follow Stollberg et al. (2000), and fix k by imputing a point estimate for the uknown number of unobserved tags, i.e., tags with y i = 0. Model (6) and (7) defines a DP mixture of Poisson distributions. Such models are popular choices for non-parametric Bayesian data analysis. See, for example, MacEachern and Müller (2000) for a review of such models, including implementation of posterior inference by MCMC simulation. Choosing the base measure G to be conjugate with the Poisson distribution we define a conjugate DP mixture, greatly facilitating the MCMC implementation. Let Ga(x; α, β) denote a Gamma distribution with mean α/β. We use G (λ) = Ga(x; α, β), with fixed hyperparameters α and β. To illustrate the model we implemented posterior inference for a SAGE library reported in Zhang et al. (1997). The same data was used in Morris et al. (2003), and is available at It records counts for k 0 = distinct transcripts, with a total number of n = y i = recorded tags. We use the estimte from Stollberg et al. (2000), and set k = 25336, with y i = 0 for i = k 0 + 1,..., k, i.e., we estimate the number of tags with censored counts y i = 0, as I(y i = 0) = Figure 5a shows a histogram of observed counts y i in the data. Figure 6 summarizes posterior inference for the transcript abundances. The figure plots posterior mean estimates E(λ i data) versus observed counts y i. The nature of the shrinkage follows the patter reported in Morris et al. (2003). For censored 9
10 frequency COUNTS Figure 5: Observed tag counts y i. The highly skewed nature is typial for SAGE data. Will still redo the figure in log-log scale. E(LAMBDA data) Frequency COUNTS y L (a) λ i vs. y i (b) p(l dta). Figure 6: Posterior means λ i = E(λ i dta) versus observed counts y i (panel a). Note the strong shrinkage for small counts. For large counts, y i > 6, posterior shrinkage quickly becomes negligible. Posterior distribution for the number of clusters L (panel b). 10
11 E(G data) LAMBDA E(G data) LAMBDA (a) E(G dta), λ < 100 (b) E(G dta) Figure 7: Estimated mixing measure G. The left panel zooms in on the lower end, λ < 100. The highly skewed nature of G = E(G dta) reflects the same feature in the recorded data y i. tags, with y i = 0, the posterior mean estimate inflates the m.l.e. and reports E(λ i data) 0.9. For rare tags with non-zero counts, posterior inference shrinks the maximum likelihood estimate. For abundant tags, posterior inference is driven only by the observed count, and E(λ i data) y i. Figure 7 shows the estimated distribution of tag abundances λ i. 5 Summary We have illustrated the use of mixture models for Bayesian inference for gene expression and proteomics. We focused on the use of mixtures as a flexible class of distributions to parametrize random distributions. Another important use of mixtures arises in models where the submodels in the mixture correspond to different biologic conditions. Such models are extensively reviewed in other chapters in this volume. We introduced DP mixtures of normal models to model microarray gene expression data, DP mixtures of Poissons to model tag counts in SAGE data, and location/scale mixtures of Beta kernels to represent mass/charge spectra in protein mass spectrometry experiments. The underlying theme in all three applications is the use of model-based inference, with a probability model on the random distribution (or mass/charge spectrum). This is in contrast to traditional, and very resonable, multi-step methods. If the goal is only a single point estimate of the gene expression in the microarray experiment, peak location and weight in the mass/charge spectrum, and abundance of a transcript in the SAGE data, then the model based methods are perhaps overly complex. The result will differ little from any reasonable ad-hoc method and might not justify the computational effort. The power of these methods lies in the full 11
12 probabilistic description of all related uncertainties. But many important inference problems go beyond point estimates. For example, consider the decision problem of flagging genes for differential expression, or the problem of identifying a set of proteins that can serve as biomarker panel, or sample size choice for a microarray experiment. A decision theoretic answer to these question relies on a description of all uncertainties in one coherent probability model. Acknowledgments Jeff Morris was supported by NCI grant CA Peter Müller were supported by NCI grant CA Michele Guindani and References Antoniak, C. E. (1974), Mixtures of Dirichlet processes with applications to Bayesian nonparametric problems, The Annals of Statistics, 2, Baggerly, K. A., Coombes, K. R., and Morris, J. S. (2006), Bayesian Inference for Gene Expression and Proteomics, in Do et al. (2006), chap. An Introduction to High-Throughput Bioinformatics Data, pp Baggerly, K. A., Morris, J. S., Wang, J., Gold, D., Xiao, L. C., and Coombes, K. R. (2003), A comprehensive approach to analysis of MALDI-TOF proteomics spectra from serum samples, Proteomics,?,?????? Broet P, Richardson S, R. F. (2002), Bayesian hierarchical model for identifying changes in gene expression from microarray experiments, J Comput Biol., 9, C. C. Holmes, A. J. and Stephens, D. A. (2005), Markov Chain Monte Carlo Methods and the Label Switching Problem in Bayesian Mixture Modeling, Statistical Science, 20, Dahl, D. (2003), Modeling differential gene expression using a Dirichlet Process mixture model, in 2003 Proceedings of the American Statistical Association, Bayesian Statistical Sciences Section, Alexandria, VA: American Statistical Association. (2006), Bayesian Inference for Gene Expression and Proteomics, in Do et al. (2006), chap. Model-Based Clustering for Expression Data via a Dirichlet Process Mixture MOdel, pp Do, K., Müller, P., and Tang, F. (2005), A Bayesian mixture model for differential gene expression, Journal of the Royal Statistical Society C, 54, Do, K.-A., Müller, P., and Vannucci, M. (eds.) (2006), Bayesian Inference for Gene Expression and Proteomics, Cambridge University Press. 12
13 Efron, B., Tibshirani, R., Storey, J. D., and Tusher, V. (2001), Empirical Bayes analysis of a microarray experiment, Journal of the American Statistical Association, 96, Ferguson, T. S. (1973), A Bayesian analysis of some nonparametric problems, The Annals of Statistics, 1, Green, P. J. (1995), Reversible jump Markov chain Monte Carlo computation and Bayesian model determination, Biometrika, 82, MacEachern, S. N. and Müller, P. (2000), Efficient MCMC Schemes for Robust Model Extensions using Encompassing Dirichlet Process Mixture Models, in Robust Bayesian Analysis, eds. Ruggeri, F. and Ríos-Insua, D., New York:Springer-Verlag, pp Morris, J. S., Baggerly, K. A., and Coombes, K. R. (2003), Bayesian Shrinkage Estimation of the Relative Abundance of MRNA Transcripts Using SAGE, Biometrics, 59, (2006), Bayesian Inference for Gene Expression and Proteomics, in Do et al. (2006), chap. Shrinkage Estimation for SAGE Data using a Mixture Dirichlet Prior, pp Müller, P. and Quintana, F. A. (2004), Nonparametric Bayesian Data Analysis, Statistical Science, 19, Newton, M. A., Kendziorsky, C. M., Richmond, C. S., R., B. F., and Tsui, K. W. (2001), On differential variability of expression ratios: improving statistical inference about gene expression changes from microarray data, Journal Computational Biology, 8, Pan, W., Lin, J., and Le, C. T. (2002), How many replicates of arrays are required to detect gene expression changes in microarray experiments? A mixture model approach, Genome Biology, 3(5), research Parmigiani, G., Garrett, E. S., Anbazhagan, R., and Gabrielson, E. (2002), A statistical framework for expression-based molecular classification in cancer, Journal of the Royal Statistical Society B, 64, Petrone, S. (1999), Bayesian density estimation using Bernstein polynomials, Canadian Journal of Statistics, 27, Richardson, S. and Green, P. J. (1997), On Bayesian Analysis of Mixtures with an Unknown Number of Components, Journal of the Royal Statistical Society B, 59, Robert, C. and Rousseau, J. (2002), A mixture approach to Bayesian goodness of fit, Tech. rep., CEREMADE. Stollberg, J. Urschitz, J., Urban, Z., and Boyd, C. (2000), A quantitative evaluation of Sage, Genome Research, 10,
14 Tadesse, M., Sha, N., Kim, S., and Vannucci, M. (2006), Bayesian Inference for Gene Expression and Proteomics, in Do et al. (2006), chap. Identification of Biomarkers in Classification and Clustering of High-Throughput Data, pp Tadesse, M., Sha, N., and Vannucci, M. (2005), Bayesian variable selection in clustering high-dimensional data, Journal of the American Statistical Association, 100, Zhang, L., Zhou, W., Velculescu, V., Kern, S., Hruban, R., Hamilton, S., Vogelstein, B., and Kinzler, K. (1997), Gene Expression Profiles in Normal and Cancer Cells, Science, 276,
29 Sample Size Choice for Microarray Experiments
 29 Sample Size Choice for Microarray Experiments Peter Müller, M.D. Anderson Cancer Center Christian Robert and Judith Rousseau CREST, Paris Abstract We review Bayesian sample size arguments for microarray
29 Sample Size Choice for Microarray Experiments Peter Müller, M.D. Anderson Cancer Center Christian Robert and Judith Rousseau CREST, Paris Abstract We review Bayesian sample size arguments for microarray
Bayesian Nonparametric Regression for Diabetes Deaths
 Bayesian Nonparametric Regression for Diabetes Deaths Brian M. Hartman PhD Student, 2010 Texas A&M University College Station, TX, USA David B. Dahl Assistant Professor Texas A&M University College Station,
Bayesian Nonparametric Regression for Diabetes Deaths Brian M. Hartman PhD Student, 2010 Texas A&M University College Station, TX, USA David B. Dahl Assistant Professor Texas A&M University College Station,
Chapter 2. Data Analysis
 Chapter 2 Data Analysis 2.1. Density Estimation and Survival Analysis The most straightforward application of BNP priors for statistical inference is in density estimation problems. Consider the generic
Chapter 2 Data Analysis 2.1. Density Estimation and Survival Analysis The most straightforward application of BNP priors for statistical inference is in density estimation problems. Consider the generic
A Bayesian Mixture Model for Differential Gene Expression
 A Bayesian Mixture Model for Differential Gene Expression Kim-Anh Do, Peter Müller and Feng Tang 1 Abstract We propose model-based inference for differential gene expression, using a non-parametric Bayesian
A Bayesian Mixture Model for Differential Gene Expression Kim-Anh Do, Peter Müller and Feng Tang 1 Abstract We propose model-based inference for differential gene expression, using a non-parametric Bayesian
Simultaneous Inference for Multiple Testing and Clustering via Dirichlet Process Mixture Models
 Simultaneous Inference for Multiple Testing and Clustering via Dirichlet Process Mixture Models David B. Dahl Department of Statistics Texas A&M University Marina Vannucci, Michael Newton, & Qianxing Mo
Simultaneous Inference for Multiple Testing and Clustering via Dirichlet Process Mixture Models David B. Dahl Department of Statistics Texas A&M University Marina Vannucci, Michael Newton, & Qianxing Mo
Distance-Based Probability Distribution for Set Partitions with Applications to Bayesian Nonparametrics
 Distance-Based Probability Distribution for Set Partitions with Applications to Bayesian Nonparametrics David B. Dahl August 5, 2008 Abstract Integration of several types of data is a burgeoning field.
Distance-Based Probability Distribution for Set Partitions with Applications to Bayesian Nonparametrics David B. Dahl August 5, 2008 Abstract Integration of several types of data is a burgeoning field.
Contents. Part I: Fundamentals of Bayesian Inference 1
 Contents Preface xiii Part I: Fundamentals of Bayesian Inference 1 1 Probability and inference 3 1.1 The three steps of Bayesian data analysis 3 1.2 General notation for statistical inference 4 1.3 Bayesian
Contents Preface xiii Part I: Fundamentals of Bayesian Inference 1 1 Probability and inference 3 1.1 The three steps of Bayesian data analysis 3 1.2 General notation for statistical inference 4 1.3 Bayesian
Bayesian Methods for Machine Learning
 Bayesian Methods for Machine Learning CS 584: Big Data Analytics Material adapted from Radford Neal s tutorial (http://ftp.cs.utoronto.ca/pub/radford/bayes-tut.pdf), Zoubin Ghahramni (http://hunch.net/~coms-4771/zoubin_ghahramani_bayesian_learning.pdf),
Bayesian Methods for Machine Learning CS 584: Big Data Analytics Material adapted from Radford Neal s tutorial (http://ftp.cs.utoronto.ca/pub/radford/bayes-tut.pdf), Zoubin Ghahramni (http://hunch.net/~coms-4771/zoubin_ghahramani_bayesian_learning.pdf),
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
Default Priors and Effcient Posterior Computation in Bayesian
 Default Priors and Effcient Posterior Computation in Bayesian Factor Analysis January 16, 2010 Presented by Eric Wang, Duke University Background and Motivation A Brief Review of Parameter Expansion Literature
Default Priors and Effcient Posterior Computation in Bayesian Factor Analysis January 16, 2010 Presented by Eric Wang, Duke University Background and Motivation A Brief Review of Parameter Expansion Literature
Simultaneous inference for multiple testing and clustering via a Dirichlet process mixture model
 Simultaneous inference for multiple testing and clustering via a Dirichlet process mixture model David B Dahl 1, Qianxing Mo 2 and Marina Vannucci 3 1 Texas A&M University, US 2 Memorial Sloan-Kettering
Simultaneous inference for multiple testing and clustering via a Dirichlet process mixture model David B Dahl 1, Qianxing Mo 2 and Marina Vannucci 3 1 Texas A&M University, US 2 Memorial Sloan-Kettering
Parametric Empirical Bayes Methods for Microarrays
 Parametric Empirical Bayes Methods for Microarrays Ming Yuan, Deepayan Sarkar, Michael Newton and Christina Kendziorski April 30, 2018 Contents 1 Introduction 1 2 General Model Structure: Two Conditions
Parametric Empirical Bayes Methods for Microarrays Ming Yuan, Deepayan Sarkar, Michael Newton and Christina Kendziorski April 30, 2018 Contents 1 Introduction 1 2 General Model Structure: Two Conditions
Optimal Sample Size for Multiple Testing: the Case of Gene Expression Microarrays
 Johns Hopkins University, Dept. of Biostatistics Working Papers 2-2-2004 Optimal Sample Size for Multiple Testing: the Case of Gene Expression Microarrays Peter Muller University of Texas M.D. Anderson
Johns Hopkins University, Dept. of Biostatistics Working Papers 2-2-2004 Optimal Sample Size for Multiple Testing: the Case of Gene Expression Microarrays Peter Muller University of Texas M.D. Anderson
Bayesian Regression Linear and Logistic Regression
 When we want more than point estimates Bayesian Regression Linear and Logistic Regression Nicole Beckage Ordinary Least Squares Regression and Lasso Regression return only point estimates But what if we
When we want more than point estimates Bayesian Regression Linear and Logistic Regression Nicole Beckage Ordinary Least Squares Regression and Lasso Regression return only point estimates But what if we
A Bayesian Nonparametric Model for Predicting Disease Status Using Longitudinal Profiles
 A Bayesian Nonparametric Model for Predicting Disease Status Using Longitudinal Profiles Jeremy Gaskins Department of Bioinformatics & Biostatistics University of Louisville Joint work with Claudio Fuentes
A Bayesian Nonparametric Model for Predicting Disease Status Using Longitudinal Profiles Jeremy Gaskins Department of Bioinformatics & Biostatistics University of Louisville Joint work with Claudio Fuentes
Nonparametric Bayesian Methods (Gaussian Processes)
 [70240413 Statistical Machine Learning, Spring, 2015] Nonparametric Bayesian Methods (Gaussian Processes) Jun Zhu dcszj@mail.tsinghua.edu.cn http://bigml.cs.tsinghua.edu.cn/~jun State Key Lab of Intelligent
[70240413 Statistical Machine Learning, Spring, 2015] Nonparametric Bayesian Methods (Gaussian Processes) Jun Zhu dcszj@mail.tsinghua.edu.cn http://bigml.cs.tsinghua.edu.cn/~jun State Key Lab of Intelligent
Petr Volf. Model for Difference of Two Series of Poisson-like Count Data
 Petr Volf Institute of Information Theory and Automation Academy of Sciences of the Czech Republic Pod vodárenskou věží 4, 182 8 Praha 8 e-mail: volf@utia.cas.cz Model for Difference of Two Series of Poisson-like
Petr Volf Institute of Information Theory and Automation Academy of Sciences of the Czech Republic Pod vodárenskou věží 4, 182 8 Praha 8 e-mail: volf@utia.cas.cz Model for Difference of Two Series of Poisson-like
Non-Parametric Bayes
 Non-Parametric Bayes Mark Schmidt UBC Machine Learning Reading Group January 2016 Current Hot Topics in Machine Learning Bayesian learning includes: Gaussian processes. Approximate inference. Bayesian
Non-Parametric Bayes Mark Schmidt UBC Machine Learning Reading Group January 2016 Current Hot Topics in Machine Learning Bayesian learning includes: Gaussian processes. Approximate inference. Bayesian
Local-Mass Preserving Prior Distributions for Nonparametric Bayesian Models
 Bayesian Analysis (2014) 9, Number 2, pp. 307 330 Local-Mass Preserving Prior Distributions for Nonparametric Bayesian Models Juhee Lee Steven N. MacEachern Yiling Lu Gordon B. Mills Abstract. We address
Bayesian Analysis (2014) 9, Number 2, pp. 307 330 Local-Mass Preserving Prior Distributions for Nonparametric Bayesian Models Juhee Lee Steven N. MacEachern Yiling Lu Gordon B. Mills Abstract. We address
Stat 542: Item Response Theory Modeling Using The Extended Rank Likelihood
 Stat 542: Item Response Theory Modeling Using The Extended Rank Likelihood Jonathan Gruhl March 18, 2010 1 Introduction Researchers commonly apply item response theory (IRT) models to binary and ordinal
Stat 542: Item Response Theory Modeling Using The Extended Rank Likelihood Jonathan Gruhl March 18, 2010 1 Introduction Researchers commonly apply item response theory (IRT) models to binary and ordinal
Hybrid Dirichlet processes for functional data
 Hybrid Dirichlet processes for functional data Sonia Petrone Università Bocconi, Milano Joint work with Michele Guindani - U.T. MD Anderson Cancer Center, Houston and Alan Gelfand - Duke University, USA
Hybrid Dirichlet processes for functional data Sonia Petrone Università Bocconi, Milano Joint work with Michele Guindani - U.T. MD Anderson Cancer Center, Houston and Alan Gelfand - Duke University, USA
STA 4273H: Sta-s-cal Machine Learning
 STA 4273H: Sta-s-cal Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistical Sciences! rsalakhu@cs.toronto.edu! h0p://www.cs.utoronto.ca/~rsalakhu/ Lecture 2 In our
STA 4273H: Sta-s-cal Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistical Sciences! rsalakhu@cs.toronto.edu! h0p://www.cs.utoronto.ca/~rsalakhu/ Lecture 2 In our
A Fully Nonparametric Modeling Approach to. BNP Binary Regression
 A Fully Nonparametric Modeling Approach to Binary Regression Maria Department of Applied Mathematics and Statistics University of California, Santa Cruz SBIES, April 27-28, 2012 Outline 1 2 3 Simulation
A Fully Nonparametric Modeling Approach to Binary Regression Maria Department of Applied Mathematics and Statistics University of California, Santa Cruz SBIES, April 27-28, 2012 Outline 1 2 3 Simulation
Flexible Regression Modeling using Bayesian Nonparametric Mixtures
 Flexible Regression Modeling using Bayesian Nonparametric Mixtures Athanasios Kottas Department of Applied Mathematics and Statistics University of California, Santa Cruz Department of Statistics Brigham
Flexible Regression Modeling using Bayesian Nonparametric Mixtures Athanasios Kottas Department of Applied Mathematics and Statistics University of California, Santa Cruz Department of Statistics Brigham
COS513 LECTURE 8 STATISTICAL CONCEPTS
 COS513 LECTURE 8 STATISTICAL CONCEPTS NIKOLAI SLAVOV AND ANKUR PARIKH 1. MAKING MEANINGFUL STATEMENTS FROM JOINT PROBABILITY DISTRIBUTIONS. A graphical model (GM) represents a family of probability distributions
COS513 LECTURE 8 STATISTICAL CONCEPTS NIKOLAI SLAVOV AND ANKUR PARIKH 1. MAKING MEANINGFUL STATEMENTS FROM JOINT PROBABILITY DISTRIBUTIONS. A graphical model (GM) represents a family of probability distributions
The Jackknife-Like Method for Assessing Uncertainty of Point Estimates for Bayesian Estimation in a Finite Gaussian Mixture Model
 Thai Journal of Mathematics : 45 58 Special Issue: Annual Meeting in Mathematics 207 http://thaijmath.in.cmu.ac.th ISSN 686-0209 The Jackknife-Like Method for Assessing Uncertainty of Point Estimates for
Thai Journal of Mathematics : 45 58 Special Issue: Annual Meeting in Mathematics 207 http://thaijmath.in.cmu.ac.th ISSN 686-0209 The Jackknife-Like Method for Assessing Uncertainty of Point Estimates for
BAYESIAN METHODS FOR VARIABLE SELECTION WITH APPLICATIONS TO HIGH-DIMENSIONAL DATA
 BAYESIAN METHODS FOR VARIABLE SELECTION WITH APPLICATIONS TO HIGH-DIMENSIONAL DATA Intro: Course Outline and Brief Intro to Marina Vannucci Rice University, USA PASI-CIMAT 04/28-30/2010 Marina Vannucci
BAYESIAN METHODS FOR VARIABLE SELECTION WITH APPLICATIONS TO HIGH-DIMENSIONAL DATA Intro: Course Outline and Brief Intro to Marina Vannucci Rice University, USA PASI-CIMAT 04/28-30/2010 Marina Vannucci
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 7 Approximate
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 7 Approximate
Multilevel Statistical Models: 3 rd edition, 2003 Contents
 Multilevel Statistical Models: 3 rd edition, 2003 Contents Preface Acknowledgements Notation Two and three level models. A general classification notation and diagram Glossary Chapter 1 An introduction
Multilevel Statistical Models: 3 rd edition, 2003 Contents Preface Acknowledgements Notation Two and three level models. A general classification notation and diagram Glossary Chapter 1 An introduction
Aspects of Feature Selection in Mass Spec Proteomic Functional Data
 Aspects of Feature Selection in Mass Spec Proteomic Functional Data Phil Brown University of Kent Newton Institute 11th December 2006 Based on work with Jeff Morris, and others at MD Anderson, Houston
Aspects of Feature Selection in Mass Spec Proteomic Functional Data Phil Brown University of Kent Newton Institute 11th December 2006 Based on work with Jeff Morris, and others at MD Anderson, Houston
A Bayesian Nonparametric Approach to Monotone Missing Data in Longitudinal Studies with Informative Missingness
 A Bayesian Nonparametric Approach to Monotone Missing Data in Longitudinal Studies with Informative Missingness A. Linero and M. Daniels UF, UT-Austin SRC 2014, Galveston, TX 1 Background 2 Working model
A Bayesian Nonparametric Approach to Monotone Missing Data in Longitudinal Studies with Informative Missingness A. Linero and M. Daniels UF, UT-Austin SRC 2014, Galveston, TX 1 Background 2 Working model
Bayesian Nonparametrics for Speech and Signal Processing
 Bayesian Nonparametrics for Speech and Signal Processing Michael I. Jordan University of California, Berkeley June 28, 2011 Acknowledgments: Emily Fox, Erik Sudderth, Yee Whye Teh, and Romain Thibaux Computer
Bayesian Nonparametrics for Speech and Signal Processing Michael I. Jordan University of California, Berkeley June 28, 2011 Acknowledgments: Emily Fox, Erik Sudderth, Yee Whye Teh, and Romain Thibaux Computer
Curve Fitting Re-visited, Bishop1.2.5
 Curve Fitting Re-visited, Bishop1.2.5 Maximum Likelihood Bishop 1.2.5 Model Likelihood differentiation p(t x, w, β) = Maximum Likelihood N N ( t n y(x n, w), β 1). (1.61) n=1 As we did in the case of the
Curve Fitting Re-visited, Bishop1.2.5 Maximum Likelihood Bishop 1.2.5 Model Likelihood differentiation p(t x, w, β) = Maximum Likelihood N N ( t n y(x n, w), β 1). (1.61) n=1 As we did in the case of the
Bayesian Learning. HT2015: SC4 Statistical Data Mining and Machine Learning. Maximum Likelihood Principle. The Bayesian Learning Framework
 HT5: SC4 Statistical Data Mining and Machine Learning Dino Sejdinovic Department of Statistics Oxford http://www.stats.ox.ac.uk/~sejdinov/sdmml.html Maximum Likelihood Principle A generative model for
HT5: SC4 Statistical Data Mining and Machine Learning Dino Sejdinovic Department of Statistics Oxford http://www.stats.ox.ac.uk/~sejdinov/sdmml.html Maximum Likelihood Principle A generative model for
Approximate Bayesian Computation
 Approximate Bayesian Computation Michael Gutmann https://sites.google.com/site/michaelgutmann University of Helsinki and Aalto University 1st December 2015 Content Two parts: 1. The basics of approximate
Approximate Bayesian Computation Michael Gutmann https://sites.google.com/site/michaelgutmann University of Helsinki and Aalto University 1st December 2015 Content Two parts: 1. The basics of approximate
Bayesian semiparametric modeling and inference with mixtures of symmetric distributions
 Bayesian semiparametric modeling and inference with mixtures of symmetric distributions Athanasios Kottas 1 and Gilbert W. Fellingham 2 1 Department of Applied Mathematics and Statistics, University of
Bayesian semiparametric modeling and inference with mixtures of symmetric distributions Athanasios Kottas 1 and Gilbert W. Fellingham 2 1 Department of Applied Mathematics and Statistics, University of
Construction of Dependent Dirichlet Processes based on Poisson Processes
 1 / 31 Construction of Dependent Dirichlet Processes based on Poisson Processes Dahua Lin Eric Grimson John Fisher CSAIL MIT NIPS 2010 Outstanding Student Paper Award Presented by Shouyuan Chen Outline
1 / 31 Construction of Dependent Dirichlet Processes based on Poisson Processes Dahua Lin Eric Grimson John Fisher CSAIL MIT NIPS 2010 Outstanding Student Paper Award Presented by Shouyuan Chen Outline
Part 8: GLMs and Hierarchical LMs and GLMs
 Part 8: GLMs and Hierarchical LMs and GLMs 1 Example: Song sparrow reproductive success Arcese et al., (1992) provide data on a sample from a population of 52 female song sparrows studied over the course
Part 8: GLMs and Hierarchical LMs and GLMs 1 Example: Song sparrow reproductive success Arcese et al., (1992) provide data on a sample from a population of 52 female song sparrows studied over the course
Nonparametric Bayesian Methods - Lecture I
 Nonparametric Bayesian Methods - Lecture I Harry van Zanten Korteweg-de Vries Institute for Mathematics CRiSM Masterclass, April 4-6, 2016 Overview of the lectures I Intro to nonparametric Bayesian statistics
Nonparametric Bayesian Methods - Lecture I Harry van Zanten Korteweg-de Vries Institute for Mathematics CRiSM Masterclass, April 4-6, 2016 Overview of the lectures I Intro to nonparametric Bayesian statistics
Computational statistics
 Computational statistics Markov Chain Monte Carlo methods Thierry Denœux March 2017 Thierry Denœux Computational statistics March 2017 1 / 71 Contents of this chapter When a target density f can be evaluated
Computational statistics Markov Chain Monte Carlo methods Thierry Denœux March 2017 Thierry Denœux Computational statistics March 2017 1 / 71 Contents of this chapter When a target density f can be evaluated
Stat 5101 Lecture Notes
 Stat 5101 Lecture Notes Charles J. Geyer Copyright 1998, 1999, 2000, 2001 by Charles J. Geyer May 7, 2001 ii Stat 5101 (Geyer) Course Notes Contents 1 Random Variables and Change of Variables 1 1.1 Random
Stat 5101 Lecture Notes Charles J. Geyer Copyright 1998, 1999, 2000, 2001 by Charles J. Geyer May 7, 2001 ii Stat 5101 (Geyer) Course Notes Contents 1 Random Variables and Change of Variables 1 1.1 Random
Pattern Recognition and Machine Learning. Bishop Chapter 2: Probability Distributions
 Pattern Recognition and Machine Learning Chapter 2: Probability Distributions Cécile Amblard Alex Kläser Jakob Verbeek October 11, 27 Probability Distributions: General Density Estimation: given a finite
Pattern Recognition and Machine Learning Chapter 2: Probability Distributions Cécile Amblard Alex Kläser Jakob Verbeek October 11, 27 Probability Distributions: General Density Estimation: given a finite
PATTERN RECOGNITION AND MACHINE LEARNING CHAPTER 2: PROBABILITY DISTRIBUTIONS
 PATTERN RECOGNITION AND MACHINE LEARNING CHAPTER 2: PROBABILITY DISTRIBUTIONS Parametric Distributions Basic building blocks: Need to determine given Representation: or? Recall Curve Fitting Binary Variables
PATTERN RECOGNITION AND MACHINE LEARNING CHAPTER 2: PROBABILITY DISTRIBUTIONS Parametric Distributions Basic building blocks: Need to determine given Representation: or? Recall Curve Fitting Binary Variables
The Mixture Approach for Simulating New Families of Bivariate Distributions with Specified Correlations
 The Mixture Approach for Simulating New Families of Bivariate Distributions with Specified Correlations John R. Michael, Significance, Inc. and William R. Schucany, Southern Methodist University The mixture
The Mixture Approach for Simulating New Families of Bivariate Distributions with Specified Correlations John R. Michael, Significance, Inc. and William R. Schucany, Southern Methodist University The mixture
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistical Sciences! rsalakhu@cs.toronto.edu! h0p://www.cs.utoronto.ca/~rsalakhu/ Lecture 7 Approximate
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistical Sciences! rsalakhu@cs.toronto.edu! h0p://www.cs.utoronto.ca/~rsalakhu/ Lecture 7 Approximate
Or How to select variables Using Bayesian LASSO
 Or How to select variables Using Bayesian LASSO x 1 x 2 x 3 x 4 Or How to select variables Using Bayesian LASSO x 1 x 2 x 3 x 4 Or How to select variables Using Bayesian LASSO On Bayesian Variable Selection
Or How to select variables Using Bayesian LASSO x 1 x 2 x 3 x 4 Or How to select variables Using Bayesian LASSO x 1 x 2 x 3 x 4 Or How to select variables Using Bayesian LASSO On Bayesian Variable Selection
Modelling geoadditive survival data
 Modelling geoadditive survival data Thomas Kneib & Ludwig Fahrmeir Department of Statistics, Ludwig-Maximilians-University Munich 1. Leukemia survival data 2. Structured hazard regression 3. Mixed model
Modelling geoadditive survival data Thomas Kneib & Ludwig Fahrmeir Department of Statistics, Ludwig-Maximilians-University Munich 1. Leukemia survival data 2. Structured hazard regression 3. Mixed model
Bayesian Wavelet-Based Functional Mixed Models
 Bayesian Wavelet-Based Functional Mixed Models Jeffrey S. Morris U M.D. Anderson Cancer Center Raymond J. Carroll exas A&M University Functional Data Analysis Functional Data: Ideal units of observation:
Bayesian Wavelet-Based Functional Mixed Models Jeffrey S. Morris U M.D. Anderson Cancer Center Raymond J. Carroll exas A&M University Functional Data Analysis Functional Data: Ideal units of observation:
Lecture: Gaussian Process Regression. STAT 6474 Instructor: Hongxiao Zhu
 Lecture: Gaussian Process Regression STAT 6474 Instructor: Hongxiao Zhu Motivation Reference: Marc Deisenroth s tutorial on Robot Learning. 2 Fast Learning for Autonomous Robots with Gaussian Processes
Lecture: Gaussian Process Regression STAT 6474 Instructor: Hongxiao Zhu Motivation Reference: Marc Deisenroth s tutorial on Robot Learning. 2 Fast Learning for Autonomous Robots with Gaussian Processes
Motivation Scale Mixutres of Normals Finite Gaussian Mixtures Skew-Normal Models. Mixture Models. Econ 690. Purdue University
 Econ 690 Purdue University In virtually all of the previous lectures, our models have made use of normality assumptions. From a computational point of view, the reason for this assumption is clear: combined
Econ 690 Purdue University In virtually all of the previous lectures, our models have made use of normality assumptions. From a computational point of view, the reason for this assumption is clear: combined
A Nonparametric Bayesian Model for Multivariate Ordinal Data
 A Nonparametric Bayesian Model for Multivariate Ordinal Data Athanasios Kottas, University of California at Santa Cruz Peter Müller, The University of Texas M. D. Anderson Cancer Center Fernando A. Quintana,
A Nonparametric Bayesian Model for Multivariate Ordinal Data Athanasios Kottas, University of California at Santa Cruz Peter Müller, The University of Texas M. D. Anderson Cancer Center Fernando A. Quintana,
A Bayesian Approach to Phylogenetics
 A Bayesian Approach to Phylogenetics Niklas Wahlberg Based largely on slides by Paul Lewis (www.eeb.uconn.edu) An Introduction to Bayesian Phylogenetics Bayesian inference in general Markov chain Monte
A Bayesian Approach to Phylogenetics Niklas Wahlberg Based largely on slides by Paul Lewis (www.eeb.uconn.edu) An Introduction to Bayesian Phylogenetics Bayesian inference in general Markov chain Monte
Time Series and Dynamic Models
 Time Series and Dynamic Models Section 1 Intro to Bayesian Inference Carlos M. Carvalho The University of Texas at Austin 1 Outline 1 1. Foundations of Bayesian Statistics 2. Bayesian Estimation 3. The
Time Series and Dynamic Models Section 1 Intro to Bayesian Inference Carlos M. Carvalho The University of Texas at Austin 1 Outline 1 1. Foundations of Bayesian Statistics 2. Bayesian Estimation 3. The
Hierarchical Mixture Models for Expression Profiles
 2 Hierarchical Mixture Models for Expression Profiles MICHAEL A. NEWTON, PING WANG, AND CHRISTINA KENDZIORSKI University of Wisconsin at Madison Abstract A class of probability models for inference about
2 Hierarchical Mixture Models for Expression Profiles MICHAEL A. NEWTON, PING WANG, AND CHRISTINA KENDZIORSKI University of Wisconsin at Madison Abstract A class of probability models for inference about
Bayesian nonparametric estimation of finite population quantities in absence of design information on nonsampled units
 Bayesian nonparametric estimation of finite population quantities in absence of design information on nonsampled units Sahar Z Zangeneh Robert W. Keener Roderick J.A. Little Abstract In Probability proportional
Bayesian nonparametric estimation of finite population quantities in absence of design information on nonsampled units Sahar Z Zangeneh Robert W. Keener Roderick J.A. Little Abstract In Probability proportional
Bayesian Mixture Modeling of Significant P Values: A Meta-Analytic Method to Estimate the Degree of Contamination from H 0 : Supplemental Material
 Bayesian Mixture Modeling of Significant P Values: A Meta-Analytic Method to Estimate the Degree of Contamination from H 0 : Supplemental Material Quentin Frederik Gronau 1, Monique Duizer 1, Marjan Bakker
Bayesian Mixture Modeling of Significant P Values: A Meta-Analytic Method to Estimate the Degree of Contamination from H 0 : Supplemental Material Quentin Frederik Gronau 1, Monique Duizer 1, Marjan Bakker
Ronald Christensen. University of New Mexico. Albuquerque, New Mexico. Wesley Johnson. University of California, Irvine. Irvine, California
 Texts in Statistical Science Bayesian Ideas and Data Analysis An Introduction for Scientists and Statisticians Ronald Christensen University of New Mexico Albuquerque, New Mexico Wesley Johnson University
Texts in Statistical Science Bayesian Ideas and Data Analysis An Introduction for Scientists and Statisticians Ronald Christensen University of New Mexico Albuquerque, New Mexico Wesley Johnson University
Bayesian non-parametric model to longitudinally predict churn
 Bayesian non-parametric model to longitudinally predict churn Bruno Scarpa Università di Padova Conference of European Statistics Stakeholders Methodologists, Producers and Users of European Statistics
Bayesian non-parametric model to longitudinally predict churn Bruno Scarpa Università di Padova Conference of European Statistics Stakeholders Methodologists, Producers and Users of European Statistics
One-shot Learning of Poisson Distributions Information Theory of Audic-Claverie Statistic for Analyzing cdna Arrays
 One-shot Learning of Poisson Distributions Information Theory of Audic-Claverie Statistic for Analyzing cdna Arrays Peter Tiňo School of Computer Science University of Birmingham, UK One-shot Learning
One-shot Learning of Poisson Distributions Information Theory of Audic-Claverie Statistic for Analyzing cdna Arrays Peter Tiňo School of Computer Science University of Birmingham, UK One-shot Learning
Density Estimation. Seungjin Choi
 Density Estimation Seungjin Choi Department of Computer Science and Engineering Pohang University of Science and Technology 77 Cheongam-ro, Nam-gu, Pohang 37673, Korea seungjin@postech.ac.kr http://mlg.postech.ac.kr/
Density Estimation Seungjin Choi Department of Computer Science and Engineering Pohang University of Science and Technology 77 Cheongam-ro, Nam-gu, Pohang 37673, Korea seungjin@postech.ac.kr http://mlg.postech.ac.kr/
Nonparametric Bayesian modeling for dynamic ordinal regression relationships
 Nonparametric Bayesian modeling for dynamic ordinal regression relationships Athanasios Kottas Department of Applied Mathematics and Statistics, University of California, Santa Cruz Joint work with Maria
Nonparametric Bayesian modeling for dynamic ordinal regression relationships Athanasios Kottas Department of Applied Mathematics and Statistics, University of California, Santa Cruz Joint work with Maria
Chris Fraley and Daniel Percival. August 22, 2008, revised May 14, 2010
 Model-Averaged l 1 Regularization using Markov Chain Monte Carlo Model Composition Technical Report No. 541 Department of Statistics, University of Washington Chris Fraley and Daniel Percival August 22,
Model-Averaged l 1 Regularization using Markov Chain Monte Carlo Model Composition Technical Report No. 541 Department of Statistics, University of Washington Chris Fraley and Daniel Percival August 22,
Bayesian Clustering with the Dirichlet Process: Issues with priors and interpreting MCMC. Shane T. Jensen
 Bayesian Clustering with the Dirichlet Process: Issues with priors and interpreting MCMC Shane T. Jensen Department of Statistics The Wharton School, University of Pennsylvania stjensen@wharton.upenn.edu
Bayesian Clustering with the Dirichlet Process: Issues with priors and interpreting MCMC Shane T. Jensen Department of Statistics The Wharton School, University of Pennsylvania stjensen@wharton.upenn.edu
Spatial Bayesian Nonparametrics for Natural Image Segmentation
 Spatial Bayesian Nonparametrics for Natural Image Segmentation Erik Sudderth Brown University Joint work with Michael Jordan University of California Soumya Ghosh Brown University Parsing Visual Scenes
Spatial Bayesian Nonparametrics for Natural Image Segmentation Erik Sudderth Brown University Joint work with Michael Jordan University of California Soumya Ghosh Brown University Parsing Visual Scenes
Supplementary Materials for Molecular QTL Discovery Incorporating Genomic Annotations using Bayesian False Discovery Rate Control
 Supplementary Materials for Molecular QTL Discovery Incorporating Genomic Annotations using Bayesian False Discovery Rate Control Xiaoquan Wen Department of Biostatistics, University of Michigan A Model
Supplementary Materials for Molecular QTL Discovery Incorporating Genomic Annotations using Bayesian False Discovery Rate Control Xiaoquan Wen Department of Biostatistics, University of Michigan A Model
Slice Sampling Mixture Models
 Slice Sampling Mixture Models Maria Kalli, Jim E. Griffin & Stephen G. Walker Centre for Health Services Studies, University of Kent Institute of Mathematics, Statistics & Actuarial Science, University
Slice Sampling Mixture Models Maria Kalli, Jim E. Griffin & Stephen G. Walker Centre for Health Services Studies, University of Kent Institute of Mathematics, Statistics & Actuarial Science, University
The Bayesian Choice. Christian P. Robert. From Decision-Theoretic Foundations to Computational Implementation. Second Edition.
 Christian P. Robert The Bayesian Choice From Decision-Theoretic Foundations to Computational Implementation Second Edition With 23 Illustrations ^Springer" Contents Preface to the Second Edition Preface
Christian P. Robert The Bayesian Choice From Decision-Theoretic Foundations to Computational Implementation Second Edition With 23 Illustrations ^Springer" Contents Preface to the Second Edition Preface
STA414/2104 Statistical Methods for Machine Learning II
 STA414/2104 Statistical Methods for Machine Learning II Murat A. Erdogdu & David Duvenaud Department of Computer Science Department of Statistical Sciences Lecture 3 Slide credits: Russ Salakhutdinov Announcements
STA414/2104 Statistical Methods for Machine Learning II Murat A. Erdogdu & David Duvenaud Department of Computer Science Department of Statistical Sciences Lecture 3 Slide credits: Russ Salakhutdinov Announcements
Introduction to Probabilistic Machine Learning
 Introduction to Probabilistic Machine Learning Piyush Rai Dept. of CSE, IIT Kanpur (Mini-course 1) Nov 03, 2015 Piyush Rai (IIT Kanpur) Introduction to Probabilistic Machine Learning 1 Machine Learning
Introduction to Probabilistic Machine Learning Piyush Rai Dept. of CSE, IIT Kanpur (Mini-course 1) Nov 03, 2015 Piyush Rai (IIT Kanpur) Introduction to Probabilistic Machine Learning 1 Machine Learning
Bayesian estimation of the discrepancy with misspecified parametric models
 Bayesian estimation of the discrepancy with misspecified parametric models Pierpaolo De Blasi University of Torino & Collegio Carlo Alberto Bayesian Nonparametrics workshop ICERM, 17-21 September 2012
Bayesian estimation of the discrepancy with misspecified parametric models Pierpaolo De Blasi University of Torino & Collegio Carlo Alberto Bayesian Nonparametrics workshop ICERM, 17-21 September 2012
Statistical Inference
 Statistical Inference Liu Yang Florida State University October 27, 2016 Liu Yang, Libo Wang (Florida State University) Statistical Inference October 27, 2016 1 / 27 Outline The Bayesian Lasso Trevor Park
Statistical Inference Liu Yang Florida State University October 27, 2016 Liu Yang, Libo Wang (Florida State University) Statistical Inference October 27, 2016 1 / 27 Outline The Bayesian Lasso Trevor Park
Markov Chain Monte Carlo methods
 Markov Chain Monte Carlo methods Tomas McKelvey and Lennart Svensson Signal Processing Group Department of Signals and Systems Chalmers University of Technology, Sweden November 26, 2012 Today s learning
Markov Chain Monte Carlo methods Tomas McKelvey and Lennart Svensson Signal Processing Group Department of Signals and Systems Chalmers University of Technology, Sweden November 26, 2012 Today s learning
Gaussian kernel GARCH models
 Gaussian kernel GARCH models Xibin (Bill) Zhang and Maxwell L. King Department of Econometrics and Business Statistics Faculty of Business and Economics 7 June 2013 Motivation A regression model is often
Gaussian kernel GARCH models Xibin (Bill) Zhang and Maxwell L. King Department of Econometrics and Business Statistics Faculty of Business and Economics 7 June 2013 Motivation A regression model is often
Wavelet-Based Preprocessing Methods for Mass Spectrometry Data
 Wavelet-Based Preprocessing Methods for Mass Spectrometry Data Jeffrey S. Morris Department of Biostatistics and Applied Mathematics UT M.D. Anderson Cancer Center Overview Background and Motivation Preprocessing
Wavelet-Based Preprocessing Methods for Mass Spectrometry Data Jeffrey S. Morris Department of Biostatistics and Applied Mathematics UT M.D. Anderson Cancer Center Overview Background and Motivation Preprocessing
Bayesian Statistics. Debdeep Pati Florida State University. April 3, 2017
 Bayesian Statistics Debdeep Pati Florida State University April 3, 2017 Finite mixture model The finite mixture of normals can be equivalently expressed as y i N(µ Si ; τ 1 S i ), S i k π h δ h h=1 δ h
Bayesian Statistics Debdeep Pati Florida State University April 3, 2017 Finite mixture model The finite mixture of normals can be equivalently expressed as y i N(µ Si ; τ 1 S i ), S i k π h δ h h=1 δ h
Semiparametric Bayesian Inference for. Multilevel Repeated Measurement Data
 Semiparametric Bayesian Inference for Multilevel Repeated Measurement Data Peter Müller, Fernando A. Quintana and Gary L. Rosner Department of Biostatistics & Applied Mathematics, The University of Texas,
Semiparametric Bayesian Inference for Multilevel Repeated Measurement Data Peter Müller, Fernando A. Quintana and Gary L. Rosner Department of Biostatistics & Applied Mathematics, The University of Texas,
State-Space Methods for Inferring Spike Trains from Calcium Imaging
 State-Space Methods for Inferring Spike Trains from Calcium Imaging Joshua Vogelstein Johns Hopkins April 23, 2009 Joshua Vogelstein (Johns Hopkins) State-Space Calcium Imaging April 23, 2009 1 / 78 Outline
State-Space Methods for Inferring Spike Trains from Calcium Imaging Joshua Vogelstein Johns Hopkins April 23, 2009 Joshua Vogelstein (Johns Hopkins) State-Space Calcium Imaging April 23, 2009 1 / 78 Outline
Variational Bayesian Dirichlet-Multinomial Allocation for Exponential Family Mixtures
 17th Europ. Conf. on Machine Learning, Berlin, Germany, 2006. Variational Bayesian Dirichlet-Multinomial Allocation for Exponential Family Mixtures Shipeng Yu 1,2, Kai Yu 2, Volker Tresp 2, and Hans-Peter
17th Europ. Conf. on Machine Learning, Berlin, Germany, 2006. Variational Bayesian Dirichlet-Multinomial Allocation for Exponential Family Mixtures Shipeng Yu 1,2, Kai Yu 2, Volker Tresp 2, and Hans-Peter
Expression Data Exploration: Association, Patterns, Factors & Regression Modelling
 Expression Data Exploration: Association, Patterns, Factors & Regression Modelling Exploring gene expression data Scale factors, median chip correlation on gene subsets for crude data quality investigation
Expression Data Exploration: Association, Patterns, Factors & Regression Modelling Exploring gene expression data Scale factors, median chip correlation on gene subsets for crude data quality investigation
NONPARAMETRIC HIERARCHICAL BAYES VIA SEQUENTIAL IMPUTATIONS 1. By Jun S. Liu Stanford University
 The Annals of Statistics 996, Vol. 24, No. 3, 9 930 NONPARAMETRIC HIERARCHICAL BAYES VIA SEQUENTIAL IMPUTATIONS By Jun S. Liu Stanford University We consider the empirical Bayes estimation of a distribution
The Annals of Statistics 996, Vol. 24, No. 3, 9 930 NONPARAMETRIC HIERARCHICAL BAYES VIA SEQUENTIAL IMPUTATIONS By Jun S. Liu Stanford University We consider the empirical Bayes estimation of a distribution
Abstract INTRODUCTION
 Nonparametric empirical Bayes for the Dirichlet process mixture model Jon D. McAuliffe David M. Blei Michael I. Jordan October 4, 2004 Abstract The Dirichlet process prior allows flexible nonparametric
Nonparametric empirical Bayes for the Dirichlet process mixture model Jon D. McAuliffe David M. Blei Michael I. Jordan October 4, 2004 Abstract The Dirichlet process prior allows flexible nonparametric
Hyperparameter estimation in Dirichlet process mixture models
 Hyperparameter estimation in Dirichlet process mixture models By MIKE WEST Institute of Statistics and Decision Sciences Duke University, Durham NC 27706, USA. SUMMARY In Bayesian density estimation and
Hyperparameter estimation in Dirichlet process mixture models By MIKE WEST Institute of Statistics and Decision Sciences Duke University, Durham NC 27706, USA. SUMMARY In Bayesian density estimation and
Spiked Dirichlet Process Prior for Bayesian Multiple Hypothesis Testing in Random Effects Models
 Bayesian Analysis (2009) 4, Number 4, pp. 707 732 Spiked Dirichlet Process Prior for Bayesian Multiple Hypothesis Testing in Random Effects Models Sinae Kim, David B. Dahl and Marina Vannucci Abstract.
Bayesian Analysis (2009) 4, Number 4, pp. 707 732 Spiked Dirichlet Process Prior for Bayesian Multiple Hypothesis Testing in Random Effects Models Sinae Kim, David B. Dahl and Marina Vannucci Abstract.
Tutorial on Approximate Bayesian Computation
 Tutorial on Approximate Bayesian Computation Michael Gutmann https://sites.google.com/site/michaelgutmann University of Helsinki Aalto University Helsinki Institute for Information Technology 16 May 2016
Tutorial on Approximate Bayesian Computation Michael Gutmann https://sites.google.com/site/michaelgutmann University of Helsinki Aalto University Helsinki Institute for Information Technology 16 May 2016
Statistics: Learning models from data
 DS-GA 1002 Lecture notes 5 October 19, 2015 Statistics: Learning models from data Learning models from data that are assumed to be generated probabilistically from a certain unknown distribution is a crucial
DS-GA 1002 Lecture notes 5 October 19, 2015 Statistics: Learning models from data Learning models from data that are assumed to be generated probabilistically from a certain unknown distribution is a crucial
Bayesian Statistical Methods. Jeff Gill. Department of Political Science, University of Florida
 Bayesian Statistical Methods Jeff Gill Department of Political Science, University of Florida 234 Anderson Hall, PO Box 117325, Gainesville, FL 32611-7325 Voice: 352-392-0262x272, Fax: 352-392-8127, Email:
Bayesian Statistical Methods Jeff Gill Department of Political Science, University of Florida 234 Anderson Hall, PO Box 117325, Gainesville, FL 32611-7325 Voice: 352-392-0262x272, Fax: 352-392-8127, Email:
Bayesian semiparametric inference for the accelerated failure time model using hierarchical mixture modeling with N-IG priors
 Bayesian semiparametric inference for the accelerated failure time model using hierarchical mixture modeling with N-IG priors Raffaele Argiento 1, Alessandra Guglielmi 2, Antonio Pievatolo 1, Fabrizio
Bayesian semiparametric inference for the accelerated failure time model using hierarchical mixture modeling with N-IG priors Raffaele Argiento 1, Alessandra Guglielmi 2, Antonio Pievatolo 1, Fabrizio
Lecture 16-17: Bayesian Nonparametrics I. STAT 6474 Instructor: Hongxiao Zhu
 Lecture 16-17: Bayesian Nonparametrics I STAT 6474 Instructor: Hongxiao Zhu Plan for today Why Bayesian Nonparametrics? Dirichlet Distribution and Dirichlet Processes. 2 Parameter and Patterns Reference:
Lecture 16-17: Bayesian Nonparametrics I STAT 6474 Instructor: Hongxiao Zhu Plan for today Why Bayesian Nonparametrics? Dirichlet Distribution and Dirichlet Processes. 2 Parameter and Patterns Reference:
Part 7: Hierarchical Modeling
 Part 7: Hierarchical Modeling!1 Nested data It is common for data to be nested: i.e., observations on subjects are organized by a hierarchy Such data are often called hierarchical or multilevel For example,
Part 7: Hierarchical Modeling!1 Nested data It is common for data to be nested: i.e., observations on subjects are organized by a hierarchy Such data are often called hierarchical or multilevel For example,
David B. Dahl. Department of Statistics, and Department of Biostatistics & Medical Informatics University of Wisconsin Madison
 AN IMPROVED MERGE-SPLIT SAMPLER FOR CONJUGATE DIRICHLET PROCESS MIXTURE MODELS David B. Dahl dbdahl@stat.wisc.edu Department of Statistics, and Department of Biostatistics & Medical Informatics University
AN IMPROVED MERGE-SPLIT SAMPLER FOR CONJUGATE DIRICHLET PROCESS MIXTURE MODELS David B. Dahl dbdahl@stat.wisc.edu Department of Statistics, and Department of Biostatistics & Medical Informatics University
Recent Advances in Bayesian Inference Techniques
 Recent Advances in Bayesian Inference Techniques Christopher M. Bishop Microsoft Research, Cambridge, U.K. research.microsoft.com/~cmbishop SIAM Conference on Data Mining, April 2004 Abstract Bayesian
Recent Advances in Bayesian Inference Techniques Christopher M. Bishop Microsoft Research, Cambridge, U.K. research.microsoft.com/~cmbishop SIAM Conference on Data Mining, April 2004 Abstract Bayesian
A Bayesian Discovery Procedure
 A Bayesian Discovery Procedure Michele Guindani University of New Mexico, Alberquerque, NM 87111, U.S.A. Peter Müller University of Texas M.D. Anderson Cancer Center, Houston, TX 77030, U.S.A. Song Zhang
A Bayesian Discovery Procedure Michele Guindani University of New Mexico, Alberquerque, NM 87111, U.S.A. Peter Müller University of Texas M.D. Anderson Cancer Center, Houston, TX 77030, U.S.A. Song Zhang
Fast Likelihood-Free Inference via Bayesian Optimization
 Fast Likelihood-Free Inference via Bayesian Optimization Michael Gutmann https://sites.google.com/site/michaelgutmann University of Helsinki Aalto University Helsinki Institute for Information Technology
Fast Likelihood-Free Inference via Bayesian Optimization Michael Gutmann https://sites.google.com/site/michaelgutmann University of Helsinki Aalto University Helsinki Institute for Information Technology
A Nonparametric Model for Stationary Time Series
 A Nonparametric Model for Stationary Time Series Isadora Antoniano-Villalobos Bocconi University, Milan, Italy. isadora.antoniano@unibocconi.it Stephen G. Walker University of Texas at Austin, USA. s.g.walker@math.utexas.edu
A Nonparametric Model for Stationary Time Series Isadora Antoniano-Villalobos Bocconi University, Milan, Italy. isadora.antoniano@unibocconi.it Stephen G. Walker University of Texas at Austin, USA. s.g.walker@math.utexas.edu
Probabilistic Inference for Multiple Testing
 This is the title page! This is the title page! Probabilistic Inference for Multiple Testing Chuanhai Liu and Jun Xie Department of Statistics, Purdue University, West Lafayette, IN 47907. E-mail: chuanhai,
This is the title page! This is the title page! Probabilistic Inference for Multiple Testing Chuanhai Liu and Jun Xie Department of Statistics, Purdue University, West Lafayette, IN 47907. E-mail: chuanhai,
Hmms with variable dimension structures and extensions
 Hmm days/enst/january 21, 2002 1 Hmms with variable dimension structures and extensions Christian P. Robert Université Paris Dauphine www.ceremade.dauphine.fr/ xian Hmm days/enst/january 21, 2002 2 1 Estimating
Hmm days/enst/january 21, 2002 1 Hmms with variable dimension structures and extensions Christian P. Robert Université Paris Dauphine www.ceremade.dauphine.fr/ xian Hmm days/enst/january 21, 2002 2 1 Estimating
eqr094: Hierarchical MCMC for Bayesian System Reliability
 eqr094: Hierarchical MCMC for Bayesian System Reliability Alyson G. Wilson Statistical Sciences Group, Los Alamos National Laboratory P.O. Box 1663, MS F600 Los Alamos, NM 87545 USA Phone: 505-667-9167
eqr094: Hierarchical MCMC for Bayesian System Reliability Alyson G. Wilson Statistical Sciences Group, Los Alamos National Laboratory P.O. Box 1663, MS F600 Los Alamos, NM 87545 USA Phone: 505-667-9167
NONINFORMATIVE NONPARAMETRIC BAYESIAN ESTIMATION OF QUANTILES
 NONINFORMATIVE NONPARAMETRIC BAYESIAN ESTIMATION OF QUANTILES Glen Meeden School of Statistics University of Minnesota Minneapolis, MN 55455 Appeared in Statistics & Probability Letters Volume 16 (1993)
NONINFORMATIVE NONPARAMETRIC BAYESIAN ESTIMATION OF QUANTILES Glen Meeden School of Statistics University of Minnesota Minneapolis, MN 55455 Appeared in Statistics & Probability Letters Volume 16 (1993)
Bayesian Nonparametric Models
 Bayesian Nonparametric Models David M. Blei Columbia University December 15, 2015 Introduction We have been looking at models that posit latent structure in high dimensional data. We use the posterior
Bayesian Nonparametric Models David M. Blei Columbia University December 15, 2015 Introduction We have been looking at models that posit latent structure in high dimensional data. We use the posterior
Proteomics and Variable Selection
 Proteomics and Variable Selection p. 1/55 Proteomics and Variable Selection Alex Lewin With thanks to Paul Kirk for some graphs Department of Epidemiology and Biostatistics, School of Public Health, Imperial
Proteomics and Variable Selection p. 1/55 Proteomics and Variable Selection Alex Lewin With thanks to Paul Kirk for some graphs Department of Epidemiology and Biostatistics, School of Public Health, Imperial
